Abstract
Rationale
We previously discovered several phosphorylations to the β subunit of the mitochondrial F1Fo ATP synthase complex in isolated rabbit myocytes upon adenosine treatment, an agent that induces cardioprotection. The role of these phosphorylations is unknown.
Objective
The current study focuses on the functional consequences of phosphorylation of the ATP synthase complex β subunit by generating non-phosphorylatable and phospho-mimetic analogs in a model system, S. cerevisiae.
Methods and Results
The four amino acid residues with homology in yeast (T58, S213, T262 and T318) were studied with respect to growth, complex and supercomplex formation, and enzymatic activity (ATPase rate). The most striking mutant was the T262 site, for which the phospho-mimetic (T262E) abolished activity, while the non-phosphorylatable strain (T262A) had an ATPase rate equivalent to wild-type. Although T262E, like all of the β subunit mutants, was able to form the intact complex (F1Fo), this strain lacked a free F1 component found in wild-type and had a corresponding increase of lower molecular weight forms of the protein, indicating an assembly/stability defect. In addition, the ATPase activity was reduced but not abolished with the phospho-mimetic mutation at T58, a site that altered the formation/maintenance of dimers of the F1Fo ATP synthase complex.
Conclusions
Taken together, these data show that pseudo-phosphorylation of specific amino acid residues can have separate and distinctive effects on the F1Fo ATP synthase complex, suggesting the possibility that several of the phosphorylations observed in the rabbit heart can have structural and functional consequences to the F1Fo ATP synthase complex.
Keywords: Mitochondria, ATP synthase, phosphorylation, preconditioning
INTRODUCTION
Preconditioning (PC) is a phenomenon by which physiological and pharmacological interventions protect the heart from damage during future ischemic episodes (for review see1). Mitochondria have long been implicated in this protective phenotype.2,3 In studying the link between PC and the mitochondria, our group observed phosphorylation of the F1Fo ATP synthase complex β subunit (ATPβ) in response to adenosine mediated PC.4 There have been several observations connecting modulation of the ATP synthase complex to PC, including its specific down-regulation to preserve ATP pools during ischemia.5–7 The goal of the current work is to gain insight into the functional aspects of phosphorylation of ATPβ by mutation of the amino acid residues in a model system.
It is becoming increasingly clear that mitochondria participate in control by kinase cascades and protein phosphorylation (for reviews see8–11). Several groups have shown that the F1Fo ATP synthase complex can be phosphorylated on subunits α,12–14 β,12,14,15 δ,16 ε,12 γ,12,14,17 4,12 OSCP,12,14 c,14 and g,12 in a broad range of species. Phosphorylation has been correlated to dimerization of the ATP synthase complex in yeast (subunit g)12 and in heart (γ subunit)17, but there is currently no evidence that phosphorylation of ATPβ regulates any aspect of the complex. Regulation of this complex could come at a variety of points: transcription, translation, import, assembly or direct functional regulation. The F1Fo ATP synthase is a well-conserved enzyme complex that is known to exist in several different assemblies, including both monomeric and dimeric forms.17–19 Its subunits have a high degree of amino acid sequence homology and have similar assembly, structure and catalytic activity from E. coli to mammals.18 Eukaryotic F1Fo ATP synthase is more sophisticated in both the number of subunits and in the specific chaperones required for assembly.20 It has been suggested that this increased complexity could provide more regulated steps in the production of the enzyme20 and phosphorylation adds yet another level of regulation.
The current study focuses on defining the individual role of each phosphorylated amino acid residue of ATPβ discovered in PC. To accomplish this, the genetically tractable model system of Saccharomyces cerevisiae was chosen so that each phosphorylation could be studied independently and in the context of a complete knock-out of the endogenous protein. The amino acid residues were mutated to non-phosphorylatable (alanine) or phospho-mimetic (aspartic acid and glutamic acid) residues. The acidic residue mutations incorporate a charged residue to mimic phosphorylation. The alanine mutations act as a control mutation by ensuring backbone spacing is retained, without the possibility of phosphorylation. The model system is an essential tool since 100% of ATPβ are modified at the same site and all ATP synthase complexes contain modified subunit. In vivo mammalian analysis would be complicated by the fact that the regulation of the ATP synthase complex could involve either a single modified subunit in each complex or multiple modifications to different sites on each of the three ATPβ in a given complex. This study expands on our cardiac proteomic findings using a model system to analyze phosphorylations independently of the complications of controlling phosphorylation and dephosphorylation in mammalian systems. The mutant strains were analyzed with respect to structure and function as compared to wild-type (WT) and an ATPβ deletion strain (atp2Δ). The data show that phosphorylation of ATPβ at unique amino acid residues could act as important regulators of complex function (ATPase rate) and structure (F1 and dimer formation), as several phospho-mimetic mutants had dramatic effects.
MATERIALS AND METHODS
Media and Genetic Methods
Yeast media included YEPD (YEP with 2% dextrose), SD (synthetic medium containing 2% dextrose), YEPD+EtBr (YEPD with 25μg/nl ethidium bromide) and SRaf (synthetic medium with 2% raffinose); yeast genetic techniques are as previously described.21 Strains. The yeast strains used in this study are listed in Table 1. See online supplement.
Table 1.
Phosphorylated amino acid residues identified in rabbit heart ATP synthase β subunit and the relationship to the yeast protein and the phospho-mimetic mutants used in this study. Acidic amino acids (A and E/D) were substituted to mimic phosphorylations and alanine residues were substituted as control mutations.
| Phosphorylated amino acid residue in rabbit (pre-protein) | Corresponding amino acid in yeast (mature protein) | Percent identity (of the tryptic peptide surrounding the residue) | Non-phosphorylatable mutant strain | Phospho-mimetic mutant strain |
|---|---|---|---|---|
| S106&T107 | T58 | 93% | T58A | T58E |
| T262/S263 | S213 | 79% | S213A | S213D |
| T312 | T262 | 95% | T262A | T262E |
| T368 | T318 | 100% | T318A | T318E |
Mitochondria and ATP synthase Isolation
Crude mitochondria were isolated from yeast homogenates22 and purified using sucrose gradients.22,23 ATP synthase was isolated using sucrose gradient centrifugation (see online supplement).24
Blue Native PAGE and 2D BN/SDS-PAGE gels
Blue Native PAGE (BN-PAGE) were used to resolve the native, intact mitochondrial protein complexes.25 Two-dimensional BN/SDS-PAGE was also performed to analyze complex subunit composition.25 See online supplement for details.
1D SDS-PAGE
1D electrophoresis, 4–12% NuPAGE Bis-Tris gels (1mm, Invitrogen), were run according to manufacturer’s protocols, using LDS sample buffer and MES running buffer and stained with silver26 or colloidal Coomassie.27
Western blotting
Gels were transferred and blotted as described in the online supplement.
ATPase Assays
In-solution ATPase assays were performed on sucrose gradient isolated complex.28 In-gel ATPase analyses were performed following BN-PAGE.19 See online supplement.
MS/MS
Protein bands from 1D BN-PAGE and 2D BN/SDS-PAGE were cut and prepared for digestion.29 See online supplement for data acquisition and protein identification details.
3D structure
The α3β3 hexamer of the S. Cerevisiae ATP synthase from Protein Data Bank (http://www.pdb.org) 2HLD structure30 was modeled using DeepView/Swiss-PdbViewer v3.7.
RESULTS
Phospho-mimetic mutations
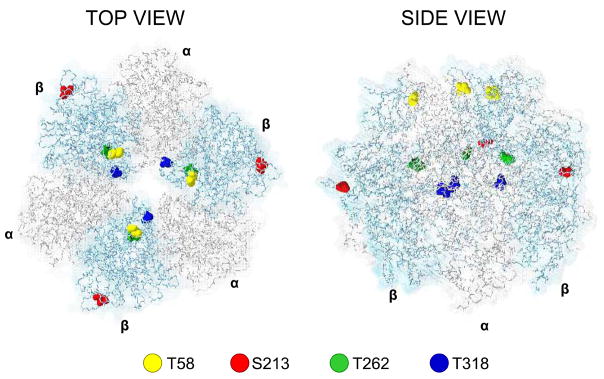
Four of the five phosphorylated residues observed in the rabbit heart protein are conserved in the yeast protein (See alignment, Online Figure I). The residues present were mutated to phospho-mimetic and non-phosphorylatable amino acids as shown in Table 1. Two residues were located on the matrix-facing surface of the subunit (T58 and S213), and two were located in the center of the complex (T262 and T318) (Figure 1). The two internal sites are located in close proximity, so double mutations were made in an attempt to determine if simultaneous phosphorylation would result in a different phenotype than the individual mutations.
Figure 1. The amino acid residues of interest mapped onto the 3D structure of the α/β hexamer of S. Cerevisiae (PDB file 2HLD)30.
Amino acid residues are color-coded, with numbers given for the mature protein (with known mitochondrial targeting sequence removed). Two of the residues (T58, yellow and S213, red) are located on the matrix-facing portion of the β subunit while the other two (T262, green and T318, blue) are located within the center of the complex. Both buried (T262) and accessible (T58) residues had observed assembly and functional differences.
Effect on growth
The effects of the mutations were assessed by examining the growth of all strains at 16, 30 and 37°C on either SD-His or YEPD+EtBr (ethidium bromide) media. These types of media were chosen to examine the growth of the mutant strains in both a selected, uninhibited manner (SD-His) and in the absence of mtDNA (YEPD+EtBr). All strains grew equivalent to WT at 30°C and 37°C, regardless of the media (data not shown). However, at 16°C the T262E strain displayed reduced growth compared on SD-His (Online Figure IIA) and did not form colonies on YEPD+EtBr media where there was no mtDNA (and thus no Fo), whereas the T262A grew comparably to WT in both instances (Online Figure IIB). This growth phenotype was mirrored by the double mutants T262A/T318A (WT-like growth) and T262E/T318E (T262E-like growth) (Online Figure IIC).
Phospho-mimetic mutants exhibit a differential arrangement of ATP synthase complex by BN-PAGE
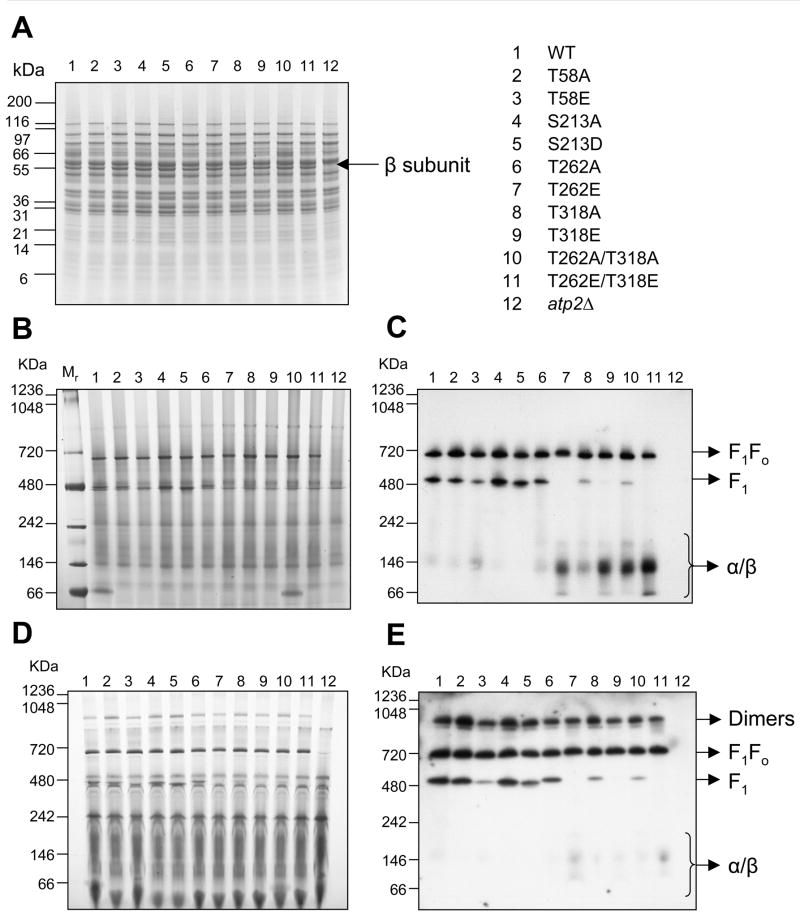
The ATP synthase assemblies present in each mutant strain, as compared to both WT and the ATPβ deletion strain (atp2Δ), were analyzed by 1D BN-PAGE. Importantly, all of the mutant strains had ATPβ protein levels equivalent to WT, as observed by SDS-PAGE of mitochondria (Figure 2A). Since detergents are known to preserve complexes to different extents31, mitochondria from all strains were solubilized in either lauryl maltoside (LM, Figure 2B, 2C) or digitonin (DIG, Figure 2D, 2E). LM disrupts protein interactions to a greater degree than DIG, which is capable of preserving more structures. Figures 2B and 2D show equal loading of gels by total protein stain (Coomassie) and Figures 2C and 2E show western blots probed with anti-ATPβ antibody (α subunit blots gave the same pattern, data not shown). Both LM-and DIG-solubilized mitochondria contain an ATPβ band at approximately 700kDa. This is identified as the intact F1Fo complex, based on apparent mass, previous observations32–35 and MS data (Figures 2C and 2E, Table 2). The F1Fo band is present at equal amounts (based on densitometry, n=3 each, Online Table II) in all mutant strains solubilized with LM or DIG. The only exception being the LM-solubilized T262E/T318E, which is reduced by 10–30% (p=0.03). Regardless of the detergent used, ATPβ was also present in a complex at 480kDa. This corresponds to the F1 portion of the complex (Table 2) which is known to assemble independently of the Fo portion.32–35 The quantity of the F1 band is markedly different between WT and several phospho-mutants in both detergent conditions (based on densitometry, n=3 each, Online Table II). In particular, T58E has much less of the F1 band than either WT or T58A and the strains T318A and T262A/T318A also have much lower levels than WT. Three of the phospho-mimetic strains (T262E, T318E and T262E/T318E) display an absence of this F1 band in both detergent conditions. Interestingly, the strains lacking the F1 band display lower molecular weight bands of ATPβ (Figure 2C and E) and α subunit (data not shown). DIG solubilization allowed for the observation of a higher molecular weight form (~1000kDa) of the ATP synthase complex which likely represent F1Fo dimers (Figure 2D). Some of the mutant strains have differences in the amount of dimer as compared to WT (22±4.9) (Online Table II). Most interestingly, T58A had a greater quantity of dimer (56±10, p<0.02) while the T58E had a WT level of dimer (27±5.1). Taken together, the data in Figure 2 imply that, though all the mutant strains have WT levels of the ATPβ and of the intact F1Fo complex, the strains differ greatly in the smaller and larger observed assemblies.
Figure 2. Blue Native PAGE (BN-PAGE) of isolated WT and mutant mitochondria.
A) 1D SDS-PAGE (4–20%), denaturing/reducing gel. The total protein stain shows equal protein loading and β subunit content of all lanes (except atp2Δ, which lacks the β subunit). Total protein stain of BN-PAGE gels shows equal protein content in all lanes for lauryl maltoside (B) and digitonin (D) solubilized mitochondria. (C) and (E) display representative western blots of the ATP synthase β subunit (n=3) for lauryl maltoside and digitonin solubilized mitochondria, respectively. Bands are labeled according to the complex that is present; F1Fo dimers, F1Fo monomers, the F1 portion of the complex and α/β lower molecular weight bands (For intensity data values see Online Table II).
Table 2.
Number of unique peptides observed by MS/MS for each subunit identified in bands from WT BN-PAGE digests with both trypsin and chymotrypsin.
| Subunit | Dimer band | F1Fo band | F1 band | ||
|---|---|---|---|---|---|
| Digitonin | Lauryl Maltoside | Digitonin | Lauryl Maltoside | Digitonin | |
| α | 8 | 30 | 23 | 40 | 17 |
| β | 9 | 30 | 18 | 25 | 18 |
| ε | 7 | 6 | 6 | 3 | |
| γ | 3 | 13 | 7 | 15 | 5 |
| d | 2 | 7 | 2 | ||
| OSCP | 6 | 12 | 5 | ||
| 4 | 2 | 9 | 3 | ||
| f | 2 | ||||
| j | 2 | ||||
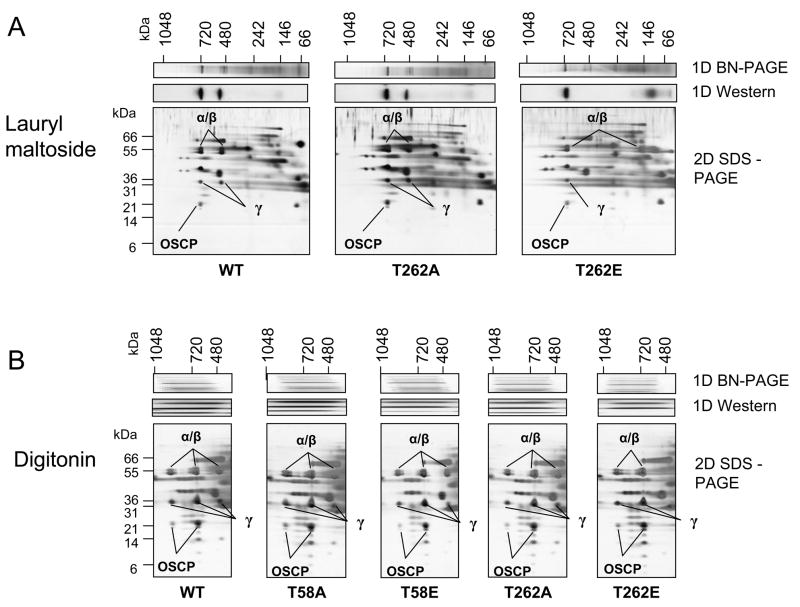
Using 2D analysis (2D BN/SDS-PAGE) in which the complexes are resolved first by BN-PAGE and then separated into subunits by reducing and denaturing SDS-PAGE, we were able to resolve the individual protein components of the 1D BN-PAGE bands (Figure 3). Figure 3A (LM solubilization) shows that the F1Fo (700kDa) band of the WT, T262A and T262E strains is made up of at least α, β, γ and OSCP subunits from both of the F1 and Fo components (based on MS, Online Figure III and Online Table III). The suspected F1 band (480kDa) in the WT and T262A strains is missing the Fo component OSCP subunit (Figure 3A). For the T262E strain, in which there is no detectable F1 band by BN-PAGE, no α, β, or γ subunits are resolved in this region. However, the T262E strain gel contains additional α and β subunit spots in the lower portion of the BN-PAGE. 2D BN/SDS-PAGE of DIG-solubilized mitochondria confirmed the subunit pattern observed in the F1Fo and F1 bands of LM-solubilized mitochondria (Figure 3B). DIG 2D gels also confirmed the presence of F1 subunits α, β, γ and the Fo subunit OSCP in the dimer area (based on MS, Online Figure III and Online Table III).
Figure 3. Two-dimensional BN/SDS-PAGE of isolated WT and mutant mitochondria.
A) Lauryl maltoside solubilized mitochondria from WT, T262A and T262E yeast strains and B) digitonin solubilized mitochondria from WT, T58A, T58E, T262A and T262E were subjected to BN-PAGE in the first dimension and SDS-PAGE in a second dimension (4–12% gels). This technique allows for the separation of complexes observed on 1D BN-PAGE into individual subunits. Circled protein gel spots were identified by MS/MS as indicated. These 2D BN/SDS-PAGE gels confirm the presence of the Fo and/or F1 subunits at the correct location in the BN-PAGE. (For MS identification data see Online Table III).
To increase the coverage of the complex subunits, MS analysis was performed directly on each 1D BN-PAGE of interest from WT mitochondria using trypsin and chymotrypsin independently. Several subunits of both the F1 and the Fo portions of the complex were observed (Table 2). F1 subunits were observed in all bands, but Fo subunits were only observed in the bands suspected to contain the full F1Fo monomer and dimers (Table 2). For MS/MS peptide identification data see Online Tables IV and V for LM and DIG, respectively.
Several phospho-mimetic mutants have decreased ATPase function
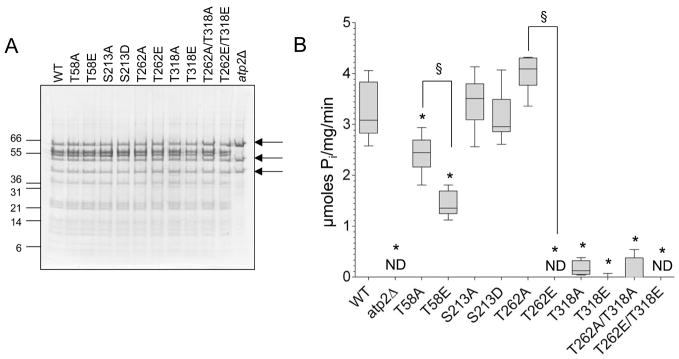
Functional assessment of the phospho-mimetic and non-phosphorylatable mutations was performed on complexes isolated from mitochondria by sucrose gradient centrifugation.24 M mitochondria isolated from the atp2Δ strain were used as a negative control. The in-solution ATPase rate for the complex with the mutants at residue S213 (S213A 3.44±0.52, S213E 3.14±0.51 μmoles Pi/mg/min) did not differ from WT (3.28±0.56 μmoles Pi/mg/min), but all other mutations caused changes with respect to WT (Figure 4B). Although the A and E mutants of the residue T58 both had reduced ATPase rates, the T58A strain had significantly better function (2.42±0.38 μmoles Pi/mg/min) than the T58E phospho-mimetic strain (1.43±0.25 μmoles Pi/mg/min, p<0.01 between T58A and T58E). Both mutations at T318 reduced ATPase rates compared to WT (T318A 0.17±0.15 μmoles Pi/mg/min, T318E no detectable activity). The T262A strain trended toward an increase in ATP hydrolysis (4.01±0.36 μmoles Pi/mg/min, p=0.054) compared to WT, whereas the phospho-mimetic (T262E) had no detectable activity. It is of note that the double mutant of T262E/T318E (0.14±0.27 μmoles Pi/mg/min) is even more impaired in its ATPase rate than the T318E mutant alone (no detectable activity).
Figure 4. In-solution ATPase assays.
A) Mitochondria were solubilized in lauryl maltoside and the complexes were separated on a sucrose gradient to isolate the ATP synthase complex. A representative gel of the combined ATP synthase fractions from each of the strains is shown. Arrows indicate the contaminating bands also present in the atp2Δ lanes. ATPase assays were performed on this deletion strain as a negative control. B) In-solution ATPase assays were performed on fractions from each of the yeast mutant strains and were compared to WT and the negative control (atp2Δ). ND = no detectable signal, *represents significant difference from WT, and §represents significant difference between the non-phosphorylatable and the phospho-mimetic strains (both based on Mann-Whitney test p<0.05). n=6 all observed values fell within the linear range of the assay.
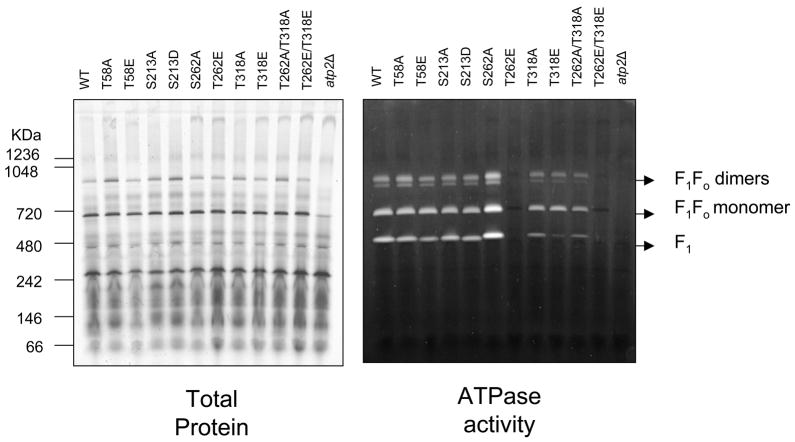
Since different quantities of the free F1 and the dimer forms of the complex were observed by BN-PAGE in some mutant strains, in-gel ATPase assays were also performed to ensure that the in-solution ATPase data were affected by these different complexes. DIG-solubilized mitochondria were used because they allowed for the observation of all ATP synthase bands (Figure 5). Although equal quantities of F1Fo ATP synthase monomer are present in all strains (except the atp2Δ control), the activity stain shows increased intrinsic activity in the T262A monomer compared to WT and no observable activity in the T262E monomer (Figure 5). This pattern was also observed in the dimer bands of the WT, T262A and T262E strains. The in-gel activity of the F1, F1Fo monomer and F1Fo dimer bands in Figure 5 show the same functional patterns as the in-solution assays, indicating that the level of intrinsic activity is driven by the ATPβ protein, not its assembled state. This is also confirmed by the fact that though T262E and T318E have similar BN-PAGE band patterns, T318E retains activity in all observed bands, whereas T262E has no activity.
Figure 5. In-gel ATPase assays.
In-gel ATPase assays were performed on BN-PAGE of digitonin solubilized mitochondria from all strains. Left panel is a representative Coomassie stained gel to indicate protein load. The right panel illustrates the ATPase activity of each of the bands as white lead phosphate precipitate on the gel. These assays mirror the results observed in Figure 4 for in-solution ATPase assays (n=3).
DISCUSSION
The current work explores the ATPβ phosphorylations, originally observed in a rabbit heart, in the model system S. Cerevisiae. Using this model system, we have shown that mutations mimicking phosphorylation of specific residues of ATPβ can have unique effects on the structure and function of the complex. The phospho-mimetic of the T262 residue blocks the ATPase function of the complex, whereas mutations at the T58 residue primarily affect dimer formation. Because the T262 residue is buried in the interior of the intact complex, it may be inaccessible to dynamic regulation (although it could be phosphorylated prior to or during complex formation). ResidueT58, which is on the surface of the complex, could be modulated in the intact ATP synthase complex in a faster time scale.
Growth of phospho-mimetic mutants
Though each strain grew well at 30°C on all media, the added stress of growth at 16°C revealed a cold-sensitive phenotype for the T262E strain. At this temperature, T262E did not grow well on SD-His and this deficiency was even more pronounced on YEPD+EtBr. As growth of yeast on media containing EtBr results in the loss of mitochondrial DNA (mtDNA),36 the phenotype of the T262E mutant on this medium implies a role of mtDNA in maintaining the viability of the T262E strain. Yeast mtDNA encodes three subunits of the Fo portion of the complex (a, c and 8).18 The F1 portion of the complex is essential to viability in the absence of the Fo section,37 possibly due to hydrolysis of ATP into ADP by the F1 component, allowing the ADP/ATP translocase to maintain mitochondrial membrane potential18,37. The inability of the T262E mutant to grow only in the absence of mtDNA at 16°C indicates that this phosphorylation most likely interferes in some aspect of the F1 component assembly or function that can be stabilized by the Fo portion of the complex.
F1Fo ATP synthase complex monomer assembly
The F1 and Fo components of the ATP synthase complex form independently in the matrix and the inner membrane.18 Different complex assemblies are observed on BN-PAGE gels depending on the type of gel used and the protein/detergent ratio.31,35,38,39 In this study, LM-solubilized WT mitochondria had two prominent ATP synthase bands: the F1Fo complex monomer (~700kDa) and the F1 portion alone (~480kDa). Solubilization with DIG allowed for the additional observation of the F1Fo complex dimer. It is possible that the independent F1 portion may be a product of the detergent extraction31 and not relevant in vivo, but the stark differences between the phospho-mimetic strains and the consistency between both LM- and DIG-solubilization give insight into the subunit interactions affected by these phosphorylations (Figure 6).
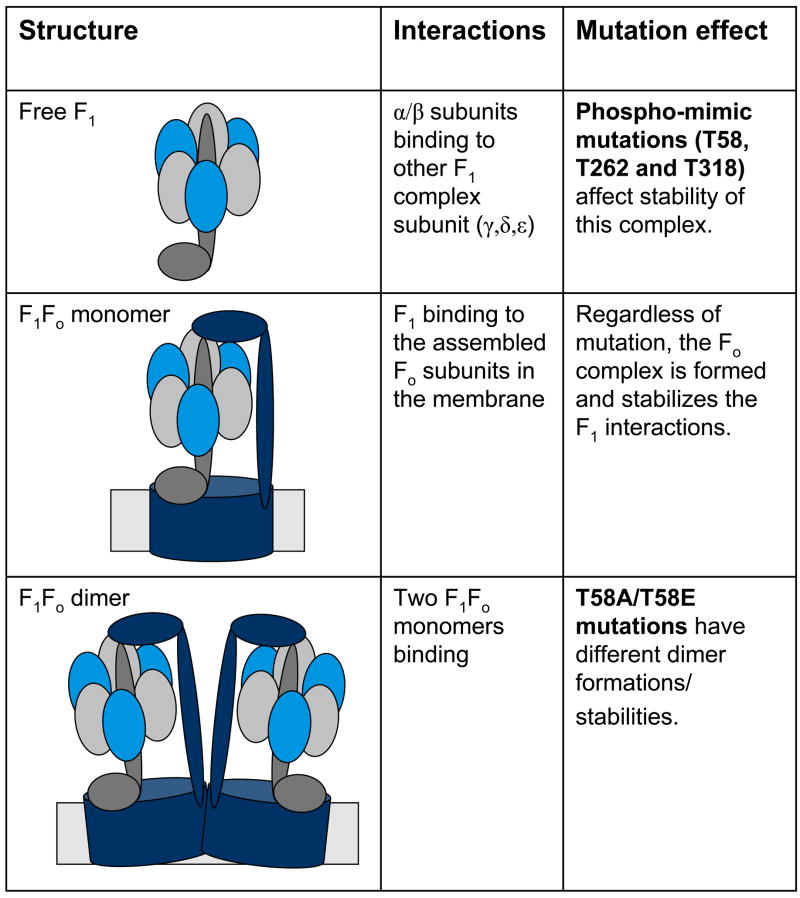
Figure 6. Schema of ATP synthase structures and the effects of the phospho-mimetic mutations.
The various structures observed by BN-PAGE are listed and the subunit interactions that occur within them. Mutants that caused defects at any stage are listed.
The F1Fo monomer is present at equal quantities in all strains, implying that intact complex monomer can assemble and is stable as the holoenzyme (Figure 2C and 2E). However, there is a striking difference in the amount of free F1 complex between WT and some mutant strains. The phospho-mimetic mutants T262E, T318E and T262E/T318E have reduced quantities of the F1 complex (Figure 2C and 2E). It is probable that the F1 portion of the complex is unstable in these phospho-mimetic strains and is either labile under the detergent extraction conditions or cannot form unless it is assembled with the Fo component. In other words, the Fo component of the complex is capable of stabilizing the F1 when the complex is fully assembled. This hypothesis is consistent with the stunted growth phenotype of the T262E mutants described above on EtBr containing media, where a functional F1 is essential in the absence of the Fo.
T262 is located on the interior of the F1 portion of ATP synthase complex (Figure 1). There are no published data to suggest that the T262 residue has either inter- or intra-subunit interactions that would explain this apparent structural interference, suggesting unknown biophysical aspects of this amino acid residue. All of the other sites that caused a decrease in the quantity of the F1 component compared to WT have known interactions. For example, T318, is in close proximity to sites known to be involved in hydrogen bonding between the α and β subunits in E.coli.40 The T58 residue is also located within a region involved in subunit interactions, the beta-barrel domain.41,42 Though T318 and T58 have less F1 by BN-PAGE and known subunit interactions, they did not show defects in growth assays, indicating that any functional effect of this F1 destabilization in the accessible T58 or buried T318 mutants is incomplete.
F1Fo ATP synthase complex dimer assembly
The other difference on the BN-PAGE gels is in dimer formation between the T58A and T58E strains (Figure 2E). Formation of the ATP synthase complex into dimers and oligomers has been observed in mammalian cells and yeast and can affect cristae formation,35 improve efficiency of the enzyme43 and even affect mitochondrial membrane potential44 (for review see45). Phosphorylation of ATP synthase complex subunits, g subunit in yeast12 and γ subunit in bovine17, have been correlated to the formation of dimers. The difference in dimer formation between the T58A and T58E strains implies an additional role for phosphorylation of ATPβ in the regulation of dimer formation or maintenance.
F1Fo ATP synthase complex function
The amino acid residues of ATPβ involved in the binding of nucleotide/phosphate and catalysis have been defined in yeast 30 None of the residues in this study are known to be a part of the catalytic site, yet it is clear that the substitution of the phospho-mimetic mutations can affect function (Figures 4 and 5), probably due to conformational or other allosteric changes. Functional consequences of ATPβ phosphorylation were examined by two ATPase assay methods and both yielded similar results. Two of the amino acid residues (S213 and T318) had no significant differences in ATPase rate between the phospho-mimetic and the non-phosphorylatable mutations. Both of the mutations of residue T318 displayed no ATPase activity, implying its importance to overall function. The double mutation of the internal sites mimicked the ablation of function observed for T318 alone and thus could not provide any insight into cooperation or inhibition of these two internally-located phosphorylations. Two of the phosphorylated amino acid residues exhibited a large difference between A and E forms, indicative of potential regulation by phosphorylation in vivo. There was a two-fold difference in ATPase rate between the T58A and T58E strains. As discussed above, the T58 amino acid residue is located within a beta-barrel domain that has structural interactions with other F1 subunits.41,42 The interactions of this domain could be disrupted by the T58E mutation (or by phosphorylation), causing an inefficient complex and producing the lower ATPase rate.
The ATPase rate of T262A trended toward being higher than WT, while the T262E strain had essentially no activity, indicating that this residue could be an important regulatory site. T262 is buried within the center of the F1 portion of the complex (Figure 1) and as such, phosphorylation would have to occur prior to assembly or by some auto-phosphorylation. This residue is located near three residues known to interact with oxygen atoms of phosphate during catalysis (βAsp256, βAsn257 and βArg260).30 Although there is no indication in the literature that T262 is important for catalysis, this study clearly indicates that modifications here have significant implications for the function of the F1Fo complex. An in vivo phosphorylation of the T262 equivalent residue in rabbit hearts would likely result in decreased function of this complex, which has been shown to occur in PC.
Implications of ATPβ phosphorylation
Though the S. cerevisiae provided an excellent model system in which to study the phosphorylation of ATPβ, there are a few important caveats to this study when considering it in the context of the mammalian heart. Most importantly, the phospho-mimetics in this study are constitutively present, mimicking universal phosphorylation at a single site in each ATPβ mutant strain. This is likely not representative of the in vivo cardiac situation, as the phosphorylated forms represent a small portion of the total ATPβ in the rabbit heart. Also, it is possible that the phosphorylations exert regulation at only discrete steps in import, assembly or function of the F1Fo complex. Since the phospho-mimetic mutations are constitutively present, it is impossible to clarify these steps and some consequences of in vivo phosphorylation may be missed. The other main difference between the model system approach and in vivo phosphorylation is that all of the ATPβ in the yeast mutants contain the same modified site. As such, the ATPβ composition in all complexes is homogenous rather than a combination of unphosphorylated or heterogeneous phosphorylation on each ATPβ in the complex. This single-site analysis is beneficial as it allows the direct assignment of function to a particular phospho-mimetic residue. However, in vivo it is possible that regulation would involve different stoichiometry at any site on each of the complex’s three β subunits and this differential residue phosphorylation could act to produce a unique phenotype. Even so, the observations of this study provide significant evidence that phosphorylation of ATPβ can have important implications to the structure and function of the F1Fo ATP synthase complex.
Here we have shown that, in a model system, mutations of ATPβ mimicking the phosphorylations observed in rabbit heart can impact both the structure and function of the F1Fo ATP synthase complex. Specifically, the T58E mutant affected dimer formation and decreased ATPase function, and the T262E mutant ablated ATPase function. The matrix-facing, accessible nature of the T58 site makes it a likely candidate for rapid regulation of the ATP synthase complex into dimers or a mild phosphorylation-dependent decrease in function. The T262 residue is buried in the center of the complex, making it likely that it is phosphorylated prior to assembly and may be involved in longer-term regulation of the complex. This would imply that in the original PC model, some portion of the ATPβ is being imported and assembled in the 60-minute timeframe. Though there is no data on the ATPβ protein specifically, measurable amounts other cardiac mitochondrial proteins can be imported, in vitro, in as quickly as three minutes46. The data from this model system analysis suggest that an increase in phosphorylation of ATPβ during PC would result in functional changes to ATP synthase complex. The kinase and phosphatase involved in the regulation of this phenomenon are still unknown and must be uncovered in order to gain a full understanding of the consequences of ATPβ phosphorylation.
NOVELTY AND SIGNIFICANCE
What is known?
Cardiac preconditioning stimuli can affect mitochondrial function and specifically the F1Fo ATP synthase enzyme complex.
Phosphorylation can occur on several subunits of the mitochondrial F1Fo ATP synthase, including five specific sites on the β subunit found in pharmacological preconditioning.
What new information does this article contribute?
In a model system, phospho-mimetic mutations of T262 ablated function, indicating that in vivo phosphorylation could result in down-modulation in ATP activity.
Phospho-mimetic mutations to the T58 site cause changes in the function and the maintenance of complex dimers, which play a large role in overall mitochondrial shape and function.
This study is an important step forward in our understanding of the post-translational regulation of the mitochondrial F1Fo ATP synthase complex. The study moved a novel proteomic discovery that the β subunit was phosphorylated to a system that allowed the functional effect of each modification to be assessed. It was shown that two of the phosphorylation sites (mimicked by pseudo-phosphorylation mutants) affect structure and function of ATP synthase. This study is the first to connect site-specific phosphorylation of an F1Fo ATP synthase subunit with modulation of the holoenzyme. This opens several new questions regarding the kinases involved in phosphorylation and the dynamic nature of these phosphorylations in vivo. Understanding the mechanisms of these phosphorylations provides a new context for mitochondrial involvement in PC.
Supplementary Material
Acknowledgments
SOURCES OF FUNDING
This work was supported by the NHLBI Proteomic Initiative (JVE: contract NO-HV-28120), the NIH (JVE: contract P01HL081427), an American Heart Association Pre-doctoral Fellowship (LK: 0715247U) and a NIH Pre-doctoral training grant (MY: 2T32-GM07445).
The authors thank the Technical Implementation and Coordination Core of The Johns Hopkins NHLBI Proteomics Center for the LTQ MS/MS analysis, Steven Elliott for 4800 MALDI TOF/TOF analysis and Cory Dunn for creation of the atp2Δ strain. We also thank Shandev Rai for editorial comments and suggestions.
NON-STANDARD ABBREVIATIONS AND ACRONYMS
- PC
preconditioning
- ATPβ
ATP synthase β subunit
- EtBr
Ethidium bromide
- BN-PAGE
Blue native polyacrylamide gel electrophoresis
- SDS-PAGE
sodium dodecyl sulfate-polyacrylamide gel electrophoresis
- LM
lauryl maltoside
- DIG
digitonin
- MS
mass spectrometry
Footnotes
DISCLOSURES
None
References
- 1.Liem DA, Honda HM, Zhang J, Woo D, Ping P. Past and present course of cardioprotection against ischemia-reperfusion injury. J Appl Physiol. 2007;103:2129–36. doi: 10.1152/japplphysiol.00383.2007. [DOI] [PubMed] [Google Scholar]
- 2.Murphy E, Steenbergen C. Preconditioning: the mitochondrial connection. Annu Rev Physiol. 2007;69:51–67. doi: 10.1146/annurev.physiol.69.031905.163645. [DOI] [PubMed] [Google Scholar]
- 3.Halestrap AP, Clarke SJ, Khaliulin I. The role of mitochondria in protection of the heart by preconditioning. Biochim Biophys Acta. 2007;1767:1007–31. doi: 10.1016/j.bbabio.2007.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Arrell DK, Elliott ST, Kane LA, Guo Y, Ko YH, Pedersen PL, Robinson J, Murata M, Murphy AM, Marban E, Van Eyk JE. Proteomic analysis of pharmacological preconditioning: novel protein targets converge to mitochondrial metabolism pathways. Circ Res. 2006;99:706–14. doi: 10.1161/01.RES.0000243995.74395.f8. [DOI] [PubMed] [Google Scholar]
- 5.Ala-Rami A, Ylitalo KV, Hassinen IE. Ischaemic preconditioning and a mitochondrial KATP channel opener both produce cardioprotection accompanied by F1F0-ATPase inhibition in early ischaemia. Basic Res Cardiol. 2003;98:250–8. doi: 10.1007/s00395-003-0413-z. [DOI] [PubMed] [Google Scholar]
- 6.Ylitalo K, Ala-Rami A, Vuorinen K, Peuhkurinen K, Lepojarvi M, Kaukoranta P, Kiviluoma K, Hassinen I. Reversible ischemic inhibition of F(1)F(0)-ATPase in rat and human myocardium. Biochim Biophys Acta. 2001;1504:329–39. doi: 10.1016/s0005-2728(00)00261-9. [DOI] [PubMed] [Google Scholar]
- 7.Penna C, Pagliaro P, Rastaldo R, Di Pancrazio F, Lippe G, Gattullo D, Mancardi D, Samaja M, Losano G, Mavelli I. F0F1 ATP synthase activity is differently modulated by coronary reactive hyperemia before and after ischemic preconditioning in the goat. Am J Physiol Heart Circ Physiol. 2004;287:H2192–200. doi: 10.1152/ajpheart.00327.2004. [DOI] [PubMed] [Google Scholar]
- 8.Huttemann M, Lee I, Samavati L, Yu H, Doan JW. Regulation of mitochondrial oxidative phosphorylation through cell signaling. Biochim Biophys Acta. 2007;1773:1701–20. doi: 10.1016/j.bbamcr.2007.10.001. [DOI] [PubMed] [Google Scholar]
- 9.Gibson BW. The human mitochondrial proteome: oxidative stress, protein modifications and oxidative phosphorylation. Int J Biochem Cell Biol. 2005;37:927–34. doi: 10.1016/j.biocel.2004.11.013. [DOI] [PubMed] [Google Scholar]
- 10.Horbinski C, Chu CT. Kinase signaling cascades in the mitochondrion: a matter of life or death. Free Radic Biol Med. 2005;38:2–11. doi: 10.1016/j.freeradbiomed.2004.09.030. [DOI] [PubMed] [Google Scholar]
- 11.Pagliarini DJ, Dixon JE. Mitochondrial modulation: reversible phosphorylation takes center stage? Trends Biochem Sci. 2006;31:26–34. doi: 10.1016/j.tibs.2005.11.005. [DOI] [PubMed] [Google Scholar]
- 12.Reinders J, Wagner K, Zahedi RP, Stojanovski D, Eyrich B, van der Laan M, Rehling P, Sickmann A, Pfanner N, Meisinger C. Profiling phosphoproteins of yeast mitochondria reveals a role of phosphorylation in assembly of the ATP synthase. Mol Cell Proteomics. 2007;6:1896–906. doi: 10.1074/mcp.M700098-MCP200. [DOI] [PubMed] [Google Scholar]
- 13.Vosseller K, Hansen KC, Chalkley RJ, Trinidad JC, Wells L, Hart GW, Burlingame AL. Quantitative analysis of both protein expression and serine/threonine post-translational modifications through stable isotope labeling with dithiothreitol. Proteomics. 2005;5:388–98. doi: 10.1002/pmic.200401066. [DOI] [PubMed] [Google Scholar]
- 14.Hopper RK, Carroll S, Aponte AM, Johnson DT, French S, Shen RF, Witzmann FA, Harris RA, Balaban RS. Mitochondrial matrix phosphoproteome: effect of extra mitochondrial calcium. Biochemistry. 2006;45:2524–36. doi: 10.1021/bi052475e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hojlund K, Wrzesinski K, Larsen PM, Fey SJ, Roepstorff P, Handberg A, Dela F, Vinten J, McCormack JG, Reynet C, Beck-Nielsen H. Proteome analysis reveals phosphorylation of ATP synthase beta -subunit in human skeletal muscle and proteins with potential roles in type 2 diabetes. J Biol Chem. 2003;278:10436–42. doi: 10.1074/jbc.M212881200. [DOI] [PubMed] [Google Scholar]
- 16.Ko YH, Pan W, Inoue C, Pedersen PL. Signal transduction to mitochondrial ATP synthase: evidence that PDGF-dependent phosphorylation of the delta-subunit occurs in several cell lines, involves tyrosine, and is modulated by lysophosphatidic acid. Mitochondrion. 2002;1:339–48. doi: 10.1016/s1567-7249(01)00036-8. [DOI] [PubMed] [Google Scholar]
- 17.Di Pancrazio F, Bisetto E, Alverdi V, Mavelli I, Esposito G, Lippe G. Differential steady-state tyrosine phosphorylation of two oligomeric forms of mitochondrial F0F1ATPsynthase: a structural proteomic analysis. Proteomics. 2006;6:921–6. doi: 10.1002/pmic.200500077. [DOI] [PubMed] [Google Scholar]
- 18.Ackerman SH, Tzagoloff A. Function, structure, and biogenesis of mitochondrial ATP synthase. Prog Nucleic Acid Res Mol Biol. 2005;80:95–133. doi: 10.1016/S0079-6603(05)80003-0. [DOI] [PubMed] [Google Scholar]
- 19.Bisetto E, Di Pancrazio F, Simula MP, Mavelli I, Lippe G. Mammalian ATPsynthase monomer versus dimer profiled by blue native PAGE and activity stain. Electrophoresis. 2007;28:3178–85. doi: 10.1002/elps.200700066. [DOI] [PubMed] [Google Scholar]
- 20.Mueller DM. Partial assembly of the yeast mitochondrial ATP synthase. J Bioenerg Biomembr. 2000;32:391–400. doi: 10.1023/a:1005532104617. [DOI] [PubMed] [Google Scholar]
- 21.Adams A, Gottschling DE, Kaiser CA, Stearns T. Methods in Yeast Genetics. Plainview, NY: Cold Spring Harbour Laboratory Press; 1997. [Google Scholar]
- 22.Daum G, Gasser SM, Schatz G. Import of proteins into mitochondria. Energy-dependent, two-step processing of the intermembrane space enzyme cytochrome b2 by isolated yeast mitochondria. J Biol Chem. 1982;257:13075–80. [PubMed] [Google Scholar]
- 23.Meisinger C, Sommer T, Pfanner N. Purification of Saccharomcyes cerevisiae mitochondria devoid of microsomal and cytosolic contaminations. Anal Biochem. 2000;287:339–42. doi: 10.1006/abio.2000.4868. [DOI] [PubMed] [Google Scholar]
- 24.Hanson BJ, Schulenberg B, Patton WF, Capaldi RA. A novel subfractionation approach for mitochondrial proteins: a three-dimensional mitochondrial proteome map. Electrophoresis. 2001;22:950–9. doi: 10.1002/1522-2683()22:5<950::AID-ELPS950>3.0.CO;2-D. [DOI] [PubMed] [Google Scholar]
- 25.Schagger H, von Jagow G. Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal Biochem. 1991;199:223–31. doi: 10.1016/0003-2697(91)90094-a. [DOI] [PubMed] [Google Scholar]
- 26.Shevchenko A, Wilm M, Vorm O, Mann M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal Chem. 1996;68:850–8. doi: 10.1021/ac950914h. [DOI] [PubMed] [Google Scholar]
- 27.Candiano G, Bruschi M, Musante L, Santucci L, Ghiggeri GM, Carnemolla B, Orecchia P, Zardi L, Righetti PG. Blue silver: a very sensitive colloidal Coomassie G-250 staining for proteome analysis. Electrophoresis. 2004;25:1327–33. doi: 10.1002/elps.200305844. [DOI] [PubMed] [Google Scholar]
- 28.Tzagoloff A. Oligomycin-sensitive ATPase of Saccharomyces cerevisiae. Methods Enzymol. 1979;55:351–8. doi: 10.1016/0076-6879(79)55044-7. [DOI] [PubMed] [Google Scholar]
- 29.Kane LA, Yung CK, Agnetti G, Neverova I, Van Eyk JE. Optimization of paper bridge loading for 2-DE analysis in the basic pH region: application to the mitochondrial subproteome. Proteomics. 2006;6:5683–7. doi: 10.1002/pmic.200600267. [DOI] [PubMed] [Google Scholar]
- 30.Kabaleeswaran V, Puri N, Walker JE, Leslie AG, Mueller DM. Novel features of the rotary catalytic mechanism revealed in the structure of yeast F1 ATPase. Embo J. 2006;25:5433–42. doi: 10.1038/sj.emboj.7601410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Grandier-Vazeille X, Guerin M. Separation by blue native and colorless native polyacrylamide gel electrophoresis of the oxidative phosphorylation complexes of yeast mitochondria solubilized by different detergents: specific staining of the different complexes. Anal Biochem. 1996;242:248–54. doi: 10.1006/abio.1996.0460. [DOI] [PubMed] [Google Scholar]
- 32.Lemaire C, Dujardin G. Preparation of respiratory chain complexes from Saccharomyces cerevisiae wild-type and mutant mitochondria: activity measurement and subunit composition analysis. Methods Mol Biol. 2008;432:65–81. doi: 10.1007/978-1-59745-028-7_5. [DOI] [PubMed] [Google Scholar]
- 33.Sabar M, Balk J, Leaver CJ. Histochemical staining and quantification of plant mitochondrial respiratory chain complexes using blue-native polyacrylamide gel electrophoresis. Plant J. 2005;44:893–901. doi: 10.1111/j.1365-313X.2005.02577.x. [DOI] [PubMed] [Google Scholar]
- 34.Smith CP, Thorsness PE. Formation of an energized inner membrane in mitochondria with a gamma-deficient F1-ATPase. Eukaryot Cell. 2005;4:2078–86. doi: 10.1128/EC.4.12.2078-2086.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Paumard P, Vaillier J, Coulary B, Schaeffer J, Soubannier V, Mueller DM, Brethes D, di Rago JP, Velours J. The ATP synthase is involved in generating mitochondrial cristae morphology. Embo J. 2002;21:221–30. doi: 10.1093/emboj/21.3.221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Goldring ES, Grossman LI, Krupnick D, Cryer DR, Marmur J. The petite mutation in yeast. Loss of mitochondrial deoxyribonucleic acid during induction of petites with ethidium bromide. J Mol Biol. 1970;52:323–35. doi: 10.1016/0022-2836(70)90033-1. [DOI] [PubMed] [Google Scholar]
- 37.Chen XJ, Clark-Walker GD. Alpha and beta subunits of F1-ATPase are required for survival of petite mutants in Saccharomyces cerevisiae. Mol Gen Genet. 1999;262:898–908. doi: 10.1007/s004380051156. [DOI] [PubMed] [Google Scholar]
- 38.Giraud MF, Paumard P, Soubannier V, Vaillier J, Arselin G, Salin B, Schaeffer J, Brethes D, di Rago JP, Velours J. Is there a relationship between the supramolecular organization of the mitochondrial ATP synthase and the formation of cristae? Biochim Biophys Acta. 2002;1555:174–80. doi: 10.1016/s0005-2728(02)00274-8. [DOI] [PubMed] [Google Scholar]
- 39.Meyer B, Wittig I, Trifilieff E, Karas M, Schagger H. Identification of two proteins associated with mammalian ATP synthase. Mol Cell Proteomics. 2007;6:1690–9. doi: 10.1074/mcp.M700097-MCP200. [DOI] [PubMed] [Google Scholar]
- 40.Boltz KW, Frasch WD. Hydrogen bonds between the alpha and beta subunits of the F1-ATPase allow communication between the catalytic site and the interface of the beta catch loop and the gamma subunit. Biochemistry. 2006;45:11190–9. doi: 10.1021/bi052592+. [DOI] [PubMed] [Google Scholar]
- 41.Kagawa Y, Hamamoto T, Endo H. The alpha/beta Interfaces of alpha1beta1, alpha3beta3, and F1: Domain Motions and Elastic Energy Stored during gamma Rotation. J Bioenerg Biomembr. 2000;32:471–84. doi: 10.1023/a:1005612923995. [DOI] [PubMed] [Google Scholar]
- 42.Bakhtiari N, Lai-Zhang J, Yao B, Mueller DM. Structure/function of the beta-barrel domain of F1-ATPase in the yeast Saccharomyces cerevisiae. J Biol Chem. 1999;274:16363–9. doi: 10.1074/jbc.274.23.16363. [DOI] [PubMed] [Google Scholar]
- 43.Strauss M, Hofhaus G, Schroder RR, Kuhlbrandt W. Dimer ribbons of ATP synthase shape the inner mitochondrial membrane. Embo J. 2008;27:1154–60. doi: 10.1038/emboj.2008.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Bornhovd C, Vogel F, Neupert W, Reichert AS. Mitochondrial membrane potential is dependent on the oligomeric state of F1F0-ATP synthase supracomplexes. J Biol Chem. 2006;281:13990–8. doi: 10.1074/jbc.M512334200. [DOI] [PubMed] [Google Scholar]
- 45.Wittig I, Schagger H. Supramolecular organization of ATP synthase and respiratory chain in mitochondrial membranes. Biochim Biophys Acta. 2009 doi: 10.1016/j.bbabio.2008.12.016. [DOI] [PubMed] [Google Scholar]
- 46.Craig EE, Hood DA. Influence of aging on protein import into cardiac mitochondria. Am J Physiol. 1997;272:H2983–8. doi: 10.1152/ajpheart.1997.272.6.H2983. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.