Abstract
Arabidopsis nonexpresser of pathogenesis-related (PR) genes (NPR1) is the sole positive regulator that has been shown to be essential for the induction of systemic acquired resistance. In npr1 mutant plants, salicylic acid (SA)–mediated PR gene expression and pathogen resistance are abolished completely. NPR1 has been shown to interact with three closely related TGA transcription factors—TGA2, TGA5, and TGA6—in yeast two-hybrid assays. To elucidate the biological functions of these three TGA transcription factors, we analyzed single and combined deletion knockout mutants of TGA2, TGA5, and TGA6 for SA-induced PR gene expression and pathogen resistance. Induction of PR gene expression and pathogen resistance by the SA analog 2,6-dichloroisonicotinic acid (INA) was blocked in tga6-1 tga2-1 tga5-1 but not in tga6-1 or tga2-1 tga5-1 plants. Loss of INA-induced resistance to Peronospora parasitica Noco2 cosegregated with the tga6-1 mutation in progeny of multiple lines that were heterozygous for tga6-1 and homozygous for tga2-1 tga5-1 and could be complemented by genomic clones of wild-type TGA2 or TGA5, indicating that TGA2, TGA5, and TGA6 encode redundant and essential functions in the positive regulation of systemic acquired resistance. In addition, tga6-1 tga2-1 tga5-1 plants had reduced tolerance to high levels of SA and accumulated higher basal levels of PR-1 under noninducing conditions, suggesting that these TGA factors also are important for SA tolerance and the negative regulation of the basal expression of PR-1.
INTRODUCTION
Systemic acquired resistance (SAR) is a general defense response that develops in the distal, uninfected parts of plants after local infection by an avirulent pathogen (Ryals et al., 1996). SAR is effective against a broad spectrum of microbial pathogens. One important signal molecule in SAR is salicylic acid (SA), which is required and sufficient for the induction of pathogenesis-related (PR) genes and pathogen resistance during SAR. When attacked by pathogens, plants synthesize and accumulate higher levels of SA in both infected and systemic tissues (Malamy et al., 1990; Métraux et al., 1990; Rasmussen et al., 1991). Application of SA or SA analogs, such as 2,6-dichloroisonicotinic acid (INA) and benzothiadiazole, also induces both PR gene expression and pathogen resistance in plants (White, 1979; Métraux et al., 1991; Görlach et al., 1996). In addition, blocking SA accumulation by expressing the bacterial SA-degrading enzyme salicylate hydroxylase prevents the induction of PR genes and SAR in transgenic plants (Gaffney et al., 1993). The importance of SA in plant defense also is confirmed by analyzing mutants that are deficient in SA synthesis. In Arabidopsis, mutations in enhanced disease susceptibility5 (EDS5) and SA induction-deficient2 (SID2) block pathogen-induced SA synthesis and render the plants more susceptible to pathogen infection (Rogers and Ausubel, 1997; Nawrath and Métraux, 1999). EDS5 encodes a member of the MATE transporter family and is likely to be involved in transporting one of the precursors for the biosynthesis of SA (Nawrath et al., 2002). SID2 encodes an isochorismate synthase, suggesting that SA accumulated during pathogen infection is derived from chorismate (Wildermuth et al., 2001).
Several different genetic screens were conducted to identify regulatory genes downstream of SA. All 12 SA-nonresponsive mutants identified contain mutations in nonexpresser of PR genes (NPR1) (also known as NIM1 and SAI1) (Cao et al., 1994; Delaney et al., 1995; Glazebrook et al., 1996; Shah et al., 1997). In npr1 plants, induction of PR genes and pathogen resistance by SA are abolished. NPR1 encodes a protein with no obvious biochemical functions except the presence of two protein–protein interaction domains, a BTB/POZ domain at the N-terminal end and an ankyrin-repeat domain in the central region (Cao et al., 1997; Ryals et al., 1997; Aravind and Koonin, 1999). The presence of protein–protein interaction domains in NPR1 suggests that NPR1 may regulate SA signaling through an association with other proteins.
Several groups have performed yeast two-hybrid screens using NPR1 as bait and found multiple TGA transcription factors that can interact with NPR1 (Zhang et al., 1999; Després et al., 2000; Niggeweg et al., 2000a; Zhou et al., 2000; Chern et al., 2001, Kim and Delaney, 2002). We showed previously that TGA2 (also known as AHBP-1b), TGA5 (also known as OBF5), and TGA6 interact with NPR1, with TGA2 and TGA6 exhibiting strong affinity and TGA5 showing weaker affinity to NPR1 in the yeast two-hybrid assay (Zhang et al., 1999). The interaction between NPR1 and TGA2 was demonstrated in vivo (Subramaniam et al., 2001; Fan and Dong, 2002), and the involvement of TGA transcription factors in SA signaling is supported further by the presence of a TGA binding site in the PR-1 promoter that is essential for SA-induced PR-1 expression (Lebel et al., 1998). However, genetic evidence for the roles of these TGA transcription factors in SAR is lacking. Here, we report that TGA2, TGA5, and TGA6 encode redundant functions and are essential for the induction of SAR.
RESULTS
Isolation of tga6-1 and Construction of the tga6-1 tga2-1 tga5-1 Triple Mutant
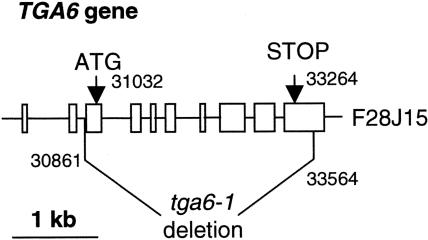
Using primers flanking TGA6 (At3g12250), we screened an Arabidopsis deletion mutant population by PCR and identified a deletion mutant for TGA6 named tga6-1. Sequence analysis of the deletion mutation revealed that a fragment of ∼2.7 kb between nucleotides 30,861 and 33,564 on BAC clone F28J15 was deleted (Figure 1). Sequence comparison between TGA6 cDNA and the genomic sequence revealed that the cDNA of TGA6 is transcribed from the region between nucleotides 30,249 and 33,651 of F28J15 and that the coding sequence is located between nucleotides 31,032 and 33,264. Thus, the deletion in tga6-1 occurred within TGA6 and removed the entire coding region. TGA2 (At5g06950) and TGA5 (At5g06960) are linked directly, and the distance between these two genes is <2 kb. We reported previously the identification of a mutant with both TGA2 and TGA5 deleted (Li et al., 2001). This mutant is named tga2-1 tga5-1. To obtain a triple mutant for TGA2, TGA5, and TGA6, we crossed tga6-1 with tga2-1 tga5-1 and screened the resulting F2 population for homozygous mutants at both loci. Two independent F2 lines that are homozygous at both loci were obtained, and the progeny of these two lines were used for subsequent phenotypic analysis. F2 lines heterozygous at the tga6-1 locus and homozygous at the tga2-1 tga5-1 locus also were obtained, and they were used later for cosegregation analysis.
Figure 1.
Structure of the Deletion in tga6-1.
The fast neutron–induced deletion in TGA6 spans the region between position 30,861 in the second intron and position 33,564 in the last exon. The numbering is based on the Arabidopsis BAC clone F28J15 that contains TGA6.
The TGA Triple Knockout Mutant Is More Sensitive to the Toxicity of SA
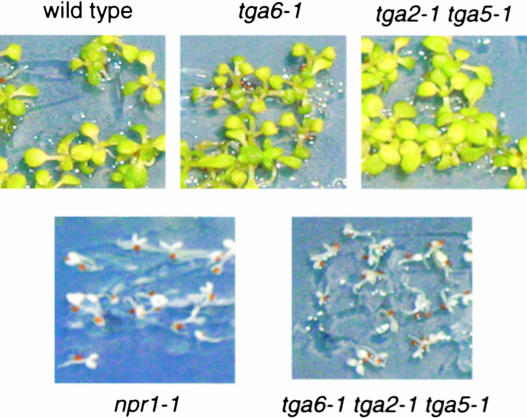
Previously, npr1-1 plants were shown to be more sensitive to high concentrations of SA (Cao et al., 1997). To determine whether the tga mutants also have altered responses to SA, the mutant seeds were plated on MS medium (Murashige and Skoog, 1962) containing 0.2 mM SA. Similar to npr1-1, tga6-1 tga2-1 tga5-1 plants were highly sensitive to SA. As shown in Figure 2, the growth of tga6-1 tga2-1 tga5-1 plants was arrested at the cotyledon stage and the seedlings were bleached, whereas tga6-1 and tga2-1 tga5-1 grew like the wild type under the same conditions. Thus, these TGA transcription factors play roles similar to that of NPR1 in the regulation of tolerance to SA.
Figure 2.
Tolerance of Wild-Type, npr1-1, tga6-1, tga2-1 tga5-1, and tga6-1 tga2-1 tga5-1 Plants to 0.2 mM SA.
Seeds were plated on MS medium containing 0.2 mM SA, and the photographs were taken 14 days after germination. This experiment was repeated twice with similar results.
Induction of PR-1 by INA Is Blocked in the TGA Triple Knockout Mutant
In Arabidopsis, PR-1 is highly induced during SAR (Uknes et al., 1992). The expression of PR-1 also can be induced by exogenous application of SA or the SA analog INA. Mutations in NPR1 completely block the induction of PR-1 by SA or INA. To determine whether TGA2, TGA5, and TGA6 encode functions similar to those of NPR1, we analyzed the expression levels of PR-1 in the TGA knockout mutants under inducing and noninducing conditions. Because SA is highly toxic to the TGA triple mutant, we used INA as the inducing agent.
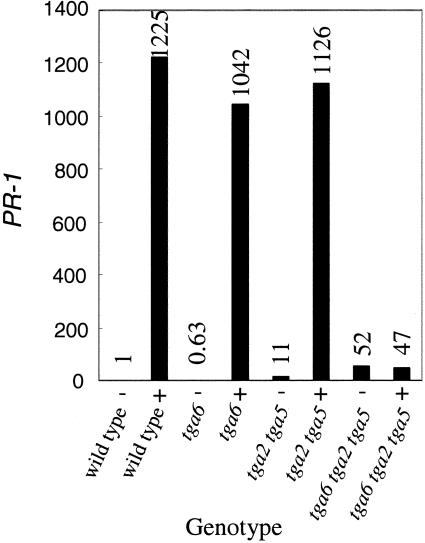
In wild-type plants, PR-1 was induced strongly by INA treatment. This induction was not affected by either tga6-1 or the tga2-1 tga5-1, because both tga6-1 and tga2-1 tga5-1 plants accumulated levels of PR-1 similar to that in wild-type plants after INA induction (Figure 3). By contrast, PR-1 was no longer induced by INA in the tga6-1 tga2-1 tga5-1 triple knockout mutant, suggesting that SAR is compromised when all three TGA factors are mutated.
Figure 3.
PR-1 Expression in Wild-Type, tga6-1, tga2-1 tga5-1, and tga6-1 tga2-1 tga5-1 Plants in Response to Treatment with INA.
Total RNA was extracted from 20-day-old seedlings grown on MS medium in the presence (+) or absence (−) of 50 μM INA. Relative levels of PR-1 were determined by real-time PCR using SYBR Green I chemistry. Values were normalized to the expression of ACTIN1 and are expressed relative to the level in wild-type plants. This experiment was repeated twice with similar results.
Interestingly, under noninducing conditions, the basal level of PR-1 was affected in the TGA knockout mutants. In tga6-1 tga2-1 tga5-1 and tga2-1 tga5-1 plants, the expression level of PR-1 was ∼50-fold and 10-fold higher than that of wild-type plants, respectively (Figure 3). No significant change in the basal level of PR-1 was observed in tga6-1 plants.
SAR Is Abolished in TGA Triple Knockout Mutants
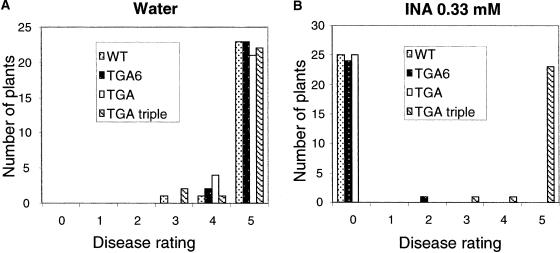
To determine whether INA-induced pathogen resistance was affected in the TGA knockout mutants, 2-week-old wild-type and mutant plants were treated with 0.33 mM INA and inoculated with the virulent oomycete pathogen Peronospora parasitica Noco2 after 3 days. The INA-treated wild-type plants were immune to P. parasitica Noco2 infection, because no conidiophores were observed on the plants 7 days after inoculation. This induced immunity was not affected by the tga6-1 or tga2-1 tga5-1 mutations, because both mutants were as resistant as the wild type (Figures 4A and 4B). By contrast, the INA-induced resistance was abolished completely in tga6-1 tga2-1 tga5-1, indicating that SAR is compromised in the TGA triple mutant.
Figure 4.
Growth of P. parasitica Noco2 on Wild-Type, tga6-1, tga2-1 tga5-1, and tga6-1 tga2-1 tga5-1 Plants.
Two-week-old seedlings were pretreated with water (A) or 0.33 mM INA (B) and sprayed with P. parasitica Noco2 spores (5 × 103 spores/mL) 3 days later. Infection was scored at 7 days after inoculation by counting the number of conidiophores per infected leaf. A total of 25 plants were scored for each treatment. Disease rating scores are as follows: 0, no conidiophores on the plants; 1, no more than 5 conidiophores per infected leaf; 2, 6 to 20 conidiophores on a few of the infected leaves; 3, 6 to 20 conidiophores on most of the infected leaves; 4, 5 or more conidiophores on all infected leaves; 5, 20 or more conidiophores on all infected leaves. This experiment was repeated once with similar results. WT, wild type.
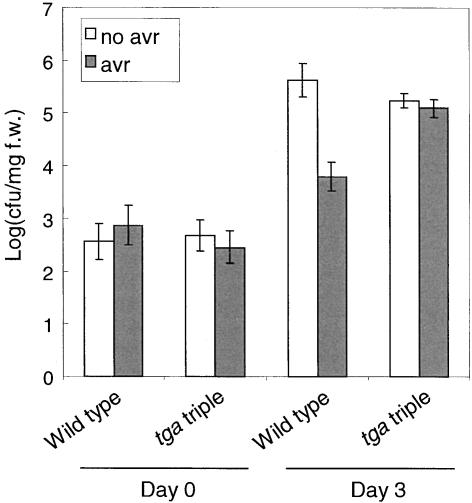
We further tested whether systemic resistance can be induced by an avirulent pathogen in tga6-1 tga2-1 tga5-1 plants. As shown in Figure 5, Pseudomonas syringae pv tomato (P.s.t.) DC3000 carrying avrRpt2 induced systemic resistance to Pseudomonas syringae pv maculicola (P.s.m.) ES4326 in wild-type plants but not in tga6-1 tga2-1 tga5-1 plants, further suggesting that SAR is compromised in the TGA triple mutant.
Figure 5.
Growth of P.s.m. ES4326 in Wild-Type and tga6-1 tga2-1 tga5-1 Plants Preinoculated with P.s.t. DC3000 avrRpt2.
Two leaves from each plant were infiltrated with P.s.t. DC3000 avrRpt2 (OD600 = 0.02) in 10 mM MgCl2 solution or with the buffer alone 3 days before P.s.m. ES4326 infection (OD600 = 0.001). Leaf discs within the inoculated area were taken at 0 and 3 days after P.s.m. ES4326 infection, and the bacterial titers were measured. Error bars represent 95% confidence limits of log-transformed data. Four samples were taken for each time point. This experiment was repeated once with similar results. avr, P.s.t. DC3000 avrRpt2; cfu, colony-forming units; f.w., fresh weight; tga triple, tga6-1 tga2-1 tga5-1.
Compromised SAR in tga6-1 tga2-1 tga5-1 Cosegregates with tga6-1
As shown in Figure 1, the deletion in tga6-1 affected only TGA6. To determine whether the loss of INA-induced resistance in the TGA triple mutant cosegregated with the tga6-1 deletion, we analyzed four independent F2 lines that were heterozygous for the tga6-1 deletion but homozygous for the tga2-1 tga5-1 deletion. The F3 plants of these lines were analyzed for susceptibility to P. parasitica Noco2 after INA treatment. As shown in Table 1, in all four lines tested, approximately one-fourth of the progeny lost INA-induced resistance to P. parasitica Noco2. To determine whether the susceptible plants were homozygous at the tga6-1 locus, DNA from each individual susceptible plant was analyzed by PCR using primers within the TGA6 deletion. All susceptible plants were found to be homozygous for the tga6-1 deletion, indicating that the loss of INA-induced resistance cosegregated with the tga6-1 deletion.
Table 1.
Segregation of the Progeny of TGA6/tga6-1 tga2-1/tga2-1 tga5-1/tga5-1 Lines
| Plant Line | No. of Resistant Plants | No. of Susceptible Plants |
|---|---|---|
| Line 1 | 36 | 11 |
| Line 2 | 30 | 9 |
| Line 3 | 31 | 12 |
| Line 4 | 40 | 12 |
Both TGA2 and TGA5 Can Restore INA-Induced PR Gene Expression and Pathogen Resistance in tga6-1 tga2-1 tga5-1 Plants
To determine whether TGA2 and TGA5 can complement the mutant phenotypes of tga6-1 tga2-1 tga5-1, we transformed tga6-1 tga2-1 tga5-1 plants with genomic clones containing either TGA2 or TGA5. Three independent lines for each construct were used for phenotypic analysis. Because similar results were obtained from all lines, only results from one line for each construct is shown in Figure 6. INA-induced PR-1 expression was restored in the TGA triple mutant plants transformed with genomic DNA clones of TGA2 or TGA5 (Figure 6A). Treatment with INA also resulted in strong resistance to P. parasitica Noco2 in these transgenic plants (Figures 6C and 6D), indicating that SAR was restored in the triple mutant expressing either of the two TGA transcription factors. Furthermore, tolerance to SA was restored in tga6-1 tga2-1 tga5-1 plants transformed with the genomic clone of TGA2 or TGA5 (Figure 6B).
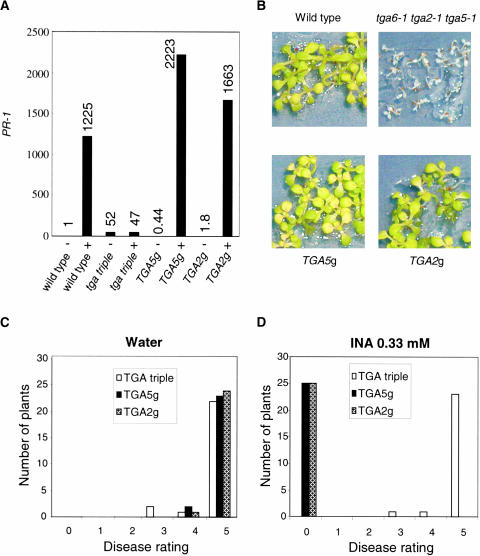
Figure 6.
Complementation of tga6-1 tga2-1 tga5-1 by Genomic Clones of TGA2 or TGA5.
tga triple, tga6-1 tga2-1 tga5-1; TGA2g, a representative tga6-1 tga2-1 tga5-1 line transformed with the genomic clone of TGA2; TGA5g, a representative tga6-1 tga2-1 tga5-1 line transformed with the genomic clone of TGA5.
(A) Complementation of tga6-1 tga2-1 tga5-1 in PR-1 expression. The relative expression levels of PR-1 in tga6-1 tga2-1 tga5-1, TGA2g, and TGA5g were determined as described for Figure 3. +, grown on MS medium + 50 mM INA; −, MS medium alone.
(B) Complementation of tga6-1 tga2-1 tga5-1 in SA tolerance. T3 seeds of tga6-1 tga2-1 tga5-1 homozygous for the TGA2 or TGA5 transgene were plated on MS medium with 0.2 mM SA along with seeds of tga6-1 tga2-1 tga5-1. The photographs were taken 14 days after gemination.
(C) and (D) Complementation of tga6-1 tga2-1 tga5-1 in response to pathogen infection. Using protocols described in Figure 4, water-treated (C) or INA-treated (D) plants were inoculated with P. parasitica Noco2. Infection was scored at 7 days after inoculation by counting the number of conidiophores per infected leaf. A total of 25 plants were scored for each treatment. Each experiment was repeated at least once with similar results.
DISCUSSION
Although TGA transcription factors have been suggested to be important regulators of SA signaling, it was unclear whether they are essential for the establishment of SAR. In previous studies, transgenic plants overexpressing dominant-negative forms of TGA2 or a tobacco TGA2 homolog were used to probe the functions of TGA transcription factors in Arabidopsis and tobacco (Niggeweg et al., 2000b; Pontier et al., 2001; Fan and Dong, 2002). Phenotypes of the transgenic plants differed dramatically depending on the specific dominant-negative mutant used. In one study, tobacco plants overexpressing a dominant-negative form of TGA2 exhibited higher levels of PR gene induction by pathogen challenge and an enhanced SAR, leading to the conclusion that TGA factors are not essential for PR gene activation or SAR (Pontier et al., 2001). In another study, INA-induced PR-1 expression was reduced in plants accumulating high levels of a truncated form of TGA2, although it is unclear whether INA-induced pathogen resistance was affected in these plants (Fan and Dong, 2002). Similarly, a reduction of SA-induced PR gene expression also was observed in transgenic plants overexpressing a dominant-negative mutant of tobacco TGA2.2 (Niggeweg et al., 2000b).
Because dominant-negative mutants most likely affect multiple independent TGA factors to various degrees, it is difficult to determine the functions of individual TGA factors using this approach. Instead, we created knockout plants of TGA2, TGA5, and TGA6 and assayed the single and combined mutants for altered regulation of SA signaling. We found that both PR gene expression and pathogen resistance cannot be induced by INA or avirulent pathogens in the tga6-1 tga2-1 tga5-1 triple knockout mutant, suggesting that these TGA transcription factors serve as essential positive regulators of SAR. We also showed that genomic clones containing either TGA2 or TGA5 can complement the loss of the SAR phenotype in the TGA triple mutant. Thus, either TGA2 or TGA5 is sufficient for INA-induced PR gene expression and pathogen resistance. Because the loss of the SAR phenotype cosegregated with the tga6-1 mutation and was observed only in the triple mutant but not in tga2-1 tga5-1, TGA6 also is sufficient for INA-induced PR gene expression and pathogen resistance. These data demonstrate that TGA2, TGA5, and TGA6 encode redundant functions in the induction of SAR.
Furthermore, we found that tga2-1 tga5-1 and tga6-1 tga2-1 tga5-1 accumulated increased levels of PR-1 under noninducing conditions, suggesting that these TGA transcription factors repress the basal expression of PR-1. The higher basal level of PR-1 in the triple mutant, compared with the double mutant, suggests that TGA6 is partially responsible for the negative regulation of basal levels of PR-1. On the other hand, either TGA2 or TGA5 appeared to be sufficient to suppress the basal expression of PR-1, because transforming the genomic clone of either TGA2 or TGA5 into the triple mutant reverted PR-1 expression to the wild-type level. The increased PR-1 expression probably is the result of the loss of binding of TGA factors to a negative element on the PR-1 promoter. This hypothesis is supported by the presence of an as-1–related TGACG element (LS5) that functions as a weak silencer in the PR-1 promoter (Lebel et al., 1998).
In addition to the loss of SAR phenotypes, npr1 plants also exhibited reduced tolerance to high concentrations of SA and enhanced susceptibility to the bacterial pathogen P.s.m. ES4326. We found that tga6-1 tga2-1 tga5-1 seedlings were very sensitive to SA toxicity, suggesting that these TGA transcription factors may work together with NPR1 to regulate tolerance to high levels of SA. Unlike npr1 plants, TGA triple knockout plants did not exhibit enhanced susceptibility to P.s.m. ES4326 (Figure 5). It is possible that the basal resistance to P.s.m. ES4326 is regulated by other mechanisms. In Arabidopsis, there are at least two other TGA transcription factors that can bind to NPR1 in the yeast two-hybrid system (Després et al., 2000; Zhou et al., 2000). Our data suggest that the other TGA transcription factors do not have the same function as TGA2, TGA5, and TGA6 in the positive regulation of SAR. Whether they serve other biological functions regulated by NPR1 remains to be determined.
METHODS
Generation of Triple Knockout Mutants for TGA2, TGA5, and TGA6
TGA2 and TGA5 are located next to each other, and the identification of tga2-1 tga5-1 was described previously (Li et al., 2001). To identify a deletion mutant for TGA6, an Arabidopsis thaliana deletion mutant population of 51,840 lines was screened by PCR using primers flanking the gene. The distance between the primers is ∼9 kb. The PCR extension time was set at 1.5 min to avoid amplification of the wild-type DNA fragment. A single deletion mutant, tga6-1, was detected initially in one of the megapools containing 2592 lines. Individual mutant plants were isolated subsequently by deconvolution as described previously (Li et al., 2001). tga6-1 tga2-1 tga5-1 was generated using pollen from tga6-1 plants to fertilize tga2-1 tga5-1 plants. The F1 plants were selfed, and tga6-1 tga2-1 tga5-1 was identified in the F2 generation by PCR using primers within the deletions to confirm homozygosity at both loci.
Analysis of PR Gene Expression in the Mutant Plants
To analyze gene expression levels by real-time reverse transcription–PCR, total RNA samples were prepared from 20-day-old plants grown on MS medium (Murashige and Skoog, 1962), with or without INA, using the Totally RNA kit from Ambion (Austin, TX). Reverse transcription of the cDNA was performed using the RT-for-PCR kit from Clontech (Palo Alto, CA). Real-time PCR was performed using the QuantiTect SYBR Green PCR kit from Qiagen (Valencia, CA). The primers used to amplify PR-1 were 5′-GTAGGTGCTCTTGTTCTTCCC-3′ and 5′-CACATAATTCCCACGAGGATC-3′. The primers used to amplify ACTIN1 were 5′-CGATGAAGCTCAATCCAAACGA-3′ and 5′-CAGAGTCGAGCACAATACCG-3′.
Pathogen Infections
Both tga6-1 and tga2-1 tga5-1 are in the Columbia ecotype background, and Peronospora parasitica Noco2 is virulent on this ecotype. Infection of wild-type and tga plants with P. parasitica Noco2 was performed by spraying a suspension of conidia (∼5 × 103 spores/mL water) on 2-week-old soil-grown plants. Inoculated plants were maintained subsequently in a Conviron TC16 growth chamber (Winnipeg, Canada) at 18°C with a 12-h photoperiod and ∼80% RH. A disease rating was determined for each plant according to Cao et al. (1998) at 7 days after inoculation. For each genotype and treatment, 25 plants were scored.
Infection with the virulent bacterial pathogen Pseudomonas syringae pv maculicola (P.s.m.) ES4326 was performed by infiltrating leaves of 4-week-old soil-grown wild-type and mutant plants with a bacterial suspension at OD600 = 0.001, which is the dose that normally causes disease in wild-type plants. Symptoms were examined 3 days after inoculation. The bacterial titer in the leaves was measured according to a previously described procedure (Cao et al., 1994).
Complementation of tga6-1 tga2-1 tga5-1 Plants by Wild-Type Genes
A 5.9-kb KpnI-BamHI fragment (MOJ9, nucleotides 28,249 to 34,162) containing TGA2 was subcloned from P1 clone MOJ9 to pGreen229 (Hellens et al., 2000) to create pG229-TGA2. A 4.6-kb EcoRI-SacI fragment (MOJ9, nucleotides 33,110 to 37,773) containing TGA5 was subcloned from MOJ9 to pGreen229 to create pG229-TGA5. pG229-TGA2 and pG229-TGA5 were transformed into tga6-1 tga2-1 tga5-1 using the floral-dip method (Clough and Bent, 1998), and transformants were selected on soil by spraying the T1 plants with the herbicide glufosinate. At least 10 transformants were obtained for each construct. All assays on the complementing lines were performed on glufosinate-resistant T2 plants.
Upon request, materials integral to the findings presented in this publication will be made available in a timely manner to all investigators on similar terms for noncommercial research purposes. To obtain materials, please contact Yuelin Zhang, yuelin@interchange.ubc.ca
Accession Number
The accession number for BAC clone F28J15 is AC069472.
Acknowledgments
We thank Brian Staskawicz for P.s.t. DC3000 avrRpt2 strain, Jian-min Zhou and Jane Parker for sending materials used in this study, and Sandra Goritschnig and Kristoffer Palma for careful reading of the manuscript. We are grateful for financial support to Y.Z. from the Natural Sciences and Engineering Council of Canada and to X.L. from the Canadian Foundation for Innovation and the Natural Sciences and Engineering Council of Canada.
Article, publication date, and citation information can be found at www.plantcell.org/cgi/doi/10.1105/tpc.014894.
References
- Aravind, L., and Koonin, E.V. (1999). Fold prediction and evolutionary analysis of the POZ domain: Structural and evolutionary relationship with the potassium channel tetramerization domain. J. Mol. Biol. 285, 1353–1361. [DOI] [PubMed] [Google Scholar]
- Cao, H., Bowling, S.A., Gordon, A.S., and Dong, X. (1994). Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 6, 1583–1592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao, H., Glazebrook, J., Clarke, J.D., Volko, S., and Dong, X. (1997). The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell 88, 57–63. [DOI] [PubMed] [Google Scholar]
- Cao, H., Li, X., and Dong, X. (1998). Generation of broad-spectrum disease resistance by overexpression of an essential regulatory gene in systemic acquired resistance. Proc. Natl. Acad. Sci. USA 95, 6531–6536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chern, M.S., Fitzgerald, H.A., Yadav, R.C., Canlas, P.E., Dong, X., and Ronald, P.C. (2001). Evidence for a disease-resistance pathway in rice similar to the NPR1-mediated signaling pathway in Arabidopsis. Plant J. 27, 101–113. [DOI] [PubMed] [Google Scholar]
- Clough, S.J., and Bent, A.F. (1998). Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743. [DOI] [PubMed] [Google Scholar]
- Delaney, T.P., Friedrich, L., and Ryals, J.A. (1995). Arabidopsis signal transduction mutant defective in chemically and biologically induced disease resistance. Proc. Natl. Acad. Sci. USA 92, 6602–6606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Després, C., DeLong, C., Glaze, S., Liu, E., and Fobert, P.R. (2000). The Arabidopsis NPR1/NIM1 protein enhances the DNA binding activity of a subgroup of the TGA family of bZIP transcription factors. Plant Cell 12, 279–290. [PMC free article] [PubMed] [Google Scholar]
- Fan, W., and Dong, X. (2002). In vivo interaction between NPR1 and transcription factor TGA2 leads to salicylic acid–mediated gene activation in Arabidopsis. Plant Cell 14, 1377–1389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaffney, T., Friedrich, L., Vernooij, B., Negrotto, D., Nye, G., Uknes, S., Ward, E., Kessmann, H., and Ryals, J. (1993). Requirement of salicylic acid for the induction of systemic acquired resistance. Science 261, 754–756. [DOI] [PubMed] [Google Scholar]
- Glazebrook, J., Rogers, E.E., and Ausubel, F.M. (1996). Isolation of Arabidopsis mutants with enhanced disease susceptibility by direct screening. Genetics 143, 973–982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Görlach, J., Volrath, S., Knauf-Beiter, G., Hengy, G., Beckhove, U., Kogel, K.-H., Oostendorp, M., Staub, T., Ward, E., Kessmann, H., and Ryals, J. (1996). Benzothiadiazole, a novel class of inducers of systemic acquired resistance, activates gene expression and disease resistance in wheat. Plant Cell 8, 629–643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hellens, R.P., Edwards, E.A., Leyland, N.R., Bean, S., and Mullineaux, P.M. (2000). pGreen: A versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol. Biol. 42, 819–832. [DOI] [PubMed] [Google Scholar]
- Kim, H.S., and Delaney, T.P. (2002). Over-expression of TGA5, which encodes a bZIP transcription factor that interacts with NIM1/NPR1, confers SAR-independent resistance in Arabidopsis thaliana to Peronospora parasitica. Plant J. 32, 151–163. [DOI] [PubMed] [Google Scholar]
- Lebel, E., Heifetz, P., Thorne, L., Uknes, S., Ryals, J., and Ward, E. (1998). Functional analysis of regulatory sequences controlling PR-1 gene expression in Arabidopsis. Plant J. 16, 223–233. [DOI] [PubMed] [Google Scholar]
- Li, X., Song, Y., Century, K., Straight, S., Ronald, P., Dong, X., Lassner, M., and Zhang, Y. (2001). A fast neutron deletion mutagenesis-based reverse genetics system for plants. Plant J. 27, 235–242. [DOI] [PubMed] [Google Scholar]
- Malamy, J., Carr, J.P., Klessig, D.F., and Raskin, I. (1990). Salicylic acid: A likely endogenous signal in the resistance response of tobacco to viral infection. Science 250, 1002–1004. [DOI] [PubMed] [Google Scholar]
- Métraux, J.P., Ahl-Goy, P., Staub, T., Speich, J., Steinemann, A., Ryals, J., and Ward, E. (1991). Induced resistance in cucumber in response to 2,6-dichloroisonicotinic acid and pathogens. In Advances in Molecular Genetics of Plant-Microbe Interactions, Vol. 1, H. Hennecke and D.P.S. Verma, eds (Dordrecht, The Netherlands: Kluwer Academic Publishers), pp. 432–439.
- Métraux, J.P., Signer, H., Ryals, J., Ward, E., Wyss-Benz, M., Gaudin, J., Raschdorf, K., Schmid, E., Blum, W., and Inverardi, B. (1990). Increase in salicylic acid at the onset of systemic acquired resistance in cucumber. Science 250, 1004–1006. [DOI] [PubMed] [Google Scholar]
- Murashige, T., and Skoog, F. (1962). A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol. Plant. 15, 473–497. [Google Scholar]
- Nawrath, C., Heck, S., Parinthawong, N., and Metraux, J.P. (2002). EDS5, an essential component of salicylic acid–dependent signaling for disease resistance in Arabidopsis, is a member of the MATE transporter family. Plant Cell 14, 275–286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nawrath, C., and Métraux, J.-P. (1999). Salicylic acid induction–deficient mutants of Arabidopsis express PR-2 and PR-5 and accumulate high levels of camalexin after pathogen inoculation. Plant Cell 11, 1393–1404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niggeweg, R., Thurow, C., Kegler, C., and Gatz, C. (2000. a). Tobacco transcription factor TGA2.2 is the main component of as-1-binding factor ASF-1 and is involved in salicylic acid- and auxin-inducible expression of as-1-containing target promoters. J. Biol. Chem. 275, 19897–19905. [DOI] [PubMed] [Google Scholar]
- Niggeweg, R., Thurow, C., Weigel, R., Pfitzner, U., and Gatz, C. (2000. b). Tobacco TGA factors differ with respect to interaction with NPR1, activation potential and DNA-binding properties. Plant Mol. Biol. 42, 775–788. [DOI] [PubMed] [Google Scholar]
- Pontier, D., Miao, Z.H., and Lam, E. (2001). Trans-dominant suppression of plant TGA factors reveals their negative and positive roles in plant defense responses. Plant J. 27, 529–538. [DOI] [PubMed] [Google Scholar]
- Rasmussen, J.B., Hammerschmidt, R., and Zook, M.N. (1991). Systemic induction of salicylic acid accumulation in cucumber after inoculation with Pseudomonas syringae pv. syringae. Plant Physiol. 97, 1342–1347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers, E.E., and Ausubel, F.M. (1997). Arabidopsis enhanced disease susceptibility mutants exhibit enhanced susceptibility to several bacterial pathogens and alterations in PR-1 gene expression. Plant Cell 9, 305–316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryals, J., Weymann, K., Lawton, K., Friedrich, L., Ellis, D., Steiner, H.-Y., Johnson, J., Delaney, T.P., Jesse, T., Vos, P., and Uknes, S. (1997). The Arabidopsis NIM1 protein shows homology to the mammalian transcription factor inhibitor IκB. Plant Cell 9, 425–439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryals, J.A., Neuenschwander, U.H., Willits, M.G., Molina, A., Steiner, H.Y., and Hunt, M.D. (1996). Systemic acquired resistance. Plant Cell 8, 1809–1819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shah, J., Tsui, F., and Klessig, D.F. (1997). Characterization of a salicylic acid-insensitive mutant (sai1) of Arabidopsis thaliana, identified in a selective screen utilizing the SA-inducible expression of the tms2 gene. Mol. Plant-Microbe Interact. 10, 69–78. [DOI] [PubMed] [Google Scholar]
- Subramaniam, R., Desveaux, D., Spickler, C., Michnick, S.W., and Brisson, N. (2001). Direct visualization of protein interactions in plant cells. Nat. Biotechnol. 19, 769–772. [DOI] [PubMed] [Google Scholar]
- Uknes, S., Mauch-Mani, B., Moyer, M., Potter, S., Williams, S., Dincher, S., Chandler, D., Slusarenko, A., Ward, E., and Ryals, J. (1992). Acquired resistance in Arabidopsis. Plant Cell 4, 645–656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White, R.F. (1979). Acetylsalicylic acid (aspirin) induces resistance to tobacco mosaic virus in tobacco. Virology 99, 410–412. [DOI] [PubMed] [Google Scholar]
- Wildermuth, M.C., Dewdney, J., Wu, G., and Ausubel, F.M. (2001). Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414, 562–565. [DOI] [PubMed] [Google Scholar]
- Zhang, Y.L., Fan, W.H., Kinkema, M., Li, X., and Dong, X. (1999). Interaction of NPR1 with basic leucine zipper protein transcription factors that bind sequences required for salicylic acid induction of the PR-1 gene. Proc. Natl. Acad. Sci. USA 96, 6523–6528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou, J.M., Trifa, Y., Silva, H., Pontier, D., Lam, E., Shah, J., and Klessig, D.F. (2000). NPR1 differentially interacts with members of the TGA/OBF family of transcription factors that bind an element of the PR-1 gene required for induction by salicylic acid. Mol. Plant-Microbe Interact. 13, 191–202. [DOI] [PubMed] [Google Scholar]