Abstract
Here, Deans and West (2009) reveal the molecular basis of the phenotypic similarities between Fanconi Anemia (FA) and Bloom's Syndrome, identifying FANCM as the anchor for both FA and Bloom's complexes at the site of the DNA interstrand crosslink.
Fanconi anemia (FA) is a rare genomic instability disorder characterized by chromosomal instability, bone marrow failure, and increased susceptibility to cancer (reviewed in Moldovan and D'Andrea, 2009). Thirteen FA genes have been cloned so far, and the encoded proteins cooperate in a DNA repair pathway designed to remove interstrand crosslinks (ICLs) during the S phase of the cell cycle. Most of the Fanconi proteins, including FANCM, form the FA core complex that functions as an E3 ligase for FANCI and FANCD2 upon DNA damage. Ubiquitinated FANCI and FANCD2 then colocalize in chromatin foci with other repair proteins. Partial colocalization with BLM, the RecQ helicase mutated in Bloom's syndrome, has also been reported. Among the FA core complex proteins, only FANCM has been reported to have DNA binding activity, with a preference for a DNA structure resembling a stalled replication fork. FANCM recruits the core complex to chromatin, and this recruitment is cell cycle regulated. Moreover, FANCM contains a DEAH helicase domain that exhibits DNA translocation and fork reversal activities, underscoring its active role in ICL repair.
FA patients exhibit partial overlapping phenotypes with Bloom's syndrome (BS) patients. BS is a rare autosomal recessive disorder associated with reduced fertility, immunodeficiency, and elevated predisposition to cancer. While FA patients tend to develop myeloid leukemias and squamous cell carcinomas, BS patients tend to have a wider range of neoplasms, with a higher incidence of lymphomas. The gene mutated in BS, BLM, encodes a DNA helicase that is found in a complex with topoisomerase IIIα, RMI1, and RMI2 (Liu and West, 2008). BLM has been implicated in the processing of diverse DNA structures, including stalled replication forks which occur when the replication machinery encounters ICLs. Interestingly, the Bloom's complex (or BS complex consisting of BLM, RMI1/2, and TopoIIIα) associates with the FA core proteins to form a supercomplex, called BRAFT, in cells treated with crosslinking agents (Meetei et al., 2003).
In the context of these clinical and biochemical similarities, the new study of Deans and West makes a key advance in unraveling the interplay between these two disorders by identifying FANCM as an anchor protein for both the FA core complex and BS complex (Deans and West, 2009). Two short motifs in FANCM, outside of the well-defined DEAH and nuclease domains, exhibit evolutionary conservation among vertebrates. These motifs, named MM1 and MM2 (FANCM motifs 1 and 2), were respectively shown to recruit the FA core complex and the BS complex. Further investigations identified FANCF as the FA protein anchoring the core complex to MM1, and TopoIIIα as well as the C-terminal region of RMI1, as the link between the BS complex and MM2. Of note, the FA core complex and BS complex do not interact biochemically in the absence of FANCM, establishing FANCM as the key bridge protein connecting them (Figure 1A).
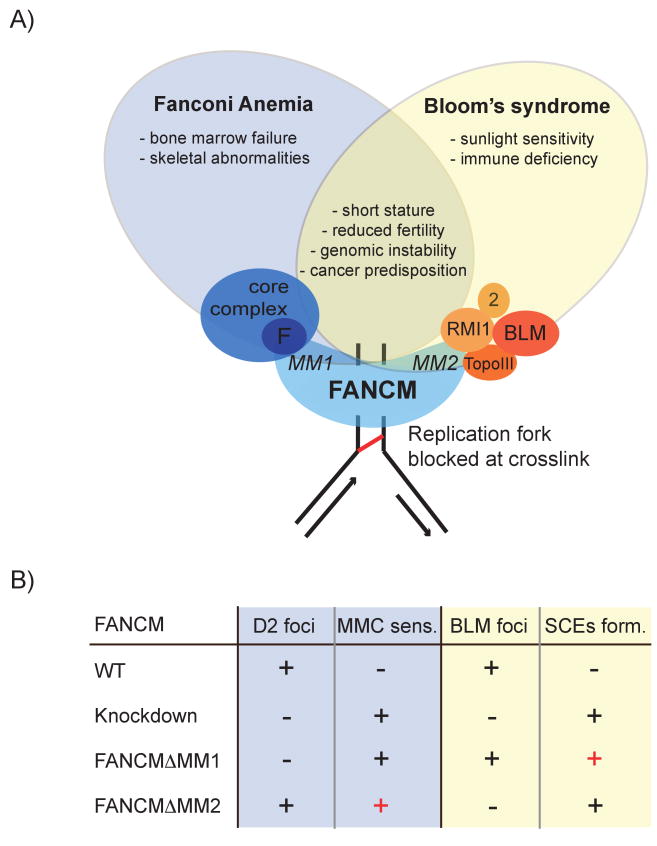
Fig 1. FANCM recruits both the FA core complex and BS complex at DNA crosslink sites and may explain the shared phenotypes of FA and BS patients.
A) This model shows FANCM binding to the FA core complex via FANCF (via the MM1 motif) and to the BS complex via RMI1 and TopoIIIα via the MM2 motif). These interactions are believed to occur at stalled replication forks, due to DNA interstrand crosslinks. In the background of the figure some of the most common phenotypes associated with Fanconi anemia and/or Bloom's syndrome are indicated.
B) In this panel, the common cellular phenotypes for Fanconi pathway deficient cells (loss of FANCD2 foci and MMC sensitivity) and BLM deficient cells (loss of BLM foci and SCEs formation) are summarized. These phenotypes were investigated in detail by Deans and West (2009) for FANCM wild type (WT) or mutants and the results are reported here. “+” indicates formation of foci and MMC sensitivity or SCEs formation. “-” indicates absence or reduction of foci and MMC resistance or absence of SCEs. The red “+” highlights interesting and unexpected new findings made in this work that further describe the interplay between FA and BS.
To complement the biochemical analysis of their work, Deans and West (2009) investigated the functional relevance of these interactions. Cells in which the FANCM-BLM interaction is abolished, through expression of a FANCM polypeptide deficient in the MM2 sequence (i.e., FANCM-MM2-deficient cells), are hypersensitive to crosslinking agents but still proficient for FANCD2 ubiquitination and foci assembly (Figure 1B). This result demonstrates that the hypersensitivity to the crosslinking agent mitomycin C (MMC) observed in BS cells (Pichierri et al., 2004) is not due to an indirect requirement of BLM in the FA pathway. Interestingly, these FANCM-MM2-deficient cells do not form BLM foci after treatment with DNA crosslinking agents, but do recruit BLM in foci after IR-irradiation. FANCM is therefore a specific anchor for the Bloom's complex in the context of an ICL-arrested replication fork but not in the setting of other kinds of DNA damage, such as DNA double strand breaks.
Cells expressing a FANCM polypeptide deficient in the MM1 sequence (i.e., FANCM-MM1-deficient cells), in which FANCM is unable to interact with the FA core complex, are crosslinker hypersensitive, as expected, but also show elevated sister chromatid exchanges (SCEs), despite the normal assembly of BLM foci. Elevated SCEs are a characteristic feature of BLM-deficient cells. This finding suggests that recruitment of both the FA complex and the Bloom's complex to sites of crosslinker damage is required to suppress sister chromatid exchange. Although SCEs are not a classical feature of human FA cells, this conclusion is consistent with studies in DT40 chicken cells where deletion of FA core complex genes does lead to elevated SCEs (Rosado et al., 2009).
The interaction of the FA and Bloom complexes via FANCM explains, at least in part, the shared phenotypes of FA and BS (Deans and West, 2009 and Figure 1A). However, other phenotypes are disease specific, suggesting that the FA and the Bloom's pathways are involved in independent functions, besides their common role at CL-arrested replication forks.
How can we reconcile the presence of both FA proteins and the BLM protein in a common complex at the stalled replication fork? On the one hand, FANCM may target the FA core complex to the site of ICL damage in order to promote FANCD2/I ubiquitination. This upstream ubiquitination step is critical for the successful lesion bypass and nuclease incision steps of crosslink repair (Knipscheer et al., 2009). On the other hand, FANCM may target the BS complex to this ICL site in order to promote the downstream step of repair after ICL excision- namely, the repair of the generated double strand break by homologous recombination. Thus, FANCM may provide the landing pad that connects these two steps.
Absence of FANCM or BLM leads to elevated levels of SCEs. However, genetic studies using DT40 cells have shown that FANCM functions in only a subset of BLM dependent non-crossover events (Rosado et al., 2009). Perhaps FANCM only regulates BLM's ability to dissolve D loops while the more central role of BLM in these events results from its direct role in decatenating Holliday junctions. How the FA core complex inhibits crossover events remains unclear. Perhaps the FA core E3 ligase monoubiquitinates another substrate protein involved in this inhibitory mechanism.
In addition to cooperating in ICL repair, the FA and Bloom's pathways may also cooperate in other DNA repair activities. Recent studies indicate that cells entering mitosis with an incompletely-replicated genome accumulate ultra-fine DNA bridges between separating sister chromatids. These bridges arise from genomic regions that are more difficult to replicate, such as common DNA fragile sites. Importantly, they are coated by BLM and tethered by foci containing ubiquitinated FANCD2/I (Chan et al., 2009; Naim and Rosselli, 2009). Therefore, these structures suggest another cooperative role for the FA and BLM pathways. Since FANCM interacts with both the FA core complex and BLM, it will be interesting to determine whether FANCM is also involved in the recruitment or regulation of BLM and FA proteins at the sites of these ultrafine bridges.
In summary, FANCM plays a central role in DNA crosslink repair, anchoring two independent complexes, FA and BS, to the site of the damage. Despite this advance, many questions regarding the actual repair of the DNA crosslink are unresolved. For example, since BLM and FANCD2 foci only partially colocalize (Pichierri et al., 2004), other damaged DNA substrates and protein factors may specifically recruit only one pathway or the other. Also, since the FANCM protein is degraded in mitosis (Kee et al, 2009), the functional interaction between the FA pathway and BLM protein in repair is likely to be confined to a specific cell cycle stage. Finally, how the FANCM protein mediates the handoff from the upstream FA pathway to the downstream events in homologous recombination repair, and whether this handoff requires the translocase activity of FANCM, remain critical unanswered questions for the field.
Acknowledgments
PV is a fellow of the Swiss Foundation for grants in biology and medicine. The work was supported by NIH grant PO1CA092584.
References
- Chan KL, Palmai-Pallag T, Ying S, Hickson ID. Nat Cell Biol. 2009;11:753–760. doi: 10.1038/ncb1882. [DOI] [PubMed] [Google Scholar]
- Deans AJ, West SC. Mol Cell this issue. 2009 doi: 10.1016/j.molcel.2009.12.006. [DOI] [PubMed] [Google Scholar]
- Kee Y, Kim JA, D'Andrea AD. Genes and Devel. 2009;23:555–560. doi: 10.1101/gad.1761309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knipscheer P, Räschle M, Smogorzewska A, Enoiu M, Ho TV, Schärer OD, Elledge SJ, Walter JC. Science. 2009 doi: 10.1126/science.1182372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, West SC. Genes Dev. 2008;22:2737–2742. doi: 10.1101/gad.1732808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meetei AR, Sechi S, Wallisch M, Yang D, Young MK, Joenje H, Hoatlin ME, Wang W. Mol Cell Biol. 2003;23:3417–3426. doi: 10.1128/MCB.23.10.3417-3426.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moldovan GL, D'Andrea AD. Annu Rev Genet. 2009 doi: 10.1146/annurev-genet-102108-134222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naim V, Rosselli F. Nat Cell Biol. 2009;11:761–768. doi: 10.1038/ncb1883. [DOI] [PubMed] [Google Scholar]
- Pichierri P, Franchitto A, Rosselli F. EMBO J. 2004;23:3154–3163. doi: 10.1038/sj.emboj.7600277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosado IV, Niedzwiedz W, Alpi AF, Patel KJ. Nucleic Acids Res. 2009;37:4360–4370. doi: 10.1093/nar/gkp365. [DOI] [PMC free article] [PubMed] [Google Scholar]