Abstract
We currently understand the mental effects of psychedelics to be caused by agonism or partial agonism of 5-HT2A (and possibly 5-HT2C) receptors, and we understand that psychedelic drugs, especially phenylalkylamines, are fairly selective for these two receptors. This manuscript is a reference work on the receptor affinity pharmacology of psychedelic drugs. New data is presented on the affinity of twenty-five psychedelic drugs at fifty-one receptors, transporters, and ion channels, assayed by the National Institute of Mental Health – Psychoactive Drug Screening Program (NIMH-PDSP). In addition, comparable data gathered from the literature on ten additional drugs is also presented (mostly assayed by the NIMH-PDSP). A new method is introduced for normalizing affinity (Ki) data that factors out potency so that the multi-receptor affinity profiles of different drugs can be directly compared and contrasted. The method is then used to compare the thirty-five drugs in graphical and tabular form. It is shown that psychedelic drugs, especially phenylalkylamines, are not as selective as generally believed, interacting with forty-two of forty-nine broadly assayed sites. The thirty-five drugs of the study have very diverse patterns of interaction with different classes of receptors, emphasizing eighteen different receptors. This diversity of receptor interaction may underlie the qualitative diversity of these drugs. It should be possible to use this diverse set of drugs as probes into the roles played by the various receptor systems in the human mind.
Introduction
We currently understand the mental effects of psychedelics to be caused by agonism or partial agonism of 5-HT2A (and possibly 5-HT2C) receptors (serotonin-2A and serotonin-2C receptors) [1]. This understanding was first developed in the 1980s [2]–[4] and has since been confirmed by a large body of evidence, as reviewed recently by Nichols [1]. However, many authors have commented that other receptors may also play a role [1], [3], [5]–[9]. In this post-genome era of high-throughput assays, it is time to take a broader view, move beyond the common-denominator approach [6], and begin to explore the role of other receptors in shaping the mental effects of psychedelics, especially the qualitative differences among them.
The objective of this paper is to present the receptor binding profiles of the thirty-five drugs (Fig. 1, Fig. 2) of this study in such a way that they can be easily compared in both their similarities and their differences. This is intended to serve as a reference work on the multi-receptor affinity pharmacology of psychedelic drugs. The tables and figures are the heart of this manuscript. Some of them have been included as “supporting information,” because they exceed the size limits of standard tables and figures. However, this supporting information is no less central to the manuscript than the standard tables and figures.
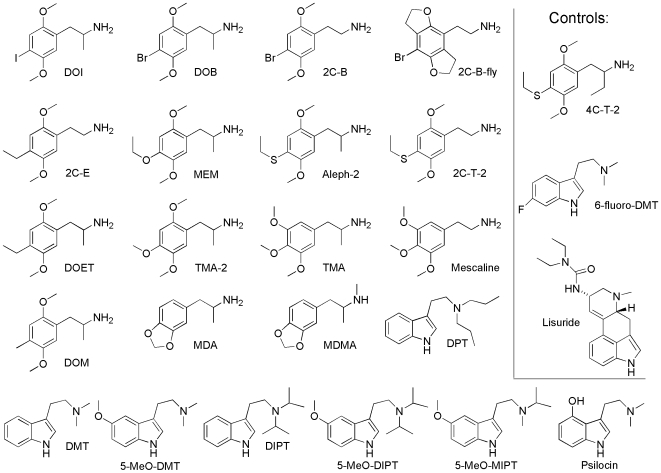
Figure 1. Twenty-five drugs assayed for this study by the NIMH-PDPS.
Twenty-five drugs assayed for this study by the NIMH-PDPS against fifty-one receptors, transporters and ion-channels. The twenty-five drugs include sixteen phenylalkylamines, eight tryptamines, and one ergoline. The three control drugs on the right include one representative from each structural class, and are believed to be non-psychedelic.
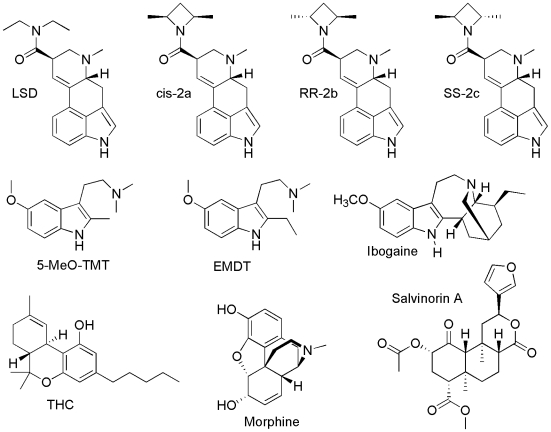
Figure 2. Ten drugs whose receptor profiles were collected from the literature.
Ten drugs whose receptor profiles were collected from the literature. All but ibogaine, THC, and morphine were assayed by the NIMH-PDSP.
Methods
Data from Literature
Data on receptor interactions of ten compounds (Fig. 2) has been collected from the literature. The four ergolines (LSD, cis-2a, RR-2b, and SS-2c) were assayed by NIMH-PDSP against forty-three receptors, transporters and ion channels [10]. Salvinorin A was assayed by NIMH-PDSP against thirty receptors and transporters [11]. EMDT and 5-MeO-TMT were assayed by NIMH-PDSP against forty receptors, transporters and ion channels [12].
Receptor data for ibogaine (Table S1), morphine (Table 1) and THC (Table 2) was collected from a variety of sources. While ibogaine has been assayed at a wide variety of receptors, morphine and THC have not, so their data should be used with caution. Although morphine is not considered to be a psychedelic, and ibogaine, THC, and salvinorin A are not considered to be “classical hallucinogens,” these four compounds are included because they provide insights into additional receptor systems (salvinorin A – κ (kappa opioid receptor), ibogaine – σ (sigma receptor) and κ, THC – CB (cannabinoid receptor), morphine – μ (mu opioid receptor)). These additional compounds could also be thought of as active controls, as compared to the three presumably inactive controls of Fig. 1.
Table 1. Receptor affinity data for morphine.
| Receptors | Hot Ligand | Source | Tissue | Ki(nM) | IC50(nM) | Reference |
| KOR | 3H-U69,593 | GUINEA PIG | ILEUM | 217±49 | PDSP; [17] | |
| 3H-U69,593 | Human | cloned | 134±22 | PDSP; [18] | ||
| [Dmt]DALDA | Human | cloned | 4.4±1.7 | [19] | ||
| DMAGO | Human | cloned | 213±28 | [19] | ||
| [3H]U69593 | rat | Brain | 113±9 | PDSP; [20] | ||
| 50 | average human | |||||
| MOR | 3H-DAMGO | GUINEA PIG | ILEUM | 1±0.04 | PDSP; [17] | |
| 3H-Diprenorphine | Human | cloned | 2.06±0.48 | PDSP; [18] | ||
| 3H-Dmt-DALDA | mouse | Brain | 5.64±0.24 | PDSP; [19] | ||
| [Dmt]DALDA | human | cloned | 0.172±0.026 | [19] | ||
| DMAGO | human | cloned | 1.180±0.120 | [19] | ||
| [3H]DAMGO | rat | Brain | 6.55±0.74 | PDSP; [20] | ||
| HEK-μ cells | 2.2±0.5 | [21] | ||||
| [3H]DAMGO | human | BE(2)-C memberanes | 1.02±0.15 | PDSP; [22] | ||
| bovine | adrenals | 1.86 | [23] | |||
| 0.81 | average human | |||||
| DOR | 3H-DSLET | MOUSE | vas deferens | 32.6±3.7 | PDSP; [17] | |
| 3H-Naltrindole | Human | cloned | >10,000 | PDSP; [18] | ||
| [Dmt]DALDA | human | cloned | 1670±40 | [19] | ||
| DMAGO | human | cloned | 1430±20 | [19] | ||
| [3H][Ile5,6]deltorphin II | rat | Brain | 217±19 | PDSP; [20] | ||
| 278±49 | [24] | |||||
| [3H]DPDPE | human | BE(2)-C memberanes | >100 | PDSP; [22] | ||
| [3H]enkephalin | rat | memberane | 69.1±3.2 | [25] | ||
| bovine | adrenals | 147.32 | [23] | |||
| 1545 | average human |
Receptor affinity data for morphine collected from the literature. The columns identify the receptor, the radioligand used in determining affinity, the source species from which the receptor was used, the tissue from which the receptor was used, the Ki value in nanomoles or the IC50 (the molar concentration of an unlabeled agonist or antagonist that inhibits the binding of a radioligand by 50%, [26]) value in nanomoles, and the literature reference from which the data was obtained.
Table 2. Receptor affinity data for THC.
| Receptors | hot ligand | source | tissue | Ki(nM) | Ref |
| CB1 | 3H-BAY 38-7271 | HUMAN | CORTICAL MEMBRANES | 13.7 | PDSP; [27] |
| 3H-BAY 38-7271 | HUMAN | CLONED | 15.3 | PDSP; [28] | |
| 3H-CP-55940 | HUMAN | CLONED | 5.05 | PDSP; [29] | |
| 10.19 | average | ||||
| CB2 | 3H-BAY 38-7271 | HUMAN | CLONED | 25.06 | PDSP; [28] |
| 3H-BAY 38-7271 | HUMAN | CLONED | 22.9 | PDSP; [27] | |
| 3H-CP-55940 | HUMAN | CLONED | 44.9 | [30] | |
| 3H-CP-55940 | HUMAN | CLONED | 3.13 | PDSP; [29] | |
| 16.85 | average | ||||
| sigma | 3H-3-PPP,(+) | RAT | BRAIN | >100,000 | PDSP; [31] |
Receptor affinity data for THC collected from the literature. The columns identify the receptor, the radioligand used in determining affinity, the source species from which the receptor was used, the tissue from which the receptor was used, the Ki value in nanomoles, and the literature reference from which the data was obtained.
New PDSP Binding Assays
For this study, the NIMH-PDSP (http://pdsp.med.unc.edu/) has assayed sixteen phenylalkylamines, eight tryptamines and one ergoline (twenty-two psychedelics and three controls, Fig. 1) against a panel of fifty-one receptors, transporters, and ion channels. The methodology has been described previously by Glennon et al. [12]. Each compound is initially assayed at 10 µM against each receptor, transporter or ion channel (primary assay). Those that induce >50% inhibition (“hit”) are then assayed at 1, 10, 100, 1,000, and 10,000 nM to determine Ki values (secondary assay). Each Ki value (equilibrium dissociation constant, concentration at which 50% of the hot ligand is displaced by the test ligand) is calculated from at least three replicated assays. Details of how individual assays were conducted can be found at the NIMH-PDSP web site: http://pdsp.med.unc.edu/pdspw/binding.php.
Table S2 shows raw Ki data for the current study combined with data collected from the literature for the ten additional compounds; a total of thirty-five drugs and sixty-seven receptors, transporters and ion channels which were assayed. The table has been divided into three sections.
The first section displays forty-two sites at which most compounds were assayed and at least one “hit” (Ki <10,000 nm) was found: 5ht1a (5-HT1A, serotonin-1A receptor), 5ht1b (5-HT1B, serotonin-1B receptor), 5ht1d (5-HT1D, serotonin-1D receptor), 5ht1e (5-HT1E, serotonin-1E receptor), 5ht2a (5-HT2A, serotonin-2A receptor), 5ht2b (5-HT2B, serotonin-2B receptor), 5ht2c (5-HT2C, serotonin-2C receptor), 5ht5a (5-HT5A, serotonin-5A receptor), 5ht6 (5-HT6, serotonin-6 receptor), 5ht7 (5-HT7, serotonin-7 receptor), D1 (D1, dopamine-1 receptor), D2 (D2, dopamine-2 receptor), D3 (D3, dopamine-3 receptor), D4 (D4, dopamine-4 receptor), D5 (D5, dopamine-5 receptor), Alpha1A (α1A, alpha-1A adrenergic receptor), Alpha1B (α1B, alpha-1B adrenergic receptor), Alpha2A (α2A, alpha-2A adrenergic receptor), Alpha2B (α2B, alpha-2B adrenergic receptor), Alpha2C (α2C, alpha-2C adrenergic receptor), Beta1 (β1, beta-1 adrenergic receptor), Beta2 (β2, beta-2 adrenergic receptor), SERT (serotonin transporter), DAT (dopamine transporter), NET (nor epinephrine transporter), Imidazoline1 (I1, imidazoline-1 receptor), Sigma1 (σ1, sigma-1 receptor), Sigma2 (σ2, sigma-2 receptor), DOR (delta opioid receptor), KOR (κ, kappa opioid receptor), MOR (μ, mu opioid receptor), M1 (M1, muscarinic-1 acetylcholine receptor), M2 (M2, muscarinic-2 acetylcholine receptor), M3 (M3, muscarinic-3 acetylcholine receptor), M4 (M4, muscarinic-4 acetylcholine receptor), M5 (M5, muscarinic-5 acetylcholine receptor), H1 (H1, histamine-1 receptor), H2 (H2, histamine-2 receptor), CB1 (CB1, cannabinoid-1 receptor), CB2 (CB2, cannabinoid-2 receptor), Ca+Channel (calcium+ ion channel), NMDA/MK801 (N-methyl D-aspartate glutamate receptor).
The second section displays seven sites at which most compounds were assayed, but at which there were no hits: 5ht3 (serotonin-3 receptor), H3 (histamine-3 receptor), H4 (histamine-4 receptor), V1 (vasopressin-1 receptor), V2 (vasopressin-2 receptor), V3 (vasopressin-3 receptor), GabaA (GABA-A receptor).
The third section displays the remaining eighteen sites, at which only a few compounds were assayed, and no hits were found: GabaB (GABA-B receptor), mGluR1a (mGluR1a metabotropic glutamate receptor), mGluR2 (mGluR2 metabotropic glutamate receptor), mGluR4 (mGluR4 metabotropic glutamate receptor), mGluR5 (mGluR5 metabotropic glutamate receptor), mGluR6 (mGluR6 metabotropic glutamate receptor), mGluR8 (mGluR8 metabotropic glutamate receptor), A2B2 (nicotinic a2/b2 acetylcholine receptor), A2B4 (nicotinic a2/b4 acetylcholine receptor), A3B2 (nicotinic a3/b2 acetylcholine receptor), A3B4 (nicotinic a3/b4 acetylcholine receptor), A4B2 (nicotinic a4/b2 acetylcholine receptor), A4B2** (nicotinic a4/b2** acetylcholine receptor), A4B4 (nicotinic a4/b4 acetylcholine receptor), BZP (a1) (GABA-BZP a1 receptor), EP3 (prostaglandin-3 receptor), MDR 1 (multidrug resistant p-Glycoprotein), PCP (PCP glutamate receptor).
Activity Assays
For the twenty-five compounds of Fig. 1, the NIMH-PDSP also performed activity assays at 5-HT2A and 5-HT2C. The Emax values (maximal activity) are relative to 5-HT (serotonin), measuring Ca++ mobilization. Ca++ flux assays were performed using a FLIPRTETRA. The activity assays were conducted with cell lines which have very high receptor expression levels (e.g. plenty of ‘spare receptors’). Under such conditions partial agonists will have considerable agonist activity. The data represent the mean ± variance of computer-derived estimates from single experiments done in quadruplicate. Thus, the four observations are averaged and a single estimate with error is provided (Table S3).
Sources
The following compounds (Fig. 1) were used in the study:
2C-B, 4-Bromo-2,5-dimethoxyphenethylamine
2C-B-fly, 1-(8-Bromo-2,3,6,7-tetrahydrobenzo[1,2-b;4,5-b′]difuran-4-yl)2-aminoethane
2C-E, 4-Ethyl-2,5-dimethoxyphenethylamine
2C-T-2, 4-Ethylthio-2,5-dimethoxyphenethylamine
ALEPH-2, (±)-4-Ethylthio-2,5-dimethoxyamphetamine
4C-T-2, 4-Ethylthio-2,5-dimethoxyphenylbutylamine
MEM, (±)-2,5-Dimethoxy-4-ethoxyamphetamine
TMA-2: (±)-2,4,5-Trimethoxamphetamine
TMA: (±)-3,4,5-Trimethoxamphetamine
mescaline: 3,4,5-Trimethoxyphenethylamine
DOB: (±)-2,5-Dimethoxy-4-bromoamphetamine
DOI: (±)-2,5-Dimethoxy-4-iodoamphetamine
DOM: (±)-2,5-Dimethoxy-4-methylamphetamine
DOET: (±)-2,5-Dimethoxy-4-ethylamphetamine
MDA: (±)-3,4-Methylenedioxyamphetamine
MDMA: (±)-3,4-Methylenedioxymethamphetamine
DMT: N,N-Dimethyltryptamine
5-MeO-DMT: 5-Methoxy-N,N-dimethyltryptamine
DPT: N,N-Dipropyltryptamine
5-MeO-MIPT: 5-Methoxy-N-methyl-N-isopropyltryptamine
DIPT: N,N-Diisopropyltryptamine
5-MeO-DIPT: 5-Methoxy-N,N-diisopropyltryptamine
6-fluoro-DMT: 6-Fluoro-N,N-dimethyltryptamine
psilocin: 4-Hydroxy-N,N-dimethyltryptamine
lisuride
5-MeO-DMT, and DOI were purchased from Sigma. DOB, DOET, mescaline, TMA, MDA, MDMA, and psilocin were provided as gifts by the National Institute on Drug Abuse Drug Supply Program. 2C-B, 2C-B-fly, MEM, 4C-T-2, 5-MeO-MIPT, 6-fluoro-DMT, TMA-2, and lisuride were provided as gifts by Dave Nichols. DMT and DOM were provided as gifts by Richard Glennon. 2C-E, 2C-T-2, Aleph-2, DIPT, 5-MeO-DIPT, and DPT were provided as gifts by Alexander Shulgin.
Normalization
The raw Ki values are distributed over several orders of magnitude, thus a log transformation is a good first step in the analysis. In addition, higher affinities produce lower Ki values, thus it is valuable to calculate: pKi = −log10(Ki). Higher affinities have higher pKi values, and each unit of pKi value corresponds to one order of magnitude of Ki value. Table S4 presents the raw data transformed into pKi values. Generally, the highest Ki value generated by NIMH-PDSP is 10,000, which produces a pKi value of −4 (although a value of 10,450 was reported for 5-MeO-TMT). For non-PDSP data gathered from the literature, some Ki values greater than 10,000 are reported (i.e. 12,500, 14,142, 22,486, 39,409 and 70,000 for ibogaine).
When the primary assay did not produce >50% inhibition, the Ki value is treated as >10,000. When the primary assay hit, but the secondary assay was not performed, the Ki value is also treated as >10,000. If a secondary assay produced a Ki value significantly greater than 10,000, it is usually also reported as >10,000. The lowest Ki value in the data set of this study is 0.3 (lisuride at 5-HT1A) and the highest value is 70,000 (ibogaine at D3), thus collectively, the data in this study cover nearly six orders of magnitude of Ki values. However, ignoring values reported as >10,000, the Ki values for a single drug in this study never exceed four orders of magnitude in range.
The goal of the normalization used in this study is to factor out potency, in order to allow easy comparison of the multi-receptor affinity profiles of different drugs. The normalization will adjust the highest pKi value for each drug to a value of 4, and set all Ki values reported as >10,000 to a value of zero. Ki values actually measured as greater than 10,000 are not set to zero (i.e. 5-MeO-TMT and ibogaine). We will call this normalized value npKi. Let the maximum pKi value for each drug be called pKiMax. For each individual drug:
If Ki treated as >10,000, then npKi = 0
npKi = 4+pKi−pKiMax
With this normalization:
higher affinities have higher values
affinities too low to be measured will be reported as zero
for each drug, the highest affinity will be set to a value of 4
each unit of npKi value represents one order of magnitude of Ki value
potency is factored out so that drugs of different potencies can be directly compared
This normalization effectively factors out the absolute potency of each drug, and allows us to focus on the relative affinities of each drug at each receptor.
Perceptibility
It will also be seen that many psychedelic drugs interact with a large number of receptors. Fig. 3 shows the ranked distributions of npKi values for DOB and DOI, and the same data is listed below in numerical form (0.00 means Ki >10,000, ND means the data is not available):
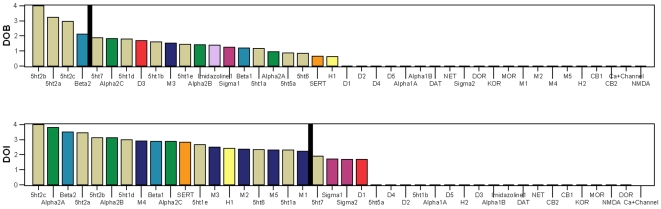
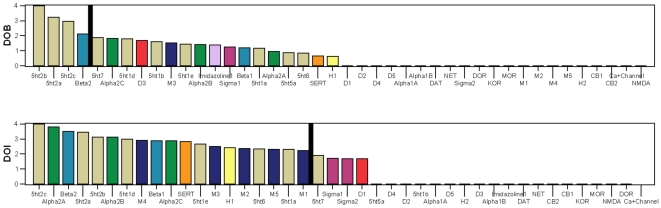
Figure 3. Receptor affinity profiles of DOB and DOI, ordered by decreasing affinity.
The vertical axis is normalized pKi (npKi). Horizontal axis is a list of forty-two receptors, arranged in order of decreasing affinity for each individual drug. Colors correspond to classes of receptors, and are the same as used in Fig. S1. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Receptors to the right of the black bar should be imperceptible, while receptors to the left of the black bar should be perceptible, increasingly so the further left they are.
DOB: 4.00 5ht2b, 3.23 5ht2a, 2.97 5ht2c, 2.11 Beta2, 1.89 5ht7, 1.82 Alpha2C, 1.79 5ht1d, 1.68 D3, 1.62 5ht1b, 1.53 M3, 1.44 5ht1e, 1.41 Alpha2B, 1.39 Imidazoline1, 1.25 Sigma1, 1.21 Beta1, 1.18 5ht1a, 0.96 Alpha2A, 0.87 5ht5a, 0.85 5ht6, 0.66 SERT, 0.63 H1; 0.00: D5, D2, D4, NET, D1, Alpha1B, Sigma2, DOR, KOR, MOR, M1, M2, DAT, M4, M5, Alpha1A, H2, CB2, CB1, Ca+Channel, NMDA
DOI: 4.00 5ht2c, 3.79 Alpha2A, 3.52 Beta2, 3.44 5ht2a, 3.13 Alpha2B, 3.13 5ht2b, 3.00 5ht1d, 2.90 M4, 2.89 Beta1, 2.88 Alpha2C, 2.83 SERT, 2.66 5ht1e, 2.51 M3, 2.42 H1, 2.36 M2, 2.34 5ht6, 2.32 M5, 2.31 5ht1a, 2.23 M1, 1.90 5ht7, 1.73 Sigma1, 1.70 Sigma2, 1.67 D1; 0.00: 5ht1b, DAT, Imidazoline1, NET, 5ht5a, DOR, KOR, MOR, Alpha1B, D2, D3, D4, D5, Alpha1A, H2, CB2, CB1, NMDA; ND: Ca+Channel
For potent compounds like DOB and DOI, it is possible to measure Ki values over nearly a full four orders of magnitude range of affinity. However, not all of these affinities are able to produce perceptible mental effects. As a rule of thumb, 100-fold affinity is considered truly selective. Thus, receptors with npKi values below about 2.0 should not have perceptible mental effects. In Fig. 3, a black vertical bar represent a 100-fold drop in affinity relative to the receptor with the highest affinity, and divides those npKi values greater than 2.0 (on the left) from those 2.0 or less (on the right). This is presumed to be the limit of perceptible receptor interaction. Receptors to the right of the black bar should be imperceptible, while receptors to the left of the black bar should be perceptible, increasingly so the further left they are. In spite of the long tail of affinities, DOB is effectively selective for the three 5-HT2 (serotonin-2) receptors (Beta2 falls at the approximate limit of perceptibility), while DOI by contrast has nineteen receptors in the presumed perceptible range, although they should not all be equally perceptible.
Breadth
An index of the breadth (or inverse of selectivity), B, of the binding profiles of the individual drugs or receptors can be constructed by summing the forty-two npKi values for each drug, or the thirty-five npKi values for each receptor. If a drug were absolutely selective, binding at only one receptor (e.g. salvinorin A), it would have the minimal B value of 4, regardless of the absolute affinity of the drug for its one receptor. If a drug bound with equal affinity to all forty-two receptors, it would have the maximum B value of 4×42 = 168, regardless of its absolute receptor affinities.
It is not clear that a simple sum of npKi values is the best index of breadth. In this method, four receptors with Ki values of 1,000 collectively carry the same weight as one receptor with a Ki value of 1. This may not be a realistic equivalence. Thus we will include three measures of breadth:
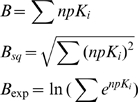 |
Bsq and Bexp give greater weight to higher affinity (lower Ki) values. Regression analysis of receptor affinity vs. potency in humans suggests that Bsq is the most meaningful breadth statistic. Table S5 and Table S6 present the raw Ki data converted into npKi values, for both the individual receptors, and groups of receptors summed using the Bsq statistic.
Proportional Breadth
In addition to looking at the breadth of interaction of individual drugs with multiple receptors, it may be of value to look at an individual drug's interaction with one receptor or group of receptors, as a proportion of the drug's total interaction with all receptors.
In order to compute the proportion for and individual receptor or a group of receptors, we divide the sum of squares of npKi values for the group of receptors, by the sum of squares of npKi values for all receptors:
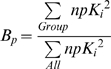 |
For example, to compute this proportion for “5-HT” receptors, we divide the squares of the values in the “5-HT” column of Table S5 (for LSD, 11.132 = 123.9), by the squares of the values in the center column (“Bsq”) of Table 3 (for LSD, 13.122 = 172.1); 123.9/172.1 = 0.719 for LSD. We will call this proportion Bp. The proportional breadth data is displayed in Table S7 and Table S8.
Table 3. Thirty-five drugs arranged in order of decreasing breadth, increasing selectivity.
| B | Drug | Bsq | Drug | Bexp | Drug |
| 67.5 | 6-F-DMT | 14.19 | 6-F-DMT | 6.22 | 6-F-DMT |
| 63.5 | DPT | 13.34 | DMT | 6.16 | DMT |
| 61.6 | DOI | 13.30 | DPT | 6.12 | LSD |
| 56.4 | LSD | 13.21 | DOI | 6.02 | DPT |
| 54.3 | DMT | 13.12 | LSD | 6.02 | DOI |
| 50.5 | lisuride | 11.88 | lisuride | 5.93 | TMA |
| 45.3 | 2C-E | 11.61 | 2C-E | 5.85 | lisuride |
| 45.2 | cis-2a | 11.55 | TMA | 5.81 | 2C-E |
| 45.2 | 5-MeO-MIPT | 11.37 | 2C-B | 5.77 | 2C-B |
| 43.9 | 2C-B | 11.29 | cis-2a | 5.75 | cis-2a |
| 42.1 | psilocin | 11.00 | 5-MeO-MIPT | 5.70 | 5-MeO-MIPT |
| 40.5 | 2C-T-2 | 10.71 | psilocin | 5.60 | psilocin |
| 38.1 | TMA | 10.21 | DIPT | 5.52 | DIPT |
| 37.6 | RR-2b | 9.85 | 5-MeO-DIPT | 5.49 | 5-MeO-DIPT |
| 34.9 | DIPT | 9.80 | RR-2b | 5.44 | 4C-T-2 |
| 34.7 | 5-MeO-DMT | 9.65 | 2C-T-2 | 5.44 | RR-2b |
| 34.5 | DOB | 9.64 | 4C-T-2 | 5.44 | MDMA |
| 33.4 | SS-2c | 9.50 | MDMA | 5.41 | DOET |
| 33.3 | DOET | 9.35 | ibogaine | 5.38 | mescaline |
| 32.5 | 2C-B-fly | 9.32 | DOET | 5.36 | 2C-T-2 |
| 32.1 | 5-MeO-DIPT | 9.00 | 5-MeO-DMT | 5.36 | 5-MeO-DMT |
| 31.2 | ibogaine | 8.85 | SS-2c | 5.35 | ibogaine |
| 31.1 | 4C-T-2 | 8.67 | 2C-B-fly | 5.29 | 2C-B-fly |
| 28.2 | MDMA | 8.67 | mescaline | 5.24 | 5-MeO-TMT |
| 28.1 | DOM | 8.44 | DOB | 5.22 | SS-2c |
| 27.0 | Aleph-2 | 8.41 | 5-MeO-TMT | 5.16 | DOB |
| 22.9 | 5-MeO-TMT | 8.29 | DOM | 5.10 | DOM |
| 21.1 | mescaline | 7.89 | MDA | 5.07 | MDA |
| 20.4 | MDA | 7.30 | Aleph-2 | 4.94 | Aleph-2 |
| 18.4 | EMDT | 7.22 | EMDT | 4.86 | EMDT |
| 13.0 | TMA-2 | 6.60 | TMA-2 | 4.78 | TMA-2 |
| 10.3 | MEM | 5.50 | THC | 4.59 | THC |
| 7.8 | THC | 5.40 | MEM | 4.37 | MEM |
| 6.9 | morphine | 4.63 | morphine | 4.19 | morphine |
| 4.0 | salvinorin A | 4.00 | salvinorin A | 4.00 | salvinorin A |
The thirty-five drugs are arranged in order of decreasing breadth and increasing selectivity, based on the breadth indices B, Bsq, and Bexp. Although the three indices provide different orderings, the orderings are quite similar at the two extremes of the table (greatest and least breadth) where most of the attention is likely to be focused. The drugs with the broadest receptor interactions (least selective) are found at the tops of the columns, and the drugs with the narrowest receptor interactions (most selective) are found at the bottoms of the columns.
Truncated Receptor Profiles
The NIMH-PDSP generally does not measure Ki values greater than 10,000 nm, because at those concentrations, there is a great deal of non-specific binding which invalidates the measurement of receptor affinity. This creates a problem for drugs whose best-hit has a Ki value of greater that 100 nm (TMA, mescaline, TMA-2, DIPT, MDMA, 5-MeO-DIPT, ibogaine). For these drugs, the range of Ki values that can be measured by the NIMH-PDSP is less than the 100-fold presumed perceptible range, and therefore, the lowest measurable npKi value is greater than the presumed limit of perceptibility at 2.00. Table 4 shows for each drug, the lowest Ki value measured (KiMin) which is the best-hit, the best-hit receptor (KiMinR), the theoretically lowest measurable npKi value (npKiLim), the lowest actually measured npKi value (npKiMin), and the receptor where the lowest npKi value was actually measured (npKiMinR). Drugs that have both a KiMin value of greater that 100 nm and an npKiMin value greater than 2.00 have truncated receptor affinity profiles.
Table 4. Truncated receptor profiles for thirty-five drugs.
| Drug | KiMin | KiMinR | npKiLim | npKiMin | npKiMinR |
| mescaline | 745.3 | Alpha2C | 2.87 | 2.92 | Alpha2A |
| TMA | 476.6 | 5ht2b | 2.68 | 2.98 | Alpha2C |
| MDMA | 219.7 | Imidazoline1 | 2.34 | 2.43 | M4 |
| ibogaine | 206 | Sigma2 | 2.31 | 1.47 | D3 |
| TMA-2 | 154.4 | 5ht2b | 2.19 | 2.58 | 5ht2c |
| 5-MeO-DIPT | 132.4 | 5ht1a | 2.12 | 2.15 | Sigma1 |
| DIPT | 120.5 | 5ht1a | 2.08 | 2.51 | 5ht1d |
| MDA | 91 | 5ht2b | 1.96 | 2.15 | 5ht2c |
| DMT | 87.5 | 5ht7 | 1.94 | 2.23 | Sigma1 |
| MEM | 64.5 | 5ht2b | 1.81 | 1.95 | 5ht7 |
| 5-MeO-TMT | 60 | 5ht6 | 1.78 | 1.76 | 5ht5a |
| 4C-T-2 | 58.1 | 5ht2b | 1.76 | 1.77 | 5ht1e |
| DOI | 45.8 | 5ht2c | 1.66 | 1.67 | D1 |
| DPT | 31.8 | 5ht1a | 1.50 | 1.54 | D2 |
| 6-F-DMT | 25.6 | 5ht6 | 1.41 | 1.56 | Sigma2 |
| 2C-E | 25.1 | 5ht2b | 1.40 | 1.88 | D2 |
| EMDT | 16 | 5ht6 | 1.20 | 1.54 | 5ht5a |
| DOET | 14.4 | 5ht1a | 1.16 | 1.17 | Sigma1 |
| 2C-B | 13.5 | 5ht2b | 1.13 | 1.28 | D3 |
| 5-MeO-MIPT | 12.3 | 5ht1a | 1.09 | 1.28 | SERT |
| DOM | 11.7 | 5ht2b | 1.07 | 1.16 | 5ht6 |
| THC | 10.19 | CB1 | 1.01 | 3.78 | CB2 |
| 2C-T-2 | 6 | 5ht2b | 0.78 | 0.81 | Beta1 |
| psilocin | 4.7 | 5ht2b | 0.67 | 1.03 | Alpha2C |
| salvinorin A | 4.3 | KOR | 0.63 | 4 | KOR |
| LSD | 3.9 | 5ht1b | 0.59 | 0.65 | Alpha1B |
| DOB | 3.9 | 5ht2b | 0.59 | 0.63 | H1 |
| cis-2a | 2.3 | 5ht1a | 0.36 | 0.7 | Beta2 |
| RR-2b | 2 | 5ht1b | 0.30 | 0.59 | Alpha1A |
| 5-MeO-DMT | 1.9 | 5ht1a | 0.28 | 0.69 | 5ht2b |
| Aleph-2 | 1.6 | 5ht2b | 0.20 | 0.3 | M5 |
| 2C-B-fly | 0.9 | 5ht2b | −0.05 | 0.12 | D2 |
| morphine | 0.81 | MOR | −0.09 | 0.72 | DOR |
| SS-2c | 0.4 | 5ht1a | −0.40 | 0.2 | H1 |
| lisuride | 0.3 | 5ht1a | −0.52 | 1.08 | Alpha1B |
Table 4 shows for each drug, the lowest Ki value measured (KiMin) which is the best-hit, the best-hit receptor (KiMinR), the theoretically lowest measurable npKi value (npKiLim), the lowest actually measured npKi value (npKiMin), and the receptor where the lowest npKi value was actually measured (npKiMinR). Drugs that have both a KiMin value of greater that 100 nm and an npKiMin value greater than 2.00, have truncated receptor affinity profiles.
Seven drugs have best-hit Ki values of greater that 100 nm: TMA, mescaline, TMA-2, DIPT, MDMA, 5-MeO-DIPT, ibogaine. For these seven drugs, the perceptible receptor profile is truncated due to the methodological limitations of the NIMH-PDSP, to the extent to which the npKiMin value is greater than 2.00. Note that for ibogaine, whose best hit is 206 nm, the npKiMin value is 1.47, indicating that the receptor profile is not fully truncated, because several Ki values above 10,000 nm were gathered from the literature (however, ibogaine has not received a full receptorome screening, and thus its receptor profile must be considered incomplete for other reasons).
Six drugs have both a KiMin value of greater that 100 nm and an npKiMin value greater than 2.00. For 5-MeO-DIPT, the npKiMin value (2.15) is close to the presumed perceptibility limit, thus we can consider its receptor profile to be complete. For TMA-2, DIPT, and MDMA, the npKiMin values (2.58, 2.51, 2.43 respectively) fall in the weak region of the presumed perceptibility range. Although these three receptor profiles are truncated, the missing data may be of little consequence. For TMA and mescaline, the npKiMin values (2.98. 2.92 respectively) fall in the moderate region of the presumed perceptibility range. We must consider these two truncated receptor profiles to be truly incomplete, with potential consequences for our interpretations of the properties of these two drugs. The receptor profiles of some other drugs are incomplete due to holes in the NIMH-PDPS data set. Morphine and THC have not been broadly assayed, and must also be considered to be incomplete.
Results
Normalized Affinity Data
Fig. S1 shows the simplest view of the normalized affinity data. The drugs in Fig. S1 are ordered to correspond roughly to similarity of structure and receptor affinity profiles. Colors correspond to classes of receptors. It can be seen at a glance that most, but not all of the drugs interact strongly with the serotonin receptors (beige), certain drugs interact strongly with the dopamine receptors (red), others with the adrenergic receptors (green), yet others with the histamine receptors (yellow), etc.
Breadth
In Table 3, the thirty-five drugs are arranged in order of decreasing breadth and increasing selectivity, based on the three breadth indices B, Bsq, and Bexp. Although the three indices provide different orderings, the orderings are quite similar at the two extremes of the table (greatest and least breadth) where most of the attention is likely to be focused. The drugs with the broadest receptor interactions (least selective) are found at the tops of the columns, and the drugs with the narrowest receptor interactions (most selective) are found at the bottoms of the columns. Regression analysis suggests that Bsq is the best statistic for combining receptors, therefore the Bsq statistic will be used in most of the breadth analyses to follow. The B, Bsq and Bexp data of Table 3 is presented graphically in Fig. 4.
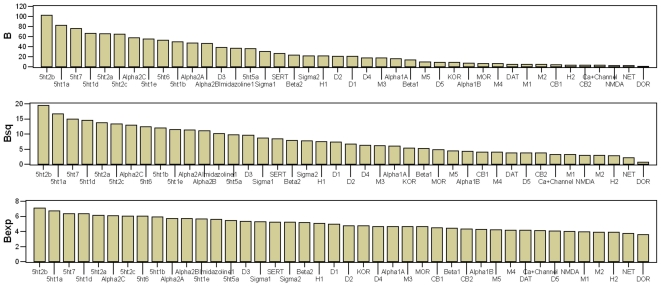
Figure 4. Thirty-five drugs arranged in order of decreasing breadth, increasing selectivity.
The thirty-five drugs are arranged in order of decreasing breadth and increasing selectivity, based on the breadth indices B, Bsq, and Bexp. Although the three indices provide different orderings, the orderings are quite similar at the two extremes of the table (greatest and least breadth) where most of the attention is likely to be focused. The drugs with the broadest receptor interactions (least selective) are found at the left of the figure, and the drugs with the narrowest receptor interactions (most selective) are found at the right of the figure.
Profiles of Drugs
The relative breadth or selectivity of the thirty-five drugs is nicely visualized in Fig. S2, in which for each drug, the bars representing forty-two receptors are arranged in order of decreasing size. The drugs are arranged in order of decreasing breadth, based on the Bsq values of Table 3 and Fig. 4. Drugs at the top of the figure have the broadest receptor interactions (least selective), while drugs at the bottom of the figure have the narrowest receptor interactions (most selective). Colors correspond to classes of receptors, and are the same as used in Fig. S1. Scanning Fig. S2 from top to bottom shows the variation in breadth of receptor interactions between drugs. It can also be seen that some distributions are relatively convex (e.g. DMT & LSD), while others are relatively concave (e.g. DOB & 5-MeO-DMT). Convexity tends to increase breadth, while concavity tends to decrease breadth.
It is also useful to present the npKi data of Fig. S2 in numerical format. In the listing below, the npKi values for each drug are arranged in decreasing order. A value of 0.00 means that the Ki value was measured as >10,000 nm. “ND” indicates that the data is not available. The 5-HT2A and 5-HT2C receptors are also highlighted in bold font for easier location. npKi values below about 2.0 should be imperceptible, while values above about 2.0 should be perceptible, and the higher the npKi value, the more perceptible a receptor should be.
6-F-DMT: 4.00 5ht6, 3.93 5ht2b, 3.80 5ht7, 3.74 H1, 3.66 5ht1d, 3.25 SERT, 3.24 Alpha2C, 3.17 Alpha1A, 3.07 5ht1b, 2.99 Alpha2B, 2.81 5ht1a, 2.74 5ht1e, 2.67 D1, 2.62 D2, 2.58 5ht2c, 2.47 5ht2a, 2.47 D3, 2.45 Imidazoline1, 2.44 H2, 2.43 5ht5a, 2.25 D4, 1.61 D5, 1.57 Sigma1, 1.56 Sigma2; 0.00: DAT, Beta1, NET, Alpha2A, CB2, CB1, Ca+Channel, Beta2, M2, M3, M4, M5, M1, Alpha1B, NMDA; ND: DOR, MOR, KOR
DMT: 4.00 5ht7, 3.97 5ht1d, 3.91 5ht2b, 3.53 Alpha2B, 3.53 Alpha2C, 3.51 D1, 3.42 5ht2c, 3.28 5ht1e, 3.25 5ht6, 3.16 5ht5a, 3.13 Imidazoline1, 2.95 Alpha1B, 2.75 Alpha2A, 2.70 Alpha1A, 2.58 5ht2a, 2.37 SERT, 2.23 Sigma1; 0.00: 5ht1a, D4, D5, Beta1, D2, D3, DAT, NET, 5ht1b, Beta2, Sigma2, CB2, KOR, Ca+Channel, M1, M2, M3, M4, M5, H2, CB1; ND: H1, DOR, MOR, NMDA
DPT: 4.00 5ht1a, 3.88 5ht2b, 3.41 H1, 3.31 SERT, 3.05 5ht7, 2.97 Imidazoline1, 2.97 Alpha2B, 2.90 Sigma1, 2.86 Alpha1B, 2.84 Alpha2A, 2.79 Alpha2C, 2.71 5ht1d, 2.57 5ht1b, 2.56 Alpha1A, 2.37 D3, 2.33 DAT, 2.31 5ht2c, 2.20 D4, 2.13 5ht1e, 2.09 5ht2a, 2.04 Sigma2, 1.86 5ht5a, 1.85 5ht6, 1.54 D2; 0.00: D1, D5, NET, Beta1, DOR, KOR, MOR, Beta2, M2, M3, M4, M5, M1, H2, CB2, CB1, Ca+Channel, NMDA
DOI: 4.00 5ht2c, 3.79 Alpha2A, 3.52 Beta2, 3.44 5ht2a, 3.13 Alpha2B, 3.13 5ht2b, 3.00 5ht1d, 2.90 M4, 2.89 Beta1, 2.88 Alpha2C, 2.83 SERT, 2.66 5ht1e, 2.51 M3, 2.42 H1, 2.36 M2, 2.34 5ht6, 2.32 M5, 2.31 5ht1a, 2.23 M1, 1.90 5ht7, 1.73 Sigma1, 1.70 Sigma2, 1.67 D1; 0.00: 5ht1b, DAT, Imidazoline1, NET, 5ht5a, DOR, KOR, MOR, Alpha1B, D2, D3, D4, D5, Alpha1A, H2, CB2, CB1, NMDA; ND: Ca+Channel
LSD: 4.00 5ht1b, 3.77 5ht7, 3.75 5ht6, 3.73 5ht1a, 3.70 5ht1d, 3.64 5ht5a, 3.54 5ht2a, 3.16 D3, 3.11 5ht2b, 3.11 5ht2c, 2.93 Alpha2A, 2.62 5ht1e, 2.55 D2, 2.39 D4, 2.34 D1, 2.05 D5, 1.54 Alpha1A, 1.40 H1, 1.39 Beta1, 1.05 Beta2, 0.65 Alpha1B; 0.00: KOR, DOR, DAT, SERT, MOR, NET; ND: Sigma2, Alpha2B, Alpha2C, Imidazoline1, M1, M2, M3, M4, M5, Sigma1, H2, CB2, CB1, Ca+Channel, NMDA
lisuride: 4.00 5ht1a, 3.88 Alpha2C, 3.78 Alpha2B, 3.22 Alpha2A, 3.01 5ht2b, 2.99 5ht5a, 2.90 D4, 2.74 5ht2a, 2.65 D2, 2.64 5ht7, 2.61 5ht6, 2.56 Beta1, 2.27 5ht1b, 2.09 Alpha1A, 1.93 Beta2, 1.83 5ht1e, 1.59 D5, 1.42 H2, 1.34 D3, 1.08 Alpha1B; 0.00: 5ht1d, 5ht2c, D1, DAT, NET, Imidazoline1, M1, SERT, CB2, KOR, MOR, M3, M2, M5, M4, CB1, Ca+Channel; ND: Sigma1, H1, Sigma2, DOR, NMDA
2C-E: 4.00 5ht2b, 3.76 5ht2a, 3.54 5ht1d, 3.44 Alpha2C, 3.38 5ht2c, 3.00 5ht1b, 2.91 Alpha2B, 2.91 5ht1a, 2.77 5ht7, 2.71 Alpha2A, 2.60 5ht1e, 2.27 D3, 2.16 M5, 1.99 M3, 1.93 5ht6, 1.88 D2; 0.00: D1, Alpha1A, Alpha1B, 5ht5a, Beta1, M1, SERT, D4, NET, Imidazoline1, H1, Sigma2, DOR, KOR, MOR, NMDA, M2, DAT, M4, D5, CB2, H2, Ca+Channel, CB1; ND: Sigma1, Beta2
TMA: 4.00 5ht2b, 3.95 Sigma2, 3.95 Sigma1, 3.80 5ht7, 3.45 5ht1a, 3.36 Alpha2A, 3.22 5ht1b, 3.20 5ht1d, 3.15 5ht1e, 3.02 5ht2c, 2.98 Alpha2C; 0.00: 5ht2a, 5ht6, D4, D1, Alpha1A, D2, D3, Alpha2B, D5, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, Alpha1B, 5ht5a, DOR, KOR, MOR, NMDA, M2, CB2, CB1, M5, H1, H2; ND: M3, M4, Ca+Channel, M1
2C-B: 4.00 5ht2b, 3.71 5ht1d, 3.69 5ht2a, 3.18 5ht2c, 3.12 Alpha2C, 3.11 5ht1b, 3.05 5ht1e, 2.81 5ht7, 2.75 5ht1a, 2.64 Alpha2A, 2.63 5ht6, 2.31 Alpha2B, 2.22 M3, 1.80 Imidazoline1, 1.60 D2, 1.28 D3; 0.00: D1, 5ht5a, Alpha1B, D5, NMDA, M1, SERT, D4, NET, Alpha1A, Sigma1, Sigma2, DOR, KOR, MOR, H1, M2, DAT, M4, M5, CB2, H2, CB1; ND: Beta2, Ca+Channel, Beta1
cis-2a: 4.00 5ht1a, 3.79 5ht1b, 3.46 D3, 3.30 5ht7, 3.25 5ht6, 3.15 5ht5a, 2.89 5ht2a, 2.72 5ht2b, 2.67 D2, 2.49 D4, 2.34 5ht2c, 2.07 5ht1e, 2.00 D1, 1.76 Beta1, 1.66 H1, 1.64 Alpha1A, 1.36 D5, 0.70 Beta2; 0.00: DOR, SERT, MOR, KOR, NET, DAT, Alpha1B; ND: 5ht1d, Alpha2A, Sigma2, Alpha2B, Alpha2C, Imidazoline1, M1, M2, M3, M4, M5, Sigma1, H2, CB2, CB1, Ca+Channel, NMDA
5-MeO-MIPT: 4.00 5ht1a, 3.79 5ht7, 3.74 5ht1d, 3.32 5ht2b, 2.98 5ht6, 2.85 Alpha2A, 2.61 5ht1b, 2.44 5ht2a, 2.29 Alpha2C, 2.15 Imidazoline1, 2.13 Sigma2, 2.11 5ht5a, 1.86 Alpha2B, 1.75 5ht2c, 1.70 D3, 1.55 5ht1e, 1.41 H1, 1.29 D4, 1.28 SERT; 0.00: D2, Alpha1B, D5, D1, Beta2, NET, DAT, Sigma1, Beta1, DOR, KOR, MOR, M1, M2, M3, M4, M5, Alpha1A, H2, CB2, NMDA, Ca+Channel; ND: CB1
Psilocin: 4.00 5ht2b, 3.40 5ht1d, 3.37 D1, 3.03 5ht1e, 2.88 5ht1a, 2.83 5ht5a, 2.82 5ht7, 2.82 5ht6, 2.67 D3, 2.52 5ht2c, 2.19 5ht1b, 2.14 5ht2a, 1.77 Imidazoline1, 1.74 SERT, 1.57 Alpha2B, 1.36 Alpha2A, 1.03 Alpha2C; 0.00: D2, Alpha1B, D5, D4, Beta2, Beta1, DAT, NET, Alpha1A, Sigma1, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, Ca+Channel, H1, H2, CB2, CB1; ND: M5, NMDA
DIPT: 4.00 5ht1a, 3.53 Imidazoline1, 3.48 5ht2b, 2.98 SERT, 2.83 Sigma1, 2.68 Alpha2C, 2.65 Sigma2, 2.62 Alpha2B, 2.56 D3, 2.55 5ht7, 2.53 H1, 2.51 5ht1d; 0.00: 5ht2a, D4, 5ht5a, D1, D2, Alpha2A, 5ht6, D5, Beta1, Beta2, 5ht2c, DAT, NET, 5ht1b, Alpha1B, 5ht1e, DOR, KOR, MOR, M1, M2, M3, M4, M5, Alpha1A, H2, CB2, CB1, Ca+Channel, NMDA
5-MeO-DIPT: 4.00 5ht1a, 3.91 5ht2b, 3.24 Imidazoline1, 3.03 5ht7, 2.89 5ht1d, 2.72 SERT, 2.66 Alpha2C, 2.64 Sigma2, 2.41 5ht1b, 2.40 Alpha2B, 2.15 Sigma1; 0.00: 5ht5a, 5ht2a, D4, D3, D1, D2, Alpha2A, 5ht6, D5, Beta1, Beta2, 5ht2c, DAT, NET, Alpha1A, Alpha1B, 5ht1e, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, CB2, CB1, Ca+Channel, NMDA
RR-2b: 4.00 5ht1b, 3.59 5ht1a, 3.20 5ht5a, 3.05 5ht1d, 2.81 5ht7, 2.52 5ht6, 2.40 D3, 2.16 5ht1e, 2.08 D4, 1.88 D2, 1.81 5ht2b, 1.77 D1, 1.63 5ht2c, 1.53 D5, 1.46 5ht2a, 1.14 H1, 0.59 Alpha1A; 0.00: SERT, DOR, KOR, MOR, Alpha1B, NET, DAT; ND: Alpha2A, Imidazoline1, Beta2, Sigma2, Alpha2B, Alpha2C, Beta1, M1, M2, M3, M4, M5, Sigma1, H2, CB2, CB1, Ca+Channel, NMDA
2C-T-2: 4.00 5ht2b, 3.18 5ht2a, 3.05 5ht2c, 2.84 5ht1d, 2.56 Alpha2C, 2.20 5ht1a, 2.16 5ht1e, 1.94 M3, 1.92 Alpha2A, 1.84 5ht1b, 1.79 Alpha2B, 1.79 5ht7, 1.70 Beta2, 1.64 5ht6, 1.60 M5, 1.51 D3, 1.46 Imidazoline1, 1.33 D2, 1.19 Sigma1, 0.81 Beta1; 0.00: D1, 5ht5a, SERT, D4, NET, Alpha1A, Alpha1B, Sigma2, DOR, KOR, MOR, M1, M2, DAT, M4, D5, H1, H2, CB2, CB1, Ca+Channel, NMDA
4C-T-2: 4.00 5ht2b, 3.67 Beta2, 3.33 5ht2a, 3.09 5ht2c, 3.05 Sigma1, 2.79 Imidazoline1, 2.66 D3, 2.56 5ht5a, 2.18 5ht7, 2.04 5ht1a, 1.77 5ht1e; 0.00: 5ht6, D1, D4, D5, Alpha1A, Alpha1B, Alpha2A, Alpha2B, Alpha2C, Beta1, D2, SERT, DAT, NET, 5ht1b, 5ht1d, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, CB2, CB1, Ca+Channel, NMDA
MDMA: 4.00 Imidazoline1, 3.64 5ht2b, 3.26 Ca+Channel, 3.21 Alpha2C, 3.09 Alpha2B, 3.07 M3, 2.94 Alpha2A, 2.54 M5, 2.43 M4; 0.00: 5ht2c, 5ht1d, D2, 5ht1e, 5ht1a, 5ht2a, Alpha1A, Alpha1B, 5ht5a, 5ht6, 5ht7, D1, Beta2, SERT, DAT, NET, 5ht1b, H1, H2, D3, KOR, Beta1, M1, M2, D5, D4, CB1, NMDA, MOR; ND: DOR, Sigma2, CB2, Sigma1
Ibogaine: 4.00 Sigma2, 3.57 SERT, 3.02 DAT, 3.01 NMDA, 2.88 KOR, 2.67 MOR, 2.55 Sigma1, 2.22 M3, 2.16 5ht2a, 1.96 M1, 1.72 M2, 1.47 D3; 0.00: DOR, 5ht1b, 5ht1d, 5ht1a, H1, 5ht2c, D2, D1, Beta1; ND: Alpha2C, D5, D4, Alpha2B, Imidazoline1, NET, Alpha2A, 5ht5a, 5ht6, 5ht7, Alpha1B, 5ht1e, 5ht2b, M4, M5, Alpha1A, H2, CB2, CB1, Ca+Channel, Beta2
DOET: 4.00 5ht1a, 3.72 5ht2a, 3.70 5ht2b, 3.13 5ht2c, 2.40 Alpha2B, 2.07 5ht7, 2.05 Alpha2A, 2.00 Alpha2C, 1.82 Beta2, 1.71 5ht1b, 1.61 5ht1e, 1.40 Beta1, 1.34 5ht1d, 1.18 Sigma2, 1.17 Sigma1; 0.00: D4, Alpha1A, D3, 5ht6, D5, D1, D2, SERT, DAT, NET, Imidazoline1, Alpha1B, 5ht5a, DOR, KOR, MOR, NMDA, M2, CB2, CB1, M5, H1, H2; ND: M3, M4, Ca+Channel, M1
5-MeO-DMT: 4.00 5ht1a, 3.69 5ht7, 3.48 5ht1d, 2.73 5ht6, 2.41 5ht1b, 2.38 D1, 1.84 5ht5a, 1.72 5ht1e, 1.58 D3, 1.57 Alpha2C, 1.55 5ht2c, 1.00 Alpha2A, 0.98 5ht2a, 0.97 SERT, 0.88 Imidazoline1, 0.86 Alpha2B, 0.82 NET, 0.78 D4, 0.73 D2, 0.69 5ht2b; 0.00: Alpha1B, Beta2, Beta1, DAT, D5, Alpha1A, Sigma1, Sigma2, CB2, KOR, Ca+Channel, M1, M2, M3, M4, M5, H2, CB1; ND: H1, DOR, MOR, NMDA
SS-2c: 4.00 5ht1a, 3.22 5ht1b, 2.82 D3, 2.45 5ht7, 2.44 5ht6, 2.32 5ht2a, 2.17 5ht5a, 2.17 5ht2b, 2.03 5ht2c, 1.74 D2, 1.72 Beta1, 1.62 D4, 1.16 5ht1e, 1.14 D1, 1.00 D5, 0.67 Alpha1A, 0.57 Beta2, 0.20 H1; 0.00: DOR, SERT, MOR, KOR, NET, DAT, Alpha1B; ND: 5ht1d, Alpha2A, Sigma2, Alpha2B, Alpha2C, Imidazoline1, M1, M2, M3, M4, M5, Sigma1, H2, CB2, CB1, Ca+Channel, NMDA
2C-B-fly: 4.00 5ht2b, 3.81 5ht1d, 2.93 5ht2c, 2.89 5ht2a, 1.91 5ht1e, 1.79 5ht1a, 1.79 Alpha2A, 1.78 5ht6, 1.69 5ht1b, 1.59 Alpha2C, 1.42 M3, 1.24 M4, 1.17 5ht7, 1.16 Alpha2B, 1.15 M1, 1.01 M5, 0.65 M2, 0.26 D1, 0.19 H1, 0.12 D2; 0.00: Beta1, Alpha1B, 5ht5a, DAT, NET, Imidazoline1, Sigma1, Sigma2, DOR, KOR, MOR, Beta2, SERT, CB2, D4, D5, Alpha1A, H2, CB1; ND: D3, Ca+Channel, NMDA
Mescaline: 4.00 Alpha2C, 3.97 5ht2b, 3.61 5ht1a, 3.44 Imidazoline1, 3.16 5ht1e, 2.92 Alpha2A; 0.00: 5ht2a, 5ht2c, 5ht6, 5ht1d, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, 5ht5a, Alpha2B, 5ht7, Beta1, Beta2, SERT, DAT, NET, 5ht1b, Sigma1, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, CB2, CB1, Ca+Channel, NMDA
DOB: 4.00 5ht2b, 3.23 5ht2a, 2.97 5ht2c, 2.11 Beta2, 1.89 5ht7, 1.82 Alpha2C, 1.79 5ht1d, 1.68 D3, 1.62 5ht1b, 1.53 M3, 1.44 5ht1e, 1.41 Alpha2B, 1.39 Imidazoline1, 1.25 Sigma1, 1.21 Beta1, 1.18 5ht1a, 0.96 Alpha2A, 0.87 5ht5a, 0.85 5ht6, 0.66 SERT, 0.63 H1; 0.00: D5, D2, D4, NET, D1, Alpha1B, Sigma2, DOR, KOR, MOR, M1, M2, DAT, M4, M5, Alpha1A, H2, CB2, CB1, Ca+Channel, NMDA
5-MeO-TMT: 4.00 5ht6, 3.62 5ht7, 3.48 5ht1a, 3.38 5ht1d, 2.52 5ht1e, 2.17 5ht2c, 1.97 NET, 1.76 5ht5a; 0.00: 5ht2a, DAT, D1, D2, D3, D4, D5, H1, H2, SERT, DOR, KOR, MOR; ND: Beta2, Alpha2A, 5ht2b, 5ht1b, Imidazoline1, Sigma1, Sigma2, Alpha2B, Alpha2C, Beta1, M1, M2, M3, M4, M5, Alpha1A, Alpha1B, CB2, CB1, Ca+Channel, NMDA
DOM: 4.00 5ht2b, 3.38 Beta2, 2.75 5ht1d, 2.36 5ht2a, 2.30 Alpha2A, 2.13 Alpha2B, 2.10 Alpha2C, 1.87 5ht7, 1.56 Alpha1A, 1.52 5ht1e, 1.51 5ht1a, 1.47 5ht2c, 1.16 5ht6; 0.00: D3, 5ht1b, D1, Alpha1B, 5ht5a, D4, D5, Beta1, D2, SERT, DAT, NET, Imidazoline1, Sigma1, Sigma2, DOR, KOR, MOR, M1, M2, CB2, CB1, M5, H1, H2, NMDA; ND: M3, Ca+Channel, M4
MDA: 4.00 5ht2b, 3.60 Alpha2C, 3.12 Alpha2B, 2.74 Alpha2A, 2.41 5ht7, 2.38 5ht1a, 2.15 5ht2c; 0.00: 5ht2a, 5ht1e, 5ht1d, D1, D2, D3, D4, 5ht1b, Alpha1A, Alpha1B, 5ht5a, 5ht6, D5, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, CB2, H2, M5, KOR, M2, CB1; ND: M1, M3, Sigma1, MOR, H1, Sigma2, DOR, M4, Ca+Channel, NMDA
Aleph-2: 4.00 5ht2b, 2.79 Beta2, 2.50 5ht2c, 2.42 5ht2a, 1.83 Sigma1, 1.70 Imidazoline1, 1.41 D3, 1.08 5ht7, 1.08 SERT, 1.06 Alpha2C, 1.02 5ht1d, 0.98 5ht1a, 0.92 M3, 0.90 5ht1b, 0.74 Alpha2B, 0.72 5ht6, 0.71 5ht1e, 0.44 Alpha2A, 0.37 Beta1, 0.30 M5; 0.00: D1, D2, 5ht5a, D4, NET, Alpha1A, Alpha1B, Sigma2, DOR, KOR, MOR, M1, M2, DAT, M4, D5, H1, H2, CB2, CB1, Ca+Channel, NMDA
EMDT: 4.00 5ht6, 2.97 5ht1a, 2.74 5ht1d, 2.73 5ht7, 2.49 5ht1e, 1.95 5ht2c, 1.54 5ht5a; 0.00: 5ht2a, DAT, D5, D1, D2, D3, D4, NET, H1, H2, SERT, DOR, KOR, MOR; ND: Beta2, Alpha2A, 5ht2b, 5ht1b, Imidazoline1, Sigma1, Sigma2, Alpha2B, Alpha2C, Beta1, M1, M2, M3, M4, M5, Alpha1A, Alpha1B, CB2, CB1, Ca+Channel, NMDA
TMA-2: 4.00 5ht2b, 3.42 5ht2a, 3.04 H1, 2.58 5ht2c; 0.00: 5ht1b, 5ht1a, 5ht1e, 5ht5a, 5ht6, 5ht7, D1, D2, D3, D4, D5, 5ht1d, Alpha1B, Alpha2A, Alpha2B, Alpha2C, Beta1, Beta2, SERT, DAT, NET, M5, Alpha1A, H2, M2; ND: KOR, DOR, M1, MOR, M3, M4, Imidazoline1, Sigma1, Sigma2, CB2, CB1, Ca+Channel, NMDA
THC: 4.00 CB1, 3.78 CB2; ND: 5ht2a, 5ht2c, 5ht1b, 5ht1d, 5ht1e, 5ht2b, 5ht1a, 5ht7, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, 5ht5a, 5ht6, Alpha2C, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, Sigma1, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, Alpha2A, Alpha2B, Ca+Channel, NMDA
MEM: 4.00 5ht2b, 2.21 5ht2a, 2.10 Sigma1, 1.95 5ht7; 0.00: 5ht2c, 5ht1a, 5ht1e, 5ht5a, 5ht6, 5ht1b, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, Alpha2A, Alpha2B, Alpha2C, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, 5ht1d, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, CB2, CB1, Ca+Channel, NMDA
Morphine: 4.00 MOR, 2.21 KOR, 0.72 DOR; ND: 5ht2c, 5ht2a, 5ht1d, 5ht1e, 5ht2b, 5ht1a, 5ht1b, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, Alpha2A, Alpha2B, Alpha2C, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, Sigma1, Sigma2, 5ht5a, 5ht6, 5ht7, M1, M2, M3, M4, M5, H1, H2, CB2, CB1, Ca+Channel, NMDA
Salvinorin A: 4.00 KOR; 0.00: 5ht2a, 5ht2b, 5ht2c, 5ht1b, 5ht1d, 5ht1e, 5ht5a, 5ht1a, 5ht7, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, SERT, DOR, 5ht6, Beta1, Beta2, M2, DAT, M4, M5, H1, M1, M3, MOR; ND: Alpha2A, Alpha2C, Sigma2, Alpha2B, NET, Imidazoline1, Sigma1, H2, CB2, CB1, Ca+Channel, NMDA
Groups of Related Receptors
In addition to looking at breadth at the full complement of forty-two receptors with which the drugs interact, we can use the Bsq statistic to look at the participation of selected groups of receptors (Table S5). 5-HT6 and 5-HT7 are grouped because they share a common G-protein, Gs [13]. The same data is also presented in Table S6, which also includes interactions with individual receptors. Tables S5 and S6 allow us to easily identify drugs with the greatest or least interaction with an individual receptor or any group of related receptors.
For example, among drugs with measurable affinity at a 5-HT2 receptor, 5-MeO-DMT has the weakest interaction with both the three 5-HT2 receptors and the two paradigmatic 5-HT2 receptors (5-HT2A/C). This can be seen in that 5-MeO-DMT has the lowest non-zero value in all four of the columns: “5-HT2,” “5-HT2max,” “5-HT2A, 5-HT2C,” and “5-HT2A/Cmax.” Meanwhile, the drugs 2C-E and DOI rise to the top of the same four columns, indicating that they have the strongest interactions with the 5-HT2 receptors (fourteen drugs tie for the top value in the 5-HT2max column because thirteen drugs have 5-HT2B as their best hit and one has 5-HT2C as its best hit). LSD has the strongest interaction collectively with the five dopamine receptors (D1, D2, D3, D4, D5), the ten assayed 5-HT receptors, and the four assayed 5-HT1 (serotonin-1) receptors. DMT has the strongest interaction with any single dopamine receptor (Dmax), and is the only drug to have its best hit at the 5-HT7 receptor (Table S6). Mescaline is the only drug to have its best hit at an adrenergic receptor (α2C, Table S6). Ibogaine is the only drug to have its best hit at a sigma receptor. MDMA is the only drug to have its best hit at an imidazoline receptor (Table S6). These observations likely provide clues to the qualitative diversity of these drugs.
Proportional Breadth
There is yet another way to look at the participation of the sub-sets of receptors. It may be relevant to consider the participation of a sub-set of receptors in proportion to the participation of all receptors (Table S7). For this we use the proportional breadth statistic (Bp) described in the methods section. The same data is also displayed in Table S8.
The proportional breadth statistic, Bp, is strongly influenced by the degree of overall breadth of the drug (Bsq 2), as this determines the denominator in calculating the proportion. Therefore, we find MEM at the top of the 5-HT column because it is almost completely selective for a single receptor, 5-HT2B. Similarly, TMA-2 is at the top of the 5-HT2 columns because it is highly selective for the 5-HT2 receptors, and EMDT is at the top of the “5-HT6, 5-HT7” column because it is highly selective for the 5-HT6 receptor. Due to their high degree of selectivity, MEM, TMA-2, and EMDT all have small denominators in calculating the Bp statistic.
More interesting cases involve less selective drugs. For example, DMT is a very promiscuous drug, yet it falls at the top of the “α1A, α1B” column. MDA falls at the top of the “Adrenergic” and “α2A, α2B, α2C” columns. 5-MeO-DMT appears third from the top of the “5-HT6, 5-HT7” column and second from the top of the “5-HT7” column (Table S8). Ibogaine appears at the top of the “σ1, σ2” column. Although fairly selective, TMA-2's position at the top of the “H1, H2” column represents an important aspect of its pharmacology, likewise for DOM's position at the top of the “β1, β2” column.
Profiles of Receptors
The B statistics can also be used to look at the relative role played by the various receptors in the pharmacology of the entire set of drugs of this study. In this case, for each receptor, we sum the npKi values at each receptor across each of the thirty-five drugs. In Table 5 we can see the rankings from the three B statistics. The data in Table 5 can also be represented graphically (Fig. 5). The three B statistics provide a very consistent ranking for the top seven receptors. In descending order of importance: 5-HT2B, 5-HT1A, 5-HT7, 5-HT1D, 5-HT2A, 5-HT2C, α2C (with some play between the sixth and seventh positions). This set of top receptors would be a good place to look for receptors other than 5-HT2A and 5-HT2C, which play an important role in the actions of psychedelic drugs.
Table 5. Forty-two receptors arranged in order of decreasing interaction with the full set of thirty-five drugs.
| B | Receptor | Bsq | Receptor | Bexp | Receptor |
| 102.400 | 5ht2b | 19.479 | 5ht2b | 7.094 | 5ht2b |
| 82.564 | 5ht1a | 16.634 | 5ht1a | 6.709 | 5ht1a |
| 75.736 | 5ht7 | 14.931 | 5ht7 | 6.348 | 5ht7 |
| 66.217 | 5ht1d | 14.574 | 5ht1d | 6.348 | 5ht1d |
| 65.462 | 5ht2a | 13.806 | 5ht2a | 6.153 | 5ht2a |
| 64.798 | 5ht2c | 13.352 | 5ht2c | 6.071 | Alpha2C |
| 58.020 | Alpha2C | 12.989 | Alpha2C | 6.037 | 5ht2c |
| 55.055 | 5ht1e | 12.430 | 5ht6 | 6.034 | 5ht6 |
| 53.236 | 5ht6 | 11.951 | 5ht1b | 5.930 | 5ht1b |
| 49.634 | 5ht1b | 11.508 | 5ht1e | 5.722 | Alpha2A |
| 47.526 | Alpha2A | 11.304 | Alpha2A | 5.709 | Alpha2B |
| 46.760 | Alpha2B | 11.138 | Alpha2B | 5.666 | 5ht1e |
| 38.823 | D3 | 10.097 | Imidazoline1 | 5.608 | Imidazoline1 |
| 36.702 | Imidazoline1 | 9.741 | 5ht5a | 5.450 | 5ht5a |
| 36.079 | 5ht5a | 9.561 | D3 | 5.348 | D3 |
| 30.487 | Sigma1 | 8.657 | Sigma1 | 5.271 | Sigma1 |
| 26.764 | SERT | 8.448 | SERT | 5.252 | SERT |
| 23.237 | Beta2 | 7.831 | Beta2 | 5.248 | Sigma2 |
| 21.839 | Sigma2 | 7.808 | Sigma2 | 5.170 | Beta2 |
| 21.769 | H1 | 7.452 | H1 | 5.080 | H1 |
| 21.302 | D2 | 7.296 | D1 | 4.985 | D1 |
| 21.107 | D1 | 6.697 | D2 | 4.732 | D2 |
| 17.995 | D4 | 6.278 | D4 | 4.732 | KOR |
| 17.821 | M3 | 6.205 | M3 | 4.649 | D4 |
| 16.510 | Alpha1A | 6.043 | Alpha1A | 4.644 | Alpha1A |
| 14.117 | Beta1 | 5.401 | KOR | 4.643 | M3 |
| 9.935 | M5 | 5.209 | Beta1 | 4.626 | MOR |
| 9.137 | D5 | 4.812 | MOR | 4.484 | CB1 |
| 9.089 | KOR | 4.492 | M5 | 4.425 | Beta1 |
| 7.540 | Alpha1B | 4.297 | Alpha1B | 4.355 | CB2 |
| 6.674 | MOR | 4.000 | CB1 | 4.282 | Alpha1B |
| 6.563 | M4 | 3.978 | M4 | 4.243 | M5 |
| 5.344 | DAT | 3.810 | DAT | 4.173 | M4 |
| 5.334 | M1 | 3.809 | D5 | 4.154 | DAT |
| 4.727 | M2 | 3.782 | CB2 | 4.097 | Ca+Channel |
| 4.000 | CB1 | 3.263 | Ca+Channel | 4.059 | D5 |
| 3.861 | H2 | 3.181 | M1 | 3.995 | NMDA |
| 3.783 | CB2 | 3.013 | NMDA | 3.942 | M1 |
| 3.263 | Ca+Channel | 2.992 | M2 | 3.914 | M2 |
| 3.013 | NMDA | 2.824 | H2 | 3.884 | H2 |
| 2.796 | NET | 2.138 | NET | 3.749 | NET |
| 0.720 | DOR | 0.720 | DOR | 3.585 | DOR |
The forty-two receptors are arranged in order of decreasing interaction with the full set of thirty-five drugs, based on the breadth statistics B, Bsq. and Bexp. The receptors with the greatest interactions are found at the tops of the columns, and the receptors with the least interactions are found at the bottoms of the columns.
Figure 5. Forty-two receptors arranged in order of decreasing interaction with the full set of thirty-five drugs.
The forty-two receptors are arranged in order of decreasing interaction with the full set of thirty-five drugs, based on the breadth statistics, B, Bsq. and Bexp. The receptors with the greatest interactions are found at the left of the figures, and the receptors with the least interactions are found at the right of the figures. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity at each drug. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Drugs to the right of the black bar should have imperceptible interactions with the receptor, while drugs to the left of the black bar should have perceptible interactions with the receptor, increasingly so the further left they are.
Fig. S3 is a more detailed graphical view of data presented in Table 5. The receptors are presented in order of decreasing breadth (Bsq). The figures for each individual receptor provide information on the relative importance of each receptor at each drug, similar to that in Table S5 and Table S6. Receptors at the top of the figure have the broadest interactions with the thirty-five drugs, while receptors at the bottom of the figure have the narrowest interactions with the thirty-five drugs. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity at each drug. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Drugs to the right of the black bar should have imperceptible interactions with the receptor, while drugs to the left of the black bar should have perceptible interactions with the receptor, increasingly so the further left they are. It is also useful to present the npKi data of Fig. S3 in numerical format. npKi values below about 2.0 should be imperceptible, while values above about 2.0 should be perceptible, and the higher the npKi value, the more perceptible a receptor should be at a particular drug.
5ht2b: 4.00 DOB, 4.00 MDA, 4.00 Aleph-2, 4.00 2C-B-fly, 4.00 2C-B, 4.00 TMA, 4.00 psilocin, 4.00 TMA-2, 4.00 2C-E, 4.00 2C-T-2, 4.00 4C-T-2, 4.00 MEM, 4.00 DOM, 3.97 mescaline, 3.93 6-F-DMT, 3.91 5-MeO-DIPT, 3.91 DMT, 3.88 DPT, 3.70 DOET, 3.64 MDMA, 3.48 DIPT, 3.32 5-MeO-MIPT, 3.13 DOI, 3.11 LSD, 3.01 lisuride, 2.72 cis-2a, 2.17 SS-2c, 1.81 RR-2b, 0.69 5-MeO-DMT; 0.00 salvinorin A; ND: 5-MeO-TMT, ibogaine, EMDT, morphine, THC
5ht1a: 4.00 5-MeO-MIPT, 4.00 lisuride, 4.00 DOET, 4.00 SS-2c, 4.00 5-MeO-DIPT, 4.00 DIPT, 4.00 5-MeO-DMT, 4.00 cis-2a, 4.00 DPT, 3.73 LSD, 3.61 mescaline, 3.59 RR-2b, 3.48 5-MeO-TMT, 3.45 TMA, 2.97 EMDT, 2.91 2C-E, 2.88 psilocin, 2.81 6-F-DMT, 2.75 2C-B, 2.38 MDA, 2.31 DOI, 2.20 2C-T-2, 2.04 4C-T-2, 1.79 2C-B-fly, 1.51 DOM, 1.18 DOB, 0.98 Aleph-2; 0.00: TMA-2, MEM, MDMA, DMT, ibogaine, salvinorin A; ND: morphine, THC
5ht7: 4.00 DMT, 3.80 6-F-DMT, 3.80 TMA, 3.79 5-MeO-MIPT, 3.77 LSD, 3.69 5-MeO-DMT, 3.62 5-MeO-TMT, 3.30 cis-2a, 3.05 DPT, 3.03 5-MeO-DIPT, 2.82 psilocin, 2.81 2C-B, 2.81 RR-2b, 2.77 2C-E, 2.73 EMDT, 2.64 lisuride, 2.55 DIPT, 2.45 SS-2c, 2.41 MDA, 2.18 4C-T-2, 2.07 DOET, 1.95 MEM, 1.90 DOI, 1.89 DOB, 1.87 DOM, 1.79 2C-T-2, 1.17 2C-B-fly, 1.08 Aleph-2; 0.00: TMA-2, MDMA, mescaline, salvinorin A; ND: ibogaine, morphine, THC
5ht1d: 3.97 DMT, 3.81 2C-B-fly, 3.74 5-MeO-MIPT, 3.71 2C-B, 3.70 LSD, 3.66 6-F-DMT, 3.54 2C-E, 3.48 5-MeO-DMT, 3.40 psilocin, 3.38 5-MeO-TMT, 3.20 TMA, 3.05 RR-2b, 3.00 DOI, 2.89 5-MeO-DIPT, 2.84 2C-T-2, 2.75 DOM, 2.74 EMDT, 2.71 DPT, 2.51 DIPT, 1.79 DOB, 1.34 DOET, 1.02 Aleph-2; 0.00: 4C-T-2, mescaline, TMA-2, MEM, lisuride, MDMA, salvinorin A, MDA, ibogaine; ND: cis-2a, SS-2c, morphine, THC
5ht2a: 3.76 2C-E, 3.72 DOET, 3.69 2C-B, 3.54 LSD, 3.44 DOI, 3.42 TMA-2, 3.33 4C-T-2, 3.23 DOB, 3.18 2C-T-2, 2.89 2C-B-fly, 2.89 cis-2a, 2.74 lisuride, 2.58 DMT, 2.47 6-F-DMT, 2.44 5-MeO-MIPT, 2.42 Aleph-2, 2.36 DOM, 2.32 SS-2c, 2.21 MEM, 2.16 ibogaine, 2.14 psilocin, 2.09 DPT, 1.46 RR-2b, 0.98 5-MeO-DMT; 0.00: TMA, DIPT, MDA, MDMA, mescaline, 5-MeO-TMT, EMDT, 5-MeO-DIPT, salvinorin A; ND: morphine, THC
5ht2c: 4.00 DOI, 3.42 DMT, 3.38 2C-E, 3.18 2C-B, 3.13 DOET, 3.11 LSD, 3.09 4C-T-2, 3.05 2C-T-2, 3.02 TMA, 2.97 DOB, 2.93 2C-B-fly, 2.58 TMA-2, 2.58 6-F-DMT, 2.52 psilocin, 2.50 Aleph-2, 2.34 cis-2a, 2.31 DPT, 2.17 5-MeO-TMT, 2.15 MDA, 2.03 SS-2c, 1.95 EMDT, 1.75 5-MeO-MIPT, 1.63 RR-2b, 1.55 5-MeO-DMT, 1.47 DOM; 0.00: 5-MeO-DIPT, mescaline, lisuride, MEM, MDMA, DIPT, ibogaine, salvinorin A; ND: morphine, THC
Alpha2C: 4.00 mescaline, 3.88 lisuride, 3.60 MDA, 3.53 DMT, 3.44 2C-E, 3.24 6-F-DMT, 3.21 MDMA, 3.12 2C-B, 2.98 TMA, 2.88 DOI, 2.79 DPT, 2.68 DIPT, 2.66 5-MeO-DIPT, 2.56 2C-T-2, 2.29 5-MeO-MIPT, 2.10 DOM, 2.00 DOET, 1.82 DOB, 1.59 2C-B-fly, 1.57 5-MeO-DMT, 1.06 Aleph-2, 1.03 psilocin; 0.00: 4C-T-2, MEM, TMA-2; ND: LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, THC
5ht6: 4.00 6-F-DMT, 4.00 EMDT, 4.00 5-MeO-TMT, 3.75 LSD, 3.25 cis-2a, 3.25 DMT, 2.98 5-MeO-MIPT, 2.82 psilocin, 2.73 5-MeO-DMT, 2.63 2C-B, 2.61 lisuride, 2.52 RR-2b, 2.44 SS-2c, 2.34 DOI, 1.93 2C-E, 1.85 DPT, 1.78 2C-B-fly, 1.64 2C-T-2, 1.16 DOM, 0.85 DOB, 0.72 Aleph-2; 0.00: 4C-T-2, 5-MeO-DIPT, mescaline, TMA, TMA-2, MDA, DOET, MEM, MDMA, DIPT, salvinorin A; ND: ibogaine, morphine, THC
5ht1b: 4.00 LSD, 4.00 RR-2b, 3.79 cis-2a, 3.22 SS-2c, 3.22 TMA, 3.11 2C-B, 3.07 6-F-DMT, 3.00 2C-E, 2.61 5-MeO-MIPT, 2.57 DPT, 2.41 5-MeO-DIPT, 2.41 5-MeO-DMT, 2.27 lisuride, 2.19 psilocin, 1.84 2C-T-2, 1.71 DOET, 1.69 2C-B-fly, 1.62 DOB, 0.90 Aleph-2; 0.00: 4C-T-2, MDMA, mescaline, DMT, DIPT, TMA-2, DOM, MDA, DOI, MEM, ibogaine, salvinorin A; ND: 5-MeO-TMT, EMDT, morphine, THC
5ht1e: 3.28 DMT, 3.16 mescaline, 3.15 TMA, 3.05 2C-B, 3.03 psilocin, 2.74 6-F-DMT, 2.66 DOI, 2.62 LSD, 2.60 2C-E, 2.52 5-MeO-TMT, 2.49 EMDT, 2.16 2C-T-2, 2.16 RR-2b, 2.13 DPT, 2.07 cis-2a, 1.91 2C-B-fly, 1.83 lisuride, 1.77 4C-T-2, 1.72 5-MeO-DMT, 1.61 DOET, 1.55 5-MeO-MIPT, 1.52 DOM, 1.44 DOB, 1.16 SS-2c, 0.71 Aleph-2; 0.00: 5-MeO-DIPT, TMA-2, MDA, MEM, MDMA, DIPT, salvinorin A; ND: ibogaine, morphine, THC
Alpha2A: 3.79 DOI, 3.36 TMA, 3.22 lisuride, 2.94 MDMA, 2.93 LSD, 2.92 mescaline, 2.85 5-MeO-MIPT, 2.84 DPT, 2.75 DMT, 2.74 MDA, 2.71 2C-E, 2.64 2C-B, 2.30 DOM, 2.05 DOET, 1.92 2C-T-2, 1.79 2C-B-fly, 1.36 psilocin, 1.00 5-MeO-DMT, 0.96 DOB, 0.44 Aleph-2; 0.00: 4C-T-2, DIPT, 5-MeO-DIPT, MEM, TMA-2, 6-F-DMT; ND: cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, THC
Alpha2B: 3.78 lisuride, 3.53 DMT, 3.13 DOI, 3.12 MDA, 3.09 MDMA, 2.99 6-F-DMT, 2.97 DPT, 2.91 2C-E, 2.62 DIPT, 2.40 DOET, 2.40 5-MeO-DIPT, 2.31 2C-B, 2.13 DOM, 1.86 5-MeO-MIPT, 1.79 2C-T-2, 1.57 psilocin, 1.41 DOB, 1.16 2C-B-fly, 0.86 5-MeO-DMT, 0.74 Aleph-2; 0.00: 4C-T-2, mescaline, TMA, MEM, TMA-2; ND: LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, THC
Imidazoline1: 4.00 MDMA, 3.53 DIPT, 3.44 mescaline, 3.24 5-MeO-DIPT, 3.13 DMT, 2.97 DPT, 2.79 4C-T-2, 2.45 6-F-DMT, 2.15 5-MeO-MIPT, 1.80 2C-B, 1.77 psilocin, 1.70 Aleph-2, 1.46 2C-T-2, 1.39 DOB, 0.88 5-MeO-DMT; 0.00: MEM, 2C-E, MDA, lisuride, DOI, 2C-B-fly, DOM, TMA, DOET; ND: TMA-2, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, THC
5ht5a: 3.64 LSD, 3.20 RR-2b, 3.16 DMT, 3.15 cis-2a, 2.99 lisuride, 2.83 psilocin, 2.56 4C-T-2, 2.43 6-F-DMT, 2.17 SS-2c, 2.11 5-MeO-MIPT, 1.86 DPT, 1.84 5-MeO-DMT, 1.76 5-MeO-TMT, 1.54 EMDT, 0.87 DOB; 0.00: MDMA, 5-MeO-DIPT, 2C-B, MDA, mescaline, 2C-B-fly, DOI, TMA, DOET, TMA-2, 2C-E, 2C-T-2, Aleph-2, MEM, DOM, DIPT, salvinorin A; ND: ibogaine, morphine, THC
D3: 3.46 cis-2a, 3.16 LSD, 2.82 SS-2c, 2.67 psilocin, 2.66 4C-T-2, 2.56 DIPT, 2.47 6-F-DMT, 2.40 RR-2b, 2.37 DPT, 2.27 2C-E, 1.70 5-MeO-MIPT, 1.68 DOB, 1.58 5-MeO-DMT, 1.51 2C-T-2, 1.47 ibogaine, 1.41 Aleph-2, 1.34 lisuride, 1.28 2C-B; 0.00: MDMA, mescaline, MEM, DMT, TMA, DOET, TMA-2, DOM, MDA, DOI, salvinorin A, 5-MeO-TMT, EMDT, 5-MeO-DIPT; ND: 2C-B-fly, morphine, THC
Sigma1: 3.95 TMA, 3.05 4C-T-2, 2.90 DPT, 2.83 DIPT, 2.55 ibogaine, 2.23 DMT, 2.15 5-MeO-DIPT, 2.10 MEM, 1.83 Aleph-2, 1.73 DOI, 1.57 6-F-DMT, 1.25 DOB, 1.19 2C-T-2, 1.17 DOET; 0.00: 2C-B, mescaline, 5-MeO-MIPT, DOM, 2C-B-fly, 5-MeO-DMT, psilocin; ND: 2C-E, lisuride, MDA, TMA-2, LSD, cis-2a, MDMA, SS-2c, 5-MeO-TMT, EMDT, RR-2b, salvinorin A, morphine, THC
SERT: 3.57 ibogaine, 3.31 DPT, 3.25 6-F-DMT, 2.98 DIPT, 2.83 DOI, 2.72 5-MeO-DIPT, 2.37 DMT, 1.74 psilocin, 1.28 5-MeO-MIPT, 1.08 Aleph-2, 0.97 5-MeO-DMT, 0.66 DOB; 0.00: 2C-E, MDA, 2C-B, mescaline, 4C-T-2, 2C-T-2, lisuride, MEM, 2C-B-fly, DOM, TMA, MDMA, TMA-2, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, DOET, salvinorin A; ND: morphine, THC
Beta2: 3.67 4C-T-2, 3.52 DOI, 3.38 DOM, 2.79 Aleph-2, 2.11 DOB, 1.93 lisuride, 1.82 DOET, 1.70 2C-T-2, 1.05 LSD, 0.70 cis-2a, 0.57 SS-2c; 0.00: 5-MeO-DMT, DMT, mescaline, DIPT, DPT, 5-MeO-MIPT, 6-F-DMT, 5-MeO-DIPT, MDMA, 2C-B-fly, psilocin, TMA, TMA-2, MDA, MEM, salvinorin A; ND: 2C-B, 2C-E, RR-2b, 5-MeO-TMT, ibogaine, EMDT, morphine, THC
Sigma2: 4.00 ibogaine, 3.95 TMA, 2.65 DIPT, 2.64 5-MeO-DIPT, 2.13 5-MeO-MIPT, 2.04 DPT, 1.70 DOI, 1.56 6-F-DMT, 1.18 DOET; 0.00: 2C-T-2, 2C-E, mescaline, 2C-B, DOB, Aleph-2, MEM, 4C-T-2, DOM, DMT, 5-MeO-DMT, 2C-B-fly, psilocin; ND: lisuride, MDA, TMA-2, LSD, cis-2a, MDMA, SS-2c, 5-MeO-TMT, EMDT, RR-2b, salvinorin A, morphine, THC
H1: 3.74 6-F-DMT, 3.41 DPT, 3.04 TMA-2, 2.53 DIPT, 2.42 DOI, 1.66 cis-2a, 1.41 5-MeO-MIPT, 1.40 LSD, 1.14 RR-2b, 0.63 DOB, 0.20 SS-2c, 0.19 2C-B-fly; 0.00: psilocin, 2C-B, 5-MeO-DIPT, TMA, 4C-T-2, 2C-E, 2C-T-2, MDMA, mescaline, DOM, EMDT, DOET, salvinorin A, 5-MeO-TMT, MEM, Aleph-2, ibogaine; ND: lisuride, DMT, MDA, 5-MeO-DMT, morphine, THC
D1: 3.51 DMT, 3.37 psilocin, 2.67 6-F-DMT, 2.38 5-MeO-DMT, 2.34 LSD, 2.00 cis-2a, 1.77 RR-2b, 1.67 DOI, 1.14 SS-2c, 0.26 2C-B-fly; 0.00: 2C-B, MDA, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, DOB, lisuride, MDMA, mescaline, DOM, TMA, DOET, TMA-2, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-TMT, EMDT, ibogaine, salvinorin A; ND: morphine, THC
D2: 2.67 cis-2a, 2.65 lisuride, 2.62 6-F-DMT, 2.55 LSD, 1.88 RR-2b, 1.88 2C-E, 1.74 SS-2c, 1.60 2C-B, 1.54 DPT, 1.33 2C-T-2, 0.73 5-MeO-DMT, 0.12 2C-B-fly; 0.00: psilocin, 5-MeO-MIPT, 5-MeO-DIPT, DMT, 4C-T-2, DIPT, MDMA, DOI, mescaline, DOM, TMA, DOET, TMA-2, DOB, MDA, Aleph-2, MEM, 5-MeO-TMT, EMDT, ibogaine, salvinorin A; ND: morphine, THC
D4: 2.90 lisuride, 2.49 cis-2a, 2.39 LSD, 2.25 6-F-DMT, 2.20 DPT, 2.08 RR-2b, 1.62 SS-2c, 1.29 5-MeO-MIPT, 0.78 5-MeO-DMT; 0.00: psilocin, MDMA, DMT, 2C-B, DIPT, 5-MeO-DIPT, mescaline, 4C-T-2, DOB, MDA, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-TMT, EMDT, salvinorin A; ND: ibogaine, morphine, THC
M3: 3.07 MDMA, 2.51 DOI, 2.22 2C-B, 2.22 ibogaine, 1.99 2C-E, 1.94 2C-T-2, 1.53 DOB, 1.42 2C-B-fly, 0.92 Aleph-2; 0.00: DMT, lisuride, 5-MeO-DMT, 5-MeO-MIPT, DIPT, psilocin, DPT, 4C-T-2, 6-F-DMT, mescaline, MEM, salvinorin A, 5-MeO-DIPT; ND: MDA, DOM, TMA, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, DOET, TMA-2, morphine, THC
Alpha1A: 3.17 6-F-DMT, 2.70 DMT, 2.56 DPT, 2.09 lisuride, 1.64 cis-2a, 1.56 DOM, 1.54 LSD, 0.67 SS-2c, 0.59 RR-2b; 0.00: psilocin, MDMA, mescaline, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, 5-MeO-DMT, 4C-T-2, DOB, MDA, DOI, 2C-B-fly, 2C-B, TMA, DOET, TMA-2, 2C-E, 2C-T-2, Aleph-2, MEM, salvinorin A; ND: 5-MeO-TMT, ibogaine, EMDT, morphine, THC
KOR: 4.00 salvinorin A, 2.88 ibogaine, 2.21 morphine; 0.00: MDA, 2C-B, DMT, MDMA, mescaline, DOB, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, psilocin, 5-MeO-DMT, 2C-E, LSD, lisuride, DOI, 2C-B-fly, DOM, TMA, DOET, DPT, 4C-T-2, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, 5-MeO-DIPT; ND: TMA-2, 6-F-DMT, THC
Beta1: 2.89 DOI, 2.56 lisuride, 1.76 cis-2a, 1.72 SS-2c, 1.40 DOET, 1.39 LSD, 1.21 DOB, 0.81 2C-T-2, 0.37 Aleph-2; 0.00: 5-MeO-DMT, DMT, mescaline, DOM, DIPT, 5-MeO-MIPT, DPT, 4C-T-2, 6-F-DMT, 5-MeO-DIPT, MDMA, 2C-B-fly, 2C-E, TMA, psilocin, TMA-2, ibogaine, MDA, MEM, salvinorin A; ND: 2C-B, 5-MeO-TMT, RR-2b, EMDT, morphine, THC
MOR: 4.00 morphine, 2.67 ibogaine; 0.00: lisuride, mescaline, 2C-B, TMA, MDMA, DPT, DOB, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, psilocin, DOET, 2C-E, LSD, cis-2a, DOI, 2C-B-fly, DOM, EMDT, 5-MeO-DIPT, salvinorin A, 4C-T-2, SS-2c, RR-2b, 5-MeO-TMT; ND: MDA, DMT, 5-MeO-DMT, TMA-2, 6-F-DMT, THC
M5: 2.54 MDMA, 2.32 DOI, 2.16 2C-E, 1.60 2C-T-2, 1.01 2C-B-fly, 0.30 Aleph-2; 0.00: 2C-B, DMT, DOB, MDA, DOET, 5-MeO-DMT, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, 6-F-DMT, lisuride, MEM, mescaline, DOM, TMA, TMA-2, salvinorin A; ND: psilocin, LSD, RR-2b, cis-2a, 5-MeO-TMT, EMDT, ibogaine, SS-2c, morphine, THC
Alpha1B: 2.95 DMT, 2.86 DPT, 1.08 lisuride, 0.65 LSD; 0.00: MDMA, 2C-B, psilocin, mescaline, DOB, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, 5-MeO-DMT, 4C-T-2, 6-F-DMT, MDA, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, 2C-E, cis-2a, RR-2b, SS-2c, salvinorin A; ND: 5-MeO-TMT, ibogaine, EMDT, morphine, THC
CB1: 4.00 THC; 0.00: MDA, MDMA, mescaline, 2C-B, DMT, psilocin, 5-MeO-DMT, 2C-E, 2C-T-2, Aleph-2, MEM, 2C-B-fly, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, DOB, lisuride, DOI, TMA, DOM, 6-F-DMT, DOET; ND: 5-MeO-MIPT, LSD, TMA-2, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, cis-2a
M4: 2.90 DOI, 2.43 MDMA, 1.24 2C-B-fly; 0.00: DOB, 2C-B, lisuride, psilocin, DMT, 2C-E, 2C-T-2, Aleph-2, 5-MeO-DMT, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, 6-F-DMT, mescaline, MEM, salvinorin A; ND: DOM, MDA, DOET, TMA, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, TMA-2, morphine, THC
DAT: 3.02 ibogaine, 2.33 DPT; 0.00: DOB, MDA, 2C-B, DMT, MDMA, mescaline, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, psilocin, 5-MeO-DMT, 4C-T-2, 6-F-DMT, lisuride, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, 5-MeO-DIPT, salvinorin A; ND: morphine, THC
D5: 2.05 LSD, 1.61 6-F-DMT, 1.59 lisuride, 1.53 RR-2b, 1.36 cis-2a, 1.00 SS-2c; 0.00: psilocin, 5-MeO-DMT, 2C-B, DMT, MDMA, mescaline, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, DOB, MDA, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-TMT, EMDT, salvinorin A; ND: ibogaine, morphine, THC
CB2: 3.78 THC; 0.00: MDA, DOI, mescaline, 2C-B, DMT, psilocin, 5-MeO-DMT, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, DOB, lisuride, DOET, 2C-B-fly, DOM, TMA, 6-F-DMT; ND: MDMA, LSD, TMA-2, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, cis-2a
Ca+Channel: 3.26 MDMA; 0.00: lisuride, DOB, mescaline, 5-MeO-MIPT, DMT, psilocin, 5-MeO-DMT, 2C-E, 2C-T-2, Aleph-2, MEM, 4C-T-2, DIPT, 5-MeO-DIPT, DPT, 6-F-DMT; ND: 2C-B, MDA, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, ibogaine, salvinorin A, morphine, THC
M1: 2.23 DOI, 1.96 ibogaine, 1.15 2C-B-fly; 0.00: lisuride, DOB, DMT, 2C-B, 5-MeO-DMT, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, psilocin, DPT, 4C-T-2, 6-F-DMT, mescaline, MDMA, salvinorin A, DOM, 5-MeO-DIPT; ND: MDA, TMA, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, DOET, TMA-2, morphine, THC
NMDA: 3.01 ibogaine; 0.00: 2C-T-2, DOB, mescaline, 2C-B, TMA, MDMA, DPT, 2C-E, 6-F-DMT, Aleph-2, MEM, 5-MeO-MIPT, DIPT, DOET, 5-MeO-DIPT, 4C-T-2, DOM, DOI; ND: lisuride, 2C-B-fly, MDA, DMT, 5-MeO-DMT, TMA-2, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-TMT, EMDT, psilocin, salvinorin A, morphine, THC
M2: 2.36 DOI, 1.72 ibogaine, 0.65 2C-B-fly; 0.00: MDA, DOB, DMT, 2C-B, 5-MeO-DMT, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, psilocin, DPT, 4C-T-2, 6-F-DMT, lisuride, MDMA, mescaline, DOM, TMA, DOET, TMA-2, 5-MeO-DIPT, salvinorin A; ND: LSD, cis-2a, 5-MeO-TMT, EMDT, RR-2b, SS-2c, morphine, THC
H2: 2.44 6-F-DMT, 1.42 lisuride; 0.00: MDMA, mescaline, 2C-B, DMT, psilocin, 5-MeO-DMT, 2C-E, 2C-T-2, Aleph-2, MEM, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, DOB, MDA, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, 5-MeO-TMT, EMDT; ND: RR-2b, SS-2c, LSD, cis-2a, ibogaine, salvinorin A, morphine, THC
NET: 1.97 5-MeO-TMT, 0.82 5-MeO-DMT; 0.00: MDMA, MDA, DOB, DMT, psilocin, mescaline, 2C-E, 2C-T-2, Aleph-2, MEM, 2C-B, DIPT, 5-MeO-DIPT, DPT, 4C-T-2, 6-F-DMT, lisuride, DOI, 2C-B-fly, DOM, TMA, DOET, TMA-2, LSD, cis-2a, RR-2b, SS-2c, 5-MeO-MIPT, EMDT; ND: ibogaine, salvinorin A, morphine, THC
DOR: 0.72 morphine; 0.00: 2C-T-2, DOI, mescaline, 2C-B, TMA, psilocin, DPT, DOB, LSD, Aleph-2, MEM, 5-MeO-MIPT, DIPT, 5-MeO-DIPT, DOET, 2C-E, 4C-T-2, cis-2a, RR-2b, 2C-B-fly, DOM, EMDT, ibogaine, salvinorin A, 5-MeO-TMT, SS-2c; ND: MDMA, lisuride, MDA, DMT, 5-MeO-DMT, TMA-2, 6-F-DMT, THC
Activity Data
The NIMH-PDSP provided activity data for the twenty-five drugs of Fig. 1, for 5-HT2A and 5-HT2C (Table S3). While most compounds appear to be full agonists at the two receptors, there are a few exceptions. Lower values of activity were reported for psilocin, MDMA, DOM and the three control drugs: 4C-T-2, 6-fluoro-DMT, and lisuride.
Discussion
Perhaps the most striking result of the NIMH-PDSP assays has been to show that the psychedelics interact with a large number of receptors (forty-two of the forty-nine sites at which most of the drugs were assayed). While the phenylalkylamines tend to be more selective than the tryptamines and ergolines, they generally can not be accurately characterized as selective for 5-HT2, as they are so widely described in the literature [1], [4], [6], [7], [14], [15]. Only DOB and MEM come close to fitting that description. Ironically, DOI has been one of the drugs of choice in studies of the molecular pharmacology of psychedelics, and has been widely assumed to be a 5-HT2-selective agent [4], [6], [7], [15], [16]. This study has revealed DOI to be one of the least selective of all psychedelics. Some of the literature on DOI may need to be reinterpreted. The same may be true of any studies whose conclusions rely on the assumption that psychedelics are selective. Studies requiring drugs selective for 5-HT2 should be conducted with DOB or MEM, and they should not be presented as typical or characteristic of psychedelics.
In addition to showing that psychedelics are not as selective as generally believed, the data presented also shows that they exhibit diverse patterns of receptor interactions. Different drugs emphasize different classes of receptors. 5-HT2B is the best hit for thirteen drugs, and 5-HT1A is the best hit for nine drugs. Five of the top six psychedelic receptors (Table 5) are 5-HT1 and 5-HT2 receptors. If we acknowledge the pervasiveness of the 5-HT1 and 5-HT2 receptors, and then look past them, we find that the set of thirty-five drugs emphasize a wide variety of receptors:
5-HT1A DOET
5-HT1D 2C-B-fly
5-HT2 DOB, 2C-T-2
5-HT2B MEM
5-HT5A RR-2b
5-HT6 EMDT, 5-MeO-TMT, 6-F-DMT
5-HT7 DMT, 5-MeO-MIPT, LSD, 5-MeO-DMT
α2A DOI
α2C mescaline, lisuride, MDA, 2C-E
β2 DOM, 2C-B, Aleph-2, 4C-T-2
H1 TMA-2, DPT
σ2 ibogaine, TMA
D1 psilocin
D3 cis-2a, SS-2c
I1 MDMA, DIPT, 5-MeO-DIPT
κ salvinorin A
μ morphine
CB1 THC
It should be possible to use this diverse set of drugs as probes into the roles played by the various receptor systems in the human mind. In the papers that follow, this possibility will be explored by synthesizing the NIMH-PDSP data together with the data on the human pharmacology of these drugs.
Supporting Information
Receptor affinity profiles of psychedelic drugs, ordered by receptor type. The vertical axis is normalized pKi (npKi). Horizontal axis is a list of forty-two receptors, grouped by receptor type. The drugs are ordered to correspond roughly to similarity of structure and receptor affinity profiles. Colors correspond to classes of receptors. It can be seen at a glance that most, but not all of the drugs interact strongly with the serotonin receptors (beige), certain drugs interact strongly with the dopamine receptors (red), others with the adrenergic receptors (green), yet others with the histamine receptors (yellow), etc.
(0.14 MB DOC)
Receptor affinity profiles of psychedelic drugs, ordered by decreasing affinity. The vertical axis is normalized pKi (npKi). Horizontal axis is a list of forty-two receptors, arranged in order of decreasing affinity for each individual drug. The thirty-five drugs are arranged in order of decreasing breadth, based on the Bsq values of Table 3 and Fig. 4. Drugs at the top of the figure have the broadest receptor interactions (least selective), while drugs at the bottom of the figure have the narrowest receptor interactions (most selective). Colors correspond to classes of receptors, and are the same as used in Fig. S1. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Receptors to the right of the black bar should be imperceptible, while receptors to the left of the black bar should be perceptible, increasingly so the further left they are.
(0.18 MB DOC)
Receptor affinities at forty-two receptors across thirty-five drugs, ordered by decreasing breadth of receptor. The vertical axis is normalized pKi (npKi). Horizontal axis is a list of thirty-five drugs, ordered by decreasing affinity at the receptor. The forty-two receptors are arranged in order of decreasing breadth, based on the Bsq values of Table 5 and Fig. 5. Receptors at the top of the figure have the broadest interactions with the thirty-five drugs, while receptors at the bottom of the figure have the narrowest interactions with the thirty-five drugs. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity at each drug. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Drugs to the right of the black bar should have imperceptible interactions with the receptor, while drugs to the left of the black bar should have perceptible interactions with the receptor, increasingly so the further left they are.
(0.20 MB DOC)
Receptor affinity data for ibogaine. Table S1 reports receptor affinity data for ibogaine collected from the literature. The columns identify the receptor, the species from which the receptor was used, the tissue from which the receptor was used, the radioligand used in determining affinity, the Ki value in nanomoles or the IC50 value in nanomoles, and the literature reference from which the data was obtained.
(0.31 MB DOC)
Raw affinity (Ki) data for thirty-five drugs at sixty-seven sites. The table has been divided into three sections. The first section displays forty-two sites at which most compounds were assayed and at least one “hit” (Ki<10,000 nm) was found (5ht1a, 5ht1b, 5ht1d, 5ht1e, 5ht2a, 5ht2b, 5ht2c, 5ht5a, 5ht6, 5ht7, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, Alpha2A, Alpha2B, Alpha2C, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, Sigma1, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, CB1, CB2, Ca+Channel, NMDA/MK801). The second section displays seven sites at which most compounds were assayed, but at which there were no hits (5ht3, H3, H4, V1, V2, V3, GabaA). The third section displays the remaining eighteen sites, at which only a few compounds were assayed, and no hits were found (GabaB, mGluR1a, mGluR2, mGluR4, mGluR5, mGluR6, mGluR8, A2B2, A2B4, A3B2, A3B4, A4B2, A4B2**, A4B4, BZP (a1), EP3, MDR 1, PCP). Missing Ki values are indicated by “ND” meaning that no data is available, or by “PH” meaning that the primary assay “hit” (>50% inhibition), but the secondary assay was not performed. For the first section of the table, an extra row and column labeled “ND/PH” provides a count of missing Ki data in the two categories, for each compound and for each receptor.
(0.05 MB XLS)
Activity data for twenty-five drugs at 5-HT2A and 5-HT2C. GF62 is the cell line that expresses the 5-HT2A receptor, and INI is the cell line that expresses the 5-HT2C receptor. The “EC50 nM” columns express the concentration that gives half of the maximal activity for that drug. The maximal activity is displayed in the “Emax±SEM” column, and represent Ca++ mobilization relative to 5-HT which should give an Emax value of 100%. Data for the drugs should produce lower Emax values. For a compound that gives, for example, 53% Emax, the EC50 is the concentration where 26.5% response occurs. Emax values above 100%±SEM are an artifact caused by extrapolation by the graphpad program when it doesn't have points at the top end to define the asymptote. The data represent the mean ± variance of computer-derived estimates from single experiments done in quadruplicate. Thus, the four observations are averaged and a single estimate with error is provided.
(0.13 MB DOC)
Affinity (Ki) data transformed into pKi values for thirty-five drugs at forty-two sites. Table S4 presents the raw Ki data transformed into pKi values. Higher affinities produce lower Ki values, thus it is valuable to calculate: pKi = −log10(Ki). Higher affinities have higher pKi values, and each unit of pKi value corresponds to one order of magnitude of Ki value. ND means the data is not available, and UM means that Ki was measured as >10,000 nm. Generally, the highest Ki value generated by NIMH-PDSP is 10,000, which produces a pKi value of −4 (although a value of 10,450 was reported for 5-MeO-TMT). For non-PDSP data gathered from the literature, some values greater than 10,000 are reported (i.e. 12,500, 14,142, 22,486, 39,409 and 70,000 for ibogaine).
(0.04 MB XLS)
Thirty-five drugs arranged in order of decreasing breadth at selected groups of receptors. The thirty-five drugs are arranged in order of decreasing breadth at groups of receptors, based on the breadth index Bsq. The drugs with the broadest receptor interactions within the group are found at the tops of the columns, and the drugs with the least receptor interactions are found at the bottoms of the columns. Some columns list the maximum npKi value for a group of receptors (e.g. 5-HT2max). In this example, the column lists the highest npKi value of the three 5-HT2 receptors (5-HT2A, 5-HT2B, and 5-HT2C). An entry of “ND” indicates that the statistic could not be calculated because some data is missing. For example, to calculate Bsq for 5-HT2, or npKi for 5-HT2max, we need npKi values for 5-HT2A, 5-HT2B, and 5-HT2C. If any one of the three values is missing, the statistics will be reported as ND. Values of 0.00 correspond to Ki values of >10,000. The same data is also presented in Table S6 Receptors in a group are listed in the column heading, or are represented with the following abbreviations: • 5-HT - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5-HT2 - 5-HT2A, 5-HT2B, 5-HT2C • 5-HT2max - maximum of 5-HT2A, 5-HT2B, 5-HT2C • 5-HT2A/Cmax - maximum of 5-HT2A, 5-HT2C • 5-HT1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5-HT1max - maximum of 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • Dmax - maximum of D1, D2, D3, D4, D5 • Adrenergic - α1A, α1B, α2A, α2B, α2C, β1, β2 • AdrenergicMax - maximum of α1A, α1B, α2A, α2B, α2C, β1, β2 • α1max - maximum of α1A, α1B • α2max - maximum of α2A, α2B, α2C • βmax - maximum of β1, β2 • Hmax - maximum of H1, H2 • σmax - maximum of σ1, σ 2 • Mmax - maximum of M1, M2, M3, M4, M5 • TransportersMax - maximum of SERT, DAT, NET • OpioidMax - maximum of DOR, KOR, MOR
(0.61 MB DOC)
Table of npKi, Bsq, or npKimax values at individual or groups of receptors for thirty-five drugs. Table S6 presents the normalized pKi data (npKi) for thirty five drugs at forty-two receptors, transporters and ion channels. In addition, it presents the three breadth statistics (B, Bsq, Bexp) for each drug across all forty-two sites, the breadth statistic Bsq for sixteen groups of related sites, and the maximum npKi value for thirteen groups of related sites. A useful way to work with the table is to choose a column, and sort the data on that column (click “Data, “Sort,” select “Header row,” choose the column from the “Sort by” drop down and select “Descending,” finally clicking “OK”). Missing values are reported as “ND.” Values of 0.0000 correspond to Ki values of >10,000. Receptors in a group are listed in the column headings using the following abbreviations: • B - sum of npKi values across forty-two receptors • Bsq - square root of sum of squares of npKi values across forty-two receptors • Bexp - log of sum of exponents of npKi values across forty-two receptors • 5ht - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5ht1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5ht1max - maximum of 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5ht2 - 5-HT2A, 5-HT2B, 5-HT2C • 5ht2max - maximum of 5-HT2A, 5-HT2B, 5-HT2C • 5ht2[ac] - 5-HT2A, 5-HT2C • 5ht2[ac]max - maximum of 5-HT2A, 5-HT2C • 5ht[67] - 5-HT6, 5-HT7 • D[1–5] - D1, D2, D3, D4, D5 • D[1–5]max - maximum of D1, D2, D3, D4, D5 • ∧[AB] - α1A, α1B, α2A, α2B, α2C, β1, β2 • ∧[AB]max - maximum of α1A, α1B, α2A, α2B, α2C, β1, β2 • Alpha1 - α1A, α1B • Alpha1max - maximum of α1A, α1B • Alpha2 - α2A, α2B, α2C • Alpha2max - maximum of α2A, α2B, α2C • Beta - β1, β2 • BetaMax - maximum of β1, β2 • M[1–5] - M1, M2, M3, M4, M5 • M[1–5]max - maximum of M1, M2, M3, M4, M5 • Sigma - σ1, σ 2 • SigmaMax - maximum of σ1, σ 2 • H[12] - H1, H2 • H[1–2]max - maximum of H1, H2 • T$ - SERT, DAT, NET • T$max - maximum of SERT, DAT, NET • [DKM]OR - DOR, KOR, MOR • [DKM]ORmax - maximum of DOR, KOR, MOR • CB[12] - CB1, CB2
(0.05 MB XLS)
Thirty-five drugs arranged in order of decreasing proportional interaction at selected groups of receptors. The thirty-five drugs are arranged in order of decreasing proportional interaction at groups of receptors, based on the proportional breadth index Bp. The drugs with the greatest proportional interactions are found at the tops of the columns, and the drugs with the least proportional interactions are found at the bottoms of the columns. An entry of “ND” indicates that the statistic could not be calculated because some data is missing. The same data is also presented in Table S8. Receptors in a group are listed in the column heading, or are represented with the following abbreviations: • 5-HT - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5-HT1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • Adrenergic - α1A, α1B, α2A, α2B, α2C, β1, β2
(0.37 MB DOC)
Table of Bp values at individual or groups of receptors for thirty-five drugs. Table S8 presents the proportional breadth statistic Bp for each drug at each of forty-two sites, and for sixteen groups of related sites. A useful way to work with the table is to choose a column, and sort the data on that column (click “Data, “Sort,” select “Header row,” choose the column from the “Sort by” drop down and select “Descending,” finally clicking “OK”). Missing values are reported as “ND.” Receptors in a group are listed in the column headings using the following abbreviations: • Bsq - square root of sum of squares of npKi values across forty-two receptors • 5ht - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5ht1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5ht2 - 5-HT2A, 5-HT2B, 5-HT2C • 5ht2[ac] - 5-HT2A, 5-HT2C • 5ht[67] - 5-HT6, 5-HT7 • D[1–5] - D1, D2, D3, D4, D5 • ∧[AB] - α1A, α1B, α2A, α2B, α2C, β1, β2 • Alpha1 - α1A, α1B • Alpha2 - α2A, α2B, α2C • Beta - β1, β2 • M[1–5] - M1, M2, M3, M4, M5 • Sigma - σ1, σ 2 • H[12] - H1, H2 • T$ - SERT, DAT, NET • [DKM]OR - DOR, KOR, MOR • CB[12] - CB1, CB2
(0.04 MB XLS)
Acknowledgments
I acknowledge Chenmei Xu for preparing the figures for this manuscript.
Footnotes
Competing Interests: The NIMH-PDSP actually produced the affinity data for twenty-five drugs, and provided it exclusively to the author. This does not alter the author's adherence to all the PLoS ONE policies on sharing data and materials.
Funding: This work resulted from a large set of receptor affinity assays performed by the NIMH-PDSP (http://pdsp.med.unc.edu/). Although the project was specifically approved for the author by the NIMH and the NIMH-PDSP, it did not result in a grant in the sense of funds that flow through the author's institution. The funding went directly to the NIMH-PDSP which was at Case Western Reserve University at the time. Also, the National Institute on Drug Abuse Drug Supply Program (http://www.nida.nih.gov/) provided many of the drugs, but they also went directly to the NIMH-PDSP at Case Western. There was no other funding for this research. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. This statement is true except that the NIMH-PDSP actually produced the affinity data for twenty-five drugs, and provided it exclusively to the author.
References
- 1.Nichols DE. Hallucinogens. Pharmacology & Therapeutics. 2004;101:131–81. doi: 10.1016/j.pharmthera.2003.11.002. [DOI] [PubMed] [Google Scholar]
- 2.Glennon RA, Young R, Rosecrans JA. Antagonism of the effects of the hallucinogen DOM and the purported 5-HT agonist quipazine by 5-HT2 antagonists. Eur J Pharmacol. 1983;91:189–96. doi: 10.1016/0014-2999(83)90464-8. [DOI] [PubMed] [Google Scholar]
- 3.Glennon RA, Titeler M, McKenney JD. Evidence for 5-HT2 involvement in the mechanism of action of hallucinogenic agents. Life Sciences. 1984;35:2505–11. doi: 10.1016/0024-3205(84)90436-3. [DOI] [PubMed] [Google Scholar]
- 4.Glennon RA, Titeler M, Young R. Structure-activity relationships and mechanism of action of hallucinogenic agents based on drug discrimination and radioligand binding studies. Psychopharmacol Bull. 1986;22:953–8. [PubMed] [Google Scholar]
- 5.Glennon RA, Darmani NA, Martin BR. Multiple populations of serotonin receptors may modulate the behavioral effects of serotonergic agents. Life Sci. 1991;48:2493–8. doi: 10.1016/0024-3205(91)90603-9. [DOI] [PubMed] [Google Scholar]
- 6.Glennon RA. Do classical hallucinogens act as 5-HT2 agonists or antagonists? Neuropsychopharmacology. 1990;3:509–17. [PubMed] [Google Scholar]
- 7.Glennon R. Classic Hallucinogens. Handbook of Experimental Pharmacology. 1996;118:343–71. [Google Scholar]
- 8.Blair JB, Kurrasch-Orbaugh D, Marona-Lewicka D, Cumbay MG, Watts VJ, et al. Effect of ring fluorination on the pharmacology of hallucinogenic tryptamines. J Med Chem. 2000;43:4701–10. doi: 10.1021/jm000339w. [DOI] [PubMed] [Google Scholar]
- 9.Nichols DE. Differences between the mechanism of action of MDMA, MBDB, and the classic hallucinogens. Identification of a new therapeutic class: entactogens. J Psychoactive Drugs. 1986;18:305–13. doi: 10.1080/02791072.1986.10472362. [DOI] [PubMed] [Google Scholar]
- 10.Nichols DE, Frescas S, Marona-Lewicka D, Kurrasch-Orbaugh DM. Lysergamides of isomeric 2,4-dimethylazetidines map the binding orientation of the diethylamide moiety in the potent hallucinogenic agent N,N-diethyllysergamide (LSD). Journal of Medicinal Chemistry. 2002;45:4344–9. doi: 10.1021/jm020153s. [DOI] [PubMed] [Google Scholar]
- 11.Roth BL, Baner K, Westkaemper R, Siebert D, Rice KC, et al. Salvinorin A: a potent naturally occurring nonnitrogenous kappa opioid selective agonist. Proc Natl Acad Sci U S A. 2002;99:11934–9. doi: 10.1073/pnas.182234399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Glennon RA, Lee M, Rangisetty JB, Dukat M, Roth BL, et al. 2-Substituted tryptamines: agents with selectivity for 5-HT(6) serotonin receptors. Journal of Medicinal Chemistry. 2000;43:1011–8. doi: 10.1021/jm990550b. [DOI] [PubMed] [Google Scholar]
- 13.Nichols DE, Nichols CD. Serotonin receptors. Chem Rev. 2008;108:1614–41. doi: 10.1021/cr078224o. [DOI] [PubMed] [Google Scholar]
- 14.Darmani NA. Further evidence for enigmas in adaptation mechanisms for the DOI-induced behaviors. Pharmacol Biochem Behav. 1992;43:765–70. doi: 10.1016/0091-3057(92)90406-6. [DOI] [PubMed] [Google Scholar]
- 15.Darmani NA, Mock OB, Towns LC, Gerdes CF. The head-twitch response in the least shrew (Cryptotis parva) is a 5-HT2- and not a 5-HT1C-mediated phenomenon. Pharmacol Biochem Behav. 1994;48:383–96. doi: 10.1016/0091-3057(94)90542-8. [DOI] [PubMed] [Google Scholar]
- 16.Glennon RA, Dukat M. Serotonin receptors and their ligands: a lack of selective agents. Pharmacol Biochem Behav. 1991;40:1009–17. doi: 10.1016/0091-3057(91)90121-h. [DOI] [PubMed] [Google Scholar]
- 17.Schiller PW, Nguyen TM, Berezowska I, Dupuis S, Weltrowska G, et al. Synthesis and in vitro opioid activity profiles of DALDA analogues. Eur J Med Chem. 2000;35:895–901. doi: 10.1016/s0223-5234(00)01171-5. [DOI] [PubMed] [Google Scholar]
- 18.Valenzano KJ, Miller W, Chen Z, Shan S, Crumley G, et al. DiPOA ([8-(3,3-diphenyl-propyl)-4-oxo-1-phenyl-1,3,8-triazaspiro[4.5]dec-3-yl]-a cetic acid), a novel, systemically available, and peripherally restricted mu opioid agonist with antihyperalgesic activity: I. In vitro pharmacological characterization and pharmacokinetic properties. J Pharmacol Exp Ther. 2004;310:783–92. doi: 10.1124/jpet.103.063313. [DOI] [PubMed] [Google Scholar]
- 19.Zhao GM, Qian X, Schiller PW, Szeto HH. Comparison of [Dmt1]DALDA and DAMGO in binding and G protein activation at mu, delta, and kappa opioid receptors. J Pharmacol Exp Ther. 2003;307:947–54. doi: 10.1124/jpet.103.054775. [DOI] [PubMed] [Google Scholar]
- 20.Spetea M, Toth F, Schutz J, Otvos F, Toth G, et al. Binding characteristics of [3H]14-methoxymetopon, a high affinity mu-opioid receptor agonist. Eur J Neurosci. 2003;18:290–5. doi: 10.1046/j.1460-9568.2003.02744.x. [DOI] [PubMed] [Google Scholar]
- 21.Gharagozlou P, Demirci H, David CJ, Lameh J. Activity of opioid ligands in cells expressing cloned mu opioid receptors. BMC Pharmacol. 2003;3:1. doi: 10.1186/1471-2210-3-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Standifer KM, Cheng J, Brooks AI, Honrado CP, Su W, et al. Biochemical and pharmacological characterization of mu, delta and kappa 3 opioid receptors expressed in BE(2)-C neuroblastoma cells. J Pharmacol Exp Ther. 1994;270:1246–55. [PubMed] [Google Scholar]
- 23.Castanas E, Bourhim N, Giraud P, Boudouresque F, Cantau P, et al. Interaction of opiates with opioid binding sites in the bovine adrenal medulla: I. Interaction with delta and mu sites. J Neurochem. 1985;45:677–87. doi: 10.1111/j.1471-4159.1985.tb04046.x. [DOI] [PubMed] [Google Scholar]
- 24.Clark JA, Houghten R, Pasternak GW. Opiate binding in calf thalamic membranes: a selective mu 1 binding assay. Mol Pharmacol. 1988;34:308–17. [PubMed] [Google Scholar]
- 25.Wolozin BL, Pasternak GW. Classification of multiple morphine and enkephalin binding sites in the central nervous system. Proc Natl Acad Sci U S A. 1981;78:6181–5. doi: 10.1073/pnas.78.10.6181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Neubig RR, Spedding M, Kenakin T, Christopoulos A. International Union of Pharmacology Committee on Receptor Nomenclature and Drug Classification. XXXVIII. Update on terms and symbols in quantitative pharmacology. Pharmacol Rev. 2003;55:597–606. doi: 10.1124/pr.55.4.4. [DOI] [PubMed] [Google Scholar]
- 27.De VJ, Denzer D, Reissmueller E, Eijckenboom M, Heil M, et al. 3-[2-cyano-3-(trifluoromethyl)phenoxy]phenyl-4,4,4-trifluoro-1-butanesulfo nate (BAY 59-3074): a novel cannabinoid Cb1/Cb2 receptor partial agonist with antihyperalgesic and antiallodynic effects. J Pharmacol Exp Ther. 2004;310:620–32. doi: 10.1124/jpet.103.062836. [DOI] [PubMed] [Google Scholar]
- 28.Mauler F, Mittendorf J, Horvath E, De VJ. Characterization of the diarylether sulfonylester (−)-(R)-3-(2-hydroxymethylindanyl-4-oxy)phenyl-4,4,4-trifluoro-1-sulfonate (BAY 38-7271) as a potent cannabinoid receptor agonist with neuroprotective properties. J Pharmacol Exp Ther. 2002;302:359–68. doi: 10.1124/jpet.302.1.359. [DOI] [PubMed] [Google Scholar]
- 29.Iwamura H, Suzuki H, Ueda Y, Kaya T, Inaba T. In vitro and in vivo pharmacological characterization of JTE-907, a novel selective ligand for cannabinoid CB2 receptor. J Pharmacol Exp Ther. 2001;296:420–5. [PubMed] [Google Scholar]
- 30.Griffin G, Tao Q, Abood ME. Cloning and pharmacological characterization of the rat CB(2) cannabinoid receptor. J Pharmacol Exp Ther. 2000;292:886–94. [PubMed] [Google Scholar]
- 31.Largent BL, Gundlach AL, Snyder SH. Psychotomimetic opiate receptors labeled and visualized with (+)-[3H]3-(3-hydroxyphenyl)-N-(1-propyl)piperidine. Proc Natl Acad Sci U S A. 1984;81:4983–7. doi: 10.1073/pnas.81.15.4983. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Receptor affinity profiles of psychedelic drugs, ordered by receptor type. The vertical axis is normalized pKi (npKi). Horizontal axis is a list of forty-two receptors, grouped by receptor type. The drugs are ordered to correspond roughly to similarity of structure and receptor affinity profiles. Colors correspond to classes of receptors. It can be seen at a glance that most, but not all of the drugs interact strongly with the serotonin receptors (beige), certain drugs interact strongly with the dopamine receptors (red), others with the adrenergic receptors (green), yet others with the histamine receptors (yellow), etc.
(0.14 MB DOC)
Receptor affinity profiles of psychedelic drugs, ordered by decreasing affinity. The vertical axis is normalized pKi (npKi). Horizontal axis is a list of forty-two receptors, arranged in order of decreasing affinity for each individual drug. The thirty-five drugs are arranged in order of decreasing breadth, based on the Bsq values of Table 3 and Fig. 4. Drugs at the top of the figure have the broadest receptor interactions (least selective), while drugs at the bottom of the figure have the narrowest receptor interactions (most selective). Colors correspond to classes of receptors, and are the same as used in Fig. S1. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Receptors to the right of the black bar should be imperceptible, while receptors to the left of the black bar should be perceptible, increasingly so the further left they are.
(0.18 MB DOC)
Receptor affinities at forty-two receptors across thirty-five drugs, ordered by decreasing breadth of receptor. The vertical axis is normalized pKi (npKi). Horizontal axis is a list of thirty-five drugs, ordered by decreasing affinity at the receptor. The forty-two receptors are arranged in order of decreasing breadth, based on the Bsq values of Table 5 and Fig. 5. Receptors at the top of the figure have the broadest interactions with the thirty-five drugs, while receptors at the bottom of the figure have the narrowest interactions with the thirty-five drugs. The black vertical bars represent a 100-fold drop in affinity relative to the receptor with the highest affinity at each drug. As a rule of thumb, this is presumed to be the limit of perceptible receptor interaction. Drugs to the right of the black bar should have imperceptible interactions with the receptor, while drugs to the left of the black bar should have perceptible interactions with the receptor, increasingly so the further left they are.
(0.20 MB DOC)
Receptor affinity data for ibogaine. Table S1 reports receptor affinity data for ibogaine collected from the literature. The columns identify the receptor, the species from which the receptor was used, the tissue from which the receptor was used, the radioligand used in determining affinity, the Ki value in nanomoles or the IC50 value in nanomoles, and the literature reference from which the data was obtained.
(0.31 MB DOC)
Raw affinity (Ki) data for thirty-five drugs at sixty-seven sites. The table has been divided into three sections. The first section displays forty-two sites at which most compounds were assayed and at least one “hit” (Ki<10,000 nm) was found (5ht1a, 5ht1b, 5ht1d, 5ht1e, 5ht2a, 5ht2b, 5ht2c, 5ht5a, 5ht6, 5ht7, D1, D2, D3, D4, D5, Alpha1A, Alpha1B, Alpha2A, Alpha2B, Alpha2C, Beta1, Beta2, SERT, DAT, NET, Imidazoline1, Sigma1, Sigma2, DOR, KOR, MOR, M1, M2, M3, M4, M5, H1, H2, CB1, CB2, Ca+Channel, NMDA/MK801). The second section displays seven sites at which most compounds were assayed, but at which there were no hits (5ht3, H3, H4, V1, V2, V3, GabaA). The third section displays the remaining eighteen sites, at which only a few compounds were assayed, and no hits were found (GabaB, mGluR1a, mGluR2, mGluR4, mGluR5, mGluR6, mGluR8, A2B2, A2B4, A3B2, A3B4, A4B2, A4B2**, A4B4, BZP (a1), EP3, MDR 1, PCP). Missing Ki values are indicated by “ND” meaning that no data is available, or by “PH” meaning that the primary assay “hit” (>50% inhibition), but the secondary assay was not performed. For the first section of the table, an extra row and column labeled “ND/PH” provides a count of missing Ki data in the two categories, for each compound and for each receptor.
(0.05 MB XLS)
Activity data for twenty-five drugs at 5-HT2A and 5-HT2C. GF62 is the cell line that expresses the 5-HT2A receptor, and INI is the cell line that expresses the 5-HT2C receptor. The “EC50 nM” columns express the concentration that gives half of the maximal activity for that drug. The maximal activity is displayed in the “Emax±SEM” column, and represent Ca++ mobilization relative to 5-HT which should give an Emax value of 100%. Data for the drugs should produce lower Emax values. For a compound that gives, for example, 53% Emax, the EC50 is the concentration where 26.5% response occurs. Emax values above 100%±SEM are an artifact caused by extrapolation by the graphpad program when it doesn't have points at the top end to define the asymptote. The data represent the mean ± variance of computer-derived estimates from single experiments done in quadruplicate. Thus, the four observations are averaged and a single estimate with error is provided.
(0.13 MB DOC)
Affinity (Ki) data transformed into pKi values for thirty-five drugs at forty-two sites. Table S4 presents the raw Ki data transformed into pKi values. Higher affinities produce lower Ki values, thus it is valuable to calculate: pKi = −log10(Ki). Higher affinities have higher pKi values, and each unit of pKi value corresponds to one order of magnitude of Ki value. ND means the data is not available, and UM means that Ki was measured as >10,000 nm. Generally, the highest Ki value generated by NIMH-PDSP is 10,000, which produces a pKi value of −4 (although a value of 10,450 was reported for 5-MeO-TMT). For non-PDSP data gathered from the literature, some values greater than 10,000 are reported (i.e. 12,500, 14,142, 22,486, 39,409 and 70,000 for ibogaine).
(0.04 MB XLS)
Thirty-five drugs arranged in order of decreasing breadth at selected groups of receptors. The thirty-five drugs are arranged in order of decreasing breadth at groups of receptors, based on the breadth index Bsq. The drugs with the broadest receptor interactions within the group are found at the tops of the columns, and the drugs with the least receptor interactions are found at the bottoms of the columns. Some columns list the maximum npKi value for a group of receptors (e.g. 5-HT2max). In this example, the column lists the highest npKi value of the three 5-HT2 receptors (5-HT2A, 5-HT2B, and 5-HT2C). An entry of “ND” indicates that the statistic could not be calculated because some data is missing. For example, to calculate Bsq for 5-HT2, or npKi for 5-HT2max, we need npKi values for 5-HT2A, 5-HT2B, and 5-HT2C. If any one of the three values is missing, the statistics will be reported as ND. Values of 0.00 correspond to Ki values of >10,000. The same data is also presented in Table S6 Receptors in a group are listed in the column heading, or are represented with the following abbreviations: • 5-HT - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5-HT2 - 5-HT2A, 5-HT2B, 5-HT2C • 5-HT2max - maximum of 5-HT2A, 5-HT2B, 5-HT2C • 5-HT2A/Cmax - maximum of 5-HT2A, 5-HT2C • 5-HT1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5-HT1max - maximum of 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • Dmax - maximum of D1, D2, D3, D4, D5 • Adrenergic - α1A, α1B, α2A, α2B, α2C, β1, β2 • AdrenergicMax - maximum of α1A, α1B, α2A, α2B, α2C, β1, β2 • α1max - maximum of α1A, α1B • α2max - maximum of α2A, α2B, α2C • βmax - maximum of β1, β2 • Hmax - maximum of H1, H2 • σmax - maximum of σ1, σ 2 • Mmax - maximum of M1, M2, M3, M4, M5 • TransportersMax - maximum of SERT, DAT, NET • OpioidMax - maximum of DOR, KOR, MOR
(0.61 MB DOC)
Table of npKi, Bsq, or npKimax values at individual or groups of receptors for thirty-five drugs. Table S6 presents the normalized pKi data (npKi) for thirty five drugs at forty-two receptors, transporters and ion channels. In addition, it presents the three breadth statistics (B, Bsq, Bexp) for each drug across all forty-two sites, the breadth statistic Bsq for sixteen groups of related sites, and the maximum npKi value for thirteen groups of related sites. A useful way to work with the table is to choose a column, and sort the data on that column (click “Data, “Sort,” select “Header row,” choose the column from the “Sort by” drop down and select “Descending,” finally clicking “OK”). Missing values are reported as “ND.” Values of 0.0000 correspond to Ki values of >10,000. Receptors in a group are listed in the column headings using the following abbreviations: • B - sum of npKi values across forty-two receptors • Bsq - square root of sum of squares of npKi values across forty-two receptors • Bexp - log of sum of exponents of npKi values across forty-two receptors • 5ht - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5ht1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5ht1max - maximum of 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5ht2 - 5-HT2A, 5-HT2B, 5-HT2C • 5ht2max - maximum of 5-HT2A, 5-HT2B, 5-HT2C • 5ht2[ac] - 5-HT2A, 5-HT2C • 5ht2[ac]max - maximum of 5-HT2A, 5-HT2C • 5ht[67] - 5-HT6, 5-HT7 • D[1–5] - D1, D2, D3, D4, D5 • D[1–5]max - maximum of D1, D2, D3, D4, D5 • ∧[AB] - α1A, α1B, α2A, α2B, α2C, β1, β2 • ∧[AB]max - maximum of α1A, α1B, α2A, α2B, α2C, β1, β2 • Alpha1 - α1A, α1B • Alpha1max - maximum of α1A, α1B • Alpha2 - α2A, α2B, α2C • Alpha2max - maximum of α2A, α2B, α2C • Beta - β1, β2 • BetaMax - maximum of β1, β2 • M[1–5] - M1, M2, M3, M4, M5 • M[1–5]max - maximum of M1, M2, M3, M4, M5 • Sigma - σ1, σ 2 • SigmaMax - maximum of σ1, σ 2 • H[12] - H1, H2 • H[1–2]max - maximum of H1, H2 • T$ - SERT, DAT, NET • T$max - maximum of SERT, DAT, NET • [DKM]OR - DOR, KOR, MOR • [DKM]ORmax - maximum of DOR, KOR, MOR • CB[12] - CB1, CB2
(0.05 MB XLS)
Thirty-five drugs arranged in order of decreasing proportional interaction at selected groups of receptors. The thirty-five drugs are arranged in order of decreasing proportional interaction at groups of receptors, based on the proportional breadth index Bp. The drugs with the greatest proportional interactions are found at the tops of the columns, and the drugs with the least proportional interactions are found at the bottoms of the columns. An entry of “ND” indicates that the statistic could not be calculated because some data is missing. The same data is also presented in Table S8. Receptors in a group are listed in the column heading, or are represented with the following abbreviations: • 5-HT - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5-HT1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • Adrenergic - α1A, α1B, α2A, α2B, α2C, β1, β2
(0.37 MB DOC)
Table of Bp values at individual or groups of receptors for thirty-five drugs. Table S8 presents the proportional breadth statistic Bp for each drug at each of forty-two sites, and for sixteen groups of related sites. A useful way to work with the table is to choose a column, and sort the data on that column (click “Data, “Sort,” select “Header row,” choose the column from the “Sort by” drop down and select “Descending,” finally clicking “OK”). Missing values are reported as “ND.” Receptors in a group are listed in the column headings using the following abbreviations: • Bsq - square root of sum of squares of npKi values across forty-two receptors • 5ht - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E, 5-HT2A, 5-HT2B, 5-HT2C, 5-HT5A, 5-HT6, 5-HT7 • 5ht1 - 5-HT1A, 5-HT1B, 5-HT1D, 5-HT1E • 5ht2 - 5-HT2A, 5-HT2B, 5-HT2C • 5ht2[ac] - 5-HT2A, 5-HT2C • 5ht[67] - 5-HT6, 5-HT7 • D[1–5] - D1, D2, D3, D4, D5 • ∧[AB] - α1A, α1B, α2A, α2B, α2C, β1, β2 • Alpha1 - α1A, α1B • Alpha2 - α2A, α2B, α2C • Beta - β1, β2 • M[1–5] - M1, M2, M3, M4, M5 • Sigma - σ1, σ 2 • H[12] - H1, H2 • T$ - SERT, DAT, NET • [DKM]OR - DOR, KOR, MOR • CB[12] - CB1, CB2
(0.04 MB XLS)