Abstract
Accumulating evidence indicates that G protein-coupled receptors can assemble as dimers/oligomers but the role of this phenomenon in G protein coupling and signaling is not yet clear. We have used the purified leukotriene B4 receptor BLT2 as a model to investigate the capacity of receptor monomers and dimers to activate the adenylyl cyclase inhibitory Gi2 protein. For this, we overexpressed the recombinant receptor as inclusion bodies in the Escherichia coli prokaryotic system, using a human α5 integrin as a fusion partner. This strategy allowed the BLT2 as well as several other G protein-coupled receptors from different families to be produced and purified in large amounts. The BLT2 receptor was then successfully refolded to its native state, as measured by high-affinity LTB4 binding in the presence of the purified G protein Gαi2. The receptor dimer, in which the two protomers displayed a well defined parallel orientation as assessed by fluorescence resonance energy transfer, was then separated from the monomer. Using two methods of receptor-catalyzed guanosine 5′-3-O-(thio)triphosphate binding assay, we clearly demonstrated that monomeric BLT2 stimulates the purified Gαi2β1γ2 protein more efficiently than the dimer. These data suggest that assembly of two BLT2 protomers into a dimer results in the reduced ability to signal.
Keywords: G Proteins/Coupled Receptors (GPCR), Protein/Folding, Protein/Ligand Binding, Protein/Purification, Receptors, Signal Transduction/G Proteins, Dimer/Monomer, Protein Expression
Introduction
G protein-coupled receptors (GPCRs),5 the largest family of integral membrane proteins (1–3), participate in regulation of most physiological functions and are the targets of 30–50% of currently marketed drugs. In light of their biological and therapeutic importance, gaining detailed knowledge of their structural organization remains one of the most crucial tasks, but also, a great challenge facing modern biomedical research.
Dimerization/oligomerization is a common phenomenon in the GPCR superfamily (4), but its role in the structure, function, and signaling of these receptors still has to be clarified. It is unambiguously evidenced that class C GPCRs exist and function as stable dimers (5). However, whether or not class A GPCR dimerization is necessary for G protein activation is still a crucial biological question (6). Indeed, for rhodopsin-like receptors, a role for monomers and dimers in signal transduction is still a matter of intense debate and investigation (7, 8). Although evidence of GPCR dimerization is accumulating even in native tissues (9), perfect functionality in terms of G protein activation has been reported so far for four different monomeric GPCRs. Indeed, monomers of rhodopsin, β2-adrenergic receptor, neurotensin NTS1 receptor, and opioid μ receptor, efficiently activate their cognate G proteins, i.e. the transducin Gt (10, 11), the stimulatory G protein Gs of adenylyl cyclase (12), the stimulatory G protein Gq of phospholipase C (PLC) (13), and the inhibitory G protein Gi of adenylyl cyclase (14), respectively.
Here, we investigated and compared the efficiency of isolated dimers and monomers of a prototypical GPCR to activate the purified Gi2 protein. As a model, we used the Gi-coupled human leukotriene B4 (LTB4) BLT2 receptor (15, 16), which plays critical roles in inflammation and immunological diseases (17, 18). To produce sufficient amounts of pure and functional BLT2, isolate monomers from dimers, and reconstitute receptor-G protein complexes, we have developed an original strategy. This approach is based first on the fusion of the receptor to an integrin fragment that allowed efficient overexpression in inclusion bodies (IBs) of the prokaryote Escherichia coli, then on refolding and functional purification of the receptor, and finally on size exclusion chromatography (SEC) of the different species. Using the purified preparations of BLT2 monomers and dimers, we provide strong evidence that BLT2 monomers activate the Gi2 protein more efficiently than dimers.
EXPERIMENTAL PROCEDURES
The construction, expression, and purification of the different GPCR fusions, as well as their thrombin cleavage and the subsequent purification of isolated GPCRs are described in the supplemental data.
Refolding of the BLT2 Receptor
The leukotriene BLT2 GPCR, purified by immobilized metal affinity chromatography (IMAC) in elution buffer (20 mm Tris-HCl, pH 8.0, 8 m urea, 0.2% SDS, 150 mm NaCl, protease inhibitors (benzamidine (10 μg/ml), leupeptine (5 μg/ml), and phenylmethylsulfonyl fluoride (10 μg/ml), 100 mm imidazole), was dialyzed overnight at 20 °C in dialysis buffer (25 mm Tris-HCl, pH 8.0, 50 mm NaCl, 0.4% SDS) to eliminate urea and imidazole using Slide-A-Lyser dialysis cassettes (Pierce, 10-kDa molecular mass cut-off). The quantity of BLT2 was determined by UV spectrophotometry using the Beer-Lambert law and calculation of molar extinction coefficients (19). Homogeneity of the preparation was checked by recording the scattering light and fluorescence emission (20). We ensured that the concentration of the GPCR was 0.1–0.5 mg/ml. Refolding of the BLT2 was first developed using a miniaturized protocol allowing comparison of many parameters such as pH, ionic strength, temperature, detergents, concentration of detergents, lipids, and additives like cholesterol. To determine whether exchange of SDS with detergents (Anatrace) and/or lipids (Fluka) could produce active receptors from the denatured purified samples, systematic ligand binding competency of the refolded fraction was measured using [3H]LTB4 (22 Ci/mmol, PerkinElmer Life Sciences). Refolding was conducted at 20 °C after binding of the His6 tag of the GPCR to the Ni-NTA superflow resin (Qiagen). Briefly, 0.5 ml of Ni-NTA superflow slurry was loaded onto a Qiaprep spin column (Qiagen) that had been equilibrated twice with the dialysis buffer. The GPCR sample (0.1–0.5 mg/ml) was loaded onto the Ni-NTA resin by a three-step centrifugation at very low speed (30 × g) for 2 min. After this step, 75–100 μg of the receptor was bound to the resin. Optimized refolding was achieved by low speed centrifugation using 4 × 0.5 ml of detergent buffer (25 mm Tris-HCl, pH 8.0, 50 mm NaCl, n-dodecyl phosphocholine (DPC), and hexadecyl-β-d-maltoside (HDM) (1:1, DPC:HDM ratio), asolectin (1:15, protein:detergent and 1:5, detergent:lipid mass ratios), 0.02% cholesteryl hemisuccinate (CHS)). Elution of the refolded GPCR was performed with 2 × 0.5 ml of elution buffer (25 mm Tris-HCl, pH 8.0, 50 mm NaCl, DPC/HDM/asolectin/CHS and 300 mm imidazole) using a final low-speed centrifugation step. The refolded GPCRs were kept on ice or at 4 °C until use. Determination of the yield of the refolding step and quantification of the protein concentration were performed by UV spectrophotometry as described above (18).
Ligand Affinity Purification of the BLT2 Receptor
To isolate the active population (corresponding to the fraction able to bind the ligand) of the BLT2 receptor, the IMAC-purified GPCR was dialyzed overnight at 20 °C in 25 mm Tris-HCl, pH 8.0, 50 mm NaCl, 0.4% SDS containing 5 mg/ml of asolectin before refolding. Refolding was typically carried out at a protein concentration of 0.1–0.5 mg/ml. The unfolded protein was immobilized onto the Ni-NTA matrix as described above. The resin was then washed with 10 column volumes of a buffer of 12.5 mm sodium borate, 10 mm NaCl, 10% glycerol, pH 9.0, containing DPC and HDM as the detergents (1:1, DPC:HDM ratio), asolectin (1:15, protein:detergent, and 1:5, detergent:lipid mass ratios), and 0.02% CHS. Elution of the refolded protein was carried out with the same buffer containing 0.3 m imidazole. All subsequent steps were carried out at 4 °C. The IMAC-purified and refolded receptor was dialyzed overnight in a buffer containing 12.5 mm sodium borate, pH 7.5, 10 mm NaCl, and DPC/HDM/asolectin/CHS (see above for detergent and lipid concentrations). The unfolded aggregated fractions were removed through a gel filtration chromatographic step on a Sephacryl S200 HR column (1.5 × 100 cm, GE Healthcare) using the same buffer. Finally, the active receptor was ligand affinity purified on a 5bα-bound affinity column. The 5bα antagonist molecule was immobilized through its free carboxylate moiety on an Affi-Gel 102 (Bio-Rad) matrix (21). Immobilization was carried out as described by the manufacturer. The refolded BLT2 receptor was recirculated (flow rate 0.5 ml/min) on the matrix for at least 12 h in the above buffer, and then washed with the same buffer. The receptor was then eluted with the buffer containing 0.1 mm 5bα. The antagonist was removed from the eluted protein by dialysis for 36 h against buffer containing 12.5 mm sodium borate, pH 7.5, 10 mm NaCl and DPC/HDM/asolectin/CHS. Following this step, the BLT2 concentration was in the 10−6–10−7 m range. When necessary, the protein preparation was concentrated by ultrafiltration (Centricon 30 device, Amicon) and subsequently dialyzed to re-equilibrate the detergent concentration.
Radiolabeled Ligand Binding Assays
Binding of a radiolabeled ligand to the purified soluble BLT2 was done by equilibrium dialysis at 18 °C for 24 h. Dialysis cassettes from Dianorm (The Nest Group, Inc.) with two 250-μl cavities separated by high permeability 10-kDa molecular mass cut-off dialysis membranes were used. Binding buffer containing 12.5 mm sodium borate, pH 7.5, 10 mm NaCl, and DPC/HDM/asolectin/CHS was used in all dialysis experiments, with protein concentrations in the 10−7 m range. 200 μl of receptor and 200 μl of tritiated ligand were put in each cavity of the cassettes at the begining of the procedure. The cassettes were held in a multiequilibrium apparatus (Dianorm) to ensure constant stirring of the samples. To measure nonspecific binding, a series of control experiments was done in parallel in which an excess of unlabeled ligand was added together with the radiolabeled ligand on one side of the cassette. Another series was included in the experiment without the receptor at each concentration to control free diffusion of the radiolabeled ligand alone through the dialysis membrane. [3H]LTB4 was used as the radioligand. Affinity of the [3H]LTB4 was directly determined in saturation experiments, with concentrations ranging from 750 to 30 nm. Unlabeled LTB4 was used in the control experiment at 100 μm to measure nonspecific binding. At the end of the equilibrium dialysis, samples were recovered from each cavity of each cassette and the radioactivity determined by scintillation counting. The ligand binding data were analyzed by nonlinear least-squares regression using the computer program Ligand (Elsevier-Biosoft).
Fluorescence-based Ligand Binding Assays
Saturation and competition ligand binding experiments with purified BLT2 monomers and dimers (see below) were performed using fluorescence anisotropy with LTB4 labeled with Alexa Fluor-568 (LTB4-568) following the method described by Sabirsh et al. (22). Briefly, LTB4-568 was produced using LTB4-aminopropylamide (Biomol International Inc.) and amine-reactive succinimidyl ester of Alexa Fluor-568 (Invitrogen). Binding buffer containing 12.5 mm sodium borate, pH 7.5, 10 mm NaCl and DPC/HDM/asolectin/CHS was used for these experiments with BLT2 monomer or dimer concentrations at 10−7 m. Saturation assays were performed using up to 0.9 μm fluorescent LTB4 with or without 100 μm LTB4. For competition assays, fluorescence experiments were carried out at a constant LTB4-568 concentration of 350 nm. The different ligands were added at increasing concentrations and binding lasted for 30 min at 15 °C. Fluorescence anisotropy associated with BLT2-bound LTB4-568 was measured using a Cary Eclipse fluorometer equipped with an anisotropy device. Data were recorded as millianisotropy units as a function of competing ligand concentration and converted as of maximum.
G Protein Coupling Assays
Two different types of experiments were carried out to demonstrate functional coupling of purified receptor monomers and dimers to the Gi protein.
First, a nucleotide exchange assay using the purified Gαi2 subunit was carried out as described by Hamm and colleagues (23). Gαi2 was prepared as already published (24). For measuring systematic activation of the G protein, the basal rate of GTPγS binding was determined by monitoring the relative increase in the intrinsic fluorescence (λexc = 300 nm, λem = 345 nm) of Gαi2 (200 nm of purified Gαi2) in the presence of BLT2 (20 nm, the final receptor to G protein molar ratio is 1:10) in buffer containing 10 mm MOPS, pH 7.2, 130 mm NaCl, and 2 mm MgCl2 for 40 min at 15 °C after the addition of 10 μm GTPγS. The detergent/lipid mixture (DPC/HDM/asolectin/CHS) was kept constant for preserving receptor structure. Similarly, the receptor-catalyzed rate was measured under the same conditions in the presence of 50 μm LTB4. The data were normalized to the baseline (buffer contribution, 0%) and the fluorescence maximum obtained with BLT1 (20 nm in the presence of 1 μm LTB4, 100%). A negative control experiment was also carried out under the same conditions with the recombinant 5-HT4(a) receptor. For kinetic studies, purified Gβ1γ2 subunits (500 nm) were added in the mixture. The β1γ2 subunits of the G protein were prepared as described previously (24). Effects on fluorescence changes were also recorded under the same conditions in the presence of 20 μm BLT2 antagonist LY255283.
Second, we directly measured the amount of GTP binding in a more classical way using radiolabeled [35S]GTPγS. Incorporation of the non-hydrolyzable [35S]GTPγS (1250 Ci/mmol, PerkinElmer Life Sciences) was done in the buffer used for monitoring the intrinsic fluorescence of Gαi2, in the presence of 10 mm MgCl2 instead of 2 mm. [35S]GTPγS was used at ∼2.5 nm (∼1,200,000 dpm/reaction). The BLT2 (monomer, dimer, or mixed populations) and the Gαi2 were added at a 1:1 molar ratio (20 nm each), with an excess of Gβ1γ2 subunits (500 nm). The detergent/lipid mixture (DPC/HDM/asolectin/CHS) was kept constant during the incubation, which was done at 25 °C for 10 min. For measuring the agonist-induced receptor-dependent activity of the Gαi2, 1 μm LTB4 was added in the mixture. Nonspecific binding was determined in the presence of 100 μm GTPγS. The reaction was stopped by putting the tubes in a ice-cold bath. Specific [35S]GTPγS binding (separation of free and bound radiolabeled GTP analogue) was then quantified by equilibrium dialysis using dialysis cassettes and membranes equivalent to those described above for radiolabeled [3H]LTB4 binding studies. To ensure complete equilibration of labeled [35S]GTPγS and cold GTPγS, the dialysis lasted for 4 h and 30 min. Samples were then recovered from each cavity of each cassette and radioactivity was determined by liquid scintillation counting.
Separation of Oligomeric States of BLT2 by Size Exclusion Chromatography
SEC experiments were carried out on a Superose 6 column (16 × 70 mm, GE Healthcare). The column was first equilibrated with 12.5 mm sodium borate, pH 7.5, 10 mm NaCl, DPC/HDM/asolectin/CHS buffer. The ligand affinity-purified BLT2 sample at a 10−6–10−7 m range concentration (1 ml in buffer 12.5 mm sodium borate, pH 7.5, 10 mm NaCl, DPC/HDM/asolectin/CHS) was then loaded on the column and gel filtration was carried out with the equilibration buffer at a 0.2 ml/min flow rate. 0.3-Milliliter fractions were collected. The oligomeric state of the receptor in the different fractions was then assessed using a chemical cross-linking approach with dithiobis(succinimidylpropionate), as previously described (25). Briefly, the different receptor fractions were submitted to cross-linking at room temperature, after addition of dithiobis(succinimidylpropionate) (125 mm stock solution in N,N-dimethylformamide) to a final protein-to-cross-linker molar ratio of 1:10. The optimal cross-linking time value was inferred from a time course analysis of the cross-linked species. The reaction was stopped by addition of glycine to a final concentration of 50 mm. The cross-linked species were then submitted to SDS-PAGE analysis under non-reducing conditions and the intensity of the electrophoretic bands determined by densitometry (public domain NIH Image software). The BLT2 monomer and dimer receptor fractions were used to catalyze exchange of GDP for GTPγS on Gαi2 as described above in the presence of Gβ1γ2 at 1 μm and increasing concentrations in Gαi2. The BLT2 receptor concentration was kept constant at 20 nm in the assays. The duration of the reaction was 15 min in all cases. GTPγS binding in the absence of LTB4 was subtracted from that in the presence of 1 μm LTB4 and the resulting data were analyzed by using a one-site binding equation (Prism Software, GraphPad) to assess the Km values for Gαi2 saturation of receptor-catalyzed GDP/GTPγS exchange. Data were normalized both to the fluorescence maximum observed for saturating G protein concentrations (in the 10 μm range) and to the BLT2 receptor concentration.
Fluorescence Resonance Energy Transfer (FRET) Studies
For the FRET experiments, the BLT2 receptor was labeled at the N or C terminus with Alexa Fluor-488 or Alexa Fluor-568 (Invitrogen) as a donor and acceptor of fluorescence, respectively. N-terminal labeling was carried out as described for the BLT1 receptor (26), using a dye:protein molar ratio of 10:1. Under the conditions used, labeling of ∼0.9 was achieved, as assessed by measuring the absorbance of the protein at 276 nm and that of the dye at its absorbance maximum. C-terminal labeling of the receptor was carried out by using the transglutaminase approach described by Jäger et al. (27). For this, the transglutaminase tag Pro-Lys-Pro-Gln-Gln-Phe-Met was appended at the C-terminal end of the protein (the construction of the expression plasmid is described in supplemental data). The fusion protein α5I-BLT2 was thus purified as described for the wild-type receptor, dialyzed as indicated above, and labeling carried out as described by Jäger et al. (27). Briefly, the protein (10 μm) in 20 mm Tris-HCl, pH 8.0, 150 mm NaCl buffer was incubated with 1 mm Alexa dyes (dye:protein molar ratio 100:1) and 0.5 units of transglutaminase (Sigma) for 24 h at 25 °C in the dark. The unreacted dye was then removed by gel filtration. Labeling yields of 85–90% were achieved under these conditions, as assessed by the absorbance of the protein at 276 nm and that of the dye at its maximum. The fusion partner was then cleaved and removed as described for the wild-type receptor. For dimer assembly, receptor labeled either with the fluorophore donor or acceptor were mixed in equivalent amounts (molar ratio 1:1) before refolding. Then, refolding and purification were carried out as described above for the unlabeled BLT2. Fluorescence emission spectra were recorded at 25 °C between 490 and 700 nm on a Cary Eclipse spectrofluorimeter with excitation at 480 or 570 nm. Buffer contributions were systematically subtracted. The FRET ratio corresponded to the ratio of the acceptor emitted fluorescence at 603 nm following excitation at two different wavelengths, 480 and 570 nm (28).
Statistical Analysis
All data are reported as group mean ± S.E. Statistical significance of the differences between independent groups was assessed by paired t test.
RESULTS
Identification of New Fusion Partners for Efficient GPCR Overexpression in E. coli IBs
To produce recombinant GPCRs in sufficient amounts to reconstitute, in vitro receptor-G protein complexes, we used an approach that had been initially described for the olfactory OR5 receptor and is based on accumulation of the receptor target in E. coli IBs as a fusion protein and on subsequent in vitro refolding (29, 30). Although the concept of this approach has been set up for overexpression and functional refolding of leukotriene BLT1 and serotonin 5HT4(a) receptors (31, 32), a generic and reliable method for GPCR accumulation in IBs has not been reported so far (33). To this end, we explored an alternative based on the identification of novel original fusion partners able to produce full-length receptors in high amounts, and that would be applicable to most GPCRs. We thus defined three criteria: 1) the fusion partner has to be targeted to IBs and highly accumulated when expressed as an isolated protein; 2) based on statistical predictions that correlate targeting of the recombinant protein to IBs with its primary sequence and physicochemical properties (34), both the partner and the fusion have to be classified as insoluble; 3) the length of the fusion partner was limited to a maximum of 600 residues. We thus selected a nonspecific PLC of Bacillus cereus (35), the glutamine phosphoribosyl pyrophosphatase amidotransferase (PurF) of E. coli (36), and a fragment of the extracellular β-propeller domain of the human α5 integrin (α5I) (37). According to statistical predictions (34), the two most important parameters controlling inclusion body formation (or insolubility) are charge average and fraction of β-turn-forming residues. As indicated in Table 1, the PLC, PurF, and α5I all possess a higher β-turn-forming residue fraction than the usual partner glutathione S-transferase (GST). Moreover, with respect to charge, both PurF and α5I contain much higher proportions of charged residues than PLC or GST. These observations suggested that PurF and α5I proteins may constitute the best potential candidates.
TABLE 1.
Physicochemical properties of the fusion partners
The physicochemical parameters of each potential fusion partner were calculated according to Wilkinson and Harrison (34). The combination of a high fraction of both charged residues and β-turn forming residues is critical for targeting a recombinant protein to E. coli IBs.
| Charge average | Fraction of β-turn forming residues | Length (amino acid residues) | |
|---|---|---|---|
| % | |||
| GST | −2 | 18.7 | 224 |
| PLC | −2 | 22.7 | 258 |
| PurF | −14 | 21.4 | 504 |
| α5I | −19 | 31.6 | 285 |
GPCR Production and Purification
Both α5I and PurF appeared as very efficient partners for accumulating GPCRs in IBs, based on the data with the human arginine-vasopressin V2 receptor used as a reference (see supplemental Fig. S1). In addition, the α5I allowed efficient overexpression in E. coli IBs and purification of several other class A GPCRs (rhodopsin-like) varying with respect to their length (from 337 to 472 amino acids), physicochemical properties, G protein coupling selectivity, and the nature of their specific ligand (Fig. 1). Although with different efficiencies, high amounts of the catecholamine β3AR, the hormone arginine-vasopressin V2 receptor, and oxytocin (OT) receptor (OTR)), the chemokine CCR5 and CXCR4 receptors or chemokine-like ChemR23 receptor, the cannabinoid CB1 receptor, and bioactive lipid leukotriene BLT2, and CysLT1 and CysLT2 receptors were overexpressed. The V2, β3AR, ChemR23, BLT2, CysLT1, CysLT2, and CB1 receptors were produced and purified as simple α5I fusions (Fig. 1A). The leukotriene BLT1 and arginine-vasopressin V1b receptors were also accumulated and purified using this strategy (data not shown). In addition, the OTR, CXCR4, and CCR5 were only produced as complex α5I-V2 fusions (Fig. 1B). Apparent molecular mass of each fusion was compatible with the corresponding calculated masses (70–75 kDa for the α5I-GPCR fusions, around 120 kDa for the α5I-V2-GPCR fusions). Moreover, the integrity of the different fusions was confirmed both by N-terminal Edman sequencing and chemiluminescence detection of the GPCR C-terminal His6 tag (data not shown).
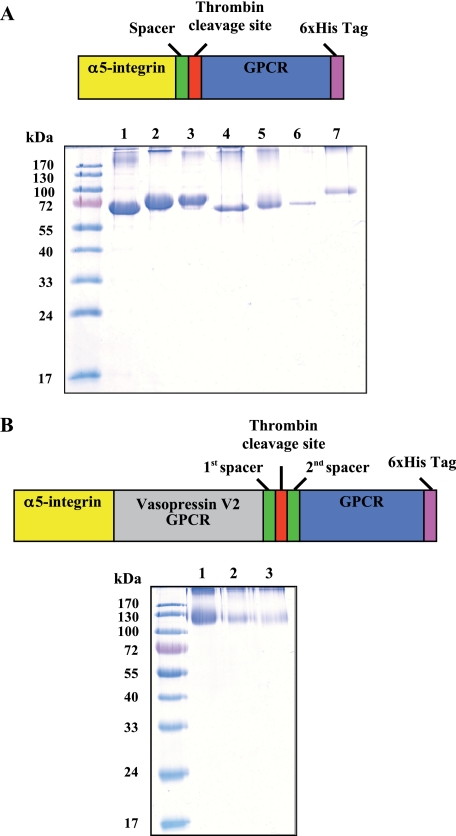
FIGURE 1.
Overexpression and purification of α5I-GPCR fusions. Samples were loaded onto 12% SDS-polyacrylamide gels and proteins stained with Coomassie Blue. To directly compare the purification yield, 10 μl of eluted fusions were put into each well. A, schematic representation of α5I-GPCR fusions and comparison of the corresponding purified proteins: V2, β3AR, ChemR23, BLT2, Cys-LT1, Cys-LT2, or CB1 receptor (lanes 1–7). B, schematic representation of α5I-V2-GPCR fusions and detection of the corresponding purified proteins. The entire α5I-V2 fusion was used as a new partner for expressing another GPCR: OTR, CXCR4, and CCR5 (lanes 1–3).
For each fusion, the α5I partner was efficiently removed by thrombin cleavage and the isolated GPCRs purified using a second IMAC. Complex fusions (e.g. α5I-V2-OTR) required an additional gel filtration step before the IMAC purification for eliminating uncleaved proteins. As illustrated in supplemental Fig. S2, representative purified OTR, ChemR23, V2, and β3AR, appeared as two major bands, corresponding to a monomer at 35–40 kDa and a dimer around 75 kDa. Identity of each receptor monomer and dimer was confirmed by direct N-terminal Edman sequencing. The quantity of each purified receptor was calculated as indicated under “Experimental Procedures”: it varied from 0.2–0.5 mg (OTR for instance) to 2–3 mg (e.g. the V2 or the BLT2) from 100 ml of bacterial cell culture (equivalent to 0.6 g wet cells).
In Vitro Refolding, Functional Purification, and Binding Properties of the Ligand-competent BLT2 Receptor
As stated above, the BLT2 receptor was efficiently accumulated in E. coli IBs and purified as a denatured protein in large quantities. Refolding conditions were subsequently explored as described for the BLT1 receptor (31). The BLT2 receptor was refolded to its native state in well defined detergent/lipid mixed micelles (DPC/HDM/asolectin/CHS). Under such conditions, the ligand-competent fraction represented around 4% of the total receptor preparation, as determined by [3H]LTB4 binding. The ligand-competent fraction of the BLT2 was then purified through a ligand-immobilized affinity chromatography procedure using the LTB4 antagonist 5bα (20). Homogeneity of the affinity-purified BLT2 was demonstrated by binding assays with [3H]LTB4 (Fig. 2). The calculated linear Scatchard plot revealed the presence of a single population of binding sites and a stoichiometric ratio of 1:1 ligand molecule:receptor was calculated (Bmax = 1.06 ± 0.02; n = 3). It has been previously shown with native BLT2 that a receptor molecule binds a single ligand (15). The occurrence of a 1:1 ligand:BLT2 molar ratio with our recombinant pure receptor therefore implies that all receptors in the preparation are in a ligand-competent state.
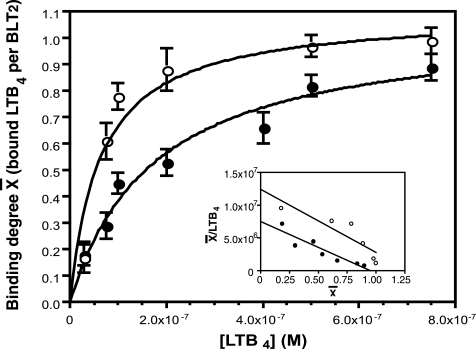
FIGURE 2.
Binding of [3H]LTB4 to the refolded affinity-purified BLT2 receptor. Three series of ligand binding experiments were carried out by equilibrium dialysis in the absence (closed circles) or presence (open circles) of the purified Gαi2 protein (inset, corresponding Scatchard plots). The binding data are presented as a plot of the binding degree x̄ as a function of the ligand concentration. x̄ is defined by bound mole of LTB4 per mol of BLT2 (49). The experiments shown are representative of three independent trials, each performed in duplicate. Kd of [3H]LTB4 was calculated as explained under “Experimental Procedures.” Mean ± S.E. are given under “Results.” The statistics of the illustrated fits were as follow. The calculated Kd in the absence of G proteins was 217.8 ± 47.7 nm (22% error) with a binding ratio of 1.07 ± 0.08 (7.5% error). The calculated Kd in the presence of G proteins was 58.1 ± 18.9 nm (29% error) with a binding ratio of 1.09 ± 0.07 (6.5% error). Error bars are S.E. (calculated as S.D./root square (n −1)).
It is to be noted that, despite this homogeneity, the [3H]LTB4 affinity for the BLT2 (Kd = 232 ± 29 nm (n = 3)) is significantly lower than that described for the receptor expressed in mammalian HEK293 cells, 22.7 nm (15). This could be due to the lack of interaction with stabilizing membrane lipids or protein partners such as G proteins, as it was demonstrated for the other LTB4 receptor BLT1 (31) or the serotonin 5HT4(a) receptor (32). Because BLT2 was demonstrated to couple to Gi protein (15–17), we reconstituted the affinity-purified receptor with the purified Gi2 subunit. A significant increase in the affinity of BLT2 for the LTB4 agonist was observed in the presence of the Gi2 protein (Fig. 2); in that case, the measured Kd was 49 ± 5 nm (n = 3). This value was much closer to that for BLT2 expressed in HEK293 cells, indicating that the agonist high affinity binding state of the refolded affinity-purified BLT2 is dependent on the presence of the G protein.
G Protein Coupling Properties of the Affinity-purified BLT2 Receptor
Functionality of the affinity-purified BLT2 was assessed by monitoring the relative increase in the intrinsic fluorescence of Gi2 after addition of GTPγS to the purified G protein subunit reconstituted with BLT2 in the absence or presence of LTB4. A significant BLT2-catalyzed GTPγS binding occurred upon binding of the LTB4 (Fig. 3A). Moreover, the amount of GTPγS bound to Gi2 was found similar to that induced by LTB4 stimulation of BLT1 receptor, as previously described (24). By contrast, a purified 5HT4(a), which is not naturally coupled to Gi, was unable to stimulate GTPγS binding (32). The kinetics of the G protein Gi2-BLT2 receptor coupling were recorded in the presence of Gβ1γ2 subunits (Fig. 3B) or in their absence. As illustrated, the LTB4-stimulated GTPγS incorporation was fast and saturated within 15 min (profile 2). It remained stable for longer periods of interaction. This result could be mimicked in the absence of the Gβγ subunits, but the rate of increase was slower and saturated within 40 min instead of 15 min (data not shown). As expected, the BLT2-specific antagonist LY255283 was unable to stimulate GTPγS binding (profile 3), and the intrinsic fluorescence signal was equivalent to that of the basal condition (profile 1). Interestingly, the time course and maximal incorporation of GTPγS measured in the presence of BLT2/G protein was equivalent to that recorded with the BLT1 receptor positive control (profile 4). These results confirmed that coupling of the affinity-purified BLT2 receptor to the G protein was functional as well in terms of kinetics, although Gβγ were necessary for rapid GTPγS binding.
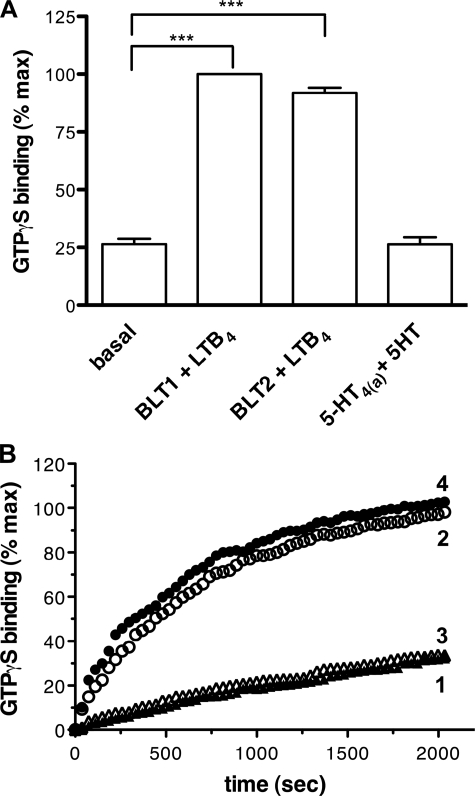
FIGURE 3.
G protein coupling properties of the refolded affinity-purified BLT2 receptor. A, BLT2-catalyzed GTPγS binding assessed by changes in the fluorescence of Gαi2. Experiments were carried out with the refolded BLT2 (20 nm) in the absence or presence of saturating concentrations of LTB4 (50 μm). For comparison, BLT1 and 5HT4(a) were also used at 20 nm and their ligands at 1 and 10 μm, respectively. Mean ± S.E. are shown. Statistics are given: ***, p < 0.01. B, time course of the relative increase in the intrinsic fluorescence of Gαi2 upon addition of GTPγS. The fluorescence was monitored as described under “Experimental Procedures” in the presence of the purified BLT2 receptor in the absence of ligand (profile 1), in the presence of the LTB4 agonist (profile 2), or in the presence of the LY255283 antagonist (profile 3). The data were normalized to the changes induced by the purified BLT1 receptor in the presence of LTB4 (profile 4). The experiment illustrated here is representative of three independent assays. Error bars are S.E. (calculated as S.D./root square (n −1)).
Oligomeric State of the Affinity-purified BLT2 Receptor
We then analyzed the oligomeric state of the ligand affinity-purified BLT2 receptor using a SEC approach. As shown in Fig. 4A, a main peak was observed that was centered at about 70 ml. No peak was observed in the dead volume of the column indicating the absence of the high molecular species that could correspond to aggregated receptor. As clearly shown in Fig. 4A, the main peak displayed a well defined shoulder at about 65 ml indicating the occurrence of different species in the eluted fractions.
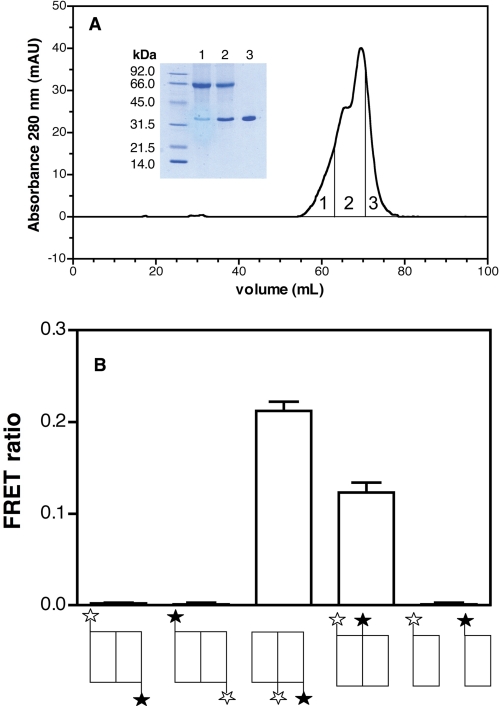
FIGURE 4.
Separation and characterization of monomeric and dimeric species of BLT2 receptor. A, separation of monomeric and dimeric species of BLT2 by SEC. The affinity-purified BLT2 preparation was loaded onto a Superose 6 column and separation of the different species was carried out as described under “Experimental Procedures.” The proteins eluted from the Superose 6 column were pooled in three fractions labeled 1, 2, and 3 as a function of their retention time, as indicated in the elution profile, and submitted to chemical cross-linking. Inset, SDS-PAGE analysis of the protein content in fractions 1–3 after chemical cross-linking. B, FRET ratio measured between Alexa Fluor-488- and Alexa Fluor-568-labeled BLT2 protomers. The species considered in each case are schematically represented below, where ☆ is the fluorescence donor and ★ the acceptor. The upper position represents N-terminal labeling, the lower position corresponds to C-terminal labeling. FRET ratios were calculated as indicated under “Experimental Procedures” (28). The experiments shown in the figure were repeated three times. Results are given as mean ± S.E. Error bars are S.E. (calculated as S.D./root square (n −1)).
To assess the oligomeric state of the different receptor populations in this elution peak, we carried out a series of chemical cross-linking experiments. The different fractions making the elution peak were pooled in three main fractions, labeled 1, 2, and 3 as a function of their elution volume. The proteins in these fractions were then submitted to chemical cross-linking using dithiobis(succinimidylpropionate) as a disulfide reagent, as previously described with the BLT1 receptor (31). The extent of cross-linking was finally assessed by SDS-PAGE under non-reducing conditions (see inset to Fig. 4A). A major band at ∼66 kDa was observed for the first protein fractions eluted from the SEC column (fraction 1). This mass value is compatible with that of the homodimeric BLT2. However, in this case, a minor band with an electrophoretic mobility compatible with that of the receptor monomer was still observed that corresponded to less than 5% of the total protein, as assessed by densitometric analysis of the SDS-PAGE gel (not shown). The occurrence of this band after cross-linking could arise either from an incomplete chemical cross-linking or from the presence of a minor fraction of monomeric receptor. In contrast, the last eluting proteins (fraction 3) strictly correspond to protein species with an electrophoretic mobility at ∼35 kDa compatible with that of the monomeric BLT2. This indicates that the receptor purified following ligand-immobilized chromatography essentially corresponds to a mixture of monomeric and dimeric species that can be separated using SEC. As expected, the protein fractions at intermediate elution volumes, i.e. fraction 2, correspond to a mixture of monomer and dimer, as assessed by chemical cross-linking.
We finally determined the topological features of the BLT2 dimer, i.e. whether the two protomers in the dimeric assembly are in parallel or antiparallel orientations. For this, we devised a FRET-based approach that consisted of labeling either the N terminus or C terminus of the receptor with a fluorescence donor or acceptor and then measuring the transfer efficiency between these two probes to assess proximity (see “Experimental Procedures”). N-terminal labeling was carried out as described for BLT1 (26). To specifically label the receptor C terminus, we introduced a transglutaminase recognition sequence that allows enzymatic modification of a reactive glutamine and incorporation of a fluorophore (27). The receptors labeled with the donor (Alexa Fluor-488) and acceptor (Alexa Fluor-568) probes were mixed in equivalent amounts before refolding and then refolded and the dimers were purified as described above. As expected, under such conditions, essentially equivalent amounts of donor- and acceptor-labeled protein were found in the dimeric fraction (fraction 1 as above), based on the UV absorption features of the proteins in this fraction (see “Experimental Procedures”). Because labeling does not affect the dimerization properties of BLT2 (similar SEC profiles were obtained for both the labeled and unlabeled proteins; not shown), one can expect the final dimeric fraction to be composed of a mixture of dimers where both protomers are labeled with the donor or acceptor molecule, and dimers where each of the protomers is labeled either with the donor or acceptor. Moreover, the distance between the extreme N and C termini of the BLT2 is expected to be significantly larger than the R0 value (∼60 Å) of the fluorophore pair used in the experiments, based on the different GPCR crystal structures obtained so far and biophysical data published for the fluorescently labeled β2-adrenergic receptor (38, 39). Consequently, FRET was expected to arise only from the latter species. As shown in Fig. 4B, a significant FRET signal was observed when protomers were labeled either both at the N terminus or both at the C terminus. This strongly suggests that the N-terminal regions are in proximity in the dimeric assembly, as well as both C termini. This is likely to be a specific effect of receptor dimerization because no signal was observed with the monomeric fractions (fraction 3 as above), ruling out possible collisional effects. In contrast, no signal was observed when one of the protomers was labeled at its N terminus and the other at the C terminus. All these data indicate that the two protomers in most, if not all dimers, are likely in a parallel orientation.
Pharmacology of the BLT2 Monomer and Dimer
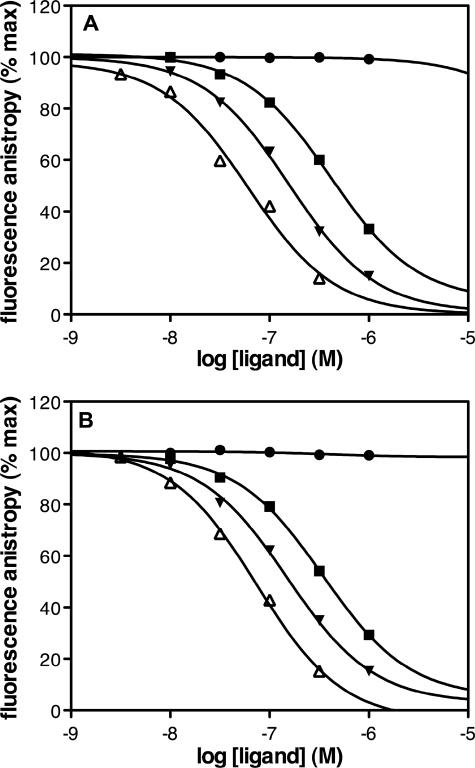
To assess that both monomeric and dimeric BLT2 populations were functional, we first measured the ligand-binding properties of the different fractions obtained after SEC. A similar Kd value for [3H]LTB4 was measured for both species, i.e. 255.8 ± 35.2 (n = 3) and 253.3 ± 38.7 nm (n = 3) for the monomeric and dimeric fractions, respectively. This is in agreement with the Kd calculated for the refolded affinity-purified BLT2 (see above). Moreover, the stoichiometric ratio of ∼1 obtained for the receptor dimer indicates that both protomers in the dimeric assembly are able to bind the LTB4 agonist. We then checked whether the receptor monomer and dimer displayed different pharmacological profiles. For this, we carried out a series of saturation and competition experiments using fluorescence anisotropy and the LTB4 derivative LTB4-568 as a fluorescent probe. Both this probe and the fluorescence anisotropy approach for monitoring ligand binding to the LTB4 receptors have been described previously (22). We first determined, by saturation fluorescence anisotropy binding assays, that affinity of the LTB4-568 for BLT2 monomers and dimers was in the same range than that defined with [3H]LTB4 for dimers and monomers, respectively. Indeed, LTB4-568 affinity was calculated to be Kd = 310 (n = 2) for monomers and 297 nm (n = 2) for dimers. We then assessed the pharmacological profile of the monomer and dimer in competition assays using LTB4-568 as the fluorescent tracer. As shown in Fig. 5, the two BLT2 populations displayed very similar pharmacological profiles, i.e. they both bound the LTB4 and 12-HHT agonists as well as the BLT2-specific LY255283 antagonist. In contrast, the BLT1-specific U75302 agonist did not significantly displaced LTB4-568 binding, considering either the monomer or dimer. Such a pharmacological profile is fully compatible with what has been reported for the BLT2 receptor expressed in neutrophils or transfected cell membrane fractions (15–18, 22, 40). Moreover, as shown in Table 2, similar IC50 values were obtained for all the ligands whether the monomer or the dimer are considered, indicating that dimerization of BLT2 does not have a significant impact on its ligand binding properties.
FIGURE 5.
Pharmacological profile of the BLT2 monomers and dimers. Fluorescence anisotropy-monitored competition experiments were carried out using the fluorescent LTB4-568 and the BLT2 monomer (A) or dimer (B) as described under “Experimental Procedures” (100 nm of monomers or dimers). Data are presented as fluorescence anisotropy (% of maximum, defined in the absence of displacing ligand) as a function of ligand concentration. Closed squares, LTB4; closed triangles, LY255283; closed circles, U75302; and open triangles, 12-HHT. The values are means from triplicates measured in an experiment representative of three independent assays, each done in triplicate.
TABLE 2.
Ligand binding properties of the BLT2 monomer and dimer fractions
The data indicated in the table are IC50 (nm) values inferred from the fluorescence anisotropy competition binding experiments reported in Fig. 5. These values are mean ± S.E. calculated from three distinct experiments carried out from two independent BLT2 monomer and dimer preparations.
| LTB4 | LY255283 | U75302 | 12-HHT | |
|---|---|---|---|---|
| BLT2 monomer | 404 ± 12 | 158 ± 13 | NMa | 61 ± 3 |
| BLT2 dimer | 344 ± 9 | 151 ± 11 | NM | 73.5 ± 5 |
a NM, not measurable.
Monomer- and Dimer-catalyzed G Protein Activation
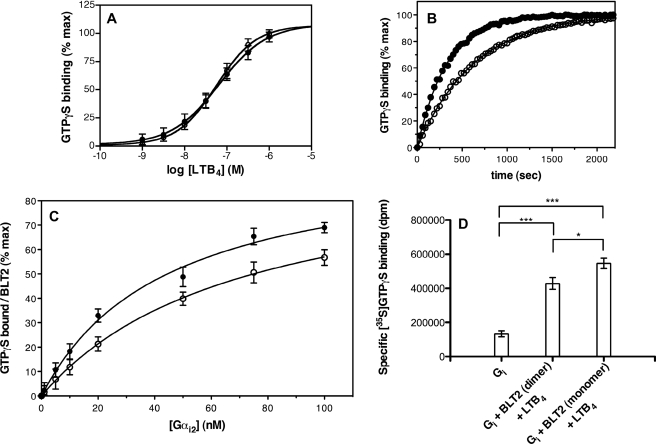
We subsequently analyzed whether the BLT2 monomer and dimer could efficiently activate the purified Gαi2β1γ2 protein. Using the fluorescence-based assay, we first assessed LTB4-catalyzed GTPγS incorporation to the Gi2 protein with the BLT2 monomers and dimers in the presence of increasing concentrations of the agonist. As shown in Fig. 6A, both fractions were able to trigger Gi2 activation. Half-saturation of LTB4-catalyzed GTPγS binding was reached at comparable agonist concentrations with monomers and dimers.
FIGURE 6.
Activation of the G protein Gαi2β1γ2 by monomeric and dimeric BLT2 receptors. A, LTB4 saturation of monomer (open circles) and dimer (closed circles) catalyzed GDP/GTP exchange. Data are presented as the percentage of maximal GTPγS binding as a function of LTB4 concentration. Results are given as mean ± S.E. calculated from three independent experiments. B, time-dependent activation of Gαi2 catalyzed by the LTB4-saturated form of the BLT2 monomer (closed circles) or dimer (open circles). Data are expressed as the percentage of maximal GTPγS binding as a function of time. The experiment shown is representative of three independent assays. In A and B, the BLT2 concentration was 20 nm, Gαi2 was 200 nm, and Gβ1γ2 were 500 nm. C, Gαi2 saturation of GDP/GTPγS exchange triggered by the BLT2 monomers (closed circles, fraction 3 in Fig. 4) and dimers (open circles, fraction 1 in Fig. 4). The contribution of the basal exchange (around 20% of the maximal receptor-catalyzed exchange) in the absence of agonist was systematically subtracted. The BLT2 concentration was 20 nm, those of Gβ1γ2 were in excess of 500 nm. Data are expressed as the percentage of maximal GTPγS binding normalized to BLT2 receptor concentration. The experiments shown in the figure were repeated three times. Results are given as mean ± S.E. D, [35S]GTPγS binding to the Gαi2 in the presence of LTB4-stimulated BLT2 monomers and dimers. BLT2 receptor preparations and Gαi2 were added at an equimolar ratio (20 nm each) in the presence of 500 nm G protein β1γ2 subunits. Data are expressed as specific dpm incorporated to the Gαi2. Mean ± S.E. are shown. Statistics are given: ***, p < 0.0001; *, p < 0.05. These experiments have been repeated at least three times, each done in triplicates. Error bars are S.E. (calculated as S.D./root square (n −1)).
We then analyzed the kinetic aspects of G protein activation triggered by the BLT2 monomer and dimer still using the fluorescence-based assay. Interestingly, when normalized to the total number of LTB4 binding sites, the reaction was approximately two times faster using the monomer compared with what was observed with the dimer (Fig. 6B, t½ = 3.6 and 7.5 min for the monomer and dimer, respectively). This suggests that the monomeric state of the receptor would represent the most active form of BLT2. To further investigate this observation on an experimental basis, we analyzed Gi2 saturation of BLT2-catalyzed GDP/GTPγS exchange with the two protein fractions, i.e. the monomeric and dimeric ones. As shown in Fig. 6C, the BLT2 monomer in the presence of saturating LTB4 concentrations was completely able to trigger GDP/GTPγS exchange at the level of the Gi2 subunit with a Km value of 44.6 ± 7.6 nm (n = 3). This clearly indicated that the receptor monomer is fully competent in terms of G protein activation. Interestingly, when the experiment was carried out with the protein fraction in which the receptor dimer was the major species, the calculated apparent Km value was significantly different by around 2-fold (82.0 ± 7.3 nm (n = 3)). This suggests that for an equivalent number of bound agonists, the BLT2 dimer is less efficient than the monomer in terms of G protein activation.
We finally confirmed the signaling reduced ability of the BLT2 dimer by directly measuring the receptor-dependent GTP binding activity of the Gi protein using radiolabeled [35S]GTPγS. To demonstrate that the receptors (monomers and dimers) are fully capable of G protein activation, each population of BLT2 and the Gi2 were added at a 1:1 molar ratio. As shown in Fig. 6D, interestingly, the amount of specifically bound [35S]GTPγS to the Gi2 following ligand stimulation of the BLT2 monomer was significantly higher than that obtained with the BLT2 dimer (545,766 ± 29,740 (n = 4) versus 428,547 ± 34,200 dpm (n = 4), p < 0.05). For comparison, incorporation of the radiolabeled nucleotide in the absence of ligand-receptor complexes (basal activity of the Gi2 protein) was much lower (131,991 ± 16,430 dpm (n = 6)). The one measured in the presence of the mixed population (BLT2 receptor before SEC separation) was intermediate (501,026 ± 61,580 dpm (n = 4)).
DISCUSSION
We analyzed and compared here the capacity of GPCR dimers and monomers to activate their cognate G protein using the purified BLT2 receptor as a model. We showed that the BLT2 monomer catalyzes GTPγS binding at the level of the Gi2 protein with higher affinity than the corresponding homodimer and with faster kinetics. We confirmed this result by directly measuring specific [35S]GTPγS incorporation to Gi2 in the presence of either the BLT2 monomer or dimer with a saturating concentration of the LTB4 agonist. These results strongly suggest that for an equivalent number of ligand-occupied binding sites, BLT2 monomers activate Gi2 protein more efficiently than dimers.
Our data indicate that the minimal BLT2 unit for Gi2 activation is the monomer and are in agreement with accumulating evidence showing that monomeric GPCRs are functional. Indeed, several studies demonstrated that monomeric rhodopsin is capable of full coupling to transducin Gt (11, 41, 42). The NTS1 neurotensin receptor monomer was also shown to activate the Gq protein better than its corresponding homodimer, although activation of Gq by monomeric and dimeric receptors has not been evaluated in the same conditions (13). Finally, the β2-adrenergic receptor and the opioid μ receptor were reconstituted into high density lipoprotein particles at one receptor per particle and were shown to efficiently activate their selective G protein Gs and Gi, respectively (12, 14). Our data with BLT2 confirm that like Gt, Gq, and Gs, Gi2 can be activated by a GPCR monomer, indicating that this is likely to be a common feature for all G protein subtypes. It is thus clear that although accumulating data reinforce the idea that GPCRs oligomerize in heterologous expression systems and in native cells (4, 9), the receptor monomer has per se all molecular determinants necessary for G protein activation.
Because the BLT2 monomer appears as the minimal unit for G protein activation (at least in detergent/lipid mixed micelles), this raises the question of what mechanism is responsible for the reduced capacity of receptor dimers to activate G proteins. Different interpretations have been proposed so far.
First, Bayburt et al. (41) have suggested that a lower efficiency of rhodopsin dimers to activate transducin could be the consequence of rhodopsin reconstitution into lipid nanodiscs that would result in a random orientation of the protomers in the dimer with two equal populations, parallel and anti-parallel, the latter being unable to activate transducin. In addition, Banerjee et al. (43) have indeed demonstrated the occurrence of anti-parallel GPCR dimers incorporated in nanoscale apolipoprotein-bound bilayers using electron microscopy of nanogold-labeled rhodopsin. However, this is not the case here for BLT2 because we directly demonstrated using an original FRET-based approach that the receptor dimer is essentially composed of a single population with both protomers in a parallel orientation. In this context, our observation means that, although established in a detergent environment far from that of a native membrane, protein:protein contacts in the BLT2 dimeric entity are of sufficient specificity and lead to correctly folded and assembled dimers.
Second, as proposed for the NTS1 receptor (13), steric hindrance between the two G protein binding sites in the dimer would be responsible for the reduced efficacy of the BLT2 dimer to activate Gi2. Such a model has also been proposed for rhodopsin for which steric constraints could prohibit interaction with more than one transducin at a time and explain why a rhodopsin dimer is less efficient than the monomer for transducin activation (41). Steric hindrance effects that would lead to G protein competition may also explain why activation by BLT2 dimers is less efficient. Although we do not so far have any direct evidence for the stoichiometric features of the BLT2·Gi2 complex, this would favor, as suggested for rhodopsin, NTS1, or BLT1 receptors (25), a complex where the BLT2 dimer would efficiently interact essentially with a single Gi2 protein.
A third possibility would be to consider a model recently proposed whereby dimerization could serve as a “desensitization” mechanism (44), rapidly suppressing G protein-mediated signaling when there are two many active receptors. In agreement with this hypothesis, dimerization could constitute a way to modulate G protein-mediated signaling. A reduced coupling efficiency of dimeric receptors to their cognate G protein, as observed in this work as well as for the neurotensin NTS1 or rhodopsin, would be consistent with this idea. Although, as proposed above, steric constraints could affect accessibility of one of the two protomers to the G protein and be responsible for this desensitization process, another possibility would be to consider a negative allostery mechanism between protomers through direct trans-conformational changes within the receptor dimer. In this case, agonist-induced conformational changes in one protomer, compatible with a complete activation of its cognate G protein, would trigger an inhibitory trans-conformational change of the second protomer. Such inhibitory cross-conformational changes have been recently proposed to occur in a μ-opioid:α2A adrenergic receptor dimer (45). This study is consistent with asymmetric roles for GPCR subunits in receptor dimers. Examples of asymmetry for different class A GPCR families in terms of G protein activation have been reported (13, 24, 41, 46). In addition, the asymmetric nature of GPCRs has also been elegantly analyzed with class C GPCRs. Very importantly, studies with metabotropic glutamate receptors mGlu1 and mGlu5 also support the conclusion that a G protein needs just one active protomer, and that two protomers in the active conformation impede signaling (47, 48).
It has to be strongly emphasized here that all the data presented on differential activation of the Gi2 protein by BLT2 monomers and dimers have been obtained because of the possibility to produce the receptor in high amounts in a functional state. This highlights the importance of the strategy we have developed that combines an original and efficient way to overexpress the receptor in E. coli IBs to the purification and refolding steps previously described (27, 29, 30). Indeed, improving the expression method allowed to obtain sufficient amounts of functional protein even with moderate refolding yields. Fusing the receptor to α5I led to high expression levels for all GPCRs tested, without any optimization of either the cell culture conditions or the extraction/purification procedures. This constitutes a significant improvement over what has been described to date with respect to expression levels in bacteria and therefore represents an important breakthrough for in vitro studies aimed at understanding the molecular bases of the function of class A GPCRs and, possibly, of other membrane proteins.
Supplementary Material
Acknowledgments
The pMR1-PLC and the pET24a-PurF vectors were provided by Drs. M. Roberts (Boston College, Chestnut Hill, MA) and H. Zalkin (Purdue University, West Lafayette, IN), respectively. Thanks to Drs. J.-D. Franssen (Euroscreen SA, Gosselies, Belgium) for providing the human ChemR23 cDNA. We are grateful to Dr. J.-P. Pin for constant support. The strategy described here has been patented by INSERM/CNRS (patents 03/07411 and WO 2004/113539 A3).
This work was supported in part by INSERM, CNRS, European community Grant LSHB-CT-2003-503337, ANRS Foundation Grant AC14, French Ministry of Research Grant ACI BCMS 328, and National Agency of Research Grant ANR 06-BLAN-0087.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1 and S2 and “Experimental Procedures.”
- GPCR
- G protein-coupled receptor
- BLT2
- leukotriene B4 receptor subtype 2
- FRET
- fluorescence resonance energy transfer
- G protein
- guanosine trisphosphate-binding protein
- Gα
- G protein α subunit
- α5I
- α5 integrin
- IBs
- inclusion bodies
- IMAC
- immobilized metal affinity chromatography
- LTB4
- leukotriene B4
- PLC
- phospholipase C
- PurF
- glutamine phosphoribosyl pyrophosphatase amidotransferase
- SEC
- size exclusion chromatography
- V2
- arginine-vasopressin receptor subtype 2
- GTPγS
- guanosine 5′-3-O-(thio)triphosphate
- LTB4-568
- LTB4 labeled with Alexa Fluor-568
- Ni-NTA
- nickel-nitrilotriacetic acid
- DPC
- n-dodecyl phosphocholine
- HDM
- hexadecyl-β-d-maltoside
- CHS
- cholesteryl hemisuccinate
- MOPS
- 4-morpholinepropanesulfonic acid
- GST
- glutathione S-transferase
- OTR
- oxytocin receptor.
REFERENCES
- 1.Bockaert J., Pin J. P. (1999) EMBO J. 18, 1723–1729 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fredriksson R., Lagerström M. C., Lundin L. G., Schiöth H. B. (2003) Mol. Pharmacol. 63, 1256–1272 [DOI] [PubMed] [Google Scholar]
- 3.Lefkowitz R. J. (2004) Trends Pharmacol. Sci. 25, 413–422 [DOI] [PubMed] [Google Scholar]
- 4.Gurevich V. V., Gurevich E. V. (2008) Trends Neurosci. 31, 74–81 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Pin J. P., Kniazeff J., Liu J., Binet V., Goudet C., Rondard P., Prézeau L. (2005) FEBS J. 272, 2947–2955 [DOI] [PubMed] [Google Scholar]
- 6.Terrillon S., Bouvier M. (2004) EMBO Rep. 5, 30–34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chabre M., le Maire M. (2005) Biochemistry 44, 9395–9403 [DOI] [PubMed] [Google Scholar]
- 8.Park P. S., Filipek S., Wells J. W., Palczewski K. (2004) Biochemistry 43, 15643–15656 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fotiadis D., Liang Y., Filipek S., Saperstein D. A., Engel A., Palczewski K. (2003) Nature 421, 127–128 [DOI] [PubMed] [Google Scholar]
- 10.Baylor D. A., Lamb T. D., Yau K. W. (1979) J. Physiol. 288, 613–634 [PMC free article] [PubMed] [Google Scholar]
- 11.Whorton M. R., Jastrzebska B., Park P. S., Fotiadis D., Engel A., Palczewski K., Sunahara R. K. (2008) J. Biol. Chem. 283, 4387–4394 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Whorton M. R., Bokoch M. P., Rasmussen S. G., Huang B., Zare R. N., Kobilka B., Sunahara R. K. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 7682–7687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.White J. F., Grodnitzky J., Louis J. M., Trinh L. B., Shiloach J., Gutierrez J., Northup J. K., Grisshammer R. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 12199–12204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kuszak A. J., Pitchiaya S., Anand J. P., Mosberg H. I., Walter N. G., Sunahara R. K. (2009) J. Biol. Chem. 284, 26732–26741 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yokomizo T., Kato K., Terawaki K., Izumi T., Shimizu T. (2000) J. Exp. Med. 192, 421–432 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kamohara M., Takasaki J., Matsumoto M., Saito T., Ohishi T., Ishii H., Furuichi K. (2000) J. Biol. Chem. 275, 27000–27004 [DOI] [PubMed] [Google Scholar]
- 17.Yokomizo T., Izumi T., Shimizu T. (2001) Arch. Biochem. Biophys. 385, 231–241 [DOI] [PubMed] [Google Scholar]
- 18.Tager A. M., Luster A. D. (2003) Prostaglandins Leukot. Essent. Fatty Acids 69, 123–134 [DOI] [PubMed] [Google Scholar]
- 19.Gill S. C., von Hippel P. H. (1989) Anal. Biochem. 182, 319–326 [DOI] [PubMed] [Google Scholar]
- 20.Nominé Y., Ristriani T., Laurent C., Lefèvre J. F., Weiss E., Travé G. (2001) Protein Eng. 14, 297–305 [DOI] [PubMed] [Google Scholar]
- 21.Poudrel J. M., Hullot P., Vidal J. P., Girard J. P., Rossi J. C., Muller A., Bonne C., Bezuglov V., Serkov I., Renard P., Pfeiffer B. (1999) J. Med. Chem. 42, 5289–5310 [DOI] [PubMed] [Google Scholar]
- 22.Sabirsh A., Wetterholm A., Bristulf J., Leffler H., Haeggström J. Z., Owman C. (2005) J. Lipid Res. 46, 1339–1346 [DOI] [PubMed] [Google Scholar]
- 23.Oldham W. M., Van Eps N., Preininger A. M., Hubbell W. L., Hamm H. E. (2006) Nat. Struct. Mol. Biol. 13, 772–777 [DOI] [PubMed] [Google Scholar]
- 24.Damian M., Martin A., Mesnier D., Pin J. P., Banères J. L. (2006) EMBO J. 25, 5693–5702 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Banères J. L., Parello J. (2003) J. Mol. Biol. 329, 815–829 [DOI] [PubMed] [Google Scholar]
- 26.Mesnier D., Banères J. L. (2004) J. Biol. Chem. 279, 49664–49670 [DOI] [PubMed] [Google Scholar]
- 27.Jäger M., Nir E., Weiss S. (2006) Protein Sci. 15, 640–646 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hickerson R., Majumdar Z. K., Baucom A., Clegg R. M., Noller H. F. (2005) J. Mol. Biol. 354, 459–472 [DOI] [PubMed] [Google Scholar]
- 29.Kiefer H., Krieger J., Olszewski J. D., Von Heijne G., Prestwich G. D., Breer H. (1996) Biochemistry 35, 16077–16084 [DOI] [PubMed] [Google Scholar]
- 30.Kiefer H. (2003) Biochim. Biophys. Acta 1610, 57–62 [DOI] [PubMed] [Google Scholar]
- 31.Baneres J. L., Martin A., Hullot P., Girard J. P., Rossi J. C., Parello J. (2003) J. Mol. Biol. 329, 801–814 [DOI] [PubMed] [Google Scholar]
- 32.Banères J. L., Mesnier D., Martin A., Joubert L., Dumuis A., Bockaert J. (2005) J. Biol. Chem. 280, 20253–20260 [DOI] [PubMed] [Google Scholar]
- 33.Mouillac B., Sen T., Durroux T., Gaibelet G., Barberis C. (2002) Prog. Brain Res. 139, 163–177 [DOI] [PubMed] [Google Scholar]
- 34.Wilkinson D. L., Harrison R. G. (1991) Biotechnology 9, 443–448 [DOI] [PubMed] [Google Scholar]
- 35.Tan C. A., Hehir M. J., Roberts M. F. (1997) Protein Expr. Purif. 10, 365–372 [DOI] [PubMed] [Google Scholar]
- 36.Tso J. Y., Zalkin H., van Cleemput M., Yanofsky C., Smith J. M. (1982) J. Biol. Chem. 257, 3525–3531 [PubMed] [Google Scholar]
- 37.Banères J. L., Roquet F., Green M., LeCalvez H., Parello J. (1998) J. Biol. Chem. 273, 24744–24753 [DOI] [PubMed] [Google Scholar]
- 38.Rosenbaum D. M., Rasmussen S. G., Kobilka B. K. (2009) Nature 459, 356–363 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Granier S., Kim S., Shafer A. M., Ratnala V. R., Fung J. J., Zare R. N., Kobilka B. K. (2007) J. Biol. Chem. 282, 13895–13905 [DOI] [PubMed] [Google Scholar]
- 40.Okuno T., Iizuka Y., Okazaki H., Yokomizo T., Taguchi R., Shimizu T. (2008) J. Exp. Med. 205, 759–766 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bayburt T. H., Leitz A. J., Xie G., Oprian D. D., Sligar S. G. (2007) J. Biol. Chem. 282, 14875–14881 [DOI] [PubMed] [Google Scholar]
- 42.Ernst O. P., Gramse V., Kolbe M., Hofmann K. P., Heck M. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 10859–10864 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Banerjee S., Huber T., Sakmar T. P. (2008) J. Mol. Biol. 377, 1067–1081 [DOI] [PubMed] [Google Scholar]
- 44.Gurevich V. V., Gurevich E. V. (2008) Trends Pharmacol. Sci. 29, 234–240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Vilardaga J. P., Nikolaev V. O., Lorenz K., Ferrandon S., Zhuang Z., Lohse M. J. (2008) Nat. Chem. Biol. 4, 126–131 [DOI] [PubMed] [Google Scholar]
- 46.Han Y., Moreira I. S., Urizar E., Weinstein H., Javitch J. A. (2009) Nat. Chem. Biol. 5, 688–695 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hlavackova V., Goudet C., Kniazeff J., Zikova A., Maurel D., Vol C., Trojanova J., Prézeau L., Pin J. P., Blahos J. (2005) EMBO J. 24, 499–509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Goudet C., Kniazeff J., Hlavackova V., Malhaire F., Maurel D., Acher F., Blahos J., Prézeau L., Pin J. P. (2005) J. Biol. Chem. 280, 24380–24385 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Wyman J., Gill S. J. (1990) in Binding and Linkage: Functional Chemistry of Biological Macromolecules (Kelly A. ed) pp. 33–61, University Science Books, Mill Valley, CA [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.