Abstract
Background
The emergence of agriculture about 10,000 years ago marks a dramatic change in human evolutionary history. The diet shift in agriculture societies might have a great impact on the genetic makeup of Neolithic human populations. The regionally restricted enrichment of the class I alcohol dehydrogenase sequence polymorphism (ADH1BArg47His) in southern China and the adjacent areas suggests Darwinian positive selection on this genetic locus during Neolithic time though the driving force is yet to be disclosed.
Results
We studied a total of 38 populations (2,275 individuals) including Han Chinese, Tibetan and other ethnic populations across China. The geographic distribution of the ADH1B*47His allele in these populations indicates a clear east-to-west cline, and it is dominant in south-eastern populations but rare in Tibetan populations. The molecular dating suggests that the emergence of the ADH1B*47His allele occurred about 10,000~7,000 years ago.
Conclusion
We present genetic evidence of selection on the ADH1BArg47His polymorphism caused by the emergence and expansion of rice domestication in East Asia. The geographic distribution of the ADH1B*47His allele in East Asia is consistent with the unearthed culture relic sites of rice domestication in China. The estimated origin time of ADH1B*47His allele in those populations coincides with the time of origin and expansion of Neolithic agriculture in southern China.
Background
The major diet shift in recent human history was caused by domestication of plants and animals[1]. During human evolution, diet shifts may create different selective pressures acting on the genetic variations of human populations. Two well-studied examples are the copy number variation of amylase gene for starchy food and the regulatory sequence variations of lactase for milk [2-4]. In southern China, the earliest agriculture started to flourish due to the domestication of rice about 10,000 years ago [5]. Hence, like the amylase gene selected for high copy numbers in agricultural societies including East Asia, the rice-culture-related selection could have been acting on populations living in southern China. Rice has been used as the material to produce fermented food and beverages for a long time in southern China since early Neolithic time. The fermentation helps to preserve and enhance the nutritional value of foods and beverages[6]. However, alcohol can lead to addiction and cause damages to human bodies, including nervous system dysfunction, tumor genesis, innate immune system modulation and fetal alcohol syndrome [7-11]. Therefore, genes involved in the ethanol metabolic pathway might become the target of selection when the ethanol-containing food and beverages had been routinely consumed by Neolithic populations in southern China.
The Class I alcohol dehydrogenase (ADH) is the major enzyme that catalyzes alcohol to acetaldehyde in liver. The Class I ADH genes (ADH1A, ADH1B, and ADH1C) encode three subunits of Class I ADH isoenzymes, i.e. α, β and γ. The well studied sequence polymorphism, ADH1BArg47His (rs1229984) is located in ADH1B. The change of amino acid from Arg to His causes enzymatic activity alteration. The derived allele, ADH1B*47His, changes the pKa of the enzyme from 8.5 to 10.0 which is associated with 40 to 100 fold increase in Km and Vmax of alcohol metabolism [12,13]. A global investigation of the ADH1B*47His allele frequency shows a strong geographic distribution. It is dominant in East Asian populations, but rare in European and African populations[14]. The molecular signature of positive selection on ADH1B have been reported [15,16], and the culture-related selective forces were proposed [17] though no correlation with rice domestication has been tested. We hypothesize that the emergence and expansion of rice domestication during Neolithic time is the driving force, leading to the current regional distribution of the ADH1BArg47His polymorphism in East Asia.
Results
ADH1B*47His allele frequency in East Asian populations
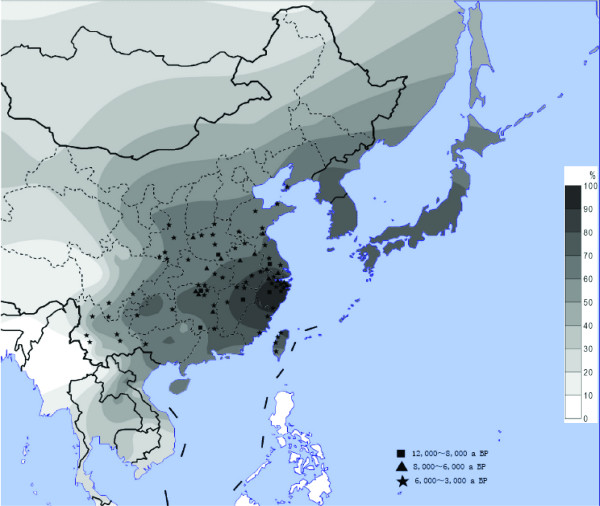
We analyzed a total of 2,275 individuals from 38 East Asian populations, especially those not included in the previous reports (northern Han Chinese, Tibetan and southern ethnic populations in China). Table 1 lists the frequencies of ADH1B*47His in the 38 populations. In general, the distribution pattern is consistent with the previous reports[14,17], and most of the populations (31/38) have frequencies higher than 50%. In Han Chinese, the highest frequency is detected in Zhejiang province of south-eastern China (98.5%), and those in the west have relatively low frequencies (60-70%). The same pattern is also observed for the other ethnic populations from China and Southeast Asia (Cambodia and Thailand) except for Tibetan (14.1% on average), Bulang (1.7%, an ethnic population from south-western China) and Cambodian (20.6%). All the five Tibetan populations from different geographic regions have low frequencies (13-21%). We created a contour map based on the data from the 38 populations and those published before (Figure. 1). The distribution of the frequencies of ADH1B*47His confirms its prevalence in East Asia and a clear east-to-west cline is observed.
Table 1.
The distribution of the ADH1B*47His allele in the 38 East Asian populations.
| Population | Long. (E) | Lat. (N) | Sample Size | Language | ADH1B*47His Frequency (%) |
|---|---|---|---|---|---|
| Han Chinese | 679 | 66.7 | |||
| Jilin | 125 | 44 | 14 | Han | 50 |
| Gansu | 105 | 36 | 174 | Han | 59.8 |
| Shanxi | 112 | 37.5 | 44 | Han | 61.4 |
| Shannxi | 108 | 34.5 | 52 | Han | 62.5 |
| Shandong | 120 | 37 | 90 | Han | 63.3 |
| Henan | 112 | 34 | 56 | Han | 64.3 |
| Anhui | 118 | 33 | 34 | Han | 66.2 |
| Liaonin | 124 | 42 | 28 | Han | 67.9 |
| Hubei | 113 | 30 | 41 | Han | 70.7 |
| Jiangsu | 119 | 34 | 24 | Han | 70.8 |
| Guangxi | 109 | 24 | 9 | Han | 72.2 |
| Guizhou | 107 | 28 | 61 | Han | 75.4 |
| Hunan | 110 | 27.5 | 20 | Han | 82.5 |
| Zhejiang | 121 | 30 | 32 | Han | 98.5 |
| Tibetan | 1088 | 14.1 | |||
| Shannan | 92 | 29 | 34 | Tibeto-Burman | 11.8 |
| Dangxiong | 91 | 30.5 | 816 | Tibeto-Burman | 13.1 |
| Changdou | 97 | 31 | 56 | Tibeto-Burman | 14.3 |
| Rikaze | 89 | 29 | 55 | Tibeto-Burman | 16.4 |
| Qinghai | 96 | 33 | 127 | Tibeto-Burman | 20.5 |
| Bulang | 100 | 23.5 | 30 | Austro-Asiatic | 1.7 |
| Cambodian | 105 | 13 | 17 | Austro-Asiatic | 20.6 |
| Hani | 102.5 | 23 | 30 | Tibeto-Burman | 41.7 |
| Bai | 100 | 25.5 | 30 | Tibeto-Burman | 50 |
| Yi | 101.5 | 24.5 | 30 | Tibeto-Burman | 55 |
| Tujia | 109 | 30 | 31 | Tibeto-Burman | 66.1 |
| Dai | 100 | 23.5 | 30 | Daic | 43.3 |
| Maonan | 108 | 25 | 15 | Daic | 60 |
| Buyi | 106.5 | 26 | 45 | Daic | 63.3 |
| Shui | 108 | 26 | 11 | Daic | 66.7 |
| Chuang | 108 | 23 | 33 | Daic | 72.7 |
| Gelo | 106 | 27 | 10 | Daic | 75 |
| Mulam | 109 | 24 | 12 | Daic | 75 |
| Dong | 58 | 65.5 | |||
| Hunan | 110 | 27.5 | 42 | Daic | 64.3 |
| Guangxi | 109 | 26 | 16 | Daic | 68.7 |
| Yao | 111 | 25 | 58 | Hmong-Mien | 64.7 |
| Miao | 109 | 26 | 29 | Hmong-Mien | 70.7 |
| She | 118 | 27 | 12 | Hmong-Mien | 91.7 |
| Manchu | 125 | 42.5 | 27 | Altai | 66.7 |
Figure 1.
The distribution of the ADH1B*47His allele and the sites of early rice relics. The contour map of the ADH1B*47His frequency in East Asian populations and the ancient sites of rice domestication in China. The allele frequency data includes the 38 populations in the present study and those published before.[17]. The geographic locations of the rice sites are from the published data [5].
Selection on the ADH1B gene
To detect the molecular signature of recent selection on the ADH1B*47 polymorphism, we applied the LRH method and the iHS statistics using the genotype data from the HapMap project. The obtained iHS value for the core SNP (rs1229984) is -2.189 (the empirical p-value is 0.0269), an indication of selection. We then define the core region of ADH1B on the basis of five SNPs (rs4147536, rs1229984, rs1353621, rs1159918 and rs6810842) which determine the East Asian-dominant haplotype. We also select the flanking SNPs, extending both upstream and downstream to 250 Kb, to study the decay of LD from the core haplotype. We plot the haplotype-bifurcation diagrams[18] for the two East Asian populations (Figure. 2) from HapMap (JPT: Japanese in Tokyo, Japan; CHB: Han Chinese in Beijing, China). At a minimum threshold of 9%, we define two core-region haplotypes in the JPT+CHB population. The haplotype CTTCG, which covers the derived variant of ADH1B*47His has an extended predominance by showing a thick branch in the haplotype-bifurcation diagram, clearly suggesting a long-range LD.
Figure 2.

The selection test for the ADH1B gene region. (A) The haplotype-bifurcation diagrams for the core haplotype with at least 9% frequency at the ADH1B gene region in the East Asian populations. The core haplotype CTTCG shows unusual long-range homozygosity. (B) The EHH and REHH plots of the core haplotype covering the candidate SNP (rs1229984) in the East Asian populations. The EHH and REHH values are plotted against the physical distances extending both upstream and downstream of the selected core region. Only the core haplotypes with frequency ≥9% are shown.
The EHH and REHH of the major core haplotypes (≥9%) are plotted against the distance away from the core for the JPT+CHB population (Figure. 2). The EHH of the CTTCG core haplotype decays more slowly than that of the other core haplotype (containing the ancestral variant ADH1B*47Arg) does. In addition, the upstream REHH value of the CTTCG is 17.329 (P = 0.01, by using 1-NORMSDIST). Again, this result is highly consistent with the previous studies, in which the molecular signature of selection was suggested in a wider genomic region containing the ADH1B locus among East Asian populations[15,17]. The selection on ADH1B was also reported previously when the global populations were screened[16]. Additionally, a strong signature of positive selection was detected for the ADH gene cluster in a genome-wide analysis[19]. Collectively, the distribution of ADH1B*47His allele frequency in the populations studied cannot be explained by random genetic drift, and recent selection needs to be invoked.
The time of selection
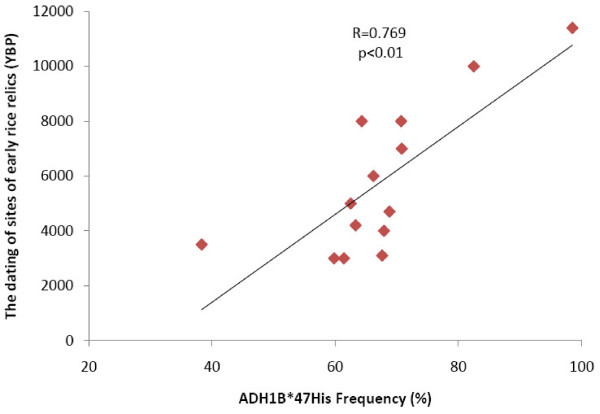
Previous studies suggested a culture-related selection on the ADH1B*47His[17]. To test this, we superimposed the unearthed culture relic sites of rice domestication in East Asia and we observed a significant correlation of the ADH1B*47His allele frequencies with the ages of rice domestication (r = 0.769, p < 0.01, two-tailed t test; Figure 3; see Additional file 1). The origin of rice domestication occurred along the Yangtze River of southern China about 10,000 years ago[20,21]. Based on the culture relics, the earliest rice sites are located in southern and south-eastern China (8,000-12,000 YBP), and then expanded to the central parts of China about 3,000-6,000 years ago, reaching Korea and Japan less than 3,000 years ago [22,23]. The spread of rice domestication agrees well with the distribution of ADH1B*47His, implying that rice domestication is likely the force driving up the frequency and expansion of ADH1B*47His in East Asia during the past 10,000 years. To see if the initial increase of ADH1B*47His in East Asia occurred during the same period as the emergence of rice domestication in early Neolithic time, we conducted molecular dating[24] by typing the nearest STR loci (a CATA repeat STR located about 14 Kb upstream to the ADH1B locus, and a ATTC repeat STR located about 35 Kb downstream to the ADH1B locus) in 598 individuals randomly selected from the 38 populations. For phase reconstruction, only homozygous individuals with the ADH1B*47His alleles are included (see Additional file 2). The estimated ages based on the STRs are 5,525 YBP (CATA repeat), and 9,200 YBP (ATTC repeat). Considering that the two STR loci are still far away from the ADH1B locus, we also estimate the age of the ADH1B*47His based on the phased SNP haplotypes from the HapMap dataset (see Additional file 3). With the fine-scale genetic map, we selected 19 contiguous polymorphic SNPs to estimate the age (Table 2). Surprisingly, the estimated ages are extremely different between the upstream SNPs (114,693-208,919 yrs, 95% confidence interval) and the downstream SNPs (7,338-9,948 yrs, 95% confidence interval), which is due to the dramatic change of recombination rates in the studied genomic region. As suggested, the method based on the moments estimator[24] is not suitable for the region of low average recombination rates. The previous genomic study based on the HapMap SNPs also excluded the regions with low average recombination rate[25]. Therefore, the age estimated based on the downstream SNPs seems to reflect the real age of ADH1B*47His allele, which is also consistent with the ages estimated from the STR variations. Taken together, the age of the derived allele at the ADH1B locus falls in the range of 10,000-7,000 years before present.
Figure 3.
The correlation test for ADH1B*47His allele frequencies with the ages of rice domestication. The correlation of the ADH1B*47His allele frequencies with the ages of rice domestication in 14 regions of China. The data of rice domestication was collected from the published study [5]. The correlation analysis was conducted with the use of SPSS13.0, and the statistical significance was accessed by t test.
Table 2.
Estimated allele ages for ADH1B*47His.
| SNP Frequency(%) | |||||
|---|---|---|---|---|---|
| Polymorphism | Distance to ADH1B*47His (bp) | Genetic distance to ADH1B*47His (cM) | G | A | Allele Age(years) |
| Upstream SNP | |||||
| rs4147531 | -27122 | 0.011537 | 80.95 | 89.86 | 164,811 |
| rs1229966 | -25886 | 0.009885 | 66.67 | 89.86 | 91,782 |
| rs12507573 | -12995 | 0.007675 | 80.95 | 89.86 | 247,742 |
| rs1042026 | -10853 | 0.007588 | 66.67 | 89.86 | 119,567 |
| rs17033 | -10374 | 0.007568 | 80.95 | 89.86 | 251,245 |
| rs2066701 | -906 | 0.007073 | 66.67 | 89.86 | 128,272 |
| rs2075633 | -321 | 0.007021 | 66.67 | 89.86 | 129,222 |
| Average | 161,806 (SD = 63,596) | ||||
| Downstream SNP | |||||
| rs1159918 | 3690 | 0.007325 | 33.33 | 98.55 | 7,494 |
| rs9307239 | 7618 | 0.053403 | 47.62 | 86.96 | 13,406 |
| rs2213041 | 8032 | 0.054139 | 90.48 | 98.55 | 7,621 |
| rs1789891 | 11100 | 0.055169 | 90.48 | 98.55 | 7,479 |
| rs1789982 | 11339 | 0.055228 | 90.48 | 98.55 | 7,470 |
| rs2173201 | 11651 | 0.055302 | 38.10 | 85.51 | 12,059 |
| rs1154433 | 14389 | 0.056568 | 90.48 | 98.55 | 7,294 |
| rs1229978 | 16880 | 0.056640 | 90.48 | 98.55 | 7,284 |
| rs2298753 | 18588 | 0.056670 | 90.48 | 98.55 | 7,281 |
| rs1614972 | 18838 | 0.056678 | 38.10 | 85.51 | 11,765 |
| rs1789898 | 19017 | 0.056679 | 90.48 | 98.55 | 7,280 |
| rs1662060 | 20522 | 0.056693 | 90.48 | 98.55 | 7,279 |
| Average | 8,643 (SD = 2,307) | ||||
Discussion
Having established that the rice culture is likely the driving force of selection on the ADH1BArg47His polymorphism, the left question would be to explain the selective advantage of the ADH1B*47His allele. In southern China, people began to make fermented beverages long time ago. The potential benefits of having fermented beverage (or foods) can be explained by ethanol's combined analgesic, disinfectant and profound mind-altering effects[26]. In addition, fermentation helps to preserve and enhance the nutritional value of foods and beverages. Chemical analyses of ancient organics absorbed into pottery jars suggests that the earliest production of rice fermentation was carried out by the Neolithic people who lived in southern China about 9,000 years ago[6], not long after the origin of rice domestication in the same region. We believe that the custom could have prevailed rapidly among those early-agriculture populations in southern China during the Neolithic time, which have lasted thousands of years.
The ADH I has a low Km for ethanol, found in the liver, which metabolizes the most part of ethanol in the body. The derived ADH1B*47His allele is known to metabolize ethanol up to 100 times quicker than the ancestral ADH1B*47Arg allele, providing support that quick eradication of ethanol, and therefore lower local exposure should be protective. The recent case-control studies also suggested that the ADH1B*47His allele is the protective variant [27-30]. The higher metabolic rate of ADH1B*47His may also lead to the accumulation of the toxic aldehyde intermediate that has been commonly associated with the flushing phenotype[31]. An association study in Han Chinese indicates that the individuals carrying ADH1B*47His have the lowest risk for alcoholism[32]. It was suggested that the flushing phenotype is biochemically equivalent to the effects of disulfiram (a drug used to prevent relapse)[33], which can influence drinking behaviour as a way of protection from over consumption of alcohol. It can also protect against the damage to human bodies caused by alcohol consumptions.
Conclusion
In summary, we provide a plausible explanation about the high frequency of the derived ADH1B*47His allele in East Asia. The distribution of the derived ADH1B*47His allele in East Asia can be well explained by the origin and expansion of the Neolithic rice culture, which is so far one of the few cases demonstrating the genetic adaptation of human populations to the dramatic change during Neolithic time. The ethanol intake increased with the origin of rice agriculture in southern China creates a selective pressure on the Neolithic populations, which is similar with the convergent adaptation of human lactase persistence in Africa and Europe along with the emergence of Neolithic cattle farming[4].
Methods
Samples
In this study, a total of 2,275 unrelated samples were collected from 38 populations (Table 1 and Figure 1). The Han Chinese samples were collected from individuals in 14 provincial areas whose geographic origins were assigned according to the birthplaces of their four grandparents, covering the major geographic regions in China. The other ethnic populations were sampled from south-western China and Qinghai-Tibet plateau where about 80% Chinese ethnic populations live with inhabited histories longer than 3,000 years[34]. All the samples were collected with informed consent. The protocol of this study was approved by the institutional review board of Kunming Institute of Zoology, Chinese Academy of Sciences.
Markers and Genotyping
Initially all the samples were genotyped for the ADH1BArg47His polymorphism (rs1229984). For genotyping by PCR-RFLP, we designed primers (forward primer, 5'FAM-GATTAGTAGCAAAACCCTCAAATAC-3'; reverse primer, 5'-CTAACCATGTGGTCATCTGCG-3') to cover this region. The restriction endonuclease used is Hin61 (Fermentas Life Sciences). The two contiguous microsatellites (CATA repeats and ATTC repeats) are from the UCSC Genome Database, located on Chr: 4 100471833-1889 and Chr: 4 100423654-3678. Both the microsatellite and the SNP genotyping were carried out by using an ABI 3130 sequencer, and the data was analyzed by using Genemapper software version 3.1 (Applied Biosystems). For haplotype analysis, we used the data from HapMap (Phase2.1)[35].
Test of Recent Selection
We used the iHS statistics[19] to detect recent positive selection. The obtained iHS value for the candidate SNP(rs1229984) and the empirical P-value for ADH1B were calculated by using Haplotter[19]. The extended haplotype homozygosity (EHH) and the relative EHH (REHH)[18] were examined by the Sweep program, using the phased haplotype data set (CHB+JPT) from the HapMap project.
Allele Age Estimation
Allele age calculations are conducted by the standard methods published previously[24,25,36]. In brief: t = [1/ln(1-c)]ln [(x(t)-y)/(1-y)], where t = allele age (in generations), c = recombination rate, x(t) = frequency in generation t, and y = frequency on ancestral chromosomes. We assume the origin of the ADH1B*47His allele is on the background of the ancestral allele haplotype, and the calculation utilizes the value of c, determined from the HapMap project recombination rate database http://hapmap.ncbi.nlm.nih.gov/downloads/recombination/latest/rates/. It should be noted that the East Asian samples from the HapMap are non-family data, therefore, limiting the estimation of recombination rates in these populations. Two types of polymorphism data have been used in our analysis. The phased haplotypes (CHB) containing a 45 Kb fragment, which includes 32 SNPs and covers the ADH1BArg47His polymorphism site, is obtained from the HapMap project website http://www.hapmap.org, and the microsatellites data is collected from the randomly selected 598 individuals. For conversion of time in generations, t, into time in years, a generation time of 25 years is assumed.
Authors' contributions
BS and HS designed the study; BS and YP analyzed data and wrote the paper; HS, YP, HZ, and XQ performed sample collection and genotyping; RM and CX provided part of the samples. All authors read and approved the final manuscript.
Supplementary Material
The ADH1B*47His allele frequencies and the ages of the rice relic sites in 14 regions of China.
The STR haplotypes in the 598 tested individuals.
The phased 32 SNP haplotypes of CHB.
Contributor Information
Yi Peng, Email: pypypy2000@163.com.
Hong Shi, Email: shihong@mail.kiz.ac.cn.
Xue-bin Qi, Email: qixuebin@post.kiz.ac.cn.
Chun-jie Xiao, Email: cjxiao@126.com.
Hua Zhong, Email: zhonghua@mail.genetics.ac.cn.
Run-lin Z Ma, Email: marunlin@mail.genetics.ac.cn.
Bing Su, Email: sub@mail.kiz.ac.cn.
Acknowledgements
We are grateful to all the voluntary donors of DNA samples in this study. We thank Hui Zhang and Yan-jiao Li for their technical help. We also thank Dr. Darren Curnoe for his critical reading of the manuscript. This study was supported by grants from the National 973 project of China (2007CB947701,2007CB815705), the Chinese Academy of Sciences (KSCX1-YW-R-34, Westlight Doctoral Program), the National Natural Science Foundation of China (30525028, 30700445, 30630013 and 30771181), and the Natural Science Foundation of Yunnan Province of China (2007C100M, 2009CD107).
References
- Diamond J. Evolution, consequences and future of plant and animal domestication. Nature. 2002;418(6898):700–707. doi: 10.1038/nature01019. [DOI] [PubMed] [Google Scholar]
- Beja-Pereira A, Luikart G, England PR, Bradley DG, Jann OC, Bertorelle G, Chamberlain AT, Nunes TP, Metodiev S, Ferrand N, Erhardt G. Gene-culture coevolution between cattle milk protein genes and human lactase genes. Nat Genet. 2003;35(4):311–313. doi: 10.1038/ng1263. [DOI] [PubMed] [Google Scholar]
- Perry GH, Dominy NJ, Claw KG, Lee AS, Fiegler H, Redon R, Werner J, Villanea FA, Mountain JL, Misra R, Carter NP, Lee C, Stone AC. Diet and the evolution of human amylase gene copy number variation. Nat Genet. 2007;39(10):1256–1260. doi: 10.1038/ng2123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tishkoff SA, Reed FA, Ranciaro A, Voight BF, Babbitt CC, Silverman JS, Powell K, Mortensen HM, Hirbo JB, Osman M, Ibrahim M, Omar SA, Lema G, Nyambo TB, Ghori J, Bumpstead S, Pritchard JK, Wray GA, Deloukas P. Convergent adaptation of human lactase persistence in Africa and Europe. Nat Genet. 2007;39(1):31–40. doi: 10.1038/ng1946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gong ZT, Chen HZ, Yuan DG, Zhao YG, Wu YJ, Zhang GL. The temporal and spatial distribution of ancient rice in China and its implications. Chinese Sci Bull. 2007;52(8):1071–1079. doi: 10.1007/s11434-007-0130-3. [DOI] [Google Scholar]
- McGovern PE, Zhang JH, Tang JG, Zhang ZQ, Hall GR, Moreau RA, Nunez A, Butrym ED, Richards MP, Wang CS, Cheng GS, Zhao ZJ, Wang CS. Fermented beverages of pre- and proto-historic China. P Natl Acad Sci USA. 2004;101(51):17593–17598. doi: 10.1073/pnas.0407921102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olive MF, Koenig HN, Nannini MA, Hodge CW. Stimulation of endorphin neurotransmission in the nucleus accumbens by ethanol, cocaine, and amphetamine. J Neurosci. 2001;21(23) doi: 10.1523/JNEUROSCI.21-23-j0002.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ridderinkhof KR, de Vlugt Y, Bramlage A, Spaan M, Elton M, Snel J, Band GPH. Alcohol consumption impairs detection of performance errors in mediofrontal cortex. Science. 2002;298(5601):2209–2211. doi: 10.1126/science.1076929. [DOI] [PubMed] [Google Scholar]
- Watanabe H, Yamazaki M, Miyazaki H, Arikawa C, Itoh K, Sasaki T, Maehama T, Frohman MA, Kanaho Y. Phospholipase D2 functions as a downstream signaling molecule of MAP kinase pathway in L1-stimulated neurite outgrowth of cerebellar granule neurons. J Neurochem. 2004;89(1):142–151. doi: 10.1111/j.1471-4159.2004.02308.x. [DOI] [PubMed] [Google Scholar]
- Gu JW, Bailey AP, Sartin A, Makey I, Brady AL. Ethanol stimulates tumor progression and expression of vascular endothelial growth factor in chick embryos. Cancer. 2005;103(2):422–431. doi: 10.1002/cncr.20781. [DOI] [PubMed] [Google Scholar]
- Goral J, Karavitis J, Kovacs EJ. Exposure-dependent effects of ethanol on the innate immune system. Alcohol. 2008;42(4):237–247. doi: 10.1016/j.alcohol.2008.02.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurley TD, Bosron WF, Stone CL, Amzel LM. Structures of 3 Human Beta-Alcohol-Dehydrogenase Variants - Correlations with Their Functional Differences. J Mol Biol. 1994;239(3):415–429. doi: 10.1006/jmbi.1994.1382. [DOI] [PubMed] [Google Scholar]
- Edenberg HJ. Regulation of the mammalian alcohol dehydrogenase genes. Progress in Nucleic Acid Research and Molecular Biology. 2000;64:295–341. doi: 10.1016/s0079-6603(00)64008-4. full_text. [DOI] [PubMed] [Google Scholar]
- Li H, Mukherjee N, Soundararajan U, Tarnok Z, Barta C, Khaliq S, Mohyuddin A, Kajuna SLB, Mehdi SQ, Kidd JR, Kidd KK. Geographically separate increases in the frequency of the derived ADH1B*47His allele in eastern and western Asia. Am J Hum Genet. 2007;81(4):842–846. doi: 10.1086/521201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han Y. Evidence of Positive Selection on a ClassI ADH Locus. The American Journal of Human Genetics. 2007;80:441–456. doi: 10.1086/512485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barreiro LB, Laval G, Quach H, Patin E, Quintana-Murci L. Natural selection has driven population differentiation in modern humans. Nat Genet. 2008;40(3):340–345. doi: 10.1038/ng.78. [DOI] [PubMed] [Google Scholar]
- Li H, Gu S, Cai X, C speed W, J Pakstis A, Golub EI, Kidd JR, kidd KK. Ethnic Related Selection for an ADH Class I Variant within East Asia. PLos one. 2008;3(4):e1881. doi: 10.1371/journal.pone.0001881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sabeti PC, Reich DE, Higgins JM, Levine HZP, Richter DJ, Schaffner SF, Gabriel SB, Platko JV, Patterson NJ, McDonald GJ, Ackerman HC, Campbell SJ, Altshuler D, Cooper R, Kwiatkowski D, Ward R, Lander ES. Detecting recent positive selection in the human genome from haplotype structure. Nature. 2002;419(6909):832–837. doi: 10.1038/nature01140. [DOI] [PubMed] [Google Scholar]
- Voight BF, Kudaravalli S, Wen XQ, Pritchard JK. A map of recent positive selection in the human genome. Plos Biol. 2006;4(3):446–458. doi: 10.1371/journal.pbio.0040072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Normile D. Archaeology - Yangtze seen as earliest rice site. Science. 1997;275(5298):309–309. doi: 10.1126/science.275.5298.309. [DOI] [Google Scholar]
- Lu HY, Liu ZX, Wu NQ, Berne S, Saito Y, Liu BZ, Wang L. Rice domestication and climatic change: phytolith evidence from East China. Boreas. 2002;31(4):378–385. doi: 10.1080/030094802320942581. [DOI] [Google Scholar]
- Akazawa. In: Advances in world Archaeology. Wendorf F, Close AF, editor. Vol. 1. New York-London: Academic press; 1982. Cultural change in prehistoric Japan: Receptivity to rice agriculture in the Japanese archipelago; pp. 151–211. [Google Scholar]
- Akazawa. Origin of rice cultivation in Korean Peninsular. J archaeol (in Japanese) 1984;2:46–49. [Google Scholar]
- Slatkin M, Rannala B. Estimating allele age. Annual Review of Genomics and Human Genetics. 2000;1:225–249. doi: 10.1146/annurev.genom.1.1.225. [DOI] [PubMed] [Google Scholar]
- Hawks J, Wang ET, Cochran GM, Harpending HC, Moyzis RK. Recent acceleration of human adaptive evolution. P Natl Acad Sci USA. 2007;104(52):20753–20758. doi: 10.1073/pnas.0707650104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vallee BL. Alcohol in the Western world. Sci Am. 1998;278(6):80–85. doi: 10.1038/scientificamerican0698-80. [DOI] [PubMed] [Google Scholar]
- Mulligan CJ, Robin RW, Osier MV, Sambuughin N, Goldfarb LG, Kittles RA, Hesselbrock D, Goldman D, Long JC. Allelic variation at alcohol metabolism genes (ADH1B, ADH1C, ALDH2) and alcohol dependence in an American Indian population. Hum Genet. 2003;113(4):325–336. doi: 10.1007/s00439-003-0971-z. [DOI] [PubMed] [Google Scholar]
- Matsuo K, Hiraki A, Hirose K, Ito H, Suzuki T, Wakai K, Tajima K. Impact of the alcohol-dehydrogenase (ADH) 1C and ADH1B polymorphisms on drinking behavior in nonalcoholic Japanese. Hum Mutat. 2007;28(5):506–510. doi: 10.1002/humu.20477. [DOI] [PubMed] [Google Scholar]
- Kim DJ, Choi IG, Park BL, Lee BC, Ham BJ, Yoon S, Bae JS, Cheong HS, Shin HD. Major genetic components underlying alcoholism in Korean population. Hum Mol Genet. 2008;17(6):854–858. doi: 10.1093/hmg/ddm357. [DOI] [PubMed] [Google Scholar]
- Hashibe M, McKay JD, Curado MP, Oliveira JC, Koifman S, Koifman R, Zaridze D, Shangina O, Wunsch V, Eluf J Levi JE, Matos E, Lagiou P, Lagiou A, Benhamou S, Bouchardy C, Szeszenia-Dabrowska N, Menezes A, Dall'Agnol MM, Merletti F, Richiardi L, Fernandez L, Lence J, Talamini R, Barzan L, Mates D, Mates IN, Kjaerheim K, Macfarlane GJ, Macfarlane TV, Simonato L, Canova C, Holcatova I, Agudo A, Castellsague X, Lowry R, Janout V, Kollarova H, Conway DI, McKinney PA, Znaor A, Fabianova E, Bencko V, Lissowska J, Chabrier A, Hung RJ, Gaborieau V, Boffetta P, Brennan P. Multiple ADH genes are associated with upper aerodigestive cancers. Nat Genet. 2008;40(6):707–709. doi: 10.1038/ng.151. [DOI] [PubMed] [Google Scholar]
- Brooks PJ, Enoch MA, Goldman D, Li TK, Yokoyama A. The Alcohol Flushing Response: An Unrecognized Risk Factor for Esophageal Cancer from Alcohol Consumption. Plos Medicine. 2009;6(3):e50. doi: 10.1371/journal.pmed.1000050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen CC, Lu RB, Chen YC, Wang MF, Chang YC, Li TK, Yin SJ. Interaction between the functional polymorphisms of the alcohol-metabolism genes in protection against alcoholism. Am J Hum Genet. 1999;65(3):795–807. doi: 10.1086/302540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oroszi G, Goldman D. Alcoholism: genes and mechanisms. Pharmacogenomics. 2004;5(8):1037–1048. doi: 10.1517/14622416.5.8.1037. [DOI] [PubMed] [Google Scholar]
- ZH W. History of nationalities in China. Beijin: China Social Science Press; 1994. [Google Scholar]
- Stephens M, Donnelly P. A comparison of Bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet. 2003;73(5):1162–1169. doi: 10.1086/379378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang E, Ding YC, Flodman P, Kidd JR, Kidd KK, Grady DL, Ryder OA, Spence MA, Swanson JM, Moyzis RK. The genetic architecture of selection at the human dopamine receptor D4 (DRD4) gene locus. Am J Hum Genet. 2004;74(5):931–944. doi: 10.1086/420854. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The ADH1B*47His allele frequencies and the ages of the rice relic sites in 14 regions of China.
The STR haplotypes in the 598 tested individuals.
The phased 32 SNP haplotypes of CHB.