Abstract
Timeless (Tim), a core circadian clock gene in Drosophila, is retained in mammals but has no apparent mammalian circadian clock function. Mammalian TIM is essential for ATR-dependent Chk1 activation and S-phase arrest. We report that TIM is likewise essential for ATM-dependent Chk2-mediated signaling of doxorubicin-induced DNA double strand breaks. TIM depletion attenuates doxorubicin-induced G2/M cell cycle arrest and sensitizes cancer cells to doxorubicin-induced cytotoxicity. TIM is, thereby, a potential novel anticancer drug target whose inhibition may enhance the therapeutic cytotoxicity of agents that activate DNA damage pathways as part of their mechanism.
Keywords: Cell Cycle, Checkpoint Control, DNA Damage, PI 3-kinase, siRNA, ATM, Chk2, Tim
Introduction
In response to DNA damage, cells activate checkpoint pathways and arrest the cell cycle at G1, S, and G2/M phases to repair the damaged DNA. If the damage is beyond repair, apoptosis is triggered (1–3). There are two major DNA damage signaling pathways. One is the ATM (ataxia telangiectasia mutated)-Chk2 pathway and the other is the ATR (ataxia telangiectasia and Rad3-related)-Chk1 pathway. Although these two pathways functionally overlap in response to DNA damage and share most downstream effectors, the DNA double strand break primarily activates the ATM-Chk2 pathway, whereas ATR-Chk1 mainly responds to DNA replication fork collapse caused by bulky DNA lesions (4–6).
DNA double strand break (DSB)2 is one of the most common DNA damages induced both endogenously and exogenously. Following DSB, the ATM protein, a phosphatidylinositol 3-kinase-like serine/threonine protein kinase, is activated by autophosphorylation at the Ser1981 site (7). The activated ATM then phosphorylates Chk2 at Thr68, which leads to Chk2 kinase activation. p53, a major effector of the DNA damage response pathway, is expressed at low levels and in an inactive form during normal conditions. Both ATM and Chk2 phosphorylate p53, causing p53 protein stabilization and activation. Activated p53 arrests the cell cycle by inducing the expression of cell cycle inhibitors such as p21. As a result, G1/S, intra-S and G2/M cell cycle checkpoints are activated to stop cell cycle progression and allow the cell to repair its damaged DNA (8–11). p53 also induces genes that will subsequently lead to apoptosis if the DNA cannot be adequately repaired (8–11). Mutations of genes in these damage response pathways can predispose to cancer (12–18). On the other hand, DNA damage-induced DNA repair mechanisms are also at least partially responsible for chemotherapeutic drug resistance of cancer cells. These processes are thereby central to both tumorigenesis and cancer therapy effectiveness.
Mammalian TIMELESS (TIM) protein shares sequence similarity with the Drosophila key circadian clock protein TIM (19). The role of mammalian TIM in circadian clock control is, however, not clear (20–23). Mammalian TIM has been shown to associate with Chk1 to transduce the replication checkpoint signal from ATR to Chk1 when DNA replication is blocked by UV irradiation or hydroxyurea treatment. Further studies have identified mammalian TIM as a replication fork-associated factor. TIM is critical for S-phase checkpoint regulation in response to UV irradiation (24–28). The role of TIM in the ATM-Chk2 pathway has not been reported. Recent screens of mutations in whole genomes of human breast and colorectal cancer have also identified TIM mutations (29).
Doxorubicin (Dox) is one of the most commonly used anticancer agents. Dox stabilizes topoisomerase II, intercalates within the DNA, and generates reactive oxygen species that induce DNA double strand breaks. Dox activates the ATM-Chk2 DNA damage response pathway and arrests the cell cycle at the G2/M phase (30, 31). We find that although TIM is not essential for ATM activation, it is required for ATM-dependent activation of Chk2 and G2/M checkpoint arrest. Down-regulation of TIM sensitizes cancer cells to doxorubicin toxicity.
EXPERIMENTAL PROCEDURES
Cell Culture and siRNA
The human colon cancer cell line HCT116 was obtained from ATCC (Manassas, VA). Cells were maintained in RPMI 1640 medium with 10% fetal bovine serum under conditions of 5% CO2 at 37 °C. For siRNA transfection, cells were plated in 6-well plates the day before to reach 30% confluency by the time of transfection. 100–200 pmol of siRNA was transfected by Lipofectamine 2000 (Invitrogen) according to the protocol provided by the manufacturer. The knockdown effects were examined after 48–72 h of incubation. The human TIM siRNA was synthesized according to Unsal-Kacmaz et al. (27). Sequences of TIM siRNA 2 and ATM were designed by Blockit siRNA Designer (Invitrogene). Sequences of RNA oligos were as follows: control (luciferase), 5′-CGUAACGCGGAAUACUUCGAdTdT-3′ and 5′-UCGAAGUAUUCCGCGUACGdTdT-3′; human TIM, 5′-GUAGCUUAGUCCUUUCAAAdTdT-3′ and 5′-UUUGAAAGGACUAAGCUACdTdT-3′; human TIM 2, 5′-CCUCUUCAAUCGUCUGCUUdTdT-3′ and 5′-AAGCAGACGAUUGAAGAGGdTdT-3′; human ATM, 5′-GCAAGCAGCUGAAACAAAUDTDT-3′ and 5′-AUUUGUUUCAGCUGCUUGCdTdT-3′. ATR siRNA was synthesized according to the sequence used by the others (32, 33): 5′-CCUCCGUGAUGUUGCUUGAdTdT-3′ and 5′-UCAAGCAACAUCACGGAGGdTdT-3′.
Western Blot and Antibodies
After 48 h of siRNA oligo transfection, cells were treated with 0.5 μm doxorubicin (Sigma) for the indicated time. Then cells were washed with phosphate-buffered saline once, and total protein extracts were obtained by incubating in Nonidet P-40 buffer (0.5% Nonidet P-40, 150 mm NaCl, 50 mm Tris-Cl, pH 7.4, protease inhibitors). Proteins were then separated by standard SDS-PAGE and transferred to nitrocellulose membranes. Antibodies against Chk2 Thr68, ATM Ser1981, λ-H2AX, and p-Ser10 H3 were from Cell Signaling Technology (Danvers, MA). Anti-TIM antibody was generated in our lab using a glutathione S-transferase-TIM C terminus (amino acids 920–1196) as an antigen. Other antibodies were from Santa Cruz Biotechnology (Santa Cruz, CA). All experiments were repeated at least three times. Images were quantified by ImageJ software.
Drug Sensitivity Assay and Flow Cytometry
Cells were treated with siRNA for 48 h followed by incubation with indicated concentrations of doxorubicin for 20 h. Cells were washed with phosphate-buffered saline and grown in drug-free normal medium for 3 days. Living cell numbers were measured by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay. For cell cycle distribution analysis, 2 days after siRNA treatment, doxorubicin was added to the medium to a final concentration of 0.5 μm and incubated for 24 h. Cells were then collected and fixed with ethanol and stained with propidium iodide for fluorescence-activated cell sorter analysis.
RESULTS AND DISCUSSION
TIM Is Required for Doxorubicin-induced Chk2 Activation
Dox-induced DNA double strand breaks activate ATM-dependent phosphorylation of its downstream targets including Chk2 and p53 in a dose- and time-dependent manner in the HCT116 colon cancer cell line (34–36). Therefore, to investigate the involvement of TIM in the ATM-Chk2 pathway, DNA damage was induced by treating HCT116 cells with doxorubicin. Phosphorylation of Chk2 Thr68 was detectable after a 1–2-h treatment with 0.5 μm Dox in the control HCT116 cells. The amount of Chk2 Thr68 phosphorylation increased in a time-dependent manner. Dox-induced Chk2 phosphorylation was, however, dramatically diminished in HCT116 cells treated with TIM siRNA, suggesting TIM was essential for Chk2 activation. Dox-induced p53 accumulation was also attenuated in TIM down-regulated cells (Fig. 1, A and B). p21 induction by doxorubicin was also slightly reduced by down-regulation of TIM.
FIGURE 1.
TIM is required for doxorubicin-induced Chk2 activation. A, HCT116 cells were treated with doxorubicin (0.5 μm) for the indicated time after 48 h of siRNA transfection. The levels of p-Thr68 Chk2, total Chk2, p53, and p21 were measured by respective antibodies. GAPDH was the loading control. B, relative abundance of Thr68- phosphorylated Chk2 (upper panel) and p53 (lower panel) after Dox treatment. Amount of p-Thr68 Chk2 was normalized to total Chk2. p53 was normalized to GAPDH. The level before doxorubicin treatment in the control cells was set as 1 (*, p < 0.01). C, human TIM was down-regulated by another siRNA oligo. Doxorubicin-induced Chk2 Thr68 phosphorylation levels in control and TIM siRNA 2-treated HCT116 cells were compared. D, mouse TIM restores Dox-induced Chk2 Thr68 phosphorylation in endogenous TIM down-regulated human cells. HCT116 cells were cotransfected with TIM siRNA oligo and empty pCDNA3 plasmids or pCDNA3-V5-mTIM plasmids. Chk2 Thr68 phosphorylation levels were measured after doxorubicin treatment for the indicated time.
To confirm that TIM-mediated Chk2 activation was specific, two more control experiments were performed. We designed another anti-human TIM siRNA oligo (TIM siRNA 2). Knocking down TIM expression by this siRNA oligo also inhibited Dox-induced Chk2T68 phosphorylation (Fig. 1C). We also cotransfected anti-human TIM siRNA with empty pCDNA3 plasmids or V5-tagged mouse TIM. Exogenous mouse TIM restored Dox-induced Chk2 Thr68 phosphorylation in endogenous TIM down-regulated HCT116 cells (Fig. 1D).
Dox-induced Chk2 Activation Depends on ATM, Not ATR
It has been reported that doxorubicin induces ATM-dependent Chk2 activation. We confirmed this by down-regulation of ATM. Dox-induced Chk2 Thr68 phosphorylation was inhibited in HCT116 cells treated with TIM siRNA (Fig. 2A) or ATM siRNA (B). Because DNA double strand breaks also lead to ATR activation (37, 38) and ATR can activate Chk2, we determined whether the effect of TIM on Chk2 activation was ATR-dependent. Down-regulation of ATR did not prevent Dox-induced Chk2 Thr68 phosphorylation, whereas down-regulation of TIM or TIM and ATR abolished Chk2 Thr68 phosphorylation (Fig. 2C). Therefore, TIM was required for ATM-mediated Chk2 activation.
FIGURE 2.
Doxorubicin-induced Chk2 activation is ATM-dependent. A and B, the expression of TIM or ATM in HCT116 cells were down-regulated by siRNA. C, TIM, ATR, or both were down-regulated by siRNA. Chk2 Thr68 phosphorylation was determined after 4 h of doxorubicin treatment (0.5 μm). GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
TIM Is Not Required for ATM Activation or H2AX Phosphorylation
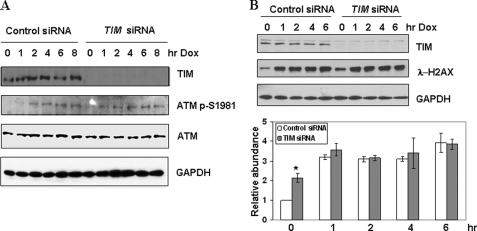
To determine whether doxorubicin could activate ATM in the absence of TIM protein, the phosphorylation of Ser1981 of ATM was examined. In both control and TIM down-regulated cells, ATM was phosphorylated at its Ser1981 site following Dox treatment, indicating that TIM was not essential for ATM activation (Fig. 3A). The result suggests that TIM is involved in ATM downstream signaling.
FIGURE 3.
TIM is not required for doxorubicin-induced phosphorylation of ATM and H2AX. HCT116 cells were treated as in Fig. 1A. The levels of Ser1981-phosphorylated ATM (A) and λ-H2AX (B) were determined by specific antibodies. The levels of λ-H2AX were quantified (B, lower panel). Its level in the control cells before doxorubicin treatment was set as 1 (*, p < 0.01). GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
Chk2 is one of the key ATM substrates. There are several other DNA damage response proteins that are phosphorylated by ATM following DNA double strand breaks. Histone H2AX is rapidly phosphorylated at serine 139 site (λ-H2AX) when DSB is introduced (39–41). Formation of λ-H2AX foci at the damage sites is critical for DSB repair. It has been shown that DNA DSB-induced λ-H2AX accumulation is inhibited in ATM-null cells (39). We observed repeatedly that the basal level of λ-H2AX was higher in TIM knockdown cells, indicating that cells might have increased spontaneous DNA damage without TIM. In both the control and TIM down-regulated cells, λ-H2AX levels accumulated following doxorubicin treatment (Fig. 3B), suggesting TIM might be required specifically for ATM-dependent Chk2 activation. It will be important to further determine whether TIM is necessary for ATM to phosphorylate other substrates such as SMC1, BRCA1, and p53 Ser15 (42).
TIM Is Involved in G2/M Checkpoint
The ATM-Chk2 pathway is crucial for DNA double strand break-induced G2/M arrest. Cell cycle progression from G2 into M depends upon the cyclin B-Cdc2 complex (43, 44). The activity of cyclin B-Cdc2 is negatively regulated by phosphorylation of Cdc2 at Thr14 and Tyr15 sites. Cyclin B is the positive regulator of Cdc2. The phosphorylated, inactive Cdc2 remains in the cytoplasm. Cdc25C phosphatase dephosphorylates and therefore activates Cdc2. WEE1 kinase phosphorylates Cdc2 and inhibits G2/M transition. Cyclin B and WEE1 protein levels are high during the G2 phase of the cell cycle and begin to decline following the completion of cell division (45–47). In control cells, cyclin B protein accumulated in a time-dependent manner following Dox treatment, whereas the levels of cyclin A remained unchanged (Fig. 4A). Doxorubicin failed to cause accumulation of cyclin B protein level in TIM knockdown cells. WEE1 protein increased modestly after doxorubicin treatment in the control cells. This Dox-induced WEE1 increase was also diminished in TIM down-regulated cells. These results suggest that doxorubicin-induced G2/M arrest, in part, requires TIM protein.
FIGURE 4.
TIM down-regulation attenuates doxorubicin-induced G2/M cell cycle arrest. After 48 h of siRNA treatment, HCT116 cells were treated with 0.5 μm of doxorubicin for the indicated time. The levels of Cyclin B1 and WEE1 (A), and H3 Ser10 phosphorylation (B, upper panel) in total cell extracts were measured by immunoblot. Levels of H3 p-Ser10 were quantified (B, lower panel). Its level in the control cells before Dox treatment was set as 1. C, TIM depletion attenuates G2/M cell cycle arrest after doxorubicin treatment. Fluorescence-activated cell sorter profile of control (left panels) and TIM siRNA (right panels)-transfected HCT116 cells before (upper panels) and after 24 h of 0.5 μm doxorubicin treatment (lower panels). GAPDH, glyceraldehyde-3-phosphate dehydrogenase.
Another marker that is related to cell cycle progression and DNA damage response is the phosphorylation of histone H3 at the serine 10 site. H3 Ser10 phosphorylation is involved in mitotic and meiotic chromosome condensation and associated with the G2/M transition (48, 49). H3 is also phosphorylated and dephosphorylated during stresses such as UV irradiation (50). Induction of H3 phosphorylation may have important roles in both transcriptional regulation and checkpoint arrest. We compared the kinetics of Dox-induced H3 Ser10 phosphorylation in the control and TIM down-regulated cells. The phosphorylated H3 reached peak levels after 4 h of Dox treatment in both the control and TIM siRNA-treated cells. However, Dox-induced H3 Ser10 phosphorylation was significantly lower in TIM down-regulated cells and it disappeared after 8 h of Dox treatment (Fig. 4B). H3 Ser10 phosphorylation was undetectable in both cell lines after 20 h of Dox treatment.
To further determine the role of TIM in Dox-induced G2/M arrest, cell cycle distributions among cells with and without TIM down-regulation after doxorubicin treatment were analyzed by flow cytometry (fluorescence-activated cell sorter). Presence or absence of TIM protein did not alter cell cycle distribution before drug treatment. After 24 h of Dox exposure, TIM down-regulated cells demonstrated lower levels of cells in G2/M compared with control cells (Fig. 4C).
TIM Depletion Increases Sensitivity to Doxorubicin
Most cytotoxic anticancer drugs damage DNA. These drugs often activate DNA checkpoints permitting attempted DNA repair, which is essential for cell survival. Therefore, the activation of DNA damage response pathways may reduce the cytotoxicity of many of these anticancer drugs. Conversely, adding checkpoint inhibitors to DNA-damaging cytotoxic drugs can enhance their cytotoxicity. Because TIM is required for doxorubicin-induced Chk2 activation, we examined the sensitivity of HCT116 cancer cells to doxorubicin when TIM was down-regulated. As expected, down-regulation of ATM or ATR sensitized cells to doxorubicin. Down-regulation of TIM by siRNA also significantly increased doxorubicin toxicity (Fig. 5). Depletion of TIM in ATR down-regulated cells rendered cells more sensitive to doxorubicin than ATR down-regulation alone. This result suggests the potential utility of TIM inhibition to enhance the cytotoxic effectiveness of chemotherapeutic drugs known to activate DNA response pathways within cancer cells. Also, mutation of TIM in human cancers may predict altered drug sensitivity. Altogether, these results suggest that the development of TIM-targeted anticancer drugs may be a novel strategy to improve cancer control.
FIGURE 5.
TIM depletion sensitizes cells to doxorubicin cytotoxicity. After 48 h of siRNA transfection, HCT116 cells were treated with the indicated concentrations of doxorubicin for 20 h. Cells were then incubated in drug-free medium for 3 days, and relative cell viability was determined by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay.
Acknowledgments
We thank Drs. Reppert and Weaver for mouse V5-Tim plasmid, Drs. Zhengguan Yang and Mike Wyatt for advice, and Dr. Dinah Quiton for help with manuscript preparation.
Footnotes
- DSB
- double strand break
- Dox
- doxorubicin
- siRNA
- small interfering RNA.
REFERENCES
- 1.Harper J. W., Elledge S. J. (2007) Mol. Cell 28, 739–745 [DOI] [PubMed] [Google Scholar]
- 2.McGowan C. H., Russell P. (2004) Curr. Opin. Cell Biol. 16, 629–633 [DOI] [PubMed] [Google Scholar]
- 3.Zhou B. B., Elledge S. J. (2000) Nature 408, 433–439 [DOI] [PubMed] [Google Scholar]
- 4.Bartek J., Lukas J. (2003) Cancer Cell 3, 421–429 [DOI] [PubMed] [Google Scholar]
- 5.Shiloh Y. (2003) Nat. Rev. Cancer 3, 155–168 [DOI] [PubMed] [Google Scholar]
- 6.Kastan M. B., Lim D. S. (2000) Nat. Rev. Mol. Cell Biol. 1, 179–186 [DOI] [PubMed] [Google Scholar]
- 7.Bakkenist C. J., Kastan M. B. (2003) Nature 421, 499–506 [DOI] [PubMed] [Google Scholar]
- 8.Ryan K. M., Phillips A. C., Vousden K. H. (2001) Curr. Opin. Cell Biol. 13, 332–337 [DOI] [PubMed] [Google Scholar]
- 9.Prives C., Hall P. A. (1999) J. Pathol. 187, 112–126 [DOI] [PubMed] [Google Scholar]
- 10.Shieh S. Y., Ahn J., Tamai K., Taya Y., Prives C. (2000) Genes Dev. 14, 289–300 [PMC free article] [PubMed] [Google Scholar]
- 11.Wahl G. M., Carr A. M. (2001) Nat. Cell Biol. 3, E277–286 [DOI] [PubMed] [Google Scholar]
- 12.van Gent D. C., Hoeijmakers J. H., Kanaar R. (2001) Nat. Rev. Genet. 2, 196–206 [DOI] [PubMed] [Google Scholar]
- 13.Reinhardt H. C., Yaffe M. B. (2009) Curr. Opin. Cell Biol. 21, 245–255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lavin M. F. (2008) Nat. Rev. Mol. Cell Biol. 9, 759–769 [DOI] [PubMed] [Google Scholar]
- 15.Perona R., Moncho-Amor V., Machado-Pinilla R., Belda-Iniesta C., Sánchez Pérez I. (2008) Clin. Transl. Oncol. 10, 538–542 [DOI] [PubMed] [Google Scholar]
- 16.Hoeijmakers J. H. (2001) Nature 411, 366–374 [DOI] [PubMed] [Google Scholar]
- 17.Gorgoulis V. G., Vassiliou L. V., Karakaidos P., Zacharatos P., Kotsinas A., Liloglou T., Venere M., Ditullio R. A., Jr., Kastrinakis N. G., Levy B., Kletsas D., Yoneta A., Herlyn M., Kittas C., Halazonetis T. D. (2005) Nature 434, 907–913 [DOI] [PubMed] [Google Scholar]
- 18.Khanna K. K., Jackson S. P. (2001) Nat. Genet. 27, 247–254 [DOI] [PubMed] [Google Scholar]
- 19.Sehgal A., Price J. L., Man B., Young M. W. (1994) Science 263, 1603–1606 [DOI] [PubMed] [Google Scholar]
- 20.Gotter A. L. (2006) Neuroreport 17, 1229–1233 [DOI] [PubMed] [Google Scholar]
- 21.Barnes J. W., Tischkau S. A., Barnes J. A., Mitchell J. W., Burgoon P. W., Hickok J. R., Gillette M. U. (2003) Science 302, 439–442 [DOI] [PubMed] [Google Scholar]
- 22.Gotter A. L., Manganaro T., Weaver D. R., Kolakowski L. F., Jr., Possidente B., Sriram S., MacLaughlin D. T., Reppert S. M. (2000) Nat. Neurosci. 3, 755–756 [DOI] [PubMed] [Google Scholar]
- 23.Zylka M. J., Shearman L. P., Levine J. D., Jin X., Weaver D. R., Reppert S. M. (1998) Neuron 21, 1115–1122 [DOI] [PubMed] [Google Scholar]
- 24.Chou D. M., Elledge S. J. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 18143–18147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gotter A. L., Suppa C., Emanuel B. S. (2007) J. Mol. Biol. 366, 36–52 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Unsal-Kaçmaz K., Chastain P. D., Qu P. P., Minoo P., Cordeiro-Stone M., Sancar A., Kaufmann W. K. (2007) Mol. Cell Biol. 27, 3131–3142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Unsal-Kaçmaz K., Mullen T. E., Kaufmann W. K., Sancar A. (2005) Mol. Cell Biol. 25, 3109–3116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yoshizawa-Sugata N., Masai H. (2007) J. Biol. Chem. 282, 2729–2740 [DOI] [PubMed] [Google Scholar]
- 29.Sjöblom T., Jones S., Wood L. D., Parsons D. W., Lin J., Barber T. D., Mandelker D., Leary R. J., Ptak J., Silliman N., Szabo S., Buckhaults P., Farrell C., Meeh P., Markowitz S. D., Willis J., Dawson D., Willson J. K., Gazdar A. F., Hartigan J., Wu L., Liu C., Parmigiani G., Park B. H., Bachman K. E., Papadopoulos N., Vogelstein B., Kinzler K. W., Velculescu V. E. (2006) Science 314, 268–274 [DOI] [PubMed] [Google Scholar]
- 30.Kurz E. U., Douglas P., Lees-Miller S. P. (2004) J. Biol. Chem. 279, 53272–53281 [DOI] [PubMed] [Google Scholar]
- 31.Mikhailov A., Shinohara M., Rieder C. L. (2004) J. Cell Biol. 166, 517–526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Casper A. M., Nghiem P., Arlt M. F., Glover T. W. (2002) Cell 111, 779–789 [DOI] [PubMed] [Google Scholar]
- 33.Kennedy D. R., Ju J., Shen B., Beerman T. A. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 17632–17637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bruno T., De Nicola F., Iezzi S., Lecis D., D'Angelo C., Di Padova M., Corbi N., Dimiziani L., Zannini L., Jekimovs C., Scarsella M., Porrello A., Chersi A., Crescenzi M., Leonetti C., Khanna K. K., Soddu S., Floridi A., Passananti C., Delia D., Fanciulli M. (2006) Cancer Cell 10, 473–486 [DOI] [PubMed] [Google Scholar]
- 35.Freiberg R. A., Hammond E. M., Dorie M. J., Welford S. M., Giaccia A. J. (2006) Mol. Cell Biol. 26, 1598–1609 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Castedo M., Perfettini J. L., Roumier T., Yakushijin K., Horne D., Medema R., Kroemer G. (2004) Oncogene 23, 4353–4361 [DOI] [PubMed] [Google Scholar]
- 37.Jazayeri A., Falck J., Lukas C., Bartek J., Smith G. C., Lukas J., Jackson S. P. (2006) Nat. Cell Biol. 8, 37–45 [DOI] [PubMed] [Google Scholar]
- 38.Shiotani B., Zou L. (2009) Mol. Cell 33, 547–558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Burma S., Chen B. P., Murphy M., Kurimasa A., Chen D. J. (2001) J. Biol. Chem. 276, 42462–42467 [DOI] [PubMed] [Google Scholar]
- 40.Foster E. R., Downs J. A. (2005) Febs J. 272, 3231–3240 [DOI] [PubMed] [Google Scholar]
- 41.Paull T. T., Rogakou E. P., Yamazaki V., Kirchgessner C. U., Gellert M., Bonner W. M. (2000) Curr. Biol. 10, 886–895 [DOI] [PubMed] [Google Scholar]
- 42.Pommier Y., Weinstein J. N., Aladjem M. I., Kohn K. W. (2006) Clin. Cancer Res. 12, 2657–2661 [DOI] [PubMed] [Google Scholar]
- 43.Toyoshima F., Moriguchi T., Wada A., Fukuda M., Nishida E. (1998) EMBO J. 17, 2728–2735 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Nigg E. A. (2001) Nat. Rev. Mol. Cell Biol. 2, 21–32 [DOI] [PubMed] [Google Scholar]
- 45.Geng L., Zhang X., Zheng S., Legerski R. J. (2007) Mol. Cell Biol. 27, 2625–2635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yarden R. I., Pardo-Reoyo S., Sgagias M., Cowan K. H., Brody L. C. (2002) Nat. Genet. 30, 285–289 [DOI] [PubMed] [Google Scholar]
- 47.Perry J. A., Kornbluth S. (2007) Cell Div 2, 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wei Y., Mizzen C. A., Cook R. G., Gorovsky M. A., Allis C. D. (1998) Proc. Natl. Acad. Sci. U.S.A. 95, 7480–7484 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Prigent C., Dimitrov S. (2003) J. Cell Sci. 116, 3677–3685 [DOI] [PubMed] [Google Scholar]
- 50.Zhong S. P., Ma W. Y., Dong Z. (2000) J. Biol. Chem. 275, 20980–20984 [DOI] [PubMed] [Google Scholar]