Abstract
Cav3.2 T-type channels contain a high affinity metal binding site for trace metals such as copper and zinc. This site is occupied at physiologically relevant concentrations of these metals, leading to decreased channel activity and pain transmission. A histidine at position 191 was recently identified as a critical determinant for both trace metal block of Cav3.2 and modulation by redox agents. His191 is found on the extracellular face of the Cav3.2 channel on the IS3-S4 linker and is not conserved in other Cav3 channels. Mutation of the corresponding residue in Cav3.1 to histidine, Gln172, significantly enhances trace metal inhibition, but not to the level observed in wild-type Cav3.2, implying that other residues also contribute to the metal binding site. The goal of the present study is to identify these other residues using a series of chimeric channels. The key findings of the study are that the metal binding site is composed of a Asp-Gly-His motif in IS3–S4 and a second aspartate residue in IS2. These results suggest that metal binding stabilizes the closed conformation of the voltage-sensor paddle in repeat I, and thereby inhibits channel opening. These studies provide insight into the structure of T-type channels, and identify an extracellular motif that could be targeted for drug development.
Keywords: Channels/Calcium, Membrane/Channels, Metals/Zinc, Cav3.2, T-type Calcium Channel, Xenopus Oocytes, Electrophysiology, Voltage-clamping
Introduction
Low voltage-activated (LVA)2 T-type Ca2+ channels have been extensively studied because of their crucial implication in neuronal excitability, hormone secretion, muscle contraction, and pacemaker activity (1, 2). Molecular cloning studies demonstrated the existence of a family of T-type Ca2+ channels that consist of the three isoforms, Cav3.1, Cav3.2, and Cav3.3. Expression studies of the T-type channel α1 subunits revealed that the activation and inactivation kinetics of Cav3.1 and Cav3.2 are much faster than that of Cav3.3, although all three isoforms are activated at low voltages. It is generally accepted that the electrophysiological properties of currents recorded through T-type channel α1 subunits are similar to those recorded from isolated cells (1, 3, 4).
Block of voltage-dependent Ca2+ channels by metal ions has been extensively studied as it provides insight into structural changes in channel conformations during gating, and serves as a pharmacological tool to distinguish the various channel types. Among trace metals, Cd2+ has been shown to selectively block high voltage-activated (HVA)2 Ca2+ channels. Combined experiments using molecular biology and electrophysiology revealed that the Cd2+ binding site in HVA Ca2+ channels is composed of a Glu-Glu-Glu-Glu (EEEE) motif in the pore loops, thereby providing insights into Ca2+ channel permeation and selectivity (5, 6). Pharmacological studies have shown that LVA T-type currents are inhibited by much higher concentrations of Cd2+ than HVA channel currents (7, 8). Sensitivity to nickel inhibition of T-type currents varies greatly between cell types, implying the existence of multiple types of T-type channels. Molecular cloning of three T-type channel α1 subunits (Cav3.1, Cav3.2, and Cav3.3) allowed a comparison of nickel sensitivities of the three T-type channel isoforms, revealing that only Cav3.2 was sensitively inhibited by low micromolar concentrations of nickel (9). All three of the T-type channel isoforms share the Glu-Glu-Asp-Asp (EEDD) motifs in their pore regions corresponding to those of HVA Ca2+ channels. Replacement of the third and fourth aspartate with glutamate residues enhances cadmium block of Cav3.1 T-type channels, suggesting that LVA and HVA channels share a similar pore structure (10). Detailed biophysical studies of metal block have also found evidence that LVA and K+ channels share an internal activation gate (11), which in the case of Zn2+ binding to Cav3.3 (12), can lead to a “foot-in-the-door” block reminiscent of Rb+ block of K+ channels (13).
Pharmacological studies of cloned T-type channels reconstituted in expression systems have shown that Cav3.2 is more sensitively inhibited by not only nickel, but also zinc and copper, relative to either Cav3.1 or Cav3.3 (9, 12, 14). Notably, Cav3.2 channels are among the most sensitive targets of ion channel targets to zinc block (12). Our recent experiments using chimeric channels between Cav3.1 and Cav3.2 channels identified His191 in the extracellular loop connecting S3 and S4 of domain I as a major structural determinant for inhibition of the Cav3.2 by nickel, zinc, copper, and redox agents (15–17).
Metal binding sites in the voltage-sensor paddle have also been reported in Cav2.3, Nav1.2, Kv2.1, and Hv1 channels (18–23). Two histidine residues in the IS3–IS4 loop of the Cav2.3 channel were identified to be critical for the nickel-sensitive inhibition (18). The interaction sites of α-scorpion and sea anemone toxins, which cause slowing of fast inactivation, were localized to the S3–S4 loop of domain IV of the Nav1.2 channel (19, 20). The ability of hanatoxin to reduce channel activity and shift channel gating was also localized to the S3–S4 loop of Kv2.1 channels (21). Hanatoxin interaction with the S3–S4 loop of the Kv2.1 channel was thought to stabilize a closed state of the channel thereby reducing channel opening (21, 22). We previously proposed that the mechanism of nickel inhibition of Cav3.2 channel activity may be similar to the mechanism of hanatoxin inhibition (15). These findings establish that not only pore regions including S5 and S6, but also voltage-sensor regions including S1–S4 and their connecting loops could be potential binding sites for inhibitors.
Although His191 was identified to be critical for rendering the nickel or zinc inhibition sensitivity to Cav3.2 T-type channels (15), reverse introduction of a histidine residue into the corresponding locus (Gln172) of Cav3.1 channels only slightly increased trace metal block, suggesting that other residue(s) are involved in metal block of Cav3.2 channels in addition to His191. Therefore, we investigated additional residue(s) involved in zinc block of Cav3.2, focusing on zinc inhibition rather than copper to avoid complications of redox reactions (16). We found that the residues that precede His191, Asp189, and Gly190, were also critical residues for determining the high zinc sensitivity of Cav3.2. Additionally, we found an important role of negatively charged residues at the outer portion of the IS2 segment in zinc block of Cav3.2. These findings provide the structural basis of the high affinity extracellular metal binding site on Cav3.2, providing a novel therapeutic target for the treatment of neuropathic pain (24).
EXPERIMENTAL PROCEDURES
Chemicals
Chemicals were purchased from either Sigma or Amresco (Solon, OH). A zinc-chloride stock solution (100 mm; Sigma) was made in deionized water, and then stored at room temperature. A series of zinc solutions (in μm: 0.3, 1, 3, 10, 30, 100, 300, 1000, and 3000) were prepared by diluting the zinc stock solution with 10 mm Ba2+ solution just before experiments and their pH were adjusted to 7.6 if necessary.
Construction of Chimeras between Cav3.1 and Cav3.2
The serial chimeric channels constructed between the rat Cav3.1 (α1G; GenBank accession number AF027984) and human Cav3.2 (α1H; GenBank accession number AF051946) channels were previously reported (15). Additional chimeric and point mutant channels were made by the same PCR method described previously (15). All PCRs were performed using Pfu DNA polymerase (Vivagen, Seoul, Korea) and amplified fragments were verified by sequencing analysis. The restriction sites were marked by numbers in parentheses by indicating the 5′-terminal nucleotide generated by cleavage. Silent and non-silent mutations for restriction sites used for construction of chimeric channels are indicated by asterisks and crosses, respectively.
Cav3.1/3.2:N-IS4
Construction of plasmid Cav3.1/3.2:N-IS4 was previously reported (15).
Cav3.1/3.2:N-IS12L
The fragments used were ClaI (5′-polylinker)-StuI (491, Cav3.2) and StuI (491, Cav3.1)-HindIII (1754, Cav3.1). The plasmid Cav3.1/3.2:N-IS12L was constructed by ligating the above fragments into the ClaI (5′-polylinker) and HindIII-digested (1755, Cav3.1) plasmid Cav3.1 pGEM-HEA.
Cav3.1/3.2:IS2-IS4
The fragments used were ClaI (5′-polylinker)-StuI (838, Cav3.1) and StuI (491, Cav3.1/3.2:N-IS4)-HindIII (1754, Cav3.1/3.2:N-IS4). The plasmid Cav3.1/3.2:IS2-IS4 was constructed by ligating the above fragments into ClaI- (5′-polylinker) and HindIII-digested (1755, Cav3.1) plasmid Cav3.1 pGEM-HEA.
Cav3.1/3.2:IS34L
The fragments used were ClaI (5′-polylinker)-PmlI* (934, Cav3.1) and PmlI (589, Cav3.1/3.2:N-IS4)-HindIII (1754, Cav3.1/3.2:N-IS4). The plasmid Cav3.1/3.2:IS34L was constructed by ligating the above fragments into ClaI- (5′-polylinker) and HindIII-digested (1755, Cav3.1) plasmid Cav3.1 pGEM-HEA.
Cav3.1/3.2:N-IS12L+L171G+Q172H
The fragments used were ClaI (5′-polylinker)-StuI (491, Cav3.2) and StuI (491, Cav3.1/L171G+Q172H)-HindIII (1754, Cav3.1/L171G+Q172H). Plasmid Cav3.1/3.2:N-IS12L+L171G+Q172H was constructed by ligating the above fragments into ClaI- (5′-polylinker) and HindIII-digested (1755, Cav3.1) plasmid Cav3.1 pGEM-HEA.
Cav3.2/D140A, Cav3.2/D140E, and Cav3.2/A141D
The forward primers used to amplify the fragments for D140A, D140E, and A141D were GAGGCCTTTGCCGCCTTCATTTTCGCCTTTTTTG, GAGGCCTTTGAGGCCTTCATTTTCGCCTTTTTTG, and GAGGCCTTTGACGACTTCATTTTCGCCTTTTTTG, respectively, and the reverse primer was CAGGATCCGCATGCTAGG. Each point mutant channel was constructed by ligating the StuI- and BamHI-digested PCR fragments and ClaI (5′-polylinker)-StuI (491, Cav3.2) fragment into ClaI- (5′-polylinker) and BamHI-digested (730, Cav3.2) plasmid Cav3.2 pGEM-HEA.
Cav3.2/G190L
The fragments used were NotI (341, Cav3.2)-BglII+ (641, Cav3.2), BglII+ (642, Cav3.2)-BamHI (729, Cav3.2), and BamHI (730, Cav3.2)-SalI (4634, Cav3.2). Plasmid Cav3.2/G190L was constructed by ligating the above fragments into NotI- (342, Cav3.2) and SalI-digested (4635, Cav3.2) plasmid Cav3.2 pGEM-HEA.
Cav3.2/D189A and Cav3.2/D189E
The forward primer to amplify the upper cassettes for D189A and D189E was TAATACGACTCACTATAGGG (T7 promoter) and the reverse primers were TGTGTCCAGCCAACGAGTACTCCATCATGCCC and TGTGTCCCTCCAACGAGTACTCCATCATGCCC, respectively. The forward primers to amplify the lower cassettes were CTCGTTGGCTGGACACAACGTGAGCCTC and CTCGTTGGAGGGACACAACGTGAGCCTC, respectively, and the reverse primer was CAGGATCCGCATGCTAGG. The upper and lower cassettes were purified using the PCR purification kit and then combined by second-step PCR. Each point mutant channel was constructed by ligating the NotI- and BamHI-digested PCR fragments and BamHI (730, Cav3.2)-SalI (4634, Cav3.2) fragment into the NotI- (342, Cav3.2) and SalI-digested (4635, Cav3.2) plasmid Cav3.2 pGEM-HEA.
Cav3.1/L171G+Q172H and Cav3.1/Q172H+F176L
Plasmid Cav3.1/Q172H was used as a PCR template, and the forward primer used to amplify the upper cassettes for L171G+Q172H and Q172H+F176L was TAATACGACTCACTATAGGG (T7 promoter). The reverse primers were GTTGTGTCCGTCCAGCGAATACTCCAG and TGCGGAGAGGCTGACGTTGTGCAGGTC, respectively. The forward primers to amplify the lower cassettes were CTGGACGGACACAACGTCAGCTTCTCC and AACGTCAGCCTCTCCGCAGTCAGGGTC, respectively, and the reverse primer was GAGAAGCTTGCCAGGGTGCTAGC. The upper and lower cassettes were purified using the PCR purification kit and then combined by second-step PCR. Each point mutant channel was constructed by ligating the ClaI- and HindIII-digested PCR fragments into ClaI- (5′-polylinker) and HindIII-digested (1755, Cav3.1) plasmid Cav3.1 pGEM-HEA.
Cav3.1/D122A+L171G+Q172H
Plasmid Cav3.1/L171G+Q172H was used as a PCR template and the forward and reverse primers to amplify the upper cassettes for D122A + L171G + Q172H were TAATACGACTCACTATAGGG (T7 promoter) and AAAGATGAAGGCATCGAAGGCCTGCAGGAT, respectively. The forward and reverse primers to amplify the lower cassettes were GCCTTCGATGCCTTCATCTTTGCCTTCTTT and GAGAAGCTTGCCAGGGTGCTAGC, respectively. The upper and lower cassettes were purified using the PCR purification kit and then combined by second-step PCR. Each point mutant channel was constructed by ligating the ClaI- and HindIII-digested PCR fragments into the ClaI- (5′-polylinker) and HindIII-digested (1755, Cav3.1) plasmid Cav3.1 pGEM-HEA.
Preparation of Oocytes and Expression of T-type Ca2+ Channels and Mutant Channels
Preparation of Xenopus oocytes was previously described (15). Briefly, ovary lobes surgically obtained from mature female Xenopus laevis (Xenopus Express, France) were torn into small clusters of 5–7 oocytes in SOS solution (in mm: 100 NaCl, 2 KCl, 1.8 CaCl2, 1 MgCl2, 5 HEPES, 2.5 pyruvic acid, and 50 μg/ml of gentamicin, pH 7.6). Isolated oocytes were then treated with collagenase (10 mg/ml, Invitrogen) and trypsin inhibitor (Type III-O, Sigma) for 50 min in Ca2+-free OR2 solution (in mm: 82.5 NaCl, 2.5 KCl, 1 MgCl2, 5 HEPES, pH 7.6) to remove follicle membranes.
To synthesize capped cRNAs, all cDNAs encoding Cav3.1, Cav3.2, and chimeric channels were linearized by AflII and in vitro transcribed using T7 RNA polymerase (Ambion, Austin, TX) in accordance with the manufacturer's instructions. The cRNAs were injected into oocytes at concentrations of 10–50 ng/50 nl using a Drummond Nanoject pipette injector (Parkway, PA) attached to a Narishige micromanipulator (Tokyo, Japan). The SOS solution was changed daily.
Electrophysiology and Data Analysis
Ba2+ currents were measured using a two-electrode voltage clamp amplifier (OC-725C, Warner Instruments, Hamden, CT) between the 3rd and 5th days after cRNA injection. Microelectrodes were pulled using a pipette puller and filled with 3 m KCl. The electrode resistance was 0.5–1.0 megohm. The bath solution contained (in mm) 10 Ba(OH)2, 90 NaOH, 1 KOH, and 5 HEPES (pH 7.4 with methanesulfonic acid). The currents were sampled at 5 kHz and low pass-filtered at 1 kHz using the pClamp system (Digidata 1322A and pClamp 8; Molecular Devices, Palo Alto, CA). Data analysis and graphs were obtained with Clampfit software and Prism software (GraphPad, San Diego, CA), respectively. Dose-response curves were fitted using the Hill equation in Prism: B = (1 + IC50/(Zn2+)n)−1, where B is the normalized block, IC50 is the concentration of Zn2+ giving half-maximal inhibition, and n is the Hill coefficient. Data are presented as mean ± S.E. Statistical significance was measured using Student's unpaired t test.
Whole cell patch clamp recordings were obtained at room temperature using an Axopatch 200A amplifier equipped with a CV201A headstage. The amplifier was connected to a computer through a Digidata 1200 A/D converter, and controlled using pCLAMP 9.2 software. Whole cell currents were recorded using the following external solution (in mm): 15 CaCl2, 155 tetraethylammonium chloride (TEA-Cl), and 10 HEPES, pH adjusted to 7.4 with TEA-OH. The internal pipette solution contained the following (in mm): 125 CsCl, 10 EGTA, 2 CaCl2, 1 MgCl2, 4 Mg-ATP, 0.3 Na3GTP, and 10 HEPES, pH adjusted to 7.2 with CsOH. Pipettes were made from TW-150-3 capillary tubing (World Precision Instruments, Inc., Sarasota, FL). Under these solution conditions the pipette resistance was ∼2.4 megaohms. Access resistance and cell capacitance were calculated using on-line exponential fits to the capacitance transient induced by a 20-mV depolarization (Membrane Test, pCLAMP software). Cell capacitance averaged 10 picofarads. Access resistance averaged 4 megaohms, and was compensated 70% using the series resistance prediction and compensation circuit. Data were filtered at 2 kHz and digitized at 5 kHz for ionic currents or filtered at 10 kHz and digitized at 20 kHz for gating currents. The voltage protocol to measure gating currents included P/−8 subtraction to remove residual capacitance. Gating currents were integrated (pA × ms), and compared using a paired Student's t test. Due to variability between cells, the data were normalized to the control value.
RESULTS
Inward Ba2+ currents were not detected from control oocytes injected with either H2O or 0.1 m KCl, whereas robust inward transient currents were recorded from oocytes injected with cRNA encoding Cav3.1, Cav3.2, or chimeric channels. We first confirmed the Xenopus oocyte expression system could be used to measure the higher potency of zinc to inhibit Cav3.2 over Cav3.1 channels, which was originally observed in mammalian cells (14, 17). Application of serial zinc solutions demonstrated that Cav3.2 currents were sensitively inhibited by low micromolar concentrations of zinc, whereas Cav3.1 currents required much higher concentrations (Fig. 1, A and B). The average IC50 values for the Cav3.1 and Cav3.2 currents were 82.2 ± 6.9 and 3.0 ± 0.2 μm, respectively (n = 13, 42; open symbols in Fig. 1, E and F), being consistent with the previously reported findings performed in HEK-293 cells (14, 17). The half-potentials for their activation and steady-state inactivation were not significantly changed by zinc (Table 1). We also repeated our recent finding that the presence of His191 in the IS3–IS4 loop is critical for determining the zinc- and nickel-sensitive inhibition of the Cav3.2 channel (15, 17). Application of serial zinc solutions inhibited Cav3.2/H191Q currents in a concentration-dependent manner (Fig. 1F, ●), and the IC50 value was 103.8 ± 14.3 μm (n = 7). This result indicated that the H191Q mutation greatly diminished the zinc sensitivity of Cav3.2, lowering it to the levels observed with Cav3.1.
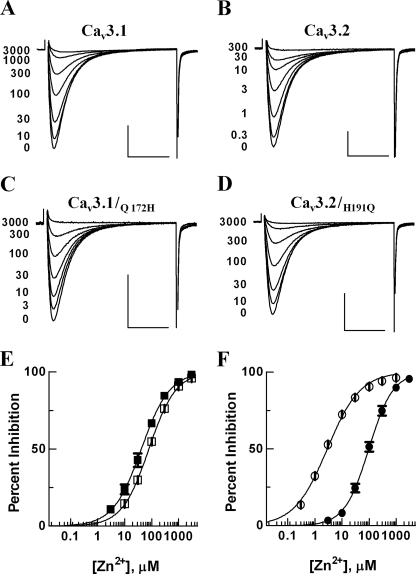
FIGURE 1.
Dose-response curves of zinc inhibition of recombinant Cav3.1, Cav3.1/Q172H, Cav3.2, and Cav3.2/H191Q expressed in Xenopus oocytes. Currents were elicited by a test potential to −20 mV from a holding potential of −90 mV every 15 s. A–D, representative current traces of Cav3.1, Cav3.2, Cav3.1/Q172H, and Cav3.2/H191Q before and after cumulative application of concentrations of zinc were superimposed. Scale bars on the x and y axes represent 20 ms and 1 μA, respectively. E, dose-response curves of zinc inhibition on Cav3.1 (□) and Cav3.1/Q172H (■). Currents were normalized to the peak current measured before application of zinc solutions, and the normalized percent inhibition was plotted against zinc concentrations. The smooth curves were obtained from fitting the average data with the Hill equation. Dose-response curves of zinc inhibition on Cav3.2 (○) and Cav3.2/H191Q (●) were obtained from analyzing data in a similar way and shown in F. All data are presented as mean ± S.E. (n = 4–42). The voltage dependence of activation and steady-state inactivation of wild-type and mutant channels are summarized in Table 1.
TABLE 1.
Summary for zinc inhibition effects on the Cav3.1, Cav3.2, and relevant mutants
IC50 values were estimated by fitting data with the Hill equation. V50 values of activation and inactivation were obtained from fitting data with a modified Boltzmann equation in the absence and presence of zinc that blocked current by about 50%. All data are shown as mean ± S.E. (n = 4–42). Slope factors and Hill coefficients were not statistically different, and thus, are not shown in the table.
| Construct | IC50 | n | V50 for activation |
V50 for inactivation |
||
|---|---|---|---|---|---|---|
| Control | Zn2+ | Control | Zn2+ | |||
| μm | mV | mV | ||||
| Cav3.1 | 82.2 ± 6.9a | 13 | −39.9 ± 1.0 | −38.0 ± 1.2 | −65.3 ± 1.1 | −63.4 ± 1.1 |
| Cav3.2 | 3.0 ± 0.2 | 42 | −35.3 ± 1.0 | −32.6 ± 0.9 | −61.0 ± 1.4 | −60.4 ± 1.7 |
| Cav3.1/3.2:IS34L | 5.6 ± 0.4a | 7 | −38.1 ± 0.5 | −35.7 ± 0.9b | −60.8 ± 1.0 | −59.9 ± 0.7 |
| Cav3.1/3.2:N-IS4 | 2.6 ± 0.3 | 6 | −40.6 ± 1.1 | −36.8 ± 1.0b | −62.9 ± 0.6 | −62.3 ± 0.7 |
| Cav3.1/3.2:N-IS2L | 85.2 ± 9.3a | 4 | −44.5 ± 2.0 | −41.3 ± 1.5 | −64.8 ± 2.0 | −64.8 ± 1.5 |
| Cav3.1/3.2:IS2-IS4 | 1.7 ± 0.2c | 13 | −32.3 ± 0.7 | −30.2 ± 1.0 | −59.3 ± 1.8 | −61.5 ± 2.2 |
| Cav3.2/D140A | 7.9 ± 1.3a | 8 | −24.6 ± 0.7 | −22.1 ± 0.8b | −50.6 ± 0.9 | −51.9 ± 1.2 |
| Cav3.2/D140E | 3.4 ± 0.7 | 10 | −30.5 ± 1.3 | −25.1 ± 1.5b | −53.8 ± 1.1 | −53.0 ± 1.0 |
| Cav3.2/A141D | 12.5 ± 2.1a | 8 | −32.5 ± 1.1 | −30.9 ± 1.1 | −56.2 ± 1.0 | −56.9 ± 0.5 |
| Cav3.2/D189A | 28.2 ± 8.8a | 13 | −38.0 ± 0.7 | −35.3 ± 0.6b | −63.8 ± 0.6 | −61.1 ± 0.5b |
| Cav3.2/D189E | 1.0 ± 0.1a | 6 | −39.3 ± 1.8 | −35.3 ± 1.5 | −65.8 ± 1.6 | −63.1 ± 1.4 |
| Cav3.2/G190L | 47.8 ± 6.5a | 12 | −32.3 ± 1.2 | −29.1 ± 1.3 | −57.0 ± 1.0 | −55.2 ± 1.2 |
| Cav3.2/H191Q | 103.8 ± 14.3a | 7 | −36.6 ± 0.7 | −34.9 ± 0.7 | −58.1 ± 0.6 | −57.5 ± 0.6 |
| Cav3.1/Q172H | 42.5 ± 7.6a | 4 | −42.8 ± 2.0 | −38.3 ± 2.0 | −66.3 ± 2.0 | −64.9 ± 2.4 |
| Cav3.1/L171G+Q172H | 6.4 ± 0.6a | 4 | −41.5 ± 1.0 | −40.7 ± 0.7 | −64.7 ± 0.8 | −64.8 ± 1.0 |
| Cav3.1/Q172H+F176L | 56.6 ± 3.3a | 10 | −42.0 ± 0.8 | −37.3 ± 1.0c | −59.4 ± 1.5 | −58.5 ± 1.6 |
| Cav3.1/D122A+L171G+Q172H | 1.4 ± 0.2a | 14 | −35.1 ± 0.5 | −30.0 ± 0.4c | −60.1 ± 0.5 | −58.9 ± 0.6 |
| Cav3.1/3.2:N-IS2L+L171G+Q172H | 5.5 ± 0.5a | 10 | −38.9 ± 1.2 | −35.5 ± 1.3b | −63.2 ± 1.0 | −63.0 ± 1.0 |
a Significant differences of the data were analyzed using Student's unpaired t test, p < 0.001.
b Significant differences of the data were analyzed using Student's unpaired t test, p < 0.05.
c Significant differences of the data were analyzed using Student's unpaired t test, p < 0.01.
We next tested how reverse mutation of the corresponding glutamine residue to histidine (Q172H) in Cav3.1 altered zinc sensitivity of Cav3.1. The zinc inhibition profile of the Cav3.1/Q172H mutant revealed an IC50 value of 42.5 ± 7.6 μm (n = 4), indicating that the mutation increased sensitivity by about 2-fold relative to wild-type Cav3.1, but still 15-fold less sensitive than Cav3.2 (Fig. 1E, ■). These results suggested that adoption of other residue(s) into Cav3.1/Q172H were required to gain the zinc-sensitive inhibition to the levels observed with Cav3.2 besides His191.
To narrow down other regions contributing to the zinc sensitivity of Cav3.2, we systematically constructed serial chimeric channels by adopting the adjacent regions(s) of His191 into either Cav3.1 or Cav3.1/Q172H. We first constructed Cav3.1/3.2:N-IS4 by introducing the region from the amino terminus to the IS4 segment of Cav3.2 into the Cav3.1. The IC50 value of Cav3.1/3.2:N-IS4 was 2.6 ± 0.3 μm (n = 6; Fig. 2), which was similar to that of Cav3.2. Consistent with our previous report (15), this result strongly suggested that residue(s) determining the zinc sensitivity of Cav3.2 were present in the voltage-sensor paddle region (IS1–IS4) of domain I of Cav3.2 rather than the pore region. Next, we constructed chimeric channels with smaller substitutions, Cav3.1/3.2:N-IS12L and Cav3.1/3.2:IS34L, by transferring the NH2 terminus to the connecting loop between IS1 and IS2 of Cav3.2 and the connecting linker between IS3 and IS4 of Cav3.2 into the Cav3.1, respectively (Fig. 2). Zinc inhibition profiles of the chimeric channels showed that the IC50 value of Cav3.1/3.2:N-IS12L was 85.2 ± 9.3 μm (n = 4), which was similar to that of Cav3.1. In contrast, zinc sensitivity of Cav3.1/3.2:IS34L (IC50 = 5.6 ± 0.4 μm, n = 7) was profoundly increased, but was still 2-fold lower than that of Cav3.2 (Student's unpaired t test, p < 0.001; Fig. 2). The 2-fold difference between the wild-type Cav3.2 and Cav3.1/3.2:IS34L implied that although residue(s) critically determining the zinc-sensitive inhibition were located in the IS3–IS4 loop of Cav3.2, additional residues might be present in its neighboring region(s), such as the region preceding the IS3–IS4 loop. Therefore, we tested the zinc sensitivity of Cav3.1/3.2:IS2-IS4, which was constructed by introducing the region from IS2 to IS4 of Cav3.2 into Cav3.1 (Fig. 2). The zinc inhibition profiles revealed that Cav3.1/3.2:IS2-IS4 was ∼3.3- and ∼1.8-fold more sensitive to zinc than Cav3.1/3.2:IS34L and wild-type Cav3.2, respectively (IC50 = 1.7 ± 0.2 μm, n = 13; p < 0.001; Fig. 2). The enhanced sensitivity suggests that residues in IS2 and IS3 that precede the IS3–IS4 loop may participate in zinc inhibition of Cav3.2 currents.
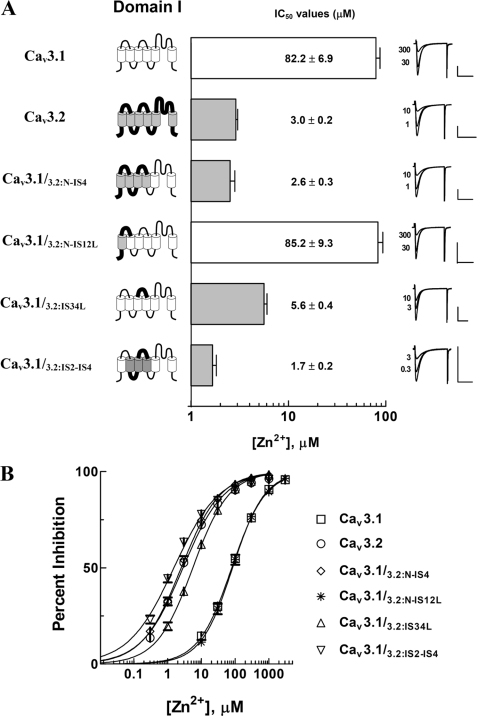
FIGURE 2.
Potency of zinc to inhibit Cav3.1, Cav3.2, and chimeric channels mutated in the domain I. A, schematic diagrams of Cav3.1, Cav3.2, and their chimeras were linearly represented for Domain I (left). The transmembrane segments and connecting loops of the T-type channels are displayed with cylinders and lines. White cylinders and thin lines represent the regions from Cav3.1, whereas gray cylinders and thick lines represent regions from Cav3.2. Domains II, III, and IV of these chimeras were from Cav3.1. IC50 values of chimeric channels were exhibited with bar graphs (middle). These values were obtained from dose-response curves (shown in B) for which the percent inhibition data were fitted with the Hill equation. Representative current traces of Cav3.1, Cav3.2, and chimeric channels before and after zinc inhibition were exhibited (right panel of A). Scale bars on the x and y axes represent 40 ms and 1 μA, respectively. B, dose-response curves of Cav3.1, Cav3.2, and their chimeric channels. The chimeric channels (Cav3.1/3.2:N-IS4 (◇), Cav3.1/3.2:IS34L (▵), and Cav3.1/3.2:IS2-IS4 (▿)), which contain the linker connecting IS3 and IS4 of Cav3.2 in common, were inhibited by low micromolar concentrations of zinc. Their zinc inhibition sensitivities were close to that of Cav3.2 (○). In contrast, Cav3.1/3.2:N-IS12L (*) required much higher concentrations to be inhibited and its zinc sensitivity was similar to that of Cav3.1 (□). All data are presented as mean ± S.E. (n = 4–42).
The reverse single mutation of Q172H increased the zinc sensitivity of Cav3.1 by about 2-fold. In contrast, adoption of the IS3–IS4 loop of Cav3.2 into the corresponding loop of Cav3.1 increased zinc sensitivity 15-fold, suggesting that not only His191, but also other residues in the IS3–IS4 loop, participate in the zinc sensitivity of Cav3.2. Comparing the amino acid sequences of the IS3–IS4 loops of Cav3.1 with those of Cav3.2 revealed that Leu171, Gln172, and Phe176 in Cav3.1 were different from the corresponding residues, Gly190, His191, and Leu195, in the Cav3.2 channel (Fig. 3A). To examine whether these residues influence zinc sensitivity, we additionally introduced individual mutations of L171G and F176L into Cav3.1/Q172H. Analysis of their zinc inhibition profiles showed that zinc sensitivity of Cav3.1/Q172H+F176L was not significantly different from that of Cav3.1/Q172H (Fig. 3, B and C). In contrast, the zinc sensitivity of Cav3.1/L171G+Q172H was significantly augmented (IC50 = 6.4 ± 0.6 μm, n = 4; p < 0.001; Fig. 3, B and C), suggesting that the double mutation of L171G as well as Q172H into Cav3.1, was critical to the increase in zinc sensitivity of wild-type Cav3.1. Consistently, reciprocal mutation of Gly190 of Cav3.2 into leucine strongly reduced the potency of zinc (Cav3.2/G190L; IC50 = 47.8 ± 6.5 μm, n = 12; Fig. 3, B and C), supporting our hypothesis that glycine as well as histidine residues in the IS3–IS4 loop are crucial elements affecting zinc sensitivity of Cav3.2.
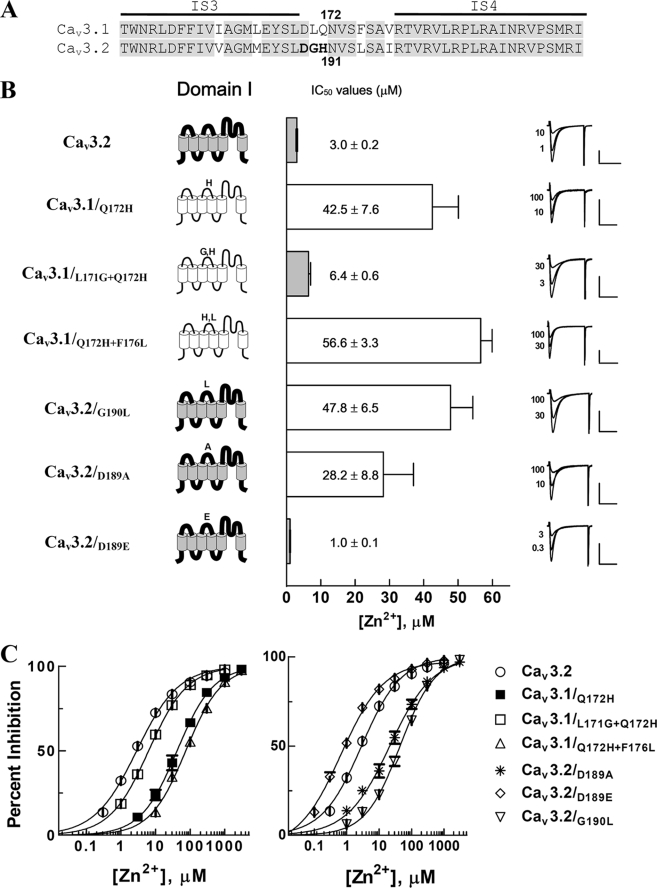
FIGURE 3.
Zinc inhibition profiles of the mutant channels in the IS3–IS4 linkers of Cav3 channels. A, the amino acid sequences between the linkers connecting S3 and S4 of domain I of Cav3.1 and Cav3.2 are aligned. The putative membrane-spanning segments are marked with horizontal bars above the sequence, and conserved residues between the Cav3.1 and Cav3.2 are highlighted in gray. The residues critically contributing to zinc inhibition sensitivity are represented by bold letters and amino acid numbers are based on the Cav3.1 and Cav3.2 sequence, respectively. B, the IC50 values of the Cav3.1 mutant channels (Cav3.1/Q172H, Cav3.1/L171G+Q172H, and Cav3.1/Q172H+F176L) and Cav3.2 mutant channels (Cav3.2/D189A, Cav3.2/D189E, and Cav3.2/G190L) are displayed with bar graphs. Representative current traces before and after zinc inhibition were superimposed for comparison. C, the dose-response curves of zinc inhibition for mutant channels were obtained from fitting the average inhibition percentages by different zinc concentrations with the Hill equation. The left panel showed dose-response curves of the Cav3.1 mutant channels (Cav3.1/Q172H (■), Cav3.1/L171G+Q172H (□), and Cav3.1/Q172H+F176L (▵)). The dose-response curves of the Cav3.2 mutant channels are exhibited in the right panel for Cav3.2/D189A (*), Cav3.2/D189E (◇), and Cav3.2/G190L (▿). All data are presented as mean ± S.E. (n = 4–42).
Sequence comparison of the IS3–IS4 loops of Cav3 isoforms identified an aspartate residue that was commonly present at the preceding position to the identified two critical residues (Gly190 and His191). Because the carboxyl acid side chains of aspartate could be another potential ligand involved in zinc binding, we tested possible participation of Asp189 in the zinc sensitivity of Cav3.2. Point mutation of D189A into Cav3.2 decreased its zinc sensitivity by about 10-fold (IC50 = 28.2 ± 8.8 μm, n = 13; Fig. 3, B and C), whereas point mutation of D189E enhanced its zinc sensitivity by about 3-fold (IC50 = 1.0 ± 0.1 μm, n = 6; p < 0.001; Fig. 3, B and C). These results indicated that an acidic residue just before the critical Gly190 and His191 residues is also required to endow the Cav3.2 with high-affinity zinc inhibition. Taken together, these findings indicate that Asp189, Gly190, and His191 compose a major part of the zinc-binding motif.
Crystallographic studies of the voltage-sensor paddle of voltage-gated K+ channels show that the S1–S2 loop and the S3 and S4 loops are in close proximity (25). Therefore, we hypothesized that the Asp-Gly-His motif in the IS3–IS4 loop is likely to be structurally close to the IS1–IS2 loop of Cav3.2 (Fig. 4A). To test the possibility that the region from the amino-terminal end to the IS1–IS2 loop influences zinc sensitivity, we constructed Cav3.1/3.2:N-IS12L+L171G+Q172H. The zinc sensitivity of this mutant was similar to that of Cav3.1/L171G+Q172H, displaying an IC50 of 5.5 ± 0.5 μm (n = 10) (Fig. 4, B and C). This finding suggests that non-conserved residues in the NH2 terminus to IS1–IS2 loop do not contribute to the zinc metal binding site.
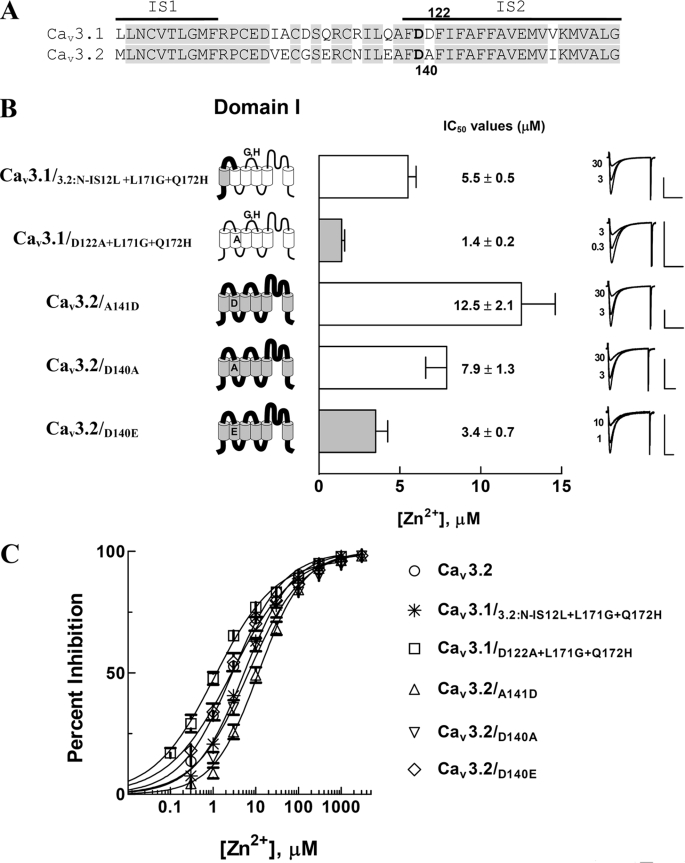
FIGURE 4.
Zinc inhibition profiles of channels mutated in transmembrane segment IS2 of Cav3. A, alignment of the amino acid sequences between the S1 and S2 of domain I of Cav3.1 and Cav3.2 channels. The horizontal bars above the sequence represent the putative membrane-spanning segments, and the highlighted portions in gray mark conserved residues between the Cav3.1 and Cav3.2. Phe-Asp-Asp is found in the outer portion of IS2 in the Cav3.1, whereas Phe-Asp-Ala in the Cav3.2. Amino acid numbers are based on the Cav3.1 and Cav3.2, respectively. B, schematic diagrams of the mutant channels (Cav3.1/3.2:N-IS12L+L171G+Q172H, Cav3.1/D122A+L171G+Q172H, Cav3.2/A141D, Cav3.2/D140A, and Cav3.2/D140E) are shown at the left. Their IC50 values and representative current traces are in the middle and right. C, the dose-response curves of zinc inhibition on individual mutant channels were obtained from fitting the average inhibition percentages by different zinc concentrations with the Hill equation for Cav3.1/3.2:N-IS12L+L171G+Q172H (*), Cav3.1/D122A+L171G+Q172H (□), Cav3.2/A141D (▵), Cav3.2/D140A (▿), and Cav3.2/D140E (◇). All data are presented as mean ± S.E. (n = 8–14).
Zinc inhibition profiles of Cav3.1/3.2:IS34L and Cav3.1/3.2:IS2-IS4 showed that, despite both mutants containing the Asp-Gly-His motif, the former was still about 2-fold less sensitive to zinc than Cav3.2, whereas the latter was about 2-fold more sensitive (Fig. 2). Compared with Cav3.1/3.2:IS34L, Cav3.1/3.2:IS2-IS4 additionally contained the IS2–IS3 and IS4 regions from Cav3.2, implying that these regions are responsible for the difference in zinc sensitivity. Sequence comparison from IS2 to IS4 revealed that “Phe-Asp-Ala141” containing one negatively charged residue was present at the outer portion of IS2 of Cav3.2, whereas “Phe-Asp-Asp122” containing two negatively charged residues was at the corresponding portion in Cav3.1 (Fig. 4A). We hypothesized that the position and number of aspartate residues at this position might alter zinc sensitivity. To test this hypothesis, we first replaced Asp122 in Cav3.1/L171G+Q172H with a non-polar alanine residue, converting “Phe-Asp-Asp” into “Phe-Asp-Ala.” Cav3.1/D122A+L171G+Q172H was about 2- and 4-fold more sensitive to zinc than wild-type Cav3.2 and Cav3.1/L171G+Q172H, respectively (IC50 = 1.4 ± 0.2 μm, n = 14; p < 0.001; Fig. 4, B and C). In contrast, Cav3.2/A141D was 4-fold less sensitive to zinc than wild-type Cav3.2 (Cav3.2/A141D, IC50 = 12.5 ± 2.1 μm, n = 8; Fig. 4, B and C). In addition, mutation of D140A (Cav3.2/D140A) reduced zinc sensitivity of Cav3.2 by 2.6-fold (IC50 = 7.9 ± 1.3 μm, n = 8; Fig. 4, B and C), whereas mutation of D140E (Cav3.2/D140E) did not significantly alter zinc sensitivity (IC50 = 3.4 ± 0.7 μm, n = 10; p = 0.4601; Fig. 4B). These consistent changes in zinc sensitivity by diverse mutations of aspartate residues in this region support the hypothesis that the acidic residue(s) in the outer portion of IS2 may also play a role in zinc binding. Another implication is that the presence of only one negatively charged residue in this region is more likely to form a structural conformation favorable to zinc block.
Based on the experimental results and potassium channel models (25, 26), we developed a model to illustrate how zinc coordinates to the Asp-Gly-His motif and the acidic residue at the outer portion of IS2 (Fig. 5). We chose a model of the closed channel (26) rather than the crystal structure of the open channel (25), because our previous studies demonstrated that nickel had a lower affinity for the open state (9). The model suggests zinc might inhibit channel opening by disrupting the function of the voltage-sensor paddle in repeat I, rather than a direct action on Ca2+ ion permeation as observed with Cd2+ block of HVA channels. We next sought biophysical data to support this hypothesis, reasoning that Zn2+ might alter charge movement that precedes channel opening. Due to limitations in the two microelectrode voltage clamp of oocytes, we used patch clamp recording of HEK-293 cells in the whole cell mode. On-gating currents can be measured at the reversal potential after series resistance compensation and proper cancellation of residual capacitance charge, and quantitated by integrating the area of the outward gating current (27). Expression of recombinant Cav3.2 in HEK-293 cells generated ∼4000 pA of inward current during step depolarizations to −20 mV and ∼750 pA outward current during step depolarizations to +55 mV (Fig. 6). Addition of 10 μm zinc inhibited the inward current by 91% (±1%, n = 6), and inhibited the outward gating current 23% (±1%, n = 6, p = 0.08, paired t test; Fig. 6, C and D). Time matched controls (Control 2) showed no change in gating current, ruling out effects due to rundown. These results indicate that zinc is capable of inhibiting gating charge movement, and is consistent with a model whereby zinc binding stabilizes the closed conformation of the voltage-sensor paddle.
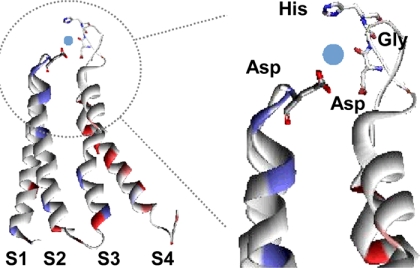
FIGURE 5.
A structural model for zinc binding to the identified elements on the Cav3.2 channel. This model is modified from the structural model of a closed voltage-gated potassium channel (26). For clarity, only the voltage-sensor paddle is shown (left panel). The zinc interacting residues are shown by individual amino acids, of which oxygen, carbon, and nitrogen atoms are marked red, white, and blue, respectively. A possible location of the zinc ion is displayed with a blue sphere. Because the amino acid sequence of the IS3–IS4 linker of Cav3.2 is profoundly different from the Shaker potassium channel, we modeled the Asp-Gly-His motif to that predicted for the albumin ATCUN motif (29).
FIGURE 6.
Zinc inhibition of ionic and gating currents of Cav3.2 expressed in HEK-293 cells. Whole cell patch recordings were made using the Hh8-5 stable cell line expressing Cav3.2 (41). Currents were measured using 10 mm Ca2+ in the external solution and 155 TEA-CI in the internal solution as charge carriers. A, ionic currents recorded during a step pulse to −20 mV from a holding potential of −100 mV. Currents were measured before (control) and after application of 10 μm ZnCl2. B, average results of the peak currents were measured in control and zinc. C, gating currents were measured from the same cell using a step depolarization to +55 mV. Each trace represents the average of 20 consecutive sweeps. As in A, data recorded in the presence of zinc is represented by a thick line. D, average gating currents were normalized to control. Control 2 is a time-matched control where cells were continuously perfused with the 10 mm external solution.
DISCUSSION
We recently identified His191 in the IS3–IS4 extracellular linker as a critical determinant of nickel, copper, zinc, and redox sensitivity of Cav3.2 (15–17). To localize other partners of His191 required for interacting with high affinity zinc, we systematically transferred various portions in domain I of Cav3.2 into Cav3.1/Q172H, which was only slightly more sensitive to zinc than wild-type Cav3.1. Analysis of their zinc dose-response relationships revealed that introduction of the region(s) only including the IS3–IS4 loop of Cav3.2 into Cav3.1/Q172H (or Cav3.1) dramatically increased its zinc sensitivity, indicating that the IS3–IS4 loop contains additional residues involved in metal binding. Consequent point mutations uncovered the role of an “Asp-Gly-His” motif involved in zinc inhibition of Cav3.2. Chimeras and point mutations also revealed an important role of aspartate residues in IS2 in zinc block, thereby providing the basis for a structural model where trace metals bind and block movements of the repeat I voltage-sensor paddle. This hypothesis was supported by measuring the ability of zinc to reduce on-gating charge movements.
The critical role of Gly in the middle of the Asp-Gly-His motif was shown by two experimental results: 1) single mutation of Q172H into Cav3.1 only partially increased zinc sensitivity, but double mutations of L171G and Q172H into Cav3.1 had a larger effect, bringing zinc potency close to that of wild-type Cav3.2, and 2) single mutation of G190L decreased the zinc sensitivity of Cav3.2. Glycine residues are known for providing flexibility for polypeptide(s) because glycine has hydrogen as its side chain, rather than a carbon, as is the case in all other amino acids (28). Based on its structural property, we simply interpret that the Gly in the Asp-Gly-His motif provides not only a deprotonated amide nitrogen atom, but also flexibility for its neighboring Asp189 and His191 residues to interact with zinc more efficiently. This may explain why introduction of histidine into the corresponding position of Cav3.1 was not sufficient to convert zinc block to the potency observed in Cav3.2. We hypothesize the bulky leucine residue in Cav3.1 interferes with zinc coordination by Asp170 and His172.
The amino-terminal Cu(II)- and Ni(II)-binding (ATCUN) motif of serum albumin is composed of an “Asp/Glu-Ala-His” (29). Based on the similarity of the Cav3.2 Asp-Gly-His motif to the ATCUN motif, we predicted that the acidic residue in the motif would also be critical for the micromolar zinc sensitivity of Cav3.2. This prediction was tested by mutating Asp189 in Cav3.2 into either a neutral residue or another acidic residue, followed by examining their zinc sensitivities. Point mutation of D189A decreased the zinc sensitivity by about 10-fold. Notably, point mutation of D189E enhanced the zinc sensitivity by about 3-fold. These results support that, in addition to Gly190 and His191, the preceding Asp189 is another requirement for the high zinc sensitivity to Cav3.2. The structure of the ATCUN motif (Asp/Glu-Ala-His) was proposed to be a pentacoordinated structure formed by the carboxyl group of the acidic residue and four nitrogen ligands (one from the amino terminus, two from the peptide backbone, and one from the imidazole nitrogen) (29). In addition to an ATCUN motif, our data support the hypothesis that Cav3.2 includes an additional ligand, coming from the carboxyl group of the aspartate residue in IS2, thereby providing a model of the zinc binding site in Cav3.2.
Previous studies have established that negatively charged residues in transmembrane segments S1, S2, and S3 also act as parts of voltage sensing machinery by electrostatically interacting with positively charged residues in the S4 segment of voltage-gated ion channels (30). The electrostatic interaction between charged residues helps fold the channel in a proper conformation that efficiently targets the channels to the plasma membrane (31, 32). Papazian and colleagues (33, 34) also found that divalent ions such as nickel and magnesium can bind to the extracellular ion binding pocket formed by the negatively charged residues at S2 and S3 in ether-à-go-go K+ channels, decelerating activation kinetics and/or inhibition of currents. Based on these previous findings, we tested whether the negatively charged residue(s) at the outer portion of IS2 were involved in zinc sensitivity. Sequence comparison showed that Cav3.1 has Phe-Asp-Asp, whereas Cav3.2 has Phe-Asp-Ala at this position. Cav3.1/D122A+L171G+Q172H, constructed by replacing Phe-Asp-Asp of Cav3.1/L171G+Q172H with Phe-Asp-Ala, was 4.6-fold more sensitive to zinc than Cav3.1/L171G+Q172H. Cav3.2/A141D constructed by mutating Phe-Asp-Ala into Phe-Asp-Asp became 4.2-fold less sensitive to zinc than wild-type Cav3.2. These data imply that the presence of one acidic residue in the middle of this triplet renders higher zinc sensitivity than that of two acidic residues. This implication is further supported by the findings that the presence or absence of a negatively charged residue in the middle of the triplet significantly affected zinc inhibition sensitivity (Cav3.2/D140E versus Cav3.2/D140A, IC50 = 3.4 ± 0.7 versus 7.9 ± 1.3 μm). These results suggest that Phe-Asp-Ala at the outer portion of IS2 is more favorable than the other sequence combinations in improving zinc inhibition sensitivity, together with the major contribution of the Asp-Gly-His motif.
All of the mutant channels where the negatively charged residue at the outer portion of IS2 was neutralized showed a common positive shift of their activation and inactivation curves compared with those of wild-type Cav3 (Table 1). For example, activation and inactivation curves of Cav3.2/D140A were positively shifted by ∼10 mV, and its functional expression in Xenopus oocytes was dramatically decreased as well (data not shown). The voltage dependence of activation and inactivation of all the other mutants were similar to their respective wild-type channels, ruling out any artifactual shift in sensitivity due to less activation under the voltage protocols used. By analogy to findings with K+ channels, the decrease in expression may suggest that Asp140 is electrostatically interacting with positively charged residues in IS4, influencing structural conformation of the channel and trafficking of the channels to the plasma membrane (30, 35).
We identified two structural elements critical for rendering the high zinc sensitivity to Cav3.2: the Asp-Gly-His motif in the IS3–IS4 loop and an Asp residue in IS2. Based on the chimeric approach used in this study, we cannot rule out possible involvement of conserved residues in other parts of the channel, such as S4 voltage-sensor paddle regions, as identified in Cu2+ block of BK channels (36). In addition, this study does not localize the lower affinity zinc binding sites involved in block of Cav3.1. Although the underlying mechanism of how the negatively charged residue in IS2 contributes to zinc sensitivity remains to be further investigated, a simple interpretation is that the acidic residue acts as another member for zinc coordination with the Asp-Gly-His motif, based on the findings that its mutation into a neutral residue decreases zinc sensitivity. The ability of zinc to decrease gating charge 25% is consistent with immobilization of one of the four voltage-sensor paddles. It is interesting to note that repeat I plays a dominant role in the opening of both HVA and LVA channels (37, 38). These studies also provide evidence for considerable structural similarity between voltage-gated K+ and Ca2+ channels, and combined with the established role of Cav3.2 in pain and epilepsy, provide a structural model for the development of novel therapeutics (17, 24, 39, 40).
Acknowledgment
We thank Benoit Roux for the file used to make Fig. 5.
This work was supported by Korea Science and Engineering Foundation Grant R11-2009-036-01003-0, Priority Research Centers Program 2009-0093822, and Basic Science Research Program 2009-0087964 through the National Research Foundation of Korea, and a grant from the University of Virginia (to E. P. R.).
- LVA
- low voltage activated
- HVA
- high voltage activated
- HEK
- human embryonic kidney
- TEA
- tetraethylammonium.
REFERENCES
- 1.Perez-Reyes E. (2003) Physiol. Rev. 83, 117–161 [DOI] [PubMed] [Google Scholar]
- 2.Iftinca M. C., Zamponi G. W. (2009) Trends Pharmacol. Sci. 30, 32–40 [DOI] [PubMed] [Google Scholar]
- 3.Kozlov A. S., McKenna F., Lee J. H., Cribbs L. L., Perez-Reyes E., Feltz A., Lambert R. C. (1999) Eur. J. Neurosci. 11, 4149–4158 [DOI] [PubMed] [Google Scholar]
- 4.Chemin J., Monteil A., Perez-Reyes E., Bourinet E., Nargeot J., Lory P. (2002) J. Physiol. 540, 3–14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ellinor P. T., Yang J., Sather W. A., Zhang J. F., Tsien R. W. (1995) Neuron 15, 1121–1132 [DOI] [PubMed] [Google Scholar]
- 6.Tsien R. W., Hess P., McCleskey E. W., Rosenberg R. L. (1987) Annu. Rev. Biophys. Biophys. Chem. 16, 265–290 [DOI] [PubMed] [Google Scholar]
- 7.Fox A. P., Nowycky M. C., Tsien R. W. (1987) J. Physiol. 394, 149–172 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Narahashi T., Tsunoo A., Yoshii M. (1987) J. Physiol. 383, 231–249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lee J. H., Gomora J. C., Cribbs L. L., Perez-Reyes E. (1999) Biophys. J. 77, 3034–3042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Talavera K., Staes M., Janssens A., Klugbauer N., Droogmans G., Hofmann F., Nilius B. (2001) J. Biol. Chem. 276, 45628–45635 [DOI] [PubMed] [Google Scholar]
- 11.Obejero-Paz C. A., Gray I. P., Jones S. W. (2004) J. Gen. Physiol. 124, 631–640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Traboulsie A., Chemin J., Chevalier M., Quignard J. F., Nargeot J., Lory P. (2007) J. Physiol. 578, 159–171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Swenson R. P., Jr., Armstrong C. M. (1981) Nature 291, 427–429 [DOI] [PubMed] [Google Scholar]
- 14.Jeong S. W., Park B. G., Park J. Y., Lee J. W., Lee J. H. (2003) Neuroreport 14, 1537–1540 [DOI] [PubMed] [Google Scholar]
- 15.Kang H. W., Park J. Y., Jeong S. W., Kim J. A., Moon H. J., Perez-Reyes E., Lee J. H. (2006) J. Biol. Chem. 281, 4823–4830 [DOI] [PubMed] [Google Scholar]
- 16.Nelson M. T., Joksovic P. M., Su P., Kang H. W., Van Deusen A., Baumgart J. P., David L. S., Snutch T. P., Barrett P. Q., Lee J. H., Zorumski C. F., Perez-Reyes E., Todorovic S. M. (2007) J. Neurosci. 27, 12577–12583 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Nelson M. T., Woo J., Kang H. W., Vitko I., Barrett P. Q., Perez-Reyes E., Lee J. H., Shin H. S., Todorovic S. M. (2007) J. Neurosci. 27, 8250–8260 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kang H. W., Moon H. J., Joo S. H., Lee J. H. (2007) FEBS Lett. 581, 5774–5780 [DOI] [PubMed] [Google Scholar]
- 19.Rogers J. C., Qu Y., Tanada T. N., Scheuer T., Catterall W. A. (1996) J. Biol. Chem. 271, 15950–15962 [DOI] [PubMed] [Google Scholar]
- 20.Chen H., Gordon D., Heinemann S. H. (2000) Pflugers Arch. 439, 423–432 [DOI] [PubMed] [Google Scholar]
- 21.Li-Smerin Y., Swartz K. J. (2000) J. Gen. Physiol. 115, 673–684 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Swartz K. J. (2007) Toxicon 49, 213–230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ramsey I. S., Moran M. M., Chong J. A., Clapham D. E. (2006) Nature 440, 1213–1216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Shin H. S., Cheong E. J., Choi S., Lee J., Na H. S. (2008) Curr. Opin. Pharmacol. 8, 33–41 [DOI] [PubMed] [Google Scholar]
- 25.Long S. B., Tao X., Campbell E. B., MacKinnon R. (2007) Nature 450, 376–382 [DOI] [PubMed] [Google Scholar]
- 26.Chanda B., Asamoah O. K., Blunck R., Roux B., Bezanilla F. (2005) Nature 436, 852–856 [DOI] [PubMed] [Google Scholar]
- 27.Vitko I., Shcheglovitov A., Baumgart J. P., Arias-Olguín I. I., Murbartián J., Arias J. M., Perez-Reyes E. (2008) PLoS ONE 3, e3560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jacob J., Duclohier H., Cafiso D. S. (1999) Biophys. J. 76, 1367–1376 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Laussac J. P., Sarkar B. (1980) J. Biol. Chem. 255, 7563–7568 [PubMed] [Google Scholar]
- 30.Planells-Cases R., Ferrer-Montiel A. V., Patten C. D., Montal M. (1995) Proc. Natl. Acad. Sci. U.S.A. 92, 9422–9426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sato Y., Sakaguchi M., Goshima S., Nakamura T., Uozumi N. (2003) J. Biol. Chem. 278, 13227–13234 [DOI] [PubMed] [Google Scholar]
- 32.Zhang L., Sato Y., Hessa T., von Heijne G., Lee J. K., Kodama I., Sakaguchi M., Uozumi N. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 8263–8268 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Silverman W. R., Roux B., Papazian D. M. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 2935–2940 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Silverman W. R., Tang C. Y., Mock A. F., Huh K. B., Papazian D. M. (2000) J. Gen. Physiol. 116, 663–678 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Tiwari-Woodruff S. K., Schulteis C. T., Mock A. F., Papazian D. M. (1997) Biophys. J. 72, 1489–1500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ma Z., Wong K. Y., Horrigan F. T. (2008) J. Gen. Physiol. 131, 483–502 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tanabe T., Adams B. A., Numa S., Beam K. G. (1991) Nature 352, 800–803 [DOI] [PubMed] [Google Scholar]
- 38.Vitko I., Bidaud I., Arias J. M., Mezghrani A., Lory P., Perez-Reyes E. (2007) J. Neurosci. 27, 322–330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bourinet E., Alloui A., Monteil A., Barrère C., Couette B., Poirot O., Pages A., McRory J., Snutch T. P., Eschalier A., Nargeot J. (2005) EMBO J. 24, 315–324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Choi S., Na H. S., Kim J., Lee J., Lee S., Kim D., Park J., Chen C. C., Campbell K. P., Shin H. S. (2007) Genes Brain Behav. 6, 425–431 [DOI] [PubMed] [Google Scholar]
- 41.Gomora J. C., Murbartián J., Arias J. M., Lee J. H., Perez-Reyes E. (2002) Biophys. J. 83, 229–241 [DOI] [PMC free article] [PubMed] [Google Scholar]