Abstract
We report that knockdown of the α1 tubulin isoform Tuba1a, but not the highly related Tuba1b, dramatically impedes nervous system formation during development and RGC axon regeneration following optic nerve injury in adults. Within the tuba1a promoter, a G/C-rich element was identified that is necessary for tuba1a induction during RGC differentiation and optic axon regeneration. KLF6a and 7a, which we previously reported are essential for optic axon regeneration (Veldman et al., 2007), bind this G/C-rich element and transactivate the tuba1a promoter. In vivo knockdown of KLF6a and 7a attenuate regeneration-dependent activation of the endogenous tuba1a and p27 genes. These results suggest tuba1a expression is necessary for CNS development and regeneration and that KLF6a and 7a mediate their effects, at least in part, via transcriptional control of tuba1a promoter activity.
Keywords: retina, retinal ganglion cell, tubulin, KLF, optic nerve, p27, promoter
INTRODUCTION
The failure of the mammalian CNS to successfully mount a regenerative response to injury is a result of a non-permissive environment at the injury site (Benfey and Aguayo, 1982; Caroni and Schwab, 1988; Garcia-Valenzuela et al., 1994; Schwab 1996; Hermanns et al., 2001) and an intrinsic inability of the adult CNS to regenerate (Chen et al., 1995; Buffo et al., 1997; Takami et al., 2002; Zheng et al., 2003; Zheng et al, 2005). In contrast, axonal regeneration within the teleost CNS is very robust, in part because glial scarring is low and the extracellular environment is favorable to regrowing axons (Bastemeyer et al., 1991), but also due to a robust intrinsic regenerative response within the injured cell. Remarkably, teleost fish recover visual and motor functions following optic nerve or spinal cord damage, respectively (Stuermer et al., 1992; Zottoli and Freemer, 2003). Therefore, teleost fish provide an excellent model system for investigating the intrinsic mechanisms contributing to successful CNS regeneration.
The success of teleost CNS regeneration may be determined, in part, by the induction of regeneration-associated genes (RAGs) such as GAP43, α1 tubulin (α1T), the cell adhesion molecule L1, and the Na, K-ATPase alpha3 subunit (Benowitz et al., 1981; Hieber et al., 1992; Liu et al., 2002; Becker et al., 1998; Becker et al., 2005) and it has been argued that the inability to appropriately regulate these genes in the mammalian central nervous system contributes to its failed regenerative response (Plunet et al, 2002). We recently showed that Kruppel-like transcription factors KLF6a and 7a are RAGs that are necessary for optic axon regeneration in zebrafish (Veldman et al., 2007). However we also found other highly induced RAGs, such as SOCS3, that were not necessary for optic axon regeneration (Veldman et al., 2007). Therefore it is important to identify which of the many induced RAGs are essential for injury-induced axon regrowth. Once these essential genes are identified they will serve as probes for mechanisms underlying their induction which may suggest new strategies for improving repair of the damaged or diseased mammalian CNS.
With this goal in mind we have begun analyzing gene expression during regeneration using microarray approaches (Veldman et al., 2007). Among the most highly induced genes are those encoding α1 and β5 tubulin, which are constituents of microtubules. Microtubules play important roles during regeneration including growth cone formation (Erturk et al, 2007) and axonal extension (Witte and Bradke, 2008). At least 8 different alpha tubulin genes have been identified in mammals (Khodiyar et al., 2007). Alpha tubulin orthologs have been identified and along with phylogenetic analysis, subgroups have been determined. This analysis indicates 3 different mouse α1T gene isoforms, Tuba1a, Tuba1b and Tuba1c, belonging to group 1. Interestingly, these isoforms are all located on the same chromosome and in the same order when comparing human, mouse and zebrafish genomes. The Tuba1a gene (formerly referred to as Tα1 in rat, mα1 in mouse and B-ALPHA-1 in human) is best correlated with axon growth during development and peripheral axon regeneration in the adult (Miller et al., 1987; Miller, et al.,1989). Missense mutations in the mouse Tuba1a gene cause abnormal neuronal migration and in humans TUBA1A missense mutations have been correlated with lissencephaly (Keays et al., 2007).
We have characterized α1T gene expression in zebrafish during optic nerve regeneration (Goldman and Ding, 2000; Goldman et al., 2001; Senut et al., 2004). Like mammals, zebrafish α1 tubulins are encoded by 3 highly related genes that show synteny with their mammalian counterparts. Although these zebrafish genes are referred to as tuba1, tuba11 and tuba1l2 in Ensembl, we will follow zebrafish convention of naming genes based on their mammalian counterparts and refer to these genes as tuba1a, tuba1b and tuba1c. In humans, mice and zebrafish these genes are arranged similarly on a single chromosome (tuba1b, tuba1a, tuba1c). Using transgenic approaches we previously demonstrated that the goldfish tuba1a promoter is activated during neural development and in adult RGCs that are regenerating damaged optic axons (Goldman and Ding, 2000; Goldman et al., 2001; Senut et al., 2004).
Although tuba1a promoter activity is increased during optic nerve regeneration, only a few regulatory elements mediating this activation have been identified (Goldman and Ding, 2000; Senut et al., 2004). In addition, it is not known if tuba1a expression is necessary for optic nerve regeneration. Indeed it is possible that the highly related tuba1b or tuba1c gene can compensate for tuba1a loss. Here we report that tuba1a expression is not only necessary for RGC axonal regeneration but also for normal development of the CNS. We found that regeneration-dependent expression is mediated in part via a G/C-rich element residing in the tuba1a promoter. DNA binding and transactivation assays suggest that previously identified RAGs, KLF6a and 7a (Veldman et al., 2007), bind this element and enhance promoter activity. Importantly we show that KLF6a and 7a expression contributes to endogenous tuba1a and p27 induction during optic nerve regeneration. These studies link KLF6a and 7a expression to tuba1a expression and establish tuba1a as an essential α1 tubulin isoform whose expression is necessary for CNS development and regeneration.
RESULTS
Organization of tuba1 genes on chromosome 23
In mammals group 1 alpha tubulin genes include Tuba1a, Tuba1b and Tuba1c. These genes localize to chromosome 12 in humans and chromosome 15 in the mouse. In both species these genes exhibit a similar arrangement along the chromosome (Tuba1b, Tuba1a, Tuba1c) with Tuba1a and Tuba1b located near each other with a similar transcriptional orientation, while Tuba1c is located over 80kb away from them and situated in the opposite orientation. Inspection of the zebrafish genome suggests a similar arrangement of these genes on chromosome 23 (Fig. 1). However, unlike their mammalian counterparts, where Tuba1a and Tuba1b are separated by about 20–50kb, zebrafish tuba1a and tuba1b are only separated by about 3kb.
Figure 1. Organization of alpha 1 tubulin genes on chromosome 23.

Three alpha 1 tubulin isoforms, tuba1a, tuba1b and tuba1c are all found on chromosome 23. Distances separating the genes and their orientation are indicated. Each gene is comprised of 4 exons (black rectangles).
Tuba1a expression is necessary for optic axon regeneration
We previously demonstrated that the goldfish tuba1a promoter is induced in zebrafish retinal ganglion cells (RGCs) that are regenerating their damaged axons (Goldman and Ding, 2000; Goldman et al., 2001; Senut et al., 2004). However, these studies did not determine if the tuba1a gene product is necessary for optic axon regeneration. To investigate if Tuba1a is necessary for optic axon regeneration we combined morpholino-modified antisense oligonucleotide (MO)-mediated RGC gene expression knockdown with retinal explants (Veldman et al., 2007). For these experiments we lesioned the optic nerve and placed Gelfoam, containing a MO targeting the tuba1 or tuba1b mRNAs or a control MO, onto the optic nerve stump. Four days later, retinas were harvested, diced and cultured as explants for 4 days prior to analysis. One advantage of studying cultured retinal explants is that they allow quantification of RGC axon growth (Veldman et al., 2007). Retinal explants prepared from untreated and control MO-treated retinas displayed robust axon outgrowth after 4 days in culture, while knockdown of Tuba1a caused a dramatic suppression of optic axon regrowth (Fig. 2). This latter result appears to be specific since knockdown of Tuba1b had little effect on RGC axonogenesis (Fig. 2). Experiments were repeated with MOs targeting different sequences and similar results were obtained (Fig. 2). tuba1a-targeting MOs were found to suppress RGC axon regeneration in a concentration-dependent manner (Supplementary Fig. 1). In addition, MOs targeting 2 different tuba1a sequences at a concentration where each had little effect on RGC axon regeneration, when combined dramatically reduced RGC axon regrowth (Supplementary Fig. 1). TUNEL stain showed Tuba1a knockdown in RGCs is not accompanied by increased RGC apoptosis (Supplementary Fig. 2).
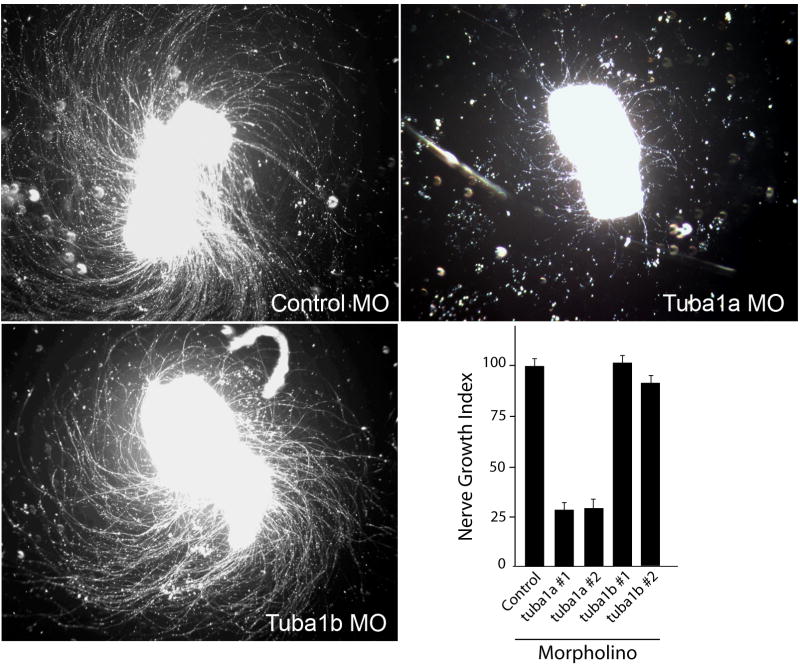
Figure 2. Knockdown of Tuba1a, but not Tuba1b, inhibits RGC axon regeneration in retinal explants.
Control, tuba1a or tuba1b-targeting morpholino-modified antisense oligonucleotides (MO) were delivered to RGCs in vivo by placing MO-soaked Gelfoam on the sectioned optic nerve stump. The next day the Gelfoam was removed and 3 days later retinas were isolated, diced and placed in explant culture for 4 days prior to analysis of neurite outgrowth. Shown are representative images of axon outgrowth. The lower right-hand graph shows Nerve Growth Index (see Materials and Methods) for control MO and 2 different MOs, #1 and #2, targeting different tuba1a and tuba1b sequences, respectively. Nerve Growth Index values were normalized to control MO treated samples that were set at 100%. Error bars are standard error of the mean. Note MOs targeting tuba1a, but not tuba1b, dramatically suppress RGC axon regeneration. Experiments were repeated at least 3 times with each MO.
We verified the effectiveness of the tuba1a- and tuba1b-targeting MOs using a GFP reporter that harbors the MO-target sequence appended upstream of the GFP initiator AUG. These reporters, along with MOs were injected into single cell zebrafish embryos and GFP expression assayed. Co-injection of each reporter plasmid with its cognate targeting MO caused a large decrease in the percentage of GFP+ embryos, while co-injection of each reporter plasmid with its non-cognate MO had little effect on reporter expression (Supplementary Table 1).
Tuba1a is necessary for CNS development and differentiation
The above experiments suggest Tuba1a is necessary for optic axon regeneration following injury. To determine if Tuba1a is also necessary for CNS development we compared Tuba1a and Tuba1b knockdown in developing embryos. Microinjection of control or experimental MOs into single cell zebrafish embryos revealed that Tuba1a expression is essential for normal development and, in particular, is necessary for formation of anterior structures such as the brain and retina (Fig. 3 and Table 1). tuba1a MO injected embryos generally do not survive beyond 40hpf. Varying the concentration of tuba1a MO (0 to 0.25mM) injected into embryos showed a dose response relationship where embryo death increased from ~7% to ~55% and mutant phenotypes increasing from ~3% to ~40% (Fig. 3). Based on these studies we chose 0.125mM tuba1a-targeting MO for further experiments. Quantification of embryo phenotypes following injection of either 0.25mM control MO, 0.125mM tuba1a- or 0.25mM tuba1b-targeting MOs showed that control and tuba1b-targeting MOs resulted in ~90% of the embryos developing normally, while injection of the tuba1a-targeting MO resulted in only ~10% developing normally (Table 1). Similar results were obtained with a second MO targeting different tuba1a and tuba1b sequences. Based on embryo morphology, it appears that tuba1a MO-treated embryos arrest development shortly after 20hpf and the majority of embryos die by 40hpf.
Figure 3. Knockdown of Tuba1a, but not Tuba1b, suppresses CNS formation in developing embryos.
Control, tuba1a or tuba1b-targeting morpholinos (MO) were injected into single cell zebrafish embryos. Top graphs show that tuba1a MO-dependent phenotypes at 30 hours post fertilization (hpf) are concentration dependent (tuba1a MO concentrations are 0, 0.065, 0.125 and 0.25mM; control MO was used to keep total MO concentrations at 0.25mM). Bottom images show representative pictures of MO-injected (0.125mM) embryo phenotypes at 30 and 52hpf. To inhibit apoptosis and examine MO specificity, some experiments included in vitro transcribed bcl2 mRNA (25pg) or tuba1a mRNA (12.5pg) that was engineered to escape MO-mediated knockdown. Note that tuba1a, but not tuba1b MO severely disrupted development, especially of anterior structures; however, inclusion of bcl2 or tuba1a mRNA diminished the severity of Tuba1a knockdown. Three different tuba1a mRNA rescue phenotypes are shown at 52hpf. Similar results were obtained with a second MO targeting different tuba1a and tuba1b sequences.
Table 1.
mRNA rescue of tuba1a MO injected embryos.
| Morpholino/mRNA | # Injected Embryos | % Dead | % Alive with Mutant Phenotype |
|---|---|---|---|
| control MO | 192 | 8 | 2 |
| tuba1a MO | 325 | 52 | 38 |
| tuba1b MO | 125 | 9 | 3 |
| tuba1a MO + tuba1a mRNA | 203 | 29 | 8 |
| tuba1a MO + bcl2 mRNA | 381 | 32 | 8 |
Single cell zebrafish embryos were injected with 0.125 mM tuba1a-targeting MO +/− 12.5pg of tuba1a mRNA or 25pg of bcl2 mRNA. Embryos were examined at 30hpf and scored as dead or alive and, of those living, the number of embryos that exhibited a typical tuba1a MO phenotype (See Fig. 3) were counted and percentages determined. Note that the tuba1a and bcl2 mRNAs significantly reduced both the number of embryos that died or that had a tuba1a MO phenotype.
Tuba1a knockdown embryos exhibited severe apoptosis which precluded analysis of CNS differentiation. Therefore we co-injected a prosurvival Bcl2 mRNA (Kratz et al., 2006) or the tuba1a mRNA (12.5pg) to rescue apoptotic and mutant cells. Bcl2 is known to antagonize p53-mediated cell death in zebrafish (Kratz et al., 2006; Chen et al., 2009). The tuba1a mRNA was engineered to escape tuba1a-targeting MO knockdown. This approach allowed embryos to develop further and survive to ~52–72hpf (52hpf shown in Fig. 3). Both bcl2 and tuba1a mRNA injections appear to rescue the mutant phenotypes to a similar extent when examined at 30hpf and suggest that a significant consequence of Tuba1a knockdown is increased apoptosis (Fig. 3 and Table 1).
To examine the consequences of Tuba1a knockdown on CNS development more closely we stained Tuba1a knockdown embryos, with and without Bcl2 or tuba1a mRNA rescue, with anti-acetylated tubulin antibody which detects differentiating neurons. When examined at 24 hours post fertilization (hpf) neuronal differentiation was readily discerned in the control and tuba1b MO-treated embryos, while very little differentiation was observed in tuba1a MO-treated embryos that were rescued with bcl2 mRNA (top panels Fig. 4). However, by 52hpf bcl2 mRNA rescued some neural development in tuba1a MO-treated embryos (bottom panels, Fig. 4), but this rescue was not as robust as that observed for the tuba1a mRNA (bottom panels, Fig. 4). These data suggest that apoptosis, resulting from Tuba1a knockdown, contributes to the neural deficits observed in Tuba1a knockdown fish.
Figure 4. Tuba1a knockdown dramatically inhibits CNS development.
Confocal images of zebrafish injected with tuba1a MO and rescue mRNAs. Top panels: Control (0.25mM), tuba1a (0.125mM) and tuba1b-targeting morpholinos (MO) (0.25mM), along with bcl2 mRNA (25pg) were injected into single cell zebrafish embryos. Embryos were analyzed at 24 hours post fertilization for differentiating neurons by whole mount immunostaining with anti-acetylated tubulin antibody. Shown are representative images of immunostained embryos. Arrowheads point to the developing post-optic commissure (POC) and the anterior commissure (AC), while arrows point to the trigeminal ganglion (TG) and the motor neurons (MN). Note the lack of differentiating neurons in the Tuba1a knockdown embryos. Bottom panels: Control (0.25mM) or tuba1a-targeting (0.125mM) MOs with bcl2 mRNA (25pg) or tuba1a mRNA (12.5pg) were injected into single cell zebrafish embryos. Embryos were analyzed for differentiating neurons by whole mount immunostaining with anti-acetylated tubulin antibody. Shown are representative images of immunostained embryos focusing on the head/eye region. The majority of immunostaining detects sensory neurons on the head (black arrows), eye (white arrows) and running over the yolk (white arrowheads). Note that both the bcl2 and tuba1a mRNA rescued embryos show significant neural differentiation; however tuba1a mRNA appears to rescue nervous system development much better than bcl2 mRNA.
Identification of a tuba1a promoter G/C-rich enhancer
The above experiments suggest that Tuba1a is an essential tubulin subunit necessary for CNS formation and regeneration. To elucidate mechanisms by which tuba1a is induced during axonogenesis we assayed tuba1a promoter in differentiating PC12 cells. Transient transfection of PC12 cells with a series of 5′ promoter deletions identified 2 regulatory regions; one, located between positions −641 and −182, mediates promoter repression while the other, located between positions −182 and −75, mediates promoter activation (Fig. 5A). Additional deletions suggested that the −182 to −75 region could be further subdivided into two regions with sequences −182 to −104 and −104 to −75 contributing to high level tuba1a promoter activity. The experiments described here focus on the −182 to −104 region.
Figure 5. Identification of a tuba1a promoter enhancer.
(A) 5′ promoter deletions identify tuba1a regulatory elements. PC12 cells were co-transfected with the indicated tuba1a (α1T)-luciferase constructs along with CMV-CAT for normalization. Two days post-transfection cells were assayed for luciferase and CAT activity. Reported are relative light units (RLU) representing luciferase activity normalized to CAT activity. (* p<0.05, t-test). (B) The −182/−104 tuba1a sequence confers activation to a heterologous minimal enkephalin (MEK) promoter in an orientation independent manner. PC12 cells were transfected and assayed, as described in (A), with the indicated constructs. (* p<0.01, t-test). (C) The −182/−104 tuba1a enhancer binds proteins present in PC12 nuclear extracts. Radiolabelled −182/−104 tuba1a enhancer was mixed with and without PC12 nuclear extract (NE) and DNA:protein complexes resolved on native acrylamide gels (EMSA). Shown is a representative autoradiogram indicating free probe (F) and 3 DNA:protein complexes (arrows). Specificity of DNA:protein binding is indicated by competition with 50X unlabeled tuba1a enhancer DNA (competitor). (D) Sequence comparison of the proximal promoter of tuba1a from goldfish (gf) and zebrafish (zf). The conserved protein binding site identified in Fig. 6 is underlined. The CAAT and TATA boxes are identified by dashed underline. An arrow identifies the transcription start site, identified in goldfish, and carrots above and below the sequence identifies the end of exon 1 and beginning of intron 1. Values in (A) and (B) are means +/− SEM.
To determine if the −182 to −104 region of the tuba1a promoter harbored an enhancer, we placed it in front of the minimal enkephalin (MEK) promoter driving luciferase expression (Walke et al., 1996). Both 5′ to 3′ and 3′ to 5′ orientations of the −182 to −104 tuba1a sequence conferred increased expression on the MEK promoter suggesting it functioned as a classical enhancer (Fig. 5B). Putative elements contributing to enhancer activity were mapped by identifying nuclear protein binding sites using gel electrophoretic mobility shift assays (EMSAs). For these experiments we generated 4 overlapping double-stranded oligonucleotides spanning nucleotides −182 to −104 and performed EMSAs with PC12 cell nuclear extracts. One of the four probes (−189 to −142) demonstrated strong and specific binding (arrows in Fig. 5C). Interestingly, a similar binding pattern was observed with zebrafish brain nuclear extracts (Fig. 6B). We further mapped protein binding to a small region spanning positions −165 to −149 (data not shown).
Figure 6. Mutation analysis of the −182/−104 tuba1a enhancer identifies a G/C-rich sequence that is necessary for nuclear protein binding and enhancer activity.
(A)DNA sequences of the oligonucleotides used for mapping the protein binding site in (B). Wt, is the wild-type sequence with the G/C-rich sequence found to be necessary for protein binding typed in bold and underlined. M1–M10, show the basepair changes made in the Wt oligonucleotides for mapping the protein-binding enhancer element. (B) EMSA demonstrates that a G/C-rich sequence is necessary for binding nuclear proteins from zebrafish brains. Nuclear protein extract was mixed with radiolabeled Wt oligonucleotides with and without 50X excess of unlabelled mutant oligonucleotides, M1–M10. Specific binding is indicated by a black arrow, non-specific binding (NS), and free probe (F). Note mutant oligonucleotide competitors M3–M8 partially or completely failed to compete for binding, indicating that these nucleotides are critical for protein binding to the −182/−104 tuba1a enhancer. (C) A single basepair mutation in the tuba1a enhancer’s G/C-rich sequence reduces enhancer activity in PC12 cells. PC12 cells were transfected with −182/−104MEK (see Fig. 5B) or −182/−104(M4)MEK (harbors the −182/−104 G/C-rich enhancer with the single basepair mutation present in the M4 oligonucleotide, shown in (A), in front of the MEK promoter) along with CMV-CAT for normalization. (D) A single basepair mutation in the G/C-rich protein binding site of the −182 tuba1a reporter (−182 α1T in Fig. 5A) reduces promoter activity in PC12 cells. PC12 cells were transfected with −182 α1T or −182 M4 α1T (harbors the −182 α1T sequence with the M4 mutation present in the G/C-rich protein binding site). (C and D) Two days post-transfection cells were assayed for luciferase and CAT activities. Luciferase activity was normalized to CAT activity and reported as relative light units (RLU). Note a single nucleotide substitution in the α1T (tuba1a) promoter’s G/C-rich element decreases enhancer activity. Values are means +/− SEM (* p<0.01, t-test).
To identify the nucleotides within the −165 to −149 region that are necessary for protein binding we generated a series of double-stranded oligonucleotides that harbored a single or double nucleotide substitution within this region (Fig. 6A) and used them as competitors in the EMSA (Fig. 6B). It was expected that mutations which disrupt protein binding would not compete with Wt radiolabeled probe for binding to nuclear proteins. These experiments identified oligonucleotides M3–M8 as harboring mutations that reduced or completely abolished their effectiveness as competitors. This defines the protein binding sequence as 5′-GGGAGGTG-3′ (bold and underlined in Fig. 6A, Wt). Due to its sequence composition we named this site the G/C-rich element. Although the G/C-rich element is conserved between the zebrafish and goldfish tuba1a promoters (underlined in Fig. 5D), there is little similarity to the rat Tuba1a sequence (Gloster et al., 1994; Hieber et al., 1998).
We verified that the G/C-rich element was necessary for enhancer activity by introducing the M4 mutation into the −182/−104 MEK:Luciferase and −182 tuba1a:Luciferase expression vectors and measuring their activity in PC12 cells. These experiments showed that the M4 mutation significantly reduces enhancer activity in both the tuba1a promoter and a heterologous promoter (Fig. 6C and D).
The G/C-rich element is necessary for developmental and regeneration-dependent expression of the tuba1a promoter in vivo
We next investigated if the G/C-rich element is important for tuba1a promoter activity in vivo. For this analysis germ-line transgenic zebrafish were created which harbor wildtype (Wt) or mutant tuba1a:GFP expression constructs (Fig. 7A). For each transgene 2 to 4 independent lines were generated and analyzed. The expression pattern for each transgene was similar, although there was some variability in expression intensity between different lines of fish harboring the same transgene. This latter variability probably results from different genomic integration sites or different transgene copy numbers. At 36–48 hours post fertilization (hpf) all lines, except Del −1696/−166, exhibited transgene expression that was restricted to the developing nervous system (Fig. 7B) indicating the G/C-rich element is not required for neural-specific expression (lines Del −166/−146 and M4). In addition, line Del −1696/−146 suggests the G/C-rich element is not sufficient for directing transgene expression to the central nervous system (Fig. 7B, panel h).
Figure 7. The G/C-rich enhancer is necessary for tuba1a promoter-transgene expression during RGC development and optic nerve regeneration.
(A) Schematic representations (not to scale) of the reporter constructs used to create the transgenic lines. Wt transgene includes the 1.696 kb tuba1a promoter with an intact G/C-rich enhancer. Del −166/−146 transgene contains an internal 20 basepair deletion removing the G/C-rich enhancer. M4 transgene contains the M4 point mutation (underlined) within the G/C-rich enhancer. Del −1696/−166 transgene contains a deletion of most of the promoter sequence upstream of the G/C-rich enhancer. (B) Representative bright field (a, c, e, and g) and fluorescent images (b, d, f, and h) of the transgenic lines at 36–48 hpf. Note the similar expression of GFP in the CNS in Wt, Del −166/−146 and M4 constructs. Del −1696/−166 exhibits no detectable GFP expression after 24 hours post fertilization. In the fluorescent images there is variable auto-fluorescence in the yolk marked with an asterisk (*). (C) Transgenic fish harboring the Wt, Del−166/−146 or M4 mutant promoters driving GFP expression were harvested at 48 hpf. Retinal sections were stained with the nuclear dye Hoechst (a, e, and i), examined for GFP expression (b, f, and j) and stained with the Zn5 antibody that identifies differentiating RGCs (c, g, and k). Merged images showing GFP and Zn5 staining are also shown (d, h, and l). Note the Wt promoter transgene is strongly expressed in the differentiating retinal ganglion cells, while Del −166/−146 and M4 promoters exhibited a significant reduction in the number of GFP positive cells in the GCL. Three fish per line and two lines per transgene were assayed with similar results. (D) Transgenic fish harboring the Wt, Del−166/−146, M4 or Del−1696/−166 promoters driving GFP expression had their right optic nerve crushed on day 0. Six days later, left (control) and right (crush) retinas were isolated and assayed for endogenous tuba1a (α1Tubulin) mRNA by in situ hybridization (a, c, e, g, i, k, m, and o) or transgene promoter activity by GFP fluorescence (b, d, f, h, j, l, n, and p), respectively. In situ hybridization for tuba1a expression and GFP images are from adjacent retinal sections. At least three independent lines were assayed for each transgene, except M4 for which only two lines were identified. Three fish from each line were tested, all of which gave similar results. Note that although endogenous tuba1a (α1Tubulin) expression is highly induced following optic nerve crush in all fish tested only transgenic fish harboring the Wt tuba1a promoter show strong transgene GFP induction following optic nerve crush. White arrows indicate GFP-positive RGCs, residing in the ganglion cell layer (GCL).
During development differentiating RGCs exhibit high levels of tuba1a expression (Gulati-Leekha and Goldman, 2006). To determine if the G/C-rich element is involved in the developmental induction of the tuba1a promoter we assayed transgene expression in zebrafish retinas at 48 hpf, a time when many RGCs are extending their axons into their brain targets. Zn5 immunostaining was used to identify differentiating RGCs and GFP expression was used to assay tuba1a promoter activity (Fig. 7C). Indeed, Wt tuba1a promoter activity is highest in differentiating RGCs with 88 +/− 5% of the Zn5-positive cells co-expressing GFP (Fig. 7C, panels a–d). In contrast, deletion of the G/C-rich element (Del −166/−146) resulted in only 3 +/−1% co-labeling (Fig. 5C, panel e–h), while a point mutation in the G/C-rich element (M4) resulted in 19 +/−3% co-labeling (Fig. 7C, panel i–l). Two independent lines for each transgene were examined with similar results. These results suggest that the G/C-rich element is important for increased promoter activity during RGC axon extension.
Because tuba1a promoter activity is induced during optic nerve regeneration (Goldman et al., 2001; Goldman and Ding, 2000; Senut et al., 2004), we were most interested in determining if the G/C-rich element was necessary for regeneration-dependent gene induction. To induce a regeneration response we crushed the right optic nerve of adult transgenic fish. The left optic nerve remained uninjured and the left retina served as a control. Fish were then sacrificed at 3 and 6 days post-injury (dpi) and their eyes fixed and sectioned. In situ hybridization was used to assay endogenous tuba1a expression, while transgene GFP expression was assayed using fluorescence microscopy. Figure 7D shows data from 6 dpi; similar results were observed at 3 dpi. All 3 fish from each transgenic line responded to optic nerve crush in an identical fashion. Endogenous tuba1a expression was used as a positive control for regeneration-dependent gene induction (Fig. 7D, compare panels a, e, i, and m with c, g, k, and o). As we previously reported (Goldman et al., 2001; Senut et al., 2004), Wt tuba1a transgenic promoter activity was highly induced following optic nerve crush as indicated by 84 +/−3% of the cells in the ganglion cell layer (GCL) expressing GFP (2 lines, n=3) (white arrows in Fig. 7D, panel d). Deletion of the G/C-rich element prevented regeneration-dependent tuba1a promoter induction (4 lines, n=3) (Fig. 7D, panel h). A single base-pair substitution within the G/C-rich element, M4, prevented regeneration-dependent transgene induction in one line (not shown) and severely reduced its response in a second line, 9 +/−1% GFP-positive cells in the GCL (n=3) (Fig. 7D, panel l). Del −1696/−166 was unresponsive to optic nerve injury in all fish tested (3 lines, n=3; Fig. 7D, panel p). These results indicate that the G/C-rich element is necessary, but not sufficient for robust regeneration-dependent induction of the tuba1a promoter.
The Sp consensus binding site cannot functionally replace the G/C-rich element in vivo
The above data demonstrate that the G/C-rich element is important for tuba1a promoter induction in differentiating RGCs and in adult RGCs that are regenerating their damaged axons. Because the G/C-rich element is similar to the G/C-rich Sp consensus sequence (GGGGCGGGG) we investigated if this latter sequence could functionally substitute for the tuba1a promoter’s G/C-rich element. This idea seemed reasonable since we were able to demonstrate that the Sp consensus sequence efficiently competed with nuclear protein binding to the tuba1a promoter’s G/C-rich element (Supplementary Fig. 3). Therefore, we generated transgenic fish harboring the −1696 tuba1a promoter with the G/C-rich element replaced by a Sp consensus sequence (SpCon). Developmentally, SpCon fish (Fig. 8A) expressed the GFP reporter in a similar fashion as transgenic fish harboring the Wt −1696 tuba1a:GFP transgene (see Fig. 7B, Wt). Surprisingly, adult SpCon fish did not induce transgene expression following optic nerve crush (Fig. 8B, compare GFP expression between Wt transgenic fish in panel (b) to SpCon transgenic fish in panel (f)). In addition, Sp1, Sp3 and Sp4 gene expression is not induced in RGCs that are regenerating their optic axons (Supplementary Fig. 4). Although Sp transcription factors may be activated in a posttranscriptional manner, our data showing the Sp consensus sequence cannot functionally replace the G/C-rich element suggests Sp proteins do not mediate activation of the tuba1a promoter via this element.
Figure 8. The Sp consensus binding site cannot functionally replace the G/C-rich enhancer in vivo.

(A and B) Transgenic zebrafish were created which harbor the Sp consensus (SpCon) binding site replacing the G/C-rich enhancer in the tuba1a:GFP transgene. (A) SpCon transgenic fish display nervous system restricted GFP expression at 48 hours post fertilization similar to Wt promoter (Fig. 8A). Auto-fluorescence in the yolk is noted with an asterisk (*). (B) SpCon transgenic fish fail to re-express GFP during optic nerve regeneration. Wt tuba1a:GFP transgenic fish exhibit strong tuba1a (a) and GFP (b) expression in the retinal ganglion cell layer (GCL) 3 days following optic nerve injury. In addition, we observed variable and weak GFP expression in inner nuclear layer cells in this particular line of fish which may reflect the transgene insertion site. Uninjured SpCon transgenic (c and d) and 3 day post-nerve injury (e and f) retinal sections were stained for tuba1a expression (c and e) or imaged for fluorescence (d and f). Note the absence of GFP fluorescence in the GCL of the injured retina (f) while endogenous tuba1a expression is highly induced (e).
KLF6a and KLF7a are induced following optic nerve injury and regulate tuba1a gene expression
We recently identified RGC genes that are regulated during optic nerve regeneration using a microarray-based screen (Veldman et al., 2007). Interestingly, two zinc finger transcription factors, KLF6a and KLF7a, were found to be induced. The KLF family of proteins is related to the Sp family through similarities in their zinc finger DNA binding domain. There are 16 described KLF proteins in mammals and an unknown number of duplicates in zebrafish (Suske et al., 2005). Due to the conserved structure of their DNA binding domain, all of these proteins have the potential to bind to a G/C-rich element. In situ hybridization assays confirmed our previous data showing KLF6a and KLF7a are induced in RGCs at 3 days post optic nerve crush (Supplemental Fig. 5A-D), while during development only KLF7a is detected in differentiating RGCs (Supplemental Fig. 5E, F). This implicates KLF7a as a potential regulator of gene expression during both development and regeneration, while KLF6a may be a regeneration-specific transcription factor.
We next explored whether KLF6a or KLF7a could bind to the tuba1a promoter’s G/C-rich element and increase promoter activity. EMSAs using in vitro translated, myc-tagged KLF6a and KLF7a showed they bind to the G/C-rich enhancer, as demonstrated by anti-myc-mediated supershifts (Fig. 9A) and competition with the Wt sequence but not the M4 mutant G/C-rich sequence (Fig. 9B). To examine if KLF6a and/or KLF7a could transactivate promoters via the G/C-rich element we constructed new luciferase reporters that harbor the minimal β-globin (MBG) promoter with 4 copies of the G/C-rich element (4xWt) or M4 mutant sequence (4xM4) upstream. These plasmids were transfected into PC12 cells with and without KLF6a or KLF7a expression vectors. The MBG promoter has very little activity in PC12 cells and it is unaffected by overexpression of KLF6a or KLF7a (Fig. 9C). In contrast, both KLF6a and KLF7a strongly induced reporter expression when 4 copies of the G/C-rich enhancer was placed in front of the MBG promoter (4xWt) (Fig. 9C) and mutation of the binding site completely abolished this transactivation (4xM4 in Fig. 9C). We also confirmed that these proteins were able to transactivate the tuba1a promoter through the G/C-rich element (Fig. 9D). G/C-rich element deletion (Del −166/−146) or mutation (M4) in the context of the full length tuba1a promoter attenuated KLF6a and KLF7a activation. The residual activation of these mutant tuba1a promoters may indicate additional KLF binding sites or collaboration with proteins bound to other elements.
Figure 9. KLF6a and KLF7a bind the tuba1a G/C-rich element and can transactivate the tuba1a promoter through the G/C-rich element.
(A) EMSAs were performed using radiolabeled G/C-rich element and in vitro synthesized KLF6a (6), KLF7a (7), myc-tag (MT), myc-tagged KLF6a (MT6), or myc-tagged KLF7a (MT7) protein. If no protein was added the lane is marked with a (-) and if the in vitro transcription translation reaction mix, lacking plasmid, was added the lane is marked with a (+). Both KLF6a and KLF7a bind to the probe (bracket) and in the cases where they harbor a myc-tag the anti-myc antibody (myc-Ab) results in a supershift of the complex (black arrow). (B) MT6 and MT7 binding is specific and can be competed by Wt oligonucleotide but not by M4 oligonucleotide (Wt and M4 EMSAs were performed on separate gels under identical conditions; M4 competitor concentrations are equal to the highest Wt competitor concentrations). Free probe (F). (C) KLF6a or KLF7a over expression can transactivate a heterologous promoter through the isolated G/C-rich element (*p<0.01, t-test). (D) KLF6a or KLF7a overexpression can transactivate the tuba1a (α1T) promoter and this transactivation is attenuated when the G/C-rich element is deleted (Del −166/−146) or mutated (M4) (*p<0.01, ANOVA with Bonferonni post-hoc test comparing tuba1a (α1T) promoter driven expression with KLF6a or KLF7a overexpression, white bars) (# p<0.05, ANOVA with Bonferroni post-hoc test comparing KLF6a or KLF7a overexpression effect on G/C-rich element deletion or mutation tuba1a (α1T):luciferase reporters, comparing grey or black bar to white bar). (C and D) PC12 cells were cotransfected with tuba1a (α1T):luciferase reporter vectors along with pCS2:β-gal and either empty pCS2 vector (Control) or pCS2-KLF6a (KLF6a) or pCS2-KLF7a (KLF7a). Reporter plasmids are: α1T, −1696 tuba1a:luciferase; Del −166/−146, internal deletion of the G/C-rich element in the −1696 tuba1a promoter; M4, point mutant in G/C-rich element of −1696 tuba1a promoter; MBG, minimal β-globin promoter; 4xWT, MBG with 4 copies of the G/C-rich enhancer cloned upstream; 4xM4, MBG with 4 copies of the M4 mutant G/C-rich enhancer cloned upstream. All transfections are normalized to pCS2:β-gal expression and luciferase values are reported as relative light units (RLU) in hundreds. Values are means +/− SEM.
The above data suggest that KLF6a and 7a bind to and regulate tuba1a promoter activity and that Tuba1a expression is necessary for optic axon regeneration. To determine if KLF6a and 7a regulate tuba1a gene expression in vivo we knocked down KLF6a and 7a in axotomized RGCs of adult tuba1a:GFP transgenic fish. Four days post optic nerve lesion and KLF knockdown, retinas were harvested and mRNA isolated for RT-PCR analysis of gene expression. Consistent with the idea that KLF6a and 7a regulate tuba1a promoter activity, we observed a dramatic suppression of regeneration-dependent tuba1a and GFP expression (Fig. 10). We also found that KLF6a and 7a regulates regeneration-dependent p27 gene expression, but has little effect on c-jun and GAPDH gene expression (Fig. 10).
Figure 10. Regeneration-dependent induction of KLF6a and 7a is necessary for tuba1a and p27 gene induction following optic nerve injury.

The optic nerve of adult tuba1a:GFP transgenic fish was lesioned and KLF6a/KLF7a targeting MO or a control MO was applied to the optic nerve stump as described in Materials and Methods. 5 days later retinas were harvested and RNA purified for use in RT-PCR reactions. Shown is an ethidium bromide stained gel showing induction of GFP, tuba1a, p27 and c-jun following optic nerve injury. GAPDH is used as a loading control. Note that KLF6a and 7a knockdown (KLF6/7 MO) caused a significant reduction in GFP, tuba1a and p27 gene induction following optic nerve injury.
DISCUSSION
The main findings from this study are: 1) tuba1a gene expression is necessary for CNS development and optic nerve regeneration, 2) a G/C-rich element participates in tuba1a promoter induction during RGC differentiation and optic nerve regeneration and 3) KLF6a and 7a bind the tuba1a promoter’s G/C-rich element and induce tuba1a gene expression during optic nerve regeneration.
The observation that Tuba1a is necessary for CNS development and optic axon regeneration suggests that other tubulin isoforms expressed in the CNS are unable to compensate for the loss of Tuba1a. This is consistent with the multi-tubulin hypothesis posited over 30 years ago which states that individual tubulin genes encode functionally divergent polypeptides that confer unique properties to the final microtubule polymer (Fulton and Simpson, 1976). Tuba1a and Tuba1b are highly related proteins differing in only 11 amino acids. Interestingly, 2 of the regions where these proteins differ are in putative protein kinase phosphorylation sites located at positions 126 and 232. In Tuba1b these residues are both serine whereas in Tuba1a these residues are alanine and glycine, respectively. It is tempting to speculate that phosphorylation of these serine residues confers properties onto Tuba1b that make it unable to compensate for Tuba1a during CNS development and regeneration.
It is also possible that differential regulation of tubulin isoforms is the predominant driving force for maintaining multiple tubulin encoding genes (Raff, 1984). In this regard, Tuba1b may not compensate for Tuba1a knockdown due to inadequate levels of expression. We have assayed tuba1b expression during optic nerve regeneration and find it is highly induced in RGCs as previously described (Bormann et al., 1998); however, it is not clear if gene induction correlates with protein level and because there are no antibodies that can distinguish the various alpha 1 tubulin isoforms, their relative protein levels remain unknown.
It is intriguing that the tuba1a and tuba1b genes are linked (~3kb separates them) in the zebrafish genome, while they are separated by over 50kb in humans and mice. The close proximity of these genes in zebrafish may allow for sharing of cis-acting regulatory elements leading to coordinated expression of these genes even though they may carry out different functions.
Like our finding for Tuba1a and Tuba1b, there is one report indicating the gamma-tubulin isoforms are not equivalent (Yuba-Kubo et al., 2005). In this study, development of Tubg1−/− mice was arrested at the blastula stage with pronounced disruption of the mitotic spindle even though TUBG2 is expressed in blasotcytes; suggesting it cannot compensate for TUBG1 loss. Interestingly Tubg2−/− mice were normal both in growth and reproduction. Thus like, Tuba1a and Tuba1b, the Tubg1 and Tubg2 gene products are not functionally equivalent in vivo.
In order to better understand the transcriptional mechanisms underlying successful central nervous system regeneration in zebrafish we used the regeneration-responsive tuba1a promoter to identify cis-acting elements necessary for regeneration-dependent promoter induction in RGCs. Previous work in our lab used sequence similarity between goldfish and zebrafish promoters to identify a tuba1a promoter E-box and homeodomain protein binding site that are necessary for regeneration-dependent promoter induction but less important for developmental expression in RGCs (Senut et al., 2004). Conversely, an internal promoter deletion spanning nucleotides −1044 to −846 prevents developmental expression while maintaining injury-dependent expression (Goldman and Ding, 2000). Consistent with this latter result, we also showed by transient expression assays that the full length −1696 tuba1a promoter provided the most robust neural-specific expression in embryos at 48hpf and that as one removes 5′ promoter sequences neural-specific expression is reduced (Hieber et al., 1998). Surprisingly in this study we found that in PC12 cells the full length promoter was less active than the 5′ deleted promoter. This discrepancy may simply reflect the difference between assaying expression in vivo where a large number of different cell types make up the nervous system and in cultured PC12 cells where the population is homogenous. Nonetheless we were able to use PC12 cells to identify a G/C-rich element that is necessary for promoter expression in both developing and adult RGCs that are regenerating their damaged axons.
The tuba1a promoter region harboring the G/C-rich element is highly conserved between zebrafish and goldfish but is not conserved with mammals and may contribute to differences in gene induction or spatial expression between species. Although there are no large regions of identity between fish and mammals, isolated G/C-rich sites in the mammalian Tuba1a promoter exist which may serve a similar function as in fish. It would be interesting to determine if deletion of these elements in the mammalian promoter or knockout of their binding proteins had an effect on promoter induction in differentiating neurons or regenerating neurons of the peripheral nervous system. Alternatively, the G/C-rich element may represent a fish-specific element which contributes to the different regenerative responses between fish and mammals.
Transgenic zebrafish were created with promoter deletions or mutations in the G/C-rich element to test its function in vivo. The Wt tuba1a:GFP transgene is pan-neuronally expressed in the developing central nervous system in a pattern very similar to the endogenous tuba1a gene (Hieber et al., 1998; Goldman et al., 2001). This expression pattern was not grossly different in transgenic fish with deletions or mutations in the G/C-rich element. However closer inspection indicates expression is no longer pan-neuronal throughout development. Unlike the Wt promoter, disruption of the G/C-rich element resulted in loss of transgene expression in cells undergoing axon elongation, such as RGCs at 48 hpf. However, expression in putative progenitors lining brain ventricles was maintained (unpublished observation). Therefore the G/C-rich element is not part of the basal promoter but rather appears to be important for tuba1a promoter activation in differentiating neurons and, in particular, differentiating RGCs. Interestingly, we found this same element is necessary for tuba1a promoter induction during optic nerve regeneration in adult animals, suggesting some of the components regulating differentiation-dependent gene induction also contribute to regeneration-dependent gene induction. We have previously shown that there are elements within the tuba1a promoter that are regeneration specific (Goldman and Ding, 2000; Senut et al., 2004), and a similar finding was reported for the GAP43 promoter (Udvadia et al, 2001). Therefore, gene regulation during axon regeneration appears to involve both novel regeneration signals and re-activation of developmental signals.
Our search for potential binding proteins to the G/C-rich element led to the identification of Sp/KLF family members. This is a large family of transcription factors with a conserved zinc finger DNA binding domain in common. There are 9 known Sp proteins and 16 KLF proteins in mammals (Suske et al., 2005). Several of the Sp proteins such as Sp1 and Sp3 are ubiquitously expressed while others such as Sp5 and Sp7 are more tissue restricted. Initial EMSA analysis of the G/C-rich enhancer indicated that protein binding could be competed with a Sp1 consensus binding site oligo. However, this Sp1 consensus sequence cannot substitute for the tuba1a promoter G/C-rich regulatory element in vivo. This surprising result could be caused by several factors. First, the EMSA is an in vitro analysis of protein binding to DNA devoid of its normal, context dependent, modifications such as methylation or histone binding. The sequence differences between the G/C-rich enhancer and the Sp1 consensus site may cause changes in these properties in vivo. Second, the binding affinity of different Sp/KLF proteins for the G/C-rich enhancer or the Sp consensus sequence may be different. Third, the protein(s) involved in regulating the G/C-rich enhancer may not be highly represented in the zebrafish brain nuclear protein extract used to determine binding, especially if the protein(s) are regeneration specific. Our results suggest the G/C-rich element is functionally distinct from the Sp1 consensus.
A microarray screen for genes induced in RGCs following optic nerve injury identified KLF6a and KLF7a (Veldman et al., 2007). Although both KLF6a and KLF7a are induced in RGCs during optic nerve regeneration, only KLF7a is expressed during RGC development. These data suggest that in the retina KLF7a is an axon growth associated gene, while KLF6a is a regeneration-specific gene.
KLF6a and KLF7a are members of a distinct sub-family of Sp/KLF proteins. These proteins are involved in cell cycle exit and differentiation. The single Drosophila homolog luna has been shown to be involved in cell differentiation (De Graeve et al., 2003). Mammalian KLF6 is expressed in restricted domains of the developing nervous system and several non-neural tissues and is a tumor suppressor (Laub et al., 2001a; Narla et al., 2001; Reeves et al, 2004). Interestingly, it is induced in mammalian dorsal root ganglia following sciatic nerve transection (Nilsson et al., 2005). KLF6 knockout mice are embryonic lethal on day E12.5, precluding examination of its role in nerve development and regeneration (Matsumoto et al., 2006). KLF7 is more broadly expressed in the nervous system and also can regulate cell cycle exit (Laub et al., 2001b). KLF7 null mice are neonatal lethal and exhibit neurite outgrowth and axon guidance problems, notably in the visual system (Laub et al., 2005). Interestingly, ~3% of KLF7 null mice escape lethality and survive to adulthood. An increase in KLF6 expression was noted in these mice indicating a possible compensatory mechanism and conserved functional properties of these two proteins ( Laub et al., 2005). Most interesting is the recent report that KLF6 and 7 stimulate optic axon regeneration in mammals (Moore et al., 2009). Thus it is likely that mechanisms underlying successful regeneration in fish will be shared with those that promote CNS regeneration in mammals and that the study of regeneration in zebrafish will suggest novel strategies for inducing regeneration in mammals.
KLF6 target genes have not been identified in the nervous system; however, several target genes for KLF7 have been identified. TrkA was identified as a KLF7 target due to its decreased expression in KLF7 null embryos dorsal root ganglia (Lei et al., 2005). This neurotrophin receptor is induced in RGCs during optic nerve regeneration in the tench, another teleost fish (Caminos et al., 1999). Induction of this receptor may be critical for cell survival and/or axon extension following nerve injury. KLF7 has also been shown to regulate the expression of cell adhesion molecule L1 and GAP43 in the olfactory bulb of mice (Kajimura et al., 2007). These two genes are well described RAGs in both mammals (Basi et al., 1987; Anderson et al., 1998) and teleosts (Bernhardt et al., 1996; Becker et al., 2005). KLF7 also regulates p27 gene expression in mammals and p27 may contribute to regeneration via regulated rho signaling (Laub et al., 2005). Our identification of the tuba1a promoter as a target of KLF6a and 7a transactivation adds to this list of RAGs and suggests that KLF6 and KLF7 play an important role in gene regulation during nerve regeneration.
In summary, we report that tuba1a gene expression is essential for CNS development and optic axon regeneration. We identified a G/C-rich enhancer within the tuba1a promoter that is necessary for tuba1a gene induction in differentiating RGCs during development and in adult RGCs that are regenerating a damaged axon. This enhancer binds KLF6a and 7a; two transcription factors previously identified as regeneration-associated genes whose expression is essential for optic axon regeneration (Veldman et al., 2007). KLF6a and 7a regulate tuba1a promoter activity in vitro via the G/C-rich enhancer and KLF6a and 7a induction in vivo contribute to regeneration-dependent tuba1a and p27 gene expression. These data suggest that KLF6a and KLF7a are important regulatory proteins necessary for the correct expression of tuba1a and other RAGs during successful nerve regeneration.
Experimental Methods
Animals, Optic nerve injury and retinal explant assay
Zebrafish were obtained from our breeding colony and maintained at 28 °C with 14/10 h light/dark cycle. Zebrafish were anesthetized and the optic nerve was lesioned as previously described (Veldman et al., 2007). Four days following optic nerve transection and morpholino treatment, retinas were isolated and placed in explant culture as previously described (Veldman et al., 2007). Briefly, fish were dark adapted, euthanized and retinas isolated. Retinas were cut into 0.5mm squares with a razor blade and digested with hyaluronidase (1mg/ml) for 15 min at room temperature. Explants were rinsed 3x with L15 culture media and plated, one retina per well, in a 6-well plate precoated with poly-L-lysine and laminin. Explants were cultured in 0.5ml L15 media containing 8% fetal calf serum, 3% zebrafish embryo extract and 1x antibiotic/antimycotic at 28 °C for 4 days in a humidified ambient air incubator. Adherent explants were quantified for axon density and length. Density was measured as the number of neurites >250 μM in length per explant. Axon length was measured with ImageJ software using the segmented line tool to determine the number of pixels from growth cone to the edge of the explants then converting the number of pixels to μm. Total neurite outgrowth was measured by calculating the “Nerve Growth Index” where density scores for each explant are multiplied by the average length. Axon density scores of 0, 1, 2, 3, and 4 correspond to 0, 1–20, 21–40, 41–100 and >100 neurites/explants, respectively. Axon length scores of 0, 1, 2, 3, and 4 correspond to 0–250, 251–500, 501–1000, 1001–1500 and >1500 mm, respectively. All animals were treated according to the guidelines of the University Committee on Use and Care of Animals at the University of Michigan.
Morpholino treatments and mRNA rescue
Morpholino-modified antisense-mediated gene knockdown in adult RGCs following optic nerve transection was accomplished by placing a small piece of Gelfoam, soaked in morpholino (MO) (unless otherwise indicated, 1 μl of a 0.25–1mM MO solution was added to the Gelfoam), onto the lesioned optic nerve stump for one day as previously described (Veldman et al., 2007). MOs were purchased from Gene Tools, LLC. The following MOs targeting the 5′ UTR or spanning the initiator AUG of the tuba1a and tuba1b transcript were used: tuba1a MO-1, 5′GAATAACGTTATAACGGACTGATC; tuba1a MO-2, 5′TCACGCATTGTTTCTAGTAGTGTTG; tuba1b MO-1, CAAATGTGTATTTTCGATGAATGGC; tuba1b MO-2 TGAATAGGATAGCCTTCGGTTTTTC; standard control MO that does not target any sequence is: 5′CCTCTTACCTCAGTTACAATTTATA. KLF6a and 7a-targeting MOs have been previously described (Veldman et al., 2007). For analysis of development tuba1a MOs (0–0.25mM) were injected (~1nl volume) into single cell zebrafish embryos and injected embryos were observed at 24–72hpf. This concentration of tuba1a MO corresponds to about 0–2.5ng of injected MO. Control MO was used at 0.25–1mM which corresponds to about 2.5–10ng of injected MO. We confirmed that tuba1a and tuba1b targeting MOs were effective by replacing the 5′ UTR of the GFP sequence with the MO target sequence. Expression plasmids harboring the sCMV promoter driving GFP expression with the appended MO tuba1a or tuba1b target sequence were injected into single cell zebrafish embryos with control, tuba1a or tuba1b-targeting MOs (0.2mM) and GFP expression assayed 1–2 days later. For mRNA rescue experiments tuba1a and bcl2 mRNA was synthesized using the mMessage mMachine kit (Ambion). tuba1a and bcl2 mRNA rescue experiments used approximately 12.5pg and 25pg of mRNA, respectively. Higher tuba1a mRNA concentrations were toxic to embryos.
Promoter analysis in vitro
Various tuba1a promoter deletions or mutations were subcloned into the pXP2 vector (Nordeen, 1988) to generate tuba1a:luciferase expression vectors. A minimal enkephalin promoter (MEK) (Walke et al., 1996) or the minimal β-globin promoter (MBG) (Woolfe et al., 2005) driving luciferase expression was used to test for enhancer activity or the function of small tuba1a regulatory elements. All plasmids were verified by sequencing. CMV:CAT (chloramphenicol acetyltransferase) or CMV: β-gal (beta-galactosidase) expression vector was co-transfected with luciferase expression vector for normalization of transfection efficiency. PC12 cells were transfected using FUGENE 6 reagent (Roche; Indianapolis, IN) as described by the manufacturer. Cells were harvested 48h post-transfection and assayed for luciferase and CAT or β-gal activity.
Cloning of zebrafish KLF6a and KLF7a
KLF6a (accession NM_201461) and KLF7a (accession BC124329) cDNAs were cloned from 3 day post-optic nerve crush retinal RNA using Superscript II and oligo dT (Invitrogen; Carlsbad, CA) for reverse transcription. Both cDNAs were cloned into plasmid pCS2+ for over expression experiments. Primers used to amplify cDNAs for cloning are KLF6a forward 5′-GAGAGACAATTGATGGATGTTCTACCAATGTGC-3′, KLF6a reverse 5′-GAGAGACTCGAGTCAGAGGTGCCTCTTCATGTG-3′, KLF7a forward 5′-GAGAGAGAATTCATGGACGTGTTGGCGAATTAC-3′, KLF7a reverse 5′-GAGAGACTCGAGTTAGATATGTCGCTTCATGTG-3′. Restriction enzyme sites added for cloning purposes are underlined in the primer sequences.
Electrophoretic mobility shift assays
EMSA was performed using standard protocols. Briefly, oligonucleotide probes were labeled with 32P-dCTP and incubated with 5 μg of protein in each binding reaction. TnT SP6 Quick kit (Promega, Madison, WI) was used to generate KLF6a and KLF7a proteins in vitro and 2μL of each synthesis reaction was used in the binding reactions. Supershift assays were performed with the addition of 1μL anti-myc 9E10 (Developmental Studies Hybridoma Bank, University of Iowa) mouse monoclonal antibody. Protein:DNA mixes were resolved on non-denaturing polyacrylamide gels and DNA migration was visualized by exposing the gel to Kodak imaging film.
Generation of transgenic zebrafish
Transgene vectors were derived from the wild type (Wt) −1696α1TIpEGFP plasmid (Goldman et al., 2001). Vectors were linearized and injected into one to two cell stage embryos as described previously (Goldman and Ding, 2000). Injected embryos were raised to adulthood and transgenic founders were identified by breeding F0 injected fish to wild types and screening for GFP-positive embryos or by PCR for GFP in genomic DNA from pools of F1 embryos. Multiple independent lines were identified for each transgene. Del −166/−146 harbors an internal 20 base pair deletion (−166 to −146) of the Wt promoter. M4 harbors a single base pair G to T mutation at position −159. Del −1696/−166 is a large 5′ deletion from position −1696 to −166. SpCon harbors the Sp consensus GC-box binding motif (5′-ACGGGGCGGGGCT-3′) in place of the endogenous protein binding site (5′-TCTGGGAGGTGTC-3′) centered on position −160 in the Wt vector. All plasmids were sequence verified prior to injection into embryos.
Immunostaining and in situ hybridization
Whole mount immunostaining was carried out as previously described (Gulati-Leekha and Goldman, 2006). Briefly, embryos were fixed in 4% paraformaldehyde at 4 °C overnight, manually dechorionated, washed in 0.1M phosphate buffer and permeabilized in water at room temperature for 5 min to 1 hr, and in acetone at −20 °C for 7 min. After blocking in 2% goat/horse serum, embryos were incubated with 1/1000 dilution of anti-acetylated tubulin antibody in 0.1M phosphate buffer and 1% DMSO at 4 °C overnight. The immune complex was visualized with a Donkey Alex555 secondary antibody at 1/500 dilution. For immunohistochemistry on tissue sections, embryos or adult eyes were fixed in 1% paraformaldahyde in PBS overnight at 4°C. Zebrafish embryos or adult eyes were prepared for cryosectioning (Barthel and Raymond, 2000) and cut at 5μm on a Leica CM3050S cryostat (Wetzlar, Germany). Strong GFP fluorescence was maintained in prepared tissue using this protocol and no immuno-labeling for GFP was necessary. Immunostaining was performed as in Senut et al., 2004. Mouse anti-zn5 (Zebrafish International Resource Center; Eugene, OR) was used at 1:500 dilution and Hoechst 33258 at 0.25μg/ml. Fluorescently labeled cells were quantified in three non-adjacent sections per animal. For cell counts in adult retina, Hoechst stained nuclei within the retinal ganglion cell layer were counted to give the total number of RGCs. The number of Hoechst-positive nuclei completely surrounded by GFP fluorescence was then counted and the percent of GFP positive RGCs calculated. For 48 hpf retinal sections, Zn5 stained Hoechst-positive RGCs were counted and the number of GFP positive cells within this group determined to give the percentage of GFP positive differentiating RGCs. In situ hybridizations were performed with digoxigenin labeled cRNA probes as described previously (Barthel and Raymond, 2000). The tuba1a anti-sense probe was 966 nucleotides encompassing the 3′ end of the transcript, accession #BC067554. KLF6a and KLF7a probes consisted of the sequence of the open reading frame for each mRNA. Axonogenesis in embryos was visualized with an anti-acetylated tubulin antibody (1:1000 dilution).
RT-PCR
Total RNA was isolated from adult retina or zebrafish embryos using Trizol reagent (Invitrogen). Reverse transcription was performed on 1μg of total RNA using oligo dT primer and Superscript II (Invitrogen; Carlsbad, CA). Radioactive PCR and autoradiography were as described in Senut et al., 2004. PCR reactions were stopped at predetermined non-saturating cycle numbers. NIH Image software was used for densitometric measurements (http://rsb.info.nih.gov/nih-image/). Primer sequences for each gene are tuba1a: 5′-TAAGTGACACAAATAACATGCAGG-3′ and 5′-CAGCTCAAAGAACTGTACC-3′; sp1: 5′-CAGATGCAGCCGATACAGATGC-3′ and 5′-GGAGCAGACGAAGGGTCTCTCG-3′; sp3: 5′CATGGGCAAGAAGAAGCAGC-3′ and 5′-GAATCTCTTCGACGACTCCG-3′; sp4: 5′ACCATCACCGGTGTACAAGG-3′ and 5′GACGAATGGTCGTTCTCCTG-3′; l24: 5′-CGACCCAGAGCAGCAAGG-3′ and 5′-AGCACATCAGAGTTTAGC-3′; gapdh: 5′-ATGACCCCTCCAGCATGA-3′ and 5′-GGCGGTGTAGGCATGAAC-3′; gfp: 5′-GCAAGCTGACCCTGAAGTTC-3′ and 5′-AACACCGCCTAGAACTTCA-3′; c-jun: 5′-TGGATACAACCACAAGGCTCT-3′ and 5′-GTCACGTTCTTGGGACACAG-3′; p21: 5′-GAAGCGCAAACAGACCAACAT-3′ and 5′-GCAGCTCAATTACGATAAAGA-3′. Primer sequences for amplifying the tuba1a mRNA used in rescue experiments: 5′-CGCGGATCCATGCGCGAGTGCATCTCTATCCACG-3′ and 5′-CCGCTCGAGCACTTAATATTCCTCGCCCTCTTCCTCTCC-3′. Rescue experiments using bcl2 mRNA used the zBlp2 prosurvival Bcl2 family member (Kratz et al., 2006).
Supplementary Material
Control or 2 different tuba1a-targeting MOs were delivered to RGCs, in vivo, by placing MO-soaked Gelfoam on the sectioned optic nerve stump. The next day the Gelfoam was removed and 3 days later retinas were isolated, diced and placed in explant culture for 4 days prior to analysis of neurite outgrowth. The graph on the right shows the Nerve Growth Index (see Materials and Methods) for each MO at the concentration indicated. Note the concentration dependent reduction in neurite outgrowth (Nerve Growth Index) as increasing concentrations of the tuba1a-targeting MO are applied to the optic nerve stump. Also note that tuba1a-targeting MOs #1 and #2 at 0.065 mM concentration have little effect on neurite outgrowth when applied individually, but synergize to significantly reduce neurite outgrowth when added together. The image on the left shows representative explants treated with tuba1a-targeting MOs #1 and/or #2 at 0.065mM concentration. Note the reduced axon outgrowth in explants from retinas that received a combination of MOs #1 and #2.
tuba1a-targeting MO was delivered to RGCs, in vivo, by placing MO-soaked Gelfoam on the sectioned optic nerve stump. The next day the Gelfoam was removed and 3 days later retinas were isolated, sectioned and assayed for cell death using TUNEL stain. Note that like the uninjured retina, MO-treated retinas showed little cell death. Positive control is a nuclease-treated retina. Brown signal indicates cells undergoing apoptosis.
Electrophoretic mobility shift assay showing the Sp1 consensus binding site probe can efficiently compete nuclear protein binding to the G/C-rich enhancer. Below the gel are shown the oligonucleotide sequences of consensus protein binding sites for myc-associated zinc finger protein (MAZ), GLI, specificity protein 1 (SP1), and smad interacting protein (SIP) used as unlabeled competitors. Consensus binding sites are underlined. The arrow marks specific protein binding. NS, Nonspecific binding; F, free probe.
Sp1, Sp3, and Sp4 expression is not changed during optic nerve regeneration in zebrafish. Quantitative RT-PCR for negative control ribosomal protein encoding transcript L24, positive control tuba1a (α1T), and Sp1, Sp3, and Sp4 using uninjured retinal RNA and 3 day post-optic nerve crush retinal RNA (ONC) as template. Bar graphs represent densitometric quantification of autoradiographic analysis of RT-PCR products from 3 separate fish.
(A and B) In situ hybridization showing KLF6a induction in retinal ganglion cells 3 days post-optic nerve crush (B) as compared to uninjured retina (A). (C and D) In situ hybridization showing KLF7a induction in retinal ganglion cells 3 days post-optic nerve crush (D) as compared to uninjured retina (C). (E) KLF6a is not expressed in the retina at 48 hours post fertilization (hpf). (F) KLF7a is expressed in the differentiating RGCs at 48 hpf. Arrows denote developing GCL. Outer nuclear layer (OLN), inner nuclear layer (INL), and ganglion cell layer (GCL).
Acknowledgments
We thank Peter Macpherson for help with confocal microscopy and members of the Goldman Lab for insightful discussions. We also thank Tori Melendez for excellent care of fish and maintenance of our fish colony. This work was supported by Grant MEDC 38 from the Michigan Economic Development Corporation, Life Sciences Corridor (DG), National Glaucoma Research (DG), NEI (DG), an NIH Organogenesis Training grant (MBV), and an NIH vision research training grant (MBV).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Anderson PN, Campbell G, Zhang Y, Lieberman AR. Cellular and molecular correlates of the regeneration of adult mammalian CNS axons into peripheral nerve grafts. Prog Brain Res. 1998;117:211–232. doi: 10.1016/s0079-6123(08)64018-2. [DOI] [PubMed] [Google Scholar]
- Barthel LK, Raymond PA. In situ hybridization studies of retinal neurons. Methods Enzymol. 2000;316:579–590. doi: 10.1016/s0076-6879(00)16751-5. [DOI] [PubMed] [Google Scholar]
- Basi GS, Jacobson RD, Virag I, Schilling J, Skene JH. Primary structure and transcriptional regulation of GAP-43, a protein associated with nerve growth. Cell. 1987;49:785–791. doi: 10.1016/0092-8674(87)90616-7. [DOI] [PubMed] [Google Scholar]
- Bastmeyer M, Beckmann M, Schwab ME, Stuermer CA. Growth of regenerating goldfish axons is inhibited by rat oligodendrocytes and CNS myelin but not but not by goldfish optic nerve tract oligodendrocytelike cells and fish CNS myelin. J Neurosci. 1991;11:626–640. doi: 10.1523/JNEUROSCI.11-03-00626.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker T, Bernhardt RR, Reinhard E, Wullimann MF, Tongiorgi E, Schachner M. Readiness of zebrafish brain neurons to regenerate a spinal axon correlates with differential expression of specific cell recognition molecules. J Neurosci. 1998;18:5789–5803. doi: 10.1523/JNEUROSCI.18-15-05789.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker T, Lieberoth BC, Becker CG, Schachner M. Differences in the regenerative response of neuronal cell populations and indications for plasticity in intraspinal neurons after spinal cord transection in adult zebrafish. Mol Cell Neurosci. 2005;30:265–278. doi: 10.1016/j.mcn.2005.07.008. [DOI] [PubMed] [Google Scholar]
- Benfey M, Aguayo AJ. Extensive elongation of axons from rat brain into peripheral nerve grafts. Nature. 1982;296:150–152. doi: 10.1038/296150a0. [DOI] [PubMed] [Google Scholar]
- Benowitz LI, Shashoua VE, Yoon MG. Specific changes in rapidly transported proteins during regeneration of the goldfish optic nerve. J Neurosci. 1981;1:300–307. doi: 10.1523/JNEUROSCI.01-03-00300.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernhardt RR, Tongiorgi E, Anzini P, Schachner M. Increased expression of specific recognition molecules by retinal ganglion cells and by optic pathway glia accompanies the successful regeneration of retinal axons in adult zebrafish. J Comp Neurol. 1996;376:253–264. doi: 10.1002/(SICI)1096-9861(19961209)376:2<253::AID-CNE7>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- Bormann P, Zumsteg VM, Roth LW, Reinhard E. Target contact regulates GAP-43 and alpha-tubulin mRNA levels in regenerating retinal ganglion cells. J Neurosci Res. 1998;52:405–419. doi: 10.1002/(SICI)1097-4547(19980515)52:4<405::AID-JNR4>3.0.CO;2-D. [DOI] [PubMed] [Google Scholar]
- Buffo A, Holtmaat AJ, Savio T, Verbeek JS, Oberdick J, Oestreicher AB, Gispen WH, Verhaagen J, Rossi F, Strata P. Targeted overexpression of the neurite growth-associated protein B-50/GAP-43 in cerebellar Purkinje cells induces sprouting after axotomy but not axon regeneration into growth-permissive transplants. J Neurosci. 1997;17:8778–8791. doi: 10.1523/JNEUROSCI.17-22-08778.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burrell HR, Dokas LA, Agranoff BW. RNA metabolism in the goldfish retina during optic nerve regeneration. J Neurochem. 1978;31:289–298. doi: 10.1111/j.1471-4159.1978.tb12462.x. [DOI] [PubMed] [Google Scholar]
- Caminos E, Becker E, Martin-Zanca D, Vecino E. Neurotrophins and their receptors in the tench retina during optic nerve regeneration. J Comp Neurol. 1999;404:321–331. doi: 10.1002/(sici)1096-9861(19990215)404:3<321::aid-cne4>3.0.co;2-y. [DOI] [PubMed] [Google Scholar]
- Caroni P, Schwab ME. Two membrane protein fractions from rat central myelin with inhibitory properties for neurite growth and fibroblast spreading. J Cell Biol. 1988;106:1281–1288. doi: 10.1083/jcb.106.4.1281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen DF, Jhaveri S, Schneider GE. Intrinsic changes in developing retinal neurons result in regenerative failure of their axons. Proc Natl Acad Sci U S A. 1995;92:7287–7291. doi: 10.1073/pnas.92.16.7287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J, Ng SM, Chang C, Zhang Z, Bourdon J-C, Lane DP, Peng J. P53 isoform d113p53 is a p53 target gene that antagonizes p53 apoptotic activity via BclxL activation in zebrafish. Genes Dev. 2009;23:278–290. doi: 10.1101/gad.1761609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Graeve F, Smaldone S, Laub F, Mlodzik M, Bhat M, Ramirez F. Identification of the Drosophila progenitor of mammalian Krüppel-like factors 6 and 7 and a determinant of fly development. Gene. 2003;314:55–62. doi: 10.1016/s0378-1119(03)00720-0. [DOI] [PubMed] [Google Scholar]
- Ertürk A, Hellal F, Enes J, Bradke F. Disorganized microtubules underlie the formation of retraction bulbs and the failure of axonal regeneration. J Neurosci. 2007;27:9169–9180. doi: 10.1523/JNEUROSCI.0612-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fulton C, Simpson PA. Selective synthesis and utilization of flagellar tubulin. The multitubulin hypothesis. In: Goldman R, Pollard T, Rosenbaum J, editors. Cell Motility. Cold Spring Harbor Publications; New York: 1976. pp. 987–1005. [Google Scholar]
- Garcia-Valenzuela E, Gorczyca W, Darzynkiewicz Z, Sharma SC. Apoptosis in adult retinal ganglion cells after axotomy. J Neurobiol. 1994;25:431–438. doi: 10.1002/neu.480250408. [DOI] [PubMed] [Google Scholar]
- Gloster A, Wu W, Speelman A, Weiss S, Causing C, Pozniak C, Reynolds B, Chang E, Toma JG, Miller FD. The T alpha 1 alpha-tubulin promoter specifies gene expression as a function of neuronal growth and regeneration in transgenic mice. J Neurosci. 1994;14:7319–7330. doi: 10.1523/JNEUROSCI.14-12-07319.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldman D, Ding J. Different regulatory elements are necessary for alpha1 tubulin induction during CNS development and regeneration. Neuroreport. 2000;11:3859–3863. doi: 10.1097/00001756-200011270-00051. [DOI] [PubMed] [Google Scholar]
- Goldman D, Hankin M, Li Z, Dai X, Ding J. Transgenic zebrafish for studying nervous system development and regeneration. Transgenic Res. 2001;10:21–33. doi: 10.1023/a:1008998832552. [DOI] [PubMed] [Google Scholar]
- Gulati-Leekha A, Goldman D. A reporter-assisted mutagenesis screen using alpha 1-tubulin-GFP transgenic zebrafish uncovers missteps during neuronal development and axonogenesis. Dev Biol. 2006;296:29–47. doi: 10.1016/j.ydbio.2006.03.024. [DOI] [PubMed] [Google Scholar]
- Hermanns S, Klapka N, Muller HW. The collagenous lesion scar--an obstacle for axonal regeneration in brain and spinal cord injury. Restor Neurol Neurosci. 2001;19:139–148. [PubMed] [Google Scholar]
- Hieber V, Agranoff BW, Goldman D. Target-dependent regulation of retinal nicotinic acetylcholine receptor and tubulin RNAs during optic nerve regeneration in goldfish. J Neurochem. 1992;58:1009–1015. doi: 10.1111/j.1471-4159.1992.tb09355.x. [DOI] [PubMed] [Google Scholar]
- Hieber V, Dai X, Foreman M, Goldman D. Induction of alpha1-tubulin gene expression during development and regeneration of the fish central nervous system. J Neurobiol. 1998;37:429–440. doi: 10.1002/(sici)1097-4695(19981115)37:3<429::aid-neu8>3.0.co;2-n. [DOI] [PubMed] [Google Scholar]
- Kajimura D, Dragomir C, Ramirez F, Laub F. Identification of genes regulated by transcription factor KLF7 in differentiating olfactory sensory neurons. Gene. 2007;388:34–42. doi: 10.1016/j.gene.2006.09.027. [DOI] [PubMed] [Google Scholar]
- Keays DA, Tian G, Poirier K, Huang GJ, Siebold C, Cleak J, Oliver PL, Fray M, Harvey RJ, Molnár Z, Piñon MC, Dear N, Valdar W, Brown SD, Davies KE, Rawlins JN, Cowan NJ, Nolan P, Chelly J, Flint J. Mutations in alpha-tubulin cause abnormal neuronal migration in mice and lissencephaly in humans. Cell. 2007;128:45–57. doi: 10.1016/j.cell.2006.12.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khodiyar VK, Maltais LJ, Sneddon KM, Smith JR, Shimoyama M, Cabral F, Dumontet C, Dutcher SK, Harvey RJ, Lafanechère L, Murray JM, Nogales E, Piquemal D, Stanchi F, Povey S, Lovering RC. A revised nomenclature for the human and rodent alpha-tubulin gene family. Genomics. 2007;90:285–289. doi: 10.1016/j.ygeno.2007.04.008. [DOI] [PubMed] [Google Scholar]
- Kratz E, Eimon PM, Mukhyala K, Stern H, Zha J, Strasser A, Hart R, Ashkenazi A. Functional characterization of the Bcl-2 gene family in the zebrafish. Cell Death Diff. 2006;13:1631–1640. doi: 10.1038/sj.cdd.4402016. [DOI] [PubMed] [Google Scholar]
- Laub F, Aldabe R, Ramirez F, Friedman S. Embryonic expression of Krüppel-like factor 6 in neural and non-neural tissues. Mech Dev. 2001a;106:167–170. doi: 10.1016/s0925-4773(01)00419-1. [DOI] [PubMed] [Google Scholar]
- Laub F, Aldabe R, Friedrich V, Jr, Ohnishi S, Yoshida T, Ramirez F. Developmental expression of mouse Krüppel-like transcription factor KLF7 suggests a potential role in neurogenesis. Dev Biol. 2001b;233:305–318. doi: 10.1006/dbio.2001.0243. [DOI] [PubMed] [Google Scholar]
- Laub F, Lei L, Sumiyoshi H, Kajimura D, Dragomir C, Smaldone S, Puche AC, Petros TJ, Mason C, Parada LF, Ramirez F. Transcription factor KLF7 is important for neuronal morphogenesis in selected regions of the nervous system. Mol Cell Biol. 2005;25:5699–5711. doi: 10.1128/MCB.25.13.5699-5711.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lei L, Laub F, Lush M, Romero M, Zhou J, Luikart B, Klesse L, Ramirez F, Parada LF. The zinc finger transcription factor Klf7 is required for TrkA gene expression and development of nociceptive sensory neurons. Genes Dev. 2005;19:1354–1364. doi: 10.1101/gad.1227705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu ZW, Matsukawa T, Arai K, Devadas M, Nakashima H, Tanaka M, Mawatari K, Kato S. Na, K-ATPase alpha3 subunit in the goldfish retina during optic nerve regeneration. J Neurochem. 2002;80:763–770. doi: 10.1046/j.0022-3042.2002.00771.x. [DOI] [PubMed] [Google Scholar]
- Matsumoto N, Kubo A, Liu H, Akita K, Laub F, Ramirez F, Keller G, Friedman SL. Developmental regulation of yolk sac hematopoiesis by Kruppel-like factor 6. Blood. 2006;107:1357–1365. doi: 10.1182/blood-2005-05-1916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller FD, Naus CC, Durand M, Bloom FE, Milner RJ. Isotypes of alpha-tubulin are differentially regulated during neuronal maturation. J Cell Biol. 1987;105:3065–3073. doi: 10.1083/jcb.105.6.3065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller FD, Tetzlaff W, Bisby MA, Fawcett JW, Milner RJ. Rapid induction of the major embryonic alpha-tubulin mRNA, T alpha 1, during nerve regneration in adult rats. J Neurosci. 1989;9:1452–1463. doi: 10.1523/JNEUROSCI.09-04-01452.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moore DL, Blackmore MG, Hu Y, Kaestner KH, Bixby JL, Lemmon VP, Goldberg JL. KLF family members regulate intrinsic axon regeneration ability. Science. 2009;326:298–301. doi: 10.1126/science.1175737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narla G, Heath KE, Reeves HL, Li D, Giono LE, Kimmelman AC, Glucksman MJ, Narla J, Eng FJ, Chan AM, Ferrari AC, Martignetti JA, Friedman SL. KLF6, a candidate tumor suppressor gene mutated in prostate cancer. Science. 2001;294:2563–2566. doi: 10.1126/science.1066326. [DOI] [PubMed] [Google Scholar]
- Nilsson A, Moller K, Dahlin L, Lundborg G, Kanje M. Early changes in gene expression in the dorsal root ganglia after transection of the sciatic nerve; effects of amphiregulin and PAI-1 on regeneration. Brain Res Mol Brain Res. 2005;136:65–74. doi: 10.1016/j.molbrainres.2005.01.008. [DOI] [PubMed] [Google Scholar]
- Nordeen SK. Luciferase reporter gene vectors for analysis of promoters and enhancers. Biotechniques. 1988;6:454–458. [PubMed] [Google Scholar]
- Plunet W, Kwon BK, Tetzlaff W. Promoting axonal regeneration in the central nervous system by enhancing the cell body response to axotomy. J Neurosci Res. 2002;68:1–6. doi: 10.1002/jnr.10176. [DOI] [PubMed] [Google Scholar]
- Raff EC. Genetics of microtubule systems. J Cell Biol. 1984;99:1–10. doi: 10.1083/jcb.99.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reeves HL, Narla G, Ogunbiyi O, Haq AI, Katz A, Benzeno S, Hod E, Harpaz N, Goldberg S, Tal-Kremer S, Eng FJ, Arthur MJ, Martignetti JA, Friedman SL. Kruppel-like factor 6 (KLF6) is a tumor-suppressor gene frequently inactivated in colorectal cancer. Gastroenterology. 2004;126:1090–1103. doi: 10.1053/j.gastro.2004.01.005. [DOI] [PubMed] [Google Scholar]
- Schwab ME. Structural plasticity of the adult CNS. Negative control by neurite growth inhibitory signals. Int J Dev Neurosci. 1996;14:379–385. [PubMed] [Google Scholar]
- Senut MC, Gulati-Leekha A, Goldman D. An element in the alpha1-tubulin promoter is necessary for retinal expression during optic nerve regeneration but not after eye injury in the adult zebrafish. J Neurosci. 2004;24:7663–7673. doi: 10.1523/JNEUROSCI.2281-04.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuermer CA, Bastmeyer M, Bahr M, Strobel G, Paschke K. Trying to understand axonal regeneration in the CNS of fish. J Neurobiol. 1992;23:537–550. doi: 10.1002/neu.480230508. [DOI] [PubMed] [Google Scholar]
- Suske G, Bruford E, Philipsen S. Mammalian SP/KLF transcription factors: bring in the family. Genomics. 2005;85:551–556. doi: 10.1016/j.ygeno.2005.01.005. [DOI] [PubMed] [Google Scholar]
- Takami T, Oudega M, Bates ML, Wood PM, Kleitman N, Bunge MB. Schwann cell but not olfactory ensheathing glia transplants improve hindlimb locomotor performance in the moderately contused adult rat thoracic spinal cord. J Neurosci. 2002;22:6670–6681. doi: 10.1523/JNEUROSCI.22-15-06670.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Udvadia AJ, Koster RW, Skene JH. GAP-43 promoter elements in transgenic zebrafish reveal a difference in signals for axon growth during CNS development and regeneration. Development. 2001;128:1175–1182. doi: 10.1242/dev.128.7.1175. [DOI] [PubMed] [Google Scholar]
- Veldman MB, Bemben MA, Thompson RC, Goldman D. Gene expression analysis of zebrafish retinal ganglion cells during optic nerve regeneration identifies KLF6a and KLF7a as important regulators of axon regeneration. Dev Biol. 2007;312:596–612. doi: 10.1016/j.ydbio.2007.09.019. [DOI] [PubMed] [Google Scholar]
- Walke W, Xiao G, Goldman D. A dual function activity-dependent, muscle-specific enhancer from rat nicotinic acetylcholine receptor delta-subunit gene. J Neurobiol. 1996;31:359–369. doi: 10.1002/(SICI)1097-4695(199611)31:3<359::AID-NEU8>3.0.CO;2-9. [DOI] [PubMed] [Google Scholar]
- Witte H, Bradke F. The role of the cytoskeleton during neuronal polarization. Curr Opin Neurobiol. 2008;18:479–487. doi: 10.1016/j.conb.2008.09.019. [DOI] [PubMed] [Google Scholar]
- Woolfe A, Goodson M, Goode DK, Snell P, McEwen GK, Vavouri T, Smith SF, North P, Callaway H, Kelly K, Walter K, Abnizova I, Gilks W, Edwards YJ, Cooke JE, Elgar G. Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biol. 2005;3:e7. doi: 10.1371/journal.pbio.0030007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuba-Kubo A, Kubo A, Hata M, Tsukita S. Gene knockout analysis of two g-tubulin isoforms in mice. Dev Biol. 2005;282:361–373. doi: 10.1016/j.ydbio.2005.03.031. [DOI] [PubMed] [Google Scholar]
- Zheng B, Atwal J, Ho C, Case L, He XL, Garcia KC, Steward O, Tessier-Lavigne M. Genetic deletion of the Nogo receptor does not reduce neurite inhibition in vitro or promote corticospinal tract regeneration in vivo. Proc Natl Acad Sci U S A. 2005;102:1205–1210. doi: 10.1073/pnas.0409026102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng B, Ho C, Li S, Keirstead H, Steward O, Tessier-Lavigne M. Lack of enhanced spinal regeneration in Nogo-deficient mice. Neuron. 2003;38:213–224. doi: 10.1016/s0896-6273(03)00225-3. [DOI] [PubMed] [Google Scholar]
- Zottoli SJ, Freemer MM. Recovery of C-starts, equilibrium and targeted feeding after whole spinal cord crush in the adult goldfish Carassius auratus. J Exp Biol. 2003;206:3015–3029. doi: 10.1242/jeb.00512. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Control or 2 different tuba1a-targeting MOs were delivered to RGCs, in vivo, by placing MO-soaked Gelfoam on the sectioned optic nerve stump. The next day the Gelfoam was removed and 3 days later retinas were isolated, diced and placed in explant culture for 4 days prior to analysis of neurite outgrowth. The graph on the right shows the Nerve Growth Index (see Materials and Methods) for each MO at the concentration indicated. Note the concentration dependent reduction in neurite outgrowth (Nerve Growth Index) as increasing concentrations of the tuba1a-targeting MO are applied to the optic nerve stump. Also note that tuba1a-targeting MOs #1 and #2 at 0.065 mM concentration have little effect on neurite outgrowth when applied individually, but synergize to significantly reduce neurite outgrowth when added together. The image on the left shows representative explants treated with tuba1a-targeting MOs #1 and/or #2 at 0.065mM concentration. Note the reduced axon outgrowth in explants from retinas that received a combination of MOs #1 and #2.
tuba1a-targeting MO was delivered to RGCs, in vivo, by placing MO-soaked Gelfoam on the sectioned optic nerve stump. The next day the Gelfoam was removed and 3 days later retinas were isolated, sectioned and assayed for cell death using TUNEL stain. Note that like the uninjured retina, MO-treated retinas showed little cell death. Positive control is a nuclease-treated retina. Brown signal indicates cells undergoing apoptosis.
Electrophoretic mobility shift assay showing the Sp1 consensus binding site probe can efficiently compete nuclear protein binding to the G/C-rich enhancer. Below the gel are shown the oligonucleotide sequences of consensus protein binding sites for myc-associated zinc finger protein (MAZ), GLI, specificity protein 1 (SP1), and smad interacting protein (SIP) used as unlabeled competitors. Consensus binding sites are underlined. The arrow marks specific protein binding. NS, Nonspecific binding; F, free probe.
Sp1, Sp3, and Sp4 expression is not changed during optic nerve regeneration in zebrafish. Quantitative RT-PCR for negative control ribosomal protein encoding transcript L24, positive control tuba1a (α1T), and Sp1, Sp3, and Sp4 using uninjured retinal RNA and 3 day post-optic nerve crush retinal RNA (ONC) as template. Bar graphs represent densitometric quantification of autoradiographic analysis of RT-PCR products from 3 separate fish.
(A and B) In situ hybridization showing KLF6a induction in retinal ganglion cells 3 days post-optic nerve crush (B) as compared to uninjured retina (A). (C and D) In situ hybridization showing KLF7a induction in retinal ganglion cells 3 days post-optic nerve crush (D) as compared to uninjured retina (C). (E) KLF6a is not expressed in the retina at 48 hours post fertilization (hpf). (F) KLF7a is expressed in the differentiating RGCs at 48 hpf. Arrows denote developing GCL. Outer nuclear layer (OLN), inner nuclear layer (INL), and ganglion cell layer (GCL).