Abstract
Nearly half of our genomes are repetitive sequences derived from retrotransposons. These repeats have accumulated by a ‘copy-and-paste’ mechanism whereby: (i.) a genomic template sequence is transcribed to RNA, (ii.) the RNA is reverse-transcribed, and (iii.) the DNA copy is inserted at a new location in the host genome. As we remain susceptible to new retrotransposition events, many of these insertions are highly polymorphic. Transposons are of interest since insertions into both coding and non-coding gene regions have been associated with a wide variety of functional sequelae and because transposable elements can be involved in genomic rearrangements in transformed cells. In this review, we highlight how expression of retrotransposons, de novo and polymorphic transposon insertions, and genomic rearrangements that these repeats potentiate contribute to both benign and neoplastic hematopoietic diseases.
Introduction
A substantial fraction of most eukaryotic genomes is comprised of repetitive sequences, many of which are transposable elements (TEs, also called mobile DNAs). The recent and rapidly increasing availability of sequence data from various genome projects is allowing ever more comprehensive analyses of TE sequences and their distributions in genomes. Although it has become clear that TEs are underrepresented in current genome assemblies, substantial evidence has accumulated that several classes of mobile DNA elements remain active in humans and can generate genetic diversity, with a strong potential to influence gene function and cause clinically significant phenotypes.
Hematologic diseases span a large spectrum of pathologies, ranging from disorders that result in failed production of blood clotting pathway components and red and white blood cell lineages, to neoplastic proliferations such as leukemias and lymphomas. The former include the hemophilias and hemoglobinopathies, fairly common and most frequently heritable diseases for which the most critical gene loci involved have been well-studied. TE insertional mutagenesis has been known for some time to be a cause of these genetic lesions – albeit infrequently. Indeed, two cases of hemophilia provided the first evidence that de novo retrotransposition events could lead to a constitutional genetic defect causing human disease [1]. Roles of many TE integrants in contributing to human diseases in general have been described previously in excellent reviews [2, 3]. In this paper, we discuss roles of TEs as mutagens in benign hematologic diseases, as well as explore ways by which polymorphic intronic insertions may contribute to more subtle genetic predispositions to disease. Finally, we will turn our focus to neoplastic hematopathologies and discuss how TE dysregulation and TE-mediated genomic rearrangements may promote some of these tumors. We likely do not know all manners in which TEs contribute to hematologic disease, or their frequency in such roles. However, given the widespread distribution of these sequences throughout our genomes, their continued ability to move, and their abilities to disrupt transcripts and chromosomes, we anticipate that many more instances of TEs contributing to blood diseases will be identified in the future.
Types of Transposable Elements (TEs)
There are two broad categories of transposable elements. Class I transposons are made up of retrotransposons which copy themselves from one genomic location to another via an RNA intermediate. The RNA is transcribed by host machinery from a genomic DNA template and then reverse-transcribed and integrated elsewhere into the genome either by enzymatic functions encoded by the TE itself (as is the case for autonomous Class I retrotransposons) or by another class of TE (as occurs in parasitic Class I retrotransposition). Class II transposons (also known as DNA transposons or colloquially as ‘jumping genes’) are more ancient and move by a ‘cut-and-paste’ mechanism not involving an RNA intermediate. Although beyond the scope of this review, V(D)J recombination mechanisms for generating antigen binding epitope diversity seem to have been co-opted from Class II type transposons [4-6]. Here, we focus on the more prevalent and polymorphic endogenous Class I transposons.
Retrotransposons (Class I TEs) are further sub-classified by sequence features, which in turn reflect their mechanism of transposition and interaction with the host genome integration site. This classification scheme distinguishes three broad groups of Class I TEs: LTR retrotransposons; non-LTR, autonomous retrotransposons; and the non-autonomous or parasitic retrotransposons (Figure 1).
Figure 1. Structural features of HERVs, LINEs, SINEs and SVAs.
(Modified from [101].) Endogenous retroviruses (ERVs) are examples of autonomous, LTR retroelements. These consist of partly overlapping coding regions bounded by long terminal repeats (LTRs). ERVs code for group-specific antigen (Gag), protease (PR), and polymerase (Pol). The LTR has promoter activity (arrow) on the 5′ as well as the 3′ end. There are also target site duplications (TSDs; thick black lines). The Pol gene contains domains for integrase (IN), reverse transcriptase (RT), and RnaseH (RH). While ERV templates are not excised in transposition, recombination between LTRs can result in proviral loss, leaving a ‘solo LTR’ at the integration site. LINE retrotransposons are a prototypical example of autonomous, non-LTR transposable elements. Human genomic L1 LINEs consist of an internal CpG-rich 5′UTR with a polymerase II promoter (arrow), two open reading frames (ORF1 and ORF2), and a poly(A) tail [An]. ORF1 encodes a 40 kDa protein involved in formation of the ribonuclear protein (RNP) complex; ORF2 encodes for reverse transcriptase (RT) and endonuclease (En) activities. Genomic L1 insertions are characteristically flanked by short target site duplications (TSD) reflecting fill in repair of staggered dsDNA breakage at the insertion site (thick black lines). Alu SINEs are examples of non-autonomous, non-LTR elements. Human Alus consist of two GC rich monomer sequences, termed the left (L Alu) and the right (R Alu), and an intervening A-rich linker. SVAs are another type of non-autonomous, non-LTR retrotransposon, and are composite structures named after constituent SINE-R (derived from HERV LTR sequence), VNTR and Alu repeats. Like LINEs and reflecting their dependence on LINE-encoded ORF2, Alus and SVAs have a poly (A) tail and are flanked by TSDs (thick black lines). Alus are transcribed by a RNA polymerase III promoter. Drawings are not to scale. Full-length HERVs approximate 10kb; L1 LINEs 6kb; Alus 300bp; SVAs 2kb.
LTR retrotransposons are named for their long terminal repeats, 300bp to 1kb sequences at their 5′ and 3′ ends, which are identical in sequence and orientation on each side of the internal proviral sequence. LTR retrotransposons are similar in structure to exogenous retroviruses and differ principally in their lack of ability to move from one cell to the other. These TEs generate dsDNA copies using self-encoded reverse transcriptase (RT) protein and template RNA as well as extrachromosomal nucleic acids, such as tRNAs, to prime the RT reaction. The complementary DNA (cDNA) is then integrated in the host genome via a recombination event mediated by an encoded integrase protein. In contrast, non-LTR retroelements, lacking long terminal repeats, couple reverse transcription of their RNA intermediate with integration into the host chromosome by a mechanism referred to as target-primed reverse transcription (TPRT; [7] Figure 2).
Figure 2. Comparison of LTR (A) and non-LTR (B) retrotransposon mechanism.
(Modified from [102].) A Retrotransposition of LTR retrotransposons starts with transcription of the TE from its genomic location (gray). This is followed by reverse transcription (RT) of the RNA template into cDNA, after which integrase (purple dots) acts to insert the cDNA into a new genomic location, generating a target site (TS) duplication at its flanks. B Reverse transcription and insertion of non-LTR retrotransposons is carried out in a coupled process involving intrachromosomal priming; this is referred to as target primed reverse transcription (TPRT). It involves ORF2p-mediated endonuclease activity at the DNA target site (TS) to expose a 3′ hydroxyl group, followed by ORF2p RT activity extension of the DNA strand to copy the retrotransposon sequence. Resolution of the resulting RNA-DNA hybrid is thought to be reliant on host DNA repair mechanisms.
LTR retrotransposons, principally families of human endogenous retroviruses (HERVs), make up about 10% of the human genome. Over time most of these elements have acquired nonsense mutations and other sequence alterations leading to a loss-of-function of genes needed for mobilization. Of the LTR retrotransposons, only the HERV-K subgroup is known to have been recently active, and there are discrepant views on whether this remains the case. Examples of polymorphic HERV-Ks have been described but are uncommon; to date, no diseases have been linked to de novo insertions of HERV-K elements [8, 9]. In the mouse genome, but not in humans, there are two highly active families of ERV autonomous LTR elements that merit mention, the intracisternal A-particle (IAP) and ETn/MusD elements [10]. IAPs are endogenous retroviruses lacking envelope sequences which are capable of very high rates of intracellular retrotransposition (about 10−4 to 10−3 [11]).
As is the case for some LTR retrotransposons, autonomous non-LTR retrotransposons encode the critical machinery needed for their reverse transcription and genomic integration. Long interspersed elements (LINEs) are a chief example of this type of TE in the human genome, as their copy number of several hundred thousand accounts for nearly 20% of the genome. While most are now non-functional sequences that have become fixed homozygous integrants across different human populations, a subset of around 100 full-length (6kb) and functional LINEs exists in the human genome. Most of these are part of the recently active L1(Ta) LINE subset, a group which has been sufficiently successful to have caused numerous polymorphic insertions. An intact L1 LINE codes for two proteins, ORF1p and ORF2p, the latter being responsible for endonuclease and reverse transcriptase enzymatic activities critical for TPRT [12, 13]. Several examples of disease-causing LINE and LINE-dependent TE insertions have been described (reviewed in [14]). We will highlight some integrants pertinent to hematology in sections to follow (see Retrotransposon-induced coagulopathies and Transposable elements and red blood cell production below.)
Non-autonomous TEs in the human genome include the Alu elements, which belong to the group of short interspersed elements (SINEs). There are close to one million, approximately 300bp Alu elements within the human genome. Of these, the AluYa5/8 and AluYb8/9 subfamilies have expanded most effectively in Homo sapiens and display significant polymorphism across demographics [15]. As we will describe in subsequent sections, Alus are important in the genetics of hematopoietic disease as both mobile DNAs and as substrates for illegitimate recombination events. Another group of non-autonomous retrotransposons in the human genome are the relatively recently described SVA elements, which are composite TEs named for their SINE-R, VNTR, and Alu-derived modules [16]. Both Alus and SVAs are thought to hijack LINE-encoded ORF2p to accomplish their retrotransposition [17].
The preceding descriptions are intended to provide a basic overview of the major classes of retroelements in mammalian genomes. For more comprehensive discussion of these TEs and their mechanisms of retrotransposition, please see [2, 18, 19] (for HERVs and IAPs); [20] (for L1 LINEs); [21] (for Alus); and [22] (for SVAs)].
Retrotransposon-induced coagulopathies
Patients with bleeding disorders known as the hemophilias have compromised blood clotting factor activities most commonly due to insufficient thrombin generation by the factor VIIIa/IXa ‘tenase’ complex of the intrinsic coagulation cascade; there are about 18,000 affected individuals in the United States. Hemophilia A is characterized by deficiency of factor VIII, commonly resulting from X-linked, inherited loss-of-function mutations at the F8 gene locus on chromosome Xq28. In 1988, Kazazian et al. reported that 2 of 240 unrelated patients with hemophilia A had 2.3 kb and 3.8 kb long insertions of L1 LINEs, respectively, positioned within a short distance of one another within exon 14 of the factor VIII gene (Figure 3A, B; [1]). These insertional mutations were ascribed to de novo (non-parental) retrotransposon integration events. This landmark report documented the first examples of human disease caused by endogenous TE retrotransposition. It has been followed more recently by a report of an L1 LINE insertion associated with a large deletion [23] and two reports of AluYb insertions at the factor VIII locus causing a similar hemophilia A phenotype. In the latter, Sukarova et al. reported a full-length AluYb8 insertion into the same exon 14 of factor VIII ([24]; Figure 3C), and Ganguly et al. [25] reported an intronic AluYb9 insertion upstream of a consequently skipped exon 19. All three of these alternations were passed by heterozygous maternal carriers to affected children.
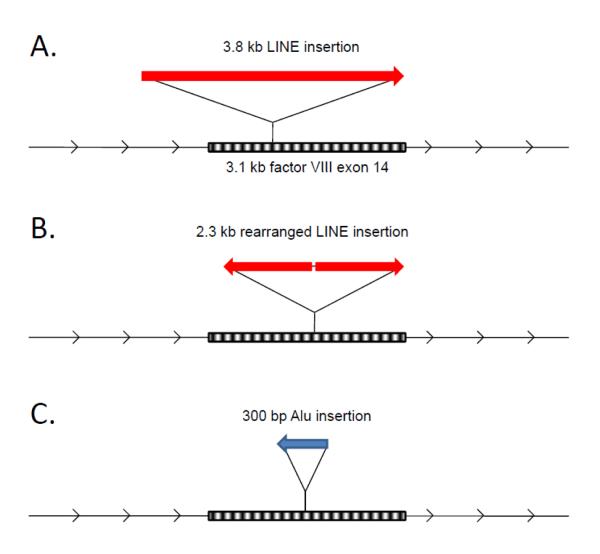
Figure 3. L1 and Alu insertions in exon 14 of F8 causing hemophila A.
Three examples of retrotransposition events causing loss-of-function of factor VIII are illustrated. All have occurred within exon 14, which is a 3.1 kb exon entirely protein coding (striped pattern). A The first is a 3.8 kb, 5′ truncated L1 LINE insertion in sense orientation that occurred with a 12 nucleotide target site duplication about 1 kb 3′ of the exon start. B The second retrotransposition event described resulted in insertion of 2.3 kb of rearranged 3′ L1 LINE sequence close to the midpoint of exon 14. It was flanked by a 13 bp target site duplication. These two L1 LINE insertions were reported by Kazazian and colleagues [1] and appeared to be new insertions in the affected probands. C The third insertion shown is an Alu insertion interrupting the same exon and oriented in antisense with respect to the F8 gene. It was reported by Sukarova et al. [24] who retrospectively identified the insertion in familial carriers, the affected patient’s mother and maternal grandmother.
Hemophilia B similarly is caused by deficiency of coagulation factor IX, and transposon insertions have been found repeatedly, although in aggregate uncommonly, to be the underlying genetic lesion at the 33 kb F9 locus on chromosome Xq27.1. Exonic insertion of partial L1 LINE sequences have been discovered to interrupt exon 5 (also known as exon E; [26]) and exon 7 (exon G) [27]. As well, three reports of exonic Alu insertions have been described by Li and colleagues [26] (exon 8, exon H), Vidaud et al. [28] (exon 5), and Wulff et al. [29] (exon 5).
As we will describe further below, retrotransposition is not the sole mechanism by which LINEs and SINEs disrupt gene function; they can also be substrates for inappropriate recombination events (see TEs as substrates for genomic instability). Alu sequences in particular may have features inherent to their structure that create a propensity for non-allelic homologous recombination (discussed by Kolomietz et al. [30]). Additionally, they may participate in rearrangements of functional consequence frequently simply because of their sheer numbers and genomic distribution within relatively GC and gene-rich regions. Such Alu-mediated recombinations have resulted in large deletions of the F8 locus causing hemophilia A in several case studies [31-33]. Another example of retrotransposon-mediated recombination events related to the hemophilias involves hemophilia C and the autosomal coagulation factor XI-encoding locus. Factor IX is upstream of the ‘tenase’ complex, and patients with loss or partial loss of F11 function can present with a spectrum of bleeding tendencies. Mitchell et al. reported two individuals hemizygous for a 31.5 kb deletion encompassing the entire factor XI coding sequence as consequence of a rearrangement between flanking Alu repeat regions [34].
Transposable elements and red blood cell production
Hemoglobin (Hb) is a major protein constituent in red blood cells and carries oxygen to tissues bound to an iron containing porphyrin ring. Adult hemoglobin is a tetramer composed of two α protein subunits and two β subunits encoded by duplicated neighboring α genes (α1 and α2) on chromosome 16p13.3 pter and a single β gene near 11p14. Hemoglobinopathies are caused by coding sequence changes or hypomorphic / loss-of-function alleles at the Hb gene loci. Respectively, these mutations result in qualitative (hemoglobin variant) or quantitative (thalassemia-type) defects in globin expression. Here we will briefly review some α-thalassemias and β-thalassemias resulting from Alu and L1 LINE-associated recombination deletion events. Genomic deletions are the predominant causes of thalassemias, though TE involvement at the breakpoints has been variably reported.
In early genetic studies of α-thalassemic patients, it became clear that Alu-repeat mediated homologous recombination events would underlie a substantial number of the pathogenic deletions [35]. Indeed, Nicholls et al. mapping various deletions in these patients were the first to describe Alu-Alu recombination in mapping a 62kb interval loss in the (α α)RA disease locus. In addition to being the first example of Alu-Alu recombination as a mechanism of genetic disease, their finding also was the first example of recombination between interspersed repeats described in higher eukaryotes. The report has been followed by other examples of deletions at this locus with loss of genetic material between two Alu breakpoints, including roughly 30 kb deletions at the α0 DUTCH allele [36] and α0 FIL allele [37] in Caucasian and Far East Asian families.
Unlike the α-globin deletions where Alu-mediated recombination seems to play a prominent role, Alus do not mediate large number of recombinations causing β-thalassemias [38]. Non-retrotransposition recombination leading to deletion of the β-globin locus do tend to involve L1 LINE sequences 3′ of the locus [39-50] although the upstream breakpoints in these cases do not share homology to L1, and ultimately roles of the L1 sequences in incurrence of the rearrangements are unclear.
To our knowledge Alu retrotransposition events (de novo insertions) have not been reported to disrupt the hemoglobin loci as has been reported for several hemophilias. In considering other red blood cell phenotypes, an exonic AluYa5 insertion in the porphobilinogen deaminase gene (PBGD) has been described to cause an acute intermittent porphyria [51]. In contrast, an intronic L1 LINE insertion (L1b-thal) within 1 kb of coding exonic sequence is documented to have compromised β-globin production in a case of β-thalassemia ([52], see also the dbRIP entry).
Finally, an intriguing case of hereditary elliptocytosis merits special mention as we consider retrotransposition events that have affected red cell production. A novel type of retroelement was found to have inserted into exon 5 of the α spectrin gene (SPTA1), which encodes an intracellular cytoskeletal protein associated closely with the red blood cell membrane and fundamental for maintaining the anucleate red cell’s biconcave disk shape. The rearranged TE, later recognizable as part of the SVA group [22, 16], caused exon skipping and a truncated spectrin protein [53]. The report represented the first case of SVA retrotransposition causing genetic disease.
Hematopoietic neoplasias
In 2007, about 45,000 new cases of leukemia and about 90,000 new cases of lymphoma and myeloma were diagnosed in the United States, accounting for about 10% of all cancer diagnoses. The overwhelming proportion is not considered heritable per se, though relative risk studies examining many of these disorders indicate familial associations, and genome-wide association studies are beginning to describe specific risk alleles in seemingly sporadic cases. There are many somatic genetic changes associated with these malignancies, and many involve chromosomal translocations and other forms of chromosomal and genetic instability. Their comprehensive molecular characterization currently is only in initial phases of meaningfully contributing to how these cancers are classified and managed. The extent to which TEs have roles in these diseases either as inherited predisposing factors or as somatic mutagens has been quite underexplored, probably because of technical limitations in identifying polymorphisms. We will begin this section by describing the understood function of transposon methylation and how this epigenetic silencing is lost in some human hematolymphoid neoplasias. Then we will describe how expressed, accessible, or otherwise inappropriately regulated TEs have been implicated in the malignant pathophysiology, focusing on their: (i.) potential effects on neighboring gene expression; (ii.) ability to serve as sites for non-homologous DNA recombination; and (iii.) ability to act as insertional mutagens in animal models.
Regulation of retrotransposons in hematopoietic neoplasias
Elaborate host defense mechanisms have developed to suppress essentially each phase of the retrotransposon life cycle, and better appreciation of these will likely be important in recognizing pathologies promoted by somatic TE expression or retrotransposition. There is intriguing evidence that some of these mechanisms are compromised with aging [54], environmental [55, 56] or chemotherapeutic exposures [57, 58] or in cancerous conditions ([59], and see below) in ways that may permit untoward retrotransposition events.
DNA methylation appears to be a cornerstone mechanism of TE silencing, and will be our focus in this section. This heritable, relatively stable epigenetic mark involves covalent attachment of a methyl group to the 5-C position of cytosine in the context of a CG dinucleotide (meCG). Most TEs are highly methylated at CpG islands in non-cancerous, appropriately differentiated somatic tissues. Mouse models indicate that the establishment and maintenance of methylation marks at TEs during early development are reliant on DNA methyltransferases DNMT3a/DNMT3b and DNMT1 [60-62]; a DNMT1 hypomorphic mouse cancer phenotype is discussed in the final section of this review ([63]; see TEs as insertional mutagens).
Recently, proteins apart from methyltransferases have been shown to be additional necessary regulators of genome-wide methylation levels in mice, and methylation of TEs specifically. An example with particular relevance to hematology is helicase, lymphoid specific (HELLS, or LSH) which belongs to the SNF2/helicase family of chromatin remodelling proteins. LSH physically associates with DNMT3a and DNMT3b [64] and LINE, SINE, and IAP genomic loci [65] in immunoprecipitation experiments in vitro. Lsh null mice die in the perinatal period [66], and tissues and embryonic fibroblasts derived from these animals show inappropriate hypomethylation of LINEs, SINEs, and IAPs [67] and derepressive histone H3 and H4 acetylation marks in association with IAPs [65]. Interestingly, hematopoietic precursors isolated from Lsh-/- fetal liver could be transplanted to irradiated recipient mice by Fan and colleagues; these cells were poor at marrow reconstitution and led in multiple instances to development of donor-derived erythroleukemias with genomic hypomethylation and increased IAP transcript expression as shown by RT-PCR [68]. If and how these derepressed TEs contribute functionally to the pathogenesis are as yet unknown.
Loss of TE methylation in various human cancers has been reported now by several groups. A handful of examples among the hematopoietic neoplasias include chronic lymphocytic leukemia (CLL), multiple myeloma (MM), and transformed or ‘blast crisis’ phase chronic myeloid leukemia (CML). Genome-wide hypomethylation and hypomethylation specifically of L1 LINEs in human B-cell CLL were reported in 1992 by two groups, Wahlfors et al. [69] and Dante et al. [70], respectively. These authors used restriction enzyme digests to differentiate between CCGG and CCmeGG sequences, with analysis of enzyme effect by gel electrophoresis alone or Southern blot. Using bisulfite treatment followed by sequencing, Bollati et al. very recently reported that both LINE and Alu sequences are hypomethylated in MM, and moreso in plasma cell leukemia cases [71]. Perhaps the most significant correlation between repeat hypomethylation and clinical parameters has been accomplished in the case of CML. Using a methylation-specific PCR, Roman-Gomez, et al. reported in 2005 that L1 LINE hypomethylation was appreciable in the majority of ‘blast crisis’ phase CML cases (about 75%) and in a subset of chronic phase CML cases (about 40%). Very importantly, in the latter, LINE hypomethylation was associated with poor cytogenetic response to therapy and worse progression-free survival [72]. The group subsequently reported hypomethylation of Alus in both CML disease phases, a finding more pronounced in blast crisis [73].
Whether hypomethylation of retrotransposons is an epiphenomenon of neoplasia or has a causal role in disease acceleration (i.e., promotes clonal evolution in these chronic malignancies) remains a subject of speculation. The answer to this and other unaddressed questions pertaining to TE expression in hematologic neoplasias can be expected to have important implications for how we view iatrogenic hypomethylation of transposable elements as epigenetic therapies are proving at least partially effective and are expanding in clinical use in myelodysplastic syndromes (MDS) and leukemias [74, 75].
Disruption of gene expression
Transposon insertions can have a variety of effects on gene expression. In the previous section reviewing hemophilias, we described several examples of loss-of-function mutations caused by disruption of open reading frame sequences. TEs can also compromise transcriptional elongation through an intron, disrupt splice site utilization around them, and create splices that inappropriately contribute TE sequence to processed mRNAs (‘exonization’; reviewed in [76] and [77]).
Several known polymorphic transposon insertions are catalogued in the database of retrotransposon insertion polymorphisms (dbRIP) [78] in gene loci with recognized roles in hematopoiesis; many of these are intronic retroelement insertions of unknown significance (Table 1). There is little doubt that the application of bench-based genome-wide transposon mapping strategies (such as TIP-chip [79] developed in yeast) to the human genome and in silico analyses of genome project data (such as reported in [80] for human and in [81] for mouse genomes) will contribute substantially to this list. In parallel, it will be key for biomedical researchers to understand functional consequences of TE insertional polymorphisms. At this point in time, early in our cataloguing of these insertions, most published TE polymorphisms are fairly high frequency alleles with no established effect on gene expression. In addition to hypothesis-driven candidate allele approaches, perhaps genome-wide association studies based on transposon detection rather than SNPs (single nucleotide polymorphisms) will be needed to glean their importance in blood dyscrasias and cancer susceptibility.
Table 1. Known transposable element insertion polymorphisms of interest in hematopathology.
Genomic locations of insertions reported to the database of retrotransposon insertion polymorphisms (dbRIP) (39) were downloaded and annotated with respect to nearby reference genes by comparison to RefSeq gene positions. Batch queries of PubMed were performed to identify genes with high numbers of citations when searched in combination with the terms ‘leukemia’, ‘lymphoma’, ‘myeloma’, ‘MDS’, or ‘coagulation’. Some insertions, many exonic, are associated with diseases as indicated above or described further in the text; functions for many intronic insertional polymorphisms, if any, are unknown.
| Gene abbreviation |
Gene name | Gene function | Transposable element (hg17) |
dbRIP ID | References |
|---|---|---|---|---|---|
| ACE | angiotensin I converting enzyme |
cardiovascular homeostasis; roles in hematopoiesis and possibly leukemia cell proliferation |
intronic Ya5 Alu chr17:58919633 |
RIP_Alu_chr17_058_01 | (1) |
| BRCA1 | breast cancer 1, early onset |
tumor suppressor gene; inherited mutations associated with increased risk of hematopoietic cancers in some studies; methylated or otherwise downregulated in several hematopoietic neoplasias |
exonic Sq Alu chr17:38499348 |
RIP_Alu_chr17_038_01 | (2, 3) |
| BRCA2 | breast cancer 2, early onset |
tumor suppressor gene; inherited mutations associated with increased risk of hematopoietic cancers in some studies |
exonic Yc1 Alu chr13:31851500; exonic Ya5 Alu chr13:31791298 |
RIP_Alu_chr13_031_01 RIP_Alu_chr13_031_02 |
(2, 3) |
| CBLB | casitas B-cell lymphoma b |
E3 ubiquitin ligase associated with tyrosine kinase degradation; mutations described in AML |
intronic L1Hs chr3:106956551- 106956812 |
RIP_L1_chr3_106_01 | (4) |
| CDH13 | cadherin 13 | cell-cell adhesion; methylated in several types of leukemias and large B-cell lymphoma |
L1HS; Ya5 Alu; Yb7 Alu; and Yb8 Alu; all intronic 16q23.3 |
RIP_L1_chr16_082_01 RIP_Alu_chr16_081_03 RIP_Alu_chr16_082_01 RIP_Alu_chr16_081_01 |
(5-8) |
| DCC | deleted in colorectal cancer |
membrane protein tumor suppressor; shows deletion or allelic loss in several hematopoietic neoplasias |
intronic Ya5 Alu; chr18:49179990 |
RIP_Alu_chr18_049_01 | (9-11) |
| F8 | coagulation factor VIII |
intrinsic coagulation cascade participant; several L1 and Alu insertions have been described to cause F8 loss-of-function mutations and hemophilia A; see text |
3 exonic and 1 intronic TE insertions; see text and Figure 3 |
RIP_L1_chrX_153_01 RIP_L1_chrX_153_02 RIP_Alu_chrX_153_01 RIP_Alu_chrX_153_02 |
(12-14) |
| F9 | coagulation factor IX |
intrinsic coagulation cascade participant; several L1 and Alu insertions have been described to cause F9 loss-of-function mutations and hemophilia B; see text |
5 exonic TE insertions; see text |
RIP_L1_chrX_138_01 RIP_L1_chrX_138_02 RIP_Alu_chrX_138_03 RIP_Alu_chrX_138_04 RIP_Alu_chrX_138_05 |
(15-18) |
| F10 | coagulation factor X |
“common pathway” clotting factor at the intersection of intrinsic and extrinsic coagulation cascades; intronic Alu insertion of no understood consequence |
intronic Ya5 Alu; chr13:112838951- 112839264 |
RIP_Alu_chr13_112_01 | (19) |
| FAS | TNF receptor superfamily, member 6; CD95 |
key component of cellular apoptotic pathways; Alu insertion causes splicing variation and autoimmune lymphoproliferative syndrome (ALPS) |
intronic Alu; chr10:90763030 |
RIP_Alu_chr10_090_01 | (20) |
| FHIT | fragile histidine triad gene |
tumor suppressor with a role in purine metabolism; the gene locus includes a fragile site on 3p14.2 rearranged in several malignancies; downregulated in ALL and AML; downregulated in large B-cell lymphoma |
intronic L1Hs; chr3:59827943- 59827944 |
RIP_L1_chr3_059_01 | (21, 22) |
| HBB | hemoglobin, β | one of two subunits in adult hemoglobin tetramers; loss-of-function mutations described to cause β-thalassemias; intronic L1 insertion (L1b-thal) associated with disease; see text |
intronic L1; chr11:5203631 |
RIP_L1_chr11_005_01 | (23) |
| MAX | MYC associated factor X |
basic helix-loop-helix leucine zipper (bHLHZ) family of transcription factors; involved in a variety of cell processes and associates with MYC; expression associated with better outcomes in small studies of pediatric ALL |
intronic Ya5; chr14:64596912- 64597181 |
RIP_Alu_chr14_064_01 | (24, 25) |
| MGMT | O6- methylguanine- DNA methyltransferase |
DNA repair of alkylated guanine residues; methylated or silenced in many malignancies: acute leukemias, large B-cell lymphomas, MDS/MPDs, and myeloma; associated with responsiveness to alkylating chemotherapies |
intronic Alu; chr10:131263887 |
RIP_Alu_chr10_131_01 | (26) |
| MYH11 | myosin, heavy chain 11, smooth muscle |
contractile protein; fused with core binding factor-β in AML associated with inv(16) |
intronic Yb8 Alu; chr16:15822993- 15823311 |
RIP_Alu_chr16_015_01 | (27) |
| NF1 | neurofibromin 1 | tumor suppressor; loss of function mutations associated with juvenile AMML and other myeloid leukemias; exonic Alu insertion causes neurofibromatosis |
intronic L1Hs; chr17:26683724; exonic Ya5 Alu chr17:26687734 |
RIP_L1_chr17_026_01 RIP_Alu_chr17_026_01 |
(28, 29) |
| NOTCH2 | Notch homolog 2 (Drosophila) |
transmembrane signalling protein; activation or gain-of-function associated with CLL and large B-cell lymphomas |
intronic Ya5 Alu; chr1:120322150 |
RIP_Alu_chr1_120_01 | (30, 31) |
| PBGD | porphobilinogen deaminase |
enzyme involved in metabolism of porphyrin; exonic insertion associated with development of acute intermittent porphyria; see text |
exonic Ya5 Alu; chr11:118465033 |
RIP_Alu_chr11_118_01 | (32) |
| PLAT | plasminogen activator, tissue |
fibrinolytic enzyme with important roles in clot degradation, and cellular migration, proliferation and apoptosis |
intronic Ya5 Alu; chr8:42159064 |
RIP_Alu_chr8_042_01 | (33, 34) |
| RAD50 | RAD50 homolog (S. cerevisiae) |
DNA damage repair and signalling; implicated in several types of neoplasia |
intronic Ya5 Alu; chr5:131988139- 131988491 |
RIP_Alu_chr5_131_01 | (35) |
| TIAM1 | T-cell lymphoma invasion and metastasis 1 |
guanine nucleotide exchange factor discovered by proviral tagging and selection for invasive T- cell lymphoma variants |
intronic Yb7 Alu; chr21:31706107 |
RIP_Alu_chr21_031_02 | (36) |
| TF | transferrin | iron binding protein; allows cellular iron uptake through CD71, a MYC-induced activation marker on many hematopoietic malignancies |
intronic Yb8 Alu; chr3:134972999- 134973307 |
RIP_Alu_chr3_134_01 | (37) |
| XG | Xg blood group | membrane glycoprotein Xg precursor; responsible for the Xg blood grouping system |
intronic Yc1 Alu; chrX:2725353 |
RIP_Alu_chrX_002_01 | (38) |
References:
Haznedaroglu IC, Ozturk MA 2003 Towards the understanding of the local hematopoietic bone marrow renin-angiotensin system. Int J Biochem Cell Biol 35:867-880
Teugels E, De Brakeleer S, Goelen G, Lissens W, Sermijn E, De Greve J 2005 De novo Alu element insertions targeted to a sequence common to the BRCA1 and BRCA2 genes. Hum Mutat 26:284
Friedenson B 2007 The BRCA1/2 pathway prevents hematologic cancers in addition to breast and ovarian cancers. BMC Cancer 7:152
Caligiuri MA, Briesewitz R, Yu J, Wang L, Wei M, Arnoczky KJ, Marburger TB, Wen J, Perrotti D, Bloomfield CD, Whitman SP 2007 Novel c-CBL and CBL-b ubiquitin ligase mutations in human acute myeloid leukemia. Blood 110:1022-1024
Kroeger H, Jelinek J, Estecio MR, He R, Kondo K, Chung W, Zhang L, Shen L, Kantarjian HM, Bueso-Ramos CE, Issa JP 2008 Aberrant CpG island methylation in acute myeloid leukemia is accentuated at relapse. Blood 112:1366-1373
Roman-Gomez J, Castillejo JA, Jimenez A, Cervantes F, Boque C, Hermosin L, Leon A, Granena A, Colomer D, Heiniger A, Torres A 2003 Cadherin-13, a mediator of calcium-dependent cell-cell adhesion, is silenced by methylation in chronic myeloid leukemia and correlates with pretreatment risk profile and cytogenetic response to interferon alfa. J Clin Oncol 21:1472-1479
Roman-Gomez J, Jimenez-Velasco A, Castillejo JA, Agirre X, Barrios M, Navarro G, Molina FJ, Calasanz MJ, Prosper F, Heiniger A, Torres A 2004 Promoter hypermethylation of cancer-related genes: a strong independent prognostic factor in acute lymphoblastic leukemia. Blood 104:2492-2498
Ogama Y, Ouchida M, Yoshino T, Ito S, Takimoto H, Shiote Y, Ishimaru F, Harada M, Tanimoto M, Shimizu K 2004 Prevalent hyper-methylation of the CDH13 gene promoter in malignant B cell lymphomas. Int J Oncol 25:685-691
Miyake K, Inokuchi K, Dan K, Nomura T 1993 Alterations in the deleted in colorectal carcinoma gene in human primary leukemia. Blood 82:927-930
Porfiri E, Secker-Walker LM, Hoffbrand AV, Hancock JF 1993 DCC tumor suppressor gene is inactivated in hematologic malignancies showing monosomy 18. Blood 81:2696-2701
Younes A, Zhao S, Jendiroba D, Kleine HD, Cabanillas F, Andreeff M 1995 Decreased expression of the deleted in colorectal carcinoma gene in non-Hodgkin’s lymphoma. Blood 85:2813-2816
Kazazian HH, Jr., Wong C, Youssoufian H, Scott AF, Phillips DG, Antonarakis SE 1988 Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 332:164-166
Sukarova E, Dimovski AJ, Tchacarova P, Petkov GH, Efremov GD 2001 An Alu insert as the cause of a severe form of hemophilia A. Acta Haematologica 106:126-129
Ganguly A, Dunbar T, Chen P, Godmilow L, Ganguly T 2003 Exon skipping caused by an intronic insertion of a young Alu Yb9 element leads to severe hemophilia A. Human Genetics 113:348-352
Li X, Scaringe WA, Hill KA, Roberts S, Mengos A, Careri D, Pinto MT, Kasper CK, Sommer SS 2001 Frequency of recent retrotransposition events in the human factor IX gene. Human Mutation 17:511-519
Mukherjee S, Mukhopadhyay A, Banerjee D, Chandak GR, Ray K 2004 Molecular pathology of haemophilia B: identification of five novel mutations including a LINE 1 insertion in Indian patients. Haemophilia 10:259-263
Vidaud D, Vidaud M, Bahnak BR, Siguret V, Gispert Sanchez S, Laurian Y, Meyer D, Goossens M, Lavergne JM 1993 Haemophilia B due to a de novo insertion of a human-specific Alu subfamily member within the coding region of the factor IX gene. European journal of human genetics : EJHG 1:30-36
Wulff K, Gazda H, Schroĺ^der W, Robicka-Milewska R, Herrmann FH 2000 Identification of a novel large F9 gene mutation-an insertion of an Alu repeated DNA element in exon e of the factor 9 gene. Human mutation 15:299
Hertzberg M 1994 Biochemistry of factor X. Blood Rev 8:56-62
Tighe PJ, Stevens SE, Dempsey S, Le Deist F, Rieux-Laucat F, Edgar JD 2002 Inactivation of the Fas gene by Alu insertion: retrotransposition in an intron causing splicing variation and autoimmune lymphoproliferative syndrome. Genes Immun 3 Suppl 1:S66-70
Iwai T, Yokota S, Nakao M, Nakazawa N, Taniwaki M, Kimura T, Sonoda Y, Kaneko H, Okuda T, Azuma H, Oka T, Takeda T, Watanabe A, Kikuta A, Asami K, Sekine I, Matsushita T, Tsuchiya T, Mimaya J, Koizumi S, Ohta S, Miyake M, Takaue Y, Iwai A, Fujimoto T 1998 Frequent aberration of FHIT gene expression in acute leukemias. Cancer Res 58:5182-5187
Chen PM, Yang MH, Hsiao LT, Yu IT, Chu CJ, Chao TC, Yen CC, Wang WS, Chiou TJ, Liu JH 2004 Decreased FHIT protein expression correlates with a worse prognosis in patients with diffuse large B-cell lymphoma. Oncol Rep 11:349-356
Divoky V, Indrak K, Mrug M, Brabec V, Huisman T, Prchal J 1996 A novel mechanism of beta thalassemia: the insertion of L1 retrotransposable element into beta globin IVS II. . Blood 88
Hurlin PJ, Huang J 2006 The MAX-interacting transcription factor network. Semin Cancer Biol 16:265-274
Yuza Y, Kawakami M, Takagi K, Yamazaki Y, Urashima M 1999 Max protein expression is associated with survival of children with lymphoblastic lymphoma. Pediatr Int 41:637-640
Esteller M 2003 Profiling aberrant DNA methylation in hematologic neoplasms: a view from the tip of the iceberg. Clin Immunol 109:80-88
Liu PP, Hajra A, Wijmenga C, Collins FS 1995 Molecular pathogenesis of the chromosome 16 inversion in the M4Eo subtype of acute myeloid leukemia. Blood 85:2289-2302
Wallace MR, Andersen LB, Saulino AM, Gregory PE, Glover TW, Collins FS 1991 A de novo Alu insertion results in neurofibromatosis type 1. Nature 353:864-866
O’Marcaigh AS, Shannon KM 1997 Role of the NF1 gene in leukemogenesis and myeloid growth control. J Pediatr Hematol Oncol 19:551-554
Lee SY, Kumano K, Nakazaki K, Sanada M, Matsumoto A, Yamamoto G, Nannya Y, Suzuki R, Ota S, Ota Y, Izutsu K, Sakata-Yanagimoto M, Hangaishi A, Yagita H, Fukayama M, Seto M, Kurokawa M, Ogawa S, Chiba S 2009 Gain-of-function mutations and copy number increases of Notch2 in diffuse large B-cell lymphoma. Cancer Sci 100:920-926
Rosati E, Sabatini R, Rampino G, Tabilio A, Di Ianni M, Fettucciari K, Bartoli A, Coaccioli S, Screpanti I, Marconi P 2009 Constitutively activated Notch signaling is involved in survival and apoptosis resistance of B-CLL cells. Blood 113:856-865
Mustajoki S, Ahola H, Mustajoki P, Kauppinen R 1999 Insertion of Alu element responsible for acute intermittent porphyria. Hum Mutat 13:431-438
Balsara RD, Ploplis VA 2008 Plasminogen activator inhibitor-1: the double-edged sword in apoptosis. Thromb Haemost 100:1029-1036
Rijken DC, Lijnen HR 2009 New insights into the molecular mechanisms of the fibrinolytic system. J Thromb Haemost 7:4-13
Williams RS, Williams JS, Tainer JA 2007 Mre11-Rad50-Nbs1 is a keystone complex connecting DNA repair machinery, double-strand break signaling, and the chromatin template. Biochem Cell Biol 85:509-520
Habets GG, Scholtes EH, Zuydgeest D, van der Kammen RA, Stam JC, Berns A, Collard JG 1994 Identification of an invasion-inducing gene, Tiam-1, that encodes a protein with homology to GDP-GTP exchangers for Rho-like proteins. Cell 77:537-549
O’Donnell KA, Yu D, Zeller KI, Kim JW, Racke F, Thomas-Tikhonenko A, Dang CV 2006 Activation of transferrin receptor 1 by c-Myc enhances cellular proliferation and tumorigenesis. Mol Cell Biol 26:2373-2386
Tippett P, Ellis NA 1998 The Xg blood group system: a review. Transfus Med Rev 12:233-257
Wang J, Song L, Grover D, Azrak S, Batzer MA, Liang P 2006 dbRIP: a highly integrated database of retrotransposon insertion polymorphisms in humans. Hum Mutat 27:323-329
Adding to the challenge of understanding the consequences of a TE insertion is that the degree to which intronic transposons affect mRNA production appears dependent on cell context. There is precedent that a transposon insertion which appears inconsequential under physiologic circumstances affects mRNA transcript structure in a pathologic condition. Amit et al. reported exonization of an AluSx element (an older Alu insertion incurred during primate evolution) located in intron 1 of a non-protein coding copy of the transcription initiation factor 1A (TIF-1A) gene known as locus 21. Transcriptional effects of this incorporated intronic AluSx sequence upon locus 21-encoded RNA were pronounced in several leukemia cell lines (particularly in 293T, HL60, Dami, U937, KMH2, MT4, CMK, Dg-75, Daudi, and REH cells), while presence of the same Alu did not result in exonization to an appreciable degree in a handful of other leukemia cell lines (Jurkat, Molt3, Raji, K562) and most normal tissues [82].
TEs as substrates for genomic instability
Many leukemias and lymphomas are associated with characteristic recurrent chromosomal abnormalities, both balanced reciprocal translocations and regional gains and losses appreciable at the cytogenetic level. Other clonal pathologies such as some treatment-related AMLs and myelodysplastic syndromes frequently have complex cytogenetics. Roles for retrotransposons in these alterations are appreciated in a few circumstances, although our understanding of the complex interactions between retroelements and DNA repair pathways in cancer development is far from complete. It is clear that retrotransposition if ongoing in human cancers can be associated with genetic instability [83]. It is also postulated that endonuclease activities of HERV proteins and L1-encoded ORF2p may initiate dsDNA breaks that can be resolved, albeit inappropriately, by genomic rearrangements, though there is little direct evidence for this in human cancers. In contrast, dsDNA breaks caused by other insults can potentiate retrotransposition in many experimental systems. Below, we focus on a third, better described relationship, how transposon sequences themselves can be involved as substrates in illegitimate recombination events as a DNA repair mechanism.
Molecular analyses of DNA breakpoints in malignancies with cytogenetic abnormalities reflect two general mechanisms of dsDNA break repair, non-homologous end-joining (NHEJ) and homologous recombinations. NHEJ is by far the most common mechanism of interchromosomal translocation in human neoplasia, though this is not reliant on repetitive sequences and so these fusions will not be considered further here.
Homologous DNA recombination normally involves several stages, namely: (i.) resection of the 5′ ends on either side of an inciting dsDNA break; (ii.) 3′ strand invasion into the homologous sequence in trans; (iii.) new DNA synthesis elongating the invader strand and the ‘captured second end’; and (iv.) resolution of the associated sequences with or without crossing over (Figure 4A and B). When DNA breaks occur within or very close to repetitive DNA, several types of illicit recombinations become possible. These occur between misaligned repeats that share features of minimal efficient processing segments (MEPS) prerequisite for activity of the cell’s homologous recombination machinery. MEPS are usually low-copy repeats (segmental duplications), though highly similar TEs can also function in this capacity. Depending on the location and direction of the ectopic repeat with respect to the invader strand, several variations on homologous recombination have been described. For example, an ectopic repeat can be on the same chromatid resulting in a type of single stranded annealing (SSA) recombination and a deletion product (Figure 4A), or on a sister chromatid or homologous chromosome in the same orientation potentiating a non-allelic homologous recombination (NAHR) with deletion and duplication products (Figure 4A and C). Essentially the same mechanisms can act on two repeats on different chromosomes with crossing over to cause chromosomal translocations, although this appears to be an uncommon mechanism of translocation in leukemias and lymphomas in general. Recombination between intrachromosomal, oppositely-oriented repeats result in intervening sequence inversions. These rearrangements are reviewed by Weinstock et al. [84] and Gu et al. [85]. In addition to dsDNA break-induced NAHR, it has also been proposed that deletions and duplications can be incurred by an intrastrand ‘slip mispairing’ model in the correction of single stranded DNA breaks (Figure 4D).
Figure 4. Repeat-associated genetic recombination events.
A Sister chromatids or homologous chromosomes are illustrated, with hypothetical repetitive sequences shown as blocks. Arrows indicate positions of a dsDNA break near or within one of the repeats. The first step in homologous repair pathways (open arrow) is a 5′ sequence resection, which prepares the 3′potential invader strands (black arrow). In this schematic, one correct (true allelic) pairing option is available to the invader (indicated by the gray arrow marked HR, homologous recombination). Erroneous pairing with a tandem repeat in the same orientation in cis, a type of single stranded annealing (SSA), results in deletion of one of the repeats and the intervening unique sequence. B Appropriate association of the invader strand with an allelic repeat is shown, with subsequent invader strand elongation. The other 3′ end left by the original dsDNA break is involved in a second strand capture that allows for elongation. After the Holiday junction (HJ) traverses sufficient distance during elongation, there is resolution of the complex (open arrow). Depending on whether there has been a cross-over event, there are one of two possible resulting products. True homologous recombination is non-mutagenic. C Misalignment of the homologs caused by repetitive sequence similarity is illustrated. The consequent non-allelic homologous recombination (NAHR) yields two products, one with deletion of one repeat and the sequence that occurred between the repeats, and one with interval duplication. If this occurs between homologous chromosomes, both deletion and duplication products can be expected to persist in the cell. If this occurs between sister chromatids, the abnormal copies are expected to separate in daughter cells, one persisting in a dominant neoplastic clone. Similar rearrangements between oppositely oriented repeats cause inversions of inter-repetitive sequence (not shown). D Intrastrand slip mispairing model for ssDNA break repair is expected to result in duplication after replication without incurrence of the corresponding deletion.
We previously reviewed how TE-mediated sequence homologies can result in deletions at the hemoglobin loci (see Transposable elements and red blood cell production above), reflecting SSA or NAHR with deletion. Several research groups have described transposons at sites of NAHR or slipped intrastrand mispair repair with tandem duplications in hematopoietic neoplasia. Some of the best characterized lesions of this type involve partial tandem duplications (PTDs) at the mixed lineage leukemia (MLL or ALL-1) locus. MLL (mixed lineage leukemia) encodes a histone methyltransferase (H3K4) which regulates expression of HOX genes and other targets; its dysfunction is caused by numerous rearrangements of the gene on 11q23 and associated with poor outcome acute leukemias with immunophenotypic features of either myeloid or lymphoid type. While a major role for TEs in MLL interchromosomal translocations has not been established, breakpoints involving Alus or in close proximity to Alus have been described [86, 87]. Several groups have reported PTDs at MLL occur between intronic Alu elements in cases with normal conventional cytogenetics. Several variant duplications create abnormal fusion MLL mRNAs; those associated with leukemias are predicted to encode an abnormal protein product with extra coding exons [88-90]. The paired Alu elements involved share short sequence identities at the breakpoint site, ranging from 8 bp to 25 bp, as reported by Strout et al. [90] and So et al., [89] respectively, but do not demonstrate very high homology overall. Interestingly, Schnittgen et al. [91] demonstrated that sensitive nested RT-PCR and seminested genomic DNA PCR techniques allow for detection of this type of MLL duplication in healthy volunteers.
In childhood T-cell lymphoblastic lymphoma/leukemia, similar tandem duplications with total gene encompassing amplification of the myeloblastosis viral oncogene homolog (the MYB proto-oncogene) frequently are mediated by hybridization of non-allelic, neighboring Alu elements 3.5 kb 5′ and 73.5 kb 3′ of the MYB gene [92]. Very interestingly, like MLL PTDs, O’Neil et al. who report this also found evidence for the rearrangement occurring somatically in peripheral blood mononuclear cells in healthy individuals. However, this was not the case in buccal swab samples. The authors suggest that something about the hematopoietic lineages or the process of T-cell maturation is responsible for susceptibility to DNA breaks or defects in repair pathways.
Despite the high incidence of Alu-mediated MLL and MYB rearrangements in healthy individuals, these are thought to directly contribute to the leukemia phenotype and not merely reflect ‘passenger’ mutations when seen in disease. Rather, it seems that the alteration is only oncogenic when it arises in a particular type of cell with ‘cancer stem cell’ potential and/or in the context of co-occurrence with other genetic lesions. Discovery of these rearrangements at low levels in normal individuals is intriguing in that they implicate TE related rearrangements in the earliest phases of neoplastic transformation.
As we mentioned previously, Alu sequences may, despite their short size, be especially efficient substrates for recombination events as compared to LINEs or other longer TEs. L1 LINEs, however, have been implicated in some genetic lesions in leukemias and lymphoma. In childhood precursor B-cell acute lymphoblastic leukemia with the TEL-AML translocation, for example, there is frequent loss of the short arm of chromosome 12. Wiemels et al. [93] characterized these breakpoints and discovered that unlike the TEL-AML reciprocal translocation, an unexpectedly high number of the del(12p) involved repetitive regions, of which 8 of 12 cases with the breakpoint involved LINEs. Non-templated nucleotides at these breakpoints are thought to reflect activity of terminal deoxynucleotidyl transferase (TdT) expressed in these leukemias. Though some chromosomal translocations associated with ALL, including TEL-AML, are detectable at high frequencies in the general population in archived cord blood samples, this does not seem true of the del(12p) lesions. Thus, the authors suggest L1 LINE-mediated loss of 12p is a secondary occurrence in childhood ALL, taking place in cells with antecedent TEL-AML translocation and adding to their commitment to transformation.
TEs as somatic insertional mutagens
Infrequent case reports have demonstrated that retrotransposons can play important roles as somatic mutagens in human cancers [94, 95], and it is speculated that TEs may similarly act as insertional mutagens to potentiate leukemias and lymphomas. Economou-Pachnis et al. [96] identified an Alu insertion at the Moloney-MuLV integration site 2 (MLVI2) locus in a human B-cell lymphoma cell line. It was not found in several other cell lines examined, and though normal tissue was not available for comparison, the authors speculated the insertion may reflect a somatic event largely because of described functions of the gene region in murine models of lymphomagenesis [97].
The question as to whether somatic TE retrotransposition events contributes to human leukemias and lymphomas has until very recently been difficult to address experimentally as conventional technologies have had limited applicability for finding new TE insertions among the hundreds of thousands of established genomic integrations. A handful of groups however are now addressing this problem by adopting high throughput genomics approaches (microarray and deep sequencing platforms) normally ineffectual or limited in efficacy for locating retrotransposons for assays specifically to detect still active TE families genome-wide.
In the meantime, the most compelling experimental data for retrotransposons as insertional mutagens in hematopoietic neoplasias come to us from studies in rodent model systems. In a recent report, Kirilyuk and colleagues describe identifying intact endogenous L1 LINE RNAs and L1 LINE-encoded ORF1p and ORF2p proteins in a rat myeloid leukemia cell line [98], corroborating earlier work with these cells [99]. Moreover, using a retrotransposition reporter assay, Kirilyuk et al. demonstrated that the leukemia line could support reverse transcription and integration of a tagged L1 element. More direct evidence for endogenous TE insertion promoting cancers is provided by a mouse model of thymic lymphomagenesis, the compound heterozygote mouse harboring a combination of one hypomorphic and one null allele for Dnmt1 [63]. The lymphomas demonstrate a profound genome-wide hypomethylation and consistently acquire an extra copy of chromosome 15. It is now clear that these tumors in many cases are also potentiated by insertion of endogenous IAP elements at the Notch1 locus [100]. This was seen in 7 of 16 lymphomas studied by Howard et al.; all were somatic IAP integrations (i.e., unique to the lymphoma in each case) occurring within a genomic interval between coding exons D and F. Notch1 transcripts were found to be overexpressed in the tumors and transcript variants were associated with several of the insertions.
While the hypothesis that retrotransposition events contribute in a recurrent manner to human cancers has yet to be meaningfully addressed, it is one of exquisite clinical relevance. Unlike many forms of mutation and cytogenetic instability described in malignant conditions that are as yet perceived as inevitable, retrotransposition processes may be targetable by agents in our increasing armamentarium of antiretroviral pharmaceuticals.
Conclusions
As we lose assumptions about ‘junk DNA’ and gain appreciation for activities of transposable elements, we are coming to understand our genomes as more animated and interesting. Work of transposon biologists and truly pioneer physician-scientists preceding us have indicated that roles of TEs are as yet understudied in clinical hematology, helped establish technologies to address some fundamental questions of clinical relevance, and captured the intrigue of young researchers. It is relatively certain that the importance of TEs in medicine is not limited to hematopoietic disorders. However, unique features of these diseases have made them amenable to study and a series of firsts in our understandings of genetic mechanisms of disease – and may also position hematology at the forefront of translational studies in transposon biology.
Acknowledgements
We apologize to those whose relevant works were not cited because of our oversights. A.M.S., K.H.B., and research in the Burns laboratory are supported by a K08 award from the National Cancer Institute and a Career Award for Medical Scientists from the Burroughs Wellcome Foundation. The investigator is mentored in her research work by Jef D. Boeke and Curt I. Civin. D.E.S. is supported by start-up funds from The Ohio State University Comprehensive Cancer Center.
Abbreviations (excluding gene names)
- TE
transposable element; refers to a mobile DNA sequence
- L1 LINE
Long Interspersed Element; a prevalent type of Class I retrotransposon capable of autonomous retrotransposition
- TPRT
Target Primed Reverse Transcription; an integration mechanism for retrotransposons that uses the 3′ hydroxyl group exposed by cleavage of a DNA strand as the primer for reverse transcription
- Retroelement
a ‘copy-paste’ type of transposon replicated via an RNA intermediate
- SVA
a composite parasitic retroelement named after its main sequence components, SINE, VNTR and Alu
- IAP
Intracisternal A-particle; a prevalent endogenous retrovirus in mouse
- LTR
Long terminal repeat; a type of autonomous retrotransposon closely related to exogenous retroviruses
- ERV/HERV
Endogenous Retrovirus/Human Endogenous Retrovirus; LTR retrotransposons
- SINE
Short Interspersed Element; includes the non-autonomous Alu and SVA retrotransposons that rely on LINE-encoded proteins for their reverse transcription
- ORF
Open Reading Frame; a stretch of DNA sequence organized into codon triplets for protein coding and uninterrupted by ‘stop’ codons
- UTR
Untranslated Region; part of a retroelement or RNA sequence that is not protein-coding
- ALL
acute lymphoblastic leukemia/lymphoma; aggressive malignancies of lymphoid (B-cell or T-cell) precursor cells, subclassified based on presentation, morphology, leukemic cell immunophenotype, and tumor-specific genetic alterations
- AML
acute myeloid leukemia; proliferation of immature myeloid cells, includes de novo cases classified by degree of cell maturation and genetic alterations as well as cases evolving from MDS/MPD spectrum pathologies
- CLL
chronic lymphocytic leukemia; a common leukemia in adults characterized by proliferation of mature B-cell lymphocytes
- CML
chronic myeloid leukemia; a neoplastic expansion of mature myeloid white blood cells (in chronic phase) with the potential to convert to an immature acute leukemia (blast phase)
- MPD
myeloproliferative disorder; a constellation of primary bone marrow disorders resulting in overproduction of blood cell lineages with variable progression to fibrosis and acute leukemia
- MDS
myelodysplastic syndromes; a constellation of primary bone marrow disorders resulting in failed hematopoiesis; clonal, pre-leukemic states
- MM
multiple myeloma; a neoplastic proliferation of plasma cells, immunoglobulin-producing cells derived from the B-cell lineage
- HR
homologous recombination; DNA repair mechanism involving interactions between two segments of similar sequence which may or may not be allelic to one another
- NAHR
non-allelic homologous recombination; results in deletions and duplications of the interval between erroneously paired sequences
- MEPS
minimal efficient processing segments; regions of similar sequence required for homologous recombination (HR) between two DNA segments
- NHEJ
non-homologous end joining; a major mechanism of interchromosomal translocation and fusion gene formation in leukemias and lymphomas that does not rely on homologous sequences
- SSA
single strand annealing; a step in homologous DNA repair pathways wherein non-duplexed DNA hybridizes to a partially complementary sequence along its strand rather than ‘invading’ another strand; typically results in deletion of intervening sequence
- dbRIP
database of retrotransposon insertion polymorphisms in humans [78]
References
- 1.Kazazian HH, Jr., Wong C, Youssoufian H, Scott AF, Phillips DG, Antonarakis SE. Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature. 1988;332:164–166. doi: 10.1038/332164a0. [DOI] [PubMed] [Google Scholar]
- 2.Belancio VP, Hedges DJ, Deininger P. Mammalian non-LTR retrotransposons: for better or worse, in sickness and in health. Genome Res. 2008;18:343–358. doi: 10.1101/gr.5558208. [DOI] [PubMed] [Google Scholar]
- 3.Chen JM, Chuzhanova N, Stenson PD, Ferec C, Cooper DN. Meta-analysis of gross insertions causing human genetic disease: novel mutational mechanisms and the role of replication slippage. Hum Mutat. 2005;25:207–221. doi: 10.1002/humu.20133. [DOI] [PubMed] [Google Scholar]
- 4.Jones JM, Gellert M. The taming of a transposon: V(D)J recombination and the immune system. Immunol Rev. 2004;200:233–248. doi: 10.1111/j.0105-2896.2004.00168.x. [DOI] [PubMed] [Google Scholar]
- 5.Kapitonov VV, Jurka J. RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLoS Biol. 2005;3:e181. doi: 10.1371/journal.pbio.0030181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhou L, Mitra R, Atkinson PW, Hickman AB, Dyda F, Craig NL. Transposition of hAT elements links transposable elements and V(D)J recombination. Nature. 2004;432:995–1001. doi: 10.1038/nature03157. [DOI] [PubMed] [Google Scholar]
- 7.Luan DD, Korman MH, Jakubczak JL, Eickbush TH. Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell. 1993;72:595–605. doi: 10.1016/0092-8674(93)90078-5. [DOI] [PubMed] [Google Scholar]
- 8.Moyes D, Griffiths DJ, Venables PJ. Insertional polymorphisms: a new lease of life for endogenous retroviruses in human disease. Trends Genet. 2007;23:326–333. doi: 10.1016/j.tig.2007.05.004. [DOI] [PubMed] [Google Scholar]
- 9.Mills RE, Bennett EA, Iskow RC, Devine SE. Which transposable elements are active in the human genome? Trends Genet. 2007;23:183–191. doi: 10.1016/j.tig.2007.02.006. [DOI] [PubMed] [Google Scholar]
- 10.Zhang Y, Maksakova IA, Gagnier L, van de Lagemaat LN, Mager DL. Genome-wide assessments reveal extremely high levels of polymorphism of two active families of mouse endogenous retroviral elements. PLoS Genet. 2008;4:e1000007. doi: 10.1371/journal.pgen.1000007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dewannieux M, Dupressoir A, Harper F, Pierron G, Heidmann T. Identification of autonomous IAP LTR retrotransposons mobile in mammalian cells. Nat Genet. 2004;36:534–539. doi: 10.1038/ng1353. [DOI] [PubMed] [Google Scholar]
- 12.Feng Q, Moran JV, Kazazian HH, Jr., Boeke JD. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell. 1996;87:905–916. doi: 10.1016/s0092-8674(00)81997-2. [DOI] [PubMed] [Google Scholar]
- 13.Mathias SL, Scott AF, Kazazian HH, Jr., Boeke JD, Gabriel A. Reverse transcriptase encoded by a human transposable element. Science. 1991;254:1808–1810. doi: 10.1126/science.1722352. [DOI] [PubMed] [Google Scholar]
- 14.Ostertag EM, Kazazian HH., Jr. Biology of mammalian L1 retrotransposons. Annu Rev Genet. 2001;35:501–538. doi: 10.1146/annurev.genet.35.102401.091032. [DOI] [PubMed] [Google Scholar]
- 15.Bennett EA, Keller H, Mills RE, Schmidt S, Moran JV, Weichenrieder O, Devine SE. Active Alu retrotransposons in the human genome. Genome Res. 2008;18:1875–1883. doi: 10.1101/gr.081737.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shen L, Wu LC, Sanlioglu S, Chen R, Mendoza AR, Dangel AW, Carroll MC, Zipf WB, Yu CY. Structure and genetics of the partially duplicated gene RP located immediately upstream of the complement C4A and the C4B genes in the HLA class III region. Molecular cloning, exon-intron structure, composite retroposon, and breakpoint of gene duplication. J Biol Chem. 1994;269:8466–8476. [PubMed] [Google Scholar]
- 17.Cost GJ, Feng Q, Jacquier A, Boeke JD. Human L1 element target-primed reverse transcription in vitro. EMBO J. 2002;21:5899–5910. doi: 10.1093/emboj/cdf592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ono M. Molecular biology of type A endogenous retrovirus. Kitasato Arch Exp Med. 1990;63:77–90. [PubMed] [Google Scholar]
- 19.Maksakova IA, Romanish MT, Gagnier L, Dunn CA, van de Lagemaat LN, Mager DL. Retroviral elements and their hosts: insertional mutagenesis in the mouse germ line. PLoS Genet. 2006;2:e2. doi: 10.1371/journal.pgen.0020002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Babushok DV, Kazazian HH., Jr. Progress in understanding the biology of the human mutagen LINE-1. Hum Mutat. 2007;28:527–539. doi: 10.1002/humu.20486. [DOI] [PubMed] [Google Scholar]
- 21.Batzer MA, Deininger PL. Alu repeats and human genomic diversity. Nat Rev Genet. 2002;3:370–379. doi: 10.1038/nrg798. [DOI] [PubMed] [Google Scholar]
- 22.Ostertag EM, Goodier JL, Zhang Y, Kazazian HH., Jr. SVA elements are nonautonomous retrotransposons that cause disease in humans. Am J Hum Genet. 2003;73:1444–1451. doi: 10.1086/380207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Van de Water N, Williams R, Ockelford P, Browett P. A 20.7 kb deletion within the factor VIII gene associated with LINE-1 element insertion. Thromb Haemost. 1998;79:938–942. [PubMed] [Google Scholar]
- 24.Sukarova E, Dimovski AJ, Tchacarova P, Petkov GH, Efremov GD. An Alu insert as the cause of a severe form of hemophilia A. Acta Haematologica. 2001;106:126–129. doi: 10.1159/000046602. [DOI] [PubMed] [Google Scholar]
- 25.Ganguly A, Dunbar T, Chen P, Godmilow L, Ganguly T. Exon skipping caused by an intronic insertion of a young Alu Yb9 element leads to severe hemophilia A. Human Genetics. 2003;113:348–352. doi: 10.1007/s00439-003-0986-5. [DOI] [PubMed] [Google Scholar]
- 26.Li X, Scaringe WA, Hill KA, Roberts S, Mengos A, Careri D, Pinto MT, Kasper CK, Sommer SS. Frequency of recent retrotransposition events in the human factor IX gene. Human Mutation. 2001;17:511–519. doi: 10.1002/humu.1134. [DOI] [PubMed] [Google Scholar]
- 27.Mukherjee S, Mukhopadhyay A, Banerjee D, Chandak GR, Ray K. Molecular pathology of haemophilia B: identification of five novel mutations including a LINE 1 insertion in Indian patients. Haemophilia. 2004;10:259–263. doi: 10.1111/j.1365-2516.2004.00895.x. [DOI] [PubMed] [Google Scholar]
- 28.Vidaud D, Vidaud M, Bahnak BR, Siguret V, Sanchez S Gispert, Laurian Y, Meyer D, Goossens M, Lavergne JM. Haemophilia B due to a de novo insertion of a human-specific Alu subfamily member within the coding region of the factor IX gene. European journal of human genetics : EJHG. 1993;1:30–36. doi: 10.1159/000472385. [DOI] [PubMed] [Google Scholar]
- 29.Wulff K, Gazda H, SchroÍ^der W, Robicka-Milewska R, Herrmann FH. Identification of a novel large F9 gene mutation-an insertion of an Alu repeated DNA element in exon e of the factor 9 gene. Human mutation. 2000;15:299. doi: 10.1002/(SICI)1098-1004(200003)15:3<299::AID-HUMU23>3.0.CO;2-1. [DOI] [PubMed] [Google Scholar]
- 30.Kolomietz E, Meyn MS, Pandita A, Squire JA. The role of Alu repeat clusters as mediators of recurrent chromosomal aberrations in tumors. Genes Chromosomes Cancer. 2002;35:97–112. doi: 10.1002/gcc.10111. [DOI] [PubMed] [Google Scholar]
- 31.Nakaya SM, Hsu TC, Geraghty SJ, Manco-Johnson MJ, Thompson AR. Severe hemophilia A due to a 1.3 kb factor VIII gene deletion including exon 24: homologous recombination between 41 bp within an Alu repeat sequence in introns 23 and 24. J Thromb Haemost. 2004;2:1941–1945. doi: 10.1111/j.1538-7836.2004.00963.x. [DOI] [PubMed] [Google Scholar]
- 32.Rossetti LC, Goodeve A, Larripa IB, De Brasi CD. Homeologous recombination between AluSx-sequences as a cause of hemophilia. Hum Mutat. 2004;24:440. doi: 10.1002/humu.9288. [DOI] [PubMed] [Google Scholar]
- 33.Vidal F, Farssac E, Tusell J, Puig L, Gallardo D. First molecular characterization of an unequal homologous alu-mediated recombination event responsible for hemophilia. Thromb Haemost. 2002;88:12–16. [PubMed] [Google Scholar]
- 34.Mitchell M, Dai L, Savidge G, Alhaq A. An Alu-mediated 31.5-kb deletion as the cause of factor XI deficiency in 2 unrelated patients. Blood. 2004;104:2394–2396. doi: 10.1182/blood-2004-04-1318. [DOI] [PubMed] [Google Scholar]
- 35.Nicholls RD, Fischel-Ghodsian N, Higgs DR. Recombination at the human alpha-globin gene cluster: sequence features and topological constraints. Cell. 1987;49:369–378. doi: 10.1016/0092-8674(87)90289-3. [DOI] [PubMed] [Google Scholar]
- 36.Harteveld KL, Losekoot M, Fodde R, Giordano PC, Bernini LF. The involvement of Alu repeats in recombination events at the alpha-globin gene cluster: characterization of two alphazero-thalassaemia deletion breakpoints. Hum Genet. 1997;99:528–534. doi: 10.1007/s004390050401. [DOI] [PubMed] [Google Scholar]
- 37.Hattori Y, Okayama N, Ohba Y, Yamashiro Y, Yamamoto K, Tsukimoto I, Kohakura M. The precise breakpoints of a Filipino-type alpha-thalassemia-1 deletion found in two Japanese. Hemoglobin. 1999;23:239–248. doi: 10.3109/03630269909005704. [DOI] [PubMed] [Google Scholar]
- 38.Henthorn PS, Smithies O, Mager DL. Molecular analysis of deletions in the human beta-globin gene cluster: deletion junctions and locations of breakpoints. Genomics. 1990;6:226–237. doi: 10.1016/0888-7543(90)90561-8. [DOI] [PubMed] [Google Scholar]
- 39.Craig JE, Kelly SJ, Barnetson R, Thein SL. Molecular characterization of a novel 10.3 kb deletion causing beta-thalassaemia with unusually high Hb A2. Br J Haematol. 1992;82:735–744. doi: 10.1111/j.1365-2141.1992.tb06952.x. [DOI] [PubMed] [Google Scholar]
- 40.Dimovski AJ, Baysal E, Efremov DG, Prior JF, Raven JL, Efremov GD, Huisman TH. A large beta-thalassemia deletion in a family of Indonesian-Malay descent. Hemoglobin. 1996;20:377–392. doi: 10.3109/03630269609005842. [DOI] [PubMed] [Google Scholar]
- 41.Game L, Bergounioux J, Close JP, Marzouka BE, Thein SL. A novel deletion causing (epsilon gamma delta beta) degrees thalassaemia in a Chilean family. Br J Haematol. 2003;123:154–159. doi: 10.1046/j.1365-2141.2003.04564.x. [DOI] [PubMed] [Google Scholar]
- 42.Gilman JG. The 12.6 kilobase DNA deletion in Dutch beta zero-thalassaemia. Br J Haematol. 1987;67:369–372. doi: 10.1111/j.1365-2141.1987.tb02360.x. [DOI] [PubMed] [Google Scholar]
- 43.Gilman JG, Brinson EC, Mishima N. The 32.6 kb Indian delta beta-thalassaemia deletion ends in a 3.4 kb L1 element downstream of the beta-globin gene. Br J Haematol. 1992;82:417–421. doi: 10.1111/j.1365-2141.1992.tb06439.x. [DOI] [PubMed] [Google Scholar]
- 44.Mishima N, Landman H, Huisman TH, Gilman JG. The DNA deletion in an Indian delta beta-thalassaemia begins one kilobase from the A gamma globin gene and ends in an L1 repetitive sequence. Br J Haematol. 1989;73:375–379. doi: 10.1111/j.1365-2141.1989.tb07756.x. [DOI] [PubMed] [Google Scholar]
- 45.Oner R, Oner C, Erdem G, Balkan H, Ozdag H, Erkan M, Gumruk F, Gurgey A, Altay C. A novel (delta beta)(0)-thalassemia due to a approximately 30-kb deletion observed in a Turkish family. Acta Haematol. 1996;96:232–236. doi: 10.1159/000203790. [DOI] [PubMed] [Google Scholar]
- 46.Rooks H, Bergounioux J, Game L, Close JP, Osborne C, Best S, Senior T, Height S, Thompson R, Hadzic N, Fraser P, Bolton-Maggs P, Thein SL. Heterogeneity of the epsilon gamma delta beta-thalassaemias: characterization of three novel English deletions. Br J Haematol. 2005;128:722–729. doi: 10.1111/j.1365-2141.2005.05368.x. [DOI] [PubMed] [Google Scholar]
- 47.Thong MK, Rudzki Z, Hall J, Tan JA, Chan LL, Yap SF. A single, large deletion accounts for all the beta-globin gene mutations in twenty families from Sabah (North Borneo), Malaysia. Mutation in brief no. 240. Online. Hum Mutat. 1999;13:413. doi: 10.1002/(sici)1098-1004(1999)13:5<413::aid-humu14>3.0.co;2-h. [DOI] [PubMed] [Google Scholar]
- 48.Yamashiro Y, Hattori Y, Okayama N, Shinoda E, Suyama N, Tanaka T, Ohi S. A novel (g)gamma(a)gamma(deltabeta)O-thalassemia with a 27 kb deletion. Hemoglobin. 2005;29:197–208. doi: 10.1081/hem-200066317. [DOI] [PubMed] [Google Scholar]
- 49.Zhang JW, Stamatoyannopoulos G, Anagnou NP. Laotian (delta beta) (0)-thalassemia: molecular characterization of a novel deletion associated with increased production of fetal hemoglobin. Blood. 1988;72:983–988. [PubMed] [Google Scholar]
- 50.Zhang XQ, Zhang JW. The 3′ breakpoint of the yunnanese (Agammadeltabeta)0-thalassemia deletion lies in an L1 family sequence: implications for the mechanism of deletion and the reactivation of the Ggamma-globin gene. Hum Genet. 1998;103:90–95. doi: 10.1007/s004390050789. [DOI] [PubMed] [Google Scholar]
- 51.Mustajoki S, Ahola H, Mustajoki P, Kauppinen R. Insertion of Alu element responsible for acute intermittent porphyria. Hum Mutat. 1999;13:431–438. doi: 10.1002/(SICI)1098-1004(1999)13:6<431::AID-HUMU2>3.0.CO;2-Y. [DOI] [PubMed] [Google Scholar]
- 52.Divoky V, Indrak K, Mrug M, Brabec V, Huisman T, Prchal J. A novel mechanism of beta thalassemia: the insertion of L1 retrotransposable element into beta globin IVS II. Blood. 1996;88 [Google Scholar]
- 53.Hassoun H, Coetzer TL, Vassiliadis JN, Sahr KE, Maalouf GJ, Saad ST, Catanzariti L, Palek J. A novel mobile element inserted in the alpha spectrin gene: spectrin dayton. A truncated alpha spectrin associated with hereditary elliptocytosis. J Clin Invest. 1994;94:643–648. doi: 10.1172/JCI117380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Barbot W, Dupressoir A, Lazar V, Heidmann T. Epigenetic regulation of an IAP retrotransposon in the aging mouse: progressive demethylation and de-silencing of the element by its repetitive induction. Nucleic Acids Res. 2002;30:2365–2373. doi: 10.1093/nar/30.11.2365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Bollati V, Baccarelli A, Hou L, Bonzini M, Fustinoni S, Cavallo D, Byun HM, Jiang J, Marinelli B, Pesatori AC, Bertazzi PA, Yang AS. Changes in DNA methylation patterns in subjects exposed to low-dose benzene. Cancer Res. 2007;67:876–880. doi: 10.1158/0008-5472.CAN-06-2995. [DOI] [PubMed] [Google Scholar]
- 56.Kale SP, Moore L, Deininger PL, Roy-Engel AM. Heavy metals stimulate human LINE-1 retrotransposition. Int J Environ Res Public Health. 2005;2:14–23. doi: 10.3390/ijerph2005010014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hagan CR, Rudin CM. Mobile genetic element activation and genotoxic cancer therapy: potential clinical implications. Am J Pharmacogenomics. 2002;2:25–35. doi: 10.2165/00129785-200202010-00003. [DOI] [PubMed] [Google Scholar]
- 58.Hagan CR, Sheffield RF, Rudin CM. Human Alu element retrotransposition induced by genotoxic stress. Nat Genet. 2003;35:219–220. doi: 10.1038/ng1259. [DOI] [PubMed] [Google Scholar]
- 59.Paz N, Levanon EY, Amariglio N, Heimberger AB, Ram Z, Constantini S, Barbash ZS, Adamsky K, Safran M, Hirschberg A, Krupsky M, Ben-Dov I, Cazacu S, Mikkelsen T, Brodie C, Eisenberg E, Rechavi G. Altered adenosine-to-inosine RNA editing in human cancer. Genome Res. 2007;17:1586–1595. doi: 10.1101/gr.6493107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Gaudet F, Rideout WM, 3rd, Meissner A, Dausman J, Leonhardt H, Jaenisch R. Dnmt1 expression in pre- and postimplantation embryogenesis and the maintenance of IAP silencing. Mol Cell Biol. 2004;24:1640–1648. doi: 10.1128/MCB.24.4.1640-1648.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sharif J, Muto M, Takebayashi S, Suetake I, Iwamatsu A, Endo TA, Shinga J, Mizutani-Koseki Y, Toyoda T, Okamura K, Tajima S, Mitsuya K, Okano M, Koseki H. The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature. 2007;450:908–912. doi: 10.1038/nature06397. [DOI] [PubMed] [Google Scholar]
- 62.Liang G, Chan MF, Tomigahara Y, Tsai YC, Gonzales FA, Li E, Laird PW, Jones PA. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol Cell Biol. 2002;22:480–491. doi: 10.1128/MCB.22.2.480-491.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Gaudet F, Hodgson JG, Eden A, Jackson-Grusby L, Dausman J, Gray JW, Leonhardt H, Jaenisch R. Induction of tumors in mice by genomic hypomethylation. Science. 2003;300:489–492. doi: 10.1126/science.1083558. [DOI] [PubMed] [Google Scholar]
- 64.Zhu H, Geiman TM, Xi S, Jiang Q, Schmidtmann A, Chen T, Li E, Muegge K. Lsh is involved in de novo methylation of DNA. EMBO J. 2006;25:335–345. doi: 10.1038/sj.emboj.7600925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Huang J, Fan T, Yan Q, Zhu H, Fox S, Issaq HJ, Best L, Gangi L, Munroe D, Muegge K. Lsh, an epigenetic guardian of repetitive elements. Nucleic Acids Res. 2004;32:5019–5028. doi: 10.1093/nar/gkh821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Geiman TM, Tessarollo L, Anver MR, Kopp JB, Ward JM, Muegge K. Lsh, a SNF2 family member, is required for normal murine development. Biochim Biophys Acta. 2001;1526:211–220. doi: 10.1016/s0304-4165(01)00129-5. [DOI] [PubMed] [Google Scholar]
- 67.Dennis K, Fan T, Geiman T, Yan Q, Muegge K. Lsh, a member of the SNF2 family, is required for genome-wide methylation. Genes Dev. 2001;15:2940–2944. doi: 10.1101/gad.929101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Fan T, Schmidtmann A, Xi S, Briones V, Zhu H, Suh HC, Gooya J, Keller JR, Xu H, Roayaei J, Anver M, Ruscetti S, Muegge K. DNA hypomethylation caused by Lsh deletion promotes erythroleukemia development. Epigenetics. 2008;3:134–142. doi: 10.4161/epi.3.3.6252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Wahlfors J, Hiltunen H, Heinonen K, Hamalainen E, Alhonen L, Janne J. Genomic hypomethylation in human chronic lymphocytic leukemia. Blood. 1992;80:2074–2080. [PubMed] [Google Scholar]
- 70.Dante R, Dante-Paire J, Rigal D, Roizes G. Methylation patterns of long interspersed repeated DNA and alphoid repetitive DNA from human cell lines and tumors. Anticancer Res. 1992;12:559–563. [PubMed] [Google Scholar]
- 71.Bollati V, Fabris S, Pegoraro V, Ronchetti D, Mosca L, Deliliers GL, Motta V, Bertazzi PA, Baccarelli A, Neri A. Differential repetitive DNA methylation in Multiple Myeloma molecular subgroups. Carcinogenesis. 2009 doi: 10.1093/carcin/bgp149. [DOI] [PubMed] [Google Scholar]
- 72.Roman-Gomez J, Jimenez-Velasco A, Agirre X, Cervantes F, Sanchez J, Garate L, Barrios M, Castillejo JA, Navarro G, Colomer D, Prosper F, Heiniger A, Torres A. Promoter hypomethylation of the LINE-1 retrotransposable elements activates sense/antisense transcription and marks the progression of chronic myeloid leukemia. Oncogene. 2005;24:7213–7223. doi: 10.1038/sj.onc.1208866. [DOI] [PubMed] [Google Scholar]
- 73.Roman-Gomez J, Jimenez-Velasco A, Agirre X, Castillejo JA, Navarro G, Jose-Eneriz E San, Garate L, Cordeu L, Cervantes F, Prosper F, Heiniger A, Torres A. Repetitive DNA hypomethylation in the advanced phase of chronic myeloid leukemia. Leuk Res. 2008;32:487–490. doi: 10.1016/j.leukres.2007.07.021. [DOI] [PubMed] [Google Scholar]
- 74.Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358:1148–1159. doi: 10.1056/NEJMra072067. [DOI] [PubMed] [Google Scholar]
- 75.Griffiths EA, Gore SD. DNA methyltransferase and histone deacetylase inhibitors in the treatment of myelodysplastic syndromes. Semin Hematol. 2008;45:23–30. doi: 10.1053/j.seminhematol.2007.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Han JS, Boeke JD. LINE-1 retrotransposons: modulators of quantity and quality of mammalian gene expression? Bioessays. 2005;27:775–784. doi: 10.1002/bies.20257. [DOI] [PubMed] [Google Scholar]
- 77.Burns KH, Boeke JD. Great exaptations. J Biol. 2008;7:5. doi: 10.1186/jbiol66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Wang J, Song L, Grover D, Azrak S, Batzer MA, Liang P. dbRIP: a highly integrated database of retrotransposon insertion polymorphisms in humans. Hum Mutat. 2006;27:323–329. doi: 10.1002/humu.20307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wheelan SJ, Scheifele LZ, Martinez-Murillo F, Irizarry RA, Boeke JD. Eukaryotic Transposable Elements and Genome Evolution Special Feature: Transposon insertion site profiling chip (TIP-chip) Proc Natl Acad Sci U S A. 2006;103:17632–17637. doi: 10.1073/pnas.0605450103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Konkel MK, Wang J, Liang P, Batzer MA. Identification and characterization of novel polymorphic LINE-1 insertions through comparison of two human genome sequence assemblies. Gene. 2007;390:28–38. doi: 10.1016/j.gene.2006.07.040. [DOI] [PubMed] [Google Scholar]
- 81.Akagi K, Li J, Stephens RM, Volfovsky N, Symer DE. Extensive variation between inbred mouse strains due to endogenous L1 retrotransposition. Genome Res. 2008;18:869–880. doi: 10.1101/gr.075770.107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Amit M, Sela N, Keren H, Melamed Z, Muler I, Shomron N, Izraeli S, Ast G. Biased exonization of transposed elements in duplicated genes: A lesson from the TIF-IA gene. BMC Molecular Biology. 2007;8 doi: 10.1186/1471-2199-8-109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Symer DE, Connelly C, Szak ST, Caputo EM, Cost GJ, Parmigiani G, Boeke JD. Human l1 retrotransposition is associated with genetic instability in vivo. Cell. 2002;110:327–338. doi: 10.1016/s0092-8674(02)00839-5. [DOI] [PubMed] [Google Scholar]
- 84.Weinstock DM, Richardson CA, Elliott B, Jasin M. Modeling oncogenic translocations: Distinct roles for double-strand break repair pathways in translocation formation in mammalian cells. DNA Repair. 2006;5:1065–1074. doi: 10.1016/j.dnarep.2006.05.028. [DOI] [PubMed] [Google Scholar]
- 85.Gu W, Zhang F, Lupski JR. Mechanisms for human genomic rearrangements. Pathogenetics. 2008;1:4. doi: 10.1186/1755-8417-1-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Megonigal MD, Rappaport EF, Jones DH, Kim CS, Nowell PC, Lange BJ, Felix CA. Panhandle PCR strategy to amplify MLL genomic breakpoints in treatment-related leukemias. Proc Natl Acad Sci U S A. 1997;94:11583–11588. doi: 10.1073/pnas.94.21.11583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Super HG, Strissel PL, Sobulo OM, Burian D, Reshmi SC, Roe B, Zeleznik-Le NJ, Diaz MO, Rowley JD. Identification of complex genomic breakpoint junctions in the t(9;11) MLL-AF9 fusion gene in acute leukemia. Genes Chromosomes Cancer. 1997;20:185–195. [PubMed] [Google Scholar]
- 88.Schichman SA, Caligiuri MA, Strout MP, Carter SL, Gu Y, Canaani E, Bloomfield CD, Croce CM. ALL-1 tandem duplication in acute myeloid leukemia with a normal karyotype involves homologous recombination between Alu elements. Cancer Research. 1994;54:4277–4280. [PubMed] [Google Scholar]
- 89.So CW, Ma ZG, Price CM, Dong S, Chen SJ, Gu LJ, So CKC, Wiedemann LM, Chan LC. MLL self fusion mediated by Alu repeat homologous recombination and prognosis of AML-M4/M5 subtypes. Cancer Research. 1997;57:117–122. [PubMed] [Google Scholar]
- 90.Strout MP, Marcucci G, Bloomfield CD, Caligiuri MA. The partial tandem duplication of ALL1 (MLL) is consistently generated by Alu-mediated homologous recombination in acute myeloid leukemia. Proc Natl Acad Sci U S A. 1998;95:2390–2395. doi: 10.1073/pnas.95.5.2390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Schnittger S, WoÍ^rmann B, Hiddemann W, Griesinger F. Partial tandem duplications of the MLL gene are detectable in peripheral blood and bone marrow of nearly all healthy donors. Blood. 1998;92:1728–1734. [PubMed] [Google Scholar]
- 92.O’Neil J, Tchinda J, Gutierrez A, Moreau L, Maser RS, Wong KK, Li W, McKenna K, Liu XS, Feng B, Neuberg D, Silverman L, DeAngelo DJ, Kutok JL, Rothstein R, DePinho RA, Chin L, Lee C, Look AT. Alu elements mediate MYB gene tandem duplication in human T-ALL. Journal of Experimental Medicine. 2007;204:3059–3066. doi: 10.1084/jem.20071637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Wiemels JL, Hofmann J, Kang M, Selzer R, Green R, Zhou M, Zhong S, Zhang L, Smith MT, Marsit C, Loh M, Buffler P, Yeh RF. Chromosome 12p deletions in TEL-AML1 childhood acute lymphoblastic leukemia are associated with retrotransposon elements and occur postnatally. Cancer Res. 2008;68:9935–9944. doi: 10.1158/0008-5472.CAN-08-2139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Miki Y, Nishisho I, Horii A, Miyoshi Y, Utsunomiya J, Kinzler KW, Vogelstein B, Nakamura Y. Disruption of the APC gene by a retrotransposal insertion of L1 sequence in a colon cancer. Cancer Res. 1992;52:643–645. [PubMed] [Google Scholar]
- 95.Morse B, Rotherg PG, South VJ, Spandorfer JM, Astrin SM. Insertional mutagenesis of the myc locus by a LINE-1 sequence in a human breast carcinoma. Nature. 1988;333:87–90. doi: 10.1038/333087a0. [DOI] [PubMed] [Google Scholar]
- 96.Economou-Pachnis A, Tsichlis PN. Insertion of an Alu SINE in the human homologue of the Mlvi-2 locus. Nucleic Acids Res. 1985;13:8379–8387. doi: 10.1093/nar/13.23.8379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Tsichlis PN, Strauss PG, Kozak CA. Cellular DNA region involved in induction of thymic lymphomas (Mlvi-2) maps to mouse chromosome 15. Mol Cell Biol. 1984;4:997–1000. doi: 10.1128/mcb.4.5.997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Kirilyuk A, Tolstonog GV, Damert A, Held U, Hahn S, Lower R, Buschmann C, Horn AV, Traub P, Schumann GG. Functional endogenous LINE-1 retrotransposons are expressed and mobilized in rat chloroleukemia cells. Nucleic Acids Res. 2008;36:648–665. doi: 10.1093/nar/gkm1045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Servomaa K, Rytomaa T. Suicidal death of rat chloroleukaemia cells by activation of the long interspersed repetitive DNA element (L1Rn) Cell Tissue Kinet. 1988;21:33–43. doi: 10.1111/j.1365-2184.1988.tb00769.x. [DOI] [PubMed] [Google Scholar]
- 100.Howard G, Eiges R, Gaudet F, Jaenisch R, Eden A. Activation and transposition of endogenous retroviral elements in hypomethylation induced tumors in mice. Oncogene. 2008;27:404–408. doi: 10.1038/sj.onc.1210631. [DOI] [PubMed] [Google Scholar]
- 101.Prak ET, Kazazian HH., Jr. Mobile elements and the human genome. Nat Rev Genet. 2000;1:134–144. doi: 10.1038/35038572. [DOI] [PubMed] [Google Scholar]
- 102.Beauregard A, Curcio MJ, Belfort M. The Take and Give Between Retrotransposable Elements and their Hosts. Annual Review of Genetics. 2008;42:587. doi: 10.1146/annurev.genet.42.110807.091549. [DOI] [PMC free article] [PubMed] [Google Scholar]