Abstract
Transcription elongation in vitro is affected by the interactions between RNA polymerase (RNAP) subunits and the nucleic acid scaffold of the ternary elongation complex (TEC, RNAP-DNA–RNA). We have investigated the role of the RNAP subunits F/E (homologous to eukaryotic RPB4/7) during transcription elongation and termination using a wholly recombinant archaeal RNAP and synthetic nucleic acid scaffolds. The F/E complex greatly stimulates the processivity of RNAP, it enhances the formation of full length products, reduces pausing, and increases transcription termination facilitated by weak termination signals. Mutant variants of F/E that are defective in RNA binding show that these activities correlate with the nucleic acid binding properties of F/E. However, a second RNA-binding independent component also contributes to the stimulatory activities of F/E. In summary, our results suggest that interactions between RNAP subunits F/E and the RNA transcript are pivotal to the molecular mechanisms of RNAP during transcription elongation and termination.
INTRODUCTION
Evolutionary related multi-subunit RNA polymerases (RNAP) facilitate DNA-dependent RNA transcription in all three domains of life (1). Transcription is often likened to a cycle where RNAPs initiates at the promoter, elongate through the gene, terminate at its 3′-end, and reinitiate at the promoter again. During the transcription cycle RNAPs are undergoing substantial conformational changes and distinct interactions are made between RNAP subunits and the promoter DNA during transcription initiation, and the DNA–RNA scaffold during transcription elongation. The overall structural layout of all multi-subunit RNAPs is conserved. However, the archaeal RNAP and eukaryotic RNAPII share a subset of subunits that are not present in the bacterial enzyme (2). The most prominent structural feature that discriminates between all bacterial and archaeal/eukaryotic enzymes is a stalk-like protrusion formed by a heterodimeric complex of RNAP subunits F/E (RPB4/7 in RNAPII). The F/E complex is stably associated with the RNAP core and binds RNA in vitro and in vivo (3–5). During transcription initiation F/E, in conjunction with the basal transcription factor TFE, is involved in DNA melting (6,7). Much research has been devoted to the understanding of the mechanisms that regulate transcription initiation, which traditionally was assumed to be the main or even sole step of transcription regulation and gene expression. Recently it has become evident that the elongation phase is another important target for the control of transcription (8). Transcription elongation is discontinuous and frequently interrupted by pausing, stalling and arrest events that are determined by the sequence of the template and can be subject to regulation by elongation factors (9). In bacteria, transcriptional pausing plays a fundamental role in the coupling of transcription and translation by allowing ribosomes to catch up with elongating RNAPs, which otherwise would be prematurely terminated by the rho factor (9). Transcription pausing allows the formation of functional secondary structures in the transcript (e.g. in ribosomal RNA), facilitates the recruitment of transcription factors (e.g. bacterial RfaH) and is a prerequisite for regulated intrinsic and rho-dependent termination in bacteria (9). The termination of transcription in the Archaea bears some resemblance to the RNAPIII system (10,11); it is independent of RNA secondary structures, such as RNA hairpins characteristic of bacterial intrinsic terminators, and exogenous transcription termination factors including the bacterial rho helicase and the eukaryotic Xrn2/Rat1 exonucleases (12–15). In eukaryotes, promoter-proximal pausing of RNAPII at ∼20–50 bases downstream of the transcription start site is a regulatory mechanism that allows rapid induction of gene expression because it does not depend on recruitment cascades of transcription factors and RNAPs. This phenomenon was initially discovered at Drosophila heat shock promoters but has subsequently been proven to be a widespread mode of regulation from yeast to man (16–18). The sequence dependent pausing of all types of RNAPs is facilitated by interactions between the RNAP and DNA template and the RNA transcript. Structural information obtained from RNAPII elongation complexes has illuminated the interactions between the downstream DNA duplex and the RNAP jaw domains, and the interactions between the DNA–RNA hybrid and the RNAP hybrid compartment (19–21). However, it has not been possible to solve the structures of the RNA transcript and the nontemplate strand (NTS) in the context of the TEC. Both the NTS and the RNA transcript are components of the protein-nucleic acid interaction network of the elongation complex, and thereby have the potential to influence transcription elongation and termination (22).
We are using a wholly recombinant hyperthermophilic archaeal in vitro transcription system to investigate the molecular mechanisms of transcription (7,23,24). This system is a genuine model system for eukaryotic RNAPII and allows us to study the functional contributions of RNAP subunits such as the F/E complex (RPB4/7) (2,3). The tractability of the RNAPII system is limited by its biochemical properties, thus it has not been possible to produce recombinant RNAPII. In addition, archaeal transcription systems allow us to uncover basic RNAP functions that are often masked by the complexity of the eukaryotic system (6,7,25).
Here we characterize the molecular mechanisms of archaeal RNAP downstream of the initiation phase, during elongation and termination of transcription. We have investigated the role of the NTS during the elongation and termination phases of transcription and characterized the functional contribution of RNAP subunits F/E during these two processes. We have discovered that transcript binding to F/E enhances the processivity and decreases transcriptional pausing, but also that the F/E complex improves the efficiency of transcription termination on weak termination signals. The modus operandi of F/E is comprised of two components, one of which depends on the RNA-binding activity of F/E. The second mechanism is likely to involve conformational changes within RNAP such as the closure of the clamp domain (26).
EXPERIMENTAL PROCEDURES
Recombinant proteins
Methanocaldococcus jannaschii RNAPs were prepared as described previously (3,4,23).
Transcription elongation assay
The template DNA strand (TS) and the 14 nucleotide (nt) RNA primer were annealed in a 1 : 5 molar ratio for 1 min at 96°C and slowly cooled to ambient temperature. In total, 50–200 nM RNAP were preincubated with 13/2.5 mM RNA/DNA scaffold at 65°C in 0.5× HNME buffer (20 mM HEPES, pH 7.3, 125 mM sodium chloride, 1.75 mM magnesium chloride, 0.05 mM EDTA, 2.5% glycerol, and 10 mM DTT). To prevent nonspecific binding of RNAP to nucleic acid scaffolds and multiround transcription by reinitiation the reaction mix was incubated for further 10 min in presence of 0.05 mg/ml Heparin at 65°C. Transcription reaction was started by addition of nucleotides (750 µM ATP/UTP/GTP, 2 µM CP, [α-32P] CTP (5000 Ci/mmol), in presence or absence of 2.5 mM NTS. The radiolabelled transcripts were separated on 16% denaturating PAGE (8M Urea, 0.5× TBE), dried for 1 h at 80°C under vacuum and visualized using an Fuji FLA 2000 reader and Image Analyser software (Fuji, Japan).
Transcript release assay
Transcription reactions were carried out as described above, the reaction was stopped by addition of 50 mM EDTA and the transcripts were separated by 10% native PAGE (Acrylamide : Bisacrylamide=19 : 1).
Electrophoretic mobility shift assay
Electrophoretic mobility shift assays (EMSAs) were performed using a 32P-labelled A2-riboprobe (4). The A2 probe was generated by in vitro transcription using the Ambion T7 transcription kit using [α-32P] UTP (5000 Ci/mmol) as tracer nucleotide. The 32P-labeled A2 riboprobe was incubated with increasing amounts of recombinant F/E variants in HNME buffer (40 mM HEPES, pH 7.3, 250 mM sodium chloride, 2.5 mM magnesium chloride, 0.1 mM EDTA, 5 % glycerol, and 10 mM dithiothreitol) containing 1 μg bovine serum albumin in a total volume of 15 μl for 20 min at 65°C. The reaction mixture was transferred into native Tris-glycine loading buffer and loaded onto a native 4–20 % Tris-glycine gradient gel (Novex, Invitrogen). The complexes were electrophoretically separated at room temperature for 1 h at 180 V, the gel was dried for 2 h at 80°C under vacuum and subjected to autoradiography (Kodak Biomax MR film) and/or phospho imager analysis (Fuji FLA 2000).
RESULTS
The nontemplate DNA strand stimulates the processivity of RNAP
In order to characterize the molecular mechanisms of transcription elongation under rigorously controlled conditions and in a promoter-independent manner, we adapted a transcription assay that utilizes synthetic elongation scaffolds consisting of custom synthesized DNA and RNA oligonucleotides (Figure 1A and B). Figure 2 shows two typical time course experiments (30 s to 60 min) that compare the transcript pattern generated by recombinant M. jannaschii RNAP on two distinct synthetic elongation scaffolds that differ with respect to the presence of the NTS. In the first experiment RNAP utilizes a template strand (TS) DNA:RNA primer scaffold (Figure 2A, left) and in the second experiment a TS/NTS DNA:RNA scaffold (Figure 2A, right). The recombinant archaeal RNAP can utilize synthetic elongation scaffolds independently of the NTS, similar to bacteriophage T7 RNAP, bacterial RNAP and eukaryotic RNAPII (Figure 2) (27,28). Under the chosen assay conditions the NTS affects transcription elongation both quantitatively and qualitatively. Wild type RNAP synthesizes fewer partial- and more runoff transcripts (71 nt), and the runoff transcript is produced earlier in the presence of the NTS (Figure 2B and C, 2 min instead of 5 min). This demonstrates that the processivity is improved by the presence of the NTS, which is likely due to an increased stability of the TEC, i.e. improved ‘traction’ of RNAP on the template. Under our experimental conditions the earliest full-length runoff transcript appears after 2 min suggesting an in vitro elongation rate of approximately 35 nt/min. This is in good agreement with the in vitro transcription elongation rate of eukaryotic RNAPII, but it is also significantly slower than elongation rates estimated in vivo (∼10–25 nt/s) (29,30). In the absence of the NTS the first runoff transcript is generated after 5 min, which suggests that the elongation rate is about two fold slower without the NTS (Figure 2A–C). However, under these conditions RNAP is also paused for a longer time period (Figure 2A, ∼23 and 33 nt) and it is therefore problematic to clearly distinguish an increased transcription elongation rate from improved processivity (Figure 2A). The total amount of radio-labelled transcripts synthesized is only marginally increased by the NTS (Figure 2C).
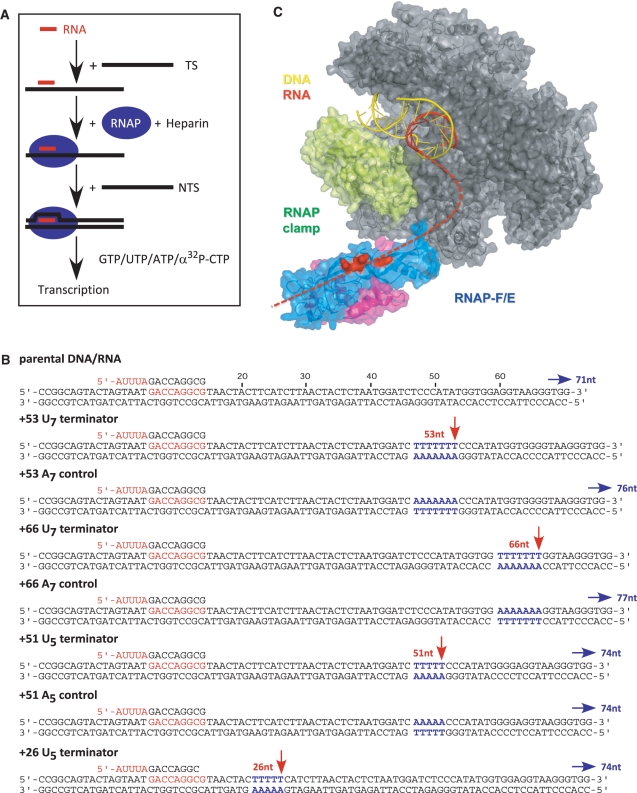
Figure 1.
Transcription elongation assay using synthetic nucleic acid scaffolds. Experimental setup (A). A 14 nt RNA oligonucleotide is pre-annealed to the DNA TS and subsequently incubated with recombinant RNAP and the NTS. Upon the addition of nucleotides RNAP extends the RNA primer completely independent of promoter elements and basal transcription factors. The templates used in this study are illustrated in (B). The RNA primer is highlighted in red and the DNA TS and NTS in black. The U7 and U5 terminator signals, and the A7 and A5 controls are highlighted in blue. Red arrows indicate the 3′ termini of terminated transcripts and blue arrows indicate the 3′ termini of transcripts generated by runoff. Structure and function of RNAP subunits F/E (C). ‘The S. shibatae RNAP structure (pdb 2WAQ) was manually superimposed on the DNA–RNA scaffold of the S. cerevisiae RNAPII elongation complex (DNA in yellow, RNA in red, pdb 1Y1W). The archaeal RNAP core is coloured grey, the two subunits F and E and highlighted in magenta and blue, and the RNAP clamp in green, respectively. Residues in subunit E that affect RNA binding are highlighted in red, and the RNA transcript that is not resolved in the crystal structures is sketched as red dotted line.
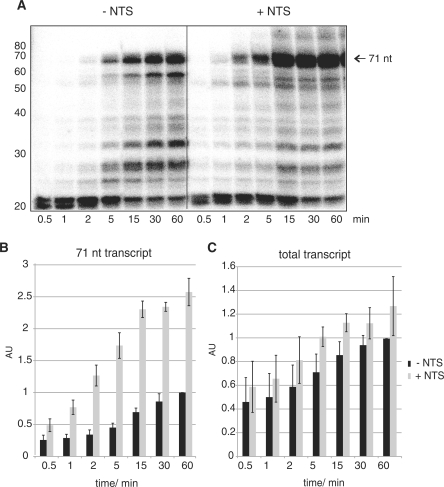
Figure 2.
The NTS stimulates the processivity. (A) The transcript pattern that was generated by in vitro transcription of the parental elongation scaffold (Figure 1B) under single-round conditions with RNAP (200 nM) in the presence (+NTS, grey bars) and absence of the NTS (−NTS, black bars). Samples were taken at the indicated time points (0.5, 1, 2, 5, 15, 30, 60 min). Histograms are based on quantization of at least three independent experiments (arbitrary units, AU). (B) The synthesis of the 71 nt-run off transcript normalized to the 60-min time point (−NTS). (C) The synthesis of total transcripts normalized to the 60-min time point (−NTS).
The RNAP subunits F/E enhance the processivity
Transcription elongation complexes are characterized by an astounding processivity in vivo, e.g. during the transcription of human muscle dystrophin transcription unit RNAPII elongates through 2.4 MB of coding sequence during which it remains associated with the DNA template for ∼16 h (31). The high stability of the elongation complex is reflected in a complex interaction network between RNAP subunits, the DNA template, the RNA-DNA hybrid and RNA transcript. We have recently solved the structure of the M. jannaschii RNAP subunits F/E and characterized its RNA-binding properties (4,32). We tested the functional contribution of F/E during transcription elongation by comparing wild type enzymes with recombinant RNAP variants lacking subunits F/E (RNAPΔF/E). The elongation properties of RNAPΔF/E are severely compromised (Figure 3A). The RNAPΔF/E is not capable of synthesizing the run off transcript in the absence of the NTS, even after 60 min of reaction time, and all RNAPs are paused on the DNA template at approximately position +33 (Figure 3A and C). The nature of this pause site is unclear and was not dependent on the CTP tracer nucleotide (data not shown). In comparison, the wild-type RNAP has already synthesized the runoff transcript at the earliest time point of 5 min (Figure 3A and C). In the presence of the NTS, the RNAPΔF/E is capable of forming runoff transcripts but the processivity of the wild-type enzyme is substantially greater (Figure 3B and D). In order to ascertain that the stimulatory activity of F/E on transcription elongation was dependent on the incorporation of F/E into the RNAP, and not due to ‘free’ F/E in solution we made use of the fact that the F/E complex can be incorporated into RNAPΔF/E prior to the reaction and that the incorporation of the F/E complex into RNAP is strictly dependent on subunit K (3). When we added F/E complex to either RNAPΔF/E or RNAPΔK/F/E only the former responded by synthesizing the runoff transcript (Figure 3E). This effect could be observed both in the absence and presence of the NTS (Figure 3E). This result demonstrates that the stimulatory properties of the F/E complex strictly depend on its association with the RNAP. We have previously generated F/E variants that are defective in RNA binding in vitro but do not affect their stability and incorporation into RNAP [(4) and data not shown]. We compared the activities of three mutant variants of F/E, F/EK33E, F/ER37E and F/Eloop (a subunit E triple mutant ER155A, K156S, R157A) in RNA binding and transcription elongation assays. Both assays were carried out with a range of F/E concentrations. Whereas the wild type F/E complex stimulates the synthesis of the runoff transcript ∼20-fold in our assay (Figure 4B and D), the RNA-binding-deficient mutant variants are impaired in a manner that correlate with their RNA-binding activity (Figure 4A, B and D). However, the F/Eloop variant that has no RNA-binding activity in vitro is still capable of stimulating the formation of the runoff transcript ∼5-fold at its highest concentration (Figure 4D). These results demonstrate that the molecular mechanisms by RNAP subunits F/E contribute to the processivity have two components. The main component is dependent on the RNA transcript binding activity of F/E, and the minor component independent of the transcript binding. In the absence of the NTS the latter mechanism accounts for about 20% of the stimulatory activity (Figure 4D). A likely mechanism for this activity is a conformational change of the RNAP, such as a closure of the RNAP clamp domain (26). The NTS improves the processivity of RNAP (Figure 2A) and accordingly the stimulatory activity of the F/E complex is weaker on duplex DNA templates. The phenotypes of the RNA-binding mutations are less pronounced under these conditions, implying that the F/E complex still stimulates elongation despite of weakened, or indeed absent interactions with the transcript. This is most obvious in a time course experiment shown in Figure 4C. At the first time point (40 s) the stimulatory activity of the RNA-binding-deficient F/Eloop variant is more than half of the wild type F/E, whereas the weaker RNA-binding mutants F/EK33E and F/ER37E have no phenotype (Figure 4C and E). After 300 s the activity of all F/E variant RNAPs have reached the same level, approximately twice the amount of transcripts generated by the RNAPΔF/E (Figure 4C and E).
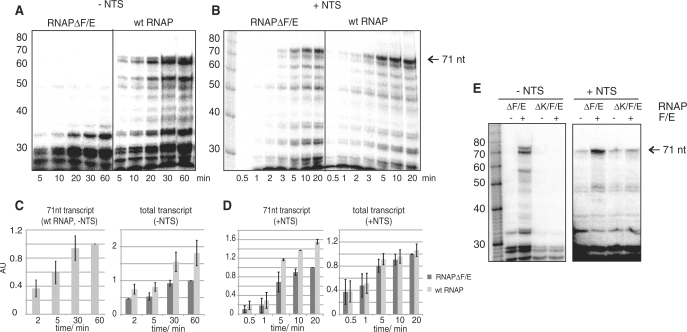
Figure 3.
The F/E complex enhances the processivity. (A) The transcript pattern generated by RNAPΔF/E (200 nM) using the parental elongation scaffold (Figure 1B) in the absence of the NTS either without (RNAPΔF/E) or following preincubation with 800 nM F/E (wt RNAP). (B) The transcript pattern generated by RNAPΔF/E (50 nM) in the presence of the NTS either without (RNAPΔF/E) or following preincubation with 200 nM F/E (wt RNAP). Samples were taken at the indicated time points. Note the difference in time scales (1–60 min −NTS and 0.5–20 min +NTS) and RNAP concentrations (200 nM −NTS and 50 nM +NTS) that are due to the higher activity level due to the presence of the NTS. Histograms are based on quantization of at least three independent experiments. (C) The synthesis of the 71 nt run off product wild type RNAP (grey bars) normalized to time point at 60 min (in arbitrary units, AU), and the total transcripts synthesized by wild type RNAP (grey bars) and RNAPΔF/E (black bars) normalized to the value at 60 min (−F/E). (D) The accumulation of the 71 nt transcript in the presence of the NTS by wild type RNAP (grey bars) and RNAPΔF/E (black bars), and the accumulation of total transcripts in the absence (black bars) or presence of F/E (grey bars). (E) RNAPΔF/E or RNAPΔK/F/E were preincubated with F/E and transcription reactions were carried out in the absence (−NTS) or presence of the NTS (+NTS).
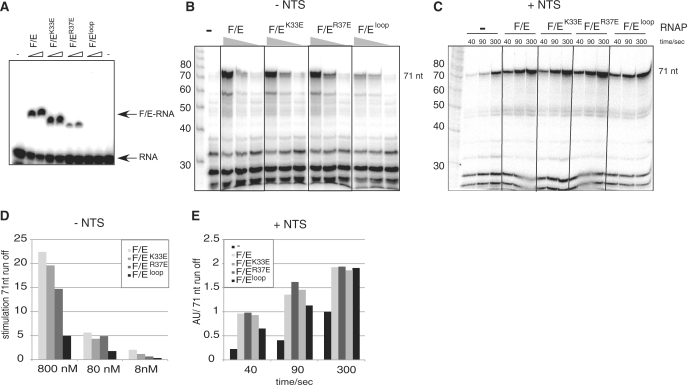
Figure 4.
The stimulatory effect of RNAP subunits F/E on transcription elongation correlates with the RNA binding activity. (A) The binding of wild type F/E and three mutant variants of the F/E complex (5 and 10 µM) to a 54 nt 32P-labelled riboprobe in EMSAs. (B) Transcription pattern generated by RNAPΔF/E (200 nM) preincubated with wild type F/E, F/EK33E, F/ER37E or F/Eloop in a dose-response experiment (8, 80 or 800 nM) using the parental elongation scaffold (Figure 1B) in the absence of the NTS (reaction time is 20 min). (C) The activity of the F/E variants in the presence of the NTS in a time course experiment. Transcript patterns generated are by RNAPΔF/E in the absence of F/E or preincubated with 80 nM F/E, F/EK33E, F/ER37E or F/Eloop. Samples were taken at the indicated time points (40, 90 and 300 s). (D) Quantization of the 71 nt transcript of panel (B) normalized to RNAPΔF/E. (E) Quantization of the 71 nt transcript of panel (C) normalized to RNAPΔF/E at 300 s (in arbitrary units).
Transcription termination of archaeal RNAP is facilitated by poly-U stretches
The 71 nt full-length transcripts in our transcription elongation assays are generated by transcription run off, i.e. spontaneous dissociation of the elongation complex at the 3′ terminus of the template. In vitro, archaeal RNAPs are capable of terminating transcription at poly-U stretches (10,14). The molecular mechanism of poly-U directed termination is distinct from run off termination. In order to investigate poly-U directed transcription termination in our recombinant system we inserted seven T-residues in the elongation templates at two different positions, +53 or +66 (Figure 1B). Recombinant archaeal RNAP terminates efficiently at both U7 termination signals generating transcripts 53 and 66 nucleotides long, respectively (Figure 5A). Close inspection of the data reveal that transcription terminates on the last U-residue of the terminator signal. The efficiency of termination is slightly higher at position +53 than compared to +66, which suggest a mild effect of the sequence context. In order to test whether the weak base pairing of the rU:dA RNA–DNA hybrid alone was sufficient to facilitate termination, we examined the elongation profiles of templates harbouring stretches of seven A instead of T residues in the NTS, resulting in rA:dT RNA–DNA hybrids. The A7-signals at +53 and +66 results in 90% and 100% read-through, respectively (Figure 5A). The results demonstrate that a weakening of the RNA–DNA hybrid by inserting an A7-signal is not sufficient to facilitate robust transcription termination, even though it is important to keep in mind that the interactions between rU:dA basepairs are less stable than rA:dU basepairs (33).
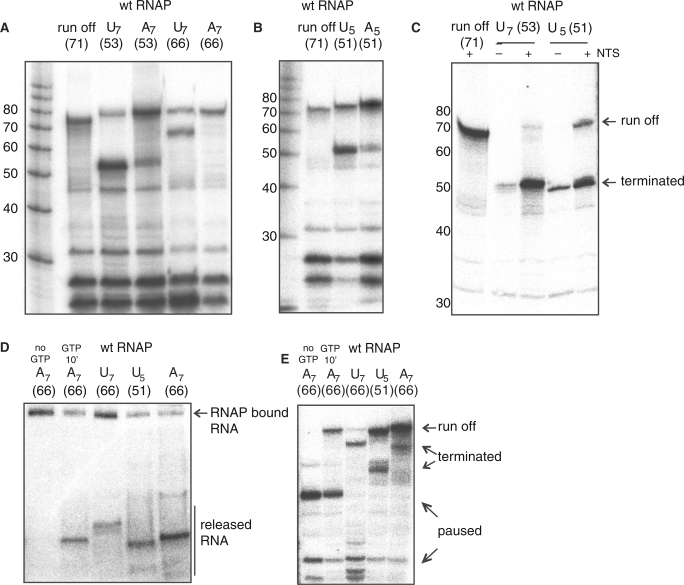
Figure 5.
Transcription termination of archaeal RNAP is facilitated by poly-U signals. (A) A stretch of seven U-residues (U7-signal) but not A-residues (A7-signals) in the template facilitate efficient transcription termination. The parental template generates a 71 nt runoff transcript, whereas the +53 and +66 U7 terminator templates generate 53 or 66 nt transcripts, respectively. The +53 and +66 A7 templates lead to the synthesis of runoff transcripts. All reactions were carried out for 5 min, include the NTS and contain 50 nM wild type RNAP. (B) Reducing the number of U-residues from seven (U7-signal) to five (U5-signal) increases transcription read through. Reactions were carried out for 5 min using 50 nM RNAP. (C) Both U7- and U5-signals can terminate transcription without the NTS. Reactions were carried out for 20 min using 50 nM RNAP. Panels (D) and (E) show that both run off- and terminated-, but not stalled-transcripts are released from the elongation complex. Products of transcription elongation reactions using the +66 A7-, +66 U7- and +51 U5-templates were separated electrophoretically under native (D) and denaturating conditions (E). As a negative control for transcript release stalled transcription elongation complexes were generated by omitting GTP from a transcription reaction (‘no GTP') using the +66 A7-template. Under these conditions a 41 nt transcript is synthesized which remains associated with RNAP. Adding GTP to this reaction after 10 min (‘GTP at 10’) and letting the reaction proceed for another 10 min generates the 66 nt run off transcript (E) that is released (D). Transcripts synthesized from U7- and U5-templates are released (D). In addition a proportion of transcripts can be found in low mobility region of the native gel that corresponds to paused complexes (D and E). For unclear reasons the U7 (+66) template leads to more far upstream pausing (at <30 nt) than the U7 (+66) and U5 (+51) templates.
Efficiency of transcription termination correlates with the number of U-residues
In order to test whether the number of U-residues in the terminator signal correlated with termination efficiency and read through, we reduced the number of T residues in the NTS of the elongation scaffolds from seven to five and compared transcription termination efficiency with the U7 template. The U5 signal resulted in 60% termination (defined as terminated RNA/terminated plus read through RNA), as compared to 95% termination facilitated by the U7 signal (Figure 5A and B). A control template harbouring five A residues led to 20% termination (Figure 5B). We conclude that the number of T residues in the termination signal plays an important role for termination and read-through. This is in agreement with observations from the Pyrococcus furiosus histone gene terminator (10). We have shown that the NTS increases the processivity, and we speculated that this could also result in increased read-through and decreased termination facilitated by U7 signals. In order to test the influence of the NTS on poly-U directed termination we carried out transcription termination assays in the absence of the NTS and on duplex DNA templates. The NTS does not qualitatively affect transcription termination directed by the strong U7-signal, however on the weaker U5-signal the NTS leads to increased read through (Figure 5C). On both U5- and U7-signals the total amount of transcripts is increased by the NTS in accordance with our previous observations (Figure 5C). The molecular mechanism of transcription termination is not well understood but is likely to require pausing of RNAP that allows conformational changes of the TEC, which eventually lead to transcript release and complex dissociation (34). Poly-U sequences are strong pause signals (35) and it is therefore exigent to demonstrate that the U7 signals lead to genuine termination of transcription, and not only pausing and possibly arrest of our elongation complexes. We ascertained that both run off and U-signal terminated transcripts were released from elongation complexes by separating free and RNAP-bound transcripts by native gel electrophoresis of the elongation reactions. Figure 5D and E show transcription elongation reactions that were electrophoretically separated under native and denaturating conditions, respectively. As a negative control for transcript release we carried out transcription reactions in the absence of GTP, which results in a paused elongation complex after synthesis of a 41 nt transcript (Figures 1B and 5E). Native gel electrophoresis of this sample demonstrates that all 32P-labelled RNA migrates as one distinct low mobility band that corresponds to the paused TEC (Figure 5D). Upon the addition of GTP after 10 min incubation of the transcription reaction the runoff transcript is formed (Figure 5E) and concomitantly a new faster migrating band appears on a native gel analysis of the reaction, which corresponds to the released transcript (Figure 5D). We subjected transcription elongation reactions with U7, A7 and U5 signal scaffolds to this comparative analysis and in all reactions free transcripts were observed (Figure 5D and E). We conclude that under our assay conditions both runoff and U-signal terminated transcripts are efficiently released from the recombinant RNAPs.
RNAP subunits F/E improve transcription termination by U5 signals
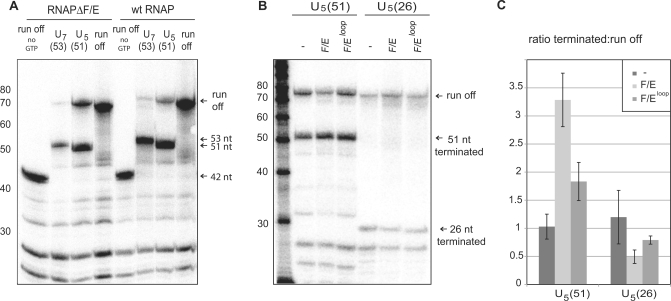
Above we have demonstrated that RNAP subunits F/E play an important role during transcription elongation that is likely to involve interactions with the transcript and conformational changes of the TEC, e.g. through a modulation of the RNAP clamp (26). In order to test whether transcription termination facilitated by poly-U signals was dependent on, or affected by the F/E complex we compared the transcript pattern of wild type RNAP and RNAPΔF/E using runoff, U7- and U5-signal elongation scaffolds. Both wild type and RNAPΔF/E generate qualitatively very similar transcript patterns on all templates tested (Figure 6A). This demonstrates that transcription termination of archaeal RNAP is not strictly dependent on the F/E subunits. However, F/E affects the termination efficiency of RNAP on weak U5 termination signals in a quantitative manner (Figure 6B and C). The ratio between terminated versus read through transcript is increased more than 3-fold by the addition of wild type F/E, and less than 2-fold by the RNA-binding deficient F/Eloop mutant. In order to elucidate the role of the RNA binding further we made use of a template containing an upstream U5 termination signal (Figure 1B, +26 U5 terminator) taking into account that the F/E complex interacts with the RNA 26–41 nt downstream of the active site (5) (Figure 1B). The ratio between terminated and read through transcript was not affected by the position of the U5-signal (+26 versus +51) in the absence of F/E (Figure 6B and C). However, on the +26 U5 template neither F/E nor F/Eloop increased transcription termination, but rather led to an increase of the runoff transcript formation congruent with F/E's; function of stimulating the processivity (Figure 6B and C). This result demonstrates that F/E only enhances transcription termination upon interaction with the RNA. If interactions between the transcript and RNAP subunits F/E are abrogated by mutations in F/E (F/Eloop) or by restricting the length of the transcript (the position of the termination signal in the +26 U5 template), F/E cannot stimulate transcription termination.
Figure 6.
RNAP subunits F/E enhance transcription termination from weak U5-terminator signals. (A) Comparison of transcript patterns generated by wild type RNAP and RNAPΔF/E using parental-(‘run off’), +53 U7- and +51 U5-templates. Reactions were carried out for 90 s using 50 nM RNAP in the presence of the NTS. RNAP subunits F/E stimulate the formation of RNA transcripts, but have no significant influence on pausing at position +41 (‘no GTP’) or on transcription termination directed by the strong +53 U7-signal. (B) The transcript pattern generated by wild type RNAP, RNAPΔF/E and RNAP F/Eloop using the +51 U5- and +26 U5-terminator templates. Reactions were carried out for 20 min including the NTS and using 50 nM RNAP. (C) Quantization of the ratio between terminated and read through transcripts in panel (B).
DISCUSSION
The elongation and termination phases of transcription are crucial for the accurate synthesis of all RNA molecules. In addition, transcription elongation has emerged as an important and widespread regulatory mechanism for gene expression by both eukaryotic and bacterial RNAPs (8,9,16–18,36). In order to thoroughly rationalize the molecular mechanisms that govern transcription elongation by RNAP from the third domain of life, the Archaea, we have used a wholly recombinant RNAP in transcription elongation experiments that utilize synthetic elongation scaffolds. This system has allowed us to perturb the RNAP and the nucleic acid scaffold of the TEC in order to investigate the influence of RNAP subunits (F/E), the TS, NTS and RNA transcript on transcription elongation and termination. Multi-subunit RNAPs from all three domains of life are capable of transcription elongation using exclusively the TS DNA (27,28); this also applies to the archaeal RNAP. However, the NTS improves the processivity, i.e. the RNAP synthesizes more full-length product and is less prone to pausing. This is in good agreement with the bacterial and yeast transcription systems where the NTS has been shown to restrict the lateral mobility of the elongating RNAP by reducing pausing and backtracking (27,37–39). Kashlev and colleagues have shown that the NTS is important for the DNA–RNA separation at the upstream edge of the transcription bubble of RNAPII; the absence of the NTS results in an overextended hybrid that drives backsliding of the enzyme and thereby impairs the processivity (37). Due to the close relationship between RNAPII and the archaeal RNAP it is not unlikely that a similar mechanism is responsible for the NTS effect on processivity we observed in this study. Several lines of evidence suggest that RNAP subunits F/E interact with the RNA transcript, (i) we and others have shown that the F/E complex bind nucleic acids in vitro (4,40), (ii) human RPB7 can be UV cross-linked to radio-labelled transcripts between RNA registers +26 and +41 (5) and (iii) structural information of RNAPII elongation complexes suggests that the RNA transcript could be directed towards RPB4/7 (20,21). However, whether this RNA binding is biologically relevant is controversial, since a FRET study using fluorescently labelled RNA and RPB7 in the RNAPII system failed to detect a proximity between RPB4/7 and the transcript in solution (41). Our results unequivocally demonstrate that the F/E complex has a profound effect on the transcription elongation properties of RNAP in vitro by enhancing the processivity. In principle, we cannot rule out that F/E leads to an increase in transcription elongation rate, but it is problematic to discern between processivity and elongation rate experimentally in our assay. Wild-type RNAPs synthesize more full-length transcripts and are less prone to pausing than RNAPΔF/E. This result is congruent with in vivo data in the rpb4Δ strain of S. cerevisiae, which shows a reduction of RNAPII levels at the 3′-end of multiple mRNA genes (42). The stimulatory activity of F/E on transcription elongation in vitro is most pronounced on synthetic scaffolds consisting of TS and RNA. Previous experimental approaches were not able to detect the effect of F/E for a number of reasons. The template in promoter-independent (‘nonspecific’) transcription assays consisted of double stranded calf thymus DNA, and the promoter-directed transcription assays carried out in our lab used supercoiled plasmid templates (23). Both experimental setups are fundamentally different from the assays reported in this manuscript, because they make use of different templates and are based on multiple round transcription endpoint assays. The elongation experiments using synthetic scaffolds are single round transcription assays following the synthesis of transcripts over a time course. It is possible that the subtle activities of F/E using double stranded templates have simply been overlooked in these less sophisticated assays employed in the past. Similarly, early studies with S. cerevisiae RNAPII were not able to detect a stimulatory effect of the homologous RPB4/7 complex on transcription elongation assays using double stranded templates (43). The M. jannaschii RNAP operates at elevated temperatures (65°C) and it is possible that under these conditions the interactions between F/E and the RNA are more important for TEC stability and elongation, compared to the low temperatures (25°C) under which the yeast transcription assays were carried out. We can observe this activity in vitro on relatively short templates; in vivo the RNAP transcribes much longer transcription units (e.g. the 2.4 Mb Dystrophin gene) and the effect of F/E is even more vital. These mechanisms are specific for the elongation phase of transcription and are unlikely to be caused by secondary effects since the absence of F/E complex does not compromise the (heat-) stability and solubility of RNAP (23) and thereby integrity of the active site (7,23). The stimulatory activity of F/E on transcription elongation depends on RNAP subunit K and thereby on the stable incorporation of F/E into the RNAP (3). Mutant variants of F/E complex that compromise its RNA binding in vitro (4) are defective in stimulating transcription elongation. We conclude that the stimulatory activity of F/E is largely based on interactions between the RNA transcript and the RNAP. The additional contacts between F/E and RNA contribute to the interaction network of the TEC and thereby stabilize it. In addition, it is possible that the F/E-RNA interactions contribute to the efficient separation of the RNA from TS at the upstream edge of the DNA–RNA hybrid (37). Both mechanisms have the potential to stimulate the processivity, and both would be dependent on the RNA-binding activity of F/E. However, even the F/E ‘loop’ mutant that cannot bind RNA retains baseline stimulatory activity. We can not rule out the possibility that the F/E ‘loop’ mutant binds RNA in the context of the TEC but would like to suggest that the activity could be due to conformational changes of the RNAP induced by F/E binding, e.g. a closure of the RNAP clamp domain (Figure 1C). This hypothesis finds support in difference maps of X-ray structures generated by comparing the 12-subunit RNAPII and 10-subunit RNAPII ΔRPB4/7, which indicate that the position of the mobile RNAP clamp is modulated by RPB4/7 (26,44). The binding of nucleic acids could also induce the closure of the RNAP clamp, as seen in the structure of the 10-subunit RNAPII elongation complex (20). Alternatively, it cannot be ruled out that the discrepancies of the clamp position between the 10- and 12-subunit RNAPII are due to differences in the crystallization conditions. Transcription termination is the least understood phase of the transcription cycle. The archaeal system is unusual in as much as efficient termination only requires a stretch of several U-residues in the transcript. Our results show that the recombinant M. jannaschii RNAP terminates efficiently when transcribing a stretch of seven U-residues (U7 signal) and this mechanism is largely independent of the sequence context. Furthermore we demonstrate that transcription termination on strong U7-signals can occur independently of the NTS, whereas the NTS mildly stimulates read through on weak U5-signals. This implies that the molecular mechanisms that are underlying robust U-stretch directed termination chiefly depend on interactions between the template DNA strand, the RNA transcript and the RNAP. But how is transcription termination facilitated? The release of the transcript and the dissociation of the RNAP–DNA complex requires disruption of approximately nine hydrogen bonds of the DNA–RNA hybrid and is likely to involve substantial conformational changes within RNAP (e.g. clamp opening). The insertion of an A7-signal into the transcript did not result in efficient termination in good agreement with previous observations from the Methanothermobacterium system (14). Even though rU:dA basepairs are less stable than rA:dT base pairs, DNA–RNA hybrid stability alone is unlikely to account for termination (33,45). This implies that interactions between the RNAP DNA–RNA hybrid binding compartment and the rU:dA hybrid, but not a rA:dT hybrid are necessary and sufficient for the termination mechanism. This result is in good agreement with observations of the bacterial RNAP system were U-rich RNA:DNA hybrids were found to destabilize the elongation complex (46). A prerequisite for efficient termination is transcriptional pausing, which allows conformational changes within the elongation complex to occur that subsequently result in termination (34,35). By reducing the number of U-residues from seven (U7-signal) to five (U5-signal) the termination efficiency was reduced by half. This could be due to shorter pausing and is in good agreement with a previous study of the P. furiosus histone gene terminator (10). The F/E complex has no significant impact on transcription termination that is facilitated by the strong U7 signals. However, termination by the weaker U5-signals is enhanced by the F/E complex and this phenomenon is dependent on its RNA binding, since the RNA-binding deficient F/Eloop mutant cannot support this activity. Even though F/Eloop does not bind RNA in vitro we cannot strictly rule out that it weakly interacts with the transcript in the context of the elongation complex due to the increased local concentration of RNA. However, by decreasing the distance between the U5-signal and the 5′ terminus of the transcript to a size that precluded its interaction with F/E (26 nt) the termination activity of F/E was abrogated. The stimulatory effect of F/E on U5-signal termination in vitro seems modest, however, the choice between elongation and termination on U5-signals in vivo could be a fine balance that is affected by even small changes in the RNAP including the F/E complex. A likely explanation for the termination activity of F/E lies in the additional interactions between F/E and the transcript, which could induce pausing and lead to termination. This mechanism is dependent on rU5:dA5 in the RNAP DNA–RNA hybrid compartment and interactions between F/E and the RNA, since the F/E complex increases the processivity in the absence of poly-U stretches.
The RNAP F/E complex, and its eukaryotic RNAPII homologue RPB4/7, plays an important role during transcription initiation by facilitating DNA melting and mediating functional interactions of RNAP with the basal factors TFE and TFIIF in Archaea and eukaryotes, respectively (6,7,25). Here we demonstrate that the F/E complex, and by inference RPB4/7, is involved in the molecular mechanisms of RNAP during transcription elongation and termination. We attribute the main function of F/E to its ability to interact with the RNA transcript, but also propose that the interaction between F/E and the RNAP core leads to conformational changes within latter that modulate its DNA binding and possibly catalytic properties. F/E-like complexes are characteristic for all eukaryotic classes of RNAP (I–V) and archaeal enzymes, but they have no homologues in bacterial RNAP. What insights can we gain into overall differences or similarities between the molecular mechanisms of bacterial and archaeo-eukaryotic RNAP, the way their respective genes are transcribed and how this process is regulated? In the context of transcription elongation it is tempting to speculate that F/E-like complexes facilitate a higher processivity and thereby enable RNAPs to transcribe longer genes, even though that is only relevant for eukaryotic, and not archaeal genes. It is noteworthy that the properties of bacteriophage antitermination factors, such as lambda Q, are reminiscent of the F/E complex. Even though Q cannot directly bind RNA, Q interacts with the beta-flap region of RNAP, which is in close proximity to the RNA-exit channel (47). Association of Q with RNAP increases its processivity and alters transcription termination patterns with the overall outcome of facilitating and thereby regulating the expression of the late genes of phage lambda (48). The RNAP subunits F/E and RPB4/7 function on a global scale, whereas Q is an operon-specific transcription factor. However, both modulate their cognate RNAP to achieve an increased processivity.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
Wellcome Trust (grant 079351/Z/06/Z to F.W.); BBSRC (grant BB/E008232/1 to F.W.). Funding for open access charge: Wellcome Trust UCL resources.
Conflict of interest statement. None declared.
Supplementary Material
ACKNOWLEDGEMENTS
The authors thank Dr Kristine Bourke Arnvig for critical reading of the manuscript.
REFERENCES
- 1.Werner F. Structural evolution of multisubunit RNA polymerases. Trends Microbiol. 2008;16:247–250. doi: 10.1016/j.tim.2008.03.008. [DOI] [PubMed] [Google Scholar]
- 2.Werner F. Structure and function of archaeal RNA polymerases. Mol. Microbiol. 2007;65:1395–1404. doi: 10.1111/j.1365-2958.2007.05876.x. [DOI] [PubMed] [Google Scholar]
- 3.Grohmann D, Hirtreiter A, Werner F. The RNAP subunits F/E (RPB4/7) are stably associated with archaeal RNA polymerase – using fluorescence anisotropy to monitor RNAP assembly in vitro. Biochem. J. 2009;421:339–343. doi: 10.1042/BJ20090782. [DOI] [PubMed] [Google Scholar]
- 4.Meka H, Werner F, Cordell SC, Onesti S, Brick P. Crystal structure and RNA binding of the Rpb4/Rpb7 subunits of human RNA polymerase II. Nucleic Acids Res. 2005;33:6435–6444. doi: 10.1093/nar/gki945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ujvari A, Luse DS. RNA emerging from the active site of RNA polymerase II interacts with the Rpb7 subunit. Nat. Struct. Mol. Biol. 2006;13:49–54. doi: 10.1038/nsmb1026. [DOI] [PubMed] [Google Scholar]
- 6.Naji S, Grunberg S, Thomm M. The RPB7 orthologue E' is required for transcriptional activity of a reconstituted archaeal core enzyme at low temperatures and stimulates open complex formation. J. Biol. Chem. 2007;282:11047–11057. doi: 10.1074/jbc.M611674200. [DOI] [PubMed] [Google Scholar]
- 7.Ouhammouch M, Werner F, Weinzierl RO, Geiduschek EP. A fully recombinant system for activator-dependent archaeal transcription. J. Biol. Chem. 2004;279:51719–51721. doi: 10.1074/jbc.C400446200. [DOI] [PubMed] [Google Scholar]
- 8.Saunders A, Core LJ, Lis JT. Breaking barriers to transcription elongation. Nat. Rev. Mol. Cell Biol. 2006;7:557–567. doi: 10.1038/nrm1981. [DOI] [PubMed] [Google Scholar]
- 9.Landick R. The regulatory roles and mechanism of transcriptional pausing. Biochem. Soc. Trans. 2006;34:1062–1066. doi: 10.1042/BST0341062. [DOI] [PubMed] [Google Scholar]
- 10.Spitalny P, Thomm M. A polymerase III-like reinitiation mechanism is operating in regulation of histone expression in archaea. Mol. Microbiol. 2008;67:958–970. doi: 10.1111/j.1365-2958.2007.06084.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hamada M, Sakulich AL, Koduru SB, Maraia RJ. Transcription termination by RNA polymerase III in fission yeast. A genetic and biochemically tractable model system. J. Biol. Chem. 2000;275:29076–29081. doi: 10.1074/jbc.M003980200. [DOI] [PubMed] [Google Scholar]
- 12.Ciampi MS. Rho-dependent terminators and transcription termination. Microbiology. 2006;152:2515–2528. doi: 10.1099/mic.0.28982-0. [DOI] [PubMed] [Google Scholar]
- 13.Henkin TM. Control of transcription termination in prokaryotes. Annu. Rev. Genet. 1996;30:35–57. doi: 10.1146/annurev.genet.30.1.35. [DOI] [PubMed] [Google Scholar]
- 14.Santangelo TJ, Reeve JN. Archaeal RNA polymerase is sensitive to intrinsic termination directed by transcribed and remote sequences. J. Mol. Biol. 2006;355:196–210. doi: 10.1016/j.jmb.2005.10.062. [DOI] [PubMed] [Google Scholar]
- 15.West S, Gromak N, Proudfoot NJ. Human 5′ → 3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature. 2004;432:522–525. doi: 10.1038/nature03035. [DOI] [PubMed] [Google Scholar]
- 16.Margaritis T, Holstege FC. Poised RNA polymerase II gives pause for thought. Cell. 2008;133:581–584. doi: 10.1016/j.cell.2008.04.027. [DOI] [PubMed] [Google Scholar]
- 17.Core LJ, Lis JT. Transcription regulation through promoter-proximal pausing of RNA polymerase II. Science. 2008;319:1791–1792. doi: 10.1126/science.1150843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Core LJ, Waterfall JJ, Lis JT. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science. 2008;322:1845–1848. doi: 10.1126/science.1162228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Brueckner F, Armache KJ, Cheung A, Damsma GE, Kettenberger H, Lehmann E, Sydow J, Cramer P. Structure-function studies of the RNA polymerase II elongation complex. Acta. Crystallogr. D. Biol. Crystallogr. 2009;65:112–120. doi: 10.1107/S0907444908039875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gnatt AL, Cramer P, Fu J, Bushnell DA, Kornberg RD. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science. 2001;292:1876–1882. doi: 10.1126/science.1059495. [DOI] [PubMed] [Google Scholar]
- 21.Kettenberger H, Armache KJ, Cramer P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell. 2004;16:955–965. doi: 10.1016/j.molcel.2004.11.040. [DOI] [PubMed] [Google Scholar]
- 22.Grohmann D, Hirtreiter A, Werner F. Molecular mechanisms of archaeal RNA polymerase. Biochem. Soc. Trans. 2009;37:12–17. doi: 10.1042/BST0370012. [DOI] [PubMed] [Google Scholar]
- 23.Werner F, Weinzierl RO. A recombinant RNA polymerase II-like enzyme capable of promoter-specific transcription. Mol. Cell. 2002;10:635–646. doi: 10.1016/s1097-2765(02)00629-9. [DOI] [PubMed] [Google Scholar]
- 24.Werner F, Weinzierl RO. Direct modulation of RNA polymerase core functions by basal transcription factors. Mol. Cell Biol. 2005;25:8344–8355. doi: 10.1128/MCB.25.18.8344-8355.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chung WH, Craighead JL, Chang WH, Ezeokonkwo C, Bareket-Samish A, Kornberg RD, Asturias FJ. RNA polymerase II/TFIIF structure and conserved organization of the initiation complex. Mol. Cell. 2003;12:1003–1013. doi: 10.1016/s1097-2765(03)00387-3. [DOI] [PubMed] [Google Scholar]
- 26.Armache KJ, Mitterweger S, Meinhart A, Cramer P. Structures of complete RNA polymerase II and its subcomplex, Rpb4/7. J. Biol. Chem. 2005;280:7131–7134. doi: 10.1074/jbc.M413038200. [DOI] [PubMed] [Google Scholar]
- 27.Kireeva ML, Komissarova N, Waugh DS, Kashlev M. The 8-nucleotide-long RNA:DNA hybrid is a primary stability determinant of the RNA polymerase II elongation complex. J. Biol. Chem. 2000;275:6530–6536. doi: 10.1074/jbc.275.9.6530. [DOI] [PubMed] [Google Scholar]
- 28.Temiakov D, Anikin M, McAllister WT. Characterization of T7 RNA polymerase transcription complexes assembled on nucleic acid scaffolds. J. Biol. Chem. 2002;277:47035–47043. doi: 10.1074/jbc.M208923200. [DOI] [PubMed] [Google Scholar]
- 29.Cheng B, Price DH. Properties of RNA polymerase II elongation complexes before and after the P-TEFb-mediated transition into productive elongation. J. Biol. Chem. 2007;282:21901–21912. doi: 10.1074/jbc.M702936200. [DOI] [PubMed] [Google Scholar]
- 30.Cheng B, Price DH. Analysis of factor interactions with RNA polymerase II elongation complexes using a new electrophoretic mobility shift assay. Nucleic Acids Res. 2008;36:e135. doi: 10.1093/nar/gkn630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tennyson CN, Klamut HJ, Worton RG. The human dystrophin gene requires 16 hours to be transcribed and is cotranscriptionally spliced. Nat. Genet. 1995;9:184–190. doi: 10.1038/ng0295-184. [DOI] [PubMed] [Google Scholar]
- 32.Todone F, Brick P, Werner F, Weinzierl RO, Onesti S. Structure of an archaeal homolog of the eukaryotic RNA polymerase II RPB4/RPB7 complex. Mol. Cell. 2001;8:1137–1143. doi: 10.1016/s1097-2765(01)00379-3. [DOI] [PubMed] [Google Scholar]
- 33.Martin FH, Tinoco I., Jr DNA-RNA hybrid duplexes containing oligo(dA:rU) sequences are exceptionally unstable and may facilitate termination of transcription. Nucleic Acids Res. 1980;8:2295–2299. doi: 10.1093/nar/8.10.2295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Epshtein V, Cardinale CJ, Ruckenstein AE, Borukhov S, Nudler E. An allosteric path to transcription termination. Mol. Cell. 2007;28:991–1001. doi: 10.1016/j.molcel.2007.10.011. [DOI] [PubMed] [Google Scholar]
- 35.Gusarov I, Nudler E. The mechanism of intrinsic transcription termination. Mol. Cell. 1999;3:495–504. doi: 10.1016/s1097-2765(00)80477-3. [DOI] [PubMed] [Google Scholar]
- 36.Borukhov S, Lee J, Laptenko O. Bacterial transcription elongation factors: new insights into molecular mechanism of action. Mol. Microbiol. 2005;55:1315–1324. doi: 10.1111/j.1365-2958.2004.04481.x. [DOI] [PubMed] [Google Scholar]
- 37.Kireeva ML, Komissarova N, Kashlev M. Overextended RNA:DNA hybrid as a negative regulator of RNA polymerase II processivity. J. Mol. Biol. 2000;299:325–335. doi: 10.1006/jmbi.2000.3755. [DOI] [PubMed] [Google Scholar]
- 38.Nudler E, Goldfarb A, Kashlev M. Discontinuous mechanism of transcription elongation. Science. 1994;265:793–796. doi: 10.1126/science.8047884. [DOI] [PubMed] [Google Scholar]
- 39.Wang D, Landick R. Nuclease cleavage of the upstream half of the nontemplate strand DNA in an Escherichia coli transcription elongation complex causes upstream translocation and transcriptional arrest. J. Biol. Chem. 1997;272:5989–5994. doi: 10.1074/jbc.272.9.5989. [DOI] [PubMed] [Google Scholar]
- 40.Orlicky SM, Tran PT, Sayre MH, Edwards AM. Dissociable Rpb4-Rpb7 subassembly of rna polymerase II binds to single-strand nucleic acid and mediates a post-recruitment step in transcription initiation. J. Biol. Chem. 2001;276:10097–10102. doi: 10.1074/jbc.M003165200. [DOI] [PubMed] [Google Scholar]
- 41.Andrecka J, Lewis R, Bruckner F, Lehmann E, Cramer P, Michaelis J. Single-molecule tracking of mRNA exiting from RNA polymerase II. Proc. Natl Acad. Sci. USA. 2008;105:135–140. doi: 10.1073/pnas.0703815105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Runner VM, Podolny V, Buratowski S. The Rpb4 subunit of RNA polymerase II contributes to cotranscriptional recruitment of 3′ processing factors. Mol. Cell Biol. 2008;28:1883–1891. doi: 10.1128/MCB.01714-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Edwards AM, Kane CM, Young RA, Kornberg RD. Two dissociable subunits of yeast RNA polymerase II stimulate the initiation of transcription at a promoter in vitro. J. Biol. Chem. 1991;266:71–75. [PubMed] [Google Scholar]
- 44.Bushnell DA, Kornberg RD. Complete, 12-subunit RNA polymerase II at 4.1-A resolution: implications for the initiation of transcription. Proc. Natl Acad. Sci. USA. 2003;100:6969–6973. doi: 10.1073/pnas.1130601100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Gil A, Branchadell V, Bertran J, Oliva A. An analysis of the different behavior of DNA and RNA through the study of the mutual relationship between stacking and hydrogen bonding. J. Phys. Chem. B. 2009;113:4907–4914. doi: 10.1021/jp809737c. [DOI] [PubMed] [Google Scholar]
- 46.Greive SJ, von Hippel PH. Thinking quantitatively about transcriptional regulation. Nat. Rev. Mol. Cell Biol. 2005;6:221–232. doi: 10.1038/nrm1588. [DOI] [PubMed] [Google Scholar]
- 47.Deighan P, Diez CM, Leibman M, Hochschild A, Nickels BE. The bacteriophage lambda Q antiterminator protein contacts the beta-flap domain of RNA polymerase. Proc. Natl Acad. Sci. USA. 2008;105:15305–15310. doi: 10.1073/pnas.0805757105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Roberts JW, Yarnell W, Bartlett E, Guo J, Marr M, Ko DC, Sun H, Roberts CW. Antitermination by bacteriophage lambda Q protein. Cold Spring Harb. Symp. Quant. Biol. 1998;63:319–325. doi: 10.1101/sqb.1998.63.319. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.