Abstract
Mrc1 (mediator of replication checkpoint), Tof1 (topoisomerase I interacting factor), and Csm3 (chromosome segregation in meiosis) are checkpoint-mediator proteins that function during DNA replication and activate the effector kinase Rad53. We reported previously that Mrc1 and Tof1 are constituents of the replication machinery and that both proteins are required for the proper arrest and stabilization of replication forks in the presence of hydroxyurea. In our current study, we show that Csm3 is a component of moving replication forks and that both Tof1 and Csm3 are specifically required for the association of Mrc1 with these structures. In contrast, the deletion of mrc1 did not affect the association of Tof1 and Csm3 with the replication fork complex. In agreement with previous observations in yeast cells, the results of a baculovirus coexpression system showed that these three proteins interact directly with each other to form a mediator complex in the absence of replication forks.
Introduction
During the progression of DNA replication, replication forks are exposed to various types of replicative stress. Many endogenous or external events have been identified to cause such stress, including template DNA damage, a reduced deoxynucleotide triphosphate (dNTP) pool as a result of hydroxyurea (HU)4 inhibition of dNTP biosynthesis, or collisions between the replication machinery and various DNA-binding proteins (1–4). The DNA replication checkpoint, a surveillance mechanism for S-phase progression, is crucial for the proper management of replicative stress stimuli and consequently for the maintenance of genome integrity.
The replication checkpoint signal transduction cascade is thought to consist of four basic steps. First, replication forks arrest in response to replicative stresses via the activity of checkpoint-mediator proteins, which are components of the replication machinery. Second, sensor kinases are recruited to arresting replication forks to phosphorylate mediator proteins that then recruit effector kinases. Third, these effector kinases are phosphorylated by sensor kinases. Finally, specific target proteins are phosphorylated, which eventually results in the stabilization of stalling replication forks, a reduction in the rate of DNA replication, and the activation of repair pathways (5, 6).
Studies in budding yeast (Saccharomyces cerevisiae) and fission yeast (Schizosaccharomyces pombe) have uncovered a protein network that constitutes a signal transduction cascade of the DNA replication checkpoint. The sensor and effector kinases that function in the replication checkpoint are well characterized and are conserved among eukaryotes. The DNA replication checkpoint sensor kinase is Mec1 (mitotic entry checkpoint) in budding yeast and Rad3 (radiation sensitive) in fission yeast, both of which are homologs of the human ATR kinase (ataxia telangiectasia-mutated) (4, 6, 7). In budding yeast, the effector kinase is Rad53, which is homologous to Cds1 (checking DNA synthesis) in fission yeast and Chk2 (checkpoint) in humans (7).
Mediator proteins have two distinct functions in the DNA replication checkpoint signal cascade. One is to monitor progression of replication forks and arrest forks in response to replicative stress, and the other is to serve as a molecular anchor for the subsequent recruitment of checkpoint sensor and effector kinases (1–4). To date, the Csm3, Tof1, and Mrc1 proteins have been identified as replication checkpoint-specific mediators in budding yeast (8–10). These three proteins were identified as components of the replisome progression complex, a large protein complex that was first identified biochemically and that includes GINS (go ichi ni san), the MCMs (minichromosome maintenance), Cdc45 (cell division cycle), and other proteins (11). Under conditions of replicative stress, activation of the effector kinase Rad53 is delayed in cells lacking any one of these mediator proteins (8, 12). Mrc1 is a substrate for the sensor kinase Mec1 and for the effector kinase Rad53 and directly interacts with Rad53 (13). Thus, Mrc1 is thought to play a central role in the DNA replication checkpoint cascade. Recently, Tof1 and Csm3, but not Mrc1, were shown to be required for proper arrest at diverse chromosomal sites where non-nucleosomal proteins bind very tightly to DNA (14). This suggests that the role of each protein during fork arrest may be different. Mrc1 may be required for maintaining the normal rate of replication fork progression as this is greatly reduced in the absence of this protein (15–18).
Although genetic and biochemical studies suggest that Tof1, Csm3, and Mrc1 have overlapping roles in activating the DNA replication checkpoint (8–10, 12, 16, 19), their relationships at the replication forks remain unclear. In our current study, we report that both Tof1 and Csm3 are specifically required for the association of Mrc1 at replication forks. Using a baculovirus expression system, we further show that recombinant Tof1, Csm3, and Mrc1 proteins form a complex in insect cells in the absence of replication forks, which is consistent with the findings of previous in vivo studies of replication checkpoint mediators (11, 15, 19, 20).
EXPERIMENTAL PROCEDURES
Yeast Strains
All yeast strains used in this study are listed in Table 1 and are derivatives of BY4741all (MATa, his3Δ1 leu2 Δ0 met15 Δ0 ura3 Δ0 trp1 Δ). In all strains, a HA3 tag or a FLAG3 tag coding sequence was fused to the sequence encoding the C terminus of the protein of interest; to achieve this, a cassette amplified from pU6H3HA or pU6H3FLAG, respectively, was used (21). All strains that expressed a tagged protein were examined for unaltered doubling times and sensitivity to HU and methyl methanesulfonate to verify the integrity of the cell phenotype. Gene deletions were performed by replacing the open reading frame of interest, including the start and stop codons, with a selective marker amplified by PCR from YIplac128, YIplac204, or YIplac211 as described previously (22).
TABLE 1.
Strains used in this study
| Strains | Genotype | Source |
|---|---|---|
| BY4741 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 | This study |
| SKY001 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::hisG | Ref. 20 |
| SKY006 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::hisG | Ref. 20 |
| TOF1-6His-3FLAG-loxP-KanMX-loxP | ||
| SKY401 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG Csm3-6His-3FLAG-loxP-KanMX-loxP | This study |
| SKY020 | MATahis3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::KanMX | Ref. 20 |
| CDC45-6HA::TRP1 AUR1::GPD-TK7X- | ||
| SKY402 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: | This study |
| KanMX csm3Δ:: LEU2 Cdc45-6HA::TRP1 AUR1::GPD-TK7X- | ||
| SKY403 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::hisG | This study |
| Csm3-3HA::TRP1 | ||
| SKY404 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::hisG | This study |
| Tof1-3HA::TRP1 | ||
| SKY405 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::hisG | This study |
| Csm3-3HA::TRP1 tof1Δ:: LEU2 | ||
| SKY406 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ::hisG | This study |
| Tof1-3HA::TRP1 csm3Δ:: LEU2 | ||
| SKY407 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Tof1-3HA::TRP1 | ||
| SKY408 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Csm3-3HA::TRP1 | ||
| SKY409 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Csm3-3HA::TRP1 tof1Δ:: LEU2 | ||
| SKY410 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Tof1-3HA::TRP1 csm3Δ:: LEU2 | ||
| SKY411 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Tof1-6His-3FLAG-loxP-KanMX-loxP Csm3-3HA::TRP1 | ||
| SKY412 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Tof1-6His-3FLAG-loxP-KanMX-loxP Csm3-3HA::TRP1 mrc1Δ:: LEU2 | ||
| SKY016 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | Ref. 20 |
| Tof1-6His-3FLAG-loxP-KanMX-loxP Cdc45-3HA::TRP1 | ||
| SKY017 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | Ref. 20 |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Cdc45-3HA::TRP1 | ||
| SKY413 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Tof1-6His-3FLAG-loxP-KanMX-loxP Cdc45-3HA::TRP1 mrc1Δ:: LEU2 | ||
| SKY414 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Cdc45-3HA::TRP1 tof1Δ:: LEU2 | ||
| SKY415 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Cdc45-3HA::TRP1 csm3Δ:: LEU2 | ||
| SKY416 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3D0 trp1Δ:: hisG | This study |
| Mrc1-6His-3FLAG-loxP-KanMX-loxP Cdc45-3HA::TRP1 tof1Δ:: LEU2 csm3Δ:: URA3 |
Antibodies
The HA epitope was detected using the mouse monoclonal antibody 12CA5 (Roche Diagnostics) or HA.11 (Covance, Princeton, NJ), and the FLAG epitope was detected using the mouse monoclonal antibody M2 (Sigma). Endogenous Mcm2 protein was detected using the goat polyclonal antibody sc-6680 (Santa Cruz Biotechnology, Santa Cruz, CA).
ChIP-Chip Analysis
Chromatin immunoprecipitations on a DNA chip (ChIP-chip) were performed as described previously (20, 23). To examine all of chromosome VI, the RikDacF DNA chip (Affymetrix, Santa Clara, CA) was used. Cells (1.5 × 108) were disrupted with a Multi-bead shocker (MB400U; Yasui Kikai, Osaka, Japan). The DNA to be used in the ChIP analysis was purified and then amplified by random-priming PCR. Hybridization, washing, and array scanning were performed using the Affymetrix GeneChip system according to the manufacturer's instructions. To discriminate between positive and negative probe binding signals, three criteria were adopted. First, the reliability of the signal strength was evaluated using the detection p value for each locus (p value ≤0.025). Second, the reliability of the binding ratio was evaluated by determining the change in the p value (for p values ≤0.025). Third, clusters consisting of at least three contiguous loci that satisfied the above two criteria in two independent experiments were selected. The latter criterion was included because a single site of protein-DNA interaction will result in immunoprecipitation of DNA fragments that hybridize not only to the locus of the actual binding site, but also to neighboring loci (20, 23). BrdUrd incorporation analyses were performed as previously described (23).
Quantitative PCR
PCR amplification was performed using the Applied Biosystems 7500 real time PCR system (ABI, Foster City, CA) according to the manufacturer's instructions. The primer pair 185-kb 5′-CCTCCAAATCCACTGTTACTGCTATCC-3′ and 5′-CCATTGTTACTGGTCTTGTCCTTCTTGC-3′ was used to amplify the region of chromosome VI encompassing positions 184962–185178, and the primer pair 196-kb 5′-TGTAATACTCCTCAAAGGTCCTCC-3′ and 5′-GAAGACAACGACGAAGAACTAGC-3′ was used for the region 196898–197077.
Yeast Cultures
G1-arrested cells were prepared by treatment with α-factor (2 μm) for 2.5 h at 23 °C. To prepare HU-treated cells, G1-arrested cells were released into medium containing 200 mm HU for the indicated times at 23 °C as previously described (20). To prepare cells in S phase without HU treatment, cells were released from G1 arrest into S phase by incubation for 40 or 60 min at 16 °C. To prepare G2-arrested cells, asynchronously growing cells were treated with 15 μg/ml of nocodazole for 2.5 h at 23 °C.
To analyze the chromosomal localization of Csm3 and Tof1 during unperturbed S phase, cells expressing a tagged version of each protein were arrested in G1 using α-factor and then released into S phase by incubation at 16 °C in the presence of 100 μg/ml of Pronase to slow the rate of DNA synthesis as described previously (24).
Co-immunoprecipitation Analyses
Synchronized cells in various phases of growth were harvested and washed twice in 0.1 mm phenylmethanesulfonyl fluoride as described previously (20). The cells were then suspended in lysis buffer (50 mm HEPES-KOH, pH 7.5, 150 mm NaCl, 3 mm MgCl2, 10% (v/v) glycerol, 0.2% Nonidet P-40) containing protease inhibitors, 1× Complete (Roche Diagnostics) and 1% protease inhibitor mixture (Sigma), and phosphatase inhibitors, 10 mm sodium fluoride and 20 mm glycerol 2-phosphate disodium salt hydrate. The cells were then disrupted using a Multi-bead shocker and the resulting lysate was treated with 200 units of DNase I (Takara Bio Inc., Shiga, Japan) at 4 °C for 15 min and then centrifuged at 20,000 × g to prepare a clear whole cell extract. The lysate was then pre-cleared by incubation with Dynabeads Protein A (Invitrogen) for 30 min. After pre-clearing, the lysate was incubated for 3 h with either anti-FLAG (M2; Sigma) or anti-HA (12CA5; Roche Diagnostics) conjugated with Dynabeads Protein A. After immunoprecipitation, samples were washed four times with lysis buffer and resuspended in SDS loading buffer (62.5 mm Tris-HCl, pH 6.8, 2% sodium lauryl sulfate, 10% (v/v) glycerol, 5% (v/v) 2-mercaptethanol).
Expression of Yeast Proteins in Insect Cells
DNA sequences encoding Mrc1 and Tof1 were cloned into the Bac-to-Bac pFastBac H/T baculovirus expression vector (Invitrogen). Sequences encoding FLAG or HA epitope tag were introduced at the N terminus of the Tof1 coding sequence. Mrc1 was tagged by the FLAG tag at its N terminus. Bacmid was prepared following the manufacturer's instructions and transfected into Sf9 insect cells (Invitrogen). The sequence encoding Csm3 with an N-terminal HA tag was cloned into the pAcUW31 vector (BD Biosciences). For expression of Csm3, BaculoGold Autographa californica nuclear polyhedrosis virus DNA (BaculoGold AcNPV; BD Biosciences) and the pAcUW31 vector containing the HA-tagged csm3 gene were co-transfected into Sf9 cells. Expression of the recombinant proteins was verified by immunoblotting. For protein expression, Sf9 cells were grown at 27 °C in Grace's insect medium (Invitrogen) supplemented with 10% fetal bovine serum and infected with the indicated viruses. Cells were harvested 2 days after infection, and extracts were prepared using lysis buffer (20 mm Tris-HCl, pH 7.5, 150 mm NaCl, 10% (v/v) glycerol, 1× Complete protease inhibitor (Roche Diagnostics) containing 0.5% Triton X-100). Extracts were cleared by centrifugation at 20,000 × g for 15 min, and the supernatant was immunoprecipitated in the presence of M2-agarose (Sigma) for 2 h. The immunoprecipitated beads were then washed four times in lysis buffer containing 300 mm NaCl and 0.1% Nonidet P-40. The elution of precipitated proteins was performed using 0.5 mg/ml FLAG-peptide (Sigma). The eluate was then immunoprecipitated again in the presence of anti-HA-agarose beads for 1 h and eluted with SDS loading buffer without boiling. The samples were separated by SDS-PAGE and stained using the Coomassie Brilliant Blue staining kit (Wako Pure Chemical Industries, Osaka, Japan).
RESULTS
Csm3 Associates with Replication Forks
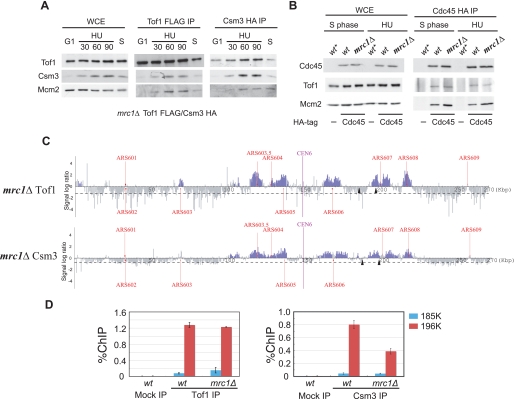
Previous biochemical studies have revealed that Tof1, Mrc1, and Csm3 are components of the replisome progression complex (11, 25). To confirm the S phase-specific association of Tof1 and Csm3 with the DNA replication forks, we performed co-immunoprecipitation experiments using cells that co-expressed Tof1-FLAG3 and Csm3-HA3 (Fig. 1, A and B). The data clearly demonstrated that these proteins physically interact with Mcm2 in vivo and do so specifically during S phase with or without HU, an inhibitor of ribonucleotide reductase (Fig. 1, A and B). To confirm the localization of Csm3 on chromosome VI during DNA replication in S phase, we performed ChIP-chip analysis (chromatin immunoprecipitation combined with DNA chip analysis) of Csm3 during S phase in the presence and absence of HU (Fig. 1, C and D). Cells were arrested in G1 using α-factor and then released into S phase in the presence or absence of 200 mm HU. Chromatin immunoprecipitation was then performed using cells expressing Csm3-FLAG3 or Tof1-HA3. The localization of Csm3 correlated well with that of Tof1, which has been reported to form a complex with Csm3 and to localize to replication forks (20, 25). In the presence of HU, both proteins were localized only at the early firing origins (Fig. 1C). In cells in S phase without HU treatment, Tof1 and Csm3 were initially localized at early firing origins and then moved away from the replication origins at a similar rate with time (Fig. 1D). These data suggest that Csm3 is a component of moving replication forks, similar to the other known mediator proteins, Tof1 and Mrc1. Interestingly, we observed similar levels of enrichment of Tof1 and Csm3 in HU-treated cells at replication forks, but when we performed ChIP-chip analysis of Csm3 during early S phase, we reproducibly observed more enrichment of Csm3 than Tof1 at the forks (Fig. 1D). This may suggest that the binding of Csm3 and Tof1 to replication forks differs between cells in normal S phase and those in S phase under HU-induced replicative stress.
FIGURE 1.
Csm3 is localized at DNA replication forks. A, Tof1 specifically interacts with Mcm2 and Csm3 during S phase. Immunoprecipitations of extracts from wild-type cells that co-expressed Tof1-FLAG3 and Csm3-HA3 were performed using anti-FLAG (FLAG IP) antibodies. Cells were grown exponentially (Asyn.), synchronized in G2 using nocodazole (Noc.), synchronized in G1 using α-factor (G1), and then released from G1 into S phase in the absence (S) or presence (HU) of 200 mm HU for 60 min. Tof1, Csm3, and Mcm2 were then detected by Western blotting analysis of immunoprecipitates (IP) and of a whole cell extract (WCE) control. B, samples were prepared as in A, except in the case of immunoprecipitations using anti-HA antibodies (HA IP). C, distribution of Tof1 and Csm3 on chromosome VI in the wild-type strain in the presence of 200 mm HU. Cells were arrested in G1 by α-factor and then released into S phase at 23 °C in the presence of 200 mm HU for 60 min. Chromatin immunoprecipitation combined with DNA chip hybridization (ChIP-chip) was carried out as described previously (23). Each blue bar represents the binding ratio of Tof1 (upper panel) or Csm3 (lower panel) at each locus and indicates a significant enrichment in some of the IP fractions. The dashed line represents the average signal ratio at the loci that were not enriched (gray bars) in each IP fraction. The scale of the vertical axis is log2. Positions of the centromere (CEN6) and all previously mapped autonomously replicating sequences (ARSs) are shown. The horizontal axis represents the length of chromosome VI in kilobases. Tof1 and Csm3 were tagged with FLAG3 and HA3, respectively. Tof1 and Csm3 were found to localize only in early firing origins of replication (ARS603.5, ARS605, ARS606, and ARS607) under our experimental conditions. D, distribution of Tof1 and Csm3 on chromosome VI in wild-type cells during normal S-phase progression. Cells expressing Tof1-FLAG3 and Csm3-HA3 were arrested in G1 using α-factor and then released into S phase for 40 or 60 min at 16 °C in the presence of Pronase to suppress the rate of DNA synthesis.
Csm3 Is Required for the Stable Arrest of Replication Forks under HU-induced Replicative Stress
We previously demonstrated that Mrc1 and Tof1 are components of the DNA replication forks and are required for the stable pausing of these forks under replicative stress induced by HU. In the absence of Mrc1 or Tof1, and in the presence of HU, replication forks do not arrest properly, and replication proteins become uncoupled from the replicated regions of the DNA (20). To address whether Csm3 functions in a similar manner, we examined the progression of DNA replication in the csm3 deletion mutant strain csm3Δ in the presence of HU. Using the ChIP-chip method, we analyzed the chromosome-wide localization of Cdc45, a component of replication forks, and measured BrdUrd incorporation to assess DNA synthesis (20). The cells were arrested by α-factor in G1 and then released into S phase in the presence of 200 mm HU. Chromatin immunoprecipitation using anti-HA (to detect Cdc45-HA3) and anti-BrdUrd was then performed. In csm3Δ cells, the distribution of the BrdUrd-incorporated regions in chromosomes was essentially equivalent (or even slightly more restricted) to that in wild-type cells, and Cdc45 progressed 1–2 kb further than in wild-type cells in the presence of HU (Fig. 2A). These results suggest that a partial uncoupling of the replication machinery took place in the replicated regions, similar to that observed in our previous report (20) using tof1Δ or mrc1Δ mutant strains in the presence of HU. The extent of this uncoupling in the csm3Δ strain was, however, less pronounced than that observed in the tof1 or mrc1 deletion mutants. In the csm3Δ mutant also, Cdc45 was detected at the late firing origin ARS608. Because the activity of the late firing origin is suppressed in response to HU-induced DNA replication checkpoint activation (26), these data suggest that csm3Δ is partially defective for DNA replication checkpoint activation in response to HU. To next examine the role of Csm3 during normal progression of S phase, we analyzed the distribution of Cdc45 and BrdUrd in the absence of HU during early DNA replication (Fig. 2B). Cells were arrested in G1 using α-factor and then released for 60 min at 16 °C to allow slow progression of S phase (20). In both wild-type and csm3Δ cells, BrdUrd incorporation and Cdc45 localization were detected around the early replicating regions and there were no differences observed in the localization patterns of Cdc45 and BrdUrd between the wild-type and csm3Δ strains (Fig. 2B). These results indicate that, similar to Mrc1 or Tof1, Csm3 is required to couple the replication machinery with the replicated DNA regions in the presence of HU.
FIGURE 2.
Distribution of Cdc45 and rate of BrdUrd incorporation on chromosome VI in wild type (wt) and csm3 deletion mutant yeast in S phase with or without HU. A, distribution of Cdc45 and BrdUrd incorporation on chromosome VI in wild-type and csm3Δ mutant cells in S phase in the presence of HU. Cells expressing a HA-tagged version of Cdc45 were arrested in G1 using α-factor, released into S phase at 23 °C in the presence of 200 mm HU for 60 min, and subjected to ChIP-chip analysis. Blue and red bars represent regions of Cdc45 binding and BrdUrd incorporation, respectively. B, distribution of Cdc45 and BrdUrd incorporation on chromosome VI in wild-type and csm3Δ cells during normal S-phase progression. Cells were arrested in G1 using α-factor, released into S phase for 60 min at 16 °C to slow DNA synthesis, and then processed for ChIP-chip analysis. Blue and red bars represent regions of Cdc45 binding and BrdUrd incorporation, respectively.
Co-dependent Association of Tof1 and Csm3 with Replication Forks
Csm3 forms a tight complex with Tof1 (27, 28). To thus examine whether these proteins are co-dependent for their association with replication forks, we analyzed the interaction of Csm3 or Tof1 with Mcm2, a member of the helicase complex, by co-immunoprecipitation analysis (Fig. 1A). A sequence encoding a HA3 epitope tag was fused to the chromosomal copy of tof1 or csm3, and each tagged protein was expressed in the absence of its counterpart to assess its interaction with endogenous Mcm2. The cells were synchronized in G1 and then released into S phase for 40 or 60 min in the presence of HU at 23 °C (Fig. 3A). We confirmed that the cells were in S phase using flow cytometric analysis (data not shown), and whole cell extracts were then prepared. Tof1 and Csm3 co-immunoprecipitated with Mcm2 in the wild-type strain. In the absence of each other, however, Tof1 and Csm3 did not interact with Mcm2 during S phase in the presence or absence of HU-induced replicative stress (Fig. 3A). In addition, the csm3 protein levels in tof1Δ whole cell extracts were reduced compared with wild-type (Fig. 3A), suggesting that the interaction between Tof1 and Csm3 is important for Csm3 stabilization.
FIGURE 3.
Co-dependent association of Tof1 and Csm3 with DNA replication forks. A, interactions among Tof1, Csm3, and Mcm2 were determined by immunoprecipitation using anti-HA antibodies. Extracts of wild-type (wt), tof1Δ, and csm3Δ cells that expressed either Tof1-HA3 or Csm3-HA3 were used in the analysis as indicated. Cells were synchronized in G1 using α-factor and then released into S phase in the absence (S phase) or presence (HU) of 200 mm HU for 60 min. Western blot analyses of Tof1, Csm3, and Mcm2 in immunoprecipitates and in whole cell extract (WCE) controls were performed using anti-HA (for Tof1 and Csm3) and anti-Mcm2 antibodies. B, distribution of Tof1 and Csm3 on chromosome VI in the csm3Δ and tof1Δ mutants under HU-induced replicative stress. Cells were synchronized in G1 using α-factor, released into S phase in the presence of 200 mm HU for 60 min, and processed for ChIP-chip analysis. For a comparison with wild type, see Fig. 1C. C, efficiency of chromatin immunoprecipitation (% ChIP; ratio of input versus IP signal) of Tofl-FLAG3 (left) or Csm3-FLAG3 (right) in wild-type, csm3Δ, and tof1Δ strains was measured by real time PCR analysis using primer pairs specific for amplification of the regions that each begin where indicated by the arrowheads in B; 185K and 196K correspond to the distances in kilobase pairs from the left end of the chromosome. Data represent the average of two independent experiments ± S.D.
We next performed ChIP-chip analysis to determine localization of Tof1 and Csm3 on chromosome VI in the absence of each other. The cells were synchronized in G1 and then released into S phase for 60 min in the presence of HU at 23 °C. In the mutant strains, Tof1 and Csm3 no longer localized at the replication forks, in contrast to the situation in wild-type cells (Fig. 3B, for comparison with wild-type, see Fig. 1C). To confirm this drastic reduction in the association of Tof1 and Csm3 with replication forks in the absence of each other, quantitative-PCR analyses were performed (Fig. 3C). A region of 196 kb from the left telomere of chromosome VI, which is next to the early firing origin ARS607 where enrichment of replication forks has been demonstrated previously (20, 29), was analyzed. As a control locus, a region of 185 kb was tested that is also from the left telomere of chromosome VI but where no replication fork is localized under the conditions studied (20, 29). The quantitative-PCR results indicated that the amount of Tof1 and Csm3 that associated with replication forks in the mutant strains was <1% of that observed in the wild-type yeast. From these data, we conclude that Tof1 interacts directly with Csm3 in vivo and that formation of a Tof1-Csm3 complex is essential for localization of these proteins at DNA replication forks.
The Association of Mrc1 with Replication Forks Is Partly Dependent on the Tof1-Csm3 Complex
Several lines of evidence have now suggested that Tof1 and Csm3 function in a pathway that is distinct from that of Mrc1 during the normal progression of S phase. Given the requirement of these three proteins for the activation of the DNA replication checkpoint, we examined whether the association of Mrc1 with the replication forks is dependent on Tof1 and Csm3. We examined the co-immunoprecipitation of Mrc1-FLAG3 with endogenous Mcm2 in the absence of Tof1 or Csm3 (Fig. 4A). Mrc1 was found to co-precipitate with Tof1, Csm3, and Mcm2 in the wild-type strain. In the absence of Tof1 or Csm3, however, Mrc1 did not co-precipitate with Mcm2. Because Tof1, Csm3, and Mrc1 are required for stabilization of replication forks, we speculated that the loss of association with Mcm2 might be due to a breakdown of fork structures in the tof1Δ and csm3Δ mutants. Hence, we examined the association of another constituent of the replication machinery, Cdc45, with Mcm2. The successful co-immunoprecipitation of Cdc45-HA3 with endogenous Mcm2 and Mcr1-FLAG3 indicates that Cdc45 interacts with Mcm2 in the absence of tof1, csm3, or both with the same efficiency as in wild-type cells (Fig. 4B). In contrast, Mrc1 co-immunoprecipitated with Cdc45 only in wild-type cells (Fig. 4B), indicating that it does not efficiently associate with the replicative helicase complex in the absence of Tof1 or Csm3, even though the fork structure appears to be intact. The localization of Mrc1 at the replication forks, as determined using ChIP-chip analysis (Fig. 4C), suggests that the levels of Mrc1 bound at the forks are reduced and partly uncoupled from the replicated DNA regions in the absence of tof1 or csm3, as we have also reported previously (20). Quantitative-PCR analyses further confirmed the reduced localization of Mrc1 at the replication forks (Fig. 4D). The localization of Mrc1 at or near chromosome site ARS607 was reduced to 30% or less of the levels observed in wild-type cells in the absence of either the tof1 or csm3 genes. However, deletion of the tof1 and csm3 genes did not have an additive effect, as had been expected from the observation that the association of Tof1 and Csm3 with Mcm2 is co-dependent (Fig. 3). These data together suggest that the efficient association of Mrc1 with replication forks is dependent on the formation of a Tof1-Csm3 complex.
FIGURE 4.
Both Tof1 and Csm3 are required for the association of Mrc1 with DNA replication forks. A, interaction between Mrc1 and Mcm2. Extracts from wild-type (wt), tof1Δ, and csm3Δ cells that expressed Mcr1-FLAG3, Tof1-HA3, or Csm3-HA3 were immunoprecipitated using anti-FLAG antibodies. Cells were synchronized in G1 and then released into S phase in the absence (S phase) or presence (HU) of 200 mm HU, followed by Western blotting using anti-FLAG (for Mrc1), anti-HA (for Tof1, Csm3), or anti-Mcm2 antibodies. Whole cell extracts (WCE) were used as a control. B, interactions among Cdc45, Mrc1, and Mcm2. Extracts from wild-type, tof1Δ, csm3Δ, and tof1Δcsm3Δ cells that expressed Cdc45-HA3 and Mcr1-FLAG3 were immunoprecipitated using anti-HA antibodies (HA IP). Cells were synchronized and analyzed as in A. Wild-type cells expressing Mcr1-FLAG3 alone were used as a control (indicated by an asterisk). C, distribution of Mrc1 on chromosome VI in wild-type (wt), tof1Δ, csm3Δ, and tof1Δcsm3Δ cells. Cells were synchronized in G1 using α-factor, released into S phase in the presence of 200 mm HU, and processed for ChIP-chip analysis. Mrc1 distribution was analyzed by ChIP-chip analysis using anti-FLAG antibodies. D, efficiency of chromatin immunoprecipitation (% ChIP; ratio of input versus IP) of Mrc1-FLAG3 in wild-type, tof1Δ, and csm3Δ cells was measured by real time PCR using primer pairs specific for amplification of the regions indicated by the arrowheads in C; 185K and 196K correspond to the distances in kilobase pairs from the left end of the chromosome. Data represent the average of two independent experiments ± S.D.
Tof1 Associates with DNA Replication Machinery in the Absence of Mrc1
To further understand the relationships between Tof1, Csm3, and Mrc1, we investigated whether Mrc1 is required for the correct localization of Tof1 and Csm3 at the replication forks in the presence or absence of the replicative stress induced by HU. We first examined the interactions among Tof1-FLAG3, Csm3-HA3, and endogenous Mcm2 and determined that the Tof1-Csm3 interaction is not affected by the deletion of mrc1 (Fig. 5A). We also examined the interactions among Tof1-FLAG3, Cdc45-HA3, and endogenous Mcm2, and found that the Tof1-Cdc45 interaction was not dependent on Mrc1 (Fig. 5B). Interactions among Tof1, Csm3, and Mcm2 were detected specifically in S phase with or without HU in the absence of Mrc1 (Fig. 5A). Similarly, interactions among Cdc45, Tof1, and Mcm2 were specifically detected in S phase with or without HU in the absence of Mrc1 (data not shown and see Fig. 5B).
FIGURE 5.
The association of Tof1 and Csm3 with DNA replication forks is unaffected by Mrc1. A, interaction of Tof1 and Csm3 with Mcm2. Extracts from Δmrc1 cells that co-expressed Tofl-FLAG3 and Csm3-HA3 were immunoprecipitated using anti-FLAG (FLAG IP) or anti-HA (HA IP) antibodies. Cells were arrested in G1 using α-factor (G1), and then released into S phase in the absence (S) or presence (HU) of 200 mm HU for the indicated times (30, 60, or 90 min). Western blotting of immunoprecipitates using anti-FLAG (Tof1), anti-HA (Csm3), and anti-Mcm2 antibodies was then performed. B, interactions among Cdc45, Mcm2, and Tof1 in mrc1Δ and wild-type cells that expressed Cdc45-HA3 and Tofl-FLAG3. Experiments were performed as in A. Wild-type cells that expressed Tofl-FLAG3 alone were used as a control (indicated by an asterisk). C, distribution of Tof1 and Csm3 on chromosome VI in mrc1Δ cells treated with HU. Cell extracts were prepared as described in the legend to Fig. 4B, and ChIP-chip analyses were performed. For a comparison with wild type, see Fig. 1C. D, efficiency of chromatin immunoprecipitation (% ChIP; ratio of input versus IP) of Tofl-FLAG3 and Csm3-FLAG3 in wild-type and mrc1Δ cells was measured by real time PCR using primer pairs specific for amplification of the regions indicated by the arrowheads in C; 185K and 196K correspond to the distances in kilobase pairs from the left end of the chromosome. Data represent the average of two independent experiments ± S.D.
We next performed ChIP-chip analyses to examine the interaction between Csm3 and Tof1 in the absence of mrc1 and in the presence of HU-induced replicative stress (Fig. 5C, for comparison with wild-type, see Fig. 1C). The data we obtained suggest that both Tof1 and Csm3 are uncoupled from replicated DNA regions in the absence of mrc1, as we had reported previously (20). Quantitative-PCR analyses further showed that the association of Csm3 with the selected sites on chromosome VI is reduced by the deletion of mrc1. Because the interaction of Csm3 with the replisome complex itself is unaffected by deletion of mrc1 (Fig. 5, A and B), this suggests that the association of Csm3 with the chromosome and binding to the replisome complex is arrested in a less synchronous manner in the absence of Mrc1, as revealed by ChIP-chip analysis (Fig. 5C).
Mrc1, Tof1, and Csm3 Form a Heterotrimeric Mediator Complex in Vitro
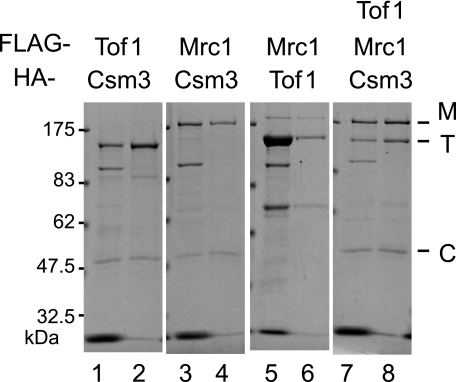
As described above, the sequential and hierarchical binding of Mrc1, Tof1, and Csm3 to DNA replication forks occurs in vivo. To evaluate this phenomenon in vitro, we adopted a biochemical strategy to examine the direct interactions between Mrc1, Tof1, and Csm3 in the mediator complex using recombinant proteins purified from baculovirus-infected insect cells. We prepared the recombinant proteins in three different combinations (Fig. 6). First, insect cells were co-infected with baculoviruses that expressed two of the three proteins, and the resultant complexes were partially purified by immunoprecipitation using anti-FLAG followed by immunoprecipitation using anti-HA. After immunoprecipitation with anti-FLAG, a few bands were detected in the eluates in addition to specific bands for each of Mrc1, Tof1, and Csm3. These extra bands were, however, mostly removed after subsequent immunoprecipitation of the anti-FLAG eluates with anti-HA. We detected the formation of complexes between Tof1 and Csm3, between Csm3 and Mrc1, and between Mrc1 and Tof1 (Fig. 6, lanes 2, 4, and 6). Tof1 was efficiently expressed when co-expressed with Mrc1 (Fig. 6, lane 5). When the cells co-expressed all three proteins, Tof1, Csm3, and Mrc1 were detected in the immunoprecipitation eluates, which suggests that Tof1, Csm3, and Mrc1 form a heterotrimeric complex in the insect cells in the absence of replication forks (Fig. 6, lane 8).
FIGURE 6.
Interactions among Tof1, Csm3, and Mrc1 in SF9 insect cells. Sf9 cells were co-infected with recombinant baculoviruses that expressed untagged, FLAG-tagged or HA-tagged Tof1 (T), HA-tagged Csm3 (C), and FLAG-tagged Mrc1 (M) in the combinations shown. Immunoprecipitation of cell lysates was performed using anti-FLAG M2-agarose beads, and elutions were performed using FLAG peptides (Lanes 1, 3, 5, and 7). The eluate was immunoprecipitated again using anti-HA-agarose beads and eluted with SDS loading buffer without boiling (lanes 2, 4, 6, and 8). Samples were separated by SDS-PAGE and stained with Coomassie Brilliant Blue solution.
DISCUSSION
Tof1, Csm3, and Mrc1 are essential for activation of the DNA replication checkpoint (8–10) and for the stabilization of DNA replication forks during pausing under replicative stress, such as that induced by HU treatment (20). Because it binds tightly to Tof1, Csm3 is thought to be a component of the replication machinery (26, 27). In our current study, we show that Csm3 is located at the replication forks and moves together with Tof1 and Mrc1 as these forks progress during S phase. We show also that the association of Tof1 and Csm3 with the replication forks is mutually dependent. Furthermore, the efficient association of Mrc1 with replication forks was found to be dependent on the presence of both Tof1 and Csm3. Partial defects in the replication checkpoint in deletion mutants tof1 and csm3 may be explained by the reduced Mrc1 association with replication forks (9, 10) as Mrc1 interacts directly with Rad53, an activator of the replication checkpoint (8). It should be noted also that association of Csm3 with replication forks and the total levels of cellular Csm3 protein were both dramatically reduced in the absence of Tof1. These results suggest that Csm3 exists in a tight complex with Tof1 in the cell and that free Csm3 is susceptible to rapid degradation, which is not the case for Tof1. The modes of association of Tof1, Csm3, and Mrc1 with the replication forks are somewhat intricate. The association of Mrc1 was found to be dependent on the Tof1-Csm3 complex but not vice versa. Recombinant protein expression in insect cells resulted in the formation of a heterotrimeric complex that contained Tof1, Csm3, and Mrc1. These results are in agreement with the observation that the efficient association of Mrc1 with replication forks is inhibited by the deletion of either tof1 or csm3. Tof1 and Csm3 could interact with Mrc1 in the absence of each other in vitro, but such interactions were not observed in vivo. The discrepancy between in vivo and in vitro results may imply the existence of inhibitory factors or protein modifications in relation to these complex formations in vivo. It would therefore be interesting to explore the role of the phosphorylation of Tof1 and Mrc1 in these processes, because both proteins are phosphorylated during S-phase with or without replicative stress (data not shown). It was recently reported also that Mrc1 directly binds DNA Pol ϵ and Mcm6 during S phase (30, 31). The weak association of Mrc1 with the replication forks that we observed in the absence of tof1 or csm3 may therefore be explained by a direct interaction of Mrc1 with these essential components of DNA replication forks.
Whether Mrc1, Tof1, and Csm3 have separate roles in controlling DNA replication has been the subject of much debate. Our current data suggest that the Tof1-Csm3 complex may directly bind to essential components of the replisome (i.e. helicase or DNA polymerases) or DNA itself and efficiently target Mrc1 at the replication forks. It will therefore be important for a fuller understanding of the molecular mechanisms underlying fork arrest and the activation of replication checkpoint to identify any interacting partners of the Tof1-Csm3 complex in future studies. Such factors will likely be essential components of the replisome and analyses of the biological significance of their interactions will be important.
Acknowledgments
We thank K. Kurihara and K. Nakagawa for technical assistance. We thank all members of the Shirahige Laboratory.
Footnotes
- HU
- hydroxyurea
- HA
- hemagglutinin
- ChIP
- chromatin immunoprecipitation
- BrdUrd
- bromodeoxyuridine.
REFERENCES
- 1.Branzei D., Foiani M. (2005) Curr. Opin. Cell Biol. 17, 568–575 [DOI] [PubMed] [Google Scholar]
- 2.Lambert S., Carr A. M. (2005) Biochimie 87, 591–602 [DOI] [PubMed] [Google Scholar]
- 3.Labib K., Hodgson B. (2007) EMBO Rep. 8, 346–853 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tourrière H., Pasero P. (2007) DNA Repair 6, 900–913 [DOI] [PubMed] [Google Scholar]
- 5.Gottifredi V., Prives C. (2005) Semin. Cell Dev. Biol. 16, 355–368 [DOI] [PubMed] [Google Scholar]
- 6.Branzei D., Foiani M. (2006) Exp. Cell Res. 312, 2654–2659 [DOI] [PubMed] [Google Scholar]
- 7.Boddy M. N., Russell P. (2001) Curr. Biol. 11, R953–R956 [DOI] [PubMed] [Google Scholar]
- 8.Alcasabas A. A., Osborn A. J., Bachant J., Hu F., Werler P. J., Bousset K., Furuya K., Diffley J. F., Carr A. M., Elledge S. J. (2001) Nat. Cell Biol. 3, 958–965 [DOI] [PubMed] [Google Scholar]
- 9.Foss E. J. (2001) Genetics 157, 567–577 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tong A. H., Lesage G., Bader G. D., Ding H, Xu H., Xin X., Young J., Berriz G. F., Brost R. L., Chang M., Chen Y., Cheng X., Chua G., Friesen H., Goldberg D. S., Haynes J., Humphries C., He G., Hussein S., Ke L., Krogan N., Li Z., Levinson J. N., Lu H., Ménard P., Munyana C., Parsons A. B., Ryan O., Tonikian R., Roberts T., Sdicu A. M., Shapiro J., Sheikh B., Suter B., Wong S. L., Zhang L. V., Zhu H., Burd C. G., Munro S., Sander C., Rine J., Greenblatt J., Peter M., Bretscher A., Bell G., Roth F. P., Brown G. W., Andrews B., Bussey H., Boone C. (2004) Science 303, 808–813 [DOI] [PubMed] [Google Scholar]
- 11.Gambus A., Jones R. C., Sanchez-Diaz A., Kanemaki M., van Deursen F., Edmondson R. D., Labib K. (2006) Nat. Cell Biol. 8, 358–366 [DOI] [PubMed] [Google Scholar]
- 12.Tanaka K., Russell P. (2001) Nat. Cell Biol. 3, 966–972 [DOI] [PubMed] [Google Scholar]
- 13.Osborn A. J., Elledge S. J. (2003) Genes Dev. 17, 1755–1767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mohanty B. K., Bairwa N. K., Bastia D. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 897–902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Calzada A., Hodgson B., Kanemaki M., Bueno A., Labib K. (2005) Genes Dev. 19, 1905–1919 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tourrière H., Versini G., Cordón-Preciado V., Alabert C., Pasero P. (2005) Mol. Cell 19, 699–706 [DOI] [PubMed] [Google Scholar]
- 17.Szyjka S. J., Viggiani C. J., Aparicio O. M. (2005) Mol. Cell 19, 691–697 [DOI] [PubMed] [Google Scholar]
- 18.Hodgson B., Calzada A., Labib K. (2007) Mol. Biol. Cell 18, 3894–3902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Noguchi E., Noguchi C., Du L. L., Russell P. (2003) Mol. Cell. Biol. 23, 7861–7874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Katou Y., Kanoh Y., Bando M., Noguchi H., Tanaka H., Ashikari T., Sugimoto K., Shirahige K. (2003) Nature 424, 1078–1083 [DOI] [PubMed] [Google Scholar]
- 21.De Antoni A., Gallwitz D. (2000) Gene 246, 179–185 [DOI] [PubMed] [Google Scholar]
- 22.Hori Y., Shirahige K., Obuse C., Tsurimoto T., Yoshikawa H. (1996) Mol. Biol. Cell 7, 409–418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Katou Y., Kaneshiro K., Aburatani H., Shirahige K. (2006) Methods Enzymol. 409, 389–410 [DOI] [PubMed] [Google Scholar]
- 24.Aparicio O. M., Weinstein D. M., Bell S. P. (1997) Cell 91, 59–69 [DOI] [PubMed] [Google Scholar]
- 25.Nedelcheva M. N., Roguev A., Dolapchiev L. B., Shevchenko A., Taskov H. B., Shevchenko A., Stewart A. F., Stoynov S. S. (2005) J. Mol. Biol. 347, 509–521 [DOI] [PubMed] [Google Scholar]
- 26.Santocanale C., Diffley J. F. (1998) Nature 395, 615–618 [DOI] [PubMed] [Google Scholar]
- 27.Mayer M. L., Pot I., Chang M., Xu H., Aneliunas V., Kwok T., Newitt R., Aebersold R., Boone C., Brown G. W., Hieter P. (2004) Mol. Biol. Cell 15, 1736–1745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Noguchi E., Noguchi C., McDonald W. H., Yates J. R., 3rd, Russell P. (2004) Mol. Cell. Biol. 24, 8342–8355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tanaka H., Katou Y., Yagura M., Saitoh K., Itoh T., Araki H., Bando M., Shirahige K. (2009) Genes Cells 14, 807–820 [DOI] [PubMed] [Google Scholar]
- 30.Komata M., Bando M., Araki H., Shirahige K. (2009) Mol. Cell. Biol. 29, 5008–5019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lou H., Komata M., Katou Y., Guan Z., Reis C. C., Budd M., Shirahige K., Campbell J. L. (2008) Mol. Cell. 32, 106–117 [DOI] [PMC free article] [PubMed] [Google Scholar]