Abstract
The carcinogenic polycyclic aromatic hydrocarbon ns (PAHs) benzo[a]pyrene (B[a]P) and dibenzo[a,l]pyrene (DB[a,l]P) are widespread environmental pollutants, however their toxicological effects within a mixture is not established. We investigated the influence of diesel exhaust (DE) on B[a]P and DB[a,l]P-induced PAH-DNA adduct formation, metabolic activation, gene expression and 8-oxo-dG adduct levels in human breast epithelial cells (MCF-10A) in culture. Following 24 and 48 h, cells co-exposed to DE plus B[a]P exhibited a significant decrease in PAH-DNA adduct levels, compared with B[a]P alone, as determined by 33P-postlabeling combined with reversed-phase high performance liquid chromatography (HPLC). Cytochrome P450 (CYP) enzyme activity, as measured by the ethoxyresorufin O-deethylase (EROD) assay and CYP1B1 expression, significantly increased with co-exposure of DE plus DB[a,l]P, compared with DB[a,l]P alone. Aldo keto-reductase (AKR)1C1, AKR1C2,and AKR1C3 expression also significantly increased in cells exposed to DE plus PAH, compared with PAH exposure alone. Cell populations exhibiting 8-oxo-dG adducts significantly increased in response to exposure to B[a]P or DE plus B[a]P for 24 h, compared with vehicle control, as quantified by flow cytometry. These results suggest that complex mixtures may modify the carcinogenic potency of PAH by shifting the metabolic activation pathway from the production of PAH diol-epoxides to AKR pathway-derived metabolites.
INTRODUCTION
Diesel engine exhaust poses as an environmental risk factor to humans, as epidemiological studies report increased morbidity and mortality due to air pollution [1,2], increased risk of lung cancer [3], and genotoxicity in various tissues [4–6]. Polycyclic aromatic hydrocarbons (PAHs) are large, planar, aromatic compounds that are the products of the incomplete combustion of fossil fuels and make up a substantial component of ambient air pollution. In addition to heterocyclics, quinones and aldehydes, PAHs are found to be adsorbed to diesel exhaust particulate matter [7].
Benzo[a]pyrene (B[a]P) and dibenzo[a,l]pyrene (DB[a,l]P) are well-studied PAH found in many environmental complex mixtures, including tobacco smoke, automobile exhaust, and air pollution [8]. B[a]P and DB[a,l]P are highly mutagenic and must be metabolically activated in order to exert their carcinogenic potential. Metabolic activation of PAH involve three distinct mechanistic pathways (as reviewed by Bolton et al. [9]): (i) the diol epoxide path, (ii) the quinone path, and (iii) the radical cation path. In the diol epoxide path, PAH are oxidized by cytochrome P450 (CYP) enzymes to a trans-dihydrodiol which is then further oxidized to highly mutagenic, electrophilic diol epoxides. Covalent DNA adduct formation by PAH-diol epoxides induces mutations in tumor suppressor genes and proto-oncogenes (e.g. [10,11]), is correlated with tumor formation [12], and linked to carcinogenic risk [13]. The quinone path involves dehydrogenation of the dihydrodiol by aldo-ketoreductase (AKR) enzymes to form a catechol that enters into a redox cycle with O2 to form a PAH o-quinone and reactive oxygen species. Nanomolar concentrations of B[a]P-7,8-dione (BPQ) has been shown to be potent mutagen; generating G-to-T transversions in the tumor suppressor gene p53 [14]. The one-electron oxidation pathway entails oxidation of PAH by peroxidase enzymes to form radical cations which react with DNA and other macromolecules, and evidence suggests that one-electron oxidation PAH metabolites produce tumors in rodent models [15]. Therefore, the mutagenic metabolites generated from all three pathways may contribute to the various stages of PAH-induced carcinogenesis.
Carcinogenic PAH metabolites have been detected in both normal and tumor human breast tissue [16,17], and have been implicated in breast tumorigenesis [18–21]. Although the mutagenicity implicated in B[a]P and DB[a,l]P associated tumorigenesis is well established, their biological effects within a complex mixture are less understood. Therefore, we investigated the influence of DE on the carcinogenic potency of these PAH through measurement of PAH-DNA adducts, metabolic activity and oxidative DNA damage in human breast epithelial cells in culture (MCF-10A). We further investigated changes in the expression of genes involved in PAH metabolism provides insight into potential mechanisms of modification by DE. Our results are consistent with the hypotheses that complex mixtures modify the potency of PAH by altering metabolic activation pathways and subsequent carcinogenic metabolites.
MATERIALS AND METHODS
Caution
B[a]P and DB[a,l]P are potent carcinogenic agents and were handled according to the NCI guidelines for the use of carcinogens. Standard Reference Material (SRM) 1975 (DE) contains known mutagenic compounds and should be handled with the same precautions. PAHs and DE were prepared and used under yellow light.
Chemicals and Reagents
Standard Reference Material (SRM) 1975, referred as diesel exhaust (DE) in this study, was obtained from the National Institute of Standards and Technology (NIST) (Gaithersberg, MD). A more detailed description of the components of SRM 1975 can be found at the following: https://srmors.nist.gov/certificates/1975.pdf. B[a]P (CAS registry number 50-32-8) and DB[a,l]P (CAS registry number 191-30-0) were purchased from Chemsyn Science Laboratories (Lenexa, KS). Dimethylsulfoxide (DMSO), tricaprylin, insulin, hydrocortisone, nuclease P1, prostatic acid phosphatase, potato apyrase (Grade VII), proteinase K, 7-ethoxyresorufin (ERES), resorufin, and NADPH were purchased from Sigma (St. Louis, MO). F12/DMEM cell growth media, horse serum, and human recombinant epidermal growth factor were purchased from Invitrogen (Carlsbad, CA). Tris-saturated phenol, RNase (DNase free), and RNase T1 were purchased from Boehringer-Mannheim (Indianapolis, IN). Cloned T4 polynucleotide kinase and snake venom phosphodiesterase were obtained from United States Biochemical Corporation (Cleveland, OH). [Y-33P] ATP was purchased from Perkin-Elmer (Wellesley, MA).
Cell culture and exposure
The MCF-10A cell line was kindly provided by Dr. Robert Pauly (Barbara Ann Karmanos Cancer Institute, Detroit, MI). This spontaneously, immortalized cell line is derived from primary, normal mammary epithelial cells. Cells were cultured in a 75 cm2 flask (Corning, Corning, NY) in a 1:1 mixture of F-12/DMEM growth media. The medium was supplemented with 5% horse serum, containing 10 mM HEPES buffer, 10 µg/ml insulin, 20 ng/ml epidermal growth factor, 100 ng/ml cholera enterotoxin (Biomol, Plymouth Meeting, PA), 0.5 µg/ml hydrocortisone. Cells were cultured at 37°C with 5% CO2. Cell cultures were subcultured at a ratio of 1:4 when flasks were 80–90% confluent. Twenty-four hours prior to exposure, cells were replenished with growth media. The cells were exposed with 75 µl DMSO (vehicle control), DE (400 µg), B[a]P (20.2 µg), DE (400 µg) plus B[a]P (20.2 µg), DB[a,l]P (0.2 µg), or DE (400 µg) plus DB[a,l]P (0.2 µg). The exposure concentrations were chosen based on previous studies in our laboratory, which indicated that these doses gave detectable levels of DNA binding [22–24]. The cells were harvested 24 and 48 h after exposure, and stored at −80°C until further analysis.
MTT Cell Viability Assay
Cell viability and proliferation following PAH or DE plus PAH exposures were measured by the reduction of yellow tetrazolium MTT (3-[4, 5-dimethylthiazolyl-2]-2, 5-diphenyltetrazolium bromide) in MCF-10A cells for 24 and 48 h, according to the manufacturer’s instructions (ATCC, Manassas, VA).
DNA Isolation
A standard DNA isolation protocol was used, as reported previously [25]. Briefly, frozen cell pellets were homogenized in a glass homogenizer with EDTA-SDS buffer [10 mM Tris, 1 mM Na2EDTA, 1% SDS (w/v), pH 8]. The homogenates were exposed with DNase-free RNase, (50 U/ml) and RNase T1 (1000 U/ml) at 37°C for 1 h, followed by exposure with proteinase K (500 µg/ml) (Sigma, St. Louis, MO) at 37°C for 1 h. The DNA was extracted with equal volumes of Tris-equilibrated phenol (Boehringer-Mannheim, Indianapolis, IN), followed by extraction with 1:1 volume of Tris-equilibrated phenol and chloroform:isoamyl alcohol (24:1) and then with equal volumes of chloroform:isoamyl alcohol (24:1). The aqueous layer was exposed with one-tenth the volume of 5 M NaCl and twice the volume of cold 100% ethanol was added to precipitate the DNA which was then dissolved in double-distilled water. The DNA quality was confirmed by at least a 1.7 260/280 nm UV absorbance, and DNA concentration was based on the absorbance at 260 nm.
33P-Postlabeling of PAH-DNA Adducts
33P-postlabeling was carried out as described previously [26]. Briefly, 10 µg DNA isolated from exposed MCF-10A cells were digested with nuclease P1 and prostatic acid phosphatase, post-labeled with [γ-33P]ATP (3,000 Ci/mmol), cleaved to adducted mononucleotides with snake venom phosphodiesterase I, and pre-purified with a Sep-Pak C18 cartridge (Waters, Milford, MA). Subsequent separation by analytical high performance liquid chromatography (HPLC-Varian system equipped with two pumps and an autosampler; Varian Systems, Walnut Creek, CA) was carried out using a 5-µm Symmetry® C18 reverse-phase column (4.6 × 250 mm; Waters, Milford, MA). DNA-adducts purified from unused 33P-ATP were resolved by elution at 1 mL/min with 0.1 M ammonium phosphate, pH 5.5 (solvent A) and 100% HPLC-grade methanol (solvent B). The HPLC elution gradient for B[a]P and DE samples was: 44–55% solvent B over 5 min, 55–60% solvent B over 20 min, and elution at 100% solvent B over 5 min. The DB[a,l]P-DNA adducts were resolved by elution at 1 mL/min with 0.1 M ammonium phosphate, pH 5.5 (solvent A) and 90% HPLC grade methanol/10% acetonitrile (solvent B). The elution gradient was as follows: 20–44% solvent B over 20 min, 44–60% solvent B over 40 min, 60–80% solvent B over 15 min, and 80–20% solvent B over 1 min. The radiolabeled nucleotides were detected by an on-line dry cell β-RAM® Model 3 (INUS, Tampa, FL) radioisotope detector, and the level of DNA binding was calculated based on the labeling efficiency of a [3H]B[a]P-7,8-dihydrodiol 9,10-epoxide standard [27]. To determine the total PAH-DNA adduct levels, three independent sets of the postlabeling reactions were carried out for every sample.
Microsome isolation
Microsomes were prepared as described previously [28], with minor modifications. Briefly, cell pellets from different exposure groups were homogenized with a steel homogenizer in microsomal homogenization buffer [0.25 M K2HPO4, 0.15 M KCl, 10 mM EDTA, and 0.25 mM phenylmethylsulfonylfluoride (PMSF)] and were centrifuged at 15,000 × g for 20 min at 4°C. The supernatant was centrifuged at 100,000 × g for 90 min at 4°C, and the pellet was resuspended in microsome dilution buffer (0.1 M KH2PO4, 20% glycerol, 10 mM EDTA, 0.1 mM DTT and 0.25 mM PMSF). Protein concentrations were determined by the BCA protein assay (Pierce, Rockford, IL).
Ethoxyresorufin O-Deethylation (EROD) Assay
Thirty micrograms of microsomal protein isolated from samples in each exposure group were added to 1 µM 7-ethoxyresorufin (ERES) in 200 µl 0.1 M Tris-HCl (pH 7.8) buffer per well in a black 96 well plate (E&K Scientific, Campbell, CA). NADPH was added to each well and the fluorescence was measured on Spectra MAX Gemini plate reader (Molecular Devices, Sunnyvale, CA). The excitation wavelength was 535 nm, emission 585 and the kinetic assay was monitored over 15 min. Each sample was assayed in triplicate and the amount of resorufin produced was calculated from the fluorescence of a known concentration of resorufin. Ten micrograms of microsomal protein from Chinese Hamster Ovary (CHO) V79 cells overexpressing human CYP1A1 were used as the positive control.
Quantitative Real-Time PCR (qPCR) Analysis
Relative qPCR was performed with SYBR Green PCR Core Reagents and an ABI PRISM 7500 Sequence Detection System (PE Applied Biosystems, Foster City, CA), using the Standard Method according to manufacturer’s instructions. Reverse transcription was performed using 10 µg RNA (Advantage RT-for-PCR Kit, Clontech-Takara Bio, Mountain View, CA). One hundred micrograms cDNA was used as a template to validate primer oligonucleotides (150 nM) with optimized PCR conditions with the Eppendorf Master Mix 2.5× (Eppendorf North America Inc., Westbury NY) and agarose gel electrophoresis. Primer sequences were as follows: CYP1B1 (NM_000104) (5’-GGGACCGTCTGCCTTGTATGG-3’ and 5’-GTGTTGGCAGTGGTGGCATGAG-3’); AKR1A1 (NM_153326) (5’-GCGGCTTCCTGTGTTCTACT and 5’-CTTCCAGGTACCCAGACCAA-3’), AKR1C1 (NM_001353) (5’-AAAGCCAGGTGAGGAAGTGA and 5’-CATGTGGCACAGAGATCCAC-3’), AKR1C2 (UO5598) (5’-GGAGGCCATGGAGAAGTGTA-3’ and 5’-AGCAGCCTGTGGTTGAAGTT-3’), AKR1C3 (NM_003739) (5’-AGCCAGGTGAGGAACTTTCA-3’ and 5’-CCTCCCAGGTGGTACAGAGA-3’), NQO1 (NM_000903) (5’-CCTGCAGTGGTTTGGAGTC-3’ and 5’-CGGAAGGGTCCTTTGTCATA-3’), ALDH3A1 (BC008892) (5’-GGGAACTCAGTGGTCCTCAA-3’ and 5’-TACTGGGGATGATGGTAGC-3’), TNFRSF21 (BC010241) (5’-TTGTCTGTGAGCTCCCTGTG-3’ and 5’-ACCGCACATCCTCAGTCTCT-3’), CYR61 (NM_001554) (5’-CAGCTGACCAGGACTGTGAA-3’ and 5’-CCAGCTCCTACTCCAGCAAC-3’), MCL1 (BF594446) (5’-TGGGTTTGTGGAGTTCTTCC-3’ and 5’-CCAGCTCCTACTCCAGCAAC-3’), LGALS8 (NM_006499) (5’-CATTAGGTCCTGCTGGGTGT-3’ and 5-‘GGCAGAGTGGCTTTAACTGG-3’), and FOS (BC004490) (5’-AGAATCCGAAGGGAAAGGAA-3’ and 5-TCTCCGCTTGGAGTGTATCA-3’). The initial melting temperature (95 °C) was held for 1 min, and was followed by 30 cycles of melting (95 °C for 5 or 30 s), annealing at optimum temperature for primers (54–66 °C for 30 s), and a final extension (72 °C for 7 min). Purified DNA was excised (MinElute gel extraction kit; Qiagen, Valencia, CA) and amplified cDNA was quantified spectrophotometrically (Nanodrop ND-100; NanDrop Technologies, Wilmington, DE). After validating primers, 1 µl of cDNA was used as a template for amplification of specific gene products in total volumes of 50 µl reactions containing: SYBR Green PCR Master Mix (1×), MgCl2 (3 mM), dNTPs (1 mM), primers (100 pmol each) (Invitrogen, Carlsbad, CA), AmpliTaq Gold (1.25 U) and Amp Erase UNG (0.25 U). qPCR consisted of initial denaturation (95°C, 10 min), then 40 cycles of denaturation (95°C, 15 s) and annealing/extension (60°C, 1 min). A standard curve of cycle number to threshold versus cDNA copy number was generated for each target gene. Each sample was assayed in triplicate and comparative threshold cycle (CT) method was used to quantify the relative differences in PCR product.
Detection of 8-oxo-dG Adducts
Measurement of 8-oxoguanine (8-oxo-dG) adduct levels was carried out using the OxyFLOW™ system by HemoGenix, Inc (Colorado Springs, CO), according to the manufacturer’s instructions. OxyFLOW™ is a flow cytometric test to determine oxidative DNA damage using a fluorochrome (FITC-conjugated protein binding reagent) against 8-oxo-dG adducts in fixed permeabilized cells. Briefly, MCF-10A cells were exposed with PAH or PAH plus DE, as described above with tricaprylin as the vehicle control (DMSO is a known hydroxyl radical scavenger [29]). After MCF-10A cells were exposed for 24 h trypsinized and harvested, they were resuspended in 2 mL PBS, centrifuged at 500 × g for 5 min, and the supernatant discarded. The cells were resuspended in 500 µl OxyFLOW™ lysis/fix reagent and incubated for 15 min at room temperature. Samples were rinsed with 3 mL PBS, mixed by inversion, centrifuged for 5 min at 500 × g, and supernatant discarded. Cells from each exposure group were exposed with 200 µl ice-cold OxyFLOW™ permeabilization reagent for 2 min on ice and rinsed with PBS. Three milliliters of OxyFLOW™ Wash Buffer was added to each sample, mixed by inversion and centrifuged for 10 min at 500 × g. After discarding the supernatant, 100 µl of OxyFLOW™ FITC-conjugated protein binding reagent was added to each sample and incubated in the dark for 45 min. Samples were then washed with OxyFLOW™ Wash Buffer, centrifuged for 10 min at 500 × g. Cells were resuspended in 1 mL PBS and analyzed by flow cytometry. At least 10,000 events were measured from unexposed controls, samples and methylene blue controls.
Statistical Analysis
DNA adduct and EROD log-transformed data from each experiment were analyzed using the Mixed procedure in SAS version 9.1 (SAS Statistical system, Cary, NC, 2005. For DNA adduct data, duplicate or triplicate measurements on each sample were first averaged prior to transformation and analysis. A priori pairwise treatment comparisons were analyzed separately using all sample sets with usable information on both treatments and with the distribution of the data determining the analysis method. For the comparison of PAH and DE plus PAH treatment groups, analysis of variance (ANOVA) was used with three replicate complete blocks (3 or 4 sets of 33P-postlabeling reactions for each sample) and a two-by-two factorial treatment structure (time and exposure as factors). No significant difference in PAH-DNA adduct levels were found between 24 and 48 h (P = 0.428); hence, the results reported here represents the main effect of PAH-DNA adduct levels averaged over both time points. For EROD data, the model had three blocks (3 sets for each sample) and a three-by-three factorial exposure structure (exposure and time as factors). The EROD data were analyzed using a linear mixed model consistent with a randomized complete block design (with replicates as random blocks). For log-scale analysis of EROD data, the two zero measurements obtained from the raw data set were replaced with one-half of the smallest observed positive response. The Unpaired Students t-Test was used to compare fold-changes in gene expression and populations with detectable levels of 8-oxo-dG with GraphPad Prism software 4.00 for Windows (GraphPad Software, San Diego, CA).
RESULTS
Cell Viability and Proliferation
The MTT assay was conducted to determine cell viability and proliferation upon exposure to both PAH and DE plus PAH exposures for 24 and 48 h. The vehicle control (DMSO) and PAH exposures did not affect cell viability and proliferation (data not shown).
Total PAH-DNA Adduct Levels
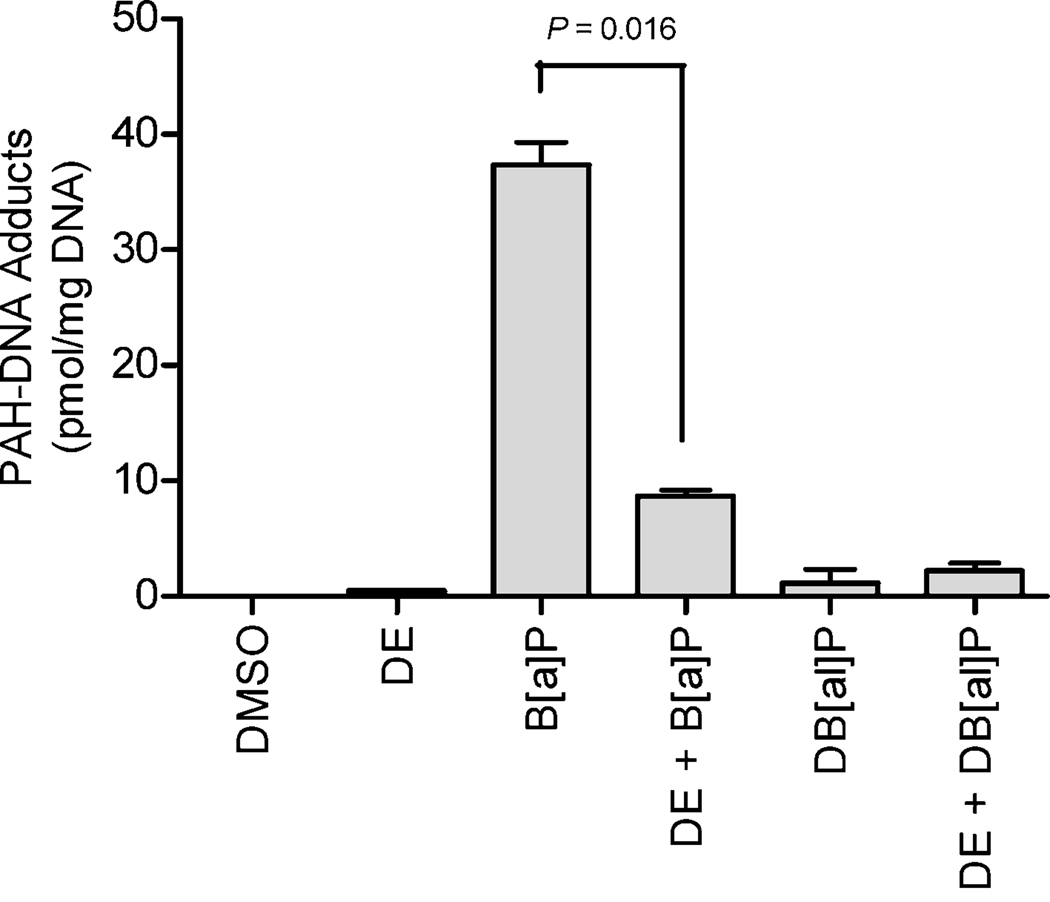
In order to determine the effects of DE on PAH-DNA adduct formation, cells were co-exposed with DE plus B[a]P or DB[a,l]P. PAH-DNA adduct levels were determined by 33P-postlabeling with reversed-phase HPLC analysis. No significant difference was observed between PAH-DNA adduct levels observed at 24 and 48 h (P > 0.05), therefore results are reported as an average of effects of each exposure group over time. Low levels of PAH-DNA adducts were observed from cells exposed with DE (0.517 adducts) (Fig.1). Cells exposed with DE plus B[a]P exhibited a significant decrease (P = 0.016) in DNA adduct levels (8.692 ± 4.873 [SEM]), compared with B[a]P exposure alone (37.41 ± 10.69), respectively (Fig.1). Conversely, no significant difference (P = 0.428) was observed in cells co-exposed with DE plus DB[a,l]P (1.180 ± 1.750), compared with DB[a,l]P alone (2.21 ± 0.692), respectively (Fig.1).
Figure 1.
The effect of DE exposure on the levels of B[a]P-diolepoxide- or DB[a,l]P-diolepoxide DNA adducts in MCF-10A cells in culture. Isolated DNA was 33P-postlabled and analyzed by reversed-phase HPLC (as described in Materials and Methods). Data points indicate means with standard error bars over n = 3 or 4 separate experiments. No effect on adduct levels were found between time of exposure (P > 0.05). DE significantly decreased the level of B[a]PDE-DNA adducts, compared with B[a]P treatment alone (P = 0.016). Whereas, DE had no significant influence on the level of DB[a,l]PDE-DNA adducts (P = 0.428).
Effect of DE on PAH-Induced CYP Metabolic Activity
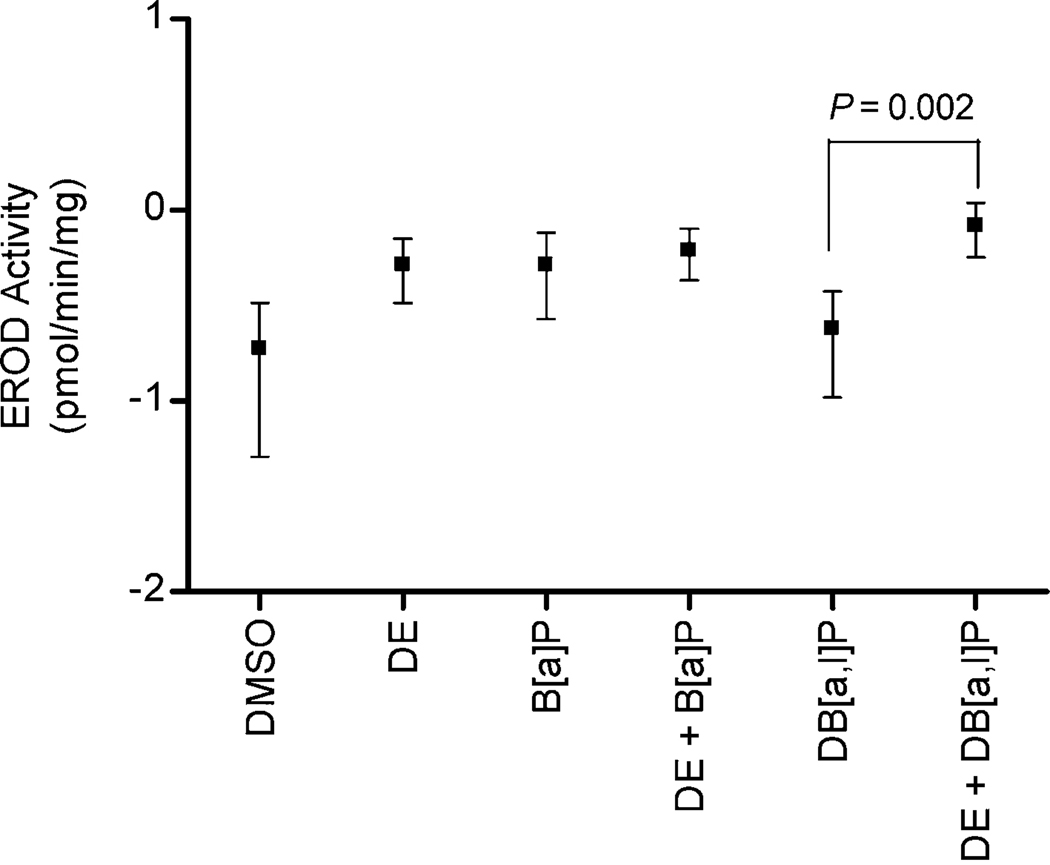
The EROD activity assay was conducted to determine the effect of DE on CYP-mediated metabolism of PAH following 24 and 48 h exposure. Over 24 and 48 h time points, no significant difference (P = 0.06) in EROD activity was observed between exposures; therefore, results are reported as an average of effects from each exposure group over time and on log-scale (Fig. 2). Cells exposed to DE exhibited a significant increase (P = 0.003) in EROD activity (0.52 ± 0.19 pmol/min/mg) compared with DMSO (vehicle control) (0.19 ± 0.14 pmol/min/mg). EROD activity significantly increased (P = 0.002) in cells co-exposed with DE plus DB[a,l]P (0.83 ± 0.26 pmol/min/mg), compared with DB[a,l]P exposure alone (0.24 ± 0.14 pmol/min/mg). No statistically significant difference was observed in cells exposed to DE plus B[a]P (0.61 ± 0.19 pmol/min/mg), compared with B[a]P alone (0.51 ± 0.25 pmol/min/mg).
Figure 2.
The influence of DE on B[a]P or DB[a,l]P-induced EROD activity in MCF-10A cells in culture. Data represents three individual experiments averaged over 24 and 48 h, presented as a simple mean ± SEM on log-scale. A statistically significant difference (P = 0.002) was observed in cells co-exposed to DE plus DB[a,l]P, compared with DB[a,l]P alone.
Effect of DE on PAH-Induced Gene Expression
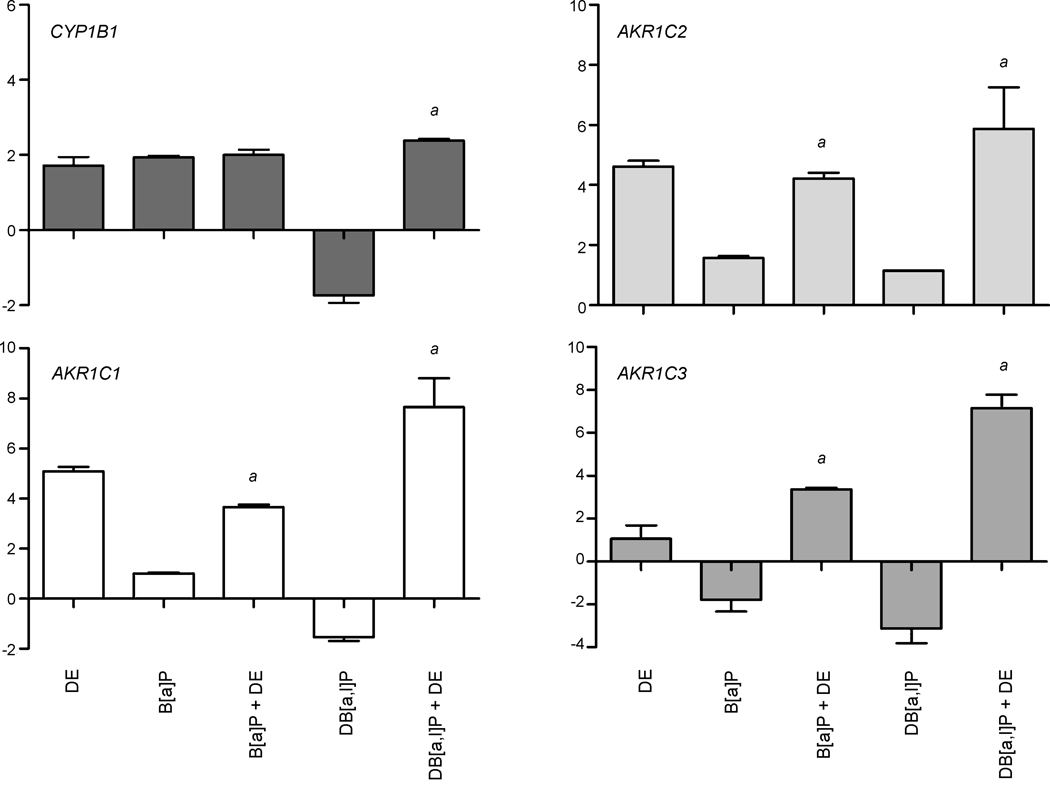
Gene expression changes in response to PAH or DE plus PAH are represented in Table 1. Candidate genes were chosen based on genes previously reported to be upregulated in a human breast carcinoma cell line (MCF-7) upon exposure to the same conditions, as measured by global gene expression analysis utilizing microarrays [30,31]. Candidate genes were also selected based on previous studies reporting effects of mixtures on the expression of PAH metabolism genes (CYP1B1, AKR1A1, AKR1C1, AKR1C2, AKR1C3, NQO1, and ALDH3A1), and those known to be overexpressed in breast tumors. (CYR61, MCL1, LGALS8, and FOS) [32,33]. CYP1B1 mRNA expression significantly increased (P ≤ 0.05) in response to DE plus DB[a,l]P, compared with DB[a,l]P alone (4.14-fold increase), respectively. Co-exposure of B[a]P or DB[a,l]P plus DE resulted in a significant increase (P ≤ 0.05) in the expression of AKR1C1, AKR1C2 and AKR1C3, compared with PAH exposure alone (Fig. 3, Table 1). AKR1C1, AKR1C2 and AKR1C3 expression significantly increased by 3.65, 2.68, and 5.15-fold, respectively, in response to B[a]P plus DE, compared with B[a]P exposure alone. AKR1C1, AKR1C2 and AKR1C3 significantly increased by 9.21, 5.10, and 10.28-fold, respectively, in cells co-exposed with DB[a,l]P plus DE, compared with DB[a,l]P exposure alone.
Table 1.
Gene Expression (Fold-Change) in Response to PAH or PAH + DE Exposure (24 h) Compared to DMSO
| PAH Exposure | ||||||
|---|---|---|---|---|---|---|
| Gene | Description | DE | B[a]P | B[a]P + DE | DB[a,l]P | DB[a,l]P + DE |
| CYP1B1 | Cytochrome P450 family 1, subfamily B, polypeptide 1 | 1.72 ± 0.40 | 1.94 ± 0.07 | 2.01 ± 0.23 | −1.75 ± 0.33 | 2.39a ± 0.07 |
| AKR1A1 | Aldo-keto reductase family 1, member A1 | −1.19 ± 0.07 | −1.94 ± 0.63 | −1.66 ± 0.10 | −1.30 ± 0.11 | −0.43 ± 1.37 |
| AKR1C1 | Aldo-keto reductase family 1, member C1 | 5.09 ± 0.30 | 1.00 ± 0.08 | 3.65a ± 0.18 | −1.55 ± 0.25 | 7.66a ± 2.00 |
| AKR1C2 | Aldo-keto reductase family 1, member C2 | 4.62 ± 0.32 | 1.57 ± 0.11 | 4.22a ± 0.33 | 1.15 ± 0.01 | 5.87a ± 2.40 |
| AKR1C3 | Aldo-keto reductase family 1, member C3 | 1.06 ± 1.07 | −1.79 ± 0.94 | 3.36a ± 0.14 | −3.13 ± 1.19 | 7.15a ±1.08 |
| NQO1 | NADPH dehydrogenase, quinone | 1.06 ± 0.04 | −1.55 ± 0.09 | −1.19 ± 0.09 | −2.25 ± 0.05 | 1.42 ± 0.10 |
| ALDH3A1 | Aldehyde dehydrogenase 3 family, member A1 | 1.53 ± 0.07 | 2.49 ± 0.38 | 2.80 ± 0.38 | 1.00 ± 0.19 | 5.21a ± 0.43 |
| CYR61 | Cysteine-rich, angiogenic inducer, 61 | 549.55 ± 442.04 | 193.23 ± 72.07 | 7.37 ± 8.74a | 442.31 ± 365.46 | 1728.49 ± 1129.13 |
| MCL1 | Myeloid cell leukemia sequence 1 (BCL2-related) | 123.75 ± 82.94 | 80.93 ± 80.92 | 20.12 ± 8.11 | −1.57 ± 0.08 | n/a |
| LGALS8 | Lectin galactoside-binding, soluble 8 (galectin 8) | −1.25 ± 0.12 | −2.00 ± 0.91 | −2.77 ± 0.41 | −3.11 ± 1.03 | −2.43 ± 0.57 |
| FOS | v-fos FJB murine osteosarcoma viral oncogene homologue | −2.28 ± 0.08 | −3.18 ± 1.38 | −2.23 ± 0.40 | −1.54 ± 0.55 | −1.44 ± 0.28 |
The data represents mean fold-change ± SD where n = 3 replicates per exposure.
Genes that were significantly altered (P ≤ 0.01) compared to B[a]P or DB[a,l]P, as determined by Student’s t-Test.
Figure 3.
Changes in expression of genes (as measured by quantitative real-time PCR) involved in PAH metabolism following 24 h exposure to PAH or PAH plus DE, compared with dimethyl sulfoxide (DMSO; vehicle control). Data represents mean ± SD (n = 3 replicates per exposure). aGenes that were significantly altered (P ≤ 0.01) compared to B[a]P or DB[a,l]P.
8-oxo-dG Adduct Levels
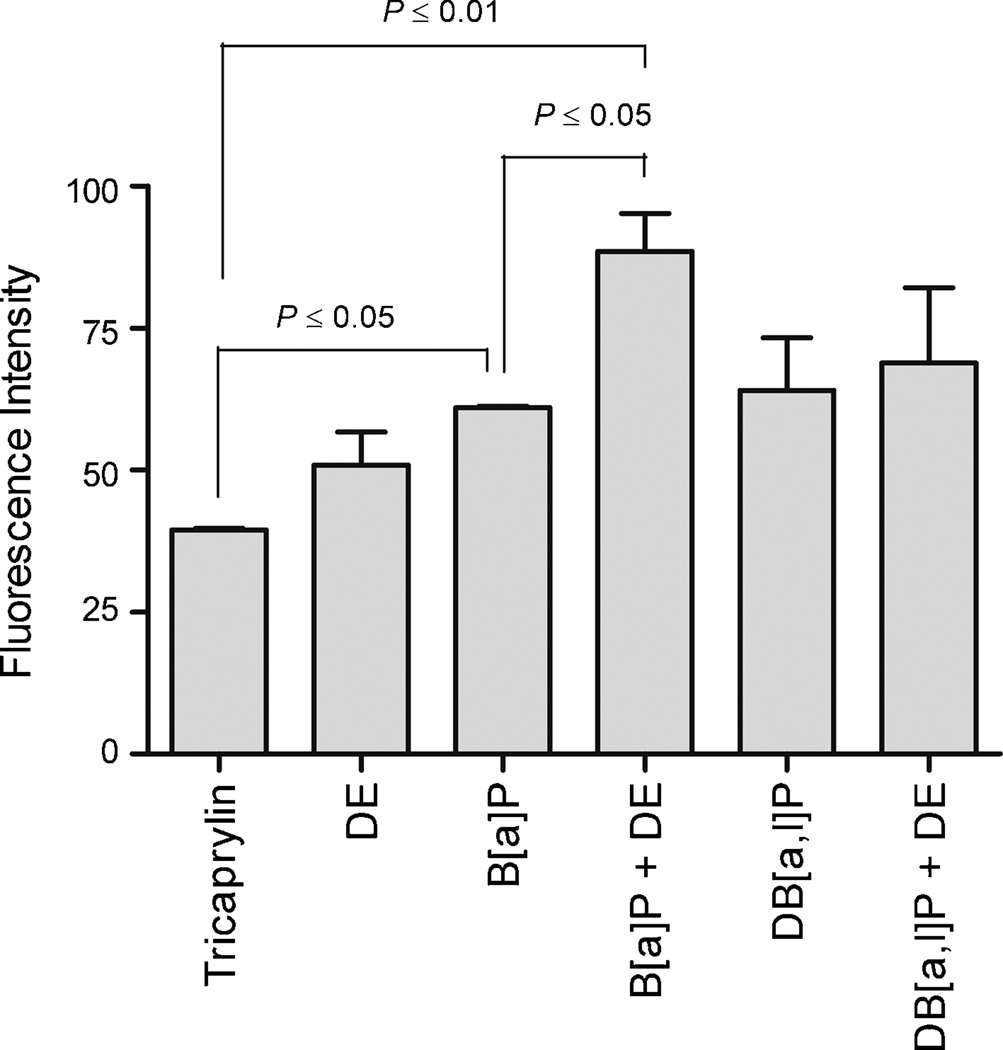
In order to determine the effects of PAH and DE plus PAH on oxidative damage, cell populations exhibiting 8-oxo-dG levels were quantified using a fluorescent, FITC-conjugated antibody and flow cytometry. A significant increase (P ≤ 0.01) in the population of cells with detectable 8-oxo-dG adducts was observed in B[a]P and B[a]P plus DE exposed cells, compared with tricaprylin exposure (vehicle control) (Fig. 4).
Figure 4.
Cell populations exhibiting 8-oxo-dG adducts, following 24 h exposure to vehicle control (tricaprylin), PAH or PAH plus DE. Adduct levels were quantified by fluorescence emitted by FITC-conjugated protein binding agent, measured by flow cytometry using the OxyFLOW® platform (HemoGenix, Inc, Colorado Springs, CO). Data represents mean ± SEM (n = 3 replicates per exposure).
DISCUSSION
PAHs are found in many environmental complex mixtures such as urban air pollution, diesel exhaust, and tobacco smoke condensates. The characterization of benzo[a]pyrene (B[a]P) and dibenzo[a,l]pyrene (DB[a,l]P), and their mechanisms of genotoxicity are well established; however limited information is available regarding their carcinogenic potency within a complex mixture. Evaluations of complex mixtures are difficult, as mixture components may have additive, synergistic or antagonistic effects on metabolic processes thereby influencing the potency of individual carcinogens [34,35]. The effects and mechanisms surrounding such interactions must be established in order to define true risk.
The selection of a cell line derived from normal human breast epithelial cells (MCF-10A) reflects our interest in the contributing role of PAH in breast cancer. Carcinogenic PAH metabolites have been detected in both normal and tumor human breast tissue [16,17], and implicated in breast tumorigenesis [18–21]. Our studies focused on the influence of DE on PAH bioactivation pathways and the genotoxic effects generated by carcinogenic metabolites in MCF-10A cells.
Bioactivation of PAHs adsorbed to the particle surface of DE results in the formation of bulky DNA adducts, and previous studies have demonstrated increased DNA adduct levels in rat lung following both short- and long-term inhalation exposure to DE [36–38]. Here we report the effect of an organic extract of DE particulate matter on PAH-DNA adduct formation, CYP-mediated bioactivation, gene expression, and oxidative DNA damage. Previous studies have detected PAH-DNA adducts following exposure to a variety of DE mixtures [36,39]; however, we report DE exposure resulted in low PAH-DNA adduct levels. This could in part be attributed to the absence of PAH commonly attributed to the mutagenicity of DE, such as 3-nitrobenzanthrone (3-NBA) [40], B[a]P and DB[a,l]P. However, we demonstrated that DE significantly decreased B[a]P-induced PAH-DNA adduct levels and no change in CYP metabolic activity levels (Fig. 2), compared with B[a]P exposure alone. These antagonistic effects may result from a reduction in the conversion of mutagens into their reactive metabolites, which could be attributed to saturation of bioactivation systems [34,41]. Conversely, no significant difference in PAH-DNA adduct levels was observed between DB[a,l]P and DE plus DB[a,l]P exposed cells; however, CYP metabolic activity and CYP1B1 expression significantly increased in response to DE plus DB[a,l]P, compared with DB[a,l]P exposure alone (Fig. 1, Fig. 2, Table 1). These results may be attributed to an upregulation of CYP-mediated bioactivation of DB[a,l]P; however, DE may influence the regio- and stereoselectivity of CYP1B1 to produce less carcinogenic stereoisomers of DB[a,l]PDE, or increase the metabolism of other PAH present in the mixture. In addition, it was recently reported that such additive effects may occur at low concentrations (0.5 µg/ml) [42]. This sub-additive effect suggests that DE may have additive or synergistic effects on CYP-mediated metabolism of PAH at low concentrations.
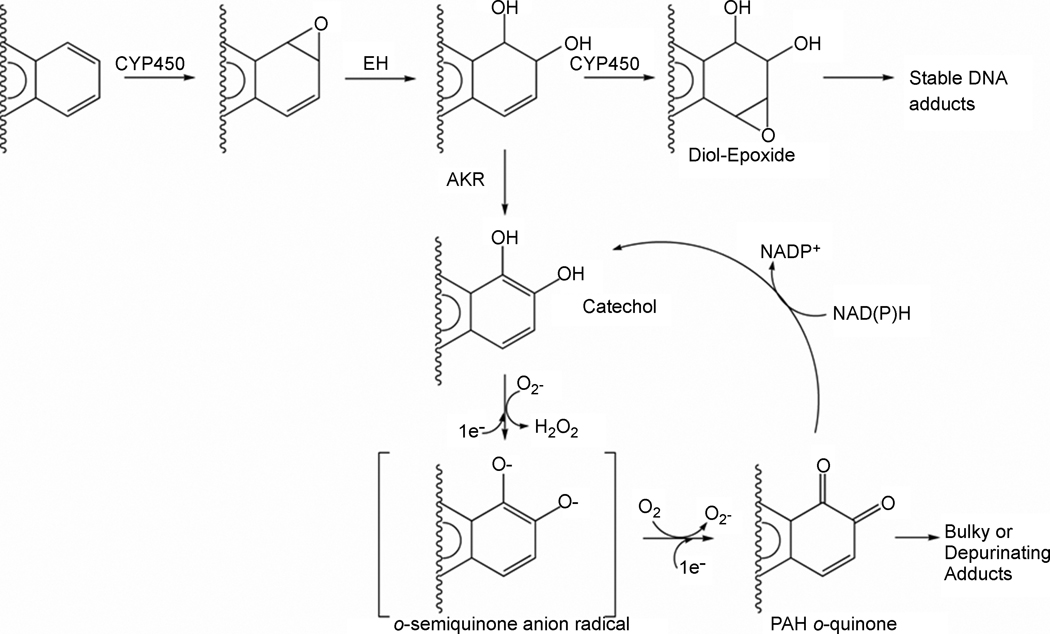
Complex mixtures have the potential to shift metabolism from bioactivation to detoxification or to another bioactivation pathway [43]. Previous studies demonstrated that complex mixtures inhibited CYP-mediated bioactivation [22,30,44,45], and suggested that alternate metabolism pathways compensate for the bioactivation of PAH [44]. The aldo keto-reductase (AKR) pathway is known to compete with CYP1A1 and CYP1B1 for B[a]P-7,8-dihydrodiol metabolism [46,47] (Fig. 5). Previous studies have also reported increased expression of AKR1C genes in human cells following exposure to urban dust particulate matter (UDPM) [30,48], DE [31], and cigarette smoke condensate [49]. We reveal that DE significantly increased the expression of AKR1C1 and AKR1C2, compared with DMSO (vehicle control) exposure. We further observed that co-exposure to DE plus PAH significantly increased the expression levels of AKR1C1, AKR1C2 and AKR1C3, compared with PAH exposure alone. Because CYP metabolic activity was not diminished and an increase in the expression of AKR genes in response to DE plus PAH was observed, it is possible that the metabolism of PAH-dihydrodiols could be shifted to the production of PAH o-quinones by AKRs, rather than PAH-diol epoxides by CYP enzymes (Fig. 5). This would further account for the significant decrease in PAH-DNA adduct levels in DE plus B[a]P exposed cells, and no change in the total PAH-DNA adduct levels resulting from DE plus DB[a,l]P exposure. These results support the hypothesis that complex mixtures influence PAH-induced genotoxicity by modulating a shift in metabolic pathways.
Figure 5.
Competing pathways for the metabolic activation of PAH by CYP, cytochrome P450, and AKR, aldo-ketoreductase (adapted from T.M. Penning [56]). Schematic also illustrates the effect of CYP inhibition by mixture components, as suggested previously [44].
Under redox conditions, ROS is a by-product of the AKR metabolic pathway. We observed a significant increase in the population of cells exhibiting 8-oxo-dG adducts in response to B[a]P and DE plus B[a]P exposures, compared with vehicle control (Fig. 4, Fig. 5). Such ROS insults generated by this pathway may contribute to the etiology of cancer through change-in-function gene mutations in critical oncogenes and tumor suppressor genes, such as p53. Yu et al [14] recently demonstrated that B[a]P-7,8-dione (BPQ) was a potent mutagen, and that the dominant G-to-T transversion mutations observed in p53 were abolished by ROS scavengers, suggesting that the mutations were oxidatively generated.
Mammary epithelial cells have a high capacity to activate B[a]P and DB[a,l]P [50], and are well-established mammary carcinogens in rodent models [51,52]. PAH exposure has been associated with human breast cancer [17,19,20], and it is suggested that environmental PAH such as B[a]P may contribute to breast carcinogenesis [53]. Although the genes known to be over-expressed in breast tumors were not altered in response to co-exposure of DE plus PAH, higher levels of polar or depurinating adducts and oxidative metabolites may be implicated in the various stages of breast cancer through mutagenesis.
This study and several in vitro and in vivo studies have examined the influence of complex mixtures on CYP-mediated bioactivation, PAH-DNA binding and tumor initiation [30,54,55]. We demonstrated that DE upregulates PAH metabolism genes specific to the AKR pathway, and is responsible for increased B[a]P-induced 8-oxo-dG adduct formation. Therefore, the carcinogenic potency of PAH within complex mixtures is dependent on the induction of metabolic enzymes involved in different PAH bioactivation pathways. These results support the idea that simply taking the sum of the actual mutagenic hazard posed by mixture constituents is not reasonable to determine human cancer risk. Further, using highly potent PAH, such as B[a]P or DB[a,l]P, as surrogates to estimate risk to environmental complex mixtures is not adequate, as bioactivation or detoxification pathways may shift. Based on the results presented here, in vivo carcinogenicity studies evaluating the effects of environmental complex mixtures on competing bioactivation pathways (e.g. AKR or radical cation pathways) and the genotoxic metabolites they produce are therefore warranted.
ACKNOWLEDGEMENTS
We wish to thank Dr. Brinda Mahadevan for critical review of this manuscript. We thank Mr. Jack Giovanini of the Integrative Health Sciences Facilities and Services Core of the Environmental Health Sciences Center at OSU for statistical assistance, supported by the National Institute of Environmental Health Sciences, National Institute of Health, P30 ES00210. This work was also supported by CA28825, Department of Health and Human Services, from the National Cancer Institute and the NIEHS Pre-doctoral Traineeship to L.A.C. (T32 ES007060).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- 1.Dockery DW, Pope CA, 3rd, Xu X, Spengler JD, Ware JH, Fay ME, Ferris BG, Jr, Speizer FE. An association between air pollution and mortality in six U.S. cities. N Engl J Med. 1993;329:1753–1759. doi: 10.1056/NEJM199312093292401. [DOI] [PubMed] [Google Scholar]
- 2.Samet JM, Dominici F, Curriero FC, Coursac I, Zeger SL. Fine particulate air pollution and mortality in 20 U.S. cities, 1987–1994. N Engl J Med. 2000;343:1742–1749. doi: 10.1056/NEJM200012143432401. [DOI] [PubMed] [Google Scholar]
- 3.Pope CA, 3rd, Burnett RT, Thun MJ, Calle EE, Krewski D, Ito K, Thurston GD. Lung cancer, cardiopulmonary mortality, and long-term exposure to fine particulate air pollution. Jama. 2002;287:1132–1141. doi: 10.1001/jama.287.9.1132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Palli D, Russo A, Masala G, Saieva C, Guarrera S, Carturan S, Munnia A, Matullo G, Peluso M. DNA adduct levels and DNA repair polymorphisms in traffic-exposed workers and a general population sample. Int J Cancer. 2001;94:121–127. doi: 10.1002/ijc.1433. [DOI] [PubMed] [Google Scholar]
- 5.Burgaz S, Demircigil GC, Karahalil B, Karakaya AE. Chromosomal damage in peripheral blood lymphocytes of traffic policemen and taxi drivers exposed to urban air pollution. Chemosphere. 2002;47:57–64. doi: 10.1016/s0045-6535(01)00185-0. [DOI] [PubMed] [Google Scholar]
- 6.Soares SR, Bueno-Guimaraes HM, Ferreira CM, Rivero DH, De Castro I, Garcia ML, Saldiva PH. Urban air pollution induces micronuclei in peripheral erythrocytes of mice in vivo. Environ Res. 2003;92:191–196. doi: 10.1016/s0013-9351(02)00061-0. [DOI] [PubMed] [Google Scholar]
- 7.Bayona JM, Markides KE, Lee ML. Characterization of polar polycyclic aromatic compounds in a heavy-duty diesel exhaust particulate by capillary column gas chromatography and high-resolution mass spectrometry. Environ. Sci. Technol. 1988;22:1440–1447. doi: 10.1021/es00177a009. [DOI] [PubMed] [Google Scholar]
- 8.De Raat WK, Kooijman SA, Gielen JW. Concentrations of polycyclic hydrocarbons in airborne particles in The Netherlands and their correlation with mutagenicity. Sci Total Environ. 1987;66:95–114. doi: 10.1016/0048-9697(87)90080-5. [DOI] [PubMed] [Google Scholar]
- 9.Bolton JL, Trush MA, Penning TM, Dryhurst G, Monks TJ. Role of quinones in toxicology. Chem Res Toxicol. 2000;13:135–160. doi: 10.1021/tx9902082. [DOI] [PubMed] [Google Scholar]
- 10.Denissenko MF, Pao A, Tang M, Pfeifer GP. Preferential formation of benzo[a]pyrene adducts at lung cancer mutational hotspots in P53. Science. 1996;274:430–432. doi: 10.1126/science.274.5286.430. [DOI] [PubMed] [Google Scholar]
- 11.Ross JA, Nesnow S. Polycyclic aromatic hydrocarbons: correlations between DNA adducts and ras oncogene mutations. Mutat Res. 1999;424:155–166. doi: 10.1016/s0027-5107(99)00016-0. [DOI] [PubMed] [Google Scholar]
- 12.Brookes P, Lawley PD. Evidence for the Binding of Polynuclear Aromatic Hydrocarbons to the Nucleic Acids of Mouse Skin: Relation between Carcinogenic Power of Hydrocarbons and Their Binding to Deoxyribonucleic Acid. Nature. 1964;202:781–784. doi: 10.1038/202781a0. [DOI] [PubMed] [Google Scholar]
- 13.Nesnow S. Mouse skin tumours and human lung cancer: relationships with complex environmental emissions. IARC Sci Publ. 1990:44–54. [PubMed] [Google Scholar]
- 14.Yu D, Berlin JA, Penning TM, Field J. Reactive oxygen species generated by PAH o-quinones can cause change-in-function mutations in p53. Chem Res Toxicol. 2002;15:832–842. doi: 10.1021/tx010177m. [DOI] [PubMed] [Google Scholar]
- 15.Cavalieri EL, Rogan EG. Role of radical cations in aromatic hydrocarbon carcinogenesis. Environ Health Perspect. 1985;64:69–84. doi: 10.1289/ehp.856469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hecht SS. Tobacco smoke carcinogens and breast cancer. Environ Mol Mutagen. 2002;39:119–126. doi: 10.1002/em.10071. [DOI] [PubMed] [Google Scholar]
- 17.el-Bayoumy K. Environmental carcinogens that may be involved in human breast cancer etiology. Chem Res Toxicol. 1992;5:585–590. doi: 10.1021/tx00029a001. [DOI] [PubMed] [Google Scholar]
- 18.Perera FP, Estabrook A, Hewer A, Channing K, Rundle A, Mooney LA, Whyatt R, Phillips DH. Carcinogen-DNA adducts in human breast tissue. Cancer Epidemiol Biomarkers Prev. 1995;4:233–238. [PubMed] [Google Scholar]
- 19.Li D, Wang M, Firozi PF, Chang P, Zhang W, Baer-Dubowska W, Moorthy B, Vulimiri SV, Goth-Goldstein R, Weyand EH, DiGiovanni J. Characterization of a major aromatic DNA adduct detected in human breast tissues. Environ Mol Mutagen. 2002;39:193–200. doi: 10.1002/em.10063. [DOI] [PubMed] [Google Scholar]
- 20.Li D, Wang M, Dhingra K, Hittelman WN. Aromatic DNA adducts in adjacent tissues of breast cancer patients: clues to breast cancer etiology. Cancer Res. 1996;56:287–293. [PubMed] [Google Scholar]
- 21.Santella RM, Gammon MD, Zhang YJ, Motykiewicz G, Young TL, Hayes SC, Terry MB, Schoenberg JB, Brinton LA, Bose S, Teitelbaum SL, Hibshoosh H. Immunohistochemical analysis of polycyclic aromatic hydrocarbon-DNA adducts in breast tumor tissue. Cancer Lett. 2000;154:143–149. doi: 10.1016/s0304-3835(00)00367-0. [DOI] [PubMed] [Google Scholar]
- 22.Mahadevan B, Parsons H, Musafia T, Sharma AK, Amin S, Pereira C, Baird WM. Effect of artificial mixtures of environmental polycyclic aromatic hydrocarbons present in coal tar, urban dust, and diesel exhaust particulates on MCF-7 cells in culture. Environ Mol Mutagen. 2004;44:99–107. doi: 10.1002/em.20039. [DOI] [PubMed] [Google Scholar]
- 23.Mahadevan B, Keshava C, Musafia-Jeknic T, Pecaj A, Weston A, Baird WM. Altered gene expression patterns in MCF-7 cells induced by the urban dust particulate complex mixture standard reference material 1649a. Cancer Res. 2005;65:1251–1258. doi: 10.1158/0008-5472.CAN-04-2357. [DOI] [PubMed] [Google Scholar]
- 24.Musafia-Jeknic T, Mahadevan B, Pereira C, Baird WM. Long term effects of a standardized complex mixture of urban dust particulate matter on the metabolic activation of carcinogenic polycyclic aromatic hydrocarbons in human cells in culture. Toxicol Sci. 2005 doi: 10.1093/toxsci/kfi329. [DOI] [PubMed] [Google Scholar]
- 25.Luch A, Coffing SL, Tang YM, Schneider A, Soballa V, Greim H, Jefcoate CR, Seidel A, Greenlee WF, Baird WM, Doehmer J. Stable expression of human cytochrome P450 1B1 in V79 Chinese hamster cells and metabolically catalyzed DNA adduct formation of dibenzo[a,l]pyrene. Chem Res Toxicol. 1998;11:686–695. doi: 10.1021/tx970236p. [DOI] [PubMed] [Google Scholar]
- 26.Ralston SL, Coffing SL, Seidel A, Luch A, Platt KL, Baird WM. Stereoselective activation of dibenzo[a,l]pyrene and its Trans-11,12-dihydrodiol to fjord-region 11,12-diol 13,14-epoxides in a human mammary carcinoma MCF-7 cell-mediated V-79 cell mutation assay. Chemical Research in Toxicology. 1997;10:687–693. doi: 10.1021/tx9700275. [DOI] [PubMed] [Google Scholar]
- 27.Lau HH, Baird WM. Detection and identification of benzo[a]pyrene-DNA adducts by [35S]phosphorothioate labeling and HPLC. Carcinogenesis. 1991;12:885–893. doi: 10.1093/carcin/12.5.885. [DOI] [PubMed] [Google Scholar]
- 28.Otto S, Marcus C, Pidgeon C, Jefcoate C. A novel adrenocorticotropin-inducible cytochrome P450 from rat adrenal microsomes catalyzes polycyclic aromatic hydrocarbon metabolism. Endocrinology. 1991;129:970–982. doi: 10.1210/endo-129-2-970. [DOI] [PubMed] [Google Scholar]
- 29.Repine JE, Eaton JW, Anders MW, Hoidal JR, Fox RB. Generation of hydroxyl radical by enzymes, chemicals, and human phagocytes in vitro. Detection with the anti-inflammatory agent, dimethyl sulfoxide. J Clin Invest. 1979;64:1642–1651. doi: 10.1172/JCI109626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Mahadevan B, Marston CP, Dashwood WM, Li Y, Pereira C, Baird WM. Effect of a standardized complex mixture derived from coal tar on the metabolic activation of carcinogenic polycyclic aromatic hydrocarbons in human cells in culture. Chem Res Toxicol. 2005;18:224–231. doi: 10.1021/tx0497604. [DOI] [PubMed] [Google Scholar]
- 31.Mahadevan B, Keshava C, Weston A, Baird WM. Altered gene expression patterns in MCF-7 cells induced by the diesel exhaust complex mixture standard reference material 1650a. doi: 10.1158/0008-5472.CAN-04-2357. 200x. [DOI] [PubMed] [Google Scholar]
- 32.Henson ES, Hu X, Gibson SB. Herceptin sensitizes ErbB2-overexpressing cells to apoptosis by reducing antiapoptotic Mcl-1 expression. Clin Cancer Res. 2006;12:845–853. doi: 10.1158/1078-0432.CCR-05-0754. [DOI] [PubMed] [Google Scholar]
- 33.Danguy A, Rorive S, Decaestecker C, Bronckart Y, Kaltner H, Hadari YR, Goren R, Zich Y, Petein M, Salmon I, Gabius HJ, Kiss R. Immunohistochemical profile of galectin-8 expression in benign and malignant tumors of epithelial, mesenchymatous and adipous origins, and of the nervous system. Histol Histopathol. 2001;16:861–868. doi: 10.14670/HH-16.861. [DOI] [PubMed] [Google Scholar]
- 34.Hermann M. Synergistic effects of individual polycyclic aromatic hydrocarbons on the mutagenicity of their mixtures. Mutat Res. 1981;90:399–409. doi: 10.1016/0165-1218(81)90062-8. [DOI] [PubMed] [Google Scholar]
- 35.Donelly K, Brown KW, Anderson VS, Barbee GC, Safe SH. Metabolism and bacterial mutagenicity of binary mixtures of benzo[a]pyrene and polychlorinated aromatic hydrocarbons. Environ. Mol. Mutagen. 1990;16:238–245. doi: 10.1002/em.2850160404. [DOI] [PubMed] [Google Scholar]
- 36.Bond JA, Mauderly JL, Wolff RK. Concentration- and time-dependent formation of DNA adducts in lungs of rats exposed to diesel exhaust. Toxicology. 1990;60:127–135. doi: 10.1016/0300-483x(90)90167-f. [DOI] [PubMed] [Google Scholar]
- 37.Wong D, Mitchell CE, Wolff RK, Mauderly JL, Jeffrey AM. Identification of DNA damage as a result of exposure of rats to diesel engine exhaust. Carcinogenesis. 1986;7:1595–1597. doi: 10.1093/carcin/7.9.1595. [DOI] [PubMed] [Google Scholar]
- 38.Sato H, Sone H, Sagai M, Suzuki KT, Aoki Y. Increase in mutation frequency in lung of Big Blue rat by exposure to diesel exhaust. Carcinogenesis. 2000;21:653–661. doi: 10.1093/carcin/21.4.653. [DOI] [PubMed] [Google Scholar]
- 39.Savela K, King L, Gallagher J, Lewtas J. 32P-postlabeling and HPLC separation of DNA adducts formed by diesel exhaust extracts in vitro and in mouse skin and lung after topical treatment. Carcinogenesis. 1995;16:2083–2089. doi: 10.1093/carcin/16.9.2083. [DOI] [PubMed] [Google Scholar]
- 40.Arlt VM, Glatt H, Muckel E, Pabel U, Sorg BL, Schmeiser HH, Phillips DH. Metabolic activation of the environmental contaminant 3-nitrobenzanthrone by human acetyltransferases and sulfotransferase. Carcinogenesis. 2002;23:1937–1945. doi: 10.1093/carcin/23.11.1937. [DOI] [PubMed] [Google Scholar]
- 41.Donelly KC, Claxton LD, Huebner HJ, Capizzi JL. Mutagenic interactions of model chemical mixtures. Chemosphere. 1998;37:1253–1261. doi: 10.1016/s0045-6535(98)00123-4. [DOI] [PubMed] [Google Scholar]
- 42.White PA. The genotoxicity of priority polycyclic aromatic hydrocarbons in complex mixtures. Mutat Res. 2002;515:85–98. doi: 10.1016/s1383-5718(02)00017-7. [DOI] [PubMed] [Google Scholar]
- 43.Kawalek JC, Andrews AW. Effect of aromatic hydrocarbons on the metabolism of 2-aminoanthracene to mutagenic products in the Ames assay. Carcinogenesis. 1981;2 doi: 10.1093/carcin/2.12.1367. 1397–1369. [DOI] [PubMed] [Google Scholar]
- 44.Shimada T, Guengerich FP. Inhibition of human cytochrome P450 1A1-, 1A2-, and 1B1-mediated activation of procarcinogens to genotoxic metabolites by polycyclic aromatic hydrocarbons. Chem Res Toxicol. 2006;19:288–294. doi: 10.1021/tx050291v. [DOI] [PubMed] [Google Scholar]
- 45.Musafia-Jeknic T, Mahadevan B, Pereira C, Baird WM. Long-term effects of a standardized complex mixture of urban dust particulate on the metabolic activation of carcinogenic polycyclic aromatic hydrocarbons in human cells in culture. Toxicol Sci. 2005;88:358–366. doi: 10.1093/toxsci/kfi329. [DOI] [PubMed] [Google Scholar]
- 46.Jiang H, Shen YM, Quinn AM, Penning TM. Competing roles of cytochrome P450 1A1/1B1 and aldo-keto reductase 1A1 in the metabolic activation of (+/−)-7,8-dihydroxy-7,8-dihydro-benzo[a]pyrene in human bronchoalveolar cell extracts. Chem Res Toxicol. 2005;18:365–374. doi: 10.1021/tx0497245. [DOI] [PubMed] [Google Scholar]
- 47.Jiang H, Vudathala DK, Blair IA, Penning TM. Competing roles of aldoketo reductase 1A1 and cytochrome P4501B1 in benzo[a]pyrene-7,8-diol activation in human bronchoalveolar H358 cells: role of AKRs in P4501B1 induction. Chem Res Toxicol. 2006;19:68–78. doi: 10.1021/tx0502488. [DOI] [PubMed] [Google Scholar]
- 48.Courter L, Musafia-Jeknic T, Giovanini J, Pereira C, Fischer K, Bildfell R, Baird WM. Urban Dust Particulate Matter Alters PAH-Induced Carcinogenesis by Inhibition of CYP1A1 and CYP1B1. Toxicol Sci. 2006 doi: 10.1093/toxsci/kfl137. in press. [DOI] [PubMed] [Google Scholar]
- 49.Nagaraj NS, Beckers S, Mensah JK, Waigel S, Vigneswaran N, Zacharias W. Cigarette smoke condensate induces cytochromes P450 and aldoketo reductases in oral cancer cells. Toxicol Lett. 2006;165:182–194. doi: 10.1016/j.toxlet.2006.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Carmichael PL, Stone EM, Grover PL, Gusterson BA, Phillips DH. Metabolic activation and DNA binding of food mutagens and other environmental carcinogens in human mammary epithelial cells. Carcinogenesis. 1996;17:1769–1772. doi: 10.1093/carcin/17.8.1769. [DOI] [PubMed] [Google Scholar]
- 51.Cavalieri EL, Higginbotham S, RamaKrishna NV, Devanesan PD, Todorovic R, Rogan EG, Salmasi S. Comparative dose-response tumorigenicity studies of dibenzo[alpha,l]pyrene versus 7,12-dimethylbenz[alpha]anthracene, benzo[alpha]pyrene and two dibenzo[alpha,l]pyrene dihydrodiols in mouse skin and rat mammary gland. Carcinogenesis. 1991;12:1939–1944. doi: 10.1093/carcin/12.10.1939. [DOI] [PubMed] [Google Scholar]
- 52.Higginbotham S, RamaKrishna NV, Johansson SL, Rogan EG, Cavalieri EL. Tumor-initiating activity and carcinogenicity of dibenzo[a,l]pyrene versus 7,12-dimethylbenz[a]anthracene and benzo[a]pyrene at low doses in mouse skin. Carcinogenesis. 1993;14:875–878. doi: 10.1093/carcin/14.5.875. [DOI] [PubMed] [Google Scholar]
- 53.Jeffy BD, Chirnomas RB, Romagnolo DF. Epigenetics of breast cancer: polycyclic aromatic hydrocarbons as risk factors. Environ Mol Mutagen. 2002;39:235–244. doi: 10.1002/em.10051. [DOI] [PubMed] [Google Scholar]
- 54.Marston CP, Pereira C, Ferguson J, Fischer K, Hedstrom O, Dashwood WM, Baird WM. Effect of a complex environmental mixture from coal tar containing polycyclic aromatic hydrocarbons (PAH) on the tumor initiation, PAH-DNA binding and metabolic activation of carcinogenic PAH in mouse epidermis. Carcinogenesis. 2001;22:1077–1086. doi: 10.1093/carcin/22.7.1077. [DOI] [PubMed] [Google Scholar]
- 55.Courter LA, Musafia-Jeknic T, Fischer K, Bildfell R, Giovanini J, Pereira C, Baird WM. Urban Dust Particulate Matter Alters PAH-Induced Carcinogenesis by Inhibition of CYP1A1 and CYP1B1. Toxicol Sci. 2007;95:63–73. doi: 10.1093/toxsci/kfl137. [DOI] [PubMed] [Google Scholar]
- 56.Penning TM. Introduction and overview of the aldo-keto reductase superfamily. In: Penning TM, Petrash JM, editors. Aldo-keto Reductases and Toxicant Metabolism. Washington D.C.: American Chemical Society; 2004. pp. 3–20. [Google Scholar]