Abstract
We describe the resolution of heparan sulfate (HS) isomers by chromatographic methods and their subsequent differentiation by mass spectrometry (MS), ion mobility, and 1H-NMR analysis. The two purified hexasaccharide isomers produced nearly identical MS spectra, quantitative disaccharide profiles, and partial enzymatic digestions. However, both tandem spectrometry (MS2) and ion mobility spectrometry (IMS) indicated structurally differences existed. All data suggested the distinction between the two hexasaccharides resided in their uronic acid stereochemistries. Glucuronic (GlcA) and iduronic acids (IdoA) were subsequently defined by 1H-NMR analysis completing the structural analysis and verifying the unique structures initially indicated by MS2 and IMS. Our results suggest that IMS may be a powerful tool in the rapid differentiation of GlcA and IdoA containing isomers in the absence of prior structural knowledge.
Keywords: heparan sulfate, glycosaminoglycan, uronic acid, sequencing, chromatography, mass spectrometry, ion mobility, nuclear magnetic resonance
INTRODUCTION
Glycosaminoglycans (GAGs) are unbranched, anionic polysaccharides predominantly present at cell surfaces and in the extracellular matrix. These polysaccharides are composed of repeating disaccharide units and are so named for the fact that one residue within each disaccharide is an amino sugar; either N-acetylglucosamine (GlcNAc) or N-acetylgalactosamine (GalNAc). With the exception of keratan sulfate, the residue to the non-reducing end of the amino sugar is an uronic acid; i.e. either of the C5 epimers iduronic acid (IdoA) or glucuronic acid (GlcA). GAGs play diverse roles in biology. Some of their functions include provision of mechanical support in the extracellular matrix, modulation of enzymatic activities, and immobilization of morphogen gradients during development. Many of the biological functions of the GAG family are carried out in some way or another through protein interactions, and physiologically the most relevant GAG involved in protein interactions is heparan sulfate (HS) 1–5.
Heparan sulfate synthesis takes place within the Golgi apparatus where the polymer is synthesized as a repeating array of [-GlcNAcα1–4GlcAβ1–4-]n. During chain elongation, and perhaps most important for protein binding, nascent HS is acted upon by a series enzymes which chemically modify the polymer. The modification reactions on GlcNAc residues include N-deacetylation/N-sulfation, 6-O, and 3-O sulfation. GlcA residues may be modified by epimerization at C5 (converting the sugar to IdoA), and 2-O sulfation. These modification reactions do not approach completion resulting in the production of HS molecules with varying sequences of acetylation, sulfation, and IdoA content 6–10. To a large extent, the average modification profile and structure of HS varies with the tissue where it was synthesized 11–13. Despite the modification content being relatively constant within a tissue, specific sequences (for example those binding to antithrombin III 14–16, basic fibroblast growth factor 17–20, or phage display antibodies 21) may only be found in select chains.
Overall HS has been implicated in a multitude physiologic and pathophysiologic processes, including blood coagulation 22–24, viral infectivity 25–27, cancer 28–33, inflammation 34–39, growth, and development 40–45. For all of these roles, only a small number of HS protein-binding sequences have been solved 2, 4, 5, 7, 46–48. The reasons for this are many and include difficulties in isolating large amounts of pure sequence, inherent difficulties of HS chemical synthesis, the fact that some proteins can bind multiple HS sequences, and that only a relatively small number of methods for defining any purified HS structure exist. Traditional methods for HS sequencing have relied upon either 2-D NMR, or incorporation of radio or chemical labels followed by extensive enzymatic processing 49–54. While such methods have utility in specific contexts, no single method is well suited for all applications. For instance, while 2-D NMR can provide 3-dimensional structural information, the technique often requires milligram quantities of material which may not be available. Labeling methods rely on the use of radioisotopes in cell culture or chemical labeling reactions which may proceed with varying efficiencies depending on the substrate and matrix.
To further facilitate HS sequence analysis, we present an efficient method of sequencing that relies heavily, but not exclusively, on mass spectrometry (MS). Our method is significantly more sensitive than 2-D NMR analysis and similarly sensitive to end labeling methods 51, 52, 54. We describe the sequencing of two HS hexasaccharide isomers whose structures had been previously resolved following different isolation and sequencing methods 49, 54, 55. We also report, for the first time, ion mobility resolution of HS sequences based solely on uronic acid identity. In our method, chromatographically-purified isomers were analyzed by nanoelectrospray MS, IMS, and MS2 time-of-flight (TOF) mass spectrometry. MS-based compositional analysis was used to identify and quantify the disaccharides comprising the hexasaccharides 56, 57, while their linear sequences were reconstructed by partial enzymatic digestions. Finally, 1-D NMR was used to verify the IMS data suggesting differentiation between the iduronic and glucuronic acid isomers, thus completing the sequence analysis. In light of the importance of HS in biology in medicine, and given that resources, materials, and technical capabilities will undoubtedly vary from one research lab to another, we present this efficient, new approach to HS sequencing. Results presented herein indicate that IMS may be a powerful, untapped resource for HS analysis.
EXPERIMENTAL SECTION
Materials
Porcine intestinal heparin sulfate was purchased from Celsus Laboratories (Cincinnati, OH). Bio-Gel® P-10 Gel was obtained from Bio-Rad (Hercules, CA). The IonPac® AS7 anion-exchange column was purchased from Dionex (Sunnyvale, CA). The heparinase III (heparitinase, EC 4.2.2.8) used for preparative digestion was a gracious gift from Professor Jian Liu, UNC School of Pharmacy, Chapel Hill, NC. Standards used in disaccharide compositional analysis were purchased from Calbiochem (La Jolla, CA), Sigma-Aldrich Corp. (St. Louis, MO), and Dextra Laboratories (Reading, UK). Heparinases I (EC 4.2.27), II (no EC number), and III (EC 4.2.2.8) used in quantitative disaccharide compositional analysis were obtained from Seikagaku Biobusiness Corp. (Tokyo, Japan). All other chemicals were purchased from Fisher Scientific or Sigma-Aldrich chemical co.
Preparative and Analytical HS Depolymerization
Heparinase III digestion of intact HS was performed essentially as previously described 58. Briefly, HS (50 mg/ml in 50 mM sodium phosphate, pH 7.5) was extensively depolymerized at 37° C 59, 60. The reaction was monitored by UV-absorption until the product concentration reached 18 mM (ε = 5,500 M−1cm−11 in 0.03 M HCl at 232 nm 61). The reaction was quenched by addition of 50% methanol, boiled for 10 minutes, and then frozen and lyophilized. Partial digests for analytical purposes were performed using approximately 0.5 mU of heparinase I, II, or III, and 1–2 nMol of each hexasaccharide in 50mM NH4OAc. These reactions were quenched by the addition of the nanoelectrospray ionization (nESI) solvent containing 1:1 MeOH: H2O and 0.1% acetic acid.
Size Exclusion Chromatography
The depolymerized HS was redissolved in 1 M NH4HCO3. The solution was centrifuged to remove any particulates and then loaded onto a 170 × 1.5 cm Econo-Column® (Bio-Rad, Hercules, CA) packed with P-10 size exclusion gel. The separation was performed under gravity flow at room temperature in 40 mM NH4HCO3 and ~1 ml fractions were collected using a BioLogic LP system (Bio-Rad, Hercules, CA). Initial separation was carried out between 0.5 and 1.0 ml/hour which served to remove an otherwise high concentration of disaccharides (dp2, degree of polymerization-2, etc.) from the rest of the oligosaccharide products. The fractions containing the larger oligosaccharides were pooled, freeze-dried, re-dissolved in 1 M NH4HCO3, re-loaded onto the column, and run at a flow rate of 3.5 ml/hour. Separation was monitored by UV absorption online at 254 nm, and later verified offline at 232 nm. Collected fractions were lyophilized using a Vacufuge™ (Eppendorf, Westbury, NY). Each of the dried fractions was then re-dissolved in 100 µl of Millipore H2O, quantified by UV-absorption, and analyzed by nESI TOF MS.
Strong Anion-Exchanged Chromatography
Fractions obtained from SEC corresponding to hexasaccharides (dp6) were pooled and re-quantified. SAX chromatography was run on a Waters Delta 600 System (Waters Corp., Milford, MA) using multistage, linear gradients as follows: solvent A, H2O (pH 3.5); solvent B, 4 M NaCl (pH 3.5); 1–5 min, 0% B; 6–75 min, 8–15% B; 75–85 min, 15–17% B; 85–100 min, 17–19% B. Eluting peaks were detected by UV-absorption at 232 nm. Between 100 and 200 µg of material were injected for individual runs, with run to run variability generally less than one minute. Peak fractions were collected, lyophilized, and re-dissolved in the smallest possible volume of H2O limited by the solubility of NaCl. Re-dissolved fractions were desalted by dialysis, at room temperature, first against degassed 500 mM NH4OAc, and then against Millipore H2O using 1kDa MWCO (molecular weight cut-off) Dispo-Biodialyzers™ (The Nest Group, inc., Southborough, MA). The desalted fractions were concentrated and quantified by spectrophotometry.
IMS and MS of Intact and Partially Digested Hexasaccharides
Borosilicate glass capillaries (# 1B120F-4) were purchased from World Precision Instruments, Inc. (Sarasota, FL). The capillaries were pulled on a Sutter P-97 Flaming/Brown micropipette puller, and gold coated using a Quorum Technologies SC7640 sputter coater (Ringmer, East Sussex, UK). MS and MS2 analyses were performed on a Waters Synapt™ HDMS™ system (Waters Corp., Milford, MA). The following parameters were used: nESI capillary voltage, 0.6 kV; sample cone, 10 V; extraction cone, 0 V; source temperature, 40°C; trap CE, 1 V; transfer CE, 3 V; trap gas flow, 1.0 ml/min; IMS gas flow, 24 ml/min; trap direct current bias, 7 V; pusher interval, 45 µs; IMS wave velocity, 400 m/s; IMS wave height, 8.0 V; transfer wave velocity, 350 m/s; transfer wave height, 5.0 V. For mass to charge-selected ions, LM and HM resolutions were each set to 14.0 or higher with the quadrupole set to isolate the m/z of interest. Collision induced dissociation (CID) of hexasaccharides was performed using a trap collision energy of 16 V, and for all experiments the time of flight analyzer was calibrated with a solution of NaI. For IMS analysis, the Synapt G2™ q-IMS-TOF mass spectrometer was used. Synapt G2™ parameters were generally similar to the parameters provided for the first generation Synapt™ mass spectrometer provided above with the following exceptions: extraction cone, 2 V; source temperature, 50°C; trap CE, 4 V; transfer CE, 0 V; trap gas flow, 1.5 ml/min; IMS gas flow, 180 ml/min; helium gas flow, 200 ml/min: trap direct current bias, 40 V; pusher interval, 50 µs; IMS wave velocity, 1600 m/s; IMS wave height, 4.7–5.1 V ramp over 10 % of IMS cycle; transfer wave velocity, 206 m/s; transfer wave height, 4.0 V.
Quantitative Disaccharide Compositional Analysis
Quantitative disaccharide compositional analysis was performed essentially as previously described 56, 57. Briefly, 5 µg of hexasaccharide from each of the fractions were fully digested to disaccharide constituents in 20 µL of 7.5 mM NH4OAc (pH 7.5) containing 1 mM Ca(OAc)2 and 2 mU each of Heparin Lyases I, II, and III. The reactions were incubated at 37° C for 16 hours and then stopped by the addition of 180 µl of quench solution to create a final solution containing 1:1 MeOH/H2O, 10 mM NH4OH, and 5 µM I-P (ΔUA2S-GlcNCOEt6S) internal standard 56, 57. Samples were analyzed by ESI-MSn without further purification. Mass spectra were collected via ESI-linear ion trap MS (LTQ, Thermo Finnigan LTQ, San Jose, CA) using Xcaliber Version 2.0 data acquisition software. All spectra were obtained in negative ion mode with a 3.5 kV spray voltage and a 225° C capillary temperature. Samples were introduced by flow injection analysis at 10 µL/min with 1:1 MeOH/H2O containing 10 mM NH4OH. MS and MS2 data acquisition and quantitative analysis of compositions were also performed as previously described 56, 57. Heparin from bovine lung and HS from bovine kidney were used as controls for all aspects of the experimental method. Measurements were obtained in triplicate and results are reported as an average ± standard deviation.
NMR Spectroscopy
NMR experiments were performed on a Bruker Avance III 600 MHz spectrometer, equipped with a cryogenically cooled triple resonance (TCI) probe. Proton chemical shifts are reported with respect to external DSS (2,2-dimethyl-2-silapentane-5-sulfonic acid). Experiments were performed at 298 K using 512 scans, a 2 s recycle delay, and a 6600 Hz sweep-width (32,000 complex points). Water suppression was accomplished using pre-saturation of the water resonance during the recycle delay. Data were processed using Bruker TopSpin software. NMR samples contained approximately 0.1 mM oligosaccharide each and were dissolved in 99.9% D2O to remove exchangeable protons.
RESULTS AND DISCUSSION
Purification of HS Hexasaccharide Isomers
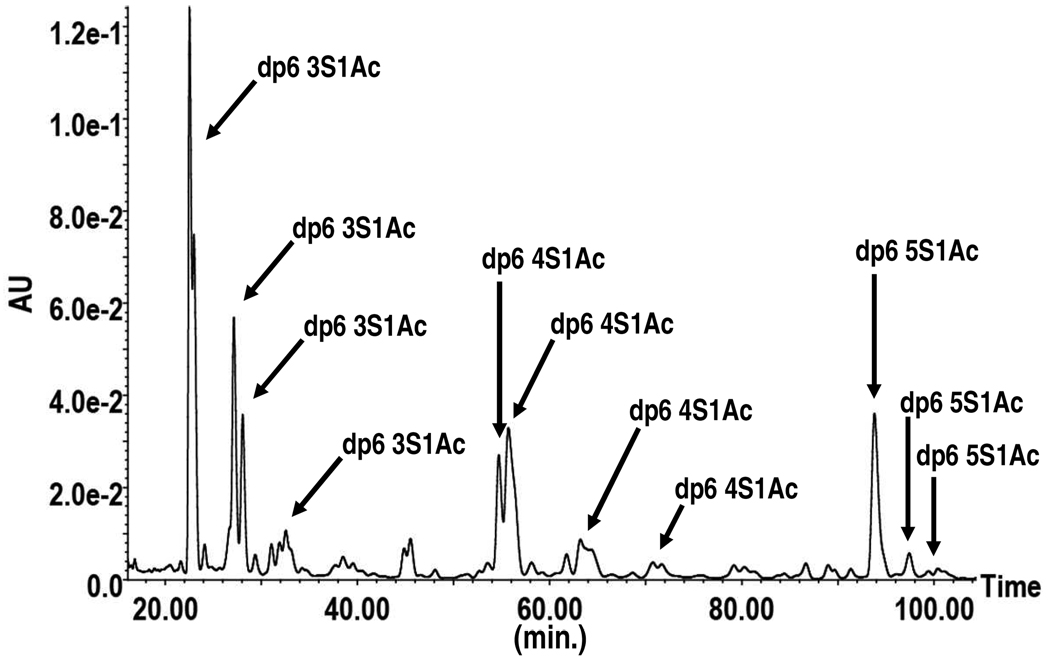
Intact HS was digested to oligomers ranging from dp2 to d12 using heparinase III 58–60. The products were subsequently separated by size exclusion chromatography (SEC), and oligosaccharides differing in length by at least one disaccharide were near baseline resolved (Figure 1). The resultant product fractions containing hexasaccharides (dp6) were pooled and subjected to strong anion exchange (SAX) chromatography using an IonPac AS7 column. This column type was chosen as it had previously been shown useful in differentiation of polyvalent anions 62, but to our knowledge had not been used in GAG analysis. When running a shallow gradient, the column proved effective in separating both isomeric hexasaccharides as well as those differing in sulfate number (Figure 2). Of particular interest is the observation that isomeric hexasaccharides with three sulfate groups and a single acetyl group eluted at distinct retentions times between 22 and 36 minutes (we use sulfate to denote either sulfate or sulfamate groups on the saccharide). A strong peak with a prominent shoulder is observed around 22 minutes along with two well-resolved peaks centered around 27 and 28 minutes, respectively. Meanwhile, a smaller series of incompletely resolved peaks are present eluting between 30 and 34 minutes. The peak at 22 minutes and its shoulder represent a single chemical entity, interconverting conformers, or anomers as evidenced by identical MS2 spectra and verified by reinjection of individually collected peaks onto the column. The peaks present at 27 and 28 minutes displayed the same relationship. Given the conformational flexibility in iduronate rings 63–65, and the fact that reducing-end residues are expected to be present as both α and β anomers 2, 5, either possibility could explain the split peaks. Taking this into consideration, the fraction at 22 minutes and its shoulder were pooled and analyzed as a single entity, as were the fractions collected at 27 and 28 minutes. For simplicity, the pooled fractions are referred to as fractions 22 and 27, respectively.
Figure 1.
SEC chromatogram of partially depolymerized HS. Peaks corresponding to di-through decasaccharides are labeled dp2 to dp10.
Figure 2.
IonPac AS7 SAX chromatogram of HS hexasaccharides (dp6). The number of sulfates and the number of acetyl groups present on the hexasaccharides in the collected peaks are represented by the numbers preceding ‘S’ and ‘Ac’, respectively.
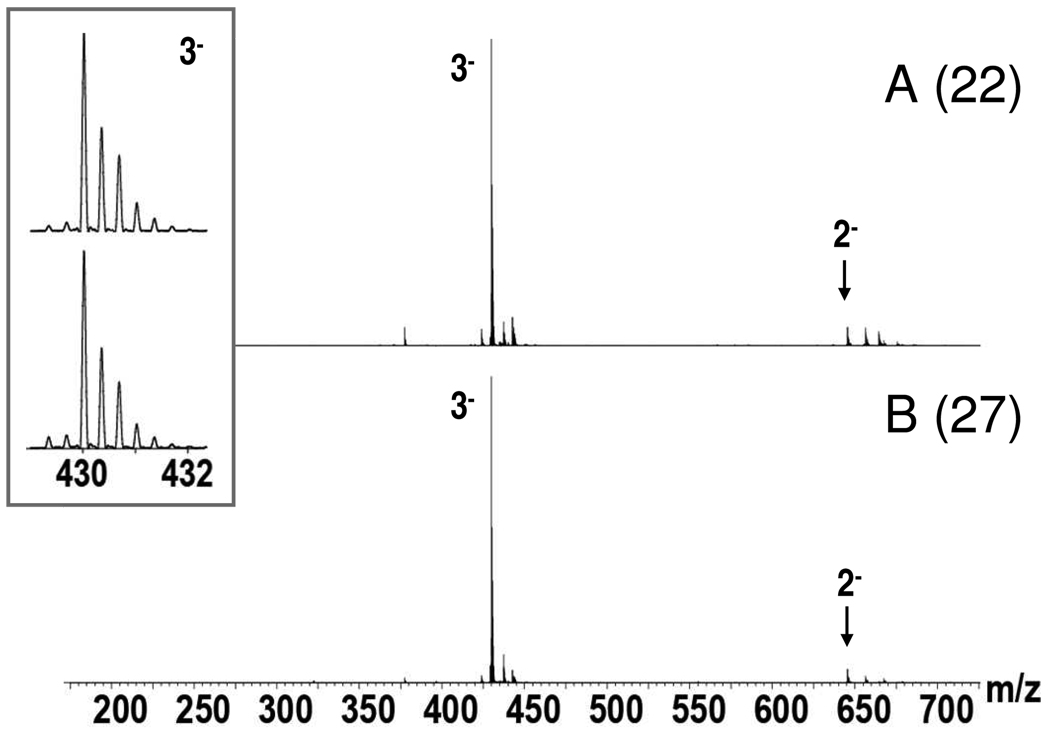
The full-scan MS spectra of fractions 22 and 27 were obtained with isotopic resolution providing a molecular mass of 1293.18 Da (Figure 3). The HOST macro was used to convert the molecular mass to the composition of a hexasaccharide with 3 sulfate groups and 1 acetyl group 66. Both purified fractions were judged to be extremely pure based on the mass spectra. It is also of interest that under the gentle nanospray conditions employed, sulfate loss, which can lead to overestimation of the components in a mixture, was non-existent.
Figure 3.
Full scan mass spectra of fractions 22 (A) and 27 (B). The inset displays the isotopic resolution with which these spectra were acquired.
MS2 and Ion Mobility Analysis of Intact Hexasaccharides
Following observation of distinct retention times, the second indicator that we had purified isomeric saccharides came from differences in the MS2 spectra of isolated species. Interestingly, the same product ions were formed for both fractions 22 and 27, but differed in relative abundances (figure 4). A similar relationship had been observed for uronic acid residue isomers in chondroitin and dermatan sulfate, but had not previously been observed in HS 67. The most intense product ions for each species are labeled in figure 4 using the Domon and Costello nomenclature 68, while all product ions present at greater than 1 % relative abundance are listed in supplemental table 1. Product ions were assigned based on cleavage patterns previously observed in HS disaccharides 56, 69 and extrapolated to the larger hexasaccharides in the present study. Given that sulfate loss is observed upon MS2 (Table S1), it is precarious to assign sulfate group positions from MS2 of an intact oligosaccharides alone. Acetyl modifications are, however, relatively stable in the gas phase and the MS2 fragments observed (X and Y ions) localize this modification to the reducing end of both hexasaccharides (Table S1). It is of particular note, that we observe differences in ion mobility arrival time distributions (ATDs) for the two purified isomers. The hexasaccharide in fraction 22 arrived at 7.02 ms while the hexasaccharide in fraction 27 arrived earlier at 6.77 ms (Figure 5). Under identical instrumental conditions, IMS arrival times are influenced by structure (as collision cross section), mass, and charge 70–72. Since both analytes have identical mass and charge, and were analyzed under identical conditions, these ATD differences are a clear indicator of their difference in molecular structure.
Figure 4.
MS2 spectra of fractions 22 (A) and 27 (B). Predominant fragment ions are labeled using the Domon and Costello nomenclature 68, whereas all fragment ions above 1% relative abundance are listed in supplementary table 1.
Figure 5.
Overlay of the ion mobility arrival time distributions for the 2− charge states of the hexasaccharide isomers in fractions 22 and 27. Arrival times are reported in milliseconds
Disaccharide Compositional Analysis
The hexasaccharides in fractions 22 and 27 were completely digested to disaccharides using a mixture of heparinases I, II, and III. Each disaccharide was then quantified via flow injection analysis using ΔUA2S-GlcNCOEt6S as an internal standard, and isomeric disaccharides were distinguished and quantified by their diagnostic ions in MS2 spectra 56, 57. The results obtained for fractions 22 and 27 are presented in table 1. Of note is that for both fractions, the disaccharides IV-A (ΔUA-GlcNAc), IV-S (ΔUA-GlcNS), and III-S (ΔUA2S-GlcNS) each make up approximately 30% of the mixture. While these data indicate the possibility of unique hexasaccharide structures in the range of 90% purity, the data provide neither the linear sequence of the disaccharides nor the stereochemistry at C5 of the uronic acids. The latter stereochemical information is lost through the use of heparinases I, II, and III which cleave via beta-elimination resulting in a C4–C5 double bond with loss of chirality at these stereogenic centers 46, 60.
Table 1.
Disaccharide compositional analysis results for fractions 22 and 27.
| Disaccharide | Fraction 22 Comp. (%) |
Fraction 27 Comp. (%) |
|
|---|---|---|---|
| IV-A | ΔUA-GlcNAc | 30.2 ± 0.2 | 31.2 ± 0.8 |
| III-A | ΔUA2S-GlcN | 0.3 ± 0.1 | 1.2 ± 0.1 |
| II-A | ΔUA-GlcNAc6S | 2.1 ± 0.5 | 2.1 ± 0.1 |
| I-A | ΔUA2S-GlcNAc6S | 0.1 ± 0.0 | 0.1 ± 0.0 |
| IV-S | ΔUA-GlcNS | 35.5 ± 0.9 | 35.8 ± 1.4 |
| III-S | ΔUA2S-GlcNS | 30.1 ± 0.3 | 27.7 ± 0.8 |
| II-S | ΔUA-GlcNS6S | 0.1 ± 0.0 | 0.1 ± 0.0 |
| I-S | ΔUA2S-GlcNS6S | 0.1 ± 0.0 | 0.1 ± 0.0 |
| I-H | ΔUA2S-GlcNH6S | 0.0 ± 0.0 | 0.0 ± 0.0 |
| II-H | ΔUA-GlcNH6S | 0.0 ± 0.0 | 0.0 ± 0.0 |
| III-H | ΔUA2S-GlcNH | 0.0 ± 0.0 | 0.0 ± 0.0 |
| IV-H | ΔUA-GlcNH | 1.5 ± 0.3 | 1.6 ± 0.3 |
Partial Enzymatic Digestion of Hexasaccharides
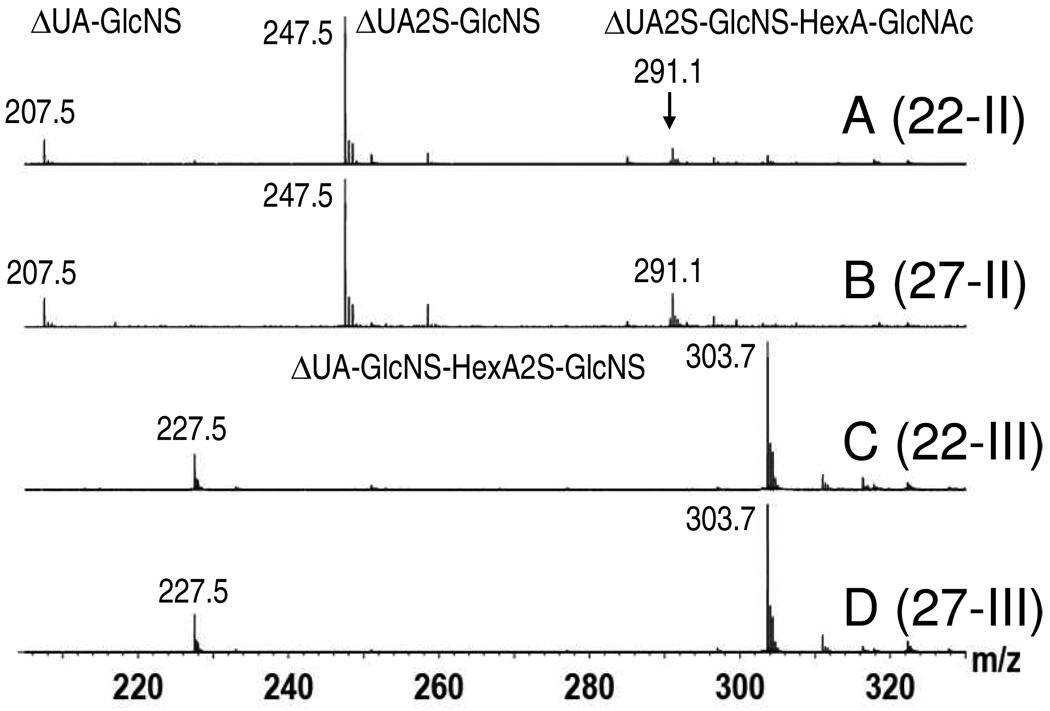
Partial enzymatic digestion is an effective way to generate sequence information between known disaccharide blocks present in longer oligosaccharides. Differential cleavage sites can be used to identify neighboring disaccharides and generate their linear sequence in the oligosaccharide. Two sets of partially cleaved hexasaccharides were generated for fractions 22 and 27 using heparinases II, and III. The results, presented in Figure 6, can be taken with the tandem MS data of the intact hexasaccharides (which localized the N-acetyl group to the reducing end) and the disaccharide compositions to generate linear sequences for each hexasaccharide, however devoid of knowledge of uronic acid identity. Interestingly the hexasaccharides in both fractions showed identical cleavage patterns (Figure 6). Heparinase II cleaved both hexasaccharides generating products at m/z 207.5 (2− charge state), 247.5 (2− charge state), and 291.1 (3− charge state, Figure 6 A and B). These mass to charge ratios deconvolute to molecular masses of 417.1, 497.5, and 876.2 Da respectively. The first two masses correspond to the disaccharides IV-S and III-S identified in the compositional analysis, but provide no new information. However, the mass of 876.2 Da corresponds to a tetrasaccharide (with 2 sulfates and 1 acetyl group) linking the III-S and IV-A disaccharides (Mtheo 876.13 Da). Since the MS2 data of the intact hexasaccharides (Figure 4) positioned the acetyl group at the reducing end, these corresponding tetrasaccharides can be sequenced as ΔUA2S-GlcNS-HexA-GlcNAc (HexA is used to indicate a lack of knowledge of C5 stereochemistry indicating either GlcA or IdoA). Meanwhile, heparinase III cleavage also generated identical cleavages between the two fractions producing ions at m/z 227.5 (4− charge state) and 303.7 (3− charge state), which both deconvolute to a molecular weight of 914.1 Da (Figure 6 C and D). This ion is indicative of a tetrasaccharide (with 3 sulfate groups and no acetyl groups) linking IV-S and III-S disaccharides (Mtheo 914.07 Da). Since the Heparinase II and MS2 data localized the HexA2S-GlcNS and HexA-GlcNAc as the central and reducing end disaccharides, respectively, the heparinase III data confirmed the sequence ΔUA-GlcNS-HexA2S-GlcNS and localized the IV−S disaccharide to the non-reducing end. Thus hexasaccharides in fraction 22 and 27 both possess linear sequences of ΔUA-GlcNS-HexA2S-GlcNS-HexA-GlcNAc. One-dimensional (1-D) 1H-NMR was subsequently used to define the uronic acid stereochemistries.
Figure 6.
Partial enzymatic digestions of HS hexasaccharides. Fractions 22 (A) and 27 (B) were treated with heparinase II. Labeled peaks at m/z 207.5, 247.5, and 291.1 correspond to the sequences presented (see text for descriptions). Fractions 22 (C) and 27 (D) were also treated with heparinase III. Peaks at m/z 227.5 and 303.7 correspond to the 4− and 3− charge states of the sequence presented. HexA denotes an uronic acid, but lack of knowledge as to whether it is GlcA or IdoA.
1-D 1H-NMR Analysis and Assignment of Uronic Acid Residues
Aliquots of fractions 22 and 27 in deionized H2O were lyophilized via freeze-drying overnight. The following morning the samples were dissolved in 99.9% D2O to remove exchangeable protons and the samples were subjected to 600 MHz 1H-NMR analysis. As for most oligosaccharides, the majority of the ring proton resonances cluster in a congested, narrow spectral range between 3.2 and 4.0 ppm (data not shown) 73–78. The data outside this range was interpreted in accordance with published 1H chemical shifts of protons in similar chemical environments to those of our saccharides 51, 52, 79–83. All signals resonating above 4 ppm were identified (Figure 7), the majority of which correspond to the anomeric protons of the individual sugar residues which are shifted downfield due to unshielding by the ring oxygen atom 78. Of note is that in both spectra of fractions 22 (Figure 7A) and 27 (Figure 7B) resonances corresponding to a 2-O-sulfated IdoA are observed. Peaks c (5.23 ppm), g (4.36 ppm), i (4.27 ppm), and j (4.08 ppm) correspond to IdoA2S H1, H2, H3, and H4 respectively 51, 52, 79, 82, 84. Given that both isomers contained one 2-O-sulfated uronic acid (the central HexA2S), these data identify the central uronic acid as IdoA2S in both hexasaccharides. Also of note is that only the spectrum of hexasaccharide-22 showed resonances for an unsulfated iduronic acid. Peaks labeled u (5.39 ppm), v (5.02 ppm), w (4.78 ppm), and x (4.15 ppm) correspond to GlcNS H1 (when linked to IdoA), IdoA H1, IdoA H5, and IdoA H3 respectively 11, 79, 80, 85. Meanwhile the spectrum of hexasaccharide-27 displays resonances unique to a saccharide containing GlcA. Peaks labeled y (5.60 ppm) and z (4.60 ppm) corresponds to GlcNS H1 (when linked to GlcA), and GlcA H1 respectively 79, 80, 82, 83. Summarizing, the common internal HexA2S in both hexasaccharide-22 and -27 is identified as IdoA2S. Conversely the uronic acid in the reducing end disaccharide is IdoA in hexasaccharide-22 and GlcA in hexasaccharide-27. All other common peaks in the reporter group region were consistent with our MS-derived structures including ΔUA H4 (peak a 5.80 ppm), GlcNS H1 (when linked to IdoA2S; peak b, 5.36, ppm), GlcNAcα H1 (d, 5.23 ppm), ΔUA H1 (e, 5.12 ppm), GlcNAcβ H1 (f, 4.73 ppm), and ΔUA H3 (h, 4.32 ppm) 79, 80, 82, 84. Thus by combining the MS data of the intact hexasaccharides, the disaccharide compositional analysis, partial enzymatic digestions, and finally 1-D 1H-NMR we were able to define the purified structures in fractions 22 and 27 as ΔUA-GlcNS-IdoA2S-GlcNS-IdoA-GlcNAc, and ΔUA-GlcNS-IdoA2S-GlcNS-GlcA-GlcNAc, respectively (inset Figure 7). In defining the full sequence for both hexasaccharides, it is interesting to note that the enzymatic cleavage product, ΔUA-GlcNS-IdoA2S-GlcNS, obtained from the hexasaccharide in fraction 22 (Figure 6C) may not have been predicted based on some published accounts of heparinase III specificity 86. Nevertheless, the use of all three heparinases (regardless of specificity), in addition to the all the supporting experiments, indicates that the two derived sequences for fractions 22 and 27 are the only reasonable possibilities. Thus, our results are consistent with studies suggesting that the GlcNS-IdoA linkage is, in fact, a viable target for the heparinase III enzyme 59, 87.
Figure 7.
1-D 1H-NMR spectra for fractions 22 (A) and 27 (B). Common peaks are labeled as a ΔUAH4; b, GlcNS H1 (to Ido2S); c, IdoA2S H1; d, GlcNAcα H1; e, ΔUA H1; f, GlcNAcβ H1; g, IdoA2S H2; h, ΔUA H3; i, IdoA2S H3; and j, IdoA2S H4. Unique peaks are labeled as u, GlcNS H1 (to IdoA); v, IdoA H1; w, IdoA H5; x, IdoA H3; y, GlcNS H1 (to GlcA); and z, GlcA H1. The derived sequences for the hexasaccharide in each fraction, based combined data, are presented (inset).
In identifying the complete structure of two highly specific hexasaccharide structural isomers, we have shown that MS and ion mobility can be powerful tools in the initial screening process of complex libraries of biologically derived GAG. IMS was capable of differentiating HS isomers with changes in the axial vs. equatorial positioning of a single carboxylate functional group. Thus, this methodology may be useful in differentiating GAG structures for which there is no knowledge uronic acid chemistry, a priori. Research from other groups has shown that MS can be used to differentiate uronic acid residue isomers when conditions are optimized from known starting structures 67, 88, although the MS2 spectra in these cases are quite complicated. IMS has been applied previously for GAG analysis, however saccharides differing in multiple chemical modifications were investigated making specific structural interpretation difficult 89. Given the data presented herein, one might envision a role for IMS-MS in the rapid, complete analysis HS oligosaccharides based on a combination of experiment and theory without prior knowledge. In such a scenario, MS-based methods, like those described in this manuscript, would supply the majority of the structural information, while molecular modeling methods 89 would correlate variations in uronic acid stereochemistry with the collision cross sections obtained by IMS. Applications such as this would represent a significant advance in HS-GAG analysis.
Conclusions
Given the vast array of HS-interacting proteins and the number of physiologic and pathophysiologic processes in which those interactions are significant, new methods to define structural motifs within HS should continue to be developed. In recent years the first therapeutic based on specific knowledge of a HS-protein interaction emerged 14, 90, 91, and more are expected 92. Knowledge of HS structure will undoubtedly contribute to these developments. In the present study we applied a combination of chromatography, MS, and 1-D 1H-NMR techniques to sequence two isomeric HS hexasaccharides. The method developed herein will be amenable to the analysis of longer and more complex saccharides as they are purified, and will thus facilitate additional HS sequence determinations. We also showed that IMS was capable of resolving HS isomers which differed at only a single stereogenic center (C5 of uronic acids). Given their abilities to rapidly distinguish closely related species, MS and IMS are expected play increasingly important roles in GAG analysis and structural elucidation.
Supplementary Material
Acknowledgements
We gratefully acknowledge Professor Jian Liu, for the gracious gift of the heparinase III, and Kevin Giles and Iain Campuzano of Waters Corporation for running the hexasaccharides on the Synapt G2™ instrument. We also acknowledge NIH grants GM47356, EY012347, and RR11973 for supporting this research.
ABBREVIATIONS
- HS
heparan sulfate
- MS
mass spectrometry
- NMR
nuclear magnetic resonance
- MS2
tandem spectrometry
- IMS
ion mobility spectrometry
- GlcA
glucuronic acid
- IdoA
iduronic acid
- HexA
either glucuronic or iduronic acid
- GAGs
glycosaminoglycans
- GlcNAc
N-acetylglucosamine
- GalNAc
N-acetylgalactosamine
- TOF
time-of-flight
- NH4OAc
ammonium acetate
- ESI
electrospray ionization
- nESI
nanoelectrospray ionization
- dp
degree of polymerization (dp2 = disaccharide, etc.)
- SEC
size exclusion chromatography
- SAX
strong anion exchange chromatography
- MWCO
molecular weight cut-off
- Ca(OAc)2
calcium acetate
- DSS
2,2-dimethyl-2-silapentane-5-sulfonic acid
- ATDs
arrival time distributions
References
- 1.Casu B, Lindahl U. Adv Carbohydr Chem Biochem. 2001;57:159–206. doi: 10.1016/s0065-2318(01)57017-1. [DOI] [PubMed] [Google Scholar]
- 2.Conrad HE. Heparin-binding proteins. San Diego: Academic Press; 1998. [Google Scholar]
- 3.Linhardt RJ, Toida T. Acc Chem Res. 2004;37:431–438. doi: 10.1021/ar030138x. [DOI] [PubMed] [Google Scholar]
- 4.Raman R, Sasisekharan V, Sasisekharan R. Chem Biol. 2005;12:267–277. doi: 10.1016/j.chembiol.2004.11.020. [DOI] [PubMed] [Google Scholar]
- 5.Varki A, Cummings RD, Esko JD, Freeze HH, Stanley P, Bertozzi CR, Hart GW, Etzler ME, editors. Essentials of Glycobiology. 2nd ed. Cold Spring Harbor: Cold Spring Harbor Laboratory Press; 2009. [PubMed] [Google Scholar]
- 6.Esko JD, Lindahl U. J Clin Invest. 2001;108:169–173. doi: 10.1172/JCI13530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Esko JD, Selleck SB. Annu Rev Biochem. 2002;71:435–471. doi: 10.1146/annurev.biochem.71.110601.135458. [DOI] [PubMed] [Google Scholar]
- 8.Lindahl U, Kusche-Gullberg M, Kjellen L. J Biol Chem. 1998;273:24979–24982. doi: 10.1074/jbc.273.39.24979. [DOI] [PubMed] [Google Scholar]
- 9.Sasisekharan R, Venkataraman G. Curr Opin Chem Biol. 2000;4:626–631. doi: 10.1016/s1367-5931(00)00145-9. [DOI] [PubMed] [Google Scholar]
- 10.Stringer SE, Gallagher JT. Int J Biochem Cell Biol. 1997;29:709–714. doi: 10.1016/s1357-2725(96)00170-7. [DOI] [PubMed] [Google Scholar]
- 11.Griffin CC, Linhardt RJ, Van Gorp CL, Toida T, Hileman RE, Schubert RL, Brown SE. Carbohydrate Research. 1995;276:183–197. doi: 10.1016/0008-6215(95)00166-q. [DOI] [PubMed] [Google Scholar]
- 12.Toida T, Yoshida H, Toyoda H, Koshiishi I, Imanari T, Hileman RE, Fromm JR, Linhardt RJ. Biochem J. 1997;322(Pt 2):499–506. doi: 10.1042/bj3220499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Vongchan P, Warda M, Toyoda H, Toida T, Marks RM, Linhardt RJ. Biochimica et Biophysica Acta (BBA) - General Subjects. 2005;1721:1–8. doi: 10.1016/j.bbagen.2004.09.007. [DOI] [PubMed] [Google Scholar]
- 14.Bauer KA, Hawkins DW, Peters PC, Petitou M, Herbert JM, van Boeckel CA, Meuleman DG. Cardiovasc Drug Rev. 2002;20:37–52. doi: 10.1111/j.1527-3466.2002.tb00081.x. [DOI] [PubMed] [Google Scholar]
- 15.Choay J, Petitou M, Lormeau JC, Sinay P, Casu B, Gatti G. Biochem Biophys Res Commun. 1983;116:492–499. doi: 10.1016/0006-291x(83)90550-8. [DOI] [PubMed] [Google Scholar]
- 16.Thunberg L, Backstrom G, Lindahl U. Carbohydr Res. 1982;100:393–410. doi: 10.1016/s0008-6215(00)81050-2. [DOI] [PubMed] [Google Scholar]
- 17.Faham S, Hileman RE, Fromm JR, Linhardt RJ, Rees DC. Science. 1996;271:1116–1120. doi: 10.1126/science.271.5252.1116. [DOI] [PubMed] [Google Scholar]
- 18.Guglieri S, Hricovini M, Raman R, Polito L, Torri G, Casu B, Sasisekharan R, Guerrini M. Biochemistry. 2008 doi: 10.1021/bi801007p. [DOI] [PubMed] [Google Scholar]
- 19.Maccarana M, Casu B, Lindahl U. J Biol Chem. 1993;268:23898–23905. [PubMed] [Google Scholar]
- 20.Plotnikov AN, Schlessinger J, Hubbard SR, Mohammadi M. Cell. 1999;98:641–650. doi: 10.1016/s0092-8674(00)80051-3. [DOI] [PubMed] [Google Scholar]
- 21.Thompson SM, Connell MG, Fernig DG, Ten Dam GB, van Kuppevelt TH, Turnbull JE, Jesudason EC, Losty PD. Pediatr Surg Int. 2007;23:411–417. doi: 10.1007/s00383-006-1864-8. [DOI] [PubMed] [Google Scholar]
- 22.Jin L, Abrahams JP, Skinner R, Petitou M, Pike RN, Carrell RW. Proc Natl Acad Sci U S A. 1997;94:14683–14688. doi: 10.1073/pnas.94.26.14683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Petitou M, Duchaussoy P, Driguez PA, Herault JP, Lormeau JC, Herbert JM. Bioorg Med Chem Lett. 1999;9:1155–1160. doi: 10.1016/s0960-894x(99)00155-9. [DOI] [PubMed] [Google Scholar]
- 24.Petitou M, Herault JP, Bernat A, Driguez PA, Duchaussoy P, Lormeau JC, Herbert JM. Nature. 1999;398:417–422. doi: 10.1038/18877. [DOI] [PubMed] [Google Scholar]
- 25.Barth H, Schafer C, Adah MI, Zhang F, Linhardt RJ, Toyoda H, Kinoshita-Toyoda A, Toida T, Van Kuppevelt TH, Depla E, Von Weizsacker F, Blum HE, Baumert TF. J Biol Chem. 2003;278:41003–41012. doi: 10.1074/jbc.M302267200. [DOI] [PubMed] [Google Scholar]
- 26.Chen Y, Maguire T, Hileman RE, Fromm JR, Esko JD, Linhardt RJ, Marks RM. Nat Med. 1997;3:866–871. doi: 10.1038/nm0897-866. [DOI] [PubMed] [Google Scholar]
- 27.WuDunn D, Spear PG. J Virol. 1989;63:52–58. doi: 10.1128/jvi.63.1.52-58.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Borsig L, Wong R, Hynes RO, Varki NM, Varki A. Proc Natl Acad Sci U S A. 2002;99:2193–2198. doi: 10.1073/pnas.261704098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liu D, Shriver Z, Venkataraman G, El Shabrawi Y, Sasisekharan R. Proc Natl Acad Sci U S A. 2002;99:568–573. doi: 10.1073/pnas.012578299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sasisekharan R, Moses MA, Nugent MA, Cooney CL, Langer R. Proc Natl Acad Sci U S A. 1994;91:1524–1528. doi: 10.1073/pnas.91.4.1524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sasisekharan R, Shriver Z, Venkataraman G, Narayanasami U. Nat Rev Cancer. 2002;2:521–528. doi: 10.1038/nrc842. [DOI] [PubMed] [Google Scholar]
- 32.Varki NM, Varki A. Semin Thromb Hemost. 2002;28:53–66. doi: 10.1055/s-2002-20564. [DOI] [PubMed] [Google Scholar]
- 33.Vlodavsky I, Friedmann Y, Elkin M, Aingorn H, Atzmon R, Ishai-Michaeli R, Bitan M, Pappo O, Peretz T, Michal I, Spector L, Pecker I. Nat Med. 1999;5:793–802. doi: 10.1038/10518. [DOI] [PubMed] [Google Scholar]
- 34.Middleton J, Neil S, Wintle J, Clark-Lewis I, Moore H, Lam C, Auer M, Hub E, Rot A. Cell. 1997;91:385–395. doi: 10.1016/s0092-8674(00)80422-5. [DOI] [PubMed] [Google Scholar]
- 35.Parish CR. Nat Rev Immunol. 2006;6:633–643. doi: 10.1038/nri1918. [DOI] [PubMed] [Google Scholar]
- 36.Proudfoot AE, Handel TM, Johnson Z, Lau EK, LiWang P, Clark-Lewis I, Borlat F, Wells TN, Kosco-Vilbois MH. Proc Natl Acad Sci U S A. 2003;100:1885–1890. doi: 10.1073/pnas.0334864100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Shamri R, Grabovsky V, Gauguet JM, Feigelson S, Manevich E, Kolanus W, Robinson MK, Staunton DE, von Andrian UH, Alon R. Nat Immunol. 2005;6:497–506. doi: 10.1038/ni1194. [DOI] [PubMed] [Google Scholar]
- 38.Wang L, Fuster M, Sriramarao P, Esko JD. Nat Immunol. 2005;6:902–910. doi: 10.1038/ni1233. [DOI] [PubMed] [Google Scholar]
- 39.Webb LM, Ehrengruber MU, Clark-Lewis I, Baggiolini M, Rot A. Proc Natl Acad Sci U S A. 1993;90:7158–7162. doi: 10.1073/pnas.90.15.7158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Belenkaya TY, Han C, Yan D, Opoka RJ, Khodoun M, Liu H, Lin X. Cell. 2004;119:231–244. doi: 10.1016/j.cell.2004.09.031. [DOI] [PubMed] [Google Scholar]
- 41.Bellaiche Y, The I, Perrimon N. Nature. 1998;394:85–88. doi: 10.1038/27932. [DOI] [PubMed] [Google Scholar]
- 42.Hacker U, Nybakken K, Perrimon N. Nat Rev Mol Cell Biol. 2005;6:530–541. doi: 10.1038/nrm1681. [DOI] [PubMed] [Google Scholar]
- 43.Rapraeger AC, Krufka A, Olwin BB. Science. 1991;252:1705–1708. doi: 10.1126/science.1646484. [DOI] [PubMed] [Google Scholar]
- 44.Tsuda M, Kamimura K, Nakato H, Archer M, Staatz W, Fox B, Humphrey M, Olson S, Futch T, Kaluza V, Siegfried E, Stam L, Selleck SB. Nature. 1999;400:276–280. doi: 10.1038/22336. [DOI] [PubMed] [Google Scholar]
- 45.Yayon A, Klagsbrun M, Esko JD, Leder P, Ornitz DM. Cell. 1991;64:841–848. doi: 10.1016/0092-8674(91)90512-w. [DOI] [PubMed] [Google Scholar]
- 46.Capila I, Linhardt RJ. Angew Chem Int Ed Engl. 2002;41:391–412. doi: 10.1002/1521-3773(20020201)41:3<390::aid-anie390>3.0.co;2-b. [DOI] [PubMed] [Google Scholar]
- 47.Noti C, Seeberger PH. Chemistry & Biology. 2005;12:731–756. doi: 10.1016/j.chembiol.2005.05.013. [DOI] [PubMed] [Google Scholar]
- 48.Lyon M, Gallagher JT. Matrix Biol. 1998;17:485–493. doi: 10.1016/s0945-053x(98)90096-8. [DOI] [PubMed] [Google Scholar]
- 49.Merry CL, Lyon M, Deakin JA, Hopwood JJ, Gallagher JT. J Biol Chem. 1999;274:18455–18462. doi: 10.1074/jbc.274.26.18455. [DOI] [PubMed] [Google Scholar]
- 50.Stringer SE, Kandola BS, Pye DA, Gallagher JT. Glycobiology. 2003;13:97–107. doi: 10.1093/glycob/cwg006. [DOI] [PubMed] [Google Scholar]
- 51.Chuang W-L, Christ MD, Rabenstein DL. Analytical Chemistry. 2001;73:2310–2316. doi: 10.1021/ac0100291. [DOI] [PubMed] [Google Scholar]
- 52.Pervin A, Gallo C, Jandik KA, Han X-J, Linhardt RJ. Glycobiology. 1995;5:83–95. doi: 10.1093/glycob/5.1.83. [DOI] [PubMed] [Google Scholar]
- 53.Liu J, Desai UR, Han XJ, Toida T, Linhardt RJ. Glycobiology. 1995;5:765–774. doi: 10.1093/glycob/5.8.765. [DOI] [PubMed] [Google Scholar]
- 54.Vives RR, Pye DA, Salmivirta M, Hopwood JJ, Lindahl U, Gallagher JT. Biochem J. 1999;339(Pt 3):767–773. [PMC free article] [PubMed] [Google Scholar]
- 55.Murphy KJ, McLay N, Pye DA. Journal of the American Chemical Society. 2008;130:12435–12444. doi: 10.1021/ja802863p. [DOI] [PubMed] [Google Scholar]
- 56.Saad OM, Leary JA. Anal Chem. 2003;75:2985–2995. doi: 10.1021/ac0340455. [DOI] [PubMed] [Google Scholar]
- 57.Saad OM, Ebel H, Uchimura K, Rosen SD, Bertozzi CR, Leary JA. Glycobiology. 2005;15:818–826. doi: 10.1093/glycob/cwi064. [DOI] [PubMed] [Google Scholar]
- 58.Schenauer MR, Yu Y, Sweeney MD, Leary JA. Journal of Biological Chemistry. 2007;282:25182–25188. doi: 10.1074/jbc.M703387200. [DOI] [PubMed] [Google Scholar]
- 59.Desai UR, Wang HM, Linhardt RJ. Biochemistry. 1993;32:8140–8145. doi: 10.1021/bi00083a012. [DOI] [PubMed] [Google Scholar]
- 60.Linhardt RJ, Galliher PM, Cooney CL. Appl Biochem Biotechnol. 1986;12:135–176. doi: 10.1007/BF02798420. [DOI] [PubMed] [Google Scholar]
- 61.Yamagata T, Saito H, Habuchi O, Suzuki SJ. Biol. Chem. 1968;243:1523–1535. [PubMed] [Google Scholar]
- 62.Weiss J. Handbook of Ion Chromatography. (3rd ed.) 2005 [Google Scholar]
- 63.Atkins ED, Nieduszynski IA. Fed Proc. 1977;36:78–83. [PubMed] [Google Scholar]
- 64.Casu B, Choay J, Ferro DR, Gatti G, Jacquinet JC, Petitou M, Provasoli A, Ragazzi M, Sinay P, Torri G. Nature. 1986;322:215–216. doi: 10.1038/322215b0. [DOI] [PubMed] [Google Scholar]
- 65.Mulloy B, Forster MJ, Jones C, Davies DB. Biochem J. 1993;293(Pt 3):849–858. doi: 10.1042/bj2930849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Saad OM, Leary JA. Anal Chem. 2005;77:5902–5911. doi: 10.1021/ac050793d. [DOI] [PubMed] [Google Scholar]
- 67.Zaia J, Li XQ, Chan SY, Costello CE. J Am Soc Mass Spectrom. 2003;14:1270–1281. doi: 10.1016/S1044-0305(03)00541-5. [DOI] [PubMed] [Google Scholar]
- 68.Domon B, Costello CE. Glycoconjugate Journal. 1988;5:397–409. [Google Scholar]
- 69.Saad OM, Leary JA. J Am Soc Mass Spectrom. 2004;15:1274–1286. doi: 10.1016/j.jasms.2004.05.008. [DOI] [PubMed] [Google Scholar]
- 70.Clemmer DE, Jarrold MF. Journal of Mass Spectrometry. 1997;32:577–592. [Google Scholar]
- 71.Jarrold MF. Annual Review of Physical Chemistry. 2000;51:179–207. doi: 10.1146/annurev.physchem.51.1.179. [DOI] [PubMed] [Google Scholar]
- 72.Wyttenbach T, Bowers MT. Modern Mass Spectrometry. 2003;Vol. 225:207–232. [Google Scholar]
- 73.Bubb WA. Concepts in Magnetic Resonance, Part A. 2003;19A:1–19. [Google Scholar]
- 74.Duus JO, Gotfredsen CH, Bock K. Chemical Reviews. 2000;100:4589–4614. doi: 10.1021/cr990302n. [DOI] [PubMed] [Google Scholar]
- 75.Jimenez-Barbero J, Peter T, editors. NMR Spectroscopy of Glycoconjugates. Weinheim: Wiley-VCH; 2003. [Google Scholar]
- 76.Mulloy B. Molecular Biotechnology. 1996;6:241–265. doi: 10.1007/BF02761706. [DOI] [PubMed] [Google Scholar]
- 77.Vliegenthart JFG, van Halbeek H, Dorland L. Pure and Applied Chemistry. 1981;53:45–77. [Google Scholar]
- 78.Vliegenthart JFG, Woods RJ, editors. NMR Spectroscopy and Computer Modeling of Carbohydrates. New York: Oxford University Press; 2006. [Google Scholar]
- 79.Hileman RE, Smith AE, Toida T, Linhardt RJ. Glycobiology. 1997;7:231–239. doi: 10.1093/glycob/7.2.231. [DOI] [PubMed] [Google Scholar]
- 80.Sugahara K, Tohno-oka R, Yamada S, Khoo K-H, Morris HR, Dell A. Glycobiology. 1994;4:535–544. doi: 10.1093/glycob/4.4.535. [DOI] [PubMed] [Google Scholar]
- 81.Yamada S, Murakami T, Tsuda H, Yoshida K, Sugahara K. J. Biol. Chem. 1995;270:8696–8705. [PubMed] [Google Scholar]
- 82.Yamada S, Sakamoto K, Tsuda H, Yoshida K, Sugahara K, Khoo K-H, Morris HR, Dell A. Glycobiology. 1994;4:69–78. doi: 10.1093/glycob/4.1.69. [DOI] [PubMed] [Google Scholar]
- 83.Yamada S, Sakamoto K, Tsuda H, Yoshida K, Sugiura M, Sugahara K. Biochemistry. 1999;38:838–847. doi: 10.1021/bi981889n. [DOI] [PubMed] [Google Scholar]
- 84.Yamada S, Murakami T, Tsuda H, Yoshida K, Sugahara K. J Biol Chem. 1995;270:8696–8705. [PubMed] [Google Scholar]
- 85.Sugahara K, Tsuda H, Yoshida K, Yamada S, de Beer T, Vliegenthart JF. J Biol Chem. 1995;270:22914–22923. doi: 10.1074/jbc.270.39.22914. [DOI] [PubMed] [Google Scholar]
- 86.Linhardt RJ, Turnbull JE, Wang HM, Loganathan D, Gallagher JT. Biochemistry. 1990;29:2611–2617. doi: 10.1021/bi00462a026. [DOI] [PubMed] [Google Scholar]
- 87.Desai UR, Wang HM, Kelly TR, Linhardt RJ. Carbohydr Res. 1993;241:249–259. doi: 10.1016/0008-6215(93)80112-r. [DOI] [PubMed] [Google Scholar]
- 88.Wolff JJ, Chi L, Linhardt RJ, Amster IJ. Anal Chem. 2007;79:2015–2022. doi: 10.1021/ac061636x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Jin L, Barran PE, Deakin JA, Lyon M, Uhrin D. Physical Chemistry Chemical Physics. 2005;7:3464–3471. doi: 10.1039/b508644b. [DOI] [PubMed] [Google Scholar]
- 90.Petitou M, Duchaussoy P, Herbert JM, Duc G, El Hajji M, Branellec JF, Donat F, Necciari J, Cariou R, Bouthier J, Garrigou E. Semin Thromb Hemost. 2002;28:393–402. doi: 10.1055/s-2002-34309. [DOI] [PubMed] [Google Scholar]
- 91.Petitou M, van Boeckel CA. Angew Chem Int Ed Engl. 2004;43:3118–3133. doi: 10.1002/anie.200300640. [DOI] [PubMed] [Google Scholar]
- 92.Lindahl U. Thromb Haemost. 2007;98:109–115. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.