Abstract
Frontotemporal dementia with ubiquitin-positive inclusions (FTLD-U) can be caused by mutations in the progranulin gene (GRN). Progranulin (PGRN) is a cysteine-rich growth factor, which is proteolytically cleaved by elastase to produce several granulins (GRNs). All FTLD-U mutations in GRN characterized to date result in reduced secreted PGRN protein. We recently reported a Spanish family with progressive nonfluent aphasia and dementia in which a novel C521Y mutation segregates with disease. A second cysteine mutation (C139R) has also been reported to be disease specific. Allele-specific mRNA expression assays in brain reveal that the C521Y mutant allele is expressed at similar levels to the wild-type allele. Furthermore, plasma PGRN levels in C521Y carriers are comparable to non-carrier family relatives, suggesting that the mutation does not affect PGRN protein expression and secretion in vivo. Despite normal PGRN levels C521Y and C139R mutant GRNs show reduced neurite growth stimulating activity in vitro. Further study revealed that these mutations also cause impaired cleavage of PGRN by elastase. Our data suggest that these mutations affect the function of full-length PGRN as well as elastase cleavage of PGRN into GRNs, leading to neurodegeneration.
Keywords: progranulin, granulin, FTD, elastase, neurite outgrowth, neuronal survival
Introduction
Frontotemporal dementia (FTD) is a set of progressive disorders affecting predominantly the frontal and/or anterior temporal lobes of the brain. Clinically, it is characterized by prominent impairment in personality, behavior and/or language (McKhann et al. 2001). The neuropathology of FTD is heterogeneous (Trojanowski and Dickson 2001) but can be classified into three major categories: FTD with tau-positive inclusions; FTD with ubiquitin-positive, tau-negative inclusions and FTD with inclusions negative for both. The genetics of FTD is complex, with multiple genes associated with disease etiology. Approximately 35-40% of autosomal dominant FTD results from mutations in the gene encoding microtubule associated protein tau (MAPT) on chromosome 17q21 (http://www.molgen.ua.ac.be/FTDMutations/) and show abnormal tau pathology in the brain (Hutton et al. 1998; Poorkaj et al. 1998; Spillantini et al. 1998). However, several FTD families with significant linkage to the same chromosomal region exhibit tau-negative, ubiquitin-positive inclusions (FTLD-U) in the brain and lack mutations in MAPT. The cause of FTD in this subset of families was shown to be mutations in the progranulin gene (GRN) (Baker et al. 2006; Cruts et al. 2006). Sixty-six pathogenic mutations have been reported (http://www.molgen.ua.ac.be/FTDMutations/) in GRN. The majority of these mutations are null mutations causing premature termination of the coding sequence leading to the degradation of mutant GRN mRNA by nonsense-mediated decay, and a 50% decrease in the GRN mRNA and protein. It has been proposed that FTD in these families results from haploinsufficiency.
Progranulin protein (PGRN) is a 68 kDa glycoprotein composed of a signal peptide followed by 7.5 tandem repeats of 12 cysteine-rich granulin motifs, separated by linker regions. Glycosylation of PGRN leads to the formation of multiple higher molecular weight forms, the most common of which is a 88 kD protein. PGRN exists as a secreted full-length precursor protein, which is converted into several 6-25 kDa fragments, called granulins (GRNs), via proteolytic processing by elastase and other extracellular proteases (He and Bateman 2003). Five possible elastase cleavage sites, all within the inter-GRN linker regions, have been identified (Zhu et al. 2002). Two-dimensional NMR spectroscopy has shown that each GRN consists of four stacked β-hairpins, with disulfide bridges forming an axial rod-like structure (Hrabal et al. 1996). Both PGRN and GRNs (GRN A-F) are growth factors, involved in a wide variety of functions including signal transduction, inflammation, proliferation, tumorigenesis and wound repair (Eriksen and Mackenzie 2008). The full-length PGRN and GRNs may have opposing functions in wound repair and inflammation (anti-inflammatory vs proinflammatory). Conversion of PGRN to GRNs by elastase, a process regulated by secretory leukocyte protease inhibitor, may act as a molecular switch in host defense and wound repair (Zhu et al. 2002). Recently, both PGRN and a recombinant GST-fusion peptide which contains the GRN E sequence (GST-GRNE) have been shown to have neurotrophic properties: promoting neuronal survival and neurite outgrowth (Van Damme et al. 2008).
We have previously reported a point mutation in the signal peptide of PGRN (A9D) that causes FTLD-U in a large kindred, HDDD2 (Mukherjee et al. 2006). This mutation did not affect PGRN mRNA levels but the mutant protein was trapped within the cell and did not undergo normal glycosylation or secretion, resulting in a functional haploinsufficiency (Mukherjee et al. 2008). Two other pathogenic missense mutations (P248L and R432C) were recently shown to have impaired secretion (Shankaran et al. 2008). It appears that all GRN mutations characterized so far cause reduced protein production or secretion.
We recently identified a novel C521Y GRN mutation in a Spanish family with progressive nonfluent aphasia (PNFA). The C521Y mutation segregates with the disease. It was present in all individuals with PNFA or dementia and in some asymptomatic individuals indicating incomplete penetrance (Cruchaga et al. 2008). This residue is located within the last GRN peptide (GRN E) and is conserved among vertebrate species, suggesting that it might be important for PGRN/GRN function. Another cysteine mutation (C139R) has been reported to be disease-specific by two groups (Bernardi et al. 2008; Brouwers et al. 2008). This cysteine residue is located within GRN F and is also conserved among different species. The plasma PGRN level in an individual who carries the C139R mutation was relatively low compare to normal individuals, but falls outside the range of the typical loss-of-function mutations (Finch et al. 2009). The program SIFT predicts that both the C521Y and the C139R mutations are likely to affect protein function (Cruchaga et al. 2008; Brouwers et al. 2008).
While it is thought that decreased PGRN protein causes FTD, the mechanism by which PGRN haploinsufficiency affects neuronal function and leads to neurodegeneration is not clear. Studying the pathogenic mechanism of the C521Y and C139R mutations may help us understand how haploinsufficiency of PGRN/GRN causes neurodegeneration.
Materials and methods
Plasmids, cells and antibodies
The full-length GRN cDNA in pCMV-SPORT6 vector was purchased from Invitrogen (Carlsbad, CA). The C521Y and C139R point mutations were introduced into the GRN cDNA using the QuickChange II site-directed mutagenesis kit (Stratagene, La Jolla, CA). Human embryonic kidney 293 (HEK) cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM, GibcoBRL, Grand Island, NY), containing 10% fetal bovine serum (FBS), 2 mM L-Glutamine, 100 μg/ml penicillin/streptomycin under a 5% CO2 atmosphere. HEK293 cells stably expressing vector control or 6xHis-GFP-tagged GRN, GRN-C139R or GRN-C521Y were grown to confluence in DMEM supplemented with 10% FBS, 100 μg/ml penicillin/streptomycin and 300 μg/ml G418. Medium was changed to 0.1%BSA DMEM for 24 hrs. Conditioned medium was collected after 48 hrs and used immediately after collection with B27 as a supplement for in vitro culture of E15 mouse cortical neurons.
The following antibodies were used: a rabbit polyclonal antibody against the C-terminus of PGRN (aa 494-594) (anti-PGRNC-term, 1:500 dilution, Zymed, San Francisco, CA); a goat polyclonal antibody against full-length recombinant PGRN (anti-PGRNFL, 1:1000 dilution, R&D Systems, Minneapolis, MN); a rabbit polyclonal antibody against GRN A (aa 281-336) (anti-PGRNGRN A, 1:500 dilution, SDI, Newark, DE); a polyclonal anti-β-actin antibody (1:1000 dilution, Sigma, St. Louis, MO); a mouse monoclonal anti-GFP antibody (1:7500 dilution). The recombinant GRN E peptide (GST-GRN E, Van Damme et al, 2008) and the C521Y GRN E peptide (GST-C521Y) were purchased from Abnova Corporation (Taipei, Taiwan). Both peptides contain a GST tag and PGRN aa 494-594, encompassing GRN E and flanking regions, with a molecular weight of 37 kDa.
GRN total gene expression
RNA from fresh blood was isolated using the “Tempus Blood RNA tube Kit” (Applied Biosystems, Foster City, USA) and single-strand cDNA was synthesized using the SuperScript III RT-PCR kit (Invitrogen, Carlsbad, USA) and a pool of random primers. RNA from brain tissue was isolated using the “RNeasy Mini Kit (Qiagen) and cDNA was synthesized using the High-Capacity cDNA Archive kit (ABI). Gene expression levels were analyzed using an ABI-7500 real-time PCR system. GRN expression levels were measured by a custom TaqMan assay for GRN exon 13 (primers and probes available upon request) using GAPDH as the endogenous control. Each real-time PCR run included within-plate duplicates and each experiment was performed, at least twice for each sample. Real-time data were analyzed using the comparative Ct method (Muller et al, 2002). The Ct values of each sample were normalized with the Ct value for the housekeeping gene, GADPH, and were corrected for the PCR efficiency of each assay (Muller et al, 2002), although the efficiency of all reactions was close to 100%. Only samples with a standard error of <0.15% were analyzed.
C521Y allele specific gene expression
We developed an allele-specific real-time PCR assay to calculate the mutant/non-mutant allelic expression ratio (AER) at the c.1562 GRN position among C521Y carriers. First, we calculated the efficiency of the allele-specific TaqMan (Applied Biosystems, Foster City) probes (sequence available upon requests) with serial dilutions of a cDNA heterozygous for the mutation. Second, we confirmed that the technique was able to quantify the amount of the mutant allele in relation to the normal allele by generating a standard curve mixing the mutant cDNA with the normal cDNA at different ratios (data not shown). Finally, the mutant/non-mutant allele expression ratio was calculated comparing the Ct value for the FAM-probe (mutant allele, A) with that of the VIC-probe (normal allele, G) using heterozygous genomic DNA as a reference by the comparative Ct method as explained above.
PGRN ELISA
PGRN protein levels in human plasma samples were measured using the human Progranulin ELISA kit (Adipogen Inc., Seoul, Korea), according to the Manufacturer’s instructions.
Transfections and immunoblotting
HEK cells were transiently transfected with wild-type (WT) or mutant GRN constructs using the FuGENE 6 HD transfection reagent (Roche, Indianapolis, IN). Approximately 18 hrs after transfection, transfection media was replaced with 0.7 ml serum-free media. 24 hrs later, the conditioned media was collected and cells were lysed in lysis buffer (50 mM Tris pH 7.6, 150 mM NaCl, 2 mM EDTA, 1% NP-40, 0.5% Triton X-100 and protease inhibitor cocktail (Roche, one tablet in 10ml lysis buffer according to Manufactorer’s instructions) for further analysis. For immunoblotting, cell lysates (20 μg) and conditioned media (15 μl) were separated in 4-20% Tris-HCl or 4-12% Bis-Tris criterion gels (Bio-Rad, Hercules, CA) under reducing (with DTT) or non-reducing (without DTT) conditions, transferred to PVDF membrane and detected by appropriate antibodies.
Deglycosylation
Cell lysates and media were treated with PNGase F (New England Biolabs, Ipswich, MA) according to the manufacturer’s instructions. For deglycosylation under non-reducing conditions, DTT was omitted from all reaction buffers.
Elastase treatment
Approximately 18 hrs after transfection, growth medium was replaced with serum-free DMEM. Cell lysates and media were collected 24 hrs later and treated with human neutrophil elastase (1 unit/ml) (Sigma) or saline as a control for various times at 37 °C.
Neuronal cultures
Cortical and motor neuron cultures were prepared from E14 Wistar rat embryos as previously described (Van Damme et al., 2007; Van Damme et al., 2008). The frontal cortex and the ventral part of the spinal cord were dissected and dissociated enzymatically and mechanically. A motor neuron-enriched fraction was obtained after centrifugation on a 6.2% Optiprep cushion (Axis-Shields PoC AS, Oslo, Norway). For cortical neurons, the density centrifugation step was omitted. Neurons were plated in culture dishes coated with poly-L-ornithine and laminin (Sigma). The purity of these cultures was previously established (Van Damme et al., 2007; Van Damme et al., 2008).
The culture medium consisted of L15 supplemented with sodium bicarbonate (0.2%), glucose (3.6 mg/ml), progesterone (20 nM), insulin (5 μg/ml), putrescine (0.1 mM), conalbumin (0.1 mg/ml), sodium selenite (30 nM), penicillin (100 U/ml), streptomycin (100μg/ml) and chick embryo extract (5%) (Invitrogen, Grand Island, NY).
Quantification of neuronal survival
Motor neuron survival was quantified by direct counting of living cells under a phase-contrast microscope as previously described (Van Den Bosch et al., 2004). A marked area of the culture dish was counted on day 1 in culture and again 24 hrs after treatment of the culture with buffer control, GST-GRN E or GST-C521Y peptides (100 ng/ml). The number of surviving cells on day 2 was normalized to the number of cells on day 1. For quantification of cortical neurons a calcein assay was used (Van Damme et al., 2008). After seeding neurons in a 96-well plate, wells were treated with buffer control, GST-GRN E or GST-C521Y peptides. Neuronal survival was quantified 24 hrs later after loading the cells with 2 μM calcein-AM in the culture medium at 37°C for 45 minutes. After washing the plate with PBS, the fluorescence was measured using a plate reader (Victor X3, Perkin Elmer). The average of at least 8 wells per condition was normalized to the signal of untreated control wells. Since the recombinant GRN peptides were resuspended in a buffer containing reduced glutathione, this was added to the corresponding control wells. A one-way analysis of variance with Tukey-Kramer multiple comparisons was used to calculate significance.
Quantification of neurite length and soma size
For quantification of neurite length and soma size, cultures were immunostained with neurofilament heavy chain (NF-H, 1:400 rabbit polyclonal, Sigma) as previously described (Van Damme et al., 2008). After seeding, cultures were treated with buffer control, GST-GRN E or GST-C521Y peptides (100 ng/ml). After 24 hrs, the cells were fixed and stained. The soma size and length of the longest neurite per cell was marked manually on digital images and measured using NIS Elements software (Nikon Instruments, Melville, NY). A one-way analysis of variance with Tukey-Kramer multiple comparisons was used to calculate significance.
Full-length PGRN axon outgrowth assay
Dissociated primary neurons were cultured essentially as described previously (Liu et al., 2004) with the following modifications. Briefly, embryos were removed from timed-pregnant mice at E15 (for cortical neurons). Mouse brains were dissected in cold Hank’s buffered salt solution (Gibco). The cortex was cut into small pieces and trypsinized for 15 min at 37°C. The dissociated neurons were resuspended in Neurobasal medium supplemented with heat-inactivated FBS and 20 Units/ml of penicillin/streptomycin. Cells were grown on dishes or coverslips coated overnight with 100 μg/ml poly-L-lysine and 5 μg/ml laminin at 37°C in a 5% CO2 incubator.
To analyze neurite outgrowth, E15 cortical neurons were cultured in Neurobasal medium with 10% FBS for 2 hrs. Neurons were then cultured for 20 hrs in the presence of the control or PGRN conditioned medium and then fixed with 4% paraformaldehyde for 10 min, preincubated with 5% BSA in PBS with 0.1% Triton X-100 and stained using a monoclonal antibody specific for neuronal βIII tubulin. Nuclei were visualized after staining with Hoechst dye (Sigma). Quantification was performed using NIH ImageJ software.
Results
The C521Y mutation does not affect GRN mRNA or PGRN protein expression and secretion in vivo
Lymphocyte mRNA was available from 7 PNFA family members, five of whom were C521Y GRN mutation carriers (Table 1A). Sequencing of GRN cDNA showed that the C521Y mutant allele was expressed in lymphocytes. Gene expression assays for GRN exon 13 showed that the level of total GRN mRNA in 2 out of the 5 mutation carriers (III.3 and III.9) was about 2.5 times higher than in non-carrier members (Table 1A). While there was variability among individuals, there was no evidence that total GRN mRNA is reduced in C521Y mutation carriers compared to family non-carrier controls.
Table 1. A. Total and allele-specific GRN gene expression in white blood cells of the PNFA family.
| Index case |
C521Y GRN DNA variant |
GRN cDNA RE1(SEM) | c.1562A/c.1562G AER 2(SEM) |
|---|---|---|---|
| II.3 | Yes | 0.94 ± 0.017 | 0.96 ± 0.026 |
| III.1 | Yes | 1.10 ± 0.007 | 1.09 ± 0.110 |
| III.3 | Yes | 2.33 ± 0.17 | 1.04 ± 0.025 |
| III.5 | No | 0.98 ± 0.039 | 0.08 ± 0.001** |
| III.6 | Yes | 1.01 ± 0.029 | 1.19 ± 0.061 |
| III.7 | No | 1.22 ± 0.040 | 0.05 ± 0.005** |
| III.9 | Yes | 2.98 ± 0.044 | 1.15 ± 0.039 |
| B. Allele-specific GRN gene expression in brain of the III.2 individual | ||
|---|---|---|
| Index case |
Brain region | c.1562A/c.1562G AER 2(SEM) |
| III.2 | Temporal | 1.13 ± 0.05 |
| Occipital | 1.07 ± 0.06 | |
Relative GRN mRNA levels measured in white blood cell. Only values with an standard error less than 15% were analyzed.
A/ G allelic expression ratio: This value indicates the ratio of the A mutant allele expressed in relation to the wild type, G allele.
SEM: Standard Error of the Mean
Values close to 0 indicate that the mutant allele is not expressed.
We examined allele-specific expression of GRN mRNA. The c.1562A/c.1562G allelic expression ratio (AER) in non-carriers was close to 0 confirming that the mutant allele was not expressed. In all of the C521Y carriers the AER was close to 1 in lymphocytes (Table 1A) and in brain (Table 1B), indicating that there was no significant differential expression of G/A alleles at the c.1562 position.
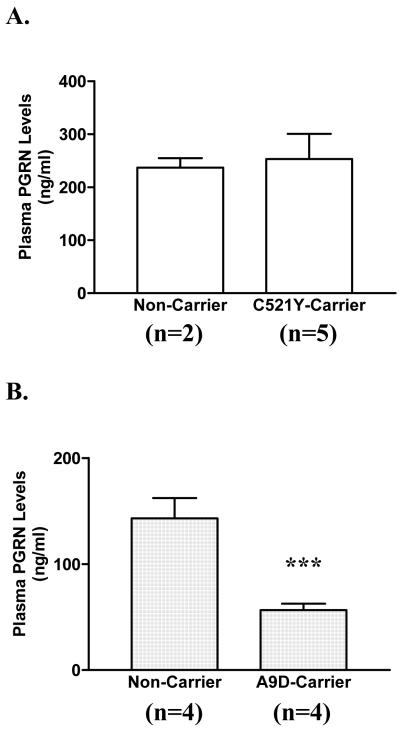
We also measured plasma PGRN protein levels in the same 7 PNFA family members using a PGRN ELISA assay (Fig. 1A). No significant differences in the levels of PGRN were found between carriers and non-carriers. While plasma PGRN protein levels in the C521Y carriers vary a lot (119 – 347 ng/ml), they all fall in the range for normal individuals. This high degree of variability in normal individuals is confirmed by other reports (Finch et al. 2009; Ghidoni et al. 2008). In contrast, the plasma PGRN levels in A9D mutation carriers were significantly lower than HDDD2 family controls (Fig. 1B). These data demonstrate that the C521Y mutation does not affect GRN mRNA or PGRN protein expression and secretion in vivo.
Figure 1. Plasma PGRN levels in GRN mutation carriers and non-carriers in PNFA (A) and HDDD2 (B) families.
PGRN levels in plasma samples from PNFA family (5 C521Y mutation carriers and 2 non-carriers) and HDDD2 family (4 A9D mutation carriers and 4 non-carries) were measured using a human Progranulin ELISA kit, according to the Manufacturer’s instructions. The bar diagram represents mean values ± SEM. Asterisks indicate a significant difference between A9D-carriers and HDDD2 family controls, determined by Student’s t-test (*** p < 0.001). There is no difference between PNFA family non-carriers and C521Y-carriers (p=0.847).
The C521Y and C139R mutations do not affect PGRN expression and secretion in a transfection system
Since human C139R samples were unavailable, we compared the effects of the C521Y and C139R mutations on PGRN function in an in vitro transfection system. We introduced the point mutations into GRN cDNA and transiently transfected WT and mutant GRN in HEK cells and analyzed PGRN protein in both cell lysates and media. As observed in vivo, the C521Y mutant protein was normally expressed and secreted, as was the C139R mutant protein (Fig. 2A). Semi-quantitative analysis of the gels showed that the mutant and wild type proteins were expressed (Fig. 2B) and secreted (Fig. 2C) at similar levels. We also compared the kinetics of protein maturation and secretion for WT and C521Y PGRN. Both showed a similar temporal secretion pattern (Supplemental Figure 1).
Figure 2. The C521Y and C139R mutations do not affect PGRN expression and secretion in a transfection system.
A. HEK cells were transfected with GFP, WT-PGRN or mutant PGRNs as indicated. Cell lysates (5 μg) and conditioned media (5 μl) were separated on a 4-12% Bis-Tris gel under reducing conditions. The membrane was western blotted with anti-PGRNC-term (upper panel). The same membrane was stripped and reblotted with an anti-actin antibody (lower panel). B, C. Semi-quantitative densitometric analysis. B. The densities of the PGRN bands (lysates) were quantified using Quantity One Software (Bio-rad) and normalized to actin levels. %WT was plotted. C. Percentages of normalized PGRN in media over total (media + lysates) were plotted.
Cysteine mutations affect protein mobility under non-reducing conditions, but not under reducing conditions
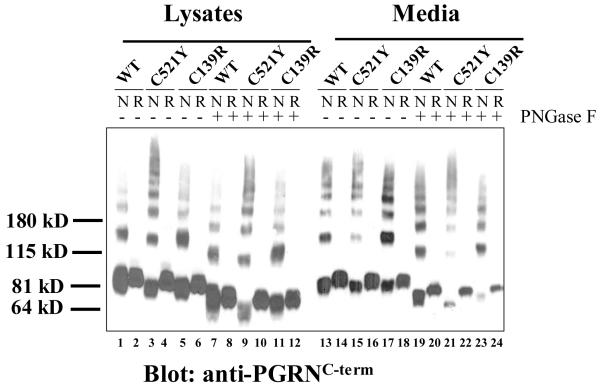
Disulfide bridges are very important for PGRN/GRN folding and structure. Losing one conserved cysteine will likely disrupt one of the disulfide bridges and therefore affect protein conformation. To examine whether protein conformation is changed, we analyzed protein mobility under reducing and non-reducing conditions in the presence and absence of PNGase F to remove all of the sugars. As shown in Fig. 3, we observed that in lysates under non-reducing conditions the C521Y and C139R mutant proteins migrated slightly faster than the WT protein (lane 1 vs lanes 3 & 5). This mobility shift was more pronounced after PNGase F treatment with the C521Y mutant (lane 7 vs lane 9), although not as prominent with the C139R mutant (lane 7 vs lane 11). In media, the mobility shift was not obvious between the WT and the mutants without PNGase F treatment (lane 13 vs lanes 15 &17). The mobility shift became evident with the C521Y mutant after PNGase F treatment (lane 19 vs 21). Importantly, the mobility shift was observed under non-reducing conditions (odd number lanes), but not reducing conditions (even number lanes), suggesting that the shift is related to disulfide bond formation. We detected less mutant protein (compared to the WT) in the media under non-reducing conditions after PNGase F treatment (lanes 21 and 23). This is most likely attributable to the loss of reactivity with the anti-C-terminal antibody (anti-PGRNC-term, Zymed) under these conditions, because similar amounts of protein were detected when the same membrane was reblotted with an anti-full-length antibody (anti-PGRNFL, R&D), which detects PGRN better under non-reducing conditions (lanes 21 and 23, Supplemental Figure 2).
Figure 3. The cysteine mutation affects protein mobility under non-reducing conditions, but not reducing conditions.
Media from HEK cells transfected with WT, C521Y or C139R were collected and subjected to PNGase F treatment. Samples were mixed with non-reducing (N) or reducing (R) sample buffers, separated on a 4-12% Bis-Tris gel and western blotted with anti-PGRNC-term.
The mutant PGRN proteins did not promote neurite outgrowth
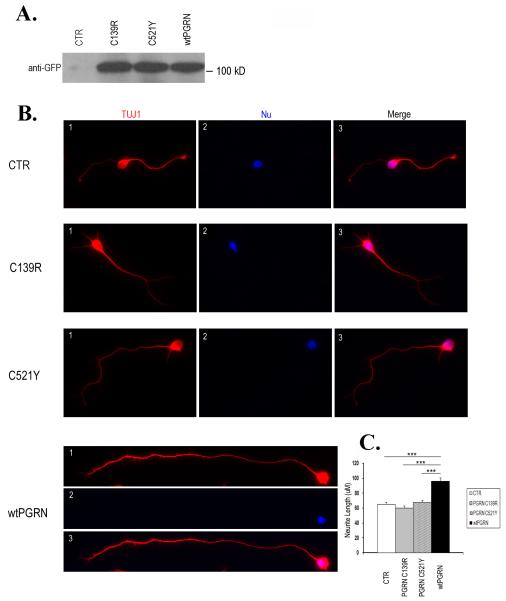
We then tested whether the mutations affect PGRN function. We have previously shown that PGRN promotes neurite outgrowth (Van Damme et al., 2008). To investigate whether this function of PGRN is compromised by the mutations we generated HEK cells stably expressing WT or mutant PGRN. Conditioned media from these cell lines were collected. The effect of WT and mutant PGRN on neurite outgrowth in cortical neurons was examined. WT and mutant PGRN were expressed and secreted into the media at equivalent levels (Fig. 4A). However, while WT PGRN promoted neurite outgrowth as previously reported, the C521Y and C139R mutant PGRNs failed to stimulate neurite outgrowth (Fig. 4B&C).
Figure 4. The C139R and C521Y mutant PGRN do not stimulate neurite outgrowth.
A. 10 μl conditioned media from HEK cells stably expressing vector control (CTR), C139R, C521Y and wtPGRN was resolved in 7.5% SDS-Tris gel and western blotted with a monoclonal anti-GFP antibody. B. Primary cultures of mouse cortical neurons were grown in the presence of 1 ml conditioned media from HEK cells expressing vector control (CTR), wtPGRN or PGRN mutants C139R and C521Y. Axonal outgrowth assays were performed as described in Materials and Methods. Shown are representative neurons visualized with neuronal βIII tubulin (TUJ1, Red, 1), nuclei (Nu, Blue, 2) and Merge (3). C. Quantification of axon outgrowth. The average length of the longest neurite was quantified using ImageJ software from 200 randomly selected neurons in each group. The data represent three independent experiments. The bar diagram represents mean values ± SEM. Asterisks indicate a significant difference from the wtPGRN, as determined by Student’s t-test (*** p < 0.001).
Cysteine mutations affect PGRN cleavage by elastase
Since PGRN and GRNs may both have important functions in the nervous system and in neurodegeneration we tested whether the cysteine mutations also affect production and activity of GRNs. We examined whether the cysteine mutations affect elastase cleavage of PGRN and GRN production.
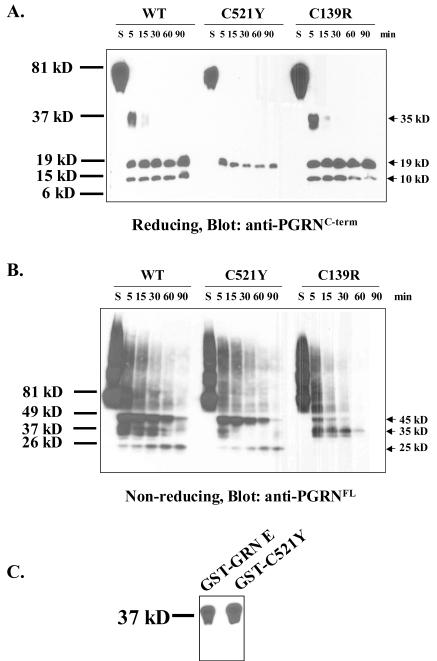
In order to detect intermediate products, secreted WT, C521Y and C139R mutant proteins were collected and treated with elastase for various times. The samples were run under both reducing and non-reducing conditions, because some of the antibodies used recognize epitopes that are sensitive to these conditions. The anti-PGRNC-term detects proteins better under reducing conditions while the anti-PGRNFL detects antigen better under non-reducing conditions. Under reducing conditions with anti-PGRNC-term, we observed three cleavage products (Fig. 5A): an intermediate fragment of ~35 kD which disappeared within 15 min post-elastase treatment and two smaller fragments of approximately 19 kD and 10 kD. All three fragments were greatly reduced or not detected in the C521Y mutant (Fig. 5A). The 35 kD fragment was completely lacking in the C521Y mutant, not showed up even after long exposure (Supplemental Figure 3A). Using this antibody we did not observe any differences in the cleavage patterns observed for the WT and C139R mutant.
Figure 5. The cysteine mutations affect PGRN cleavage by elastase.
A. Serum-free media collected from HEK cells expressing WT, C521Y or C139R PGRN were treated with saline (S) for 90 min or elastase for various times as indicated and resolved on a 4-20% Tris-HCl gel under reducing conditions. The membrane was blotted with anti-PGRNC-term. Arrows indicate the 35 kD, 19 kD and 10 kD C-terminal fragments detected by anti-PGRNC-term. B. Same as A, except that the samples were run under non-reducing conditions and immunoblotted with anti-PGRNFL. Arrows indicate the 45 kD, 35 kD and 25 kD fragments detected by this antibody. C. 1.25 μg of recombinant GST tagged WT or C521Y GRN E peptides (GST-GRN E and GST-C521Y) were run on a 4-20% Tris-HCl gel under reducing conditions and western blotted with anti-PGRNC-term.
When the samples were run under non-reducing conditions and detected with anti-PGRNFL, we observed another intermediate fragment of 45 kD (Fig. 5B), in addition to the 35 kD band detected with anti-PGRNC-term (Fig. 5A). We also detected a 25 kD fragment which was normally produced by the C521Y mutant. This fragment is however absent with the C139R mutant (Fig. 5B). The 45 kD fragment was greatly reduced with the C139R mutant, but not the C521Y mutant, suggesting that it is an N-terminal intermediate. When blotted with an antibody raised against GRN A (SDI), we observed a 45 kD fragment in the WT and the C521Y mutant, but not the C139R mutant (Supplemental Figure 3B), confirming that this is an N-terminal intermediate.
Failure to detect certain fragments could be due to the absence of these fragments or the loss of the epitopes detected by the antibodies (due to the mutations). To distinguish between these two possibilities, we immunoblotted GST-GRN E and GST-C521Y peptides with anti-PGRNC-term (Fig. 5C). Both peptides are readily detected by this antibody, suggesting that the C-terminal fragments are indeed absent in cultures expressing the C521Y cDNA.
C521Y mutant peptide promotes neuronal survival and neurite outgrowth
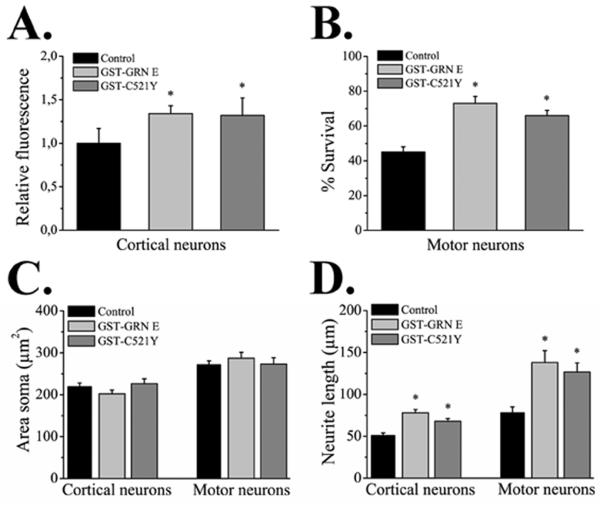
As previously shown, GST-GRN E has neurotrophic properties in primary rat neuronal cultures (Van Damme et al., 2008). Since the C521Y mutation is expected to alter the disulfide bonding and tertiary structure of the GRN E domain, we examined the effect of GST-GRN E or GST-C521Y peptides on neuronal survival and neurite length. As expected, treatment with 100 ng/ml of GST-GRN E enhanced neuronal survival. Surprisingly, the GST-C521Y also improved neuronal survival, both in cortical neuronal cultures (Fig. 6A) and motor neuron cultures (Fig. 6B). We then examined the effect of the GST-GRN E and GST-C521Y peptides on neurite outgrowth in cortical and motor neurons. As previously observed (Van Damme et al., 2008), the GST-GRN E peptide stimulated neurite outgrowth in both neuronal cultures (Fig. 6D), but had no effect on the size of the cell soma (Fig. 6C). The GST-C521Y mutant peptide behaved exactly the same as the WT peptide (Fig. 6C & 6D) indicating that the C521Y mutation does not affect this activity of the GST-GRN E peptide.
Figure 6. The C521Y mutant peptide promotes neuronal survival and neurite outgrowth.
A. Effect of GST-GRN E and GST-C521Y peptides on survival of cortical neurons on day 2 in culture measured by a calcein assay, normalized to untreated controls (control, n = 10, * different from control p < 0.03). B. Effect of GST-GRN E and GST-C521Y peptides on survival of motor neurons on day 2 in culture (n = 6, * different from control p < 0.005). C. Effect of GST-GRN E and GST-C521Y peptides on soma size in cortical and motor neurons (n = 58-106, p > 0.3). D. Effect of GST-GRN E and GST-C521Y peptides on neurite length of longest neurite per cell (n = 58-106, * different from control p ≤ 0.001). Error bars show mean ± SEM. There is no significant difference between GST-GRN E and GST-C521Y.
Discussion
Since the discovery of GRN as one of the disease-causing genes in FTLD-U, 66 pathogenic mutations have been identified in both families and sporadic cases. These pathogenic mutations affect PGRN expression at different levels. Most of the mutations introduce premature termination codons into the mutant allele resulting in null alleles that show decreased mRNA stability suggesting a nonsense mediated decay mechanism with no mutant protein translated (Baker et al., 2006; Cruts et al., 2006). Three missense mutations have been biochemically characterized. The A9D substitution is within the signal peptide, resulting in low protein expression levels and no maturation and secretion of the mutant protein (Mukherjee et al., 2008; Shankaran et al., 2008). P248L and R432C also show defects in secretion, resulting in reduced protein secretion (Shankaran et al., 2008). Thus all of the mutations characterized to date result in reduced secreted PGRN. While these mutations support the haploinsufficiency model, they were not helpful in providing further insight into the role of PGRN/GRN in FTLD-U pathogenesis.
Here we have characterized two pathogenic missense mutations at cysteine residues within PGRN. The C521Y mutation has no effect on GRN mRNA, protein expression, maturation, secretion or stability in vivo and in vitro (Table 1, Fig. 1). Although the C139R was previously reported to be associated with lower plasma PGRN levels than normal, it was expressed at a higher level than other pathogenic mutations, suggesting an intermediate effect on risk for FTLD (Finch et al., 2009). Expression of the C139R mutant in a transfection system did not result in an obvious difference in protein expression and secretion. In contrast to other pathogenic variants in PGRN, these two mutations result in normal levels of secreted PGRN (Fig. 2).
While the amount of secreted full-length PGRN is normal, the activity of the mutant PGRN is compromised. Both C521Y and C139R mutant PGRN failed to induce neurite outgrowth in cortical neurons (Fig. 4), in contrast to the WT PGRN as previous reported (Van Damme et al., 2008). Several lines of evidence suggest that the mutations cause an altered conformation of progranulin that therefore causes a loss of function of full-length progranulin as well as reduced cleavage by elastase to generate GRNs. First, C521Y and C139R mutations alter protein mobility under non-reducing conditions, but not reducing conditions (Fig. 3). This is consistent with the hypothesis that these mutations alter protein conformation possibly by affecting disulfide bond formation. Furthermore, we observed that these mutations affect PGRN proteolytic cleavage by elastase, which is required to produce mature GRNs (Fig. 5). The C521Y and C139R mutations are located within different GRNs (GRN E and GRN F respectively). Accordingly, we observed distinctive cleavage patterns, which are consistent with the location of the mutations.
The amount of the 45 kD and 35 kD fragments decrease with time and correlate with an increase in the 25 kD and 19 kD peptides, suggesting a substrate-product relationship. The 35 kD, 19 kD and 10 kD fragments are detectable with anti-PGRNC-term, suggesting that they are C-terminal intermediates and products. The C521Y mutation, but not the C139R mutation, affects the levels of these fragments. The 45 kD fragment is detected by the GRN A antibody and anti-PGRNFL, while the 25 kD fragment is only detected by anti-PGRNFL, suggesting that these are N-terminal intermediates and products. The level of these fragments is affected by the C139R mutation, but not the C521Y mutation. The 19 kD and the 10 kD peptides are produced at the same time and are stable over time, suggesting that they are not substrate and product, but rather the products of two simultaneous cleavage events.
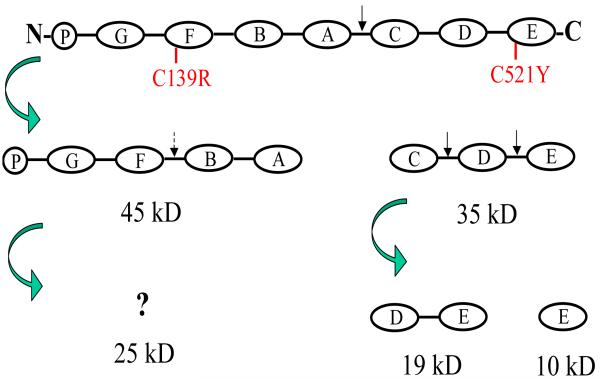
Our data suggest that elastase does not cleave PGRN at multiple sites to produce different GRNs simultaneously. Instead, it is a multi-step process. As elastase only cleaves PGRN at inter-linker regions, we propose the following elastase cleavage scheme for PGRN (Fig. 7): PGRN is initially cleaved, probably at the linker region between GRN A and GRN C (linker AC), to produce a 45 kD N-terminal intermediate and a 35 kD C-terminal intermediate. The 35 kD C-terminal intermediate is further cleaved, at both linker region CD and DE, to produce a 19 kD peptide (containing GRN D-E) and a 10 kD peptide (containing GRN E). The 45 kD intermediate is further cleaved to produce a 25 kD fragment. According to a previous study (Zhu et al., 2002), there are cleavage sites at linkers PG and GF, but none at linkers FB and BA. This cleavage scheme should produce a fragment containing GRN F-B-A. However, since the SDI antibody, which detects GRN A, did not recognize the 25 kD fragment (Supplemental Figure 3B), it is unresolved at this point where the cleavage occurs and what part of PGRN is present in the 25 kD fragment. Although not previously identified (Zhu et al., 2002), it is possible that another cleavage occurs at linker FB and the 25 kD fragment represents GRN P-G-F. Since P only contains half a GRN, the molecular weight of GRN P-G-F should be close to 25 kD.
Figure 7. A model of PGRN processing by elastase.
Only fragments detected in this study are shown. Full-length PGRN protein is initially cleaved by elastase at linker AC (solid arrow) to yield a 45 kD N-terminal intermediate fragment (encompassing GRNs P-G-F-B-A) and a 35 kD C-terminal intermediate fragment (encompassing GRNs C-D-E). The 35 kD intermediate fragment is further cleaved at linkers CD and DE (solid arrows) to yield a 19 kD fragment (GRNs D-E) and a 10 kD fragment (GRN E) that are detected with anti-PGRNC-term. The 45 kD intermediate fragment is further cleaved possibly at linker FB (dashed arrow), to produce a 25 kD fragment detected by anti-PGRNFL. A dashed arrow is used because the exact site of this cleavage is not confirmed in this study.
The C521Y mutation resulted in greatly reduced levels of all C-terminal fragments, most likely due to affecting both peptide stability and generation of the C-terminal fragments by proteolytic processing. Similarly, the C139R mutation dramatically decreased the amount of N-terminal fragments, but not the C-terminal fragments, indicating that this mutation affects the stability of the 45 kD intermediate.
A recombinant GRN E peptide harboring the C521Y mutation is functional in promoting neuronal survival and neurite outgrowth, suggesting that the mutation does not affect GRN E peptide function. Instead the mutation disrupts cleavage of PGRN by elastase, which affects the production of mature GRNs. The lower levels of GRNs as a result of reduced elastase cleavage could contribute to disease pathogenesis.
Hence the cysteine mutations affect both the activity of full-length PGRN and elastase cleavage of PGRN to GRNs. These results are consistent with either or both PGRN and GRNs contributing to neuronal development and neurodegeneration. While both PGRN and GRNs may be important in the nervous system, the simplest explanation of our data is that impaired PGRN activity causes FTLD-U because the two mutations reduce production of different GRNs. For lower GRN levels rather than lower PGRN levels to be the cause of the neurodegeneration it is necessary to postulate that both GRN F and GRN E affect neuronal survival. This is not impossible, but loss of PGRN activity provides a simpler explanation.
In summary, we provide evidence that two disease-causing mutations in GRN, which replace cysteine residues in PGRN, show normal levels of PGRN secretion in vivo and in vitro but show altered full-length PGRN function and abnormal protease cleavage to generate GRNs. Our data suggest that in addition to the levels of PGRN protein, the activity of full-length PGRN and cleavage of PGRN to generate mature GRNs are also essential for normal neuronal function and that perturbation of PGRN activity and/or GRN production could lead to neurodegeneration.
Supplementary Material
Acknowledgements
This work is supported by grants from the Alzheimer’s Association (IIRG-07-60299) and the Barnes-Jewish Foundation. JYW is supported by NIH grants EY014576, GM070967, CA107193. CC was a Government of Navarra Postdoctoral Fellow (2005-2007). The authors would like to thank Dr. Pau Pastor for providing PNFA family samples, Dr. Nupur Ghoshal for helpful discussions and Sumi Chakraverty for technical assistance.
References
- Baker M, Mackenzie IR, Pickering-Brown SM, et al. Mutations in progranulin cause tau-negative frontotemporal dementia linked to chromosome 17. Nature. 2006;442:916–919. doi: 10.1038/nature05016. [DOI] [PubMed] [Google Scholar]
- Bernardi L, Tomaino C, Anfossi M, et al. Novel PSEN1 and PGRN mutations in early-onset familial frontotemporal dementia. Neurobiol. Aging. 2009;30:1825–1833. doi: 10.1016/j.neurobiolaging.2008.01.005. [DOI] [PubMed] [Google Scholar]
- Brouwers N, Sleegers K, Engelborghs S, et al. Genetic variability in progranulin contributes to risk for clinically diagnosed Alzheimer disease. Neurology. 2008;71:656–664. doi: 10.1212/01.wnl.0000319688.89790.7a. [DOI] [PubMed] [Google Scholar]
- Cruchaga C, Fernandez-Seara MA, Seijo-Martinez M, et al. Cortical Atrophy and Language Network Reorganization Associated with a Novel Progranulin Mutation. Cereb. Cortex. 2009;19:1751–1760. doi: 10.1093/cercor/bhn202. [DOI] [PubMed] [Google Scholar]
- Cruts M, Gijselinck I, van der Zee J, et al. Null mutations in progranulin cause ubiquitin-positive frontotemporal dementia linked to chromosome 17q21. Nature. 2006;442:920–924. doi: 10.1038/nature05017. [DOI] [PubMed] [Google Scholar]
- Eriksen JL, Mackenzie IR. Progranulin: normal function and role in neurodegeneration. J. Neurochem. 2008;104:287–297. doi: 10.1111/j.1471-4159.2007.04968.x. [DOI] [PubMed] [Google Scholar]
- Finch N, Baker M, Crook R, et al. Plasma progranulin levels predict progranulin mutation status in frontotemporal dementia patients and asymptomatic family members. Brain. 2009;132:583–591. doi: 10.1093/brain/awn352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghidoni R, Benussi M, Glionna M, Franzoni M, Binetti G. Low plasma progranulin levels predict progranulin mutations in frontotemporal lobar degeneration. Neurology. 2008;71:1235–1239. doi: 10.1212/01.wnl.0000325058.10218.fc. [DOI] [PubMed] [Google Scholar]
- He Z, Bateman A. Progranulin (granulin-epithelin precursor, PC-cell-derived growth factor, acrogranin) mediates tissue repair and tumorigenesis. J. Mol. Med. 2003;81:600–612. doi: 10.1007/s00109-003-0474-3. [DOI] [PubMed] [Google Scholar]
- Hrabal R, Chen Z, James S, Bennett HP, Ni F. The hairpin stack fold, a novel protein architecture for a new family of protein growth factors. Nat. Struct. Biol. 1996;3:747–752. doi: 10.1038/nsb0996-747. [DOI] [PubMed] [Google Scholar]
- Hutton M, Lendon CL, Rizzu P, et al. Association of missense and 5′-splice-site mutations in tau with the inherited dementia FTDP-17. Nature. 1998;393:702–705. doi: 10.1038/31508. [DOI] [PubMed] [Google Scholar]
- Liu G, Beggs H, Jürgensen C, Park HT, Tang H, Gorski J, Jones KR, Reichardt LF, Wu J, Rao Y. Netrin requires focal adhesion kinase and Src family kinases for axon outgrowth and attraction. Nat. Neurosci. 2004;7:1222–1232. doi: 10.1038/nn1331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKhann GM, Albert MS, Grossman M, Miller B, Dickson D, Trojanowski JQ. Clinical and pathological diagnosis of frontotemporal dementia: report of the Work Group on Frontotemporal Dementia and Pick’s Disease. Arch. Neurol. 2001;58:1803–1809. doi: 10.1001/archneur.58.11.1803. [DOI] [PubMed] [Google Scholar]
- Mukherjee O, Pastor P, Cairns NJ, et al. HDDD2 is a familial frontotemporal lobar degeneration with ubiquitin-positive, tau-negative inclusions caused by a missense mutation in the signal peptide of progranulin. Ann. Neurol. 2006;60:314–322. doi: 10.1002/ana.20963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mukherjee O, Wang J, Gitcho M, et al. Molecular characterization of novel progranulin (GRN) mutations in frontotemporal dementia. Hum. Mutat. 2008;29:512–521. doi: 10.1002/humu.20681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muller PY, Janovjak H, Miserez AR, Dobbie Z. Processing of gene expression data generated by quantitative real-time RT-PCR. Biotechniques. 2002;32:1372–1374. 1376, 1378–1379. [PubMed] [Google Scholar]
- Poorkaj P, Bird TD, Wijsman E, Nemens E, Garruto RM, Anderson L, Andreadis A, Wiederholt WC, Raskind M, Schellenberg GD. Tau is a candidate gene for chromosome 17 frontotemporal dementia. Ann. Neurol. 1998;43:815–825. doi: 10.1002/ana.410430617. [DOI] [PubMed] [Google Scholar]
- Shankaran SS, Capell A, Hruscha AT, Fellerer K, Neumann M, Schmid B, Haass C. Missense mutations in the progranulin gene linked to frontotemporal lobar degeneration with ubiquitin-immunoreactive inclusions reduce progranulin production and secretion. J. Biol. Chem. 2008;283:1744–1753. doi: 10.1074/jbc.M705115200. [DOI] [PubMed] [Google Scholar]
- Spillantini MG, Murrell JR, Goedert M, Farlow MR, Klug A, Ghetti B. Mutation in the tau gene in familial multiple system tauopathy with presenile dementia. Proc. Natl. Acad. Sci. U S A. 1998;95:7737–7741. doi: 10.1073/pnas.95.13.7737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trojanowski JQ, Dickson D. Update on the neuropathological diagnosis of frontotemporal dementias. J. Neuropathol. Exp. Neurol. 2001;60:1123–1126. doi: 10.1093/jnen/60.12.1123. [DOI] [PubMed] [Google Scholar]
- Van Damme P, Bogaert E, Dewil M, et al. Astrocytes regulate GluR2 expression in motor neurons and their vulnerability to excitotoxicity. Proc. Natl. Acad. Sci. U S A. 2007;104:14825–14830. doi: 10.1073/pnas.0705046104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Damme P, Van Hoecke A, Lambrechts D, Vanacker P, Bogaert E, van Swieten J, Carmeliet P, Van Den Bosch L, Robberecht W. Progranulin functions as a neurotrophic factor to regulate neurite outgrowth and enhance neuronal survival. J. Cell Biol. 2008;181:37–41. doi: 10.1083/jcb.200712039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Den Bosch L, Storkebaum E, Vleminckx V, Moons L, Vanopdenbosch L, Scheveneels W, Carmeliet P, Robberecht W. Effects of vascular endothelial growth factor (VEGF) on motor neuron degeneration. Neurobiol. Dis. 2004;17:21–28. doi: 10.1016/j.nbd.2004.06.004. [DOI] [PubMed] [Google Scholar]
- Zhu J, Nathan C, Jin W, et al. Conversion of proepithelin to epithelins: roles of SLPI and elastase in host defense and wound repair. Cell. 2002;111:867–878. doi: 10.1016/s0092-8674(02)01141-8. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.