Abstract
Increased O-linked β-N-acetylglucosamine (O-GlcNAc) is associated with insulin resistance in muscle and adipocytes. Upon insulin treatment of insulin-responsive adipocytes, O-GlcNAcylation of several proteins is increased. Key insulin signaling proteins, including IRS-1, IRS-2, and PDK1, are substrates for OGT, suggesting potential O-GlcNAc control points within the pathway. To elucidate the roles of O-GlcNAc in dampening insulin signaling (Vosseller, K., Wells, L., Lane, M. D., and Hart, G. W. (2002) Proc. Natl. Acad. Sci. U. S. A. 99, 5313–5318), we focused on the pathway upstream of AKT. Increasing O-GlcNAc in 3T3-L1 adipocytes decreases phosphoinositide 3-kinase (PI3K) interactions with both IRS-1 and IRS-2. Elevated O-GlcNAc also reduces phosphorylation of the PI3K p85 binding motifs (YXXM) of IRS-1 and results in a concomitant reduction in tyrosine phosphorylation of Y608XXM in IRS-1, one of the two main PI3K p85 binding motifs. Additionally, insulin signaling stimulates the interaction of OGT with PDK1. We conclude that one of the steps at which O-GlcNAc contributes to insulin resistance is by inhibiting phosphorylation at the Y608XXM PI3K p85 binding motif in IRS-1 and possibly at PDK1 as well.
Keywords: Carbohydrate/Function, Carbohydrate/Glycoconjugate, Carbohydrate/Glycoprotein, Diseases/Diabetes, Glycosylation, Metabolism/Metabolic Syndrome, Signal Transduction/Protein Kinases/Serine/Threonine, O-GlcNAc
Introduction
O-Linked β-N-acetylglucosamine (O-GlcNAc)5 is a single sugar modification that is found on both serine and threonine residues, similar to phosphorylation, throughout the nucleocytoplasm of the plant and animal kingdom (1, 2). O-GlcNAc is often found attached at or adjacent to phosphorylation sites, where in several cases the sugar residue plays a competitive role in cell signaling pathways (3). A global reciprocal relationship between O-phosphorylation and O-GlcNAcylation has been observed (4–6) in diverse cell types as well as a dynamic interplay at specific sites (4, 7–10). O-GlcNAc transferase (OGT) mediates the addition of O-GlcNAc, whereas β-d-N-acetylglucosaminidase (O-GlcNAcase) removes the sugar residue (11, 12). Interestingly, OGT activity has recently been shown to be responsive to cell signaling events (13, 14).
It is hypothesized that abnormal increases in O-GlcNAc disrupt phosphorylation-mediated insulin signaling by direct competition at the same or adjacent serine and threonine residues. A few studies have shown that increased O-GlcNAc dampens AKT activity in response to insulin stimulation of insulin target tissues, such as adipocytes and muscle and endothelial cells (15–18). Recent studies have demonstrated that a specific increase in global O-GlcNAc levels, induced by inhibiting O-GlcNAcase, in the absence of hyperglycemia, causes insulin resistance directly (16, 19, 20). However, another study in 3T3-L1 adipocytes suggests that increased O-GlcNAc is not required for insulin resistance, supporting the long held view that many different independent mechanisms can lead to insulin resistance (21).
Recently, O-GlcNAc sites were mapped to a number of key residues on IRS-1 (insulin receptor substrate 1) that are involved in different cell signaling events (22). In addition, insulin resistance in adipocytes is attributed to postinsulin receptor defects (23), making IRS-1 and IRS-2 excellent candidates to study early steps in the pathway that might explain how O-GlcNAc inhibits insulin signaling. In addition, OGT has been shown to play an important role in nematode insulin signaling (24), and its activity is also responsive to insulin signaling in both adipocytes (13) and liver cells (25). Therefore, one of the primary goals of our studies is to understand the impact of O-GlcNAcylation on the function of proteins in the PI3K/AKT signaling pathways.
Binding of insulin to its receptor initiates a well studied signaling cascade; activated insulin receptor phosphorylates the insulin receptor substrate proteins (IRS-1, IRS-2, IRS-3, and IRS-4) (26). IRS proteins bind to the regulatory subunit (p85) of PI3K via its Src homology 2 domains, binding predominantly to two YXXM motifs, Y612XXM and Y632XXM (mouse Y608XXM and Y628XXM) (27, 28). Activated PI3K phosphorylates phosphoinositides on the inositol ring, thus activating PDK1 (phosphoatidylinositol 3,4-bisphosphate/phosphoatidylinositol 3,4,5-trisphosphate-dependent kinase 1) (29, 30). PDK1 then phosphorylates Thr308 of AKT/PKB, causing its activation (31, 32). The PI3K pathway also activates atypical protein kinase C (PKCλ/ζ) (33). The activated AKT/PKB inactivates GSK3β (glycogen synthase kinase-3β), by phosphorylating Ser9 of GSKβ, hence activating glycogen synthesis (34). AKT/PKB and PKCλ/ζ as well as other proteins phosphorylate proteins that increase GLUT-4 (glucose transporter-4) vesicular trafficking to the plasma membrane (35–38).
In this study, we show that increased O-GlcNAcylation in 3T3-L1 adipocytes and on IRS-1 correlates with the down-regulation of phosphorylation at the Y608XXM PI3K p85 binding motif, which in turn is responsible for the decrease in AKT activation and GLUT-4 glucose uptake. Additionally, IRS-1, IRS-2, and PDK1 are excellent in vitro substrates for OGT. Insulin signaling stimulates interaction of OGT with PDK1 and increases its O-GlcNAc modification.
EXPERIMENTAL PROCEDURES
Reagents
Antibodies used in this study are against IRS-1, IRS-2, PI3K p85 (Upstate Biotechnology), Y612IRS-1 human (antibody will be referred to as Y608IRS-1 mouse throughout this work), PKCζ, GSK3β (Santa Cruz Biotechnology, Inc. (Santa Cruz, CA)), phospho-YXXM p85 motif, PDK1, AKT, phospho-Thr308 AKT (Cell Signaling), actin (Sigma), α-tubulin (T5168, Sigma), AL-28 OGT (Hart laboratory), anti-O-GlcNAc 110.6 CTD (Hart laboratory and Covance), and RL-2 (Affinity BioReagents, Golden, CO). Horseradish peroxidase-conjugated secondary antibodies were from Amersham Biosciences, and anti-IgM was from Sigma. Blots were developed with ECL reagent and Hyper-film (GE Healthcare). O-(2-Acetamido-2-deoxy-d-glucopyranosylidene)amino-N-phenylcarbamate (PUGNAc) was purchased from Carbogen (Switzerland) or synthesized in our laboratory (T. Lakshmanan) (20). Recombinant human insulin was from Roche Applied Science. All other reagents used were from Sigma unless otherwise indicated.
Cell Culture
3T3-L1 adipocytes were grown and differentiated into adipocytes, as described (39) with minor modifications (13, 39). Preadipocytes were grown to confluence in Dulbecco's modified Eagle's medium plus 10% calf serum containing penicillin/streptomycin and biotin; were differentiated in Dulbecco's modified Eagle's medium plus 10% fetal bovine serum with dexamethasone (390 ng/ml), insulin (1 μg/ml), and methylisobutylxanthine (115 μg/ml) for 48 h; and then treated with insulin (1 μg/ml) for an additional 48 h. Adipocytes were used for experiments between 8 and 11 days after differentiation.
Western Blotting and Immunoprecipitation
Whole cell lysates were harvested by washing 3T3-L1 three times in ice-cold phosphate-buffered saline and scraping into Nonidet P-40 buffer (1% Nonidet P-40, 15 mm Tris-HCl (pH 7.5), 150 mm NaCl, 1 mm EDTA, protease inhibitor mixture 1, 40 mm GlcNAc (GlcNAc prevents O-GlcNAcase from removing O-GlcNAc from proteins during cell lysis), and 1 mm NaVO4 as described (13). In most cases, the whole cell extract was sonicated two times for 10 s each with a 30-s pause between on ice. Lipids were removed by adding an equal volume (1:1) of ice-cold Freon (1,1,2-trichlorotrifluoroethane), centrifuged at 15,000 × g, and the protein concentration was determined by Bio-Rad protein reagent. For immunoprecipitations, cell lysates were rocked at 4 °C for 2 h to gently lyse cells and solubilize proteins in cell membranes before treating samples in a manner similar to that stated above. Immunoprecipitations were then performed with the indicated antibodies overnight at 4 °C, captured with Protein A/G-Sepharose (Santa Cruz Biotechnology, Inc.), and washed gently four times in phosphate-buffered saline (shorter incubations in some instances were necessary when trying to preserve post-translational modifications of proteins). Sepharose beads were boiled with modified Laemmli buffer to solubilize protein. Precast Criterion gels (Bio-Rad) were used for SDS-PAGE, transferred to polyvinylidene difluoride membrane (Millipore), blocked with 3% (w/v) bovine serum albumin (Sigma) in Tris-buffered saline plus 0.1% Tween 20 (TBST), and probed with the indicated antibodies overnight at 4 °C. The respective horseradish peroxidase-conjugated secondary antibodies were incubated with the blots, and ECL (Pierce) detection was used according to the manufacturer's directions. When necessary, blots were stripped with 100 mm glycine, pH 2.8, for 30 min at room temperature, washed with TBST, and reprobed with antibodies.
β-1,4-Galactosyltransferase
Immunoprecipitated proteins were rinsed twice with phosphate-buffered saline and then twice with 1× GalT1 (galactosyltransferase) labeling buffer (10×: 100 mm HEPES, pH 7.5, 100 mm galactose, 50 mm MnCl2) and then incubated in GalT1 buffer with 1 μCi/reaction of UDP-[3H]galactose, 1 unit of calf intestinal alkaline phosphatase at 4 °C overnight (40). Reactions were stopped by boiling samples with modified Laemmli buffer.
Enzymatic Labeling of O-GlcNAc Sites
The immunoprecipitated IRS-1 was labeled with N-azidogalactosamine and tagged with biotin as described previously (41). The enzymatic reactions were eluted with Laemmli buffer for Western blot analysis using streptavidin-horseradish peroxidase.
Kinase Assay
Insulin receptor substrate was immunoprecipitated from 3T3-L1 adipocytes and labeled in vitro with UDP-[3H]GlcNAc by incubating with or without OGT (bacterially expressed (42)), Sepharose A/G beads were washed extensively, and then 1 μm insulin-stimulated insulin receptor (Sigma I9266 protocol followed) was added for 20 min in kinase buffer (20 μm ATP, 5 mm Mn(CH3CO2)2, 50 mm HEPES, 0.1% Triton X-100, pH 6.9) (13, 43). The level of phosphorylation of the phospho-Tyr608 IRS-1 YXXM site was measured by Western blot analysis.
RESULTS
IRS-1, IRS-2, and PDK1 of the PI3K/AKT Insulin Signaling Cascade Are Substrates for OGT
To determine which proteins in the insulin signaling cascade might be substrates of OGT, we immunopurified several proteins in the pathway, including IRS-1, IRS-2, PI3K p85, PDK1, AKT, PKCζ, and GSK3β from 3T3-L1 adipocytes stimulated or not with 100 nm insulin. Each immunopurified protein was then incubated for 4 h with recombinant OGT and UDP-[3H]GlcNAc. All seven insulin signaling proteins were subjected to SDS-PAGE, stained with Coomassie Brilliant Blue, treated with EnHance, dried, and subjected to autoradiography (Fig. 1). IRS-1 is a good substrate of OGT and became highly modified with O-GlcNAc in both control and insulin-activated samples. Several groups have shown that IRS-1 is modified by O-GlcNAc in vivo (17, 18, 22). IRS-2 is also a substrate for OGT. However, IRS-2 isolated from cells not treated with insulin was a better substrate in vitro than that isolated from insulin-treated cells, suggesting that insulin-induced modification of IRS-2 by phosphate or another modification (like O-GlcNAc) may be blocking in vitro O-GlcNAcylation. PDK1 was also a good substrate for OGT and was modified in a pattern similar to that of IRS-2, with the non-activated PDK1 from control cells undergoing higher incorporation of O-GlcNAc than PDK1 from insulin-activated cells. In the insulin-stimulated PI3K p85 immunoprecipitation, IRS-1 co-immunoprecipitated and was also modified in vitro with O-GlcNAc, suggesting that the O-GlcNAc modification site is not modified by either O-GlcNAc or phosphorylation when IRS-1 is interacting with PI3K p85. OGT did not appear to modify PI3K, AKT, PKCζ, or GSK3β while slightly modifying the insulin receptor (data not shown) in vitro. Western blot analysis was conducted on all insulin-signaling proteins to confirm their presence in the immunoprecipitations.
FIGURE 1.
IRS-1, IRS-2, and PDK1 of the PI3K/AKT insulin signaling cascade are OGT substrates. Proteins from the insulin signaling cascade, IRS-1, IRS-2, PI3K p85, PDK, AKT, PKCζ, and GSK3β were immunoprecipitated from 3T3-L1 adipocytes stimulated or not with 100 nm insulin and incubated for 4 h while shaking with 0.5 μg of OGT and OGT reaction buffer containing UDP-[3H]GlcNAc. The reaction was stopped with modified Laemmli buffer, subjected to 10% SDS-PAGE, stained with Coomassie Brilliant Blue G-250, treated with EnHance, dried, and subjected to autoradiography at −80 °C for 4 days. Western blots (WB) of each immunoprecipitated protein were conducted.
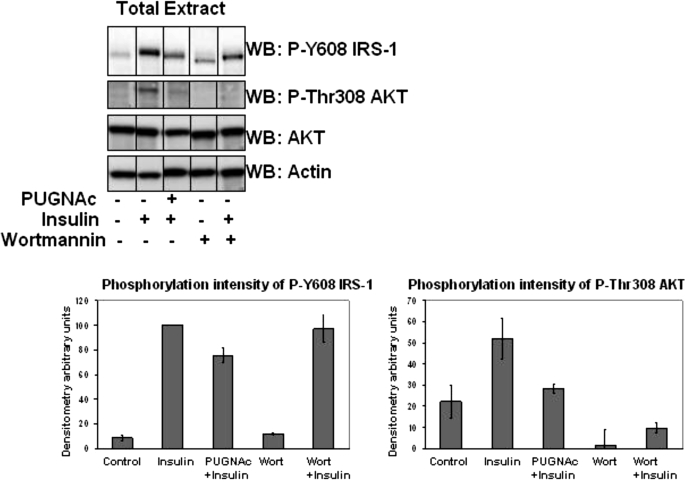
Increased O-GlcNAc in 3T3-L1 Adipocytes Inhibits Tyrosine Phosphorylation of the Tyr608 IRS-1 PI3K p85 YXXM Motif and Downstream AKT Phosphorylation
Previously, Vosseller et al. (16) showed that increasing O-GlcNAc levels in 3T3-L1 adipocytes, by using a selective inhibitor of O-GlcNAcase, dampened insulin-induced AKT Thr308 phosphorylation, an indicator of AKT activity, and concomitantly reduced insulin-stimulated glucose uptake. Multiple groups have demonstrated that IRS-1 and IRS-2 are modified by O-GlcNAc, suggesting that this may play a role in the downstream AKT activity. However, Vosseller et al. (16), using anti-phosphotyrosine antibodies, did not detect O-GlcNAc-modulated changes in insulin-stimulated tyrosine phosphorylation of IRS-1 or IRS-2. Because IRS-1 and IRS-2 contain over 20 potential tyrosine phosphorylation sites, it is unlikely that the two key tyrosine phosphorylation sites in the YXXM PI3K p85 binding motif, which are responsible for greater than 80% of PI3K p85 activation (27), would be seen by probing with a pan-specific phosphotyrosine antibody. Thus, to further examine the point at which O-GlcNAc plays a role upstream of the dampened AKT activity, we specifically looked at the phosphorylation of one of the key YXXM motifs of IRS-1 Tyr(P)608 in whole cell extracts of mouse 3T3-L1 adipocytes (Fig. 2). 3T3-L1 cells were treated with 100 μm PUGNAc overnight for 12 h and then retreated with PUGNAc for 4 h before insulin stimulation with 5 nm insulin for 10 min. The retreatment of 3T3-L1 adipocytes with PUGNAc insured increased global O-GlcNAc and significantly dampened the response of AKT activation by insulin (Fig. 2). Quantification of the Thr(P)308 AKT phosphorylation indicated that O-GlcNAc had reduced AKT activity by about 45%. Quantification of Tyr(P)608 IRS-1 indicated that O-GlcNAc reduced this key YXXM phosphorylation site by about 25%. Treatment of 3T3-L1 adipocytes with a general PI3K inhibitor, wortmannin, had no effect on Tyr(P)608 IRS-1 insulin-induced phosphorylation, whereas blocking Thr(P)308 AKT phosphorylation. The biological significance of the reduction in Tyr(P)608 and Thr(P)308 AKT phosphorylation was determined by the reduction in GLUT-4 glucose uptake (data not shown), consistent with earlier findings by Vosseller et al. (16).
FIGURE 2.
Elevations of O-GlcNAc in total cell extracts of 3T3-L1 adipocytes inhibit tyrosine phosphorylation of the Tyr(P)608 IRS-1 YXXM motif and downstream AKT phosphorylation. Total cell extracts from 3T3-L1 adipocytes pretreated or not with 100 μm PUGNAc, pretreated or not with 200 nm wortmannin, and stimulated or not with 5 nm insulin were Western blotted with Tyr(P)608 IRS-1, Thr(P)308 AKT, AKT, and actin antibodies. Data are representative of at least six experiments. Equal protein loading was demonstrated by Western blotting (WB) with anti-AKT and anti-actin antibody. The intensity of Tyr(P)608 IRS-1 Western blot analysis was normalized by equal protein loading and normalized to 100% of insulin stimulation using ImageJ. The intensity of Thr(P)308 AKT Western blot intensity was also quantitated with ImageJ.
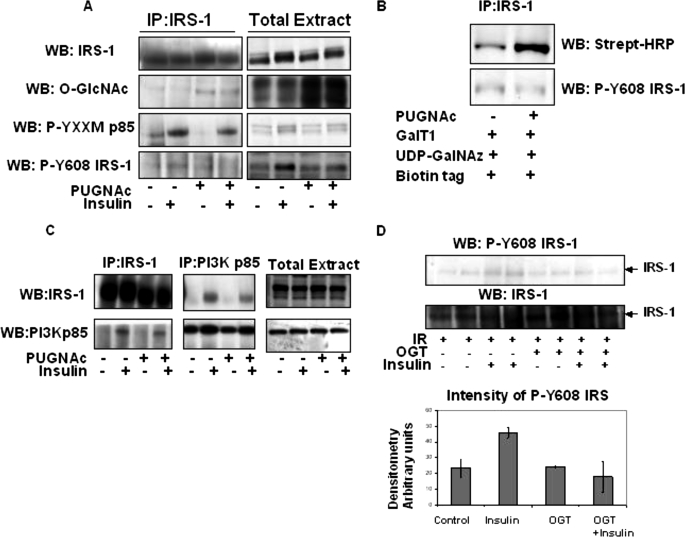
We then demonstrated that immunoprecipitated IRS-1 from whole cell extracts of 3T3-L1 adipocytes treated with PUGNAc had increased levels of O-GlcNAc modification (Fig. 3A) compared with controls. It is important to point out that the RL2 antibody is more immunoreactive to the O-GlcNAc-modified IRS-1 than the 110.6 O-GlcNAc antibody. In addition, we also used a far more sensitive GalT1, UDP-N-azidogalactosamine, biotin tag, streptavidin-horseradish peroxidase method to identify the O-GlcNAc modification on IRS-1 (Fig. 3B). This method clearly allows us to see that treatment of 3T3-L1 adipocytes with PUGNAc significantly increases O-GlcNAc modification on IRS-1. Western blot analysis of the IRS-1 with both a general phospho-YXXM motif antibody and the specific Tyr(P)608 IRS-1 antibody revealed that increased O-GlcNAc reduced insulin-induced phosphorylation of this motif in IRS-1 (Fig. 3A). All Western blots were also conducted with total cell extracts as controls (Fig. 3A, right panels). IRS-2 does not appear to be immunoreactive with either the general phospho-YXXM motif or Tyr(P)608 IRS-1 antibody (data not shown).
FIGURE 3.
IRS-1 is modified with O-GlcNAc, and elevations in O-GlcNAc reduce IRS-1 interaction with PI3K p85 and inhibit phosphorylation of the Tyr(P)608 IRS-1 YXXM motif. A, IRS-1 was immunoprecipitated (IP) from 3T3-L1 adipocytes pretreated or not with PUGNAc and then stimulated or not with 5 nm insulin and probed for IRS-1, O-GlcNAc RL-2, phospho-YXXM p85, and Tyr(P)612 IRS-1. The total cell extract was also Western blotted (WB) for all antibodies as a control. B, IRS-1 was immunoprecipitated from 3T3- L1 adipocytes pretreated or not with PUGNAc and labeled with GalT1, UDP-N-azidogalactosamine (GalNAz), and biotin tag and subjected to SDS-PAGE and Western blotting with streptavidin-horseradish peroxidase. C, IRS-1 and PI3K p85 were immunoprecipitated from PUGNAc-treated or -untreated and 5 nm insulin-stimulated or -unstimulated 3T3-L1 adipocytes and Western blotted for PI3K p85 and IRS-1 interaction, respectively. Total extracts were also Western blotted with IRS-1 and PI3K p85 as controls. Data are representative of at least three experiments IR, insulin receptor. D, immunopurified IRS-1 was labeled in vitro with UDP-[3H]GlcNAc by incubating with or without OGT, Sepharose A/G and washed, and then insulin receptor, stimulated or not with 1 μm insulin, was added for 20 min, and insulin-stimulated insulin receptor phosphorylation of Tyr(P)608 IRS-1 was measured by Western blot analysis using Tyr(P)608 IRS-1 antibody. Western blot intensity was quantitated using ImageJ.
Increased O-GlcNAc on IRS-1 reduced insulin induced interaction of PI3K p85 with IRS-1 immunoprecipitations (Fig. 3C) and reduced IRS-1 interaction with PI3K p85 immunoprecipitations, strongly suggesting that O-GlcNAc modification of IRS-1 dampens its interaction with PI3K p85. We were unable to detect O-GlcNAc on IRS-2, the level of IRS-2 protein or stoichiometry of O-GlcNAc may be too low (data not shown). However, PUGNAc treatment of 3T3-L1 adipocytes also resulted in reduced insulin-stimulated binding of PI3K p85 to IRS-2 (supplemental Fig. 1).
To further test the hypothesis that O-GlcNAc interferes with IRS-1 activity, we O-GlcNAcylated IRS-1 immunoprecipitates in vitro with recombinant OGT and UDP-GlcNAc. Untreated and O-GlcNAcylated IRS-1 were then mixed with insulin receptor stimulated or not with insulin, allowed to react, and immunoblotted for phospho-Y608IRS-1 (Fig. 3D). We found that modifying IRS-1 with O-GlcNAc substantially reduced insulin-stimulated insulin receptor phosphorylation of IRS-1 Tyr608 by about 50%.
PI3K p85-associated IRS-1 Does Not Appear to Be Modified by O-GlcNAc
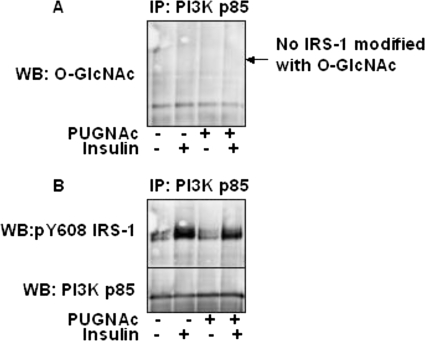
Interestingly, when the PI3K p85 immunoprecipitations were probed for O-GlcNAc at the molecular weight of IRS-1, no signal was found (Fig. 4A). When we probed PI3K p85 immunoprecipitations with Tyr(P)608 IRS-1 antibody, as expected, we saw that insulin induced an increase in phosphorylated Tyr608 IRS-1 interaction with PI3K p85 (Fig. 4B). We also saw that pretreating 3T3-L1 adipocytes with PUGNAc reduced the amount of phosphorylated Tyr608 IRS-1 associated with PI3K p85 (Fig. 4B). Interestingly, when the PI3K p85 immunoprecipitations were probed for O-GlcNAc at the molecular weight of IRS-1, no signal was found (Fig. 4B).
FIGURE 4.
O-GlcNAc-modified IRS-1 does not interact with PI3K p85. PI3K p85 was immunoprecipitated (IP) from 3T3-L1 adipocytes and (A) probed (WB) for O-GlcNAc and then stripped and (B) reprobed for Tyr(P)608 (pY608) IRS-1 and for PI3K p85 (to demonstrate equivalent protein).
OGT Interacts with PDK1 upon Insulin Stimulation of 3T3-L1 Adipocytes
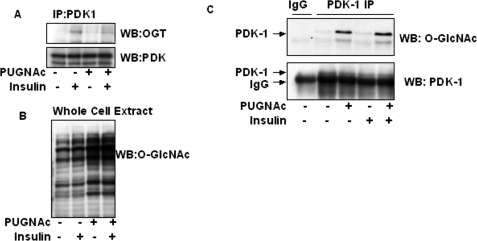
Previously, we documented that insulin treatment of 3T3-L1 adipocytes induces insulin receptor interaction and activation of OGT activity (13). In addition, Yang et al. (25) also showed that OGT was responsive to the insulin signaling. We therefore wanted to test if OGT interacts with any other major insulin signaling proteins that would affect downstream AKT activity. IRS-1, IRS-2, PI3K p85, PDK1, and AKT were each immunoprecipitated, subjected to SDS-PAGE, and probed by Western blotting to determine if OGT interacts with any of the proteins in the insulin signaling pathway besides the insulin receptor. We found that OGT interacts with PDK1 upon insulin treatment and that in 3T3-L1 adipocytes pretreated with PUGNAc, the PDK1-OGT association is significantly reduced (Fig. 5A; whole Western blot is shown in supplemental Fig. 2). The effect of PUGNAc on whole cell O-GlcNAc levels is displayed in Fig. 5B. Interestingly, 3T3-L1 adipocytes treated with PUGNAc significantly elevated O-GlcNAc on PDK1 (Fig. 5C). In addition, insulin treatment of these cells slightly increased O-GlcNAc modification on PDK1. PUGNAc may be dampening the interaction of OGT with PDK1 by a multitude of potential mechanisms. One possible mechanism of protein-protein interaction may be that the O-GlcNAc modification on PDK1 is inhibiting OGT interaction. Insulin stimulation of OGT activity (13) may feed back and down-regulate PDK1 activity and hence AKT activity. We did not detect any O-GlcNAc modification on AKT; nor did we see any OGT interaction with AKT (supplemental Fig. 3). The two anti-O-GlcNAc immunoblot bands in the PUGNAc-treated AKT immunoprecipitation do not line up with the anti-AKT immunoblot bands. We also did not detect OGT interacting with other O-GlcNAc-modified insulin signaling proteins (data not shown). Perhaps the on/off rate is too quick, or interactions with these substrates may occur transiently at time points different from those studied here. However, identification of elevated O-GlcNAc on PDK1 may, in addition to the dampening of Tyr(P)608 IRS-1 phosphorylation, have an effect on dampening AKT activity and GLUT-4 glucose transport.
FIGURE 5.
PDK1 interacts with OGT upon insulin treatment, and elevated O-GlcNAc dampens this response. A, PDK was immunoprecipitated (IP) from 3T3-L1 adipocytes pretreated or not with PUGNAc, stimulated or not with 5 nm insulin, and then Western blotted (WB) for the presence of OGT and PDK1. See supplemental Fig. 2 for whole Western blot. B, total cell extracts were Western blotted for O-GlcNAc. C, immunoprecipitates of PDK1 pretreated or not with PUGNAc and stimulated or not with insulin were also Western blotted for O-GlcNAc and for PDK1. IgG immunoprecipitation was used as a negative control.
DISCUSSION
The effect of O-GlcNAc on protein phosphorylation in cell signaling, cell cycle control, transcriptional regulation, cellular stress, and protein degradation, as well as many other cellular processes, is just beginning to be revealed (1–3, 44–48). Several groups have shown that O-GlcNAc directly inhibits insulin signaling in several cell lines (16–18). In addition, there are a number of other publications that suggest that the role of O-GlcNAc in Type II diabetes may be cell and protein cell signaling specific to adipocytes (16, 25, 49), muscles (18), and human coronary artery endothelial cells (17) because others have found that O-GlcNAc did not have any affect on AKT phosphorylation in neuroblastoma cells (50), neural precursor cells (51), and skeletal muscle cells (52). Interestingly, OGT itself, the enzyme responsible for mediating the addition of O-GlcNAc, is known to be involved in insulin signaling (13, 24, 25), activated by insulin signaling (13), and involved in insulin resistance (24, 25).
The significant increase of O-GlcNAc modification on IRS-1, IRS-2, and PDK1 in vitro when incubated with OGT suggests that this modification may play a role in insulin resistance with these proteins. Interestingly, IRS-2 and PDK1 from insulin-stimulated samples were poorer substrates for OGT then the same proteins from control samples, suggesting that insulin-stimulated phosphorylation or other modification may be inhibiting the O-GlcNAc modification of these proteins. AKT, PKCζ, PI3K, and GSK3β were not good substrates for OGT in vitro. In addition, analyses of immunoprecipitations of AKT (supplemental Fig. 3) and PKCζ (data not shown) were unable to detect O-GlcNAc on these proteins. We not only found that IRS-2, and especially IRS-1, are excellent substrates for OGT, but we also found that elevation of O-GlcNAc in whole cell extracts and on IRS-1 correlated with inhibition of PI3K p85 binding to the YXXM motif, inhibition of the phosphorylation of the YXXM motifs, and prevention of phosphorylation of Y608IRS-1.
Elevations of O-GlcNAc modification on IRS-1 may play a direct role in the decreased phosphorylation of the YXXM motifs. Both sites Tyr608 and Tyr628 are necessary for full activation of PI3K p85 (27), so reduced phosphorylation of either site would result in down-regulation of insulin-stimulated AKT activity. Interestingly, there is an O-GlcNAc consensus PVPS site between the two YXXM motifs (608ympmspgvapvpsgrkgsgd628ympms) in IRS-1 that may be a site of regulation for the tyrosine phosphorylation of the YXXM motifs. Recently, it has been shown that an IRS-1 Ser1011 site close to one of nine putative YXXM PI3K p85 sites is modified with O-GlcNAc (53). However, the putative YXXM PI3K p85 binding site adjacent to the O-GlcNAc-modified IRS-1 Ser1011 site would not be expected to have much of a role in PI3K interaction and activity because PI3K is predominantly activated by sites Tyr608 and Tyr628 in adipocytes. Although a number of O-GlcNAc modifications were site-mapped on IRS-1 (22, 53), none were close to the Tyr608 IRS-1 site. Therefore, we hypothesize that there may also be other potential serine O-GlcNAc sites in the vicinity of the major YXXM motifs, Tyr608 and Tyr628, that have not been site-mapped. Further analyses of IRS-1 and IRS-2 serine/threonine residues near the Tyr608 and Tyr628 sites, especially at the PVPS620 site, are necessary to determine if O-GlcNAc plays a role in their phosphorylation. Because insulin is capable of stimulating OGT activity and increases O-GlcNAc modification, it is plausible that hyperinsulemia, in conjunction with hyperglycemia, which increases UDP-GlcNAc pools, may be synergistically activating OGT, leading to abnormally increased O-GlcNAc levels in cells and contributing to insulin resistance (Fig. 6). Consistent with this model, IRS-1, IRS-2, and PDK1 are all substrates of OGT, and increased O-GlcNAc specifically inhibits tyrosine phosphorylation of IRS-1 at Tyr608 of the YXXM PI3K p85 binding motif, leading to downstream inactivation of AKT and dampened GLUT-4-mediated glucose uptake (data not shown). In addition, insulin stimulation of OGT and/or inhibition of O-GlcNAcase may also be regulating PDK1 regulation of AKT activity by elevation of O-GlcNAc on PDK1.
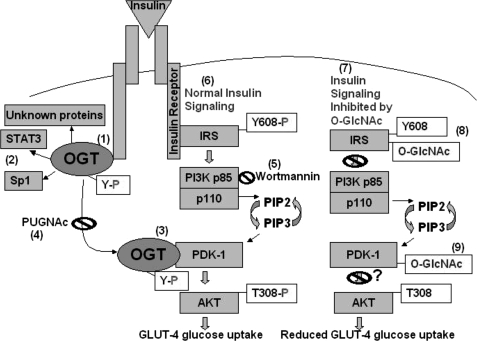
FIGURE 6.
Sites within the early steps of the insulin signaling pathway affected by OGT and O-GlcNAcylation. Upon insulin treatment of 3T3-L1 adipocytes, OGT is recruited to the insulin receptor (IR), tyrosine-phosphorylated, and catalytically activated (13) (1). OGT in turn modifies proteins, such as transcription factors (Sp1 and STAT3) (2) and kinases (PDK1), as well as a potential host of other unknown proteins (3). Acute insulin stimulation induces OGT interaction with at least PDK1 in the insulin signaling pathway and an increase in O-GlcNAc modification (4). Elevation of O-GlcNAc induced by PUGNAc inhibits OGT interaction with PDK1. Wortmannin (5) is a general inhibitor of PI3K, which blocks downstream AKT activation (6). During normal insulin-stimulated insulin receptor/AKT signaling, the insulin receptor becomes activated and phosphorylates the IRS-1 and IRS-2 YXXM motifs (Y608XXM and Y628XXM), which recruit and activate PI3K p85, resulting in downstream activation of PDK1, which in turn phosphorylates AKT (Thr308), resulting in GLUT-4 glucose uptake (7). Elevations in O-GlcNAc specifically induced by the O-GlcNAcase inhibitor PUGNAc result in O-GlcNAc modification of IRS-1 and decreased phosphorylation of Y608XXM (8), decreased PI3K p85 interaction with IRS-1, and downstream reduced phosphorylation of Thr308 of AKT, resulting in reduced GLUT-4 glucose uptake. IRS-1, IRS-2, and PDK1 were all excellent substrates for OGT in vitro (9). O-GlcNAc modification of PDK1 may play a role in reduced AKT activation.
OGT interaction with PDK1 was found to increase upon insulin stimulation, and PDK1 appears to become modified with O-GlcNAc. Further studies are necessary to elucidate the role of insulin-stimulated OGT interaction with PDK1. There could be many potential mechanisms for elevated O-GlcNAc blocking OGT and PDK1 interaction, such as modification of phosphatases, scaffolding proteins, or modification of sites on OGT and PDK1 themselves.
A growing number of publications are beginning to elucidate the role that O-GlcNAc plays in the cellular function of Type II diabetes (13, 22, 24, 53–58). Here we have shown that elevations of O-GlcNAc in 3T3-L1 cells inhibit phosphorylation of Tyr608 of IRS-1, hence down-regulating AKT activity, and that insulin stimulates OGT interaction with PDK1 as well as its O-GlcNAc modification in adipocytes. This study further documents that O-GlcNAc is an important post-translational modification in insulin signaling and probably plays a key role in the etiology of diabetes.
Acknowledgments
We thank Chad Slawson for critical reading of and suggestions for the manuscript, and we thank the Hart laboratory.
This work was supported, in whole or in part, by National Institutes of Health (NIH) Grant DK61671 and NIH Contract N01-HV-28180. G. W. H. receives a share of royalty received by the university on sales of the CTD 110.6 antibody. The terms of this arrangement are being managed by The Johns Hopkins University in accordance with its conflict of interest policies.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. 1–3.
- O-GlcNAc
- O-linked GlcNAc
- OGT
- O-GlcNAc transferase
- O-GlcNAcase
- O-GlcNAc-specific β-d-N-acetylglucosaminidase
- PI3K
- phosphoinositide 3-kinase (composed of regulatory unit p85 and catalytic subunit p110)
- PKB or AKT
- protein kinase B
- PKC
- protein kinase C
- PUGNAc
- O-(2-acetamido-2-deoxy-d-glucopyranosylidene)amino-N-phenylcarbamate.
REFERENCES
- 1.Whelan S. A., Hart G. W. (2003) Circ. Res. 93, 1047–1058 [DOI] [PubMed] [Google Scholar]
- 2.Wells L., Vosseller K., Hart G. W. (2001) Science 291, 2376–2378 [DOI] [PubMed] [Google Scholar]
- 3.Zachara N. E., Hart G. W. (2006) Biochim. Biophys. Acta 1761, 599–617 [DOI] [PubMed] [Google Scholar]
- 4.Comer F. I., Hart G. W. (2001) Biochemistry 40, 7845–7852 [DOI] [PubMed] [Google Scholar]
- 5.Lefebvre T., Alonso C., Mahboub S., Dupire M. J., Zanetta J. P., Caillet-Boudin M. L., Michalski J. C. (1999) Biochim. Biophys. Acta 1472, 71–81 [DOI] [PubMed] [Google Scholar]
- 6.Wang Z., Gucek M., Hart G. W. (2008) Proc. Natl. Acad. Sci. U.S.A. 105, 13793–13798 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chou T. Y., Hart G. W., Dang C. V. (1995) J. Biol. Chem. 270, 18961–18965 [DOI] [PubMed] [Google Scholar]
- 8.Kelly W. G., Dahmus M. E., Hart G. W. (1993) J. Biol. Chem. 268, 10416–10424 [PubMed] [Google Scholar]
- 9.Arnold C. S., Johnson G. V., Cole R. N., Dong D. L., Lee M., Hart G. W. (1996) J. Biol. Chem. 271, 28741–28744 [DOI] [PubMed] [Google Scholar]
- 10.Tao G. Z., Kirby C., Whelan S. A., Rossi F., Bi X., MacLaren M., Gentalen E., O'Neill R. A., Hart G. W., Omary M. B. (2006) Biochem. Biophys. Res. Commun. 351, 708–712 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kreppel L. K., Blomberg M. A., Hart G. W. (1997) J. Biol. Chem. 272, 9308–9315 [DOI] [PubMed] [Google Scholar]
- 12.Lubas W. A., Frank D. W., Krause M., Hanover J. A. (1997) J. Biol. Chem. 272, 9316–9324 [DOI] [PubMed] [Google Scholar]
- 13.Whelan S. A., Lane M. D., Hart G. W. (2008) J. Biol. Chem. 283, 21411–21417 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Song M., Kim H. S., Park J. M., Kim S. H., Kim I. H., Ryu S. H., Suh P. G. (2008) Cell. Signal. 20, 94–104 [DOI] [PubMed] [Google Scholar]
- 15.Yki-Järvinen H., Virkamäki A., Daniels M. C., McClain D., Gottschalk W. K. (1998) Metabolism 47, 449–455 [DOI] [PubMed] [Google Scholar]
- 16.Vosseller K., Wells L., Lane M. D., Hart G. W. (2002) Proc. Natl. Acad. Sci. U. S. A. 99, 5313–5318 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Federici M., Menghini R., Mauriello A., Hribal M. L., Ferrelli F., Lauro D., Sbraccia P., Spagnoli L. G., Sesti G., Lauro R. (2002) Circulation 106, 466–472 [DOI] [PubMed] [Google Scholar]
- 18.Patti M. E., Virkamäki A., Landaker E. J., Kahn C. R., Yki-Järvinen H. (1999) Diabetes 48, 1562–1571 [DOI] [PubMed] [Google Scholar]
- 19.Haltiwanger R. S., Grove K., Philipsberg G. A. (1998) J. Biol. Chem. 273, 3611–3617 [DOI] [PubMed] [Google Scholar]
- 20.Horsch M., Hoesch L., Vasella A., Rast D. M. (1991) Eur. J. Biochem. 197, 815–818 [DOI] [PubMed] [Google Scholar]
- 21.Buse M. G. (2006) Am. J. Physiol. Endocrinol. Metab. 290, E1–E8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ball L. E., Berkaw M. N., Buse M. G. (2006) Mol. Cell. Proteomics 5, 313–323 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lima F. B., Thies R. S., Garvey W. T. (1991) Endocrinology 128, 2415–2426 [DOI] [PubMed] [Google Scholar]
- 24.Forsythe M. E., Love D. C., Lazarus B. D., Kim E. J., Prinz W. A., Ashwell G., Krause M. W., Hanover J. A. (2006) Proc. Natl. Acad. Sci. U. S. A. 103, 11952–11957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yang X., Ongusaha P. P., Miles P. D., Havstad J. C., Zhang F., So W. V., Kudlow J. E., Michell R. H., Olefsky J. M., Field S. J., Evans R. M. (2008) Nature 451, 964–969 [DOI] [PubMed] [Google Scholar]
- 26.White M. F. (1997) Diabetologia 40, Suppl. 2, S2–S17 [DOI] [PubMed] [Google Scholar]
- 27.Esposito D. L., Li Y., Cama A., Quon M. J. (2001) Endocrinology 142, 2833–2840 [DOI] [PubMed] [Google Scholar]
- 28.Rordorf-Nikolic T., Van Horn D. J., Chen D., White M. F., Backer J. M. (1995) J. Biol. Chem. 270, 3662–3666 [DOI] [PubMed] [Google Scholar]
- 29.Storz P., Toker A. (2002) Front. Biosci. 7, d886–d902 [DOI] [PubMed] [Google Scholar]
- 30.Toker A., Newton A. C. (2000) Cell 103, 185–188 [DOI] [PubMed] [Google Scholar]
- 31.Nicholson K. M., Anderson N. G. (2002) Cell. Signal. 14, 381–395 [DOI] [PubMed] [Google Scholar]
- 32.Stokoe D., Stephens L. R., Copeland T., Gaffney P. R., Reese C. B., Painter G. F., Holmes A. B., McCormick F., Hawkins P. T. (1997) Science 277, 567–570 [DOI] [PubMed] [Google Scholar]
- 33.Le Good J. A., Ziegler W. H., Parekh D. B., Alessi D. R., Cohen P., Parker P. J. (1998) Science 281, 2042–2045 [DOI] [PubMed] [Google Scholar]
- 34.Shaw M., Cohen P., Alessi D. R. (1997) FEBS Lett. 416, 307–311 [DOI] [PubMed] [Google Scholar]
- 35.Calera M. R., Martinez C., Liu H., Jack A. K., Birnbaum M. J., Pilch P. F. (1998) J. Biol. Chem. 273, 7201–7204 [DOI] [PubMed] [Google Scholar]
- 36.Kupriyanova T. A., Kandror K. V. (1999) J. Biol. Chem. 274, 1458–1464 [DOI] [PubMed] [Google Scholar]
- 37.Standaert M. L., Bandyopadhyay G., Sajan M. P., Cong L., Quon M. J., Farese R. V. (1999) J. Biol. Chem. 274, 14074–14078 [DOI] [PubMed] [Google Scholar]
- 38.Kotani K., Ogawa W., Matsumoto M., Kitamura T., Sakaue H., Hino Y., Miyake K., Sano W., Akimoto K., Ohno S., Kasuga M. (1998) Mol. Cell Biol. 18, 6971–6982 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Student A. K., Hsu R. Y., Lane M. D. (1980) J. Biol. Chem. 255, 4745–4750 [PubMed] [Google Scholar]
- 40.Whelan S. A., Hart G. W. (2006) Methods Enzymol. 415, 113–133 [DOI] [PubMed] [Google Scholar]
- 41.Dias W. B., Cheung W. D., Wang Z., Hart G. W. (2009) J. Biol. Chem. 284, 21327–21337 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Gross B. J., Kraybill B. C., Walker S. (2005) J. Am. Chem. Soc. 127, 14588–14589 [DOI] [PubMed] [Google Scholar]
- 43.Kohanski R. A., Frost S. C., Lane M. D. (1986) J. Biol. Chem. 261, 12272–12281 [PubMed] [Google Scholar]
- 44.Kudlow J. E. (2006) J. Cell Biochem. 98, 1062–1075 [DOI] [PubMed] [Google Scholar]
- 45.Hanover J. A. (2001) FASEB J. 15, 1865–1876 [DOI] [PubMed] [Google Scholar]
- 46.Love D. C., Hanover J. A. (2005) Sci. STKE 2005, re13. [DOI] [PubMed] [Google Scholar]
- 47.Wells L., Whelan S. A., Hart G. W. (2003) Biochem. Biophys. Res. Commun. 302, 435–441 [DOI] [PubMed] [Google Scholar]
- 48.Slawson C., Housley M. P., Hart G. W. (2006) J. Cell Biochem. 97, 71–83 [DOI] [PubMed] [Google Scholar]
- 49.Park S. Y., Ryu J., Lee W. (2005) Exp. Mol. Med. 37, 220–229 [DOI] [PubMed] [Google Scholar]
- 50.Gandy J. C., Rountree A. E., Bijur G. N. (2006) FEBS Lett. 580, 3051–3058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yanagisawa M., Yu R. K. (2009) J. Neurosci. Res. 87, 3535–3545 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Arias E. B., Kim J., Cartee G. D. (2004) Diabetes 53, 921–930 [DOI] [PubMed] [Google Scholar]
- 53.Klein A. L., Berkaw M. N., Buse M. G., Ball L. E. (2009) Mol. Cell Proteomics 8, 2733–2745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Wells L., Vosseller K., Hart G. W. (2003) Cell Mol. Life Sci. 60, 222–228 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Robinson K. A., Ball L. E., Buse M. G. (2007) Am. J. Physiol. Endocrinol. Metab. 292, E884–E890 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Nelson B. A., Robinson K. A., Buse M. G. (2000) Diabetes 49, 981–991 [DOI] [PubMed] [Google Scholar]
- 57.Housley M. P., Rodgers J. T., Udeshi N. D., Kelly T. J., Shabanowitz J., Hunt D. F., Puigserver P., Hart G. W. (2008) J. Biol. Chem. 283, 16283–16292 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Housley M. P., Udeshi N. D., Rodgers J. T., Shabanowitz J., Puigserver P., Hunt D. F., Hart G. W. (2009) J. Biol. Chem. 284, 5148–5157 [DOI] [PMC free article] [PubMed] [Google Scholar]