Abstract
Serine-threonine kinase receptor-associated protein (STRAP) interacts with transforming growth factor β (TGF-β) receptors and inhibits TGF-β signaling. Here, we identify STRAP as an interacting partner of ASK1 (apoptosis signal-regulating kinase 1). The association between ASK1 and STRAP is mediated through the C-terminal domain of ASK1 and the fourth and sixth WD40 repeats of STRAP. Using cysteine-to-serine amino acid substitution mutants of ASK1 (C1005S, C1351S, C1360S, and C1351S/C1360S) and STRAP (C152S, C270S, and C152S/C270S), we demonstrated that Cys1351 and Cys1360 of ASK1 and Cys152 and Cys270 of STRAP are required for ASK1-STRAP binding. ASK1 phosphorylated STRAP at Thr175 and Ser179, suggesting a potential role for STRAP phosphorylation in ASK1 activity regulation. Expression of wild-type STRAP, but not STRAP mutants (C152S/C270S and T175A/S179A), inhibited ASK1-mediated signaling to both JNK and p38 kinases by stabilizing complex formation between ASK1 and its negative regulators, thioredoxin and 14-3-3, or decreasing complex formation between ASK1 and its substrate MKK3. In addition, STRAP suppressed H2O2-mediated apoptosis in a dose-dependent manner by inhibiting ASK1 activity through direct interaction. These results suggest that STRAP can act as a negative regulator of ASK1.
Introduction
ASK1 (apoptosis signal-regulating kinase 1) is a mitogen-activated protein kinase kinase kinase that activates the c-Jun N-terminal kinase (JNK)3/p38 signaling cascade by directly phosphorylating and activating mitogen-activated protein kinase kinases, such as MKK3, -4, -6, and -7, which in turn induce proapoptotic function (1–3). ASK1 activity is stimulated by various stress-related stimuli, including oxidative stress, tumor necrosis factor-α, Fas ligand, endoplasmic reticulum stress, calcium overload, and anti-tumor agents (1–8). ASK1 activity is reportedly regulated through protein-protein interactions; is stimulated by tumor necrosis factor receptor-associated factor, Daxx, JNK/stress-activated protein kinase-associated protein 1/JNK-interacting protein 3, calcineurin, or MPK38 (murine protein serine/threonine kinase 38); and is suppressed by thioredoxin (Trx), glutaredoxin, glutathione S-transferase μ (GSTμ), heat shock protein 72, 14-3-3, Akt/protein kinase B, p21Cip1, or protein serine/threonine phosphatase 5 (1, 4, 8–18). These findings suggest that unidentified ASK1-interacting proteins may function as potential regulators in controlling ASK1 function.
Serine-threonine kinase receptor-associated protein (STRAP) is a transforming growth factor-β (TGF-β) receptor-interacting protein that inhibits TGF-β signaling by stabilizing the association between TGF-β receptors and Smad7 and is localized in both the cytoplasm and nucleus (19, 20). STRAP reportedly acts as an activator of PDK1 (3-phosphoinositide-dependent protein kinase-1) by dissociating 14-3-3, a negative regulator of PDK1, from the PDK1–14-3-3 complex (21). STRAP also contributes to tumor progression by blocking TGF-β-mediated signaling, especially in colon and lung carcinomas, indicating the possible involvement of STRAP in tumorigenesis (20, 22). STRAP can also interact directly with p53 and stimulate p53-mediated signaling, implying that STRAP may function as a tumor suppressor in cells (23).
In the present study, we found that STRAP, an ASK1-interacting partner, may play an important role in the regulation of the ASK1-mediated signaling pathway, in which it acts as a negative regulator of ASK1 activity by stabilizing complex formation between ASK1 and Trx or 14-3-3. We also demonstrated that ASK1 phosphorylates STRAP at Thr175 and Ser179 within a region between the WD4 and WD5 domains, which subsequently leads to ASK1 activity inhibition. Moreover, this association suppressed JNK-mediated transactivation and H2O2-induced apoptosis.
MATERIALS AND METHODS
Antibodies and Plasmids
Anti-FLAG (M2), anti-His, anti-hemagglutinin (HA), anti-ASK1, anti-MPK38, anti-GST, anti-MKK3, anti-ATF2, anti-poly(ADP-ribose) polymerase (PARP), anti-caspase-3, anti-phospho-MKK3/6(Ser189/Ser207), anti- p38, anti-phospho-p38(Thr180/Tyr182), anti-phospho-ATF2 (Thr71), anti-phospho-ASK1(Thr845), anti-phospho-ASK1(Ser83), anti-phospho-ASK1(Ser967), and anti-β-actin antibodies were described previously (11). Anti-PDK1, anti-phospho-AKT(Thr308), anti-AKT, anti-phospho-Bad(Ser136), anti-Bad, and anti-phospho-Ser/Thr antibodies were described previously (21, 24). Anti-phospho-PDK1(Ser241) antibody was from Cell Signaling Technology (Danvers, MA). Wild-type and kinase-dead ASK1, ASK1-N, ASK1-K, ASK1-C, wild-type STRAP and its mutants (WD1–3, WD4–6, WD4, C152S, WD5, WD6, C270S, C152S/C270S, and CT), activator protein 1 (AP-1)-Luc reporter, pSuper vector, GST-tagged MKK6(K82A) and p38, and c-fos were also previously described (11, 25).
Cell Culture and Stable Cell Line Constructions
HEK293, NIH 3T3, HeLa, HCT116, and 293T cells were grown in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum (Invitrogen) as described previously (11). HCT116 cells stably expressing pcDNA3-His-STRAP plasmid (STRAP(OE)) were screened in the presence of 850 μg/ml G418 until control parental HCT116 cells completely died. To prepare HCT116 cells stably expressing a STRAP-specific short hairpin RNA (shRNA) (STRAP(KD)), double-stranded oligonucleotides (forward, 5′-GATCCCCTTACGCATATATGACTTGATTCAAGAGATCAAGTCATATATGCGTAATTTTTGGAAA-3′; reverse, 5′-AGCTTTTCCAAAAATTACGCATATATGACTTGATCTCTTGAATCAAGTCATATATGCGTAAGGG-3′; STRAP sequence underlined) were cloned into the pSuper vector. STRAP(KD) cells were screened in the presence of 1.5 μg/ml puromycin until all control parental HCT116 cells died, as described previously (11). The presence of endogenous and exogenous STRAP proteins was confirmed by immunoblot analysis.
Construction of an Inducible STRAP shRNA Cell Line
An inducible STRAP shRNA HEK293 cell line was generated using the following oligonucleotides as described previously (26). The double-stranded oligonucleotides (forward, 5′-TCGAGGTTACGCATATATGACTTGATTCAAGAGATCAAGTCATATATGCGTAACTTTTTTA-3′; reverse, 5′-AGCTTAAAAAAGTTACGCATATATGACTTGATCTCTTGAATCAAGTCATATATGCGTAACC-3′; STRAP sequence underlined) were cloned into the XhoI/HindIII sites of pSingle-tTS-shRNA vector (Clontech). HEK293 cells were transfected with pSingle-tTS-shRNA harboring STRAP-specific shRNA, pSingle-tTS-shRNA harboring scrambled shRNA as a nonspecific control (24), or pSingle-tTS-shRNA empty vector using WelFect-ExTM Plus (11). Inducible STRAP shRNA stable clones were screened in the presence of 450 μg/ml G418 for 14 days until all control parental HEK293 cells had died. To confirm the knockdown of endogenous STRAP, stable clones were isolated, treated with 1 μg/ml doxycycline (Sigma), a tetracycline analogue, for 72 h, and subjected to Western blotting by anti-STRAP antibody.
Construction of STRAP and ASK1-C Mutants
STRAP mutants were generated by PCR as described previously (11). In brief, wild-type STRAP was used as the template for amplification with either the STRAP forward T3 5′-ATTAACCCTCACTAAAG-3′ or reverse M13 5′-GTAAAACGACGGCCAGT-3′ primer, in conjunction with one of the following mutant primers containing alterations in the nucleotide sequence of STRAP: for T175A/S179A, forward (5′-GATCATGCTACTATGGCAGAAGTGAAAGCTCTAAATTTTAATATG-3′) and reverse (5′-CATATTAAAATTTAGAGCTTTCACTTCTGCCATAGTAGCATGATC-3′); for T201A/S205A, forward (5′-GAGATTTTGGTTATAGCTTATGGACGAGCTATTGCTTTTCATAGT-3′) and reverse (5′-ACTATGAAAAGCAATAGCTCGTCCATAAGCTATAACCAAAATCTC-3′); for S284A/T288A, forward (5′-CTCTATGCCAGTGGTGCAGAAGATGGAGCATTGAGACTATGGCAA-3′) and reverse (5′-TTGCCATAGTCTCAATGCTCCATCTTCTGCACCACTGGCATAGAG-3′). The reaction parameters were 2 min at 95 °C, 1 cycle; 20 s at 95 °C, 40 s at 42–60 °C, 1 min at 72 °C, 18 cycles; 5 min at 72 °C, 1 cycle. Amplified PCR mixtures were subjected to a second round of PCR amplification in the absence of primers using the following parameters: 2 min at 95 °C, 1 cycle; 20 s at 95 °C, 40 s at 42 °C, 1 min at 72 °C, 7 cycles. A third round of PCR amplification was then performed using STRAP forward T3 and reverse M13 primers and the following parameters: 2 min at 95 °C, 1 cycle; 20 s at 95 °C, 40 s at 50 °C, 1 min at 72 °C, 18 cycles. To generate GST-tagged recombinant proteins for ASK1-C mutants (C1351S, C1360S, and C1351S/C1360S), amplified PCR products were digested with EcoRI and XhoI and ligated into pGEX4T-1 (Amersham Biosciences). PCR was performed using FLAG-tagged wild-type ASK1 as the template. The following primers were used: ASK1-C forward primer (5′-GCGAATTCTCAAGCAAAAAGAAAAAG-3′) containing an EcoRI site (underlined), ASK1-C reverse primer (5′-GCCTCGAGTCAAGTCTGTTTGTTTCG-3′) containing an XhoI site (underlined), in conjunction with one of the following mutant primers containing alterations in the nucleotide sequence of ASK1: for C1005S, forward (5′-GCCAAGTCCTCAGGAGAAAGA-3′) and reverse (5′-TCTTTCTCCTGAGGACTTGGC-3′); for C1351S, forward (5′-GACTTAAAATCATTGAGACTA-3′) and reverse (5′-TAGTCTCAATGATTTTAAGTC-3′); for C1360S, forward (5′-GGGATGCTGTCAACACTGTGG-3′), reverse (5′-CCACAGTGTTGACAGCATCCC-3′). To generate the C1351S/C1360S mutant, pGEX4T-1-C1351S was used as the template, and the C1360S forward and reverse primers were used for PCR amplification. GST-tagged wild-type ASK1 was generated by PCR with the following primers: forward primer (5′-GCGTCGACATGAGCACGGAGGCGGACGAGGGC-3′) containing a SalI site (underlined) and reverse primer (5′-GCGCGGCCTCAAGTCTGTTTGTTTCGAAAGTC-3′) containing a NotI site (underlined). The amplified PCR products were digested with SalI and NotI and ligated into pGEX4T-3 (Amersham Biosciences).
In Vivo and in Vitro Binding Assays
In vivo binding assays were performed as previously described using HEK293, NIH 3T3, or HeLa cells transiently transfected with the indicated expression vectors (11, 23–25). To determine the direct interaction between ASK1 and STRAP, a GST pull-down assay was also performed using recombinant ASK1 and STRAP proteins as described (27). Native PAGE for determining the in vitro binding between ASK1 and STRAP was performed using in vitro-translated 35S-labeled STRAP, prepared with the TNT reticulocyte lysate system (Promega) as described previously (23) or autophosphorylated ASK1, prepared in the presence of kinase buffer (20 mm Tris-HCl, pH 7.5, 0.1 mm sodium orthovanadate, 1 mm dithiothreitol, and 20 mm MgCl2) containing 1 μm [γ-32P]ATP (0.2 mCi/ml).
In Vitro Kinase Assays
Assays were performed as described earlier (14). Briefly, the cleared lysates from HEK293 cells transiently transfected with the indicated expression vectors were subjected to immunoprecipitation with appropriate antibodies. After washing the immunoprecipitate three times with lysis buffer and then twice with each kinase buffer (11), the immunoprecipitates were incubated for 30 min at 30 °C in the presence of each kinase buffer containing 5 μg of recombinant GST-tagged substrates. The reaction mixtures were separated by SDS-PAGE, transferred to polyvinylidene difluoride membranes, and detected by autoradiography. Protein concentrations were determined by the Bradford assay.
Assay for JNK Kinase Activity
HEK293 cells were transiently transfected with GST-tagged JNK, along with the indicated expression vectors, and solubilized with lysis buffer (20 mm HEPES, pH 7.9, 10 mm EDTA, 0.1 m KCl, and 0.3 m NaCl). The cleared lysates were precipitated by glutathione-Sepharose beads. The GST precipitates were washed three times with lysis buffer and twice with kinase buffer (20 mm HEPES, pH 7.6, 0.1 mm sodium orthovanadate, 25 mm sodium β-glycerophosphate, 2 mm dithiothreitol, and 20 mm MgCl2) and then subjected to an in vitro kinase assay using recombinant c-Jun as a substrate in the presence of 5 μCi of [γ-32P]ATP, followed by SDS-PAGE and autoradiography.
Small Interfering RNA (siRNA) Experiments
RNA interference experiments were performed as described previously (21). The STRAP-specific siRNA (number 515) of the oligonucleotides (5′-UUACGCAUAUAUGACUUGA-3′) targeting a coding region (aa 124–129) on human STRAP (GenBankTM accession number BC000162) was used for transfection experiments. The ASK1-specific siRNA (5′-GGUAUACAUGAGUGGAAUUTT-3′) was described previously (11).
Luciferase Reporter Assay
293T cells were transfected with the indicated expression and AP-1-Luc reporter vectors by WelFect-ExTM Plus reagent (WelGENE, Daegu, Korea). After 48 h, the cells were harvested, and luciferase activity was measured by a luciferase assay system (Promega). The total DNA concentration was kept constant by supplementation with empty vector DNAs. The values were adjusted relative to the expression levels of a cotransfected β-galactosidase reporter control. For each experiment, at least three independent transfections were performed.
Apoptosis Assay
Apoptosis experiments were performed as described previously using the green fluorescent protein system and terminal deoxynucleotide transferase-mediated dUTP nick end labeling staining in HEK293 cells transfected with the indicated expression vectors (11). Cells exposed to 1 mm H2O2 for 9 h were used as a positive control.
Statistical Analysis
All values are represented as the means ± S.E. A p value relative to control of <0.05, as calculated by Student's t test, was considered statistically significant.
RESULTS
STRAP Is a Functional Linker between the PDK1 and ASK1 Signaling Pathways in Cells
Given that Akt, a downstream target of PDK1, inhibits signaling downstream of ASK1 through direct interaction and phosphorylation (16), it is possible that PDK1 is associated with ASK1-mediated signaling. Therefore, we reasoned that STRAP, which was previously identified as a positive regulator of PDK1 (21), might be involved in the regulation of ASK1-mediated signaling. To determine whether STRAP can be a potential modulator for ASK1 in cells, we initially employed an inducible STRAP-specific shRNA system to investigate the effect of STRAP on ASK1 function. Knockdown of endogenous STRAP significantly increased signaling downstream of ASK1 as determined by immunoblot analysis using anti-phospho-specific antibodies for ASK1 Thr845, MKK3/6, p38, and ATF2 (Fig. 1A, left). As expected, an opposite trend was clearly observed in a stable system for tetracycline-inducible expression of STRAP-specific shRNA when we assessed the effect of STRAP on PDK1-mediated signaling (Fig. 1A, right). These results suggest that STRAP can act as an intermediating molecule linking PDK1- and ASK1-mediated signaling.
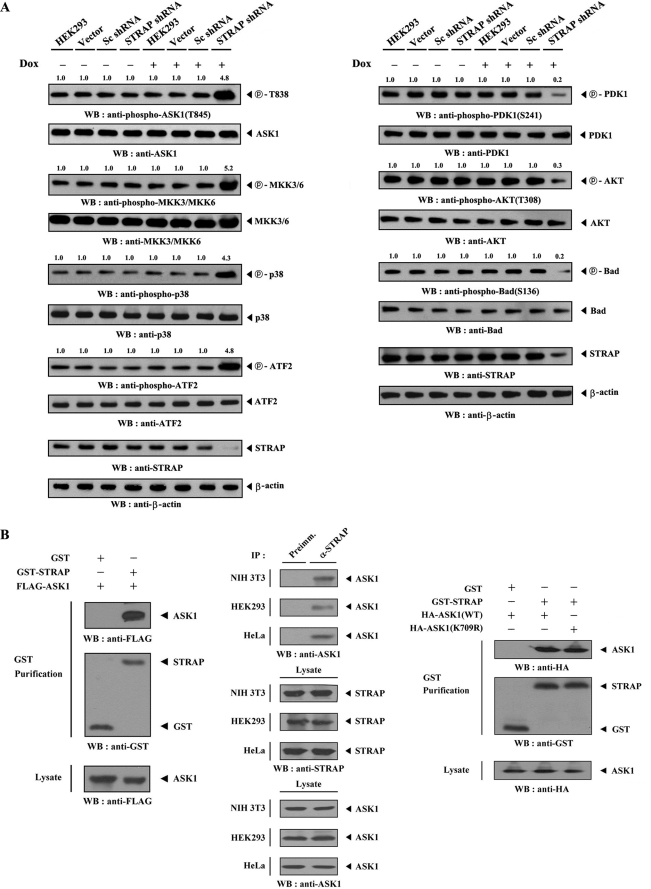
FIGURE 1.
Direct interaction of ASK1 with STRAP in vivo and in vitro. A, regulation of PDK1 and ASK1 activity by STRAP. HEK293 cells harboring stably integrated pSingle-tTS-shRNA empty vector (Vector), pSingle-tTS-shRNA vector containing scrambled shRNA (Sc shRNA), or pSingle-tTS-shRNA vector containing STRAP-specific shRNA (STRAP shRNA) were cultured in the presence or absence of 1 μg/ml doxycycline (Dox) for 72 h to determine the activities of ASK1 (left) or PDK1 (right). Inducible silencing of endogenous STRAP expression by doxycycline was assessed by immunoblotting (WB) using an anti-STRAP antibody. β-Actin was used as a loading control. The relative level of phosphorylation was quantitated by densitometric analyses, and -fold increase relative to control parental cells was calculated. Data are representative of at least three independent experiments. B, physical association of ASK1 with STRAP. GST alone and GST-STRAP were cotransfected with FLAG- or HA-tagged ASK1 (wild-type (WT) or kinase-dead (K709R) form) into HEK293 cells. GST fusion proteins were purified on glutathione-Sepharose beads (GST Purification), and the amounts of complex formation were determined by anti-FLAG or anti-HA antibody immunoblot (left and right). Cell lysates from NIH 3T3, HEK293, or HeLa cells were subjected to immunoprecipitation (IP) using either rabbit preimmune serum (Preimm.) or anti-STRAP antibody (α-STRAP), followed by immunoblot analysis using an anti-ASK1 antibody to determine the complex formation between endogenous ASK1 and STRAP (middle). As a control, the expression levels of STRAP and ASK1 in the total cell lysate were analyzed by immunoblot using anti-STRAP and anti-ASK1 antibodies, respectively.
To test whether STRAP is physically associated with ASK1 in cells, we performed cotransfection experiments using GST-STRAP and FLAG-ASK1. ASK1 was detected in the GST precipitate only when coexpressed with GST-STRAP and not with control GST alone, demonstrating that STRAP physically interacts with ASK1 (Fig. 1B, left). We also analyzed the direct association of purified, recombinant ASK1 with STRAP using a GST pull-down assay. ASK1 was found to be associated with STRAP in vitro (supplemental Fig. S1). To confirm the endogenous interaction between ASK1 and STRAP, we performed coimmunoprecipitation experiments in NIH 3T3, HEK293, and HeLa cells. Immunoprecipitation of endogenous STRAP by an anti-STRAP antibody and then immunoblotting with an anti-ASK1 antibody clearly revealed an interaction between the two endogenous ASK1 and STRAP proteins (Fig. 1B, middle). However, the kinase activity of ASK1 had no effect on the association between ASK1 and STRAP (Fig. 1B, right). These results indicate that ASK1 binds to STRAP in cells.
Dissociation of ASK1-STRAP Complex by ASK1 Stimuli
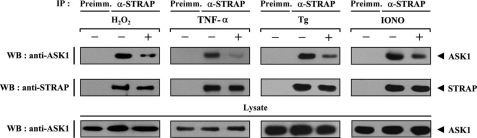
We examined whether H2O2, a stimulator of ASK1, can influence the ASK1-STRAP complex formation in HEK293 and NIH 3T3 cells (Fig. 2 and supplemental Fig. S2). Cells were incubated in media with or without 2 mm H2O2 for 30 min. Endogenous STRAP was immunoprecipitated by anti-STRAP antibody, and the presence of ASK1 was then determined by anti-ASK1 immunoblotting. ASK1-STRAP complex formation was significantly decreased by H2O2 treatment compared with untreated control (Fig. 2, first panel). Similarly, the endogenous interaction of ASK1 with STRAP appeared to be decreased by other ASK1 stimuli, including tumor necrosis factor-α, endoplasmic reticulum stress (thapsigargin), and calcium overload (ionomycin) (Fig. 2, second through fourth panels), suggesting the involvement of STRAP in the ASK1 signaling pathway.
FIGURE 2.
Modulation of the ASK1-STRAP interaction by ASK1 stimuli. HEK293 cell lysates were treated with or without the following stimuli: H2O2 (2 mm, 30 min), tumor necrosis factor-α (500 ng/ml, 30 min), thapsigargin (Tg; 20 μm, 30 min), or ionomycin (IONO; 1 μm, 24 h). They were then immunoprecipitated (IP) with either rabbit preimmune serum (Preimm.) or anti-STRAP antibody (α-STRAP), followed by immunoblotting (WB) with an anti-ASK1 antibody to determine the endogenous association between ASK1 and STRAP.
Mapping of the Interaction Domains between ASK1 and STRAP
We first performed in vivo binding assays to determine which ASK1 domain(s) contributes to STRAP binding. The C-terminal regulatory domain (ASK1-C, aa 941–1375) of ASK1 was found to be responsible for STRAP binding, whereas the N-terminal (ASK1-N, aa 1–648) and kinase (ASK1-K, aa 649–940) domains were unable to bind with STRAP (Fig. 3A, lower left), indicating that ASK1 physically interacts with STRAP through its C-terminal regulatory domain.
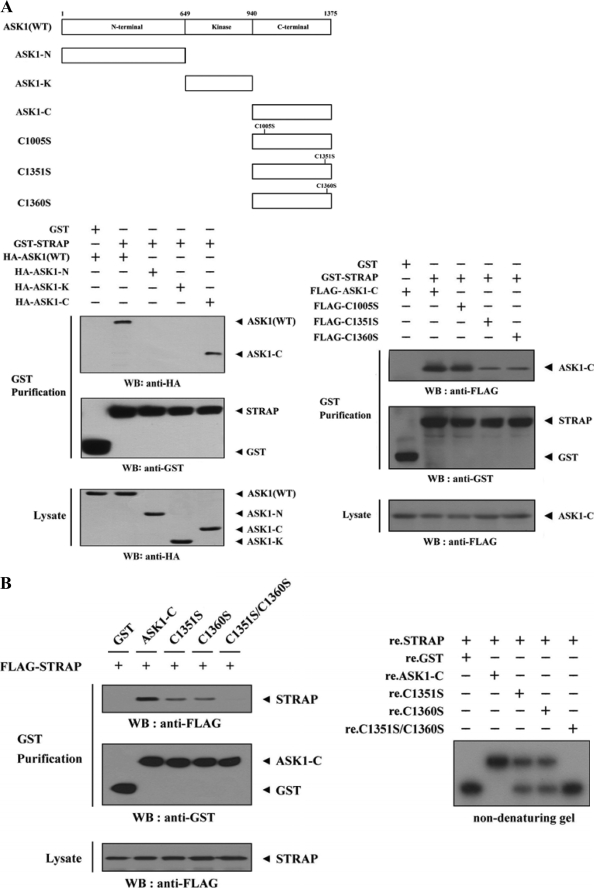
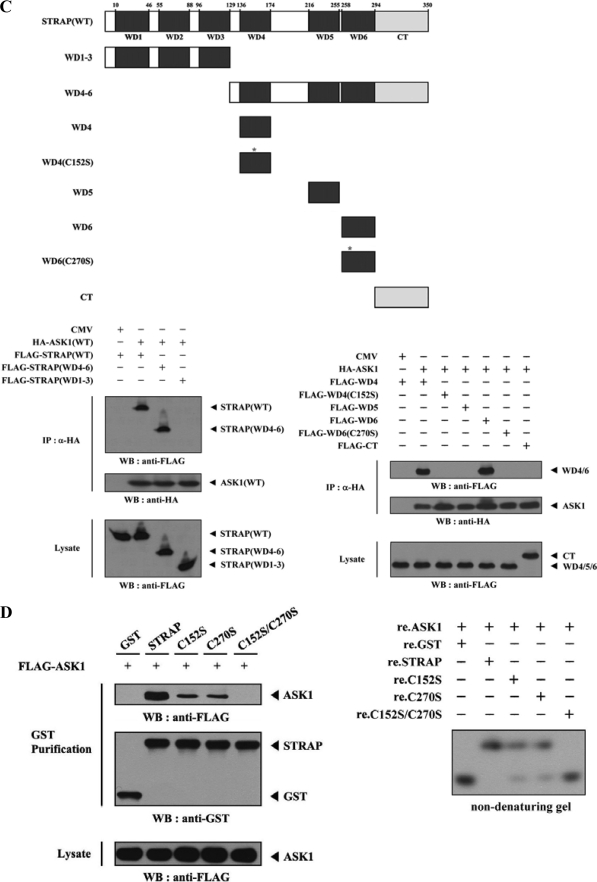
FIGURE 3.
Mapping of the ASK1 and STRAP interaction domains. A and C, schematic structures of wild-type and deletion constructs of ASK1 (A, top) and STRAP (C, top). The numbers indicate the amino acid residues corresponding to the domain boundaries. HEK293 cells transiently transfected with the indicated expression vectors were lysed and then precipitated using glutathione-Sepharose beads (GST Purification) or immunoprecipitated (IP) with an anti-HA antibody (α-HA) and immunoblotted (WB) with anti-HA or anti-FLAG antibody to determine the level of ASK1-STRAP complex formation. B, effect of ASK1-C and its substitution mutants (C1351S, C1360S, or C1351S/C1360S) on ASK1-STRAP complex formation. HEK293 cells, transfected with the expression vectors indicated, were lysed, and the GST precipitates were analyzed for complex formation between ASK1 and STRAP by immunoblot analysis with an anti-FLAG antibody (left). For native PAGE (8%) of the ASK1-STRAP complex, in vitro-translated 35S-labeled STRAP was incubated with unlabeled recombinant wild-type and mutant forms of ASK1-C at room temperature for 1 h (right). D, effect of STRAP and its substitution mutants (C152S, C270S, or C152S/C270S) on ASK1-STRAP complex formation. Complex formation between ASK1 and STRAP was analyzed by immunoblot analysis with an anti-FLAG antibody as described for B (left). For native PAGE (8%) of the ASK1-STRAP complex, purified recombinant ASK1 was autophosphorylated, and 2–3 μg was incubated with unlabeled recombinant GST, as a control, or wild-type and mutant forms of STRAP (5 μg of each) as indicated at room temperature for 1 h (right). Experiments were independently performed at least three times with similar results. re., recombinant.
To further define the region of ASK1 that interacts with STRAP, we generated four cysteine-to-serine amino acid substitution mutants of ASK1-C (C1005S, C1351S, C1360S, and C1351S/C1360S). This was done because the physical association of STRAP with its interacting partners, such as Nm23-H1 and p53, is mediated by cysteine residues in these proteins (23, 25). STRAP interacted with ASK1-C and its mutant C1005S. STRAP also interacted weakly with the ASK1-C mutants C1351S and C1360S (Fig. 3A, lower right), but this interaction was not observed in the presence of the C1351S/C1360S mutant (Fig. 3B, left), indicating that both Cys1351 and Cys1360 of ASK1 play a critical role in its association with STRAP.
We also confirmed the effect of the cysteine residues of ASK1-C on the association between ASK1 and STRAP using non-denaturing PAGE. Consistently, a shift in the mobility of 35S-labeled STRAP was evident upon incubation in the presence of ASK1-C and was weakly detectable in the presence of the ASK1-C mutants, C1351S and C1360S (Fig. 3B, right, second lane versus the third and fourth lanes). Additionally, the band shift was not observed in the presence of the C1351S/C1360S mutant (Fig. 3B, right, second lane versus fifth lane), again supporting a physical association between ASK1 and STRAP through cysteine residues.
To identify the STRAP region required for ASK1 binding, we first performed in vivo binding assays with HEK293 cells transfected with two deletion constructs of STRAP, STRAP(WD1–3) and STRAP(WD4–6). ASK1 interacted with STRAP(WD4–6), which contained the three C-terminal WD40 repeats (aa 129–350), but not with STRAP(WD1–3), which harbored the three N-terminal WD40 repeats (aa 1–129) (Fig. 3C, lower left). These results indicate that the interaction of STRAP with ASK1 is mediated via the C-terminal WD40 repeats of STRAP.
To more precisely define the ASK1-binding motif of STRAP, we examined the binding properties of six additional STRAP deletion constructs: STRAP-WD4, -WD4(C152S), -WD5, -WD6, -WD6(C270S), and -CT. The binding of ASK1 to STRAP(WD4) or STRAP(WD6) was readily detectable, whereas ASK1 did not interact with other STRAP mutants tested, including WD4(C152S), WD5, WD6(C270S), and CT (Fig. 3C, lower right). This was also confirmed by cotransfection experiments (Fig. 3D, left) and by non-denaturing PAGE analysis with recombinant ASK1 and wild-type and mutant forms of STRAP proteins (Fig. 3D, right). Together, these results clearly demonstrate that the physical association between ASK1 and STRAP requires the participation of cysteine residues present in the respective proteins: Cys1351 and Cys1360 of ASK1 and Cys152 and Cys270 of STRAP.
STRAP Inhibits ASK1 Kinase Activity by Direct Interaction
To assess the effects of an ASK1-STRAP interaction on ASK1 activity, we first performed in vitro kinase assays to determine ASK1 kinase activity using HEK293 cells transfected with ASK1 alone or cotransfected with STRAP. ASK1 kinase activity markedly decreased when ASK1 was coexpressed with STRAP (Fig. 4A, first lane versus second lane), indicating that STRAP is apparently involved in the negative regulation of ASK1 activity. In a separate experiment with recombinant ASK1 and STRAP proteins, we also confirmed that STRAP inhibited ASK1 kinase activity in a dose-dependent manner (supplemental Fig. S3).
FIGURE 4.
STRAP inhibits ASK1 kinase activity. A and B, HEK293 cells were transiently transfected with HA-ASK1 in the presence or absence of the FLAG-tagged wild-type (WT) or mutant form (C152S/C270S) of STRAP. Cell lysates were then subjected to immunoprecipitation (IP) with an anti-HA antibody, and the resulting immunoprecipitates were subjected to an in vitro kinase assay using MKK6(K82A) as the substrate to determine ASK1 kinase activity. The circled P followed by MKK6(K82A) indicates phosphorylated MKK6(K82A). The same blot was stripped and reprobed with an anti-HA antibody to determine the expression level of immunoprecipitated ASK1 (third panels), and the expression level of FLAG-STRAP in total cell lysates was examined by immunoblot analysis (WB) using an anti-FLAG antibody (fourth panels). The presence of equivalent amounts of substrate was verified by immunoblotting with an anti-GST antibody (bottom panels).
To examine whether the direct binding property of STRAP plays a key role in the negative regulation of ASK1 kinase activity, we analyzed the effect of the C152S/C270S mutant, which was unable to bind with ASK1 (see Fig. 3D), on ASK1 kinase activity using an in vitro kinase assay. Coexpression of C152S/C270S had no effect on ASK1 kinase activity compared with control expressing ASK1 alone (Fig. 4B, first lane versus third lane), suggesting a critical role for the direct interaction between ASK1 and STRAP in the regulation of ASK1 kinase activity. As a control, the levels of immunoprecipitated ASK1 proteins were analyzed, and similar expression levels were found for the ASK1 construct (Fig. 4, A and B, third panels), indicating that the observed differences in phosphorylated MKK6(K82A) were not due to differences in ASK1 expression levels of HA immunoprecipitates. The STRAP-mediated inhibition of ASK1 kinase activity was also confirmed by immunoblot analysis using an anti-phospho-specific antibody for ASK1 Thr845 (Fig. 4, A and B, second panels). These results suggest that STRAP may be a negative regulator of ASK1.
ASK1 Phosphorylates STRAP at Thr175 and Ser179
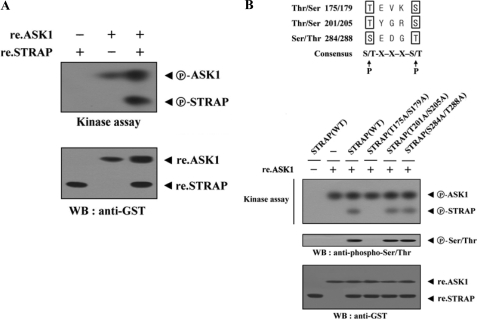
Because STRAP interacts with ASK1 (Fig. 1B), we next determined whether STRAP can be a direct substrate for ASK1. Recombinant ASK1 proteins were incubated with [γ-32P]ATP to allow phosphorylation of the recombinant STRAP proteins, which were used as substrates for the ASK1 kinase assay. STRAP phosphorylation was clearly observed in the presence of ASK1 (Fig. 5A, lane 3), indicating STRAP as a substrate for ASK1.
FIGURE 5.
Identification of ASK1 phosphorylation sites on STRAP. A, purified recombinant ASK1 was assayed for its kinase activity in the presence of kinase buffer containing ∼3–4 μg of recombinant STRAP as a substrate. B, recombinant ASK1 was analyzed for its kinase activity by an in vitro kinase assay using recombinant wild-type (WT) STRAP and one of its substitution mutants, T175A/S179A, T201A/S205A, or S284A/T288A, as substrate. The phosphorylation of STRAP was also confirmed by immunoblotting (WB) with an anti-phospho-Ser/Thr antibody (second panel). re., recombinant.
To more precisely define the ASK1 phosphorylation sites of STRAP, we searched consensus ASK1 phosphorylation sites ((S/T)XXX(S/T)) in STRAP and performed in vitro kinase assays using three amino acid substitution STRAP mutants (T175A/S179A, T201A/S205A, and S284A/T288A). The T175A/S179A mutant completely abolished ASK1-mediated phosphorylation (Fig. 5B, third lane versus fourth lane). However, ASK1-mediated phosphorylation occurred in the presence of other STRAP substitution mutants (T201A/S205A and S284A/T288A) (Fig. 5B, third lane versus fifth and sixth lanes), indicating that Thr175 and Ser179 of STRAP are potential phosphorylation sites for ASK1. These results suggest that ASK1 directly phosphorylates STRAP at Thr175 and Ser179 through a physical interaction.
ASK1-mediated STRAP Phosphorylation Is Important for ASK1-p38 Signaling Inhibition
We next examined whether STRAP affects signaling downstream of ASK1. HEK293 cells transfected with an empty vector, wild-type STRAP, C152S/C270S mutant, or T175A/S179A mutant were treated with H2O2 and subsequently immunoprecipitated using anti-ASK1, anti-MKK3, and anti-p38 antibodies. Endogenous ASK1, MKK3, and p38 kinase activities were evaluated using in vitro kinase assays. As expected, H2O2 sufficiently stimulated the kinase activities of ASK1, MKK3, and p38 in HEK293 cells expressing an empty vector (Vector in Fig. 6). However, wild-type STRAP, but not the C152S/C270S or T175A/S179A mutants, significantly decreased these kinase activities (Fig. 6A). Consistent with this, overexpression of STRAP (STRAP(OE)) showed a similar trend in the modulation of ASK1, MKK3, and p38 kinase activities (Fig. 6B, upper panels), suggesting that STRAP inhibits ASK1-mediated signaling to p38 kinase.
FIGURE 6.
STRAP inhibits ASK1 downstream signaling. HEK293 cells transiently transfected with an empty vector (Vector), wild-type STRAP (WT), or mutant forms of STRAP (C152S/C270S and T175A/S179A) (A); parental HCT116 cells (B and C); and HCT116 cells stably overexpressing STRAP (STRAP(OE)) or HCT116 cells stably expressing STRAP-specific shRNA (STRAP(KD)) (B and C) were incubated with or without 2 mm H2O2 for 30 min. The endogenous activities of ASK1, MKK3, or p38 were determined by in vitro kinase assays or immunoblot analysis (WB) using the immunoprecipitates (IP) indicated or total cell lysates. The presence of equal amounts of each substrate in these assays was confirmed by immunoblotting with an anti-GST antibody. Overexpression or knockdown level of endogenous STRAP was determined by anti-STRAP immunoblotting. The relative level of kinase activity was quantified by densitometric analyses. The -fold increase relative to untreated samples in control HEK293 cells containing an empty vector alone or parental HCT116 cells was calculated.
We confirmed these results in HCT116 cells stably expressing shRNA targeting STRAP (STRAP(KD)). Knockdown of endogenous STRAP significantly increased ASK1, MKK3, and p38 kinase activities compared with the control parent HCT116 cells, regardless of H2O2 treatment (Fig. 6C, upper panels). These effects were also confirmed by immunoblot analysis using anti-phospho-specific antibodies for ASK1 Thr845, MKK3/6, p38, and ATF2 (Fig. 6, B and C, lower panels). Together, these results suggest that STRAP phosphorylation at Thr175 and Ser179 through direct interaction with ASK1 is critical for the negative regulation of ASK1-mediated signaling.
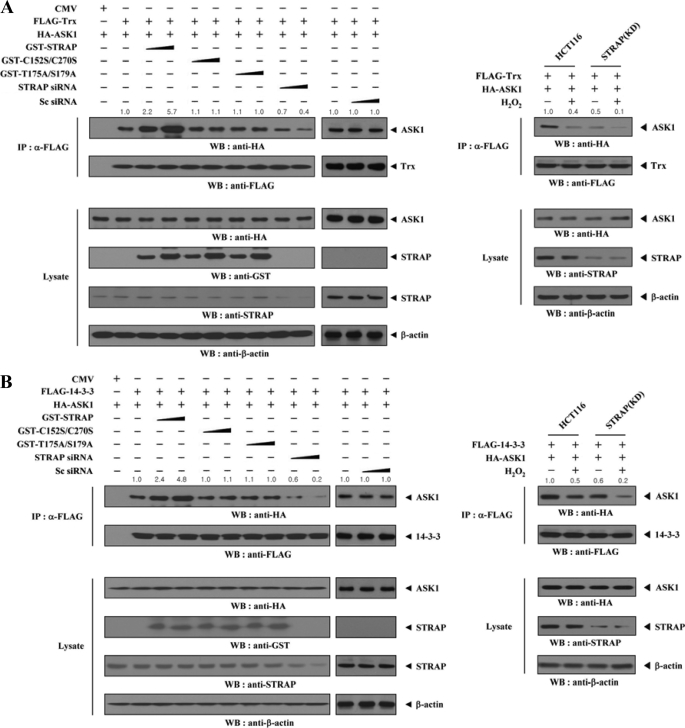
STRAP Stabilizes Complex Formation between ASK1 and Its Negative Regulators, Trx and 14-3-3, and Interferes with the Association between ASK1 and MKK3
To investigate the mechanism of STRAP-mediated inhibition of ASK1 activity, we examined whether STRAP contributes to the interaction between ASK1 and its negative regulator Trx. HEK293 cells transfected with vectors expressing FLAG-Trx and HA-ASK1 in the presence or absence of wild-type STRAP, C152S/C270S mutant, T175A/S179A mutant, scrambled siRNA, or STRAP-specific siRNA were subjected to immunoprecipitation using an anti-FLAG antibody, followed by immunoblot analysis with an anti-HA antibody. There was a dose-dependent increase in complex formation between ASK1 and Trx in cells expressing wild-type STRAP compared with expression in the absence of STRAP (Fig. 7A, left). However, the C152S/C270S and T175A/S179A mutants had no effect on the association between ASK1 and Trx, consistent with the above result (Fig. 6A). This indicates that the direct interaction and phosphorylation between ASK1 and STRAP is important for STRAP-mediated regulation of the association between ASK1 and Trx. We also observed a similar trend showing the importance of the direct interaction and phosphorylation between ASK1 and STRAP in STRAP-mediated regulation of complex formation between ASK1 and 14-3-3 (Fig. 7B, left). These results were verified in STRAP-knocked down HCT116 cells (Fig. 7, A and B, right panels).
FIGURE 7.
STRAP-mediated modulation of the association between ASK1 and its regulators (Trx and 14-3-3) or substrate (MKK3) and the homo-oligomerization of ASK1. A and B, effect of STRAP on the association between ASK1 and its negative regulators Trx (A) and 14-3-3 (B). HEK293 cells were transfected with the indicated combinations of expression vectors and STRAP-specific siRNA. Cell lysates were then subjected to immunoprecipitation (IP) with an anti-FLAG antibody, and the resulting immunoprecipitates were analyzed by immunoblot analysis (WB) with an anti-HA antibody to determine the association of ASK1 with Trx or 14-3-3 (left panels). The effect of STRAP knockdown on the association between ASK1 with Trx or 14-3-3 was determined by immunoblotting with the indicated antibodies using HCT116 cells (STRAP(KD)) stably expressing STRAP-specific shRNA (right panels). C and D, effect of STRAP on the association between ASK1 and its substrate MKK3 (C) and the homo-oligomerization of ASK1 (D). Complex formation between ASK1 and MKK3 or ASK1 was determined by immunoblotting with the antibodies indicated, as described in A and B. The relative level of complex formation was quantified by densitometric analyses. The -fold increase relative to control HEK293 cells expressing ASK1 and its regulators (Trx, 14-3-3, MKK3, or ASK1) alone or untreated HCT116 cells expressing ASK1 and its regulators was calculated. Sc siRNA, scrambled siRNA. CMV, cytomegalovirus.
Because ASK1 was previously shown to form active complexes with its substrate MKK3 (11), and because the homo-oligomerization of ASK1 is apparently involved in its kinase activity (4, 28), we next investigated the effects of STRAP on MKK3 binding to ASK1 and ASK1 homo-oligomerization. STRAP inhibited ASK1 activity by inhibiting ASK1-MKK3 complex formation and ASK1 homo-oligomerization (Fig. 7, C and D). In contrast, the C152S/C270S and T175A/S179A mutants had no effect on these associations.
STRAP Suppresses ASK1-dependent AP-1 Transcriptional Activity
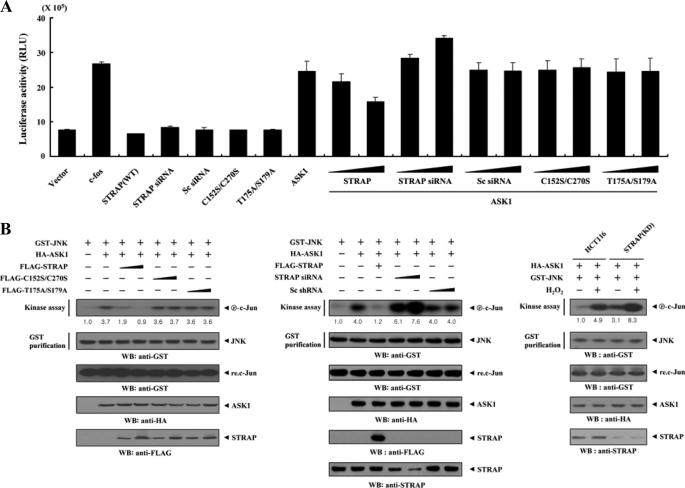
To investigate whether STRAP influences downstream gene transcriptional events, we examined the transcription factor AP-1 activated by JNK and p38 kinases using a luciferase reporter assay. Wild-type STRAP significantly decreased ASK1-dependent AP-1 transcriptional activity in a dose-dependent manner, whereas C152S/C270S and T175A/S179A mutants had no effect (Fig. 8A), again suggesting a critical role of direct interaction and phosphorylation between ASK1 and STRAP in the regulation of ASK1-mediated transactivation.
FIGURE 8.
Suppression of JNK-mediated transcription by STRAP. A, 293T cells were transfected with 0.2 μg of AP-1 luciferase plasmid and increasing amounts of wild-type (WT) and mutant forms (C152S/C270S and T175A/S179A) of STRAP (6 and 9 μg), STRAP-specific siRNA (50 and 200 nm), and scrambled siRNA (50 and 200 nm), as indicated, in the presence or absence of c-fos (0.6 μg). Data shown are means ± S.E. of three independent experiments. B, inhibition of JNK activity by STRAP. HEK293 cells were cotransfected using GST-JNK (1.5 μg) and HA-ASK1 (2 μg) in the presence or absence of increasing amounts of wild-type and mutant forms (C152S/C270S and T175A/S179A) of STRAP (6 and 9 μg), STRAP-specific siRNA (100 and 200 nm), or scrambled siRNA (100 and 200 nm). An in vitro kinase assay for JNK activity was performed as described, and the amounts of precipitated JNK and expression levels of ASK1 and STRAP in total cell lysates were determined by immunoblot analysis (WB) using the indicated antibodies (left and middle). STRAP(KD) cells transfected with GST-JNK and HA-ASK1 were also precipitated using glutathione-Sepharose beads (GST purification), and the precipitates were subjected to an in vitro kinase assay using c-Jun as a substrate to determine JNK activity (right). The relative level of JNK activity was quantified by densitometric analyses. The -fold increase relative to control HEK293 cells expressing JNK alone or untreated HCT116 cells expressing JNK and ASK1 was calculated. Sc siRNA, scrambled siRNA.
We also confirmed the effect of STRAP on ASK1-dependent AP-1 transcriptional activity using a STRAP knockdown system. Transfection of STRAP-specific siRNA significantly increased AP-1 transcriptional activity in a dose-dependent manner (Fig. 8A, eighth lane versus the eleventh and twelfth lanes).
To examine whether the STRAP-mediated suppression of AP-1 transcriptional activity was associated with JNK inactivation, we performed in vitro kinase assays using c-Jun as a substrate. As expected, ASK1-mediated JNK activation was markedly decreased by wild-type STRAP in a dose-dependent manner, whereas the C152S/C270S and T175A/S179A mutants had no effect on JNK activity (Fig. 8B, left). We confirmed this result in STRAP-knocked down cells. The introduction of STRAP-specific siRNA significantly increased ASK1-mediated JNK activation in a dose-dependent manner (Fig. 8B, middle). Consistently, a similar result was also observed in HCT116 cells stably expressing an shRNA targeting STRAP (STRAP(KD)) (Fig. 8B, right). Together, these results support the conclusion that the direct phosphorylation of STRAP at Thr175 and Ser179 through physical interaction with ASK1 plays a key role in STRAP-mediated inhibition of ASK1-mediated signaling to both JNK and p38 kinases.
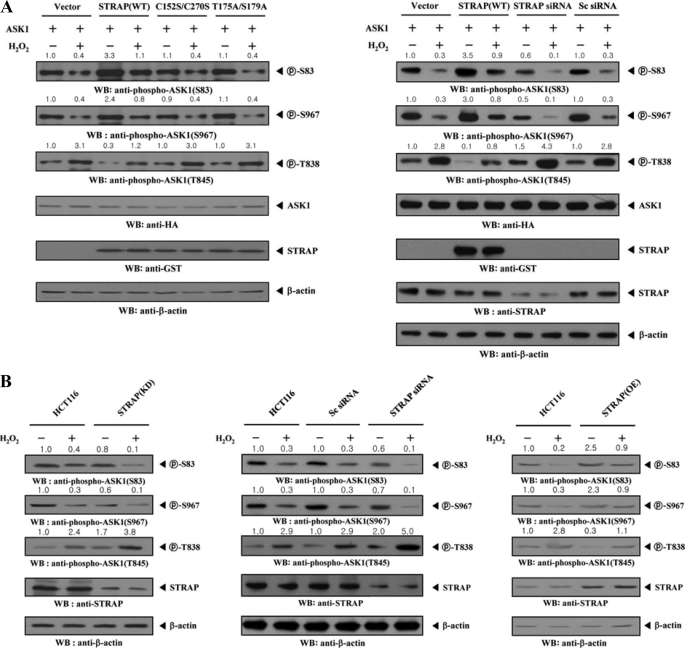
STRAP-mediated Inhibition of ASK1 Activity Is Accompanied by ASK1 Phosphorylation at Multiple Sites
ASK1 is reportedly positively or negatively regulated, depending on its phosphorylation status at Ser83, Thr838 (corresponding to Thr845 in mice), or Ser967 (29). To determine if the inhibitory effect of STRAP on ASK1-p38/JNK signaling is closely associated with ASK1 phosphorylation at these sites, we performed immunoblot analysis with anti-phospho-specific antibodies for ASK1 Ser83 and Ser967 (for inactive forms of ASK1), as well as Thr845 (for an active form of ASK1), in HEK293 and NIH 3T3 cells (Fig. 9A and data not shown). Expression of wild-type STRAP markedly alleviated the H2O2-induced reduction of ASK1 phosphorylation at Ser83 and Ser967, whereas expression of C152S/C270S and T175A/S179A mutants had no effect (Fig. 9A, top and second panels). As expected, H2O2-induced stimulation of ASK1 phosphorylation at Thr838 was significantly decreased by wild-type STRAP expression compared with the expression of an empty vector (Fig. 9A, third panels). Consistently, expression of C152S/C270S and T175A/S179A mutants had no effect on H2O2-induced stimulation of ASK1 phosphorylation at Thr838.
FIGURE 9.
STRAP-mediated inhibition of ASK1 activity is dependent on ASK1 phosphorylation. A, HEK293 cells were transiently transfected with empty vector (Vector), wild-type (WT), or mutant forms (C152S/C270S and T175A/S179A) of STRAP, scrambled siRNA (200 nm), or STRAP-specific siRNA (200 nm), in the presence of ASK1. Cells were then treated with or without 2 mm H2O2 for 30 min. ASK1 phosphorylation was determined by immunoblot analysis (WB) using anti-phospho-ASK1(Ser83), anti-phospho-ASK1(Thr845), and anti-phospho-ASK1(Ser967) antibodies. B, parental HCT116 cells (HCT116), HCT116 cells stably expressing STRAP-specific shRNA (STRAP(KD)), HCT116 cells transiently transfected with STRAP-specific siRNA (STRAP siRNA), HCT116 cells transiently transfected with scrambled siRNA (Sc siRNA), or HCT116 cells stably overexpressing STRAP (STRAP(OE)) were incubated in the presence or absence of 2 mm H2O2 for 30 min. Cell lysates were examined for ASK1 phosphorylation (Ser83, Thr838, and Ser967) by immunoblot analysis using the antibodies indicated. The knockdown and overexpression levels of STRAP were determined by anti-STRAP immunoblotting (fourth panels). The relative level of phosphorylation was quantified by densitometric analyses, and the -fold increase relative to untreated samples in parental HCT116 cells was calculated.
Endogenous STRAP knockdown using STRAP-specific siRNA clearly showed an opposite trend in the modulation of Ser83, Ser967, and Thr838 phosphorylations (Fig. 9A, right). To further verify this in a physiological context, we determined the effect of STRAP on H2O2-induced activation of ASK1 using HCT116 cells stably expressing an shRNA targeting STRAP (STRAP(KD)). Similar to the above results, in parental HCT116 cells, H2O2 treatment induced ASK1 activity by decreasing Ser83 and Ser967 phosphorylation and increasing Thr838 phosphorylation (Fig. 9B, HCT116). However, in STRAP(KD) cells, Ser83 and Ser967 phosphorylation was decreased and Thr838 phosphorylation was increased compared with control parental HCT116 cells (Fig. 9B, left). Consistently, a similar result was obtained in the STRAP knockdown system via STRAP-specific siRNA (Fig. 9B, middle). In contrast, overexpression of endogenous STRAP (STRAP(OE)) increased Ser83 and Ser967 phosphorylation and decreased Thr838 phosphorylation (Fig. 9B, right). These results suggest that STRAP-mediated inhibition of ASK1 function apparently involves modulation of the phosphorylation levels of ASK1 at Ser83, Thr838, and Ser967.
STRAP Suppresses H2O2-mediated Apoptosis through Caspase-3 Inactivation
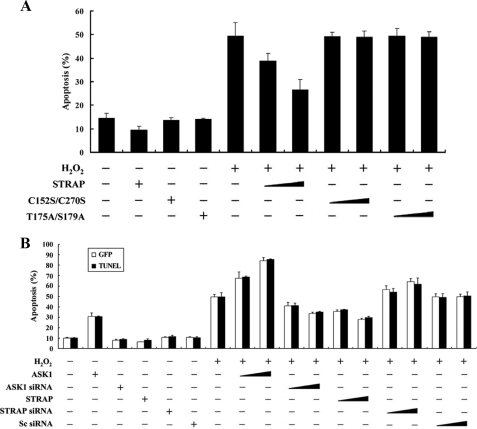
Because ASK1 activation induces apoptotic cell death under various conditions (2, 3), we next examined whether STRAP could influence H2O2-mediated cell death in HEK293 cells, as determined by a green fluorescent protein system and terminal deoxynucleotide transferase-mediated dUTP nick end labeling staining (11). Expression of wild-type STRAP significantly decreased H2O2-induced cell death in a dose-dependent manner (Fig. 10A, fifth lane versus sixth and seventh lanes), indicating that STRAP is involved in the regulation of H2O2-mediated cell death. We also determined whether the ASK1-mediated phosphorylation of STRAP was necessary for regulation of H2O2-mediated cell death, because wild-type STRAP, but not C152S/C270S or T175A/S179A mutants, inhibited ASK1 activity (Figs. 4, 6, and 8). No difference in the H2O2-mediated cell death was found in the presence of C152S/C270S and T175A/S179A mutants compared with the control treated with H2O2 in the absence of STRAP (Fig. 10A, fifth lane versus the eighth through eleventh lanes), indicating an important role for the direct interaction and phosphorylation between ASK1 and STRAP in the regulation of H2O2-mediated cell death.
FIGURE 10.
STRAP suppresses H2O2-induced apoptosis. A and B, HEK293 cells were transiently transfected with the indicated combinations of expression vectors encoding wild-type (WT) and mutant forms (C152S/C270S and T175A/S179A) of STRAP (1 and 3 μg), wild-type ASK1 (1 and 3 μg), scrambled siRNA (100 and 200 nm), and ASK1- and STRAP-specific siRNAs (100 and 200 nm) in the presence or absence of H2O2. Apoptotic cell death was determined using the green fluorescent protein expression system (GFP) or terminal deoxynucleotide transferase-mediated dUTP nick end labeling (TUNEL) (11). Cells exposed to 1 mm H2O2 for 9 h were used as a positive control. Data shown represent the means ± S.E. of three independent experiments. C, inhibition of caspase-3 and PARP degradation by STRAP. HEK293 cells were transiently transfected with increasing amounts of wild-type (WT) and mutant forms (C152S/C270S and T175A/S179A) of STRAP (0.8 and 1.6 μg) or wild-type ASK1 (1 μg) in the presence or absence of H2O2. Cell lysates were then analyzed by immunoblot analysis (WB) with an anti-caspase-3 or anti-PARP antibody to determine the degradation of caspase-3 and PARP, and the expression levels of ASK1 and STRAP were determined by anti-HA and anti-GST immunoblotting, respectively (left panels). HCT116 cells stably expressing STRAP-specific shRNA (STRAP(KD)) were incubated with 2 mm H2O2 for 30 min. Cells were lysed and then subjected to immunoblot analysis using the indicated antibodies (right panels). The data are representative of at least three independent experiments.
We also performed knockdown experiments of endogenous STRAP using STRAP-specific siRNA. Transfection of HEK293 cells with siRNA duplexes targeting STRAP considerably increased H2O2-mediated apoptosis proportional to the amount of siRNA used in the transfection (Fig. 10B, seventh lane versus the fourteenth and fifteenth lanes). These data suggest that STRAP contributes to the negative regulation of H2O2-mediated apoptosis by inhibiting ASK1 activity through a physical interaction.
ASK1 has previously been implicated in caspase-3 and PARP activation (30–32). Therefore, we assessed the effects of STRAP on H2O2-induced caspase-3 activity and PARP cleavage in HEK293 cells transfected with the indicated expression vectors (Fig. 10C, left panels) and HCT116 cells stably expressing an shRNA targeting STRAP (STRAP(KD)) (Fig. 10C, right panels) by immunoblotting with anti-caspase-3 and anti-PARP antibodies, respectively. STRAP inhibited H2O2-mediated apoptosis by suppressing in vivo caspase-3 activity.
DISCUSSION
In the present study, we investigated the regulatory mechanism of ASK1 activity and found that STRAP, an interacting partner of TGF-β receptors (33), physically interacted with ASK1 and inhibited ASK1 function, suggesting that STRAP functions as a negative regulator of ASK1-mediated signaling.
To gain insight into the mechanism(s) by which STRAP inhibits ASK1 activity, we examined whether the direct interaction between ASK1 and STRAP is important for STRAP-mediated inhibition of ASK1 activity using in vitro kinase assays and immunoblot analyses. A C152S/C270S mutant of STRAP, which is unable to bind with ASK1, had no effect on the ASK1-mediated signaling to both JNK and p38 kinases (Figs. 6, 8, and 10) or ASK1 phosphorylation (Fig. 9), whereas a marked inhibition of ASK1 function was detected in the presence of wild-type STRAP. These results strongly indicate that STRAP inhibits ASK1 function through its direct interaction with ASK1.
The next step was to investigate whether ASK1-mediated phosphorylation of STRAP affects the activity of ASK1 and its downstream targets, such as MKK3/6 and p38 (11). We first determined whether STRAP could act as a direct substrate for ASK1, because STRAP physically interacts with ASK1 (Fig. 1B). ASK1 phosphorylated STRAP at Thr175 and Ser179 through direct interaction (Fig. 5B), suggesting that Thr175 and Ser179 of STRAP, potential phosphorylation sites for ASK1, might play important roles in the negative regulation of ASK1 activity.
To examine this hypothesis, we performed in vitro kinase assays to assess the endogenous kinase activities of ASK1, MKK3, and p38 in the presence of the T175A/S179A mutant, which is defective in ASK1-mediated phosphorylation (see Fig. 5B), together with wild-type STRAP and the C152S/C270S mutant. Wild-type STRAP, but not STRAP mutants (C152S/C270S and T175A/S179A), significantly decreased the endogenous kinase activities for ASK1, MKK3, and p38 compared with an empty vector-expressing control (Fig. 6A). This result strongly supports a critical role for direct interaction and phosphorylation between ASK1 and STRAP in the regulation of ASK1 function. Thus, it is possible that ASK1-mediated phosphorylation of STRAP at Thr175 and Ser179 may affect the physical association between ASK1 and STRAP. The T175A/S179A mutant, like the C152S/C270S mutant, failed to complex with ASK1 (supplemental Fig. S4, left), suggesting that STRAP phosphorylation at Thr175 and Ser179 is required for its interaction with ASK1. To further investigate whether STRAP phosphorylation plays an important role in ASK1 activity regulation, we also analyzed the effect of the T175A/S179A mutant on ASK1 kinase activity using an in vitro kinase assay. Coexpression of the T175A/S179A mutant, like the C152S/C270S mutant, had no effect on ASK1 kinase activity (supplemental Fig. S4, right). These results, together with other data (Figs. 6, 8, and 10), clearly indicate that STRAP phosphorylation at Thr175 and Ser179 is critical for its binding to ASK1 and ASK1 activity regulation.
However, the question remains of whether STRAP is directly or indirectly responsible for inhibiting ASK1 activity. As shown in Fig. 7, our data show that the interaction and phosphorylation between ASK1 and STRAP have an effect on the association between ASK1 and its known negative regulators, Trx and 14-3-3 (3, 9, 15), suggesting that STRAP indirectly induces the inactivation of ASK1. However, we cannot rule out the possibility that STRAP is directly coupled to the inactivation of ASK1 because STRAP was shown to inhibit directly the ASK1 activity when the kinase activity of ASK1 was determined by an in vitro kinase assay using recombinant ASK1 and STRAP proteins (supplemental Fig. S3).
The physical association of STRAP with TGF-β receptors raised the possibility that STRAP is a substrate of the serine/threonine kinase TGF-β receptor (19). STRAP phosphorylation at the C-terminal 57 amino acids was mediated by type 1 TGF-β receptor, a serine/threonine kinase. However, the observation of STRAP phosphorylation in epithelial cells deficient in type 1 TGF-β receptor strongly suggested the presence of other protein kinases, probably associated with STRAP, for STRAP phosphorylation. We used an in vitro kinase assay to investigate whether ASK1 can act as an alternative kinase responsible for STRAP phosphorylation and observed that ASK1 phosphorylated STRAP at Thr175 and Ser179 (Fig. 5B). This result suggests that ASK1 is a putative kinase for STRAP phosphorylation. In addition, the serine/threonine kinase TGF-β receptor and ASK1 are not alike in terms of STRAP phosphorylation because ASK1, unlike the serine/threonine kinase TGF-β receptor responsible for the C-terminal region of STRAP, phosphorylates the Thr175 and Ser179 residues in the middle region of STRAP.
ASK1 is reportedly regulated by phosphorylation and dephosphorylation at four different phosphorylation sites (29). ASK1 activation is accompanied by phosphorylation of Thr845 and dephosphorylation of Ser83, Ser967, and Ser1034. STRAP actually decreased Thr838 (corresponding to Thr845 in mice) phosphorylation but increased Ser83 and Ser967 phosphorylation, an effect that would inactivate ASK1 (Fig. 9). These data suggest that STRAP mediates the control of ASK1 phosphorylation, which is correlated with ASK1 activation and inactivation. Therefore, we postulated that STRAP probably plays an important role in the regulation of identified (Akt for Ser83; ASK2/MPK38 for Thr845) and unidentified kinases, which are responsible for ASK1 phosphorylation at Ser967 and Ser1034, probably by modulating kinase activities through physical interactions. This is because our previous studies demonstrated that STRAP could stimulate PDK1 kinase activity through its direct binding with PDK1 (21).
ASK1 is uniquely regulated by oxidative stress through direct interaction with Trx (3, 9), indicating the importance of the redox potential in defining the overall effect of stimuli that result in ASK1 activation. We previously showed that STRAP physically interacts with Nm23-H1 (25) and p53 (23) through cysteine residues and regulates their activities. Therefore, using in vivo binding assays, we investigated whether the physical interaction between ASK1 and STRAP could be mediated by the cysteine residues present in each of these two proteins and whether it was dependent on the redox state. STRAP, unlike Trx, which binds to the N-terminal region of ASK1, was shown to redox-dependently interact with ASK1 through the C-terminal region of ASK1, thereby inhibiting ASK1 activity (Fig. 3 and supplemental Fig. S5). This finding suggests that STRAP may act as an intermediate of reactive oxygen species-mediated signaling.
In conclusion, our present results demonstrate that the TGF-β receptor-interacting protein STRAP (33) inhibits the activation of ASK1-mediated signaling to both JNK and p38 kinases through its phosphorylation at Thr175 and Ser179 induced by direct binding with ASK1. Moreover, the potential role of STRAP as a novel inhibitor of ASK1-mediated signaling may help to clarify the regulatory mechanism(s) of ASK1-mediated signaling in cells. Further investigation of a possible functional link between ASK1 and TGF-β signaling pathways is necessary, because STRAP acts as a regulator in the modulation of the TGF-β signaling pathway (19).
Supplementary Material
This work was supported by Korea Science and Engineering Foundation Grant R0A-2007-000-20006-0 and in part by Chungbuk National University Grant 2008.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S5.
- JNK
- c-Jun N-terminal kinase
- STRAP
- serine-threonine kinase receptor-associated protein
- TGF-β
- transforming growth factor-β
- siRNA
- small interfering RNA
- shRNA
- short hairpin RNA
- Trx
- thioredoxin
- GST
- glutathione S-transferase
- PARP
- poly(ADP-ribose) polymerase
- HA
- hemagglutinin
- AP-1
- activator protein 1
- aa
- amino acids.
REFERENCES
- 1.Chang H. Y., Nishitoh H., Yang X., Ichijo H., Baltimore D. (1998) Science 281, 1860–1863 [DOI] [PubMed] [Google Scholar]
- 2.Ichijo H., Nishida E., Irie K., ten Dijke P., Saitoh M., Moriguchi T., Takagi M., Matsumoto K., Miyazono K., Gotoh Y. (1997) Science 275, 90–94 [DOI] [PubMed] [Google Scholar]
- 3.Saitoh M., Nishitoh H., Fujii M., Takeda K., Tobiume K., Sawada Y., Kawabata M., Miyazono K., Ichijo H. (1998) EMBO J. 17, 2596–2606 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gotoh Y., Cooper J. A. (1998) J. Biol. Chem. 273, 17477–17482 [DOI] [PubMed] [Google Scholar]
- 5.Chen Z., Seimiya H., Naito M., Mashima T., Kizaki A., Dan S., Imaizumi M., Ichijo H., Miyazono K., Tsuruo T. (1999) Oncogene 18, 173–180 [DOI] [PubMed] [Google Scholar]
- 6.Wang T. H., Wang H. S., Ichijo H., Giannakakou P., Foster J. S., Fojo T., Wimalasena J. (1998) J. Biol. Chem. 273, 4928–4936 [DOI] [PubMed] [Google Scholar]
- 7.Nishitoh H., Matsuzawa A., Tobiume K., Saegusa K., Takeda K., Inoue K., Hori S., Kakizuka A., Ichijo H. (2002) Genes Dev. 16, 1345–1355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu Q., Wilkins B. J., Lee Y. J., Ichijo H., Molkentin J. D. (2006) Mol. Cell. Biol. 26, 3785–3797 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Liu H., Nishitoh H., Ichijo H., Kyriakis J. M. (2000) Mol. Cell. Biol. 20, 2198–2208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Matsuura H., Nishitoh H., Takeda K., Matsuzawa A., Amagasa T., Ito M., Yoshioka K., Ichijo H. (2002) J. Biol. Chem. 277, 40703–40709 [DOI] [PubMed] [Google Scholar]
- 11.Jung H., Seong H. A., Ha H. (2008) J. Biol. Chem. 283, 34541–34553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Song J. J., Rhee J. G., Suntharalingam M., Walsh S. A., Spitz D. R., Lee Y. J. (2002) J. Biol. Chem. 277, 46566–46575 [DOI] [PubMed] [Google Scholar]
- 13.Cho S. G., Lee Y. H., Park H. S., Ryoo K., Kang K. W., Park J., Eom S. J., Kim M. J., Chang T. S., Choi S. Y., Shim J., Kim Y., Dong M. S., Lee M. J., Kim S. G., Ichijo H., Choi E. J. (2001) J. Biol. Chem. 276, 12749–12755 [DOI] [PubMed] [Google Scholar]
- 14.Park H. S., Cho S. G., Kim C. K., Hwang H. S., Noh K. T., Kim M. S., Huh S. H., Kim M. J., Ryoo K., Kim E. K., Kang W. J., Lee J. S., Seo J. S., Ko Y. G., Kim S., Choi E. J. (2002) Mol. Cell. Biol. 22, 7721–7730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhang L., Chen J., Fu H. (1999) Proc. Natl. Acad. Sci. U.S.A. 96, 8511–8515 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kim A. H., Khursigara G., Sun X., Franke T. F., Chao M. V. (2001) Mol. Cell. Biol. 21, 893–901 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhan J., Easton J. B., Huang S., Mishra A., Xiao L., Lacy E. R., Kriwacki R. W., Houghton P. J. (2007) Mol. Cell. Biol. 27, 3530–3541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Morita K., Saitoh M., Tobiume K., Matsuura H., Enomoto S., Nishitoh H., Ichijo H. (2001) EMBO J. 20, 6028–6036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Datta P. K., Moses H. L. (2000) Mol. Cell. Biol. 20, 3157–3167 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Halder S. K., Anumanthan G., Maddula R., Mann J., Chytil A., Gonzalez A. L., Washington M. K., Moses H. L., Beauchamp R. D., Datta P. K. (2006) Cancer Res. 66, 6156–6166 [DOI] [PubMed] [Google Scholar]
- 21.Seong H. A., Jung H., Choi H. S., Kim K. T., Ha H. (2005) J. Biol. Chem. 280, 42897–42908 [DOI] [PubMed] [Google Scholar]
- 22.Anumanthan G., Halder S. K., Friedman D. B., Datta P. K. (2006) Cancer Res. 66, 10824–10832 [DOI] [PubMed] [Google Scholar]
- 23.Jung H., Seong H. A., Ha H. (2007) J. Biol. Chem. 282, 35293–35307 [DOI] [PubMed] [Google Scholar]
- 24.Seong H. A., Jung H., Kim K. T., Ha H. (2007) J. Biol. Chem. 282, 12272–12289 [DOI] [PubMed] [Google Scholar]
- 25.Seong H. A., Jung H., Ha H. (2007) J. Biol. Chem. 282, 12075–12096 [DOI] [PubMed] [Google Scholar]
- 26.Jung H., Seong H. A., Ha H. (2008) J. Biol. Chem. 283, 20383–20396 [DOI] [PubMed] [Google Scholar]
- 27.Garg S., Alam M. S., Bajpai R., Kishan K. R., Agrawal P. (2009) BMC Biochem. 10, 1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tobiume K., Saitoh M., Ichijo H. (2002) J. Cell. Physiol. 191, 95–104 [DOI] [PubMed] [Google Scholar]
- 29.Fujii K., Goldman E. H., Park H. R., Zhang L., Chen J., Fu H. (2004) Oncogene 23, 5099–5104 [DOI] [PubMed] [Google Scholar]
- 30.Hatai T., Matsuzawa A., Inoshita S., Mochida Y., Kuroda T., Sakamaki K., Kuida K., Yonehara S., Ichijo H., Takeda K. (2000) J. Biol. Chem. 275, 26576–26581 [DOI] [PubMed] [Google Scholar]
- 31.Patel T., Gores G. J., Kaufmann S. H. (1996) FASEB J. 10, 587–597 [DOI] [PubMed] [Google Scholar]
- 32.Chinnaiyan A. M., Orth K., O'Rourke K., Duan H., Poirier G. G., Dixit V. M. (1996) J. Biol. Chem. 271, 4573–4576 [DOI] [PubMed] [Google Scholar]
- 33.Datta P. K., Chytil A., Gorska A. E., Moses H. L. (1998) J. Biol. Chem. 273, 34671–34674 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.