Abstract
Deoxyhypusine synthase, an NAD+-dependent enzyme, catalyzes the first step in the post-translational synthesis of an unusual amino acid, hypusine (Nϵ-(4-amino-2-hydroxybutyl)lysine), in the eukaryotic initiation factor 5A precursor protein. Two putative deoxyhypusine synthase (DHS) sequences have been identified in the Leishmania donovani genome, which are present on chromosomes 20: DHSL20 (DHS-like gene from chromosome 20) and DHS34 (DHS from chromosome 34). Although both sequences exhibit an overall conservation of key residues, DHSL20 protein lacks a critical lysine residue, and the recombinant protein showed no DHS activity in vitro. However, DHS34 contains the critical lysine residue, and the recombinant DHS34 effectively catalyzed deoxyhypusine synthesis. Furthermore, in vivo labeling confirmed that hypusination of eukaryotic initiation factor 5A occurs in intact Leishmania parasites. Interestingly, the DHS34 is much longer, with 601 amino acids, compared with the human DHS enzyme (369 amino acids) and contains several unique insertions. To study the physiological role of DHS34 in Leishmania, gene deletion mutations were attempted via targeted gene replacement. However, chromosomal null mutants of DHS34 could only be obtained in the presence of a DHS34-containing episome. The present data provide evidence that DHS34 is essential for L. donovani and that structural differences in the human and leishmanial DHS enzyme may be exploited for designing selective inhibitors against the parasite.
Introduction
Leishmania is a protozoan parasite that results in a wide spectrum of diseases ranging from simple cutaneous lesions to the usually fatal visceral form. Leishmania donovani is the major causative agent of the visceral leishmaniasis. Pentavalent antimonials are the standard first line of treatment for leishmaniasis, but due to toxicity and increasing resistance of the present repertoire of drugs (1–3), an urgent need exists for newer drugs and drug targets.
The polyamine pathway is one such pathway that has been exploited successfully for antiparasitic chemotherapy (4). The enzymes ornithine decarboxylase, adenosylmethionine decarboxylase, and spermidine synthase (SPDS) have been identified as important drug targets (4). Spermidine, the final product of the polyamine biosynthetic pathway in kinetoplastids, acts as a precursor for trypanothione (a key molecule in defense against oxidative stress) (5) and hypusine (6). Hypusine (Nϵ-(4-amino-2-hydroxybutyl)lysine) is an unusual amino acid named after its two structural components, hydroxyputrescine and lysine (7). Hypusine synthesis is the result of a post-translational modification occurring exclusively in one cellular protein, eukaryotic initiation factor 5A (eIF5A),3 by the transfer of the 4-aminobutyl moiety from spermidine to its lysine side chain (8). Hypusine formation occurs mainly in two enzymatic steps (9). The first step is catalyzed by the enzyme deoxyhypusine synthase (DHS), which catalyzes the NAD+-dependent transfer of the 4-aminobutyl moiety of spermidine to a specific lysine residue of the eIF5A precursor protein to form an intermediate, deoxyhypusine (10, 11). This intermediate is subsequently hydroxylated by the enzyme deoxyhypusine hydroxylase (12), which completes the synthesis of hypusine and maturation of eIF5A.
Biosynthesis of hypusine represents a novel, unique, and the most specific post-translational modification known in literature. In mammalian cells, inhibitors of spermidine biosynthesis (13) and hypusine biosynthetic enzymes, DHS and deoxyhypusine hydroxylase, exert antiproliferative effects (14–16), including in cancer cell lines, and cause arrest of cell cycle progression (17). DHS has also been suggested as the potential target for antiretroviral therapy (18). Direct evidence showing the essential nature of eIF5A and DHS comes from gene inactivation studies in Saccharomyces cerevisiae. Disruption of the two eIF5A genes (TIF51A and TIF51B) (19) and the deoxyhypusine synthase gene (20) produces a lethal phenotype.
Deoxyhypusine synthase is found in archaea and all eukaryotes. It is a highly conserved protein and exists as a single gene product of 300–400 amino acids (33–48 kDa) in most eukaryotes. The native enzymes of human and yeast recombinant proteins exist as a ∼160-kDa molecule, a tetramer of four identical subunits of ∼40 kDa (21, 22). Crystal structures of the human DHS tetrameric enzyme in complex with a cofactor NAD and its inhibitor have been reported (23, 24).
In the present study, we, for the first time, report the identification, molecular cloning, expression, and characterization of the enzyme deoxyhypusine synthase from L. donovani. Two DHS-like genes have been identified in L. donovani, which show low homology with the human DHS. Both genes have been cloned and expressed, but only one, DHS34, exhibits deoxyhypusine synthase activity. Gene replacement studies for DHS34 indicate that the enzyme deoxyhypusine synthase and eIF5A modification play an essential role in cell viability of this pathogenic organism.
EXPERIMENTAL PROCEDURES
Materials
Radiolabeled spermidine trihydrochloride [1,8-3H]spermidine (16.6 Ci/mmol) and radiolabeled [1,4-14C]putrescine (110 mCi/mmol) were purchased from PerkinElmer Life Sciences. [α-32P]dCTP (3000 Ci/mmol) was acquired from Amersham Biosciences. All restriction enzymes and DNA-modifying enzymes were obtained from MBI Fermentas (Germany). Neomycin, phleomycin, and hygromycin were obtained from Sigma. N1-guanyl-1,7-diaminoheptane was synthesized as described earlier (14). Recombinant human DHS was purified from Escherichia coli cells as reported earlier (21). The other materials used in this study were of analytical grade and were commercially available.
Parasite and Culture Conditions
L. donovani AG83 (MHOM/IN/1983/AG83) promastigotes were cultured at 22 °C in modified M199 medium (Sigma) supplemented with 100 units/ml penicillin (Sigma), 100 μg/ml streptomycin (Sigma), and 10% heat-inactivated fetal bovine serum (Hyclone). The genetically manipulated parasites were derived from the wild type LdBob strain of L. donovani (25) that was originally obtained from Dr. Stephen Beverley (Washington University, St. Louis, MO). LdBob promastigotes and axenic amastigotes were routinely cultivated at 28 °C, pH 7.4, and 37 °C, pH 5.5, respectively, in the culture media previously reported (25, 26). All parasite strains were cycled back and forth between the promastigote and axenic amastigote forms. Wild type parasites were routinely cultured in media with no drug supplementations, whereas the DHS heterozygotes were maintained in either 50 μg/ml hygromycin (DHS34/HYG::dhs) or 50 μg/ml phleomycin (DHS34/PHLEO::dhs). The DHS34/HYG::dhs strain containing the pX63PAC-DHS episome was grown in 50 μg/ml hygromycin and 20 μm puromycin, and the putative Δdhs34[pX63PAC-DHS34] clones were kept in 50 μg/ml hygromycin, 50 μg/ml phleomycin, and 20 μm puromycin.
Cloning of Putative DHS Genes from L. donovani
A 1.14-kb DNA fragment encompassing the whole open reading frame (ORF) of the L. donovani DHS-like-gene DHSL20 was amplified from genomic DNA using a sense primer with a flanking BamHI site, 5′-CGCGGATCCATGCTTGCCTCTGTCCCAG-3′ and an antisense primer with a flanking HindIII site, 5′-CCCAAGCTTGCGGCGCTCTGCGCTAC-3′. A 1.8-kb DNA fragment encompassing the whole ORF of the L. donovani DHS34 gene was also amplified from genomic DNA using the sense primer with a flanking BamHI site, 5′-CGGGATCCATGGCGAATATTGCGGAATC-3′, and the antisense primer with the flanking HindIII site, 5′-CCCAAGCTTGAGTGCCCGGAGACGTCTTG-3′. PCR was performed in 50-μl reaction volumes containing 100 ng of genomic DNA, 25 pmol each of the gene-specific forward and reverse primers, 200 μm each of dNTPs, 2 mm MgCl2, and 5 units of Pfu DNA polymerase (MBI Fermentas). The conditions of the PCR were as follows: 94 °C for 10 min, 30 cycles of 94 °C for 1 min, 60 °C for 30 s, and 72 °C for 1.5 min for DHSL20 and 72 °C for 3 min for DHS34. Final extension was carried out for 10 min at 72 °C. A single ∼1140-bp (DHSL20) and 1806-bp (DHS34) PCR products were obtained. Each gene was then cloned into the pTZ57R/T (MBI Fermentas) vector and sequenced.
Sequence and Structural Analysis
Sequence comparison of DHSL20 and DHS34 with other sequences of the data base was done using the search algorithm BLAST (available on the NCBI Web site). Multiple alignment of amino acid sequences was performed using the program ClustalW (available on the World Wide Web) The bootstrap analysis was done and the phylogenetic tree was constructed using the software MEGA version 4.0 (27). DHSL20 and DHS34 protein sequences were searched against the multiple position-specific substitution matrix data base (MulPSSM) (28) of Pfam protein domain families (29) using an RPS-BLAST approach (30). Local alignments of DHS34 and DHSL20 protein were compared with human DHS to analyze the conservation of important amino acid residues. Subsequently, DHS34 was searched against the Protein Data Bank (31) using the BLAST program (32) for closely related DHS homologs with known three-dimensional structure. The DHS34 protein sequence was modeled on the basis of the tertiary structure of the human DHS template (Protein Data Bank code 1dhs) (23) using MODELLER version 8.0 (available on the World Wide Web) (33). The generated model was energy-minimized using Kollman united atom forced field in SYBYL (Tripos Inc., St. Louis, MO) to relieve short contacts if any. The overall fit of the sequence to the modeled structure was checked using PROSA (34, 35). The stereochemistry of the models was evaluated using PROCHECK (36, 37).
Construction of DHSL20 and DHS34 Expression Vectors and Purification of the Proteins
The coding regions of the DHSL20 gene and DHS34 gene were subcloned into the BamHI-HindIII site of pET30a vector (Novagen). The fidelity of the PCR amplification of L. donovani DHSL20 and DHS34 was confirmed by automated DNA sequencing. The recombinant constructs were transformed into the BL21 (DE3) strain of E. coli. Expression from the constructs DHSL20-pET30a and DHS34-pET30a was induced at an A600 of 0.6 with 0.25 mm isopropyl-1-β-d-galactopyranoside at 16 °C for 12 h. Bacteria were then harvested by centrifugation at 5000 × g for 10 min, and the cell pellet was resuspended in binding buffer (50 mm Tris-Cl, pH 8.0, 10 mm imidazole, 300 mm sodium chloride, 1 mm phenylmethylsulfonyl fluoride, and protease inhibitor mixture). The resulting cell suspension was sonicated six times for 30 s with 1-min intervals. The lysate was centrifuged at 10,000 × g for 30 min at 4 °C. The resulting supernatant, which contained the protein, was loaded onto pre-equilibrated Ni2+-nitrilotriacetic-agarose resin (Qiagen). The mixture was kept on a rocking platform for 2 h at 4 °C. It was centrifuged at 400 × g for 1 min at 4 °C. The supernatant was removed, and the resin was washed three times with wash buffer (50 mm Tris-Cl, pH 8.0, 20 mm imidazole, 300 mm NaCl, 1 mm phenylmethylsulfonyl fluoride, and protease inhibitor mixture). The protein was eluted with increasing concentrations of imidazole. Imidazole was removed, and the purified protein was found to be >95% pure as judged by SDS-PAGE. The purified protein was aliquoted and stored at −80 °C in 50 mm Tris-Cl, pH 8.0, 300 mm NaCl buffer.
Deoxyhypusine Synthase Assay
Deoxyhypusine synthase activity was assayed by measuring incorporation of radioactivity from [1,8-3H]spermidine into L. donovani eIF5A (accession number ABO34138.1) precursor protein (38, 39). A typical reaction mixture contained, in a total volume of 50 μl, 0.2 m glycine-NaOH buffer, pH 9.2, 1 mm dithiothreitol, 1 mm NAD, 200 μg of bovine serum albumin, ∼5 μm (4 μCi) [1,8-3H]spermidine, 5 μm eIF5A, and the indicated amount of recombinant DHS. The reaction was incubated at 37 °C for 60 min. The reaction mixture was terminated by the addition of 10% trichloroacetic acid containing putrescine, spermidine, and spermine (1 mm each). The samples were then centrifuged at 10,000 × g for 5 min at 4 °C. The precipitate obtained was washed repeatedly with 10% trichloroacetic acid containing polyamines to remove the [3H]spermidine bound to the pellet. The [3H]deoxyhypusine formed was measured by dissolving the pellet in 100 μl of 0.1 n NaOH, and the counts were taken. One unit of enzyme activity is defined as the amount of enzyme catalyzing the formation of one pmol/h of deoxyhypusine. Deoxyhypusine formation was also confirmed by detection of radiolabeled eIF5A by fluorography after SDS-PAGE.
In Vivo Labeling of eIF5A
Wild type and spermidine synthase-deficient (Δspdsyn) parasites (40) were incubated in growth medium containing radiolabeled [1,4-14C]putrescine and 100 μm unlabeled spermidine for 4 days. Subsequently, parasites were harvested and sonicated, and lysates were fractionated by SDS-PAGE. The gel was stained with Coomassie Blue and subsequently dried and exposed for autoradiography.
Molecular Constructs for the Replacement of the DHS34 Alleles
To construct drug resistance cassettes for targeted DHS34 gene replacement, the 5′- and 3′-flanking regions of DHS34 were amplified by PCR. The 5′-flank was amplified using genomic DNA as a template and the following primers: AAGCTTCCAGAGTACACTGGATAGGC (HindIII site underlined) and GTCGACTCTCTTGTCTCACCGCC (SalI site underlined). The 423-bp 5′-flanking region was then cloned into the HindIII/SalI sites of pX63HYG to generate pX63HYG-5′DHS34. The 3′-flank was amplified using the following primer pair: CCCGGGGGAGTGGGCTGGTGGCG (SmaI site underlined) and AGATCTCGCGCCGCAACCTCGTAAAAG (BglII site underlined). The 584-bp 3′-flanking region was cloned into the SmaI/BglII site of pX63HYG-5′DHS34 to generate the pX63HYGΔdhs34 gene replacement construct. The same 5′-flank was cloned into the HindIII/SalI sites of the pX63PHLEO vector to generate pX63PHLEO-5′DHS34. Because of the presence of a SmaI site within the coding region of PHLEO, a different strategy was employed to insert the DHS34 3′-flanking region into the pX63PHLEO-5′DHS34. For the generation of the 3′-flanking fragment, the following primers were used: GGATCCGGAGTGGGCTGGTGGCG (BamHI underlined) and AGATCTCGCGCCGCAACCTCGTAAAAG (BglII underlined) to generate a 580-bp PCR fragment. The fragment was cloned into pX63PHLEO-5′DHS34 to generate the gene replacement construct pX63PHLEOΔdhs34. PCRs were performed using the AdvantageTM PCR kit (BD) at the following conditions: 94 °C for 10 min, 30 cycles of 94 °C for 1 min, 60 °C for 1 min, 72 °C for 1 min, and a final extension at 72 °C for 10 min. Restriction mapping and limited nucleotide sequence analysis was used to confirm the correct orientation of the 5′ and 3′ DHS34 flanking region within the pX63HYGΔdhs34 and pX63PHLEOΔdhs34 vectors.
Cloning of the pX63PAC-DHS34 Episome
To generate the pX63PAC-DHS34 complementation construct, the following primers were used to amplify the DHS34 coding region by PCR: GGATCCATGGCGAATATTGCGGAATCTG (BamHI site underlined) and AGATCTTCATTCTCCAAGACACGGGTGG (BglII site underlined). PCRs were performed using the high fidelity HFAdvantageTM PCR kit (BD Biosciences) under the following conditions: 94 °C for 10 min, 30 cycles of 94 °C for 1 min, 60 °C for 2 min, 72 °C for 1 min, and a final extension at 72 °C for 10 min. The 1.8-kb BamHI-BglII fragment was subcloned into the BamHI/BglII sites of pX63PAC. The correct orientation and sequence fidelity of the insert was verified by nucleotide sequence analysis.
Generation of Genetically Manipulated Parasites
The pX63HYGΔdhs34 and pX63PHLEOΔdhs34 plasmids were digested with HindIII and BglII to liberate the deletion fragments containing the drug resistance cassette and DHS34 flanking regions (designated HYGΔdhs34 and PHLEOΔdhs34). These fragments were gel-purified and transfected into wild type LdBob parasites according to standard protocols (26, 41). The HYGΔdhs34 and PHLEOΔdhs34 gene replacement cassettes were individually transfected into wild type parasites, and colonies were selected from semisolid agar plates in the presence of 50 μg/ml hygromycin or 50 μg/ml phleomycin, respectively. The genotype of the putative heterozygous parasites, DHS34/HYG::dhs34 and DHS34/PHLEO::dhs34, was confirmed by Southern blot analysis using the 5′- and 3′-flanking regions as probes. The growth phenotype of heterozygous parasites was established by seeding 2 × 105 cells/ml, and parasites were counted after 4 days on a hemocytometer. The heterozygous parasites, DHS34/HYG::dhs34 and DHS34/PHLEO::dhs34, were subsequently transfected with the PHLEOΔdhs34 and HYGΔdhs34 replacement cassettes, respectively, in an attempt to generate homozygous gene deletion mutants. Colonies were selected on semisolid agar plates containing 50 μg/ml hygromycin and 50 μg/ml phleomycin, and parasite genotypes were established by Southern blotting. The pX63PAC-DHS34 episome was transfected into DHS34/HYG::dhs34 heterozygous parasites, and parasites containing the plasmid were selected in 20 μm puromycin. These DHS34/HYG::dhs34[pX63PAC-DHS34] parasites were subsequently transfected with the purified PHLEOΔdhs34 gene replacement construct, and colonies were selected on semisolid agar plates containing 50 μg/ml hygromycin, 20 μm puromycin, and 50 μg/ml phleomycin to isolate triple drug-resistant parasites. The genotypes of several clones were established by Southern blotting.
Molecular Characterization of Genetically Manipulated Parasites
Genomic DNA from wild type and genetically manipulated parasites was prepared for Southern blot and PCR analysis using the DNAeasy kit (Qiagen Inc., Valencia, CA). DNA was digested with the appropriate restriction enzymes and probed with either 5′- or 3′-flanking regions or the DHS34 coding region probe using highly stringent hybridization conditions. Furthermore, a PCR strategy was devised to confirm the results obtained by Southern blotting. The following primers were designed to amplify DNA fragments within the DHS34 coding region: the top primer 1 (ATGGCGAATATTGCGGAGTCT) and the bottom primers 7 (CAACACCCTCCTGCACAAGAA), 8 (GGCGTAGTAGTGCAGTGTG), and 10 (CACAGCCCTCAGCGCCTGCGC). Another set of primers was designed to amplify a fragment spanning from the 5′-flanking region into the coding region, the top primers 2 (cctcagtctgctatgctccc) and 4 (agaaagagcgatgtcgttgcctc) to hybridize to the 5′-flanking region, and the bottom primers 5 (cagattccgcaatattcgcCAT) and 6 (caccctcctgcacaagaaagcg) to hybridize to the DHS34 coding region. PCRs were performed using the AdvantageTM PCR kit (BD Biosciences) at the following conditions: 94 °C for 10 min, 30 cycles of 94 °C for 1 min, 58 °C for 1 min, 72 °C for 1 min, and a final extension at 72 °C for 10 min.
RESULTS
Sequence Analysis and Genomic Organization
Two DHS sequences have been identified in each of the Leishmania infantum and Leishmania major genomes (see the EMBL-EBI Web site). Two DHS-like sequences have also been identified in the Trypanosoma brucei (Tb927.1.870 and Tb10.70.4900) and Trypanosoma cruzi genomes (Tc00.1047053511421.60 and Tc00.1047053504119.29), whereas most other organisms have only one DHS gene copy. BLAST analysis of L. major and L. infantum genomes revealed that the deoxyhypusine synthase-encoding genes are present on chromosomes 20 (DHSL20) and 34 (DHS34). Southern blot analysis of the DHSL20 and DHS34 showed that these were single copy genes (data not shown). Sequence analysis, data base search, and alignment of the L. donovani DHSL20 and DHS34 amino acid sequences were performed. The DHSL20 amino acid sequence had a single open reading frame consisting of 1140 bp (L. donovani DHS, EF512031). The ORF encoded a putative polypeptide of 379 amino acids, with a predicted molecular mass of ∼42 kDa. Amino acid sequence alignment of DHSL20 protein with homologous sequences from other species revealed that it shares 63.4% identity with the T. cruzi (Tc00.1047053511421.60), 56.1% identity with T. brucei (Tb927.1.870), and 30.7% with S. cerevisiae (P38791) proteins (Fig. 1).
FIGURE 1.
Multiple sequence alignment of DHS-like protein (DHSL20) sequences from L. donovani (ABP65295), T. cruzi (Tc00.1047053511421.60), T. brucei (Tb927.1.870), S. cerevisiae (P38791), and human (P49366). Conserved residues are shaded in black, showing complete identity, and residues shaded in gray are similar. NAD+ binding regions are enclosed in yellow rectangular boxes. The catalytic lysine residue is shown on a cyan background. Other residues important for the overall functioning of enzyme are highlighted on a red background.
Furthermore, we proceeded with the characterization of DHS34, which is located on chromosome 34. The sequence identity between DHSL20 protein and DHS34 is only 21.7%. A single open reading frame consisting of 1806 bp was identified (L. donovani DHS, EU864544). The ORF encoded a putative polypeptide of 601 amino acids, with a predicted molecular mass of ∼66 kDa. Amino acid sequence alignment of DHS34 with homologous sequences from other species revealed that it is 46.5% identical to T. cruzi (Tc00.1047053504119.29), 45% with T. brucei (Tb10.70.4900), and 33.3% with S. cerevisiae (P38791). Interestingly, DHSL20 protein has closer identity with the T. brucei (56.1%) and T. cruzi genes (63.4%) compared with DHS34 (45% with T. brucei and 46.5% with T. cruzi).
Alignment of DHSL20 and DHS34 protein with the human DHS (P49366) shows conservation of important residues, which are essential for the overall functioning of the enzyme (Figs. 1 and 2). The leishmanial DHS34 and DHSL20 protein share 32.8% (Fig. 2) and 34.7% (Fig. 1) sequence identity with the human DHS sequence, respectively.
FIGURE 2.
Multiple sequence alignment of DHS34 protein sequences from L. donovani (ACF75531), T. cruzi (Tc00.1047053504119.29), T. brucei (Tb10.70.4900), S. cerevisiae (P38791), and human (P49366). Conserved residues are shaded in black, showing complete identity, and residues shaded in gray are similar. NAD+ binding regions are enclosed in yellow rectangular boxes. The catalytic lysine residue is shown on a cyan background. Other residues important for the overall functioning of enzyme are highlighted on a red background.
Interestingly, although the DHSL20 protein with 379 amino acid residues is of similar length as the human enzyme, the DHS34 is much longer with 601 amino acids. Alignment of DHS34 with the human DHS shows that there is a big insertion in DHS34 from residue 130 to 354 (Fig. 3). No homologs were found for this region using standard homology detection methods like PSI-BLAST. A reverse transcriptase PCR on DHS34 (parasite RNA as template) confirmed that the entire gene (with all insertions) was transcribed (data not shown).
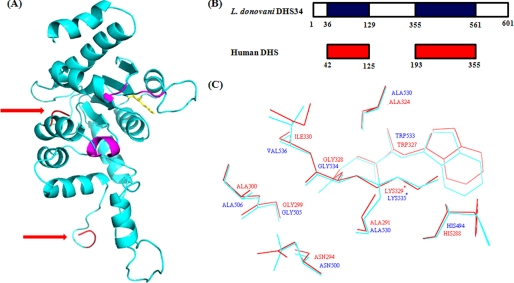
FIGURE 3.
Structural modeling of DHS34. A, the DHS34 model was generated using MODELLER (version 8.0). The catalytic lysine residue is shown in yellow sticks. The regions of insertion are depicted by red arrows. The probable NAD+ binding region is shown in magenta. B, alignment showing homologous residues from L. donovani DHS34 (blue) and human DHS (red). C, residues around active site lysine (indicated by asterisks) of both human DHS (shown in red) and modeled DHS34 (shown in cyan).
The residues that have been identified as important for NAD+ binding in the human DHS, Asn106, Asp238, His288, and Asp313, are conserved in DHS34 (42). Residues equivalent to human DHS, His288, Trp327, Lys329, Asp316, and Glu323, which participate in spermidine binding and reaction mechanism, are also conserved (23, 42–44). Amino acids corresponding to Lys287 (important for covalent intermediate formation) (45) and the catalytic center (Lys329) (43) of the human DHS and the L. donovani DHS34 (Lys535) show the conservation.
In the case of the DHSL20 protein, some additional residues corresponding to Glu137, Asp243 (important for spermidine binding), and Ser317 (mutation of which leads to reduced NAD+ binding) are also conserved (42). Most importantly, however, the catalytic lysine (Lys329 in humans), which is indispensable for the enzymatic reaction and is present in the active center for catalysis (enzyme-substrate intermediate formation), is substituted by a non-polar amino acid, leucine (Leu344 in Leishmania). This leucine residue, Leu344 (Lys329 in humans), in DHSL20 is highlighted in cyan (Fig. 1). Such a mutation is likely to render the enzyme inactive (43, 46).
The computer model that we generated for DHS34 shows that the probable NAD+ binding regions are spatially closer in the tertiary structure. Alignment of DHS34 with human DHS shows that there is a big insertion in DHS34 from residues 130–354 (total 225 residues) (Fig. 3B). The large 225-residue insertion is marked in Fig. 3A. Fig. 3B shows the homologous regions between DHS34 of L. donovani and the human DHS. Region 36–129 (94 residues) of L. donovani DHS34 is homologous to region 42–125 of the human DHS (84 residues), and region 355–561 (total 207 residues) is homologous to region 193–355 (163 residues) of the human DHS. However, residues 1–36 and 562–601 (40 residues) of DHS34 have no homology to the human DHS. There are four NAD+ binding regions in the human DHS: 1) SNLIS (residues 105–109), 2) TAG (residues 131–133), 3) TA (residues 308 and 309), and 4) DA (residues 342 and 343). The alignment of the human DHS and DHS34 shows the presence of SGG of DHS34 aligning with TAG of human DHS, which is the nucleotide binding motif. This region matches with a very small fragment of human DHS and lies within a big insertion. The remaining three regions come together to form the NAD+ binding region in DHS34; however, the NAD+ binding regions are interrupted by the large insertion of 225 residues.
Analysis of residues around the catalytic center (Lys329) (within a 4-Å radius) of the human DHS and the modeled L. donovani DHS34 (Lys535) shows the conservation of all of the residues except Ile330 of human DHS, which is substituted by Val536 in the case of Leishmania (Fig. 3C).
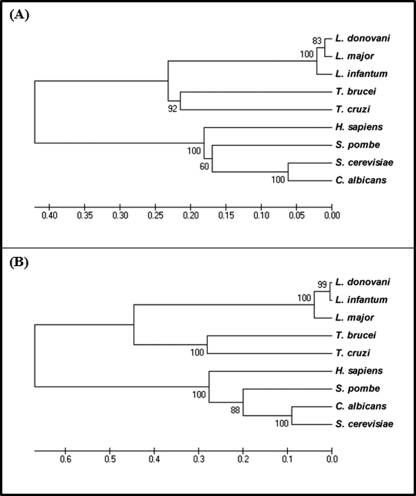
A phylogenetic tree has been constructed for both putative DHS sequences (Fig. 4) using representative DHS sequences. The tree for DHSL20 sequence indicates a close evolutionary relationship of L. donovani with trypanosome species among the kinetoplastid protozoa (Fig. 4A). Another phylogenetic tree has been constructed (Fig. 4B) using the L. donovani DHS34 sequence and other representative DHS sequences. This tree also indicates a close evolutionary relationship of L. donovani and T. cruzi among the kinetoplastid protozoa. In both cases, the kinetoplastid DHS sequences are conserved among each other and are distantly related with human DHS.
FIGURE 4.
A, phylogenetic analysis by Unweighted Pair Group Method with Arithmetic Mean of DHS-like gene, DHSL20. The phylogram presented was a consensus of 500 bootstrap replicates constructed using the MEGA program (version 4.0). The numbers at the node present the percentage of trees with the same node among all of the bootstraps. The accession numbers of the gene sequences used are as follows: L. donovani (ABP65295), L. major (LmjF20.0250), L. infantum (LinJ20.0270), T. brucei (Tb927.1.870), T. cruzi (Tc00.1047053511421.60), Homo sapiens (P49366), Schizosaccharomyces pombe (CAA22194), S. cerevisiae (P38791), Candida albicans (XM_716612.1). B, phylogenetic analysis by Unweighted Pair Group Method with Arithmetic Mean of the DHS34 gene. The phylogram presented was a consensus of 500 bootstrap replicates constructed using the MEGA program (version 4.0). The numbers at the node present the percentage of trees with the same node among all the bootstraps. The accession numbers of the gene sequences used are as follows: L. donovani (ACF75531), L. major (LmjF34.0330), L. infantum (LinJ34.0350), T. brucei (Tb10.70.4900), T. cruzi (Tc00.1047053504119.29), H. sapiens (P49366), S. pombe (NP_595146.1), C. albicans (XP_721705.1), S. cerevisiae (P38791).
Overexpression and Purification of the Deoxyhypusine Synthase Enzymes in E. coli
Both DHSL20 and DHS34 genes were overexpressed in E. coli cells. A protein with an estimated molecular mass of ∼48 kDa was induced; the size correlated well with the amino acid composition of DHSL20 protein (∼42 kDa) with a His6 tag (∼6 kDa) at the C terminus (Fig. 5A). Purification of DHSL20 by metal affinity chromatography yielded ∼1 mg of purified protein/liter of bacterial culture. The recombinant DHSL20 from L. donovani did not show any activity under varying pH, eIF5A, and spermidine concentrations (data not shown). This is to be expected because DHSL20 lacks the catalytic lysine residue at residue 344 and contains leucine instead. In an effort to determine whether the lack of the catalytic lysine is solely responsible for the lack of its activity, we have converted this leucine residue (Leu-344) to lysine by site-directed mutagenesis. This mutant enzyme of DHSL20 (L344K) was still devoid of any DHS activity (data not shown), suggesting that other differences in amino acid sequences also contribute to its inactivity.
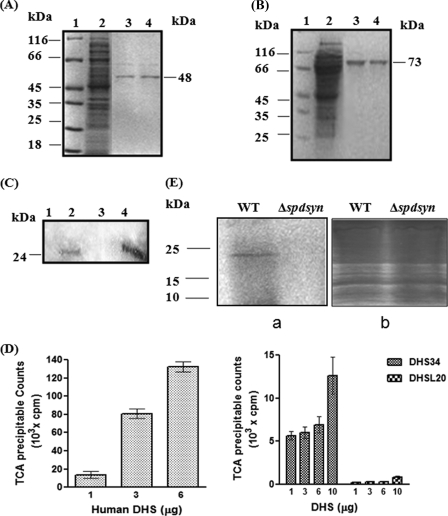
FIGURE 5.
Characterization of DHSL20 and DHS34 proteins. A, purification of DHSL20 protein on Ni2+-nitrilotriacetic acid affinity resin. Lane 1, molecular weight protein marker (MBI Fermentas); lane 2, flow-through; lanes 3 and 4, eluted fractions showing purified protein with buffer containing 300 mm imidazole. B, purification of DHS34 protein on Ni2+-NTA affinity resin. Lane 1, molecular weight marker; lane 2, flow-through; lanes 3 and 4, eluted fractions showing purified DHS34 protein with buffer containing 300 mm imidazole from the affinity column. C, identification of 3H-labeled deoxyhypusine-containing eIF5A product from DHS reaction mixture by SDS-PAGE and fluorography. Lane 1, DHS34 minus eIF5A; lane 2, DHS34 plus eIF5A; lane 3, human recombinant DHS minus eIF5A; lane 4, human recombinant DHS plus eIF5A. D, comparison of trichloroacetic acid-precipitable counts obtained in the DHS enzyme assay using different concentrations of recombinant enzymes, human DHS (hDHS), DHS34, or DHSL20. A DHS enzyme assay was performed as reported under “Experimental Procedures.” Results are mean ± S.D. of triplicate samples. E, in vivo metabolic labeling of 18-kDa protein in the wild type cells (WT) but not in the (Δspdsyn) mutant after culture with [1, 4-14C]putrescine. The gel was stained with Coomassie Blue (b) and subsequently dried and exposed to autoradiography (a).
Upon induction of E. coli transformed with the DHS34 expression vector, a protein that matched the estimated ∼73 kDa of DHS34 with a His tag was induced (Fig. 5B). Purification yielded ∼3 mg of purified protein per liter of bacterial culture.
The kinetic parameters of the enzyme DHS34 were determined using [1,8-3H]spermidine and L. donovani eIF5A as the substrates. The enzyme showed a specific activity of 569 units/mg of protein, which is much lower than the specific activity of the human recombinant DHS. Low specific activity of DHS34 could be due to the fact that one of the NAD+ binding regions in the DHS34 is present in the big long insertion when compared with human DHS, and how this region positions itself to carry out this reaction is not clear. The Km value of DHS34 for spermidine was estimated to be 23.2 μm, which was much higher than that of human (7.2 μm) (21) and S. cerevisiae (12.5 μm) (22), whereas the Km value of DHS34 for eIF5A was 1.77 μm, higher than human (0.6 μm) (21) but comparable with that of S. cerevisiae (1.01 μm) (22). Deoxyhypusine formation was also confirmed by running the reaction mixture on an SDS-polyacrylamide gel, and the labeled eIF5A was detected by fluorography. Fig. 5C shows ∼24-kDa radiolabeled protein in lane 2, confirming the deoxyhypusine formation on L. donovani eIF5A by DHS34. A similar ∼24-kDa labeled protein was confirmed in the reaction mixture containing human DHS (Fig. 5C, lane 4). Fig. 5D shows a comparison of trichloroacetic acid-precipitable radioactivity obtained in the DHS reaction mixture with different concentrations of the recombinant human DHS, DHS34, and DHSL20. Interestingly, N1-guanyl-1,7-diaminoheptane, a potent and competitive inhibitor of spermidine binding for human DHS, had little inhibitory effect on the leishmanial recombinant DHS34 (data not shown).
Human DHS native enzyme is a homotetramer consisting of two dimers of 40-kDa subunits. We questioned if Leishmania DHS34 native enzyme also exists as a tetramer. Gel exclusion chromatography of DHS34 on a Sephacryl S-300 column resulted in a peak corresponding to a molecular mass of 137 ± 7.6 kDa (supplemental Fig. 1), suggesting that DHS34 exists as a dimer in its native form.
In Vivo Activity of Deoxyhypusine Synthase
To assess if the hypusination of eIF5A occurs in intact Leishmania parasites, a labeling experiment was performed in both wild type and spermidine synthase-deficient parasites (Δspdsyn) using radiolabeled putrescine (40). The Δspdsyn parasites are not able to convert putrescine to spermidine, which donates the aminobutyl group for the hypusination of eIF5A, and depend on exogenous supply of spermidine for growth. Fig. 5E shows that 18-kDa eIF5A was radiolabeled upon culture with 14C-labeled putrescine in the wild type parasite but not in the Δspdsyn mutant.
Gene Deletion Studies for Deoxyhypusine Synthase 34
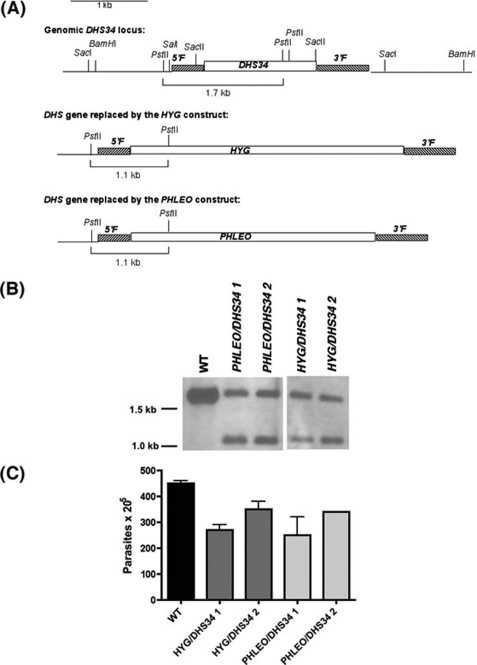
We attempted to delete the DHS34 gene by targeted gene replacement techniques to determine if this enzyme is vital for the parasite. Heterozygous parasites were generated, in which one DHS34 gene was replaced with either the phleomycin drug resistance gene or the hygromycin drug resistance gene: PHLEO::dhs34/DHS34 (PHLEO/DHS34) and HYG::dhs34/ DHS34 (HYG/DHS34) (Fig. 6B). Interestingly, the heterozygous parasites consistently showed a slight growth delay compared with wild type parasites (Fig. 6C). It is reasonable to assume that a gene dosage effect resulted in the production of less DHS protein, and such a conjecture would suggest that DHS34 is involved in optimal cell proliferation.
FIGURE 6.
A, restriction map of the DHS34 locus and the predicted rearrangements occurring during HYG and PHLEO replacement of DHS34 in the chromosomal locus. B, Southern blot analysis of wild type (WT) and heterozygous parasites. Genomic DNA from wild type parasites, two PHLEO/DHS34 (1 and 2), and two HYG/DHS34 (1 and 2) heterozygous cell lines were digested with PstI and probed with a PCR fragment corresponding to the 5′-flanking sequence. Molecular weight markers are indicated to the left of the blot. C, proliferation of heterozygous parasites. Wild type parasites, two HYG/DHS34 (1 and 2), and two PHLEO/DHS34 (1 and 2) heterozygous cell lines were incubated at 2 × 105 parasites/ml and counted after 4 days on a hemocytometer. The experiment was set up in duplicates and repeated several times with similar outcomes.
Several attempts to replace both DHS34 gene copies to generate homozygous gene deletion mutants (Δdhs34) failed. Although several clones resistant to both drugs were obtained, Southern blot analyses demonstrated that the DHS34 gene was still present in the genome of these parasite lines (data not shown), indicating that DHS34 may be an essential gene.
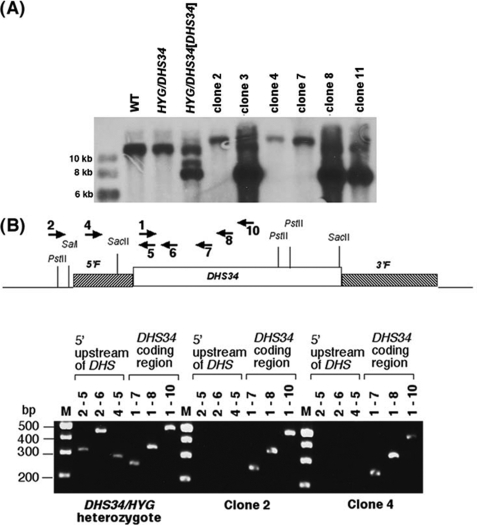
To further test this conjecture, a chromosomal DHS34 gene deletion was attempted in the presence of a rescuing episome that contained the leishmanial DHS34 gene (pX63PAC-DHS34). HYG/DHS34 heterozygous parasites, containing pX63PAC-DHS34, were then transfected with the PHLEOΔdhs34 gene deletion constructs. Southern blot and PCR analysis was performed to investigate the genotype of these triple drug-resistant parasites. Genomic DNA derived from wild type, HYG/DHS34 heterozygotes, HYG/DHS34 heterozygotes transfected with pX63PAC-DHS34, and several triple drug-resistant clones was probed with the DHS34 coding region probe (Fig. 7A). DNA from wild type and heterozygous parasites showed a ∼11 kb band hybridizing with DHS34, corresponding to the chromosomal DHS34 gene. This ∼11-kb DHS34 fragment was replaced with a higher molecular weight band, suggesting that some clones had indeed lost the chromosomal DHS34 gene (clones 2, 4, and 7). The hybridizing intensity of this band was similar to that of wild type DNA, suggesting that the DHS34 gene was present as a single copy, either on a single episome or integrated into the genome. Since the size of pX63PAC-DHS34 was predicted to be ∼8 kb and contained a single BamHI restriction site, the ∼12 kb band hybridizing to DHS34 was indicative of a random integration into the chromosome. Other clones (3, 8, 11) showed an intense band at ∼8 kb, suggesting the presence of several copies of the pX63PAC-DHS34 episome in these parasites, in addition to the ∼11-kb chromosomal DHS34 fragment. A control cell line, the heterozygous HYG/DHS34 parasites transfected with pX63PAC-DHS34, exhibited the ∼11-kb DHS34 chromosomal fragment and a ∼9 kb and an intense ∼8 kb episomal band hybridizing to the DHS34 coding region probe. To further bolster the premise that a chromosomal DHS34 knock-out had occurred in complemented clones 2, 4, and 7, a PCR strategy was applied (Fig. 7B). Primer pairs designed to the DHS34 coding region (primers 1 and 7, 1 and 8, and 1 and 10) generated fragments when genomic template DNA was used derived from DHS34/HYG heterozygous parasites and clones 2 and 4, as expected. Additional primer pairs were designed to generate fragments only if the DHS34 gene was present in its chromosomal context because the upstream primers (primers 2 and 4) were complementary to a region upstream of the gene, and the downstream primers were complementary to the coding region (primers 5 and 6). The primer combinations 2 and 5, 2 and 6, and 4 and 5 generated fragments only from DNA derived from the heterozygous cell line but not from clones 2 and 4. The PCR analysis demonstrated that DHS34 in the chromosomal location could only be found in the heterozygous cell line. Thus, although heterozygous parasites could be created for both the HYGΔdhs34 and PHLEOΔdhs34 gene deletion constructs, a chromosomal knock-out of DHS34 was only achieved when the gene was provided on a complementing plasmid. Taken together, these data provide strong evidence that DHS34 is indeed an essential gene.
FIGURE 7.
A, Southern blot of genetically manipulated parasites. Genomic DNA was extracted from the following cell lines: wild type parasites (WT), heterozygous parasites (HYG/DHS34), heterozygous parasites transfected with pX63PAC-DHS34 (HYG/DHS34[DHS34]), and six clonal cell lines (clones 1, 3, 4, 7, 8, and 11) derived from the HYG/DHS34[DHS34] strain transfected with the PHLEOΔdhs34 gene replacement construct. B, the top panel shows a map of the DHS34 genomic locus and the locations of the primers used for PCR analysis. Genomic DNA from HYG/DHS34 heterozygous parasites (first panel) and from clones 2 and 4 was used as a template for PCR analysis. Primers 2 and 4 were designed to match the upstream region of the DHS34 gene, with primer 4 corresponding to a region (5′F) that was present in the gene deletion construct used for the chromosomal DHS34 replacements. Primer 1 was designed as a forward primer corresponding to the early coding region. Primers 5–10 were reverse primers corresponding to various regions of the DHS34 coding region. Molecular size markers (in kb) are indicated on the left.
DISCUSSION
In the present study, we, for the first time, report the identification, cloning, and characterization of DHS from L. donovani. Two DHS-like genes have been identified in L. donovani, but only one, DHS34, exhibits deoxyhypusine synthase activity. The ORF of this leishmanial DHS encodes a putative polypeptide of 601 amino acids and contains long intervening sequences compared with the human DHS, which is only 369 amino acids long (Fig. 2). DHS is an essential enzyme in mammalian and yeast cells and also appears to be vital for the parasite. Attempts to delete the DHS34 gene from the parasite genome failed, and chromosomal null mutants of DHS34 could only be obtained in the presence of an episome that contained the DHS34 gene (Figs. 6 and 7). Thus, the L. donovani DHS34 appears to be an essential protein, whose structure is expected be significantly different from its human counterpart.
Leishmania has two distantly related putative DHS genes. One, DHS34, is a true DHS because its recombinant protein showed DHS activity (Fig. 5), whereas the other, DHSL20 protein, does not function as such but may carry out a different unknown function. The phylogenetic tree analysis showed a close evolutionary relationship of both DHSL20 and DHS34 protein with trypanosome species among kinetoplastid protozoa. It also illustrates their common evolutionary origin and that they are distantly related to the higher eukaryotic DHS sequences. In most eukaryotes, a single DHS gene has been found. Only kinetoplastid genomes contain two DHS-like genes, and the DHS34 sequence with its extensive insertions appears to be unique to these organisms. The question that needs to be addressed is why would an organism have two of the DHS-like genes when only one of them shows DHS activity? It is possible that the DHSL20 protein performs a different, as of yet unknown function. The sequence identity between DHSL20 and DHS34 is only 21.7%. These two distantly related copies might have originated by a gene duplication event followed possibly by functional divergence of one of the copies. Alternatively, it is also plausible that one of the genes is a DHS cognate gene like that of cyanobacteria, myxobacteria, etc. and has been laterally transferred from the archaea (47). In certain plants, it has been proposed that homospermidine synthase evolved from DHS by gene duplication (48). However, amino acid sequence alignment of DHSL20 and DHS34 protein showed only 35.5 and 22.6% identity, respectively, with the homospermidine synthase from Senecio vernalis. Because of the recent discovery that the kinetoplastid AdoMetDC requires an inactive AdoMetDC-like protein for dimerization and activation (49), we tested the possibility of DHSL20 having a regulatory role in the activity of DHS34. DHS enzyme assays were performed in the presence of both DHSL20 and DHS34 at varying ratios; however, no significant increase in DHS34 enzyme activity was observed (data not shown).
DHS34 shares only 34.7% sequence identity with the human DHS. DHS34 exists as a dimer in its native form in contrast to human DHS, which is a tetramer. The residues that have been found to be important for NAD+ binding in the human DHS are conserved in both DHSL20 and DHS34 proteins. Furthermore, the residues that take part in spermidine binding and reaction mechanism are also conserved in the leishmanial proteins. However, a critical lysine residue (Lys329), which is the site of the enzyme-intermediate formation in human DHS (43), is replaced by a non-polar amino acid leucine (Leu344) in the case of the DHSL20 protein. It has been shown earlier in the human DHS that a Lys329 → Arg or Ala substitution abolishes the covalent enzyme-substrate intermediate formation and DHS activity (45). Site-directed mutagenesis of leucine residue to lysine (L344K) of DHSL20 showed no activity, thereby suggesting that other differences in amino acid sequences also contribute to its inactivity.
The DHS34 sequence codes for 601 amino acids and is much longer than the human DHS (369 amino acids) and DHSL20 (379 amino acids). The deoxyhypusine synthase domain is interrupted by a long intervening sequence that spans from amino acid 130 to 354, which does not show any significant homology with any other sequence. It is unknown whether this intervening sequence of DHS34 adds a special function unique to the parasite DHS. The computer-generated model of DHS34 shows the conservation of residues around the catalytic center. The amino acid sequence alignment of DHS34 with human DHS shows that one of the NAD+ binding regions is present in the 225-amino acid-long insertion of DHS34 that might interrupt the NAD+ binding regions as they come together spatially to form the perfect NAD+ binding site.
Disruption of eIF5A genes (19, 50) and DHS genes (20, 51) in S. cerevisiae leads to a lethal phenotype, demonstrating the importance of deoxyhypusine modification in eukaryotic cell survival. Deoxyhypusine hydroxylase is an essential gene in the fruit fly and C. elegans but not in yeast, suggesting an evolutionary progression of the requirement of hypusine modification in eukaryotes. In Leishmania, chromosomal null mutants of DHS34 could only be obtained in the presence of a rescuing episome that contained the DHS34 gene, suggesting that DHS34 is essential for L. donovani and that the hypusination pathway also plays a vital role in L. donovani survival and proliferation.
Despite conservation of some of the active site amino acid residues between the human and leishmanial DHS (Fig. 3), a potent inhibitor of human DHS, N1-guanyl-1,7-diaminoheptane, had little inhibitory effect on either L. donovani proliferation or recombinant DHS34. This finding suggests a topological difference in the spermidine binding sites between the human and the leishmanial enzymes and opens the possibility that the differences between the two enzymes could be exploited for drug development for visceral leishmaniasis. Inhibitors of hypusine biosynthetic enzymes like spermidine analogs (for DHS) and metal ion chelators (for deoxyhypusine hydroxylase) have been shown to exert antiproliferative effects (9, 15), making the pathway a potential target. Further studies are required to elucidate the leishmanial enzyme structure, its interaction with its substrates, and its reaction mechanism. This will pave the way to conclusively establish the hypusine pathway as a chemotherapeutic target for the treatment of leishmaniasis and perhaps even African sleeping sickness and Chagas disease.
Supplementary Material
This work was authored, in whole or in part, by National Institutes of Health staff. This work is supported by a grant from the Council of Scientific and Industrial Research, India (to R. M.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Fig. 1.
- eIF5A
- eukaryotic initiation factor 5A
- DHS
- deoxyhypusine synthase
- ORF
- open reading frame.
REFERENCES
- 1.Sundar S., More D. K., Singh M. K., Singh V. P., Sharma S., Makharia A., Kumar P. C., Murray H. W. (2000) Clin. Infect. Dis. 31, 1104–1107 [DOI] [PubMed] [Google Scholar]
- 2.Murray H. W. (2001) Antimicrob. Agents Chemother. 45, 2185–2197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sundar S. (2001) Trop. Med. Int. Health 6, 849–854 [DOI] [PubMed] [Google Scholar]
- 4.Müller S., Coombs G. H., Walter R. D. (2001) Trends Parasitol. 17, 242–249 [DOI] [PubMed] [Google Scholar]
- 5.Fairlamb A. H., Cerami A. (1992) Annu. Rev. Microbiol. 46, 695–729 [DOI] [PubMed] [Google Scholar]
- 6.Park M. H., Cooper H. L., Folk J. E. (1981) Proc. Natl. Acad. Sci. U.S.A. 78, 2869–2873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Shiba T., Mizote H., Kaneko T., Nakajima T., Kakimoto Y. (1971) Biochim. Biophys. Acta 244, 523–531 [DOI] [PubMed] [Google Scholar]
- 8.Cooper H. L., Park M. H., Folk J. E., Safer B., Braverman R. (1983) Proc. Natl. Acad. Sci. U.S.A. 80, 1854–1857 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Park M. H., Cooper H. L., Folk J. E. (1982) J. Biol. Chem. 257, 7217–7222 [PubMed] [Google Scholar]
- 10.Chen K. Y., Dou Q. P. (1988) FEBS Lett. 229, 325–328 [DOI] [PubMed] [Google Scholar]
- 11.Murphey R. J., Gerner E. W. (1987) J. Biol. Chem. 262, 15033–15036 [PubMed] [Google Scholar]
- 12.Abbruzzese A., Park M. H., Folk J. E. (1986) J. Biol. Chem. 261, 3085–3089 [PubMed] [Google Scholar]
- 13.Byers T. L., Lakanen J. R., Coward J. K., Pegg A. E. (1994) Biochem. J. 303, 363–368 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jakus J., Wolff E. C., Park M. H., Folk J. E. (1993) J. Biol. Chem. 268, 13151–13159 [PubMed] [Google Scholar]
- 15.Park M. H., Wolff E. C., Lee Y. B., Folk J. E. (1994) J. Biol. Chem. 269, 27827–27832 [PubMed] [Google Scholar]
- 16.Nishimura K., Murozumi K., Shirahata A., Park M. H., Kashiwagi K., Igarashi K. (2005) Biochem. J. 385, 779–785 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hanauske-Abel H. M., Park M. H., Hanauske A. R., Popowicz A. M., Lalande M., Folk J. E. (1994) Biochim. Biophys. Acta 1221, 115–124 [DOI] [PubMed] [Google Scholar]
- 18.Hauber I., Bevec D., Heukeshoven J., Krätzer F., Horn F., Choidas A., Harrer T., Hauber J. (2005) J. Clin. Invest. 115, 76–85 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Schnier J., Schwelberger H. G., Smit-McBride Z., Kang H. A., Hershey J. W. (1991) Mol. Cell. Biol. 11, 3105–3114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Park M. H., Joe Y. A., Kang K. R. (1998) J. Biol. Chem. 273, 1677–1683 [DOI] [PubMed] [Google Scholar]
- 21.Joe Y. A., Wolff E. C., Park M. H. (1995) J. Biol. Chem. 270, 22386–22392 [DOI] [PubMed] [Google Scholar]
- 22.Kang K. R., Wolff E. C., Park M. H., Folk J. E., Chung S. I. (1995) J. Biol. Chem. 270, 18408–18412 [DOI] [PubMed] [Google Scholar]
- 23.Liao D. I., Wolff E. C., Park M. H., Davies D. R. (1998) Structure 6, 23–32 [DOI] [PubMed] [Google Scholar]
- 24.Umland T. C., Wolff E. C., Park M. H., Davies D. R. (2004) J. Biol. Chem. 279, 28697–28705 [DOI] [PubMed] [Google Scholar]
- 25.Goyard S., Segawa H., Gordon J., Showalter M., Duncan R., Turco S. J., Beverley S. M. (2003) Mol. Biochem. Parasitol. 130, 31–42 [DOI] [PubMed] [Google Scholar]
- 26.Boitz J. M., Yates P. A., Kline C., Gaur U., Wilson M. E., Ullman B., Roberts S. C. (2009) Infect. Immun. 77, 756–763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Tamura K., Dudley J., Nei M., Kumar S. (2007) Mol. Biol. Evol. 24, 1596–1599 [DOI] [PubMed] [Google Scholar]
- 28.Gowri V. S., Krishnadev O., Swamy C. S., Srinivasan N. (2006) Nucleic Acids Res. 34, D243–D246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Finn R. D., Tate J., Mistry J., Coggill P. C., Sammut S. J., Hotz H. R., Ceric G., Forslund K., Eddy S. R., Sonnhammer E. L., Bateman A. (2008) Nucleic Acids Res. 36, D281–D288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Marchler-Bauer A., Anderson J. B., DeWeese-Scott C., Fedorova N. D., Geer L. Y., He S., Hurwitz D. I., Jackson J. D., Jacobs A. R., Lanczycki C. J., Liebert C. A., Liu C., Madej T., Marchler G. H., Mazumder R., Nikolskaya A. N., Panchenko A. R., Rao B. S., Shoemaker B. A., Simonyan V., Song J. S., Thiessen P. A., Vasudevan S., Wang Y., Yamashita R. A., Yin J. J., Bryant S. H. (2003) Nucleic Acids Res. 31, 383–387 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Berman H. M., Westbrook J., Feng Z., Gilliland G., Bhat T. N., Weissig H., Shindyalov I. N., Bourne P. E. (2000) Nucleic Acids Res. 28, 235–242 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Altschul S. F., Madden T. L., Schäffer A. A., Zhang J., Zhang Z., Miller W., Lipman D. J. (1997) Nucleic Acids Res. 25, 3389–3402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sali A. (1995) Mol. Med. Today 1, 270–277 [DOI] [PubMed] [Google Scholar]
- 34.Wiederstein M., Sippl M. J. (2007) Nucleic Acids Res. 35, W407–W410 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sippl M. J. (1993) Proteins 17, 355–362 [DOI] [PubMed] [Google Scholar]
- 36.Morris A. L., MacArthur M. W., Hutchinson E. G., Thornton J. M. (1992) Proteins 12, 345–364 [DOI] [PubMed] [Google Scholar]
- 37.Laskowski R. A., MacArthur M. W., Moss D. S., Thornton J. M. (1993) J. Appl. Crystallogr. 26, 283–291 [Google Scholar]
- 38.Park M. H., Wolff E. C. (1988) J. Biol. Chem. 263, 15264–15269 [PubMed] [Google Scholar]
- 39.Wolff E. C., Park M. H., Folk J. E. (1990) J. Biol. Chem. 265, 4793–4799 [PubMed] [Google Scholar]
- 40.Roberts S. C., Jiang Y., Jardim A., Carter N. S., Heby O., Ullman B. (2001) Mol. Biochem. Parasitol. 115, 217–226 [DOI] [PubMed] [Google Scholar]
- 41.Robinson K. A., Beverley S. M. (2003) Mol. Biochem. Parasitol. 128, 217–228 [DOI] [PubMed] [Google Scholar]
- 42.Lee C. H., Um P. Y., Park M. H. (2001) Biochem. J. 355, 841–849 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wolff E. C., Folk J. E., Park M. H. (1997) J. Biol. Chem. 272, 15865–15871 [DOI] [PubMed] [Google Scholar]
- 44.Wolff E. C., Wolff J., Park M. H. (2000) J. Biol. Chem. 275, 9170–9177 [DOI] [PubMed] [Google Scholar]
- 45.Joe Y. A., Wolff E. C., Lee Y. B., Park M. H. (1997) J. Biol. Chem. 272, 32679–32685 [DOI] [PubMed] [Google Scholar]
- 46.Wolff E. C., Park M. H. (1999) Yeast 15, 43–50 [DOI] [PubMed] [Google Scholar]
- 47.Brochier C., López-García P., Moreira D. (2004) Gene 330, 169–176 [DOI] [PubMed] [Google Scholar]
- 48.Ober D., Harms R., Witte L., Hartmann T. (2003) J. Biol. Chem. 278, 12805–12812 [DOI] [PubMed] [Google Scholar]
- 49.Willert E. K., Fitzpatrick R., Phillips M. A. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 8275–8280 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wöhl T., Klier H., Ammer H., Lottspeich F., Magdolen V. (1993) Mol. Gen. Genet. 241, 305–311 [DOI] [PubMed] [Google Scholar]
- 51.Sasaki K., Abid M. R., Miyazaki M. (1996) FEBS Lett. 384, 151–154 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.