Abstract
PKR (protein kinase R) is induced by interferon and is a key component of the innate immunity antiviral pathway. Upon binding dsRNA or dimerization in the absence of dsRNA, PKR undergoes autophosphorylation at multiple serines and threonines that activate the kinase. Although it has previously been demonstrated that phosphorylation enhances PKR dimerization, gel filtration analysis reveals a second monomeric phosphorylated form. These forms are referred to as phosphorylated dimeric PKR (pPKRd) and phosphorylated monomeric PKR (pPKRm). These two forms do not reversibly interconvert. Sedimentation equilibrium measurements reveal that pPKRm dimerizes weakly with a Kd similar to unphosphorylated PKR. Isoelectric focusing and mass spectrometry demonstrate that both pPKRm and pPKRd are heterogeneous in their phosphorylation states, with an average of 9-10 phosphates. Equilibrium chemical denaturation analysis indicates that phosphorylation destabilizes the kinase domain by about 1.5 kcal/mol in the dimeric form but not in the monomeric form. Limited proteolysis also reveals that phosphorylation induces a conformational change in pPKRd that is not detected in pPKRm. pPKRm binds dsRNA with similar affinity as unphosphorylated PKR whereas binding cannot be detected with pPKRd. Despite these substantial differences in biophysical properties, both pPKRm and pPKRd are catalytically competent and are activated to phosphorylate the PKR substrate eIF2α in the absence of dsRNA. Thus, both monomeric and dimeric forms of phosphorylated PKR may participate in the interferon antiviral pathway.
Protein kinase R (PKR) is an interferon-induced kinase that plays a key role in the innate immunity response to viral infection (1, 2). PKR also functions in the control of cell growth and proliferation and has been implicated as a tumor suppressor protein (3, 4). The form of PKR induced by interferon is not functional but it is activated by binding dsRNA to undergo autophosphorylation. The most well characterized cellular substrate of PKR is the alpha subunit of eukaryotic initiation factor eIF2. Phosphorylation of eIF2α at serine 51 blocks the recycling of eIF2 between GTP- and GDP-bound states, thereby inhibiting translation initiation. Thus, production of dsRNA during viral infection (5) results in PKR activation and inhibition of viral and host protein synthesis.
PKR contains two tandem copies of the dsRNA binding motif (6) at the N-terminus and a C-terminal kinase, with a ~90 amino acid linker lying between these domains. Each of the dsRNA binding motifs has the typical αβββα fold (7) with a short unstructured region between the folded regions. In the crystal structure of a complex of the PKR kinase domain with an N-terminal fragment of eIF2α, the catalytic domain adopts a bilobal structure typical of protein kinases (8). The kinase forms a dimer mediated by interactions between residues within the C-terminal lobes. The large linker lying between the dsRNA binding motifs and the kinase is unstructured (9) and small angle scattering measurements indicate that the enzyme is flexible in solution and adopts multiple conformations (10).
Although a variety of molecular models for the activation of PKR have been put forth, a consensus has emerged that dimerization plays a key role in the activation mechanism (11). A defining feature of PKR is the “bell-shaped” curve for activation, where low concentrations of dsRNA activate but higher concentrations are inhibitory (12, 13). These data have generally been interpreted to indicate that low concentrations of dsRNA drive assembly of PKR dimers on a single dsRNA whereas higher dsRNA concentrations dilute PKR monomers onto separate molecules of dsRNA (14). Although PKR can bind to dsRNAs as short as 15 bp (15-17), a minimum of 30 bp is required for activation (13). The ability of dsRNAs to function as PKR activators is correlated with binding of two or more PKR monomers (18), supporting a model where the role of the dsRNA is to bring two or more PKR monomers in close proximity to enhance dimerization via the kinase domain. PKR alone exists in a weak monomer-dimer equilibrium and dimerization of PKR in the absence of dsRNA is sufficient to activate PKR to undergo autophosphorylation (19).
PKR autophosphorylates at multiple serine and threonine sites in a highly complex process. The activity of protein kinases is often modulated by autophosphorylation of one or more residues within the activation loop that lies outside of the catalytic site (20, 21). In the PKR activation loop, Thr 446 is phosphorylated in vivo and in vitro (22, 23). Mutation of either Thr 446 or 451 to Ala impairs PKR activity (23). However, phosphorylation of Thr 451 has not been detected by mass spectrometry. In addition to the activation loop, three phosphorylation sites (Ser 252, Thr 255 and Thr 258) lie within the linker region between the dsRNA binding domain and kinase (24) and one (Thr 33) lies within the first dsRNA binding domain (22). Six phosphorylation sites have been localized to the region between the first and second dsRNA binding domain (22, 25) and another cluster of six sites exists in the kinase insert region between kinase subdomains IV and V (22). Overall, about 15 serines and threonines are believed to be phosphorylated in PKR (22, 26). Outside of the activation loop, the only phosphorylation site that appears functionally important is Thr 258 (24); however, others report that only Thr 446 and Thr 451 were critical for PKR activity in yeast (26). In addition to these serine and threonine sites, it has been reported that PKR also autophosphorylates at three tyrosines (27).
The structural and functional consequences of PKR autophosphorylation are not well understood. Once autophosphorylated, PKR is constitutively activated and is only minimally stimulated by further addition of dsRNA (14, 28). PKR autophosphorylation reduces the binding affinity for dsRNA (9, 29). In the absence of dsRNA, PKR dimerizes weakly, with Kd ~ 500 μM (19). Remarkably, dimer stability is enhanced (30) by ~500-fold upon autophosphorylation (19). In the crystal structure of the kinase domain, phosphothreonine 446 interacts with basic residues within helix αC which in turn interacts with the substrate binding site and forms a portion of the dimer interface, suggesting a possible mechanism linking dimerization to enzymatic activation (8). Autophosphorylation induces a conformation change in the activation loop that blocks trypsin proteolysis and slightly reduces the thermodynamic stability of the kinase domain (31). NMR chemical shift changes observed upon PKR phosphorylation are localized within the kinase domain (9).
In the present study we have investigated the biophysical properties of autophosphorylated PKR. Gel filtration analysis reveals that phosphorylated PKR can be separated into two distinct forms: one that dimerizes strongly and one that dimerizes weakly with affinity similar to unphosphorylated enzyme. We characterize the biophysical properties of these two forms of phosphorylated PKR and correlate them with enzymatic activity. Our results indicate that both forms are catalytically competent and are activated to phosphorylate the PKR substrate eIF2α in the absence of dsRNA. Thus, both monomeric and dimeric forms of phosphorylated PKR may participate in the interferon antiviral pathway.
Materials and Methods
Expression and purification of unphosphorylated PKR was performed as previously described (19). PKR undergoes dsRNA-independent autophosphorylation upon incubation at protein concentrations greater than 0.5 μM in the presence of ATP (19) and this reaction was used to generate phosphorylated enzyme for biophysical analysis. PKR was exchanged into 50 mM Tris, 100 mM NaCl, 12 mM MgCl2, 0.1 mM EDTA, 10 mM BME, pH 8.0 using either disposable spin columns containing Biogel P6 (BioRad), or dialysis, and concentrated to 80 μM. The reaction was initiated by addition of 6 mM ATP and proceeded at room temperature for 4 hours. The resulting material was centrifuged at 15,000 g for 10 min. to remove aggregates that form during phosphorylation and applied onto a Superdex 200 16/60 preparative gel filtration column (GE Healthcare) equilibrated in AU200 buffer (20 mM HEPES, 200 mM NaCl, 0.1 mM EDTA, 0.1 mM TCEP, pH 7.5). Two fractions were isolated, with ~80% of the material eluting at the volume corresponding to PKR monomer and ~20% eluting at a volume corresponding to dimer. These fractions are designed pPKRm and pPKRd, respectively.
The homogeneity of pPKRm and pPKRd phosphorylation was analyzed by isoelectric focusing using pH 3-7 gels (Invitrogen) and IEF gel standards with pIs ranging from 3.5-10.7 (Serva). Based on its amino acid composition, unphosphorylated PKR is predicted by the program SEDNTERP (32) to have a pI of 8.76 and does not enter the gel. Control experiments using pH 3-10 gels confirmed that the unphosphorylated enzyme has a pI between 8 and 9 (data not shown). The phosphorylation states of pPKRm and pPKRd were analyzed by LC-MS electrospray mass spectrometry. Samples were prepared by buffer exchange into a solution containing 0.5% acetic acid and 50% methanol. HPLC was performed using a 10ADvp system (Shimadzu, Columbia, MD) and a reversed-phase column (Hypersil GOLD, 1.9 μm, 100 mm × 1.0 mm, Thermo Scientific, Waltham, MA). Solvent A was formic acid/acetonitrile/H2O [2:10:988 (v/v/v)] and solvent B was formic acid/H2O/acetonitrile [2:10:988 (v/v/v)]. Electrospray mass spectrometry was performed on a QSTAR Elite (Applied Biosystems, Foster City, CA). The deconvoluted spectra were obtained by Bayesian Protein Reconstruction in Analyst QS 2.0.
Equilibrium chemical denaturation experiments were performed and analyzed as previously described (31) using CD to detect global unfolding and tryptophan fluorescence to selectively monitor the kinase domain. The data were fit using the program Savuka (33, 34). Limited trypsin proteolysis measurements were performed as previously described (31). Sedimentation velocity analysis was performed at 20°C and 50,000 RPM using absorbance optics at 260 nm with a Beckman-Coulter XL-I analytical ultracentrifuge. Standard two-channel cells were used with quartz windows. Initial analysis was performed using DCDT+ (35) to obtain g(s*) distributions. Multiple datasets were globally fit to hetero-association models using SEDANAL (36). Sedimentation equilibrium experiments were performed at 20°C and rotor speeds of 18,000 and 24,000 RPM using interference optics. Samples were loaded into aged, six-channel external loading cells equipped with sapphire windows (37). Water blanks were taken before and after the run at each speed. The match routine in the program HETEROANALYSIS was used to determine when equilibrium was established. The data was analyzed using HETEROANALYSIS (38). Extinction coefficients, molecular masses, partial specific volumes and solvent densities were calculated using Sednterp (39).
Enzymatic activity assays were performed at 32°C in 20 mM HEPES, 50 mM KCl, 5 mM MgCl2, 0.1 mM TCEP, pH 7.5. A construct encoding residues 3-183 of the PKR substrate eIF2α cloned into pET-30a, which provides a C-terminal histidine tag. eIF2α was purified by metal ion affinity chromatography on His-trap FF and gel filtration on Superdex 75 (GE Healthcare). Phosphorylation reactions contained 100 nM PKR, 385 nM eIF2α, variable concentrations of poly (rI): poly (rC) dsRNA (GE Healthcare) and 40-400 μM ATP containing 1 μCi [γ-32P] ATP/sample. The reaction was quenched by addition of SDS-PAGE loading buffer and heating at 70°C for 10 minutes. The samples were run on 12% SDS-PAGE polyacrylamide gels (Invitrogen) for 75 min at 200V. Incorporation of 32P into PKR and eIF2α was visualized and quantitated using a PMI phosphorimager (BioRad).
Results
Detection of a monomeric form of phosphorylated PKR
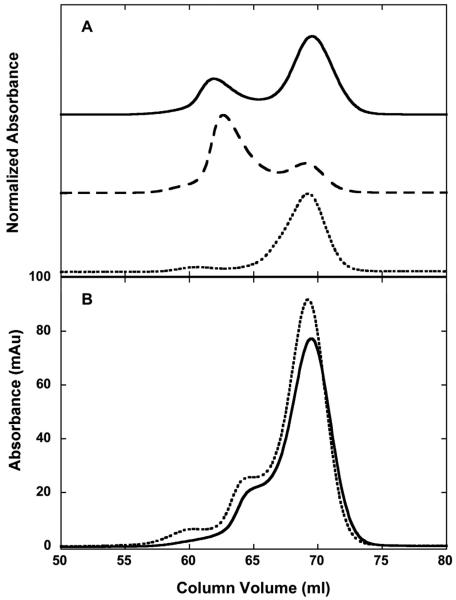
PKR was autophosphorylated in the absence of dsRNA by incubation at high protein concentration in the presence of ATP. Figure 1 shows that two forms are resolved by gel filtration. The larger peak has the same elution volume as monomeric PKR whereas the smaller feature corresponds to a PKR dimer (19). Previously, we demonstrated that this PKR dimer fraction is phosphorylated and characterized its dimerization affinity, functional activity and stability (19, 31). Data shown below demonstrates that the larger peak at 70 mL also corresponds to a phosphorylated form of PKR and not to residual unphosphorylated enzyme. Here, we characterize this monomeric, phosphorylated fraction and compare its properties to the unphosphorylated and dimeric phosphorylated forms of PKR.
Figure 1.
Gel filtration analysis of phosphorylated PKR. A) Overplot of initial reaction mixture (solid) and reapplication of purified pPKRd (dashed) and pPKRm (dotted) . The pPKRm and pPKRd fractions were isolated by preparative gel filtration, concentrated to about 5 mg/ml and equilibrated for 5 days at 4°C. The chromatograms are vertically offset for clarity. B) Overplot of pPKRm (solid) and pPKRm after re-reaction with ATP (dotted). pPKRm isolated from initial preparative gel filtration was concentrated to 80 μM and equilibrated into 50 mM Tris, 100 mM NaCl, 12 mM MgCl2, 0.1 mM EDTA, 10 mM BME, pH 8.0. Half of the sample was incubated with ATP and MgCl for 4 hours at 20°C. Both samples were separately reapplied to the column. Analytical gel filtration chromatography was performed on a GE Healthcare Superdex 200 10/300 column in AU200 buffer at 0.5 ml/min and 4°C.
There are two alternative explanations for the monomer and dimer forms of PKR detected in gel filtration: 1) two distinct molecular components that do not interconvert, or 2) two species representing a single component engaged in a reversible monomer-dimer equilibrium where the rate of interconversion is slow on the timescale of the experiment. To discriminate between these possibilities, each fraction was collected, reconcentrated to the original concentration, allowed to equilibrate for five days and reapplied to the Superdex 200 column (Figure 1A). The dimer fraction gives rise to a predominant dimer peak with a smaller monomer feature. Given the limited resolution of gel filtration, it is not possible to baseline-separate the two forms and presence of a monomer peak in the dimer fraction is at least partially associated with cross-contamination with monomer. In addition, we have previously used analytical ultracentrifugation to demonstrate that the fraction isolated from the dimer peak itself exists in a reversible monomer monomer-dimer equilibrium with Kd = 0.95 μM (19). Thus, the dimer fraction will produce a peak corresponding to the monomer due to dissociation during the dilution that occurs upon gel filtration. In sedimentation velocity experiments, where dilution is minimal, a monomer peak near 3.5 S increases in area as the sample concentration decreases from 0.6 to 0.1 mg/ml (Figure S1). However, at the highest concentration the monomer peak is less than 5% of the total area. Like the dimer fraction, the monomeric fraction retains its chromatographic behavior after prolonged incubation and elutes as a monomer with a trace of dimer (Figure 1A). The amplitude of the dimer peak corresponds to about 6% of the total area. Similarly, sedimentation equilibrium analysis (vide infra) indicates about a 5% dimer contaminant. Thus, in summary, the monomer and dimer fractions isolated by gel filtration represent distinct molecular components with a low level of cross-contamination. The monomeric fraction will be referred to as pPKRm and the dimer fraction as pPKRd. However, as demonstrated below, each of these components engages in reversible dimerization with pPKRm exhibiting weak self-association and pPKRd forming a stronger dimer.
Analysis of phosphorylation states of PKR
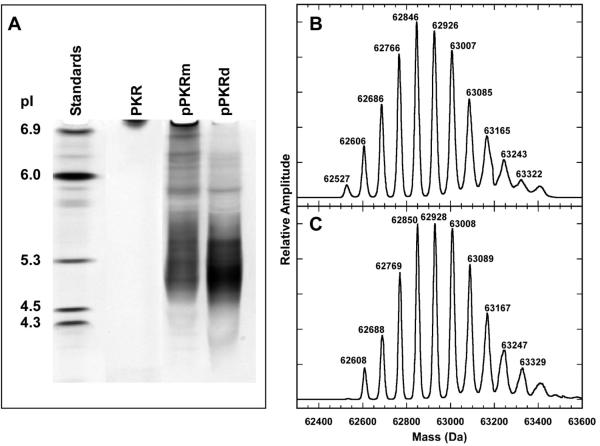
The phosphorylation states of pPKRm and pPKRd were evaluated by isoelectric focusing and mass spectrometry. Based on amino acid composition, the isoelectric point of unphosphorylated PKR is predicted be 8.76 and addition of acidic phosphate groups is expected to reduce the pI. In isoelectric focusing carried out with a pH range of 3-7, unphosphorylated PKR essentially does not enter the gel and produces a band at the well (Figure 2A). In contrast, both phosphorylated forms give rise to a broad range of bands pI~ 6– 4.5 with the most intense bands centered at pI~5. Thus, pPKRm and pPKRd are considerably more acidic than unphosphorylated PKR, consistent with a high degree of phosphorylation. The average level of phosphorylation appears to be slightly higher in pPKRd than pPKRm. In addition the pPKRm band has a fraction that does not migrate into the gel, possibly indicating the presence of some portion of this fraction that has little or no phosphorylation. However, it is not possible to distinguish aggregate from unphosphorylated PKR in this gel system, and this material may correspond to an aggregate.
Figure 2.
Analysis of pPKRm and pPKRd phosphorylation states. A) IEF gel of unphosphorylated and phosphorylated PKR forms. Samples were loaded at a concentration of 3 mg/ml and analyzed on a pH 3-7 IEF gel (Invitrogen ) with SERVA markers according to the manufacturers' instructions. Gels were stained with Coomassie blue. B) Deconvoluted electrospray ionization mass spectrum of pPKRm. C) Deconvoluted electrospray ionization mass spectrum of pPKRd. See Materials and Methods for experimental details.
Electrospray ionization mass spectrometry of intact proteins was used to characterize the phosphorylation states of the pPKRm and pPKRd. A series of peaks are detected in the deconvoluted mass spectrum of pPKRm that range from 62,527 Da to above 63,222 Da (Figure 2B). The interval between successive peaks is 80 Da, corresponding to single phosphate group. Based on the mass of unphosphorylated PKR of 62,095 Da (19), pPKRm consists of a distribution corresponding to 6-16 phosphates/PKR with the most populated form containing 9 phosphates. A similar distribution is observed for pPKRd (Figure 2C) but the most populated form is slightly shifted to 10 phosphates / PKR. These data confirm the IEF results and demonstrate that both pPKRm and pPKRd are both heterogeneously phosphorylated and show that pPKRd is slightly more heavily phosphorylated than pPKRm. The IEF analysis appears to resolve a larger number of phosphorylated forms than mass spectrometry. Given the presence of as many as 15 potential phosphorylation sites in PKR, it is likely each peak in Figure 2B actually consists of a population contain the same total number of phosphates distributed on different sites in the protein. These forms would likely have different pI values, giving rise to additional bands in the IEF analysis. We do not detect any unphosphorylated material in the mass spectrum of pPKRm. We have observed small quantities of unphosphorylated PKR in some preparations of pPKRm by IEF analysis on pH 7-10 gels (data not shown).
The heterogeneity in phosphorylation states is not kinetically controlled. Control experiment using trace quantities of [γ32P] ATP to quantitate PKR labeling demonstrate that the extent of PKR labeling is saturated over the four hour time interval used to prepare phosphorylated PKR (C. Quartararo, E. Anderson and J. Cole, unpublished observations). Given the large number of phosphates incorporated into PKR, it has not proven feasible to separate the individual phosphorylation states present in either pPKRm or pPKRd by ion exchange chromatography and related methods (E. Anderson and J. Cole, unpublished observations).
We considered the possibility that pPKRm represents an intermediate in the formation of pPKRd. After isolation of pPKRm from gel filtration, the protein was concentrated, re-equilibrated into the phosphorylation buffer and was incubated in the presence of ATP under the same conditions that we used originally to prepare the phosphorylated protein. Figure 1B shows that this material coelutes on gel filtration with pPKRm that was not subject to a rephosphorylation, with the major peak eluting at a volume corresponding to PKR monomer. The formation of a shoulder eluting at 60 mL likely corresponds to formation of aggregate. These data indicate that pPKRm cannot be converting into pPKRd by additional phosphorylation and thus does not represent an intermediate in the formation of pPKRd. Similar to the human enzyme, mouse PKR also forms pPKRm and pPKRd upon in vitro autophosphorylation, demonstrating that these forms are not specific to the human enzyme (Brown R. and Cole, J., unpublished observations). Monomeric and dimeric phosphorylated forms of human PKR are also produced upon dsRNA-dependent autophosphorylation of PKR at lower protein concentration (0.25 μM) the presence of poly (rI): poly (rC) dsRNA (Figure S1). IEF analysis of the monomeric fraction reveals that the distribution of phosphorylation states is similar to pPKRm and pPKRd generated in the absence of dsRNA. Thus, production of both pPKRm and pPKRd occurs with multiple PKR constructs and is independent of the mechanism used to induce autophosphorylation. This heterogeneity in phosphorylated PKR may be physiologically significant. Thus, we have characterized the biophysical and enzymatic properties of pPKRm and compare the properties of this form with previous studies of pPKRd (9, 18, 19, 29, 31).
Stability and conformation of pPKRm
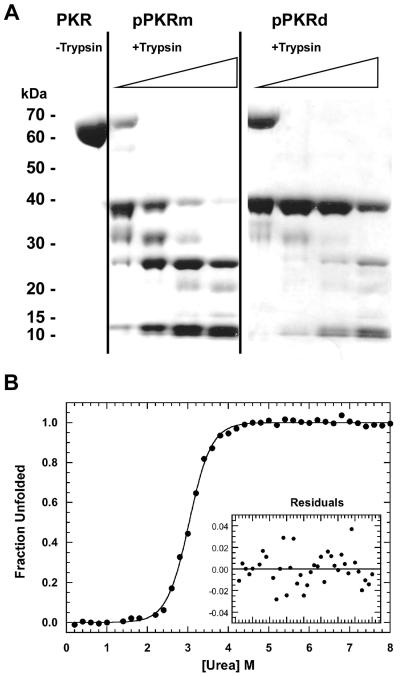
We have previously used limited proteolysis to define the accessible and unstructured regions of PKR and to monitor conformational changes associated with ligand binding and activation (31). Figure 3A compares the trypsin proteolysis patterns of pPKRm and pPKRd. At the lowest protease concentration, both phosphorylated forms are cleaved to generate peptides of about 40 and 30 kDa. This cleavage occurs near the N terminus of the kinase domain and the 40 kDa peptide corresponds to the kinase domain (31). In the case of pPKRm, at higher [trypsin], the 40 kDa kinase domain peptide is cleaved and additional bands appear at 25 kDa and 11 kDa. This trypsin proteolysis pattern is similar to unphosphorylated PKR where the second cleavage occurs within the activation loop (31). In contrast, as we previously reported, the 40 kDa band in pPKRd is dramatically stabilized to cleavage by trypsin (31).
Figure 3.
Analysis of the stability of phosphorylated PKR. A) Limited proteolysis of pPKRm and pPKRd. Samples were incubated at 1 mg/ml at several ratios of PKR: trypsin at 20 °C for 30 minutes. Reactions were quenched by the addition of SDS sample loading buffer and heating to 90°C for 10 minutes. The samples were run on a 4-12% acrylamide bis-tris gel under denaturing conditions and visualized with Coomassie blue staining. Lane 1 contains unphosphorylated PKR that has not been digested. Lanes 2-5 contain pPKRm with increasing concentrations of trypsin. Lanes 6-9 contain pPKRd and increasing concentrations of trypsin. B) Analysis of the stability of pPKRm by equilibrium urea unfolding titration. pPKRm was prepared at 0.15 mg/ml and variable urea concentrations and equilibrated at 20°C for 3 hours. Unfolding was monitored by tryptophan fluorescence emission with excitation at 295 and emission at 340 nm. The data are shown as filled circles and the fit to a two-state model using SAVUKA is shown as a solid line. The best-fit parameters are ΔG° = 6.41 ± 0.31 kcal mol−1 and m = 2.11 ± 0.01 kcal mol−1 M−1. Inset: residuals of the fit.
PKR contains three tryptophans, all localized within the kinase domain, and we have previously used intrinsic tryptophan fluorescence to selectively probe the stability of the kinase domain and CD to measure global stability (31). Urea denaturation measurements indicated that unfolding of unphosphorylated PKR monitored by tryptophan fluorescence occurs as a single transition with ΔG° = 6.66 ± 0.53 kcal/mol. Figure 3B shows that the unfolding of pPKRm monitored by tryptophan fluorescence also occurs as a single transition with ΔG° = 6.41 ± 0.31 kcal/mol. Thus, phosphorylation to form pPKRm does not perturb the stability of the PKR kinase domain. In contrast, the stability of the kinase domain in pPKRd is reduced by about 1 kcal/mol to 5.45 ± 0.28 kcal/mol (31).
We have also compared the CD spectra of PKR, pPKRm and pPKRd to detect potential differences in secondary structure and conformation. The spectra of the two phosphorylated forms overlay with latent PKR, indicating the absence of significant differences in secondary structure (data not shown).
Dimerization of pPKRm
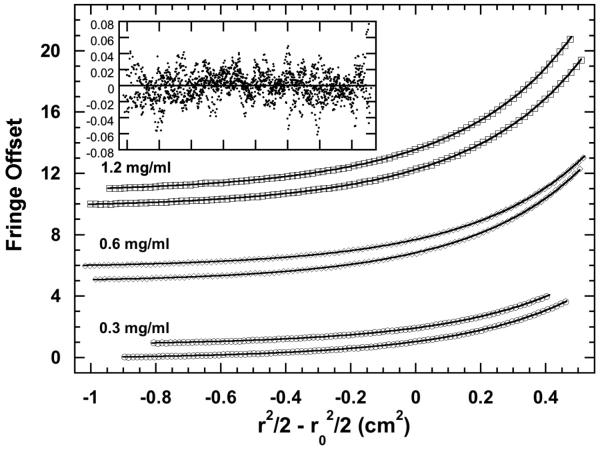
Latent PKR exists in a weak monomer-dimer equilibrium with Kd = 450 μM and formation of pPKRd results in 500-fold enhanced dimerization affinity such that Kd = 0.95 μM. Here, we define the self-association properties of pPKRm. In our preliminary analysis of sedimentation equilibrium data obtained with pPKRm using a monomer-dimer model we found that it was necessary to incorporate the presence of incompetent dimer to obtain satisfactory fits. In this model, the incompetent dimer corresponds to a nonequilibrating component with a mass corresponding to a dimer of pPKRm. As shown in figure 1, it is not possible to baseline-separate the two phosphorylated forms of PKR and the dimer contaminant found in our preparation of pPKRm is likely pPKRd. Figure 4 show a fit of the pPKRm sedimentation equilibrium to a monomer-dimer model with Kd = 473 (377, 624) μM incorporating 5% incompetent dimer (the values in parentheses refer to the 1 standard deviation joint confidence intervals). Thus, the Kd of pPKRm is similar to unphosphorylated enzyme and is significantly lower than pPKRd.
Figure 4.
Sedimentation equilibrium analysis of pPKRm dimerization. The open symbols are the data and the solid lines are a global fit using a monomer-dimer equilibrium model incorporating incompetent dimer. For clarity, only every third data point is shown. The inset shows a superposition of the residuals from the global fit. The best-fit parameters are: Kd = 473 (377, 624) μM with 5% incompetent dimer and RMS = 0.0158 fringes.
Binding of pPKRm and pPKRd to dsRNA
Sedimentation velocity analytical ultracentrifugation monitored by UV absorbance at 260 nm was used to define the dsRNA binding affinity of monomeric and dimeric phosphorylated PKR. We have previously demonstrated that PKR binds to a 20 bp dsRNA with Kd = 860 nM (18). We were unable to detect any binding of pPKRd to the same dsRNA under similar conditions (data not shown). These results are consistent with previous observations indicating that phosphorylation of PKR interferes with RNA binding (9, 29). In contrast, pPKRm bound readily as revealed in an overlay of normalized g(s*) sedimentation coefficient distribution functions for a titration of the 20 bp dsRNA with pPKRm (Figure S2). The apparent sedimentation coefficient increases continuously upon addition of pPKRm from about 2.5 S to about 4.5 S in the presence of a 6-fold molar excess of protein. This behavior is consistent with a simple 1:1 binding model. We previously reported a similar shift upon binding of unphosphorylated PKR to the same dsRNA (18). We globally fit these data using the program SEDANAL with this model to extract Kd. Figure 5 shows that the data fit very well as indicated by minimal systematic deviations in the residuals and a low RMS deviation of 0.0071 OD. The best fit Kd = 3.10 (1.67, 5.49) μM. Thus, pPKRm binds to dsRNA about 3-4 fold weaker than unphosphorylated enzyme.
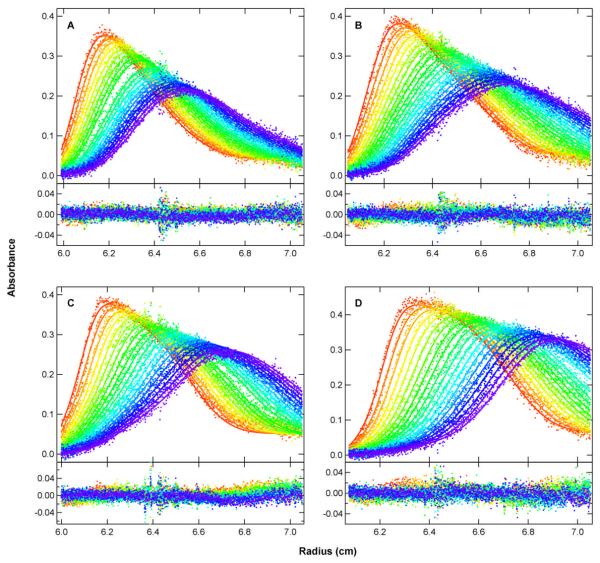
Figure 5.
Sedimentation velocity analysis of pPKRm binding to a 20 bp dsRNA. Samples were prepared at 1 μM dsRNA + 0.5 eq. pPKRm (A), dsRNA + 1 eq. pPKRm (B), dsRNA +2 eq. pPKRm (C) and dsRNA + 6 eq. pPKRm (D). The sedimentation velocity profiles were subtracted in pairs to remove time-invariant noise and the four data sets were globally fit to a 1:1 binding stoichiometry model using SEDANAL (36). The top panels show the data (points) and fit (solid lines) and the bottom panels show the residuals (points). The best-fit parameters are: Kd = 3.10 (1.67, 5.49) μM and RMS deviations = 0.0071 OD.
Functional activity of pPKRm and pPKRd
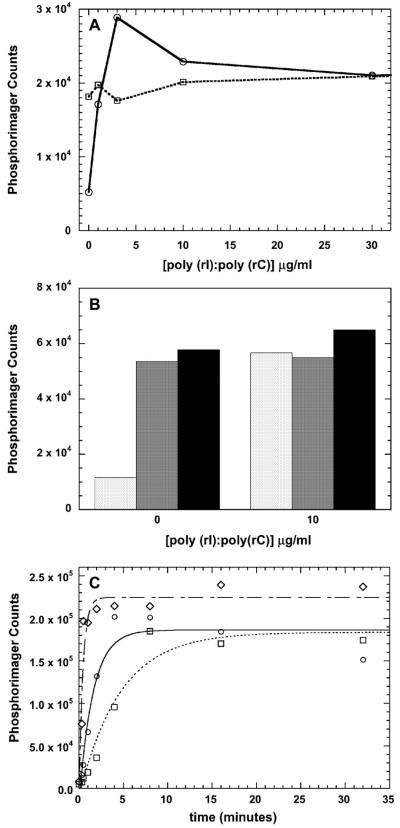
PKR is activated upon binding to dsRNA to undergo autophosphorylation and subsequently phosphorylate the physiological substrate, eIF2α. Typically, these reactions are assayed by monitoring the transfer of 32P from [γ-32P]ATP into PKR and eIF2α. We have used this assay to measure the ability of pPKRm and pPKRd prepared using unlabeled ATP to subsequently catalyze phosphorylation eIF2α with [γ-32P] ATP. In control experiments, the ability of unphosphorylated PKR to phosphorylate eIF2α is dependent on dsRNA (Figure 6A). Interestingly, pPKRm is also capable of catalyzing phosphorylation of eIF2α. Although pPKRm binds dsRNA (Figure 5), eIF2α phosphorylation does not require dsRNA and is essentially constant up to a concentration of 30 μg/ml (Figure 6A).
Figure 6.
eIF2α kinase activity of pPKRm and pPKRd. A) Effects of dsRNA on eIF2α phosphorylation. Unphosphorylated PKR control ( ) and pPKRm (
) and pPKRm ( ). B) Comparison of unphosphorylated PKR (white), pPKRm (grey) and pPKRd (black). For both A and B, samples contained 400 μM ATP and were quenched after 20 minutes. C) Kinetics of eIF2α phosphorylation catalyzed by unphosphorylated PKR (
). B) Comparison of unphosphorylated PKR (white), pPKRm (grey) and pPKRd (black). For both A and B, samples contained 400 μM ATP and were quenched after 20 minutes. C) Kinetics of eIF2α phosphorylation catalyzed by unphosphorylated PKR ( ), pPKRm (
), pPKRm ( ) and pPKRd (
) and pPKRd ( ). All samples contained 40 μM ATP and the unphosphorylated PKR sample contained 10 μg/ml dsRNA. The best fit line for a single exponential fit is shown for each reaction, with rate constants of 0.64 min−1 (PKR), 0.22 min−1 (pPKRm) and 2.2 min−1 (pPKRd).
). All samples contained 40 μM ATP and the unphosphorylated PKR sample contained 10 μg/ml dsRNA. The best fit line for a single exponential fit is shown for each reaction, with rate constants of 0.64 min−1 (PKR), 0.22 min−1 (pPKRm) and 2.2 min−1 (pPKRd).
Several lines of evidence indicate that the eIF2α kinase activity of pPKRm is not due to pPKRd contamination. First, we have consistently observed that all preparations of pPKRm are able to phosphorylate eIF2α with similar rates, which suggests that this reaction is not due to contamination. Second, additional data from the experiment depicted in Figure 6B that strongly argue that both phosphorylated forms are active. Even though pPKRm and pPKRd are produced in a reaction using unlabeled ATP, we have consistently observed that both of these preparations also undergo low levels of autophosphorylation upon subsequent incubation with γ-32P-ATP. Presumably, this reaction occurs because of the incomplete incorporation of phosphates in pPKRm and pPKRd as indicated by the IEF and mass spectrometry analysis in Figure 2. We measured the extent of PKR autophosphorylation in the experiment in figure 6B in addition to the phosphorylation of eIF2α. Importantly, the amount of 32P incorporated into pPKRm is 92% of the amount incorporated into pPKRd. If the activity of pPKRm were associated with contaminating pPKRd, this amount of 32P incorporation would be much lower, on the order of 5%.
pPKRd is capable of phosphorylating eIF2α (Figure 6B). pPKRm and pPKRd phosphorylate eIF2α to the same extent independent of the presence of dsRNA. Under the same conditions, complete phosphorylation of the substrate by PKR requires dsRNA. Although both pPKRm and pPKRd are active enzymes, the kinetics of eIF2α phosphorylation are different (figure 6C). Under the same conditions, the reaction catalyzed by pPKRd is about 10-fold faster than pPKRm. In the presence of saturating dsRNA, the rate of eIF2α phosphorylation catalyzed by PKR is intermediate between the phosphorylated enzymes. These results demonstrate that both pPKRm and pPKRd are both catalytically competent and do not require dsRNA to trans-phosphorylate substrate.
Discussion
We (19) and others (9, 30) have previously reported that autophosphorylation of PKR results in enhancement of dimerization affinity. In these studies, the phosphorylated enzyme was purified by collecting the fraction eluting from a gel filtration column as a dimer (pPKRd). However, biophysical analysis of the monomer fraction reveals that component is also phosphorylated and we refer to this form as pPKRm. These two forms do not reversibly interconvert and thus represent distinct molecular components. pPKRm cannot be converted into pPKRd upon additional phosphorylation, suggesting that it does not represent an intermediate in the activation process. Furthermore, both forms are generated upon activation of human PKR in the absence and presence of dsRNA as well as upon activation of mouse PKR, suggesting this heterogeneity in self-association affinity is an intrinsic property of the activated enzyme. Based on these observations, we have undertaken to characterize the biophysical and functional properties of pPKRm to compare with previous analysis of pPKRd (9, 18, 19, 29, 31).
IEF and mass spectrometry indicate that both pPKRm and pPKRd are heterogeneously phosphorylated, with each containing a range of about 6-16 phosphates with a maximum population near 9 phosphates/ enzyme for pPKRm and 10 for pPKRd. Thus, the overall level of phosphorylation is similar for the forms. The maximal number of phosphates we observe is close to the 14-15 phosphorylation sites previously detected in PKR (22, 26). The heterogeneity in phosphorylation state was previously observed in IEF gels of wild-type PKR expressed in E. coli in the absence of a phosphatase (40). In contrast, NMR analysis of phosphorylated PKR (presumably pPKRd) suggested a homogeneous population (9); however, this preparation was not characterized by methods that directly probe the distribution of phosphorylation states. The origin of the heterogeneity in phosphate incorporation in pPKRm and pPKRd is not known. The autophosphorylation reaction reaches a plateau level, suggesting that phosphatate incorporation is not kinetically controlled. There is no apparent consensus sequence flanking the serine and threonine phosphorylation sites in PKR. Possibly, some of them are poorly recognized by the kinase and thus only rarely phosphorylated, resulting in an incompletely phosphorylated product.
Although both pPKRm and pPKRd exhibit a similar overall level of phosphorylation they have distinct biophysical properties. The sensitivity of pPKRm to limited trypsin proteolysis and the thermodynamic stability of the kinase domain are similar to unphosphorylated PKR. In contrast, the activation loop in pPKRd is stabilized against proteolysis, suggesting that it adopts a more ordered conformation upon phosphorylation (31). The stability of the kinase domain is reduced in pPKRd (31). pPKRm exists in a monomer dimer equilibrium with a dissociation constant that is similar to unphosphorylated enzyme whereas the dimer affinity is enhanced about 500-fold in pPKRd (19). Finally, pPKRm binds to a 20 bp dsRNA about 3-4 fold weaker than unphosphorylated PKR whereas binding is undetectable for pPKRd. Thus, with the exception of the greatly decreased pI induced by phosphorylation, the biophysical properties of pPKRm are very similar to the unphosphorylated enzyme and differ dramatically from pPKRd.
Despite the considerable differences in the biophysical properties of pPKRm and pPKRd, both forms are functional and are capable of phosphorylating eIF2α. Both pPKRm and pPKRd are constitutively activated and are not stimulated by addition of dsRNA, as has previously been reported for activated forms of PKR (14, 28). Thus, the conformational changes and enhanced dimerization in pPKRd are not required for activating PKR. Both pPKRm and pPKRd are produced under a variety of conditions and our results suggest that both monomeric and dimeric forms of phosphorylated PKR may be physiologically relevant and participate in the interferon antiviral pathway. PKR isolated from the cytosolic fraction from interferon-induced cells exists predominantly as a dimer whereas PKR isolated from a ribosomal salt wash is mostly dimeric (30). Both forms are partially phosphorylated, suggesting that pPKRm and pPKRd may exist in vivo. One difference between the two activated forms is that pPKRd phosphorylates eIF2α more rapidly than pPKRm. This difference may be associated with the enhanced dimerization of pPKRd. The heterogeneity in PKR phosphorylation may serve to confound viral countermeasures to evade PKR.
Supplementary Material
Acknowledgement
We thank Carolyn Teschke for use of the PMI phosphorimager, Gerhard Wagner for providing a plasmid that encodes human eIF2α and the reviewers for useful comments.
Glossary
1Abbreviations
- Bis-tris
Bis(2-hydroxyethyl)-imino-tris(hydroxymethyl)-methane
- BME
β mercaptoethanol
- EDTA
ethylenediaminetetraacetic acid
- CD
circular dichroism
- eIF2α
eukaryotic initiation factor 2α
- HEPES
4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
- IEF
isoelectric focusing
- MS
mass spectrometry
- pPKRm
monomer form of phosphorylated PKR
- pPKRd
dimeric form of phosphorylated PKR
- TCEP
Tris[2-carboxyethyl] phosphine
- Tris
Tris(hydroxymethyl)aminomethane
Footnotes
This work was supported by grant AI-53615 from NIH (J.L.C.)
Supporting Information
This material is available free of charge via the internet at http://pubs.acs.org.
References
- 1.Kaufman RJ. The double stranded RNA-activated protein kinase PKR. In: Sonenberg N, Hershey JWB, Mathews MB, editors. Translational Control of Gene Expression. Cold Spring Harbor Laboratory Press; Cold Spring Harbor: 2000. pp. 503–528. [Google Scholar]
- 2.Toth AM, Zhang P, Das S, George CX, Samuel CE. Interferon action and the double-stranded RNA-dependent enzymes ADAR1 adenosine deaminase and PKR protein kinase. Prog. Nucleic Acid Res. Mol. Biol. 2006;81:369–434. doi: 10.1016/S0079-6603(06)81010-X. [DOI] [PubMed] [Google Scholar]
- 3.Koromilas AE, Roy S, Barber GN, Katze MG, Sonenberg N. Malignant transformation by a mutant of the IFN-inducible dsRNA-dependent protein kinase. Science. 1992;257:1685–1689. doi: 10.1126/science.1382315. [DOI] [PubMed] [Google Scholar]
- 4.Meurs E, Galabru J, Barber GN, Katze MG, Hovanessian AG. Tumor supressor function of interferon-induced double-stranded RNA activated protein kinase. Proc. Natl. Acad. Sci. USA. 1993;90:232–236. doi: 10.1073/pnas.90.1.232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Weber F, Wagner V, Rasmussen SB, Hartmann R, Paludan SR. Double-stranded RNA is produced by positive-strand RNA viruses and DNA viruses but not in detectable amounts by negative-strand RNA viruses. J. Virol. 2006;80:5059–5064. doi: 10.1128/JVI.80.10.5059-5064.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tian B, Bevilacqua PC, Diegelman-Parente A, Mathews MB. The double-stranded RNA binding motif: Interference and much more. Nature Rev. Mol. Cell Biol. 2004;5:1013–1023. doi: 10.1038/nrm1528. [DOI] [PubMed] [Google Scholar]
- 7.Nanduri S, Carpick BW, Yang Y, Williams BR, Qin J. Structure of the double-stranded RNA binding domain of the protein kinase PKR reveals the molecular basis of its dsRNA-mediated activation. EMBO J. 1998;17:5458–5465. doi: 10.1093/emboj/17.18.5458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dar AC, Dever TE, Sicheri F. Higher-order substrate recognition of eIF2alpha by the RNA-dependent protein kinase PKR. Cell. 2005;122:887–900. doi: 10.1016/j.cell.2005.06.044. [DOI] [PubMed] [Google Scholar]
- 9.McKenna SA, Lindhout DA, Kim I, Liu CW, Gelev VM, Wagner G, Puglisi JD. Molecular framework for the activation of RNA-dependent protein kinase. J. Biol. Chem. 2007;282:11474–11486. doi: 10.1074/jbc.M700301200. [DOI] [PubMed] [Google Scholar]
- 10.VanOudenhove J, Anderson E, Kreuger S, Cole JL. Analysis of PKR structure by small angle scattering. J. Mol. Biol. 2009 doi: 10.1016/j.jmb.2009.02.019. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cole JL. Activation of PKR: an open and shut case? Trends Biochem. Sci. 2007;32:57–62. doi: 10.1016/j.tibs.2006.12.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Minks MA, West DK, Benvin S, Baglioni C. Structural requirements of double-stranded RNA for the activation of 2′-5′-Oligo(A) polymerase and protein kinase of interferon-treated HeLa cells. J. Biol. Chem. 1979;254:10180–10183. [PubMed] [Google Scholar]
- 13.Manche L, Green SR, Schmedt C, Mathews MB. Interactions between double-stranded RNA regulators and the protein kinase DAI. Mol. Cell Biol. 1992;12:5238–5248. doi: 10.1128/mcb.12.11.5238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kostura M, Mathews MB. Purification and activation of the double-stranded RNA-dependent eIF-2 kinase DAI. Mol. Cell. Biol. 1989;9:1576–1586. doi: 10.1128/mcb.9.4.1576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Schmedt C, Green SR, Manche L, Taylor DR, Ma Y, Mathews MB. Functional characterization of the RNA-binding domain and motif of the double-stranded RNA-dependent protein kinase DAI (PKR) J. Mol. Biol. 1995;249:29–44. doi: 10.1006/jmbi.1995.0278. [DOI] [PubMed] [Google Scholar]
- 16.Bevilacqua PC, Cech TR. Minor-groove recognition of double-stranded RNA by the double-stranded RNA-binding domain of the RNA-activated protein kinase PKR. Biochemistry. 1996;35:9983–9994. doi: 10.1021/bi9607259. [DOI] [PubMed] [Google Scholar]
- 17.Ucci JW, Kobayashi Y, Choi G, Alexandrescu AT, Cole JL. Mechanism of interaction of the double-stranded RNA (dsRNA) binding domain of protein kinase R with short dsRNA sequences. Biochemistry. 2007;46:55–65. doi: 10.1021/bi061531o. [DOI] [PubMed] [Google Scholar]
- 18.Lemaire PA, Anderson E, Lary J, Cole JL. Mechanism of PKR Activation by dsRNA. J. Mol. Biol. 2008;381:351–360. doi: 10.1016/j.jmb.2008.05.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lemaire PA, Lary J, Cole JL. Mechanism of PKR activation: dimerization and kinase activation in the absence of double-stranded RNA. J. Mol. Biol. 2005;345:81–90. doi: 10.1016/j.jmb.2004.10.031. [DOI] [PubMed] [Google Scholar]
- 20.Adams JA. Activation loop phosphorylation and catalysis in protein kinases: is there functional evidence for the autoinhibitor model? Biochemistry. 2003;42:601–607. doi: 10.1021/bi020617o. [DOI] [PubMed] [Google Scholar]
- 21.Nolen B, Taylor S, Ghosh G. Regulation of protein kinases; controlling activity through activation segment conformation. Mol. Cell. 2004;15:661–675. doi: 10.1016/j.molcel.2004.08.024. [DOI] [PubMed] [Google Scholar]
- 22.Zhang X, Herring CJ, Romano PR, Szczepanowska J, Brzeska H, Hinnebusch AG, Qin J. Identification of phosphorylation sites in proteins separated by polyacrylamide gel electrophoresis. Anal. Chem. 1998;70:2050–2059. doi: 10.1021/ac971207m. [DOI] [PubMed] [Google Scholar]
- 23.Romano PR, Garcia-Barrio MT, Zhang X, Wang Q, Taylor DR, Zhang F, Herring C, Mathews MB, Qin J, Hinnebusch AG. Autophosphorylation in the activation loop is required for full kinase activity in vivo of human and yeast eukaryotic initiation factor 2α kinases PKR and GCN2. Mol. Cell. Biol. 1998;18:2282–2297. doi: 10.1128/mcb.18.4.2282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Taylor DR, Lee SB, Romano PR, Marshak DR, Hinnebusch AG, Esteban M, Mathews MB. Autophosphorylation sites participate in the activation of the double-stranded-RNA-activated protein kinase PKR. Mol. Cell. Biol. 1996;16:6295–6302. doi: 10.1128/mcb.16.11.6295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Taylor DR, Tian B, Romano PR, Hinnebusch AG, Lai MM, Mathews MB. Hepatitis C virus envelope protein E2 does not inhibit PKR by simple competition with autophosphorylation sites in the RNA-binding domain. J. Virol. 2001;75:1265–1273. doi: 10.1128/JVI.75.3.1265-1273.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhang F, Romano PR, Nagamura-Inoue T, Tian B, Dever TE, Mathews MB, Ozato K, Hinnebusch AG. Binding of double-stranded RNA to protein kinase PKR is required for dimerization and promotes critical autophosphorylation events in the activation loop. J. Biol. Chem. 2001;276:24946–24958. doi: 10.1074/jbc.M102108200. [DOI] [PubMed] [Google Scholar]
- 27.Su Q, Wang S, Baltzis D, Qu LK, Wong AH, Koromilas AE. Tyrosine phosphorylation acts as a molecular switch to full-scale activation of the eIF2{alpha} RNA-dependent protein kinase. Proc. Natl. Acad. Sci. U S A. 2006;103:63–68. doi: 10.1073/pnas.0508207103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Galabru J, Hovanessian A. Autophosphorylation of the protein kinase dependent on double-stranded RNA. J. Biol. Chem. 1987;262:15538–15544. [PubMed] [Google Scholar]
- 29.Jammi NV, Beal PA. Phosphorylation of the RNA-dependent protein kinase regulates its RNA-binding activity. Nucleic Acids Res. 2001;29:3020–3029. doi: 10.1093/nar/29.14.3020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Langland JO, Jacobs BL. Cytosolic double-stranded RNA-dependent protein kinase is likely a dimer of partially phosphorylated Mr=66,000 subunits. J. Biol. Chem. 1992;267:10729–10736. [PubMed] [Google Scholar]
- 31.Anderson E, Cole JL. Domain Stabilities in Protein Kinase R (PKR): Evidence for Weak Interdomain Interactions. Biochemistry. 2008;47:4887–4897. doi: 10.1021/bi702211j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Laue TM. Short column sedimentation equilibrium analysis for rapid characterization of macromolecules in solution. Beckman Coulter Application Information. 1992 [Google Scholar]
- 33.Gualfetti PJ, Bilsel O, Matthews CR. The progressive development of structure and stability during the equilibrium folding of the alpha subunit of tryptophan synthase from Escherichia coli. Protein Sci. 1999;8:1623–1635. doi: 10.1110/ps.8.8.1623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bilsel O, Zitzewitz JA, Bowers KE, Matthews CR. Folding mechanism of the alpha-subunit of tryptophan synthase, an alpha/beta barrel protein: global analysis highlights the interconversion of multiple native, intermediate, and unfolded forms through parallel channels. Biochemistry. 1999;38:1018–1029. doi: 10.1021/bi982365q. [DOI] [PubMed] [Google Scholar]
- 35.Philo JS. Improved methods for fitting sedimentation coefficient distributions derived by time-derivative techniques. Anal. Biochem. 2006;354:238–246. doi: 10.1016/j.ab.2006.04.053. [DOI] [PubMed] [Google Scholar]
- 36.Stafford WF, Sherwood PJ. Analysis of heterologous interacting systems by sedimentation velocity: curve fitting algorithms for estimation of sedimentation coefficients, equilibrium and kinetic constants. Biophys. Chem. 2004;108:231–243. doi: 10.1016/j.bpc.2003.10.028. [DOI] [PubMed] [Google Scholar]
- 37.Ansevin AT, Roark DE, Yphantis DA. Improved Ultracentrifuge Cells for High-Speed Sedimentation Equilibrium Studies with Interference Optics. Anal.Biochem. 1970;34:237–261. doi: 10.1016/0003-2697(70)90103-x. [DOI] [PubMed] [Google Scholar]
- 38.Cole JL. Analysis of heterogeneous interactions. Methods Enzymol. 2004;384:212–232. doi: 10.1016/S0076-6879(04)84013-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Laue TM, Shah BD, Ridgeway TM, Pelletier SL. Computer-aided interpretation of analytical sedimentation data for proteins. In: Harding S, Rowe A, Horton J, editors. Analytical Ultracentrifugation in Biochemistry and Polymer Science. Royal Society of Chemistry; Cambridge: 1992. pp. 90–125. [Google Scholar]
- 40.Xu Z, Wang D, Lee X, Williams BR. Biochemical Analyses of Multiple Fractions of PKR Purified from Escherichia coli. J. Interferon Cytokine Res. 2004;24:522–535. doi: 10.1089/jir.2004.24.522. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.