Abstract
Copy number variations (CNVs) are universal genetic variations, and their association with disease has been increasingly recognized. We designed high-density microarrays for CNVs, and detected 3000–4000 CNVs (4–6% of the genomic sequence) per population that included CNVs previously missed because of smaller sizes and residing in segmental duplications. The patterns of CNVs across individuals were surprisingly simple at the kilo-base scale, suggesting the applicability of a simple genetic analysis for these genetic loci. We utilized the probabilistic theory to determine integer copy numbers of CNVs and employed a recently developed phasing tool to estimate the population frequencies of integer copy number alleles and CNV–SNP haplotypes. The results showed a tendency toward a lower frequency of CNV alleles and that most of our CNVs were explained only by zero-, one- and two-copy alleles. Using the estimated population frequencies, we found several CNV regions with exceptionally high population differentiation. Investigation of CNV–SNP linkage disequilibrium (LD) for 500–900 bi- and multi-allelic CNVs per population revealed that previous conflicting reports on bi-allelic LD were unexpectedly consistent and explained by an LD increase correlated with deletion-allele frequencies. Typically, the bi-allelic LD was lower than SNP–SNP LD, whereas the multi-allelic LD was somewhat stronger than the bi-allelic LD. After further investigation of tag SNPs for CNVs, we conclude that the customary tagging strategy for disease association studies can be applicable for common deletion CNVs, but direct interrogation is needed for other types of CNVs.
INTRODUCTION
Copy number variations (CNVs), which occupy about 5–12% of the human genome (1,2), greatly influence phenotypic traits and disease susceptibility, such as in HIV infection, autoimmunity and autism (3). A global CNV profile was recently reported (1) in HapMap samples using high-density oligonucleotide microarrays (Affymetrix 500KEA arrays) that we designed. However, the non-uniform probe density compromised the precise detection of copy number changes. Those probes were designed at SNP positions, where the signal intensities differed depending on the SNP genotypes of samples, which led to non-optimal copy number judgment when the signals of samples with different SNP genotypes were compared (4). Also, the probe density was low in chromosomal regions with segmental duplications, where SNP probes were hard to design (1,4). Furthermore, an inability to precisely distinguish the genotypes of CNVs (e.g. the genotype of one and one copies from that of zero and two copies, when two copies were experimentally observed) may have hindered the understanding of the allelic nature of CNVs (5), particularly when CNVs were multi-allelic. These earlier generation platforms and analyses may have contributed to conflicting results explaining the linkage disequilibrium (LD) between CNVs and SNPs, causing some to report that CNVs had strong LD (2,6–8), whereas others reported that they did not (1,9).
Here, we report a global population-genetics study of CNVs, using microarrays that we specially designed to detect CNVs and an algorithm that we developed to precisely analyze CNV alleles (10,11). The Affymetrix custom microarrays (Nsp1.3M arrays) that we designed use non-SNP probes to capture 1.3 million NspI restriction enzyme fragments across the entire genome. The power of these high-density non-SNP-probe arrays was demonstrated in a recent study, although only two individuals were analyzed (12). In our current report, we investigated CNVs from 90 individuals of European descent from Utah, USA (CEU) and 90 Yoruba individuals from Nigeria (YRI) in the HapMap populations (13,14). For these samples, we applied the multiple-reference method (4) to minimize noise and reliably detect smaller CNVs and their boundaries. These more global and precise analyses revealed the bi-allelic and multi-allelic nature of CNVs and further resolved the previous conflicting reports on CNV–SNP LD.
RESULTS
Genomic nature of CNVs
We used the Nsp1.3M arrays to obtain signal intensities for the CEU and YRI samples. To determine CNVs from the data, we used essentially the same methods as before (1,4) (Materials and Methods). Briefly, using the array data for every pair of test and reference samples, we executed GIM (15,16) to reduce noise, correct biases arising from probes and restriction enzyme fragments, and normalize the signal intensities. We then executed SW-ARRAY (17) for all the pair-wise signal-intensity ratios to determine continuous chromosomal segments with CNVs for a single reference sample. We removed unusually long chromosomal segments putatively associated with cell-line artifacts (1) and only examined sub-microscopic CNVs (<3 Mbp) (18,19) on autosomal chromosomes. For each SW-ARRAY segment determined by the pair-wise comparison, we identified multiple reference samples that were thought to have two copies (4). Using these multiple reference samples, we identified chromosomal segments with CNVs. To precisely describe CNV spans on chromosomes, we defined several terms, such as CNV ‘segments’ and ‘regions’; see Figure 1A for these terms.
Figure 1.
Definitions of CNVs and the typical observed pattern. (A) Definitions of CNVs. CNV segments are the chromosomal segments with CNVs for each individual (blue). CNV regions are the union of overlapping CNV segments (red). CNV events are the union of CNV segments that have the same start and end positions (black). CNV fragments are the parts of CNV segments that are divided with the start and end positions of any CNV segments (red circles). CNV fragment-sites are the union of CNV fragments (green). A fragment-segment rate is the proportion of the number of individuals with CNV fragments to the number of individuals with any CNV segments at a CNV fragment-site. (B) The typical segment pattern in a CNV region (chr 18: 45 938 595 to 45 956 033 for CEU). The red and four blue lines indicate a region (17 kb) and segments, respectively. Most CNV regions (89–90%) had a simple segment pattern characterized by two features: no individual with multiple segments and only one ‘core’ fragment-site, which was a fragment-site with a 100% fragment-segment rate.
When we performed quantitative PCR (qPCR) experiments for 90 randomly selected chromosomal locations, which consist of locations both with and without CNV segments, overall 93.3% (84/90) were consistent in the results of CNV presence or absence between microarray and qPCR (Supplementary Material, Table S1). A false-discovery rate and a sensitivity about CNV presence were estimated to be 9.5% (2/21) and 82.6% (19/23). As a computational approach, we performed random permutation of the probe data (Materials and Methods). The estimated false-discovery rate of identified CNV segments was 4.9%. The previous study (1) confirmed 66 CNV segments by qPCR or Mass Spectrometry for multiple references. When we examined the locations of those validated CNV segments, 44 out of the 66 were overlapped with segments detected by our microarrays, which indicates an estimated sensitivity of 67% (44/66). From these rates, it is indicated that the false-discovery rate was about 5–10% and the sensitivity was about 70–80%.
For the two populations together, we found 6184 CNV regions, which covered 224 Mbp (7.9%) of the autosomal genome (Table 1). These were more than the previous 1081 regions (699 for 500KEA; 669 for WGTP) (1), but the genomic coverage was less than the previous 253 Mbp (72 M for 500KEA; 240 M for WGTP). The median CNV region length was 12 700 bp, which was shorter than the previous 162 586 bp (31 367 for 500KEA; 228 858 for WGTP). Table 1 shows the statistics for CEU and YRI separately. The distribution of CNV region lengths was close to the exponential distribution (Supplementary Material, Fig. S1).
Table 1.
Statistics of CNV regions
| CEU and YRI together |
CEU |
YRI |
|||||||
|---|---|---|---|---|---|---|---|---|---|
| Nsp1.3M | 500KEA | WGTP | Nsp1.3M | 500KEA | WGTP | Nsp1.3M | 500KEA | WGTP | |
| Count | 6184 | 699 | 669 | 2986 | 379 | 484 | 4083 | 417 | 469 |
| Genomic coverage (bp) | 224 M | 72 M | 240 M | 123 M | 47 M | 176 M | 156 M | 40 M | 168 M |
| Median length (bp) | 12 700 | 31 367 | 228 858 | 12 241 | 37 270 | 224 588 | 12 700 | 30 990 | 233 836 |
These statistics pertain to CNV regions on the autosomal chromosomes. The statistics of 500KEA and WGTP were calculated from Redon et al. (1).
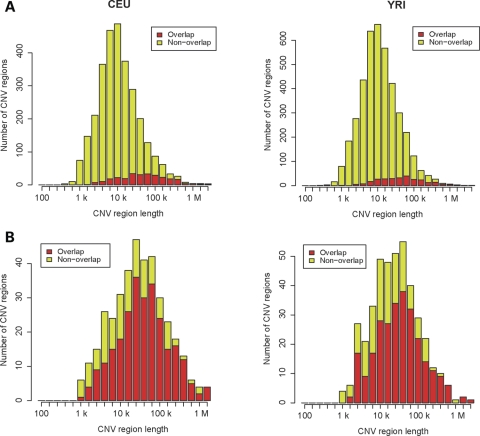
For CEU and YRI together, 1291 (covering 88 Mbp, 3.1% of the genome) of CNV regions we detected were not reported in the Database of Genomic Variants (ver. 5) (20), which extensively collects CNV regions from different sources. Meanwhile, we did not observe 3885 (covering 573 Mbp, 20%) CNV regions that were reported in this database. Compared with the previous 500KEA platform, we identified 2713 and 3809 new regions, which covered 93 and 131 Mbp (3.3 and 4.6% of the genome) for CEU and YRI, respectively. At the shorter lengths, the proportion of the number of newly found regions and that of their bases were larger (Fig. 2A and Supplementary Material, Fig. S2), suggesting that smaller regions tended to be newly detected. The new regions resided in chromosomal positions where 500KEA probes had previously been sparse (on average, 3.6 times higher density in Nsp1.3M), which suggests that the finding of new regions was due to the higher resolution of our current platform. The regions previously detected in 500KEA were mostly found in Nsp1.3M as well (Fig. 2B and Supplementary Material, Fig. S2). Compared with the WGTP arrays, we found 2562 and 3650 new regions, covering 60 and 97 Mbp (2.1 and 3.4%) for CEU and YRI, respectively. Typically, the previous regions were split into smaller regions (1:1.7 on average) in Nsp1.3M.
Figure 2.
Comparison of CNV regions in Nsp1.3M with those in 500KEA. The length of CNV regions detected with one platform versus the number of regions. The number of regions that did and did not overlap with those from the other platform is shown in red and olive, respectively. (A) CNV regions detected with Nsp1.3M. (B) CNV regions previously detected with 500KEA.
The frequency of individuals with any CNV segment per region was generally low (Supplementary Material, Fig. S3). Almost 40% of the regions (39% for CEU; 37% for YRI) had a frequency of ≤2%. When we classified all regions by 1–10, 10–100 and 100–1000 kb lengths, we found that a longer length was associated with a less steep frequency distribution (Supplementary Material, Fig. S3). For a whole-genome view of CNVs, we drew CNV lengths and the number of individuals with CNV segments on the chromosome map (Supplementary Material, Fig. S4). Supplementary Material, Figure S5 shows that the number of CNVs was roughly proportional to chromosome length; however, the chromosome coverage by CNVs was not proportional to chromosome length. In particular, chromosomes 15, 16 and 22 were highly covered by CNVs because of the presence of many large CNVs. Presumably, CNVs occur along chromosomes with similar probability, but the lengths are not equally distributed across chromosomes. The median number of segments and base coverage for one individual over the (autosomal) genome were 184 and 0.3% for CEU, and 244 and 0.4% for YRI, respectively.
The Nsp arrays used non-SNP probes and were expected to capture more CNVs that overlapped with segmental duplications. We found 349 and 454 such CNV regions for CEU and YRI, respectively; of these, 261 and 365 regions were newly found. This finding is due to the increased probe density in the regions of segmental duplications (21) (13–17 times higher in those regions than for the 500 KEA). Although our probes excluded SNPs, the detected CNV regions overlapped with a large number of HapMap SNPs (13,14) (121 994 and 150 516 SNPs for CEU and YRI, respectively), and almost all of the regions (96% for CEU and YRI) overlapped with at least one SNP, which may complicate disease-association studies, as described in Discussion.
Most CNV regions had simple patterns of segments across individuals on chromosomes, as illustrated in Figure 1B. For example, within most regions (96% for CEU and YRI), no individual had multiple segments (Fig. 1B and Supplementary Material, Fig. S6). This feature was observed not only in regions where the number of individuals with any segment was only one, but also in regions where that number was two or more. Moreover, most regions (89–90%) had only one ‘core’ fragment-site with a 100% fragment-segment rate (Fig. 1B and Supplementary Material, Fig. S6), where a fragment-segment rate is the proportion of the number of individuals with CNV fragments to the number of individuals with any CNV segment at a CNV fragment-site (Fig. 1A). This feature was observed also in regions where the number of individuals with any segment was two or more. Most regions (90% for CEU; 89% for YRI) had both of these features. These simple segment patterns would simplify CNV analyses because it would be difficult to determine which chromosomal location should be compared across individuals within a region, if a region did not have these features.
Allelic nature of CNVs
We used supervised signal-intensity data on one to five copies for the signal intensities of CNV segments and determined the integer copy numbers of the segments (Materials and Methods and Supplementary Material, Table S2). Because these copy numbers represent the total number of copies over two homologous chromosomes, we term these numbers diploid copy numbers and denote them with the prefix ‘D’ (e.g. D2 for the two copies). For each CNV region with the two simple segment features (described above), we used the computational tool MOCSphaser (10) and observed diploid copy numbers at the core site to estimate the population frequencies of allelic copy numbers, which represent the number of copies on one homologous chromosome, denoted with the prefix ‘A’ (e.g. A1 for the one copy).
We found that the frequency of allelic copy numbers other than A1 was quite low for most CNV regions (Supplementary Material, Fig. S7). When we classified CNVs by the allele frequency into rare (the frequency <1%), relatively common (1–5%) and common (≥5%), most CNVs were rare, and only a small percentage were common (Table 2). YRI had a lower percentage of rare CNVs than CEU. We plotted an allele frequency spectrum (Fig. 3A), which is the histogram of all alleles classified by their population frequency. This spectrum also showed that non-one-copy alleles tended towards small frequencies and further showed that the most frequent non-one-copy alleles were A0 and A2. The longer the region, the more gradual was the slope of the distribution (Supplementary Material, Fig. S8). The proportion of the number of the deletion allele (A0) to the duplication alleles (A2 or more) was 1.3 for CEU and 2.2 for YRI. This proportion decreased with increasing region length, which may be related to natural selection: larger deletions are deleterious, whereas duplications are generally permissible. Regarding the number of alleles, if we suppose that alleles with a frequency of less than 0.1% practically do not exist in the population, most (96–98%) of the CNV regions with two or more alleles were bi-allelic CNVs, and only a small percentage of them had three or more alleles. Most bi-allelic CNVs were composed of A0 and A1, or A1 and A2; most tri-allelic CNVs were composed of A0, A1 and A2. With respect to diploid copy number, the frequency spectrum (Fig. 3B) showed a larger number of D1 than D0 among the loss-type (<D2) diploid copy numbers and dominance of D3 among the gain-type (>D2) diploid copy numbers. When we classified CNV regions into loss CNV regions, which is composed only of the loss-type and standard-type (D2) segments, gain CNV regions, which is composed only of the gain- and standard-type segments, and mixture CNV regions, which is composed of all the loss-, gain- and standard-type segments, the proportion of the three was 57%:40%:3% (1073:754:59) for CEU and 69%:29%:2% (1773:751:46) for YRI.
Table 2.
Number of common, relatively common and rare CNV regions
| CEU | YRI | |
|---|---|---|
| Common | 133 (5.7%) | 187 (6.0%) |
| Relatively common | 427 (18.4%) | 729 (23.5%) |
| Rare | 1,760 (75.9%) | 2,185 (70.5%) |
The common, relatively common and rare CNV regions were CNV regions for which one minus the frequency of A1 was ≥5%, 1–5% and <1%, respectively.
Figure 3.
Frequency spectrums. These counts are based on allelic copy numbers and the derived diploid copy numbers that are classified by their population frequency. The width of each bin is 2%. Alleles with a very small or large frequency of <0.1% or >99.9% are excluded from the counts. (A) The allele frequency spectrum. (B) The frequency spectrum of diploid copy numbers.
In terms of genotypes, the frequency of genotypes other than the standard genotype A1/A1 was quite low (Supplementary Material, Fig. S7), although the frequency distribution was not quite as steep as in the allele case. Across CNV regions, the most frequent genotypes other than the standard genotype were A1/A0, A1/A2 and A0/A0 in order of decreasing frequency. Within CNV regions, the most standard diploid copy number, D2, can theoretically take two forms: A1/A1 or A2/A0. Our results, however, showed that D2 practically took only one form, A1/A1 (i.e. the proportion of the A1/A1 frequency to the A1/A1 plus A2/A0 frequency ≥99.9% for a region), in most CNV regions (99% of all regions for both CEU and YRI).
We assessed the heritability of CNVs within trio families, checking the consistency between the diploid copy number of a child and the copy-number genotypes of the parents (Supplementary Material, Table S3). The Mendelian discordance rates (Materials and Methods) were 1.04 and 1.18% for CEU and YRI, respectively (Supplementary Material, Table S4). Note that Mendelian discordance arises from two possibilities: observation errors of copy numbers of a child or parents, and copy number change that occurs in a child's chromosomes when the chromosomes are transmitted from parents to a child. We cannot readily distinguish these possibilities. Although these rates were based on CNV detection using multiple reference samples, we also calculated rates using the conventional single reference method. The discordance rates were 1.85 and 1.77% for CEU and YRI, respectively (Supplementary Material, Table S4). The multiple reference method showed lower discordance rates, implying that this method more precisely detected CNVs relative to the single reference method.
We compared CNV allele frequencies between CEU and YRI. We found that the chromosomal locations of many CNV regions (75% for CEU; 83% for YRI) did not overlap between the two populations. This finding suggests that the large majority were monomorphic (i.e. not CNVs) in either population, which is consistent with our observed tendency toward the rareness of non-one-copy alleles. For CNV regions that overlapped one-to-one with another between both populations (371 regions), we plotted dots representing the allele frequencies in CEU and YRI (Supplementary Material, Fig. S9). The dots were broadly spread around the y = x line, indicating discordance of the allele frequencies between the populations. For each one-to-one overlapping CNV, we calculated Fst (Supplementary Material, Methods), which represents population differentiation that implies recent population-specific positive selection. We found several CNV regions with high population differentiation, including six regions that were not reported in the previous study (1) (Supplementary Material, Fig. S10, and Table 3). For example, one such region overlapped with CLDN5, which encodes an integral membrane protein involved in epithelial or endothelial cell sheets. This result implies the population-specific selection of this CNV/gene between CEU and YRI.
Table 3.
CNV regions with high population differentiation (Fst > 0.1)
| Chr | Start | End | Fst | Overlapping gene |
|---|---|---|---|---|
| 1 | 149365093 | 149419009 | 0.427 | LCE3A/3B/3C/3D/3E, late cornified envelope |
| 2 | 3701609 | 3727783 | 0.121 | None |
| 2 | 34576366 | 34662239 | 0.469 | None |
| 4 | 10063092 | 10086289 | 0.106 | ZNF518B, zinc finger protein 518B |
| 4 | 34595900 | 34663168 | 0.485 | None |
| 4 | 187464847 | 187498012 | 0.373 | CYP4V2, cytochrome P450 |
| 6 | 32060463 | 32136004 | 0.209 | CYP21A2, cytochrome P450 |
| 8 | 120216553 | 120271645 | 0.106 | Collectin sub-family member 10 |
| 14 | 81562743 | 81591452 | 0.588 | None |
| 15 | 25588301 | 25606455 | 0.116 | None |
| 22 | 17921878 | 18002715 | 0.618 | CLDN5, claudin 5 transcript variant 2 |
Fst is a commonly used statistic to estimate population differentiation, ranging from 0 (undifferentiated) to 1 (population-specific). The start and end positions in the table indicate the boundaries of the union of the CEU and YRI CNV regions. Bold letters indicate that those CNV regions were not reported for either population in the previous study (1).
We examined disease-related genes that overlapped with CNVs. A total of 165 and 226 OMIM (Online Mendelian Inheritance in Man) genes overlapped with CNV regions in CEU and YRI, respectively (Supplementary Material, Table S5). Of these, 114 and 164 genes overlapped with regions not reported in the previous study (1). These included FRAS1 (susceptible for Fraser Syndrome 1) and UBE3A (Angelman Syndrome), which overlapped with non-rare deletion CNV regions in both CEU and YRI. CYP4V2 (related to Bietti crystalline corneoretinal dystrophy), FOXN1 (T-cell immunodeficiency) and TJP2 (Hypercholanemia) overlapped with multi-allelic (tri-allelic or more) CNVs in either CEU or YRI. Dozens of OMIM genes (18 for CEU and 27 for YRI) overlapped with multiple CNV regions that were all newly detected in this study. For example, CTNND2 (Mental retardation in cri-du-chat syndrome) overlapped with three rare and one relatively common CNV regions in CEU and with two rare CNV regions in YRI. We confirmed that CCL3L1 (22), FCGR3B (23), BTNL2 (24), AMY1 (25) and CYP2D6 (26), for which CNVs are suggested to be associated with complex diseases and human phenotypic variation, overlapped with our CNV regions.
For common, relatively common and rare CNV regions, we looked into the association of these regions and the flanking regions (up to 10 000 bp from the CNV boundaries) with sequence features: segmental duplications, genes and repetitive elements (Table 4). Segmental duplications were the most associated with and enriched in those CNV regions. The association was found for not only the CNV regions but the flanking regions, though the latter association was weaker. The association clearly increased according to the increase of the population frequencies of CNVs. These results indicate either that segmental duplications are involved in the recurrent generation of CNVs, or that some segmental duplications are not fixed in the population (1). Genic regions were significantly deficient in CNV regions, which is consistent with the hypothesis that CNVs present in genic regions are deleterious and removed by purifying selection (5). Repetitive elements were enriched in CNV regions but the association was not so strong, which suggests that repetitive elements do not greatly contribute to the generation of CNVs. We further investigated repetitive element sub-classes (SINE Alu/MIR, LINE L1/L2 and LTR ERV1/MaLR), for which the associations were also not strong, although LINE L1 had a somewhat strong association (Supplementary Material, Table S6).
Table 4.
Association of CNV and flanking regions with sequence features
| CNV region or flanking region | Sequence feature | Odds ratio, CEU | Odds ratio, YRI |
|---|---|---|---|
| Common | Segmental duplication | 6.93 | 4.56 |
| Relatively common | 3.88 | 2.80 | |
| Rare | 1.92 | 2.00 | |
| Flanking around common | 5.23 | 3.51 | |
| Flanking around relatively common | 2.83 | 2.03 | |
| Flanking around rare | 1.41 | 1.47 | |
| Common | Gene | 2.15a | 1.83a |
| Relatively common | 1.34a | 1.14a | |
| Rare | 1.17a | 1.24a | |
| Flanking around common | 2.00a | 1.79a | |
| Flanking around relatively common | 1.38a | 1.23a | |
| Flanking around rare | 1.25a | 1.25a | |
| Common | Repetitive element | 1.45 | 1.57 |
| Relatively common | 1.30 | 1.29 | |
| Rare | 1.27 | 1.24 | |
| Flanking around common | 1.39 | 1.42 | |
| Flanking around relatively common | 1.27 | 1.25 | |
| Flanking around rare | 1.22 | 1.21 |
Flanking regions are regions up to 10 000 bp from the boundaries of common, relatively common or rare CNV regions. For an odds ratio, we first calculated the summed number of bases that overlapped with CNV regions and regions of a sequence feature, that of bases that overlapped with only CNV regions, that of bases that overlapped with only sequence feature regions, and that of bases that did not overlap with either of them. We used these four summed numbers to calculate an odds ratio. The superscript ‘a’ represents the reciprocal number of a calculated odds ratio with the value of below one so that this number can be easily comparable with other odds ratios. All the odds ratios were significant (P < 10−6) in the Fisher's exact test.
LD nature of CNVs
We examined pair-wise LD between every CNV site (region) and the neighboring SNP sites, classifying CNVs into bi-allelic CNVs and tri-allelic CNVs (determined as described above). For bi-allelic CNVs (mostly A1 and A0, or A1 and A2), we found that the LD index R2 between CNV and SNP sites was almost zero on average all along the distance between the two sites (i.e. the distance from either boundary of a CNV region to a SNP position) (Fig. 4A). This CNV–SNP LD was clearly lower than the SNP–SNP LD. We plotted this SNP–SNP LD, adjusting the minor allele frequency of one half of a SNP pair to the frequency of a CNV allele (see Materials and Methods); hence, this difference would not result from the frequencies of CNV alleles, but would reflect the relationship between the CNV and SNP sites. This difference was also confirmed with LD between CNV and SNP sites with similar allele frequencies (Supplementary Material, Fig. S11). Despite the almost-zero LD tendency, not all CNV–SNP pairs had low LD; 233 and 115 CNV–SNP pairs (39 and 42 unique CNVs) had R2 values of ≥0.8 among all 79 159 and 143 463 pairs (503 and 875 unique CNVs) within an 80-kb distance for CEU and YRI, respectively. For another LD index, D′ [and multi-allelic D′ (27)], the estimated values were unreliably inflated because of small allele frequencies (28); we did not use these values.
Figure 4.
CNV–SNP LD. LD versus distance for (A) bi-allelic CNVs and (B) two-way tri-allelic CNVs. The numbers of bi-allelic CNVs and tri-allelic CNVs were 503 and 875 and 32 and 22 for CEU and YRI, respectively. The distance between a CNV and a SNP was measured from either boundary of a CNV region to a SNP position. The distances were binned in a 10-kb width, and the median of (≥10) LD values was plotted against the middle distance of the bin. ‘SNP (permutated)’ indicates that the SNP genotype data were permutated across individuals, and the error bars indicate the standard deviation. ‘SNP (adjusted)’ indicates that the minor allele frequencies of one half of the SNP pairs in SNP–SNP LD were adjusted to those of CNVs. The larger and relatively larger frequencies indicate ≥10% and 1–10% frequencies of the deletion/duplication alleles, respectively.
We further examined LD by classifying CNVs according to several features as indicated in Figure 4A and Supplementary Material, Figure S11. Interestingly, CNVs with a larger frequency of the deletion allele (A0) showed remarkably higher LD, but the other CNVs, including those with a larger frequency of the duplication allele (any of A2, A3, A4, >A4), had almost zero LD (Fig. 4A). The LD related to the larger deletion frequency was lower in YRI than in CEU. Irrespective of CNV length, LD was always close to zero, whether CNVs were common between CEU and YRI or whether CNVs overlapped with genes, segmental duplications or repetitive elements (Supplementary Material, Fig. S11). We confirmed the same tendencies by multiple regression analysis (29) using these features (data not shown).
In terms of tri-allelic CNVs, we used the LD index V2 for ‘two-way’ (deletion and duplication) tri-allelic CNVs, which were composed of the deletion allele (A0), the standard allele (A1) and the duplication allele (any of A2, A3, A4, >A4). In contrast to bi-allelic CNVs, the LD values were somewhat higher than zero, up to ∼20 kb, and showed decreasing dependence on the distance (Fig. 4B). This non-zero tendency was statistically stronger than that of bi-allelic CNVs (the Z-scores measured with the permutated data were larger in the tri-allelic CNVs, 1.25 and 0.62 on average over ≤20-kb distances, than that in the bi-allelic CNVs, 0.88 and 0.39 for CEU and YRI, respectively). The LD values were lower in YRI than in CEU. To examine the contribution of each allele to the LD, we decomposed V2 into components (w2, see Supplementary Material Methods) for each of the deletion, standard and duplication alleles. The proportion of the decomposed LD (w2) to the total LD (V2) was only high for the deletion and duplication alleles (on average, 43.5, 8.7 and 47.8% for the deletion, standard and duplication alleles, respectively), and only the pair of the deletion and duplication alleles had a strong (negative) correlation coefficient (−0.89) between the decomposed LD values. This result suggests that in a two-way tri-allelic CNV, most of the association measured by V2 is explained by the association between SNP alleles and either of the deletion or duplication alleles, and that the other two CNV alleles do not have an association with the SNP alleles.
Tag SNPs for CNVs
We searched for SNPs that had high LD with CNVs. Such SNPs (tag SNPs for CNVs), when genotyped, would serve to predict CNV alleles and surrogate CNVs in investigation of the associations between CNVs and disease. This methodology reduces costs because SNPs are currently much easier to genotype than CNVs. For bi-allelic CNVs, the definition of such tag SNPs is relatively straightforward and has been used in several studies (1,2,7–9), but it is challenging for multi-allelic CNVs. Here, we have considered whether CNV alleles can be predicted from SNP genotypes, and have defined a tag SNP for each allele of a CNV. To measure the prediction ability, we used the LD index R2 and the conditional probability given genotyped SNP alleles (Materials and Methods). We defined a CNV as tagged when both alleles were tagged in bi-allelic CNVs. In multi-allelic CNVs, we defined a CNV as all tagged when all alleles were tagged and defined as partly tagged when at least one or two alleles were tagged with R2 or the conditional probability, respectively (these different counts are consistent because when one allele is tagged with R2, there is always another allele or allele group that can be tagged).
With respect to bi-allelic CNVs, we found only a small number of CNVs tagged by SNPs when using 0.8 or even 0.6 as a cutoff value for R2 and the conditional probability (Fig. 5A). This small number is consistent with the weak LD tendency in the above-described results. Common CNVs were more frequently tagged than relatively common CNVs. There was no clear difference in the number of tagged CNVs between the two tagging methods, but the conditional probability method tagged a greater proportion of common CNVs. At the 0.8 cutoff, the number of tagged CNVs was almost saturated at a distance of 80–100 kb to SNPs (Supplementary Material, Fig. S12). Tagged alleles were primarily A0 among non-A1 alleles (Supplementary Material, Fig. S12). Multi-allelic (two-way tri-allelic) CNVs were also rarely tagged, although the number of multi-allelic CNVs was originally small (Fig. 5B). The method using conditional probability tagged more CNVs than the method using R2. The number of partly or all tagged CNVs at the 0.8 cutoff was not more than 10 or 5, respectively. Tagged alleles were A0 and A2 among non-A1 alleles (Supplementary Material, Fig. S12).
Figure 5.
Number of CNVs tagged by SNPs. Number of tagged CNVs versus the cutoff association values (R2 and conditional probability). We searched for tag SNPs up to 200 kb from the boundaries of each CNV region. ‘c.p.’ indicates conditional probability. (A) For bi-allelic CNVs. (B) For two-way tri-allelic CNVs.
DISCUSSION
We classified CNVs by integer values, which is a more detailed classification than the customary ‘gain’ and ‘loss’ (1,30). Recent studies (2,31) used SNP arrays (SNP6.0) and determined copy numbers as single integer values. Meanwhile, we used Nsp arrays and supervised data on different numbers of DNA copies to determine copy numbers in more extended representations such as ‘3 or 4’ copies or ‘>4’ copies (10) [ambiguity in the ‘or’ representation was resolved by use of the phasing tool (10) in a subsequent procedure]. This is a practical way to handle copy numbers as integer values on array platforms because the signal intensities of different copies are usually not clearly separated from each other. In the future, the probability distribution of copy numbers [e.g. via the emission probability in Birdsuite (31)] will be useful for a more detailed representation (11). When we compared our copy-number frequencies with copy-number frequencies in a study (2) reporting integer copy numbers for each of the overlapping CNV regions, the frequencies were fairly close to each other [60% of the CNV regions had lower than 0.05 in the total variation distance between population frequencies (11,32)]. CNV regions with a bad concordance tended to have lower CNV frequencies in our study than in their study, indicating that we might miss CNVs within some samples or that they might excessively detect CNVs.
The multiple-reference method assumes that the largest group with the same copy number is the two-copy group (4). Although we made adjustments for the case where this assumption does not hold, some exceptional CNVs may escape such an adjustment. In this case, the largest group with non-two copies is regarded as the two-copy or the reference group, so that the multiple-reference method might mistake non-CNV segments for CNV segments within a CNV region. This would result in the confusion of a high-frequency CNV region with a low-frequency CNV region. However, it is difficult to know how many high-frequency CNV regions are mistakenly regarded as low-frequency CNV regions. In fact, it is considered difficult to precisely detect high-frequency CNV regions as long as a reference sample (or samples) is used (33), and their fraction to all CNV regions remains unknown. However, taking into consideration that almost half of CNV regions are found in only one sample in other studies (1,2), the proportion of high-frequency CNVs is expected to be relatively small, and the influence on our results would be relatively minor. Better detection of high-frequency CNVs would require some absolute measurement without using a reference sample or samples.
Using a phasing tool (10), we obtained the population frequencies of allelic copy numbers and their genotypes. Note that this phasing assumes Hardy–Weinberg equilibrium; although nothing is known about the effect of deviation from this equilibrium, it is known that the deviation does not significantly affect SNP haplotype phasing based on the same basic algorithm (11,34). We presented the allele frequency spectrum of CNVs, which is a basic graph that summarizes their population-genetic nature. Among non-one-copy alleles, the only alleles with a large frequency were the zero- and two-copy alleles. This finding indicates that the allele types of CNV regions we detected were mostly simple. When examining the segment patterns, most CNV regions were also simple; they had only one segment for each individual and only one ‘core’ site across individuals. This was surprising because we did not perform the strong operation of aligning segments across individuals, as was done in recent studies (2,31). Our result justifies such operation and encourages the use of simple methods for analysis of the allelic nature of CNVs.
The comparison of the whole genome or chromosomal distributions of CNVs between CEU and YRI populations did not show clear differences between the populations. Furthermore, the allele frequencies of CNVs in both populations were mostly very low. One interpretation is that CNVs are relatively deleterious (compared with SNPs) and that natural selection often removes them from a population, following which little genetic variation in the form of CNVs may remain to characterize population differences. Nevertheless, we found some exceptional CNVs with high Fst, which showed high population differentiations between CEU and YRI. Some of these CNVs were located in cytochrome P450, in which population differences were previously reported (35). Another CNV was located in LCE, a keratin protein of skin. These results support the previous findings of human population differences (36). In addition, we found several other CNVs with high Fst, which would be interesting to closely investigate as to what roles those regions played in human evolution. In particular, CNVs that did not overlap genic regions but had high Fst (Table 3) would likely have roles as regulatory elements or non-coding RNAs.
We calculated CNV–SNP LD for a large number (1432) of CNVs, which are almost 20 times as many as those in the previous study (1). This large-scale investigation provided us with a comprehensive view of CNV–SNP LD for bi-allelic and multi-allelic CNVs. The LD for bi-allelic CNVs tended to be extremely low, even compared with the previous study (1). This low LD might be associated with high mutation rates [10−4–10−7 mutations per locus per generation for CNVs versus 10−7–10−8 for SNPs (1)] or recurrent mutations in CNVs (5,37). In contrast, common deletion CNVs tended to have high LD, although other types of CNVs did not, including common duplication CNVs. This high LD might be related to LD inflation by negative selection, as indicated for SNP–SNP LD (38,39). In previous reports, there was conflicting evidence that LD was high enough to be well tagged by SNPs in some reports (6–8), but not in others (1,9). Our results suggest that these reports are consistent because the former studies reported LD for common deletion CNVs. A recent large-scale report (2) suggested a high degree of LD, and this was also explained by the high LD for common deletion CNVs because the reported CNV set was primarily composed of such CNVs. Unlike the analysis of multi-allelic CNVs based on continuous signal intensities (1), we addressed multi-allelic CNVs using discrete alleles and examined LD between the CNVs and SNPs. This canonical method revealed that this LD had a different nature from that of bi-allelic CNVs; for example, the LD tended to be somewhat stronger.
As is often the case with CNV studies (1), the CNV set in that recent report (2) was not well concordant with our set: 87% of our CNVs were not in their set, and 59% of theirs were not in ours. When we also looked into the concordance of our CNV set with the CNV set in another large-scale study (40) using tiling microarrays, the consistency was also not good: 76% of our CNVs were not in their set and 88% vice versa. These differences would presumably result from differences in the platforms (e.g. SNP arrays versus Nsp arrays) and the detection procedures (e.g. using across-sample information versus not using it, possibly influencing the commonness of detected CNVs across samples).
Our research has several implications for disease-association studies of CNVs. (i) Because most of the CNVs that we detected had simple segment patterns across individuals at the kilo-base level, the part of the CNV region that is compared between case and control individuals is not a concern, as long as the CNVs are examined in such arrays at this scale. (ii) Because common deletion CNVs were in high LD with SNPs, conventional association studies based on LD and the common disease-common variant hypothesis would be applicable for this type of CNV. However, considering that the number of CNVs classified in other types was much larger, direct interrogation not using LD information would be needed for a comprehensive search. In particular, the presence of many rare CNVs would underline the necessity of finding disease-causal alleles among rare alleles. In addition, CNVs would have to be examined separately from population to population. (iii) On the basis of the observation that different methodologies continue to identify a non-mutually overlapping set of CNVs, one could conclude that the current methodologies are complementary to each other and utilization of multiple methods in order to maximize the CNV set under investigation could be considered. (iv) Finally, although our platform did not use SNP probes, most CNVs overlapped with HapMap SNPs, many of which would be SNVCs [single nucleotide variation on copy units (11)]. In this case, as demonstrated in a previous study (11), association analyses based on differences in integer copy numbers are insufficient, and finer analyses based on differences in the bases/lengths of copy units, which are the units of DNA sequence that are duplicated within a CNV region (11), might be necessary for detecting differences in genetic variation between two population groups. Considering the possible universality of SNVCs, those finer analyses would be needed in a full search for disease-causal alleles.
MATERIALS AND METHODS
Samples
DNA samples from cell lines derived from 90 CEU (30 trios) and 90 YRI (30 trios) individuals in the HapMap project (13,14), as well as samples with different numbers of X chromosomes, were purchased from Coriell Cell Repositories (http://ccr.coriell.org/).
Microarrays
Probes on the chip were designed using NspI fragment sequences with lengths between 200 and 1000 bp in the human genome build (UCSC version hg17, NCBI version v35). For each fragment, a probe set of 10 perfect-match 25-mer oligonucleotide probes was designed. The probes avoided repetitive elements listed in the REPBASE database (http://www.girinst.org/repbase/index.html) and avoided SNPs present in dbSNP (release 123), the latter of which should minimize a bias due to polymorphisms in restriction enzyme sites. In total, the designed probe sets represented 1 330 354 fragments with an average and median spacing of 2271 and 776 bp, respectively. About 90% of the segmental duplications had at least one of these fragments within their boundaries. The details of the array design and its quality control are described elsewhere (12). The experimental protocol and the quality control were the same as that described for the Affymetrix 500K arrays. About 90 μg of the target DNA was hybridized to the arrays overnight (http://www.affymetrix.com/products/arrays/specific/500k.affx). The signal-intensity ratios of test samples to reference samples for probe sets (fragments) were obtained with GIM (15,16).
Signal intensities of 1X to 5X
To obtain supervised data on signal-intensity ratios for different copy numbers, as in the previous study (4), we measured the signal intensities of probes for samples with different numbers of X chromosomes in triplicate and used GIM to obtain the signal-intensity ratios of nX (representing each of the X chromosomes, where n = 1, 2, … , 5) to 2X. We excluded the ratios of probe sets located in segmental duplications or known CNV regions because n copies of X might not be guaranteed in these regions. We observed that the histogram of the log of the ratios (to the base 2) for each nX was approximately close to the normal distribution with a mean and variance of the log ratios. Hence, we used the normal distribution to analyze the log ratios for each nX.
CNV detection by SW-ARRAY
We used SW-ARRAY (17) to determine CNV segments for each sample pair. Considering the previous tool settings (4), we used signal-intensity ratios of 1.05 and 0.91 as the upper and lower thresholds, respectively. These values correspond to the false-negative rate 2.5% (97.5% of the area of the 1X and 3X distributions) for detection of loss and gain CNVs. We required that CNV segments should have at least four successive probe-sets that were called (i.e. ≥4 called restriction enzyme fragments). We adopted 0.05 as the P-value (false-positive rate) cutoff for determining continuous CNV segments. We excluded chromosomal regions with karyotypic abnormalities identified in the previous study (1) for particular samples; otherwise, their inclusion would result in very long CNV segments. We did not use sex chromosomes in our analyses because many segments of these chromosomes were very long and difficult to interpret.
CNV determination using multiple references
From the SW-ARRAY results of all pair-wise samples, we used the previous algorithm (4) to determine CNV segments on the basis of multiple reference samples. In brief, the algorithm first identifies the largest group with the same copy number; namely, it identifies reference samples for which the frequency of test samples with segments identified by SW-ARRAY is the smallest. Generally speaking, this group is thought to contain samples with two copies (4). Then, the algorithm incorporates multiple-reference information to determine CNV segments for each sample. See the reference (4) and Supplementary Material Methods for the detailed procedure. Finally, we visually inspected CNV regions identified by the automatic procedures to exclude aberrant CNV segments (in a CNV region, only one conspicuously long segment with almost the same signal intensity as that of two copies). We also excluded CNV regions that were composed only of CNV segments that had almost the same signal intensity (the representative signal ratio, below) as that of two copies.
PCR validation of CNVs
CNV validation was done by digital PCR using a Fluidigm Digital Array and the BioMark System (Fluidigm CA, USA). FAM-labeled TaqMan PCR assays for CNV regions (rg4537-P:CCATCCTCGCAGCTC, rg4537-F:GGCCCCACTGAGTGTTTGAT, rg4537-R:CCGCCAACTCTGGTCCTCTA, rg5228-P:CTCTAGATTCTCAGGAGAGAT, rg5228-F:TGCCTGTAGCCAACTGATCCT, rg5228-R:ACCAAAGAGAGAGCCAAGTCAGA) were ordered from Applied Biosystems (CA, USA) with VIC labeled RNase P gene assay control (product #4316844). The assay methods are essentially as indicated in a study (41). Briefly, 4 µl reaction mixes were prepared for each assay, containing 1 × TaqMan gene expression master mix, 1 × RNase P-VIC TaqMan assay, 1 × TaqMan assay for the target CNV, 1 × sample loading reagent (Fluidigm CA, USA) and genomic DNA with 10 ng/μl concentration. The reaction mix was uniformly partitioned into the 770 reaction chambers of each panel and the digital array was thermocycled on the BioMark System. Molecules of the two genes were independently amplified, and FAM and VIC signals of all chambers were recorded at the end of each PCR cycle. The numbers of both FAM-positive chambers (target CNV) and VIC-positive chambers (RNase P) in each panel was counted and copy number of target CNV was calculated as in a study (42).
Random permutation to estimate the false-discovery rate
To estimate the false discovery rate of identified CNV segments, we randomly permutated the chromosomal positions of microarray probe sets with their signal intensities. We then performed the same procedures as in the normal (non-permutated) data to determine CNV segments based on multiple reference samples. The false-discovery rate was calculated as the number of CNV segments obtained from the permutated data divided by the number of CNV segments obtained from the normal data.
Estimation of diploid copy numbers
Considering the previous study (4), we used both the distribution of signal ratios (in log) for nX and the representative signal ratio (in log) of a CNV segment to determine the diploid copy number of the segment. The representative signal ratio was the median of the signal ratios across the reference samples over the probe sets within a segment. When the Z-score of the log representative ratio of a segment was within ±1.96 under the nX normal distribution (meaning <95% of the area of the distribution), we determined the diploid copy number to be n. This corresponds to a theoretical false-negative rate of 5%. If the log ratio was within the threshold for the multiple nX distributions, we treated the diploid copy number as any of the multiple ns and denoted it by concatenating all candidate copy numbers by ‘or’, such as ‘D2 or D3’. We observed that high copy numbers were difficult to discern in the case of >4 copies from the nX distributions. Hence, when the log representative ratio was the (lower) threshold or more for the 5X distribution, we treated those high copy numbers together and denoted them as ‘>D4’. Zero copies were determined as the log ratio with smaller than the lower threshold for the 1X distribution. See Supplementary Material Methods for the specific ratio ranges to determine diploid copy numbers.
Population frequencies of CNV alleles and CNV–SNP haplotypes
In each population, we used diploid copy numbers and SNP genotypes as the input for a phasing tool, MOCSphaser (10), to infer the population frequencies of allelic copy numbers and of two-site haplotypes composed of allelic copy numbers and SNP alleles. The SNP genotypes were downloaded from the HapMap project (13,14). Since the tool uses unrelated samples to estimate the population frequencies, we used parents in the trios for the input. The tool also output the frequencies of genotypes/diplotypes. The population frequencies of diploid copy numbers were calculated from those of the genotypes.
Mendelian discordance
We assessed the heritability of CNVs within trio families. For each family, we checked the consistency between the diploid copy number of a child and the genotypes of the copy numbers of the parents in phased CNV regions. For the parents’ genotypes, we used the most probable genotype determined by the phasing tool (10). If it was impossible to generate the child's diploid copy number from any combination of the parents’ alleles, we defined this state as Mendelian discordance. For example, when the genotypes of a father and a mother were both A1/A1, D2 would be the only consistent diploid copy number of the child. Therefore, if the child did not have D2, it was considered discordant. We defined the discordance rate as the total number of discordants of all the families across all the regions divided by the number of all the families times the number of all the regions.
Linkage disequilibrium
Using the haplotype frequencies estimated by MOCSphaser (10), we calculated the two-locus LD of R2 for bi-allelic data and that of the square of Cramer's V (43,44) for multi-allelic data. The mathematical definitions are given in Supplementary Material, Methods. By definition, V2 is a natural extension of R2. These LD indices range from 0 to 1. In the LD calculation, we excluded CNV alleles for which the allele frequencies were less than 0.1%; we then determined the number of CNV alleles and standardized the allele frequencies. We calculated LD only when all allele frequencies at both loci were 1% or more. In the comparison of CNV–SNP LD with SNP–SNP LD, we adjusted allele frequencies in SNP–SNP LD to exclude the influence of allele frequencies on the LD values. Specifically, we first grouped SNPs and bi-allelic CNVs by the minor allele frequency of 1% on a chromosome; for each frequency group, we next randomly selected SNPs from the SNPs up to the same number as the CNVs. Then, we calculated the LD for every selected SNP against other SNPs. Using these procedures, we adjusted the minor allele frequencies (and the number) of both the selected and the other SNPs in SNP–SNP LD to those of both CNVs and SNPs in CNV–SNP LD.
Tag SNPs for CNVs
We used two methods to select SNPs that were statistically related to CNV alleles. In the first method, for each CNV locus we selected neighboring SNPs up to 200 kb from the boundaries of the CNV region and calculated the LD of R2 for a haplotype frequency table that contained the two rows of a CNV allele (or multiple alleles) and the other remaining CNV alleles versus the two columns of a SNP allele and the other SNP allele. When either row was a row for multiple CNV alleles, we summed the haplotype frequencies over the multiple alleles to construct a 2 × 2 table. We calculated LD only when all marginal frequencies in a table were 1% or more. If we found a SNP with a high R2 value (e.g. >0.8) for a CNV allele (alleles), we considered the CNV allele (alleles) to be associated with that SNP. From such SNPs, we selected the SNP with the largest R2 value as the tag SNP for the CNV allele (alleles).
The first method depends on the familiar LD index R2, but R2 arranges both an allele of interest and the other remaining allele (or alleles) together in the 2 × 2 table. To treat each allele independently, particularly for multi-allelic CNVs, we used the conditional probability of a CNV allele (or alleles) given a SNP allele. The equation for the conditional probability is given in Supplementary Material Methods. We calculated the conditional probability only when the frequency of a CNV allele (alleles) was 1% or more and the SNP allele frequency was 5% or more. If we found a SNP allele for which the conditional probability of a CNV allele (alleles) was high (e.g. >0.8), we considered that we could predict the CNV allele (alleles) from the SNP allele. From SNPs with such alleles, we selected a SNP with the largest probability value as the tag SNP for the CNV allele (alleles).
SUPPLEMENTARY MATERIAL
FUNDING
This work was supported by Japan Society for the Promotion of Science (JSPS.KAKENHI 20790269 to M.K.); the National Cancer Institute (Award Number P30CA045508 to M.K.); Ministry of Education, Culture, Sports, Science and Technology (Applied Genomics 20018005 to S.I.); New Energy and Industrial Technology Development Organization (NEDO) of Japan (Industrial Technology Research Grant Program to S.I.). Funding to pay the open access charge was provided by RIKEN.
Supplementary Material
ACKNOWLEDGEMENTS
We thank Takashi Morizono for coding computer programs and formatting the figures and Tetsuo Abe for coding programs.
Conflict of Interest statement. M.S. and K.J. declare competing financial interests. They are employees of Affymetrix, Inc.
REFERENCES
- 1.Redon R., Ishikawa S., Fitch K.R., Feuk L., Perry G.H., Andrews T.D., Fiegler H., Shapero M.H., Carson A.R., Chen W., et al. Global variation in copy number in the human genome. Nature. 2006;444:444–454. doi: 10.1038/nature05329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.McCarroll S.A., Kuruvilla F.G., Korn J.M., Cawley S., Nemesh J., Wysoker A., Shapero M.H., de Bakker P.I., Maller J.B., Kirby A., et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 2008;40:1166–1174. doi: 10.1038/ng.238. [DOI] [PubMed] [Google Scholar]
- 3.Sebat J. Major changes in our DNA lead to major changes in our thinking. Nat. Genet. 2007;39:S3–S5. doi: 10.1038/ng2095. [DOI] [PubMed] [Google Scholar]
- 4.Komura D., Shen F., Ishikawa S., Fitch K.R., Chen W., Zhang J., Liu G., Ihara S., Nakamura H., Hurles M.E., et al. Genome-wide detection of human copy number variations using high-density DNA oligonucleotide arrays. Genome Res. 2006;16:1575–1584. doi: 10.1101/gr.5629106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Conrad D.F., Hurles M.E. The population genetics of structural variation. Nat. Genet. 2007;39:S30–S36. doi: 10.1038/ng2042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hinds D.A., Kloek A.P., Jen M., Chen X., Frazer K.A. Common deletions and SNPs are in linkage disequilibrium in the human genome. Nat. Genet. 2006;38:82–85. doi: 10.1038/ng1695. [DOI] [PubMed] [Google Scholar]
- 7.McCarroll S.A., Hadnott T.N., Perry G.H., Sabeti P.C., Zody M.C., Barrett J.C., Dallaire S., Gabriel S.B., Lee C., Daly M.J., et al. Common deletion polymorphisms in the human genome. Nat. Genet. 2006;38:86–92. doi: 10.1038/ng1696. [DOI] [PubMed] [Google Scholar]
- 8.Cooper G.M., Zerr T., Kidd J.M., Eichler E.E., Nickerson D.A. Systematic assessment of copy number variant detection via genome-wide SNP genotyping. Nat. Genet. 2008;40:1199–1203. doi: 10.1038/ng.236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Locke D.P., Sharp A.J., McCarroll S.A., McGrath S.D., Newman T.L., Cheng Z., Schwartz S., Albertson D.G., Pinkel D., Altshuler D.M., et al. Linkage disequilibrium and heritability of copy-number polymorphisms within duplicated regions of the human genome. Am. J. Hum. Genet. 2006;79:275–290. doi: 10.1086/505653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kato M., Nakamura Y., Tsunoda T. MOCSphaser: a haplotype inference tool from a mixture of copy number variation and single nucleotide polymorphism data. Bioinformatics. 2008;24:1645–1646. doi: 10.1093/bioinformatics/btn242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kato M., Nakamura Y., Tsunoda T. An algorithm for inferring complex haplotypes in a region of copy-number variation. Am. J. Hum. Genet. 2008;83:157–169. doi: 10.1016/j.ajhg.2008.06.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Shen F., Huang J., Fitch K.R., Truong V.B., Kirby A., Chen W., Zhang J., Liu G., McCarroll S.A., Jones K.W., et al. Improved detection of global copy number variation using high density, non-polymorphic oligonucleotide probes. BMC Genet. 2008;9:27.21–27.18. doi: 10.1186/1471-2156-9-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.The International HapMap Consortium. A haplotype map of the human genome. Nature. 2005;437:1299–1320. doi: 10.1038/nature04226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.The International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature. 2007;449:851–861. doi: 10.1038/nature06258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ishikawa S., Komura D., Tsuji S., Nishimura K., Yamamoto S., Panda B., Huang J., Fukayama M., Jones K.W., Aburatani H. Allelic dosage analysis with genotyping microarrays. Biochem. Biophys. Res. Commun. 2005;333:1309–1314. doi: 10.1016/j.bbrc.2005.06.040. [DOI] [PubMed] [Google Scholar]
- 16.Komura D., Nishimura K., Ishikawa S., Panda B., Huang J., Nakamura H., Ihara S., Hirose M., Jones K.W., Aburatani H. Noise reduction from genotyping microarrays using probe level information. In Silico Biol. 2006;6:0009. [PubMed] [Google Scholar]
- 17.Price T.S., Regan R., Mott R., Hedman A., Honey B., Daniels R.J., Smith L., Greenfield A., Tiganescu A., Buckle V., et al. SW-ARRAY: a dynamic programming solution for the identification of copy-number changes in genomic DNA using array comparative genome hybridization data. Nucleic Acids Res. 2005;33:3455–3464. doi: 10.1093/nar/gki643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Feuk L., Carson A.R., Scherer S.W. Structural variation in the human genome. Nat. Rev. Genet. 2006;7:85–97. doi: 10.1038/nrg1767. [DOI] [PubMed] [Google Scholar]
- 19.Feuk L., Marshall C.R., Wintle R.F., Scherer S.W. Structural variants: changing the landscape of chromosomes and design of disease studies. Hum. Mol. Genet. 2006;15(Spec No 1):R57–R66. doi: 10.1093/hmg/ddl057. [DOI] [PubMed] [Google Scholar]
- 20.Zhang J., Feuk L., Duggan G.E., Khaja R., Scherer S.W. Development of bioinformatics resources for display and analysis of copy number and other structural variants in the human genome. Cytogenet. Genome Res. 2006;115:205–214. doi: 10.1159/000095916. [DOI] [PubMed] [Google Scholar]
- 21.Cheung J., Estivill X., Khaja R., MacDonald J.R., Lau K., Tsui L.C., Scherer S.W. Genome-wide detection of segmental duplications and potential assembly errors in the human genome sequence. Genome Biol. 2003;4:R25. doi: 10.1186/gb-2003-4-4-r25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gonzalez E., Kulkarni H., Bolivar H., Mangano A., Sanchez R., Catano G., Nibbs R.J., Freedman B.I., Quinones M.P., Bamshad M.J., et al. The influence of CCL3L1 gene-containing segmental duplications on HIV-1/AIDS susceptibility. Science. 2005;307:1434–1440. doi: 10.1126/science.1101160. [DOI] [PubMed] [Google Scholar]
- 23.Aitman T.J., Dong R., Vyse T.J., Norsworthy P.J., Johnson M.D., Smith J., Mangion J., Roberton-Lowe C., Marshall A.J., Petretto E., et al. Copy number polymorphism in Fcgr3 predisposes to glomerulonephritis in rats and humans. Nature. 2006;439:851–855. doi: 10.1038/nature04489. [DOI] [PubMed] [Google Scholar]
- 24.Valentonyte R., Hampe J., Huse K., Rosenstiel P., Albrecht M., Stenzel A., Nagy M., Gaede K.I., Franke A., Haesler R., et al. Sarcoidosis is associated with a truncating splice site mutation in BTNL2. Nat. Genet. 2005;37:357–364. doi: 10.1038/ng1519. [DOI] [PubMed] [Google Scholar]
- 25.Perry G.H., Dominy N.J., Claw K.G., Lee A.S., Fiegler H., Redon R., Werner J., Villanea F.A., Mountain J.L., Misra R., et al. Diet and the evolution of human amylase gene copy number variation. Nat. Genet. 2007;39:1256–1260. doi: 10.1038/ng2123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gasche Y., Daali Y., Fathi M., Chiappe A., Cottini S., Dayer P., Desmeules J. Codeine intoxication associated with ultrarapid CYP2D6 metabolism. N. Engl. J. Med. 2004;351:2827–2831. doi: 10.1056/NEJMoa041888. [DOI] [PubMed] [Google Scholar]
- 27.Hedrick P.W. Gametic disequilibrium measures: proceed with caution. Genetics. 1987;117:331–341. doi: 10.1093/genetics/117.2.331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Mueller J.C. Linkage disequilibrium for different scales and applications. Brief Bioinform. 2004;5:355–364. doi: 10.1093/bib/5.4.355. [DOI] [PubMed] [Google Scholar]
- 29.Smith A.V., Thomas D.J., Munro H.M., Abecasis G.R. Sequence features in regions of weak and strong linkage disequilibrium. Genome Res. 2005;15:1519–1534. doi: 10.1101/gr.4421405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.McCarroll S.A., Altshuler D.M. Copy-number variation and association studies of human disease. Nat. Genet. 2007;39:S37–S42. doi: 10.1038/ng2080. [DOI] [PubMed] [Google Scholar]
- 31.Korn J.M., Kuruvilla F.G., McCarroll S.A., Wysoker A., Nemesh J., Cawley S., Hubbell E., Veitch J., Collins P.J., Darvishi K., et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 2008;40:1253–1260. doi: 10.1038/ng.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Excoffier L., Slatkin M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 1995;12:921–927. doi: 10.1093/oxfordjournals.molbev.a040269. [DOI] [PubMed] [Google Scholar]
- 33.Jakobsson M., Scholz S.W., Scheet P., Gibbs J.R., VanLiere J.M., Fung H.C., Szpiech Z.A., Degnan J.H., Wang K., Guerreiro R., et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature. 2008;451:998–1003. doi: 10.1038/nature06742. [DOI] [PubMed] [Google Scholar]
- 34.Niu T., Qin Z.S., Xu X., Liu J.S. Bayesian haplotype inference for multiple linked single-nucleotide polymorphisms. Am. J. Hum. Genet. 2002;70:157–169. doi: 10.1086/338446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nebert D.W., Dalton T.P. The role of cytochrome P450 enzymes in endogenous signalling pathways and environmental carcinogenesis. Nat. Rev. Cancer. 2006;6:947–960. doi: 10.1038/nrc2015. [DOI] [PubMed] [Google Scholar]
- 36.de Cid R., Riveira-Munoz E., Zeeuwen P.L., Robarge J., Liao W., Dannhauser E.N., Giardina E., Stuart P.E., Nair R., Helms C., et al. Deletion of the late cornified envelope LCE3B and LCE3C genes as a susceptibility factor for psoriasis. Nat. Genet. 2009;41:211–215. doi: 10.1038/ng.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kidd J.M., Cooper G.M., Donahue W.F., Hayden H.S., Sampas N., Graves T., Hansen N., Teague B., Alkan C., Antonacci F., et al. Mapping and sequencing of structural variation from eight human genomes. Nature. 2008;453:56–64. doi: 10.1038/nature06862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kato M., Sekine A., Ohnishi Y., Johnson T.A., Tanaka T., Nakamura Y., Tsunoda T. Linkage disequilibrium of evolutionarily conserved regions in the human genome. BMC Genomics. 2006;7:326.321–326.328. doi: 10.1186/1471-2164-7-326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kato M., Miya F., Kanemura Y., Tanaka T., Nakamura Y., Tsunoda T. Recombination rates of genes expressed in human tissues. Hum. Mol. Genet. 2008;17:577–586. doi: 10.1093/hmg/ddm332. [DOI] [PubMed] [Google Scholar]
- 40.Conrad D.F., Pinto D., Redon R., Feuk L., Gokcumen O., Zhang Y., Aerts J., Andrews T.D., Barnes C., Campbell P., et al. Origins and functional impact of copy number variation in the human genome. Nature. 2009 doi: 10.1038/nature08516. doi:10.1038/nature08516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Qin J., Jones R.C., Ramakrishnan R. Studying copy number variations using a nanofluidic platform. Nucleic Acids Res. 2008;36:e116. doi: 10.1093/nar/gkn518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dube S., Qin J., Ramakrishnan R. Mathematical analysis of copy number variation in a DNA sample using digital PCR on a nanofluidic device. PLoS One. 2008;3:e2876. doi: 10.1371/journal.pone.0002876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Abecasis G.R., Cookson W.O. GOLD—graphical overview of linkage disequilibrium. Bioinformatics. 2000;16:182–183. doi: 10.1093/bioinformatics/16.2.182. [DOI] [PubMed] [Google Scholar]
- 44.Dudbridge F. Pedigree disequilibrium tests for multilocus haplotypes. Genet. Epidemiol. 2003;25:115–121. doi: 10.1002/gepi.10252. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.