Abstract
MicroRNAs (miRNAs), a class of small noncoding RNAs that regulate gene expression, have fundamental roles in biological processes, including cell differentiation and proliferation. These small molecules mainly direct either target messenger RNA (mRNA) degradation or translational repression, thereby functioning as gene silencers. Epithelial cells of the uterine lumen and glands undergo cyclic changes under the influence of the sex steroid hormones estradiol-17beta and progesterone. Because the expression of miRNAs in human endometrium has been established, it is important to understand whether miRNAs have a physiological role in modulating the expression of hormonally induced genes. The studies herein establish concomitant differential miRNA and mRNA expression profiles of uterine epithelial cells purified from endometrial biopsy specimens in the late proliferative and midsecretory phases. Bioinformatics analysis of differentially expressed mRNAs revealed cell cycle regulation as the most significantly enriched pathway in the late proliferative-phase endometrial epithelium (P = 5.7 × 10−15). In addition, the WNT signaling pathway was enriched in the proliferative phase. The 12 miRNAs (MIR29B, MIR29C, MIR30B, MIR30D, MIR31, MIR193A-3P, MIR203, MIR204, MIR200C, MIR210, MIR582-5P, and MIR345) whose expression was significantly up-regulated in the midsecretory-phase samples were predicted to target many cell cycle genes. Consistent with the role of miRNAs in suppressing their target mRNA expression, the transcript abundance of predicted targets, including cyclins and cyclin-dependent kinases, as well as E2F3 (a known target of MIR210), was decreased. Thus, our findings suggest a role for miRNAs in down-regulating the expression of some cell cycle genes in the secretory-phase endometrial epithelium, thereby suppressing cell proliferation.
Keywords: cell cycle, endometrium, estradiol, microarray, miRNA, progesterone
Global miRNA and mRNA expression arrays demonstrate distinct gene expression signatures in human late proliferative-phase and midsecretory-phase endometrial epithelium.
INTRODUCTION
Human endometrium undergoes cyclic changes regulated by the female sex steroid hormones estradiol-17β (E2) and progesterone (P4). E2 elicits a wave of uterine epithelial cell proliferation, whereas P4 inhibits this E2-induced epithelial cell proliferation, promotes differentiation, and has decidualizing effects on endometrial stroma later in the secretory phase. E2 and P4 act through their cognate nuclear receptor transcription factors, estrogen receptor 1 and P4 receptor. Results of experiments using tissue recombinants of uterine epithelium and stroma, whose estrogen status differ because of inactivating mutation, suggest that E2 stimulates uterine epithelial cell proliferation through epithelial estrogen receptor 1 (ESR1) [1]. This contrasts with the situation in mice, in which E2 acts on epithelial cell proliferation via paracrine influences downstream of E2 binding to stromal ESR1 and E2-induced uterine epithelial cell proliferation is inhibited by P4 actions mediated via P4 receptor in stroma [1, 2]. Coordinated and synchronized action of E2 and P4 is essential for controlled proliferation of endometrium and for uterine receptivity at the time of implantation. Identification of molecular mechanisms involved in normal hormonal regulation of human endometrium is therefore an important step toward understanding molecular deregulation occurring in pathological situations and may expose novel therapeutic opportunities.
E2 elicits rapid responses in human and mouse uterus by activating the so-called canonical cell cycle pathway [2–4]. In mouse uterus, paracrine signaling through the epithelially expressed insulin-like growth factor 1 receptor leads to the activation of the phosphoinositide 3-kinase pathway, followed by AKT activation that results in inhibitory phosphorylation of glycogen synthase kinase 3 beta (GSK3B), thereby allowing accumulation of cyclin D1 in the nucleus, retinoblastoma protein phosphorylation, and progress of cells through the restriction point into S-phase [2, 4, 5]. In addition, parallel activation of DNA replication licensing involving the minichromosome maintenance proteins (MCMs) is necessary for cell cycle progression into S-phase in mouse endometrial endometrium [6]. However, in the more complex regenerative human endometrium, molecular mechanisms underlying its cyclic changes are not as clearly elucidated. Our previous investigations on human endometrial biopsy specimens demonstrated increased epithelial cell expression of markers of proliferation (antigen identified by monoclonal antibody Ki-67 [MKI67], proliferating cell nuclear antigen [PCNA], cyclin A1 [CCNA], and cyclin E [CCNE]) during the proliferative phase of the menstrual cycle compared with low expression in the secretory-phase epithelium [7]. Further inhibition of GSK3B by lithium chloride in reconstructed human endometrial biopsy specimens xenotransplanted into immunocompromised mice resulted in epithelial cell entry into S-phase [8]. These results provide evidence that similar cell cycle regulators are activated by E2 and inhibited by P4 in human endometrium as in mouse [7].
miRNAs, a class of small noncoding RNAs, function as posttranscriptional regulators of gene expression. They bind through complementary base pairing to the 3′ untranslated regions of their target mRNAs, leading to mRNA degradation or translational repression [9]. The 5′ sequences of miRNAs, particularly those at nucleotide positions 2 through 7 relative to the 5′ end of miRNA, are important for binding to the target. Individual targets of miRNAs responsible for the phenotypes have been proposed in experimental settings, although it is likely that many miRNAs function through cooperative regulation of multiple mRNAs [10].
While it is established that miRNAs are expressed in human endometrium [11, 12], hormonal regulation of miRNAs in the function of human endometrial epithelium and their role in differentiation of endometrium into its receptive state remain unknown. miRNAs have been reported to regulate many cellular processes that are known to occur during cyclic changes in endometrium such as cell proliferation and differentiation [13]. Notably, aberrant miRNA expression has been associated with human endometrial disorders such as endometriosis, endometrial hyperplasia, and carcinoma [11, 12, 14, 15]. In addition, miRNAs were recently described as having a role in postnatal development of mouse uterus and oviducts [16–18] and in mouse embryo implantation [16, 19, 20]. Taken together, results of previous human and mouse studies strongly suggest that in human endometrium miRNAs are likely to have a regulatory function during the physiological cycle phases.
In the present study, high-density gene expression arrays were utilized to identify specific miRNAs involved in hormonal regulation of normal human endometrium by E2 and P4. Well-characterized endometrial biopsy samples were obtained from eight healthy midreproductive-aged women, and simultaneous mRNA and miRNA profiles were established for endometrial epithelial cell preparations either during the late proliferative phase or midsecretory phase of the menstrual cycle. The rationale for examining endometrial epithelium during the time of maximum proliferation (late proliferative phase) and maximum P4 action (midsecretory phase) was to elucidate the action of P4 in opposing the effects of estrogen. The expression profiling of mRNAs and miRNAs in the same samples allowed us to identify specific mRNAs that may be posttranscriptionally repressed by miRNAs in human endometrial epithelium and to determine some cellular functions and molecular pathways targeted by these differentially expressed miRNAs.
MATERIALS AND METHODS
General Participants
This study was approved by The Clinical Investigations/Institutional Review Board at Albert Einstein College of Medicine and was conducted in accord with the Declaration of Helsinki for medical research involving human subjects at the General Clinical Research Center. Written informed consent was obtained from all participants of this study.
Participants were recruited from the community and were healthy volunteer women aged 18 through 36 yr with no history of infertility who met the following criteria: 1) regular 25- to 35-day menstrual cycles, 2) no use of hormonal contraception within 3 mo, and 3) at least 90% normal weight for height [21]. Screening done solely for study purposes included history and physical examination, negative cervical cytology, negative urine pregnancy test, and normal saline hysterosonogram to rule out any intrauterine pathology. Sixteen women attended the General Clinical Research Center for endometrial biopsy, and 14 biopsies were performed. Endometrial biopsy specimens from eight women, four late proliferative phase and four midsecretory phase, met the criteria and were analyzed during the course of the study. Characteristics of the study participants are given in Supplemental Table S1 (available at www.biolreprod.org).
Endometrial Biopsy Protocol
Endometrial biopsy samplings from the fundal area, as outlined previously [7], were performed using a Pipelle catheter (Unimar, Inc., Wilton, CT). Portions of tissues were saved in formalin, 10% vol/vol solution, for hematoxylin-eosin staining and for histological evaluation. The main portion of the tissues was further processed for endometrial epithelial cell isolation.
Late proliferative-phase biopsies were performed on the mean ± SEM cycle day (CD) 12 ± 1 to target the time of maximal endometrial response to E2 [22]. Serum E2 and P4 samples were obtained on the day of the biopsy; the sample was excluded if the P4 level was at least 3.5 ng/ml, indicative of ovulation [23]. To target the endometrial window of receptivity and maximum P4 action, secretory biopsy specimens were obtained during the midluteal phase on CDs 19 through 23. Midsecretory phase was confirmed after biopsy by serum P4 level of at least 3.5 ng/ml and by histology [23, 24]. Histological evaluation of hematoxylin-eosin-stained tissue specimens was performed blindly by two persons, and cycle phases were assigned according to published criteria [24]. Histological dating was correlated with CD and serum E2 and P4 levels on the day of endometrial biopsy. Dating of all biopsy specimens was within a 2-day window from the CD as calculated from the last menstrual period.
Serum Hormone Assays
Fluoroimmunoassays were used to measure serum E2 and P4 levels (Wallac; PerkinElmer Life and Analytical Sciences, Turku, Finland). This technique was described previously [7, 25].
Epithelial Cell Isolation
After sampling, endometrial biopsy specimens were transported to the laboratory in McCoy modified 5A medium on ice. Isolation of epithelial cells from stromal cells was performed according to the modified protocol by Satyaswaroop et al. [26]. Briefly, after removal of blood and mucus, the endometrial biopsy specimen was minced into 1-mm pieces. Minced pieces were digested with type I collagenase (230 U/mg; Worthington, Lakewood, NJ) in McCoy modified 5A medium containing 20 000 IU/ml of DNase I (Sigma, St. Louis, MO) at 37°C for 1 h. Collagenase-treated cells were centrifuged and suspended into RNase-free PBS. The suspension was strained through an 80-μm nylon mesh, in which epithelial cell microaggregates remain in the mesh, whereas stromal cells and red blood cells filter through. Isolated epithelial cells were then washed with RNase-free PBS, centrifuged, and stored in TRIzol reagent (Invitrogen, Carlsbad, CA) at −80°C. Purity of the epithelial cell preparation was determined by intracellular immunohistochemistry (IHC) using anti-Pan cytokeratin antibody (clone PCK-26; Sigma) on an isolated epithelial cell preparation. Epithelial cells were attached and fixed with ethanol onto Superfrost Plus slides (Thermo Fisher Scientific, Waltham, MA). The fixative was rinsed off with PBS, and slides were incubated with Triton X-100, 0.3% vol/vol solution, in PBS for 10 min to permeabilize the membranes, followed by IHC. The stained cells were observed under light microscopy. The number of cells showing cytoplasmic cytokeratin staining (only epithelial cells [and not stromal cells] stain positive for cytokeratin) indicated that the isolation procedure yielded a greater than 90% pure endometrial epithelial cell preparation (data not shown).
RNA Extraction
Total RNA and miRNA-enriched RNA were extracted from isolated uterine epithelial cells using the miRVana miRNA Isolation Kit (Ambion Inc., Austin, TX) according to the manufacturer's protocol. RNA yield and quality were assessed by electrophoresis using the Agilent Technologies Inc. (Santa Clara, CA) Bioanalyzer 2100 and spectrophotometric analysis.
Sample and Microarray Processing, Human Genome U133 Plus 2.0 Array
Eight endometrial epithelial cell samples were used for microarray analysis. Forty nanograms of total RNA from each sample was used to generate cDNA, which was amplified using Ovation RNA Amplification version 2 (product 3100; NuGen Inc., San Carlos, CA). The resulting cDNA was purified using Research DNA Clean Concentration 25 (Zymo Research Inc., Orange, CA), and the cDNA quality was assessed using the Bioanalyzer 2100. Following cDNA amplification and purification, the product was fragmented (50–100 bp) and labeled with biotin using FL-Ovation cDNA Biotin Module version 2 (product 4200; NuGen Inc.) in preparation for hybridization to Affymetrix (Santa Clara, CA) GeneChip. We used 3.75 μg of amplified and labeled cDNA in the hybridization cocktail for GeneChip analysis. Detailed protocols for sample preparation using Ovation RNA Amplification and labeling protocols can be found at http://www.nugeninc.com. All samples were subjected to gene expression analysis using Affymetrix human genome U133 Plus 2.0 arrays. Hybridization, staining, and washing of all arrays were performed in the Affymetrix fluidics module. Streptavidin-phycoerythrin stain (SAPE; Molecular Probes, Eugene, OR) is a fluorescent conjugate used to detect hybridized target sequences. Detection and quantification of target hybridization were performed using an Affymetrix GeneChip scanner. Data were assessed for array performance before analysis. The probe-level intensities were processed using the robust multiarray average method for background correction, normalization, and log2 transformation of perfect match values [27].
Microarray Data Analysis
mRNA microarray data were analyzed using ArrayAssist 5.0.1 microarray data analysis software (Stratagene, La Jolla, CA). Data were log2 transformed. Two-tailed Student t-test was performed to compare the proliferative-phase and midsecretory-phase groups for each transcript, and fold change was determined. Raw P values were adjusted using the Benjamini-Hochberg false discovery rate to yield adjusted P values. The criteria for significance of differentially regulated genes were established as a greater than 2-fold change with an adjusted P < 0.05. We performed unsupervised two-way hierarchical clustering analysis (Pearson-centered distant metrics and centroid linkage rule) on eight samples in each data set with all probe sets included.
Sample and Microarray Processing, miRCHIP V1 Array
A custom-manufactured Affymetrix GeneChip from Ambion Inc. was designed for miRNA probes derived from miRBase (http://www.mirbase.org) and published reports by Asuragen (Austin, TX) [28–31]. Antigenomic probe sequences were provided by Affymetrix and were derived from a larger set of controls used on the Affymetrix human exon array for estimating background signal, as described herein. Other non-miRNA control probes on the array were designed to lack sequence to the human genome and can be used for spike-in external reference controls.
Samples for miRNA profiling studies were processed by Asuragen according to the company's standard operating procedures. miRNA-enriched RNA samples were obtained using the miRVana miRNA Isolation Kit and were provided to Asuragen. The 3′ ends of the RNA molecules were tailed and biotin labeled using the miRVana miRNA Labeling Kit (Ambion Inc.). The kit's deoxyribonucleotide triphosphate mixture in the tailing reaction was replaced with a proprietary mixture containing biotin-modified nucleotides (PerkinElmer, Waltham, MA). Hybridization, washing, staining, imaging, and signal extraction were performed according to Affymetrix-recommended procedures except that the 20× GeneChip eukaryotic hybridization control cocktail was omitted from the hybridization. The signal processing implemented for the Ambion Inc. miRCHIP V1 array was a multistep process and involved probe-specific signal detection calls, background estimates, and correction. For each probe, an estimated background value was subtracted that was derived from the median signal of a set of GC content-matched antigenomic controls. Arrays within a specific experiment were normalized together according to the variance stabilization method described by Huber et al. [32]. Detection calls were based on Wilcoxon rank sum test of miRNA probe signal compared with the distribution of signals from GC content-matched antigenomic probes. For statistical hypothesis testing, two-sample t-test with the assumption of equal variance was applied, and this test defined which probes were considered significantly differentially expressed based on cutoff values of P < 0.05 and a greater than 2-fold or less than 2-fold difference in expression. To reduce the false discovery rate of miRNAs, we excluded miRNAs whose expression was detected in fewer than three of four specimens in either the late proliferative-phase group or the midsecretory-phase group. We performed unsupervised two-way hierarchical clustering analysis (Pearson-centered distant metrics and centroid linkage rule) on eight samples in each data set with differentially expressed miRNAs (>1.5-fold or <1.5-fold difference in expression with P < 0.05). A heatmap was generated, and the dendrogram shows relationships between specimens.
Quantitative Real-Time PCR
One microgram of total RNA from each sample was used for the RT reaction to generate cDNA using SuperScript II RT (Invitrogen) and random hexamers. Quantitative real-time PCR was performed in triplicate using SYBR Green PCR Master Mix (Applied Biosystems, Norwalk, CT) according to the manufacture's instructions and using an ABI Prism 7900HT (Amersham-Pharmacia, Piscataway, NJ). The PCR primers were designed to be intron spanning and to amplify 68- to 300-bp fragments (Supplemental Table S2). mRNA expression of TGF3B was analyzed using TaqMan assay (Applied Biosystems). The data were normalized to expression levels of the housekeeping gene GAPDH, and the relative expression was calculated using the 2−ΔΔCT method [33].
TaqMan miRNA assays (Applied Biosystems) were used to quantify the expression of miRNAs according to the manufacturer's instructions. The mean cycle threshold (CT) value was determined from four PCR replicates. The data were normalized to expression levels of SNORD48 (also known as RNU48). Relative gene expression of the late proliferative-phase vs. midsecretory-phase samples was assessed using the 2−ΔΔCT method [33].
Bioinformatics
The Database for Annotation, Visualization and Integrated Discovery (DAVID 2008; http://david.abcc.ncifcrf.gov) and Ingenuity Pathway Analysis software (IPA 7.6; Ingenuity Systems, Redwood City, CA) were used to identify enriched cellular and molecular functions among differentially expressed gene transcripts. DAVID 2008 and the Gene Ontology project (http://www.geneontology.org) were used to classify differentially expressed genes into functionally related groups of genes. Target genes of miRNAs were predicted using the following algorithms: miRanda (http://www.ebi.ac.uk/ena/), PicTar (http://pictar.mdc-berlin.de/), and TargetScan (http://www.targetscan.org).
RESULTS
Endometrial biopsy specimens were obtained from eight midreproductive-aged women. The ages of these women ranged from 19 to 36 yr (mean age, 29.0 yr) in the late proliferative-phase group and from 22 to 30 yr (mean age, 24.3 yr) in the midsecretory-phase group. The mean ± SEM serum E2 levels were 170.43 ± 36.17 pg/ml in the late proliferative-phase group and 121.55 ± 43.56 pg/ml in the midsecretory-phase group. The mean ± SEM serum P4 levels were 0.39 ± 0.19 ng/ml in the late proliferative-phase group and 12.20 ± 1.20 ng/ml in the midsecretory-phase group. General data on the women in the study are given in Supplemental Table S1.
Late Proliferative-Phase and Midsecretory-Phase Endometrial Epithelia Exhibit Distinct miRNA and mRNA Expression Profiles
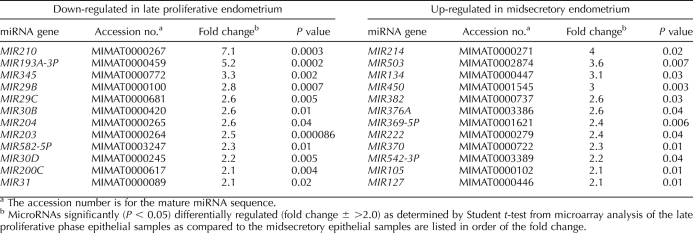
To establish miRNA and mRNA profiles of endometrial epithelium, microarray analyses were performed on sets of four late proliferative-phase and four midsecretory-phase epithelial samples of endometrium. To identify the most informative set of differentially expressed genes between the late proliferative-phase and midsecretory-phase groups, we ranked each gene by the probability that the mean of its expression values is statistically distinct between the two groups using Student t-test. We focused our attention on genes meeting our designated criteria of P < 0.05 and a greater than 2-fold or less than 2-fold change. In this fashion, we identified 3244 differentially expressed mRNAs between the late proliferative-phase vs. midsecretory-phase epithelium of endometrium; 2206 genes were up-regulated, and 1038 genes were down-regulated (Supplemental Fig. S1). Among human miRNAs on miRCHIP V1, 49 microarray probes were differentially expressed between the two groups (24 are published miRNAs, and 25 represent novel predicted miRNA sequences). The transcript abundance of 12 published miRNAs was increased and of 12 published miRNAs decreased in the late proliferative-phase vs. midsecretory-phase epithelial samples (Table 1).
TABLE 1.
MicroRNAs differentially expressed in the late proliferative phase compared to the midsecretory phase endometrial epithelium.
To visually assess differentially expressed gene profiles, we performed unsupervised hierarchical clustering analysis separately for mRNAs and miRNAs using eight well-characterized endometrial epithelial samples. Dendrograms show complete segregation of the late proliferative-phase and midsecretory-phase samples into two groups based on their mRNA (Fig. 1A) and miRNA (Fig. 1B) expression patterns. Notably, all four late proliferative-phase samples cluster robustly together, as do the midsecretory-phase samples, even when the expression data from all of the more than 48 000 probes of the human genome U133 Plus 2.0 array are utilized for the analysis. These data suggest that the late proliferative-phase endometrial epithelium exhibits unique mRNA and miRNA expression signatures compared with the midsecretory-phase endometrial epithelium.
FIG. 1.
Dendrogram and unsupervised hierarchical clustering. Expression data from all mRNA probes (A) and expression data from differentially expressed miRNAs showing a greater than 1.5-fold or less than 1.5-fold change in expression with P < 0.05 (B) in four late proliferative-phase (pp) and four midsecretory-phase (sp) samples. The top dendrogram shows the relationship between the samples based on gene expression patterns. Hierarchical clustering analyses robustly separate the late proliferative-phase samples from the midsecretory-phase samples and assign each sample to the correct menstrual cycle phase. The expression intensity of each gene in each sample varies from high (red) to low (green).
Late Proliferative-Phase Endometrial Epithelium Expresses Increased Transcript Abundance of Cell Cycle Regulators
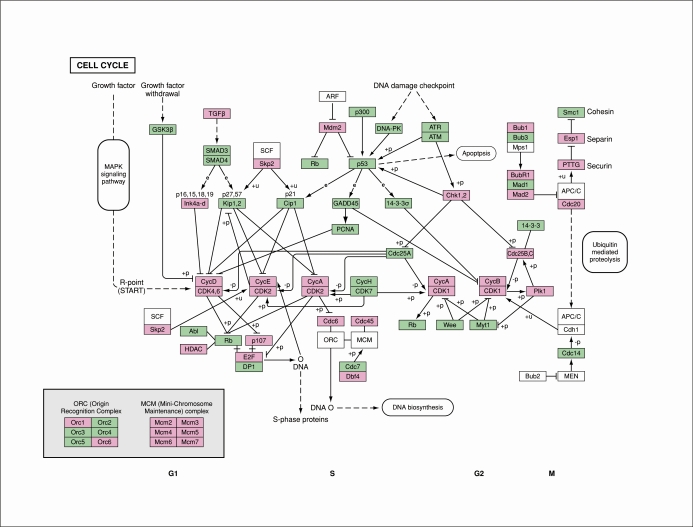
Next, we analyzed the transcriptome of the late proliferative-phase endometrial epithelium using the DAVID 2008 bioinformatics program. Genes that showed significantly increased transcript abundance in the late proliferative-phase samples were enriched for the following functional Gene Ontology groups: cell cycle (Gene Ontology enrichment score, 25.32; P = 7.5 × 10−29), cell cycle phase, M-phase, mitosis, cell division, regulation of cell cycle, and DNA replication. Also, bioinformatics analysis revealed cell cycle regulation as the most significantly enriched functional pathway in the late proliferative-phase endometrial epithelium (KEGG pathway [http://www.genome.jp/kegg/pathway.html] cell cycle, P = 5.7 × 10−15) (Fig. 2) [34, 35]. These results are in agreement with a previous global gene expression study [36] of the proliferative-phase vs. secretory-phase total endometrium. Differentially expressed transcripts were uploaded to the IPA 7.6 database to explore for enriched biological functions and pathways. As previously reported in investigations using whole endometrial specimens [36], the biological functions of cell cycle, DNA replication, recombination and repair, and cellular growth and proliferation were differentially regulated between late proliferative-phase and midsecretory-phase endometrial epithelium (Fig. 3). The enriched canonical pathways were mitotic roles of polo-like kinases, WNT/β-catenin and sonic hedgehog signaling, cell cycle regulation by the B-cell translocation gene (BTG) proteins, and checkpoint cell cycle regulation (G1/S and G2/M).
FIG. 2.
Transcripts regulating the mammalian cell cycle are up-regulated in the proliferative-phase endometrial epithelium. Genes with a greater than 2-fold increase in transcript abundance with P < 0.05 in the late proliferative-phase samples compared with the midsecretory-phase samples from the human genome U133 Plus 2.0 arrays are indicated in purple boxes and represent 38 up-regulated genes. Green and white boxes indicate cell cycle genes that showed no differential expression between the late proliferative-phase samples compared with the midsecretory-phase samples. The diagram is based on the cell cycle pathway in the KEGG pathway database [34, 35].
FIG. 3.

Transcripts mediating cell cycle regulation are differentially regulated between the late proliferative-phase vs. midsecretory-phase endometrial epithelium. Differentially expressed transcripts were subjected to bioinformatics analysis using IPA 7.6. A) Most significantly enriched groups relating to molecular and cellular functions. B) Most significantly enriched canonical signaling pathways. The total number of genes in a pathway is boldfaced. For each pathway, the percentage of genes that are of decreased (green) or increased (red) transcript abundance is indicated at the top. Portions of pathway genes that are not significantly changed are also shown (white). The yellow square for each pathway indicates –log(P value). The yellow arrow indicates P = 0.05.
miRNA Gene Signatures of Late Proliferative-Phase and Midsecretory-Phase Endometrial Epithelia
We then explored the specific miRNAs that showed differential expression between the late proliferative-phase and midsecretory-phase epithelial cell samples. As summarized in Table 1, the expression of several miRNAs was altered in epithelial cells of endometrium between the two physiological phases of the menstrual cycle. Among miRNAs with lower transcript level in the late proliferative-phase samples than in the midsecretory-phase samples, MIR210 and MIR193A-3P were the most highly decreased by microarray analysis. The other notable miRNAs in this list are MIR29B and MIR29C, which belong to the MIR29 miRNA cluster, as well as MIR30B and MIR30D, which belong to the MIR30 miRNA cluster. Among significantly up-regulated late proliferative-phase miRNAs were MIR503 and MIR450, which belong to the MIR542-3P miRNA cluster.
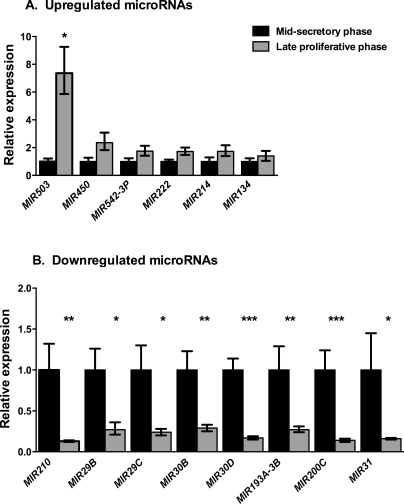
Using quantitative real-time PCR, we validated the expression of select miRNAs. As shown in Figure 4, the expression level of MIR210, MIR29B, MIR29C, MIR30B, MIR30D, MIR193A-3P, MIR200C, and MIR31 was significantly decreased in the late proliferative-phase vs. midsecretory-phase endometrial epithelium, thus validating our miRCHIP V1 data. Among up-regulated miRNAs, the expression of MIR503 showed significantly increased expression in the late proliferative-phase samples than in the midsecretory-phase samples. The expression levels of MIR405, MIR542-3P, MIR214, and MIR134 did not reach statistical significance, although the trend of increased expression level in the late proliferative-phase vs. midsecretory-phase endometrial epithelium was consistent with the microarray data. Surprisingly, MIR214 demonstrated the highest fold change expression between the two groups by microarray analysis, but quantitative real-time PCR did not validate this result. MIR222 showed 2.4-fold and 1.7-fold increases in the transcript abundance by miRNA microarray and quantitative real-time PCR, respectively.
FIG. 4.
Expression of selected up-regulated (A) and down-regulated (B) miRNAs in the late proliferative-phase endometrial samples relative to the midsecretory-phase endometrial samples by quantitative real-time PCR. All samples were normalized to RNU48. Relative gene expression of the late proliferative-phase vs. midsecretory-phase samples was assessed using the 2−ΔΔCT method [33]. Data shown indicate relative expression of the late proliferative-phase samples (gray bar) with respect to the midsecretory-phase samples (black bars) set to 1. The error bars show the SEM. *P < 0.05, **P < 0.01, and ***P < 0.001.
Cell Cycle Genes Are Targeted by miRNAs Exhibiting Increased Transcript Abundance in Midsecretory-Phase Endometrial Epithelium
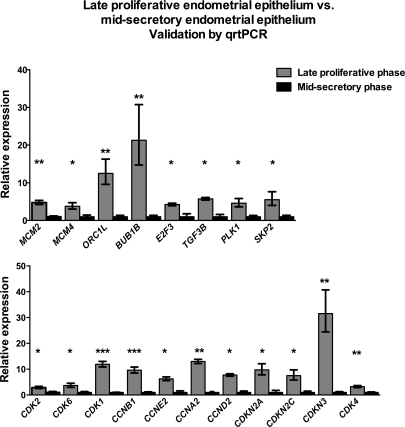
For further study, we investigated miRNAs whose transcript abundance was increased in the midsecretory-phase samples compared with the late proliferative-phase samples, as these miRNAs are likely to negatively modulate the expression of cell cycle regulators. Using bioinformatics online sites (miRanda, TargetScan, and PicTar), we identified predicted target mRNAs of these specific miRNAs. Predicted targets were then further characterized using Gene Ontology analysis (DAVID 2008) to identify targets that are functionally involved in the cell cycle pathway (KEGG pathway). Table 2 gives predicted cell cycle targets of each miRNA. With the mRNA expression results, it was striking to note that the transcript abundance of 19 predicted cell cycle genes was decreased in the midsecretory-phase samples by the mRNA array data, as would be expected if their expression was modulated by the respective miRNAs. Quantitative real-time PCR of these 19 cell cycle genes robustly validated results from the array (Fig. 5). We previously reported increased transcript abundance of cyclin A, cyclin E, and MCM2 in the late proliferative-phase vs. midsecretory-phase laser capture microdissected glandular epithelial samples, thus validating the expression of these transcripts in independent biological repeats [7].
TABLE 2.
MicroRNAs differentially expressed in the late proliferative phase compared to the midsecretory phase endometrial epithelium.
FIG. 5.
Quantitative real-time PCR validates gene expression array data from the cell cycle regulation ontological group. Nineteen genes were validated as differentially regulated cell cycle members from the human genome U133 Plus 2.0 array using quantitative real-time PCR. All samples were normalized to the housekeeping gene GAPDH. Data shown indicate relative expression of the late proliferative-phase samples (gray bar) with respect to the midsecretory-phase samples (black bars) set to 1. The error bars show the SEM. ***P < 0.001, **P < 0.01, and *P < 0.05.
DISCUSSION
In the present study, we demonstrate that the mRNA transcript profile of the late proliferative-phase endometrial epithelium is enriched for genes involved in cell cycle regulation. Moreover, we show that miRNAs are differentially expressed during the physiological phases of the menstrual cycle, suggesting that they are hormonally regulated in human endometrial epithelium. miRNAs that show increased transcript abundance during the midsecretory phase are predicted to target several cell cycle regulators. Consistent with the miRNA data, the transcript levels of several of these cell cycle regulators are lower during the midsecretory phase than during the late proliferative phase. These results suggest that miRNAs may posttranscriptionally down-regulate the expression of cell cycle genes, thereby suppressing cell cycle progression and cell proliferation in human secretory-phase endometrial epithelium.
Our global mRNA and miRNA microarray analysis yielded specific gene expression signatures for the late proliferative-phase and midsecretory-phase epithelial samples. To our knowledge, this is the first study to demonstrate differential expression profiles of miRNAs in human endometrial epithelium according to menstrual cycle phases in midreproductive-aged women who had been extensively screened for hormonal status and absence of intrauterine pathologies. Grouping of endometrial samples based solely on their mRNA expression profiles and histological menstrual cycle stage has been previously described for entire endometrial samples, including epithelium and stroma [36]. Consistent with previous expression profiles of whole endometrium [36, 38], our present data define the proliferative-phase transcriptome as one that is characterized by cell cycle regulatory functions reflecting the active mitogenic state of cells. Furthermore, two separate pathway analysis programs identified cell cycle regulation as the most significantly enriched pathway. Also, it is notable that our findings of increased transcript levels of DNA replication licensing members (such as MCM2, MCM3, MCM4, MCM5, MCM6, MCM7, and origin recognition complex proteins) in the late proliferative-phase samples are in agreement with prior expression results [39]. These data suggest that, in addition to cell cycle regulation, DNA replication licensing is likely another key regulatory point in hormonal regulation of epithelial cell proliferation in human endometrium, as previously demonstrated in mouse endometrium [6]. Furthermore, bioinformatics analysis of the mRNA array data indicates that WNT/β-catenin signaling, previously implicated in hormonal regulation of the proliferative-phase and secretory-phase endometrium [36, 38], is a significantly regulated canonical pathway in endometrial epithelium across the menstrual cycle. Specifically, we observed up-regulation of WNT ligands (WNT5A and WNT7A) and lymphoid enhance binding factor 1 (LEF1) in the late proliferative-phase vs. midsecretory-phase endometrial epithelium, whereas the transcript abundance of WNT inhibitors dikkopf homolog (DKK1) and forkhead box O1 (FOXO1) was increased in the midsecretory-phase epithelial cells. Notably, using in situ hybridization, Tulac et al. [40] have shown that WNT7A mRNA expression is restricted exclusively to luminal epithelium of human endometrium. Furthermore, using WNT-activated Ishikawa cells, Wang et al. [41] recently demonstrated that P4 induction of DKK1 and FOXO1 resulted in inhibition of WNT signaling. While WNT signaling is a well-known pathway in hormonal regulation of human endometrium based on gene expression data [36, 38] and from investigations of mouse uterus [1, 42], we recognize that molecular mechanisms of sex steroid regulation remain obscure. Because these are secreted molecules, they may mediate epithelial and stromal interactions, mediating hormonal effects on endometrial cells.
Most important, specific miRNAs with increased transcript abundance in the midsecretory-phase samples were predicted to target several genes involved in DNA replication licensing and in cell cycle regulation (Table 2). For example, key cell cycle regulators, cyclins and their partners cyclin-dependent kinases (CDKs), were predicted to be targeted by MIR31, MIR29B, MIR29C, MIR30B, and MIR30D, whereas the expression of MCM2 was predicted to be regulated by MIR31, MIR30B, and MIR30D and that of MCM4 by MIR210 (Table 2). MIR210 has been reported to down-regulate the expression of E2F transcription factor 3 (E2F3) [37], an important transcription factor that induces the expression of cell cycle-regulated genes and promotes cell cycle progression. The transcript levels of several putative cell cycle targets were decreased in the midsecretory-phase samples, as would be expected if their expression was down-regulated by miRNAs, whereas the transcript levels of others were unchanged. For instance, cell division cycle 7 (CDC7) was predicted to be targeted by at least two miRNAs, yet its mRNA level showed no significant change between the cycle phases. However, miRNA regulation of this target may occur through translational repression, in which case only the protein level would be altered. Of note is that the same miRNAs that are putative regulators of genes involved in DNA replication licensing may also regulate cyclins and CDKs. Recently, tumor-suppressive miRNAs (MIR16 and MIR34A families) have been reported to modulate the expression of multiple cell cycle regulators [43–45]. To this end, our observations (although only based on gene expression data) perfectly fit a view of miRNA-mediated regulation of gene expression in which a single miRNA can regulate multiple genes that have related functions.
Two miRNAs, MIR221 and MIR222, are known to negatively regulate the expression of CDK inhibitor 1B (CDKN1B, also known as p27/KIP1) by suppressing translation of mRNA into protein [46]. CDKN1B is an important cell cycle inhibitor, and it executes this function by binding and inhibiting CCNE/CDK2 and CCNA/CDK2 complexes in early G1-phase [47]. Recent IHC investigations by Niklaus et al. [7] demonstrated that CDKN1B protein expression is lower in epithelial cells of human endometrium in the late proliferative phase than in the midsecretory phase. Herein, we observed increased transcript abundance of MIR222 in the late proliferative-phase samples, whereas no significant change was found in CDKN1B mRNA level, as expected. Qian et al. [48] recently reported down-regulation of MIR222 expression in endometrial stromal cells, permitting CDK inhibitor 1C (CDKN1C, also known as p57/KIP2) expression and thereby suppressing stromal cell proliferation in vitro. Increased expression of MIR221 and MIR222 has been observed in several human cancers, supporting their function as oncogenes [49–51]. Notably, aberrant MIR221 expression was recently reported in atypical endometrial hyperplasia and endometrial cancer [12]. Although further validation studies are needed, our results suggest that MIR222 may have a cell cycle regulatory role in human endometrial epithelium.
A prior study [11] explored differential expression of miRNAs in human endometrium according to menstrual cycle phases but failed to show any significant differences in miRNA profiles. Of note is that this study limited research to entire endometrium, including glandular and luminal epithelium, as well as stroma, as the study was primarily designed to explore the role of miRNAs in endometriosis. There is evidence that uterus is a complex tissue comprising different resident cell types that respond differently to hormonal exposure [7]. Two markers of proliferation, MKI67 and MCM2, are expressed at peak levels in luminal and glandular epithelial cells during the proliferative phase, whereas the expression of these proliferative markers in stromal cells shows much less change across the two cycle phases [7, 52]. Collectively, these findings indicate that responses to sex steroid hormones are cell-type specific; therefore, to understand the mechanism of action of these hormones, it is essential to study responses in particular cell types. However, separating the two cellular compartments for molecular studies precludes any opportunities to investigate the mechanistic basis of epithelial and stromal cell interactions and thus paracrine influences in mediating hormonal effects on these two cell types.
Identification of protein-coding genes that are regulated by a specific miRNA has proved difficult, despite development of computational approaches to predict miRNA targets. The ability to identify target mRNA is further hampered by the fact that only partial complementary sequence between miRNA and the 3′ untranslated regions of target mRNA is needed and that target selectivity of miRNAs may depend on the cellular environment. To circumvent some of these difficulties, we studied the expression profiles of miRNA and mRNA in the same epithelial cell samples of endometrium to identify differentially expressed miRNAs and to investigate the influence of these miRNAs on putative target gene transcript levels. This approach can be successful if miRNAs of interest affect the transcript abundance of target mRNAs but fails if the target genes are regulated only by translational suppression. Most miRNAs are believed to negatively regulate their target gene expression by causing degradation of mRNA transcripts or translational repression, which would lead to an inverse expression relationship between miRNA and its target. However, this relationship may be more complex than previously thought, miRNAs were recently described as being able to activate translation of their targets [53].
The array data presented herein revealed several miRNAs whose transcript abundance is significantly increased in endometrial epithelium during the midsecretory phase compared with the late proliferative phase. Prediction analysis identified many cell cycle regulators as putative target mRNAs of these up-regulated miRNAs. These results suggest a new level of gene expression regulation that may be involved in hormonal regulation of epithelial cell proliferation in human endometrium by E2 and P4. Ongoing studies are focusing on validating the true cell cycle targets of differentially expressed miRNAs and on defining the functions of these miRNAs in human endometrial epithelium.
Acknowledgments
We thank Drs. Andy Brooks and Qi Wang in the Bionomics Research and Technology Center, University of Medicine and Dentistry of New Jersey, for excellent technical assistance with Affymetrix arrays and data analysis. In addition, we thank Ms. Donna Gerardi for support with figure design. We also thank the women who participated in the study.
Footnotes
Supported by the Eunice Kennedy Shriver NICHD/NIH through cooperative agreement (U54 HD058155) as part of the Specialized Cooperative Centers Program in Reproduction and Infertility Research and NIH R01 HD050614 (to J.W.P.), K24CD-41978 (to N.S.), and P30 CA13330 (to Albert Einstein Cancer Center), and by the American Association of Obstetricians and Gynecologists Foundation scholarship (to S.K.). J.W.P. is the Louis Goldstein Swan Chair in Women's Cancer Research.
REFERENCES
- Kurita T, Medina R, Schabel AB, Young P, Gama P, Parekh TV, Brody J, Cunha GR, Osteen KG, Bruner-Tran KL, Gold LI.The activation function-1 domain of estrogen receptor alpha in uterine stromal cells is required for mouse but not human uterine epithelial response to estrogen. Differentiation 2005; 73: 313–322. [DOI] [PubMed] [Google Scholar]
- Zhu L, Pollard JW.Estradiol-17beta regulates mouse uterine epithelial cell proliferation through insulin-like growth factor 1 signaling. Proc Natl Acad Sci U S A 2007; 104: 15847–15851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guzeloglu Kayisli O, Kayisli UA, Luleci G, Arici A.In vivo and in vitro regulation of Akt activation in human endometrial cells is estrogen dependent. Biol Reprod 2004; 71: 714–721. [DOI] [PubMed] [Google Scholar]
- Tong W, Pollard JW.Progesterone inhibits estrogen-induced cyclin D1 and cdk4 nuclear translocation, cyclin E- and cyclin A-cdk2 kinase activation, and cell proliferation in uterine epithelial cells in mice. Mol Cell Biol 1999; 19: 2251–2264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen B, Pan H, Zhu L, Deng Y, Pollard JW.Progesterone inhibits the estrogen-induced phosphoinositide 3-kinase–>AKT–>GSK-3beta–>cyclin D1–>pRB pathway to block uterine epithelial cell proliferation. Mol Endocrinol 2005; 19: 1978–1990. [DOI] [PubMed] [Google Scholar]
- Pan H, Deng Y, Pollard JW.Progesterone blocks estrogen-induced DNA synthesis through the inhibition of replication licensing. Proc Natl Acad Sci U S A 2006; 103: 14021–14026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niklaus AL, Aubuchon M, Zapantis G, Li P, Qian H, Isaac B, Kim MY, Adel G, Pollard JW, Santoro NF.Assessment of the proliferative status of epithelial cell types in the endometrium of young and menopausal transition women. Hum Reprod 2007; 22: 1778–1788. [DOI] [PubMed] [Google Scholar]
- Polotsky AJ, Zhu L, Santoro N, Pollard JW.Lithium chloride treatment induces epithelial cell proliferation in xenografted human endometrium. Hum Reprod 2009; 24: 1960–1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartel DP.MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297. [DOI] [PubMed] [Google Scholar]
- Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM.Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005; 433: 769–773. [DOI] [PubMed] [Google Scholar]
- Ohlsson Teague EM, Van der Hoek KH, Van der Hoek MB, Perry N, Wagaarachchi P, Robertson SA, Print CG, Hull LM.MicroRNA-regulated pathways associated with endometriosis. Mol Endocrinol 2009; 23: 265–275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boren T, Xiong Y, Hakam A, Wenham R, Apte S, Wei Z, Kamath S, Chen DT, Dressman H, Lancaster JM.MicroRNAs and their target messenger RNAs associated with endometrial carcinogenesis. Gynecol Oncol 2008; 110: 206–215. [DOI] [PubMed] [Google Scholar]
- Bueno MJ, de Castro IP, Malumbres M.Control of cell proliferation pathways by microRNAs. Cell Cycle 2008; 7: 3143–3148. [DOI] [PubMed] [Google Scholar]
- Pan Q, Luo X, Toloubeydokhti T, Chegini N.The expression profile of micro-RNA in endometrium and endometriosis and the influence of ovarian steroids on their expression. Mol Hum Reprod 2007; 13: 797–806. [DOI] [PubMed] [Google Scholar]
- Burney RO, Hamilton AE, Aghajanova L, Vo KC, Nezhat CN, Lessey BA, Giudice LC.MicroRNA expression profiling of eutopic secretory endometrium in women with versus without endometriosis. Mol Hum Reprod 2009; 15: 625–631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez G, Behringer RR.Dicer is required for female reproductive tract development and fertility in the mouse. Mol Reprod Dev 2009; 76: 678–688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagaraja AK, Andreu-Vieyra C, Franco HL, Ma L, Chen R, Han DY, Zhu H, Agno JE, Gunaratne PH, DeMayo FJ, Matzuk MM.Deletion of Dicer in somatic cells of the female reproductive tract causes sterility. Mol Endocrinol 2008; 22: 2336–2352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong X, Luense LJ, McGinnis LK, Nothnick WB, Christenson LK.Dicer1 is essential for female fertility and normal development of the female reproductive system. Endocrinology 2008; 149: 6207–6212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu SJ, Ren G, Liu JL, Zhao ZA, Yu YS, Su RW, Ma XH, Ni H, Lei W, Yang ZM.MicroRNA expression and regulation in mouse uterus during embryo implantation. J Biol Chem 2008; 283: 23473–23484. [DOI] [PubMed] [Google Scholar]
- Chakrabarty A, Tranguch S, Daikoku T, Jensen K, Furneaux H, Dey SK.MicroRNA regulation of cyclooxygenase-2 during embryo implantation. Proc Natl Acad Sci U S A 2007; 104: 15144–15149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Company MLI.New weight standards for men and women. Stat Bull Metrop Life Found 1983; 64: 1–4. [Google Scholar]
- Lessey BA, Killam AP, Metzger DA, Haney AF, Greene GL, McCarty KSJ.Immunohistochemical analysis of human uterine estrogen and progesterone receptors throughout the menstrual cycle. J Clin Endocrinol Metab 1988; 67: 334–340. [DOI] [PubMed] [Google Scholar]
- Israel R, Mishell DR, Stone SS, Thorneycroft IH, Moyer DL.Single luteal phase serum progesterone assay as an indicator of ovulation. Am J Obstet Gynecol 1972; 112: 1043–1046. [DOI] [PubMed] [Google Scholar]
- Noyes RW, Haman JO.Accuracy of endometrial dating: correlation of endometrial dating with basal body temperature and menses. Fertil Steril 1953; 4: 504–517. [DOI] [PubMed] [Google Scholar]
- Santoro N, Isaac B, Neal-Perry G, Weingart L, Nussbaum A, Thakur S, Jinnai H, Khosla N, Barad D.Impaired folliculogenesis and ovulation in older reproductive aged women. J Clin Endocrinol Metab 2003; 88: 5502–5509. [DOI] [PubMed] [Google Scholar]
- Satyaswaroop PG, Fleming H, Bressler RS, Gurpide E.Human endometrial cancer cell cultures for hormonal studies. Cancer Res 1978; 38: 4367–4375. [PubMed] [Google Scholar]
- Gautier L, Cope L, Bolstad BM, Irizarry RA.affy: Analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 2004; 20: 307–315. [DOI] [PubMed] [Google Scholar]
- Xie X, Lu J, Kulbokas EJ, Golub TR, Mootha V, Lindblad-Toh K, Lander ES, Kellis M.Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 2005; 434: 338–345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berezikov E, Guryev V, van de Belt J, Wienholds E, Plasterk RH, Cuppen E.Phylogenetic shadowing and computational identification of human microRNA genes. Cell 2005; 120: 21–24. [DOI] [PubMed] [Google Scholar]
- Cummins JM, He Y, Leary RJ, Pagliarini R, Diaz LA, Jr, Sjoblom T, Barad O, Bentwich Z, Szafranska AE, Labourier E, Raymond CK, Roberts BS, et al. The colorectal microRNAome. Proc Natl Acad Sci U S A 2006; 103: 3687–3692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentwich I, Avniel A, Karov Y, Aharonov R, Gilad S, Barad O, Barzilai A, Einat P, Einav U, Meiri E, Sharon E, Spector Y, et al. Identification of hundreds of conserved and nonconserved human microRNAs. Nat Genet 2005; 37: 766–770. [DOI] [PubMed] [Google Scholar]
- Huber W, von Heydebreck A, Sultmann H, Poustka A, Vingron M.Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 2002; 18(suppl 1):S96–S104. [DOI] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD.Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001; 25: 402–408. [DOI] [PubMed] [Google Scholar]
- Talbi S, Hamilton AE, Vo KC, Tulac S, Overgaard MT, Dosiou C, Le Shay N, Nezhat CN, Kempson R, Lessey BA, Nayak NR, Giudice LC.Molecular phenotyping of human endometrium distinguishes menstrual cycle phases and underlying biological processes in normo-ovulatory women. Endocrinology 2006; 147: 1097–1121. [DOI] [PubMed] [Google Scholar]
- Giannakakis A, Sandaltzopoulos R, Greshock J, Liang S, Huang J, Hasegawa K, Li C, O'Brien-Jenkins A, Katsaros D, Weber BL, Simon C, Coukos G, et al. miR-210 links hypoxia with cell cycle regulation and is deleted in human epithelial ovarian cancer. Cancer Biol Ther 2008; 7: 255–264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dennis G, Jr, Shernan BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA.DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol 2003; 4: eP3 [PubMed] [Google Scholar]
- Huang DW, Sherman BT, Lempicki RA.Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4: 44–57. [DOI] [PubMed] [Google Scholar]
- Borthwick JM, Charnock-Jones DS, Tom BD, Hull ML, Teirney R, Phillips SC, Smith SK.Determination of the transcript profile of human endometrium. Mol Hum Reprod 2003; 9: 19–33. [DOI] [PubMed] [Google Scholar]
- Burney RO, Talbi S, Hamilton AE, Vo KC, Nyegaard M, Nezhat CR, Lessey BA, Giudice LC.Gene expression analysis of endometrium reveals progesterone resistance and candidate susceptibility genes in women with endometriosis. Endocrinology 2007; 148: 3814–3826. [DOI] [PubMed] [Google Scholar]
- Tulac S, Nayak NR, Kao LC, Van Waes M, Huang J, Lobo S, Germeyer A, Lessey BA, Taylor RN, Suchanek E, Giudice LC.Identification, characterization, and regulation of the canonical Wnt signaling pathway in human endometrium. J Clin Endocrinol Metab 2003; 88: 3860–3866. [DOI] [PubMed] [Google Scholar]
- Wang Y, Hanifi-Moghaddam P, Hanekamp EE, Kloosterboer HJ, Franken P, Veldscholte J, van Doorn HC, Ewing PC, Kim JJ, Grootegoed JA, Burger CW, Fodde R, et al. Progesterone inhibition of Wnt/beta-catenin signaling in normal endometrium and endometrial cancer. Clin Cancer Res 2009; 15: 5784–5793. [DOI] [PubMed] [Google Scholar]
- Hou X, Tan Y, Li M, Dey SK, Das SK.Canonical Wnt signaling is critical to estrogen-mediated uterine growth. Mol Endocrinol 2004; 18: 3035–3049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Q, Fu H, Sun F, Zhang H, Tie Y, Zhu J, Xing R, Sun Z, Zheng X.miR-16 family induces cell cycle arrest by regulating multiple cell cycle genes. Nucleic Acids Res 2008; 36: 5391–5404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linsley PS, Schelter J, Burchard J, Kibukawa M, Martin MM, Bartz SR, Johnson JM, Cummins JM, Raymond CK, Dai H, Chau N, Cleary M, et al. Transcripts targeted by the microRNA-16 family cooperatively regulate cell cycle progression. Mol Cell Biol 2007; 27: 2240–2252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun F, Fu H, Liu Q, Tie Y, Zhu J, Xing R, Sun Z, Zheng X.Downregulation of CCND1 and CDK6 by miR-34a induces cell cycle arrest. FEBS Lett 2008; 582: 1564–1568. [DOI] [PubMed] [Google Scholar]
- le Sage C, Nagel R, Egan DA, Schrier M, Mesman E, Mangiola A, Anile C, Maira G, Mercatelli N, Ciafre SA, Farace MG, Agami R.Regulation of the p27(Kip1) tumor suppressor by miR-221 and miR-222 promotes cancer cell proliferation. EMBO J 2007; 26: 3699–3708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sherr CJ, Roberts JM.Inhibitors of mammalian G1 cyclin-dependent kinases. Genes Dev 1995; 9: 1149–1163. [DOI] [PubMed] [Google Scholar]
- Qian K, Hu L, Chen H, Li H, Liu N, Li Y, Ai J, Zhu G, Tang Z, Zhang H.Hsa-miR-222 is involved in differentiation of endometrial stromal cells in vitro. Endocrinology 2009; 150: 4734–4743. [DOI] [PubMed] [Google Scholar]
- Gillies JK, Lorimer IA.Regulation of p27Kip1 by miRNA 221/222 in glioblastoma. Cell Cycle 2007; 6: 2005–2009. [DOI] [PubMed] [Google Scholar]
- Galardi S, Mercatelli N, Giorda E, Massalini S, Frajese GV, Ciafre SA, Farace MG.miR-221 and miR-222 expression affects the proliferation potential of human prostate carcinoma cell lines by targeting p27Kip1. J Biol Chem 2007; 282: 23716–23724. [DOI] [PubMed] [Google Scholar]
- Visone R, Russo L, Pallante P, De Martino I, Ferraro A, Leone V, Borbone E, Petrocca F, Alder H, Croce CM, Fusco A.MicroRNAs (miR)-221 and miR-222, both overexpressed in human thyroid papillary carcinomas, regulate p27Kip1 protein levels and cell cycle. Endocr Relat Cancer 2007; 14: 791–798. [DOI] [PubMed] [Google Scholar]
- Kato K, Toki T, Shimizu M, Shiozawa T, Fujii S, Nikaido T, Konishi I.Expression of replication-licensing factors MCM2 and MCM3 in normal, hyperplastic, and carcinomatous endometrium: correlation with expression of Ki-67 and estrogen and progesterone receptors. Int J Gynecol Pathol 2003; 22: 334–340. [DOI] [PubMed] [Google Scholar]
- Vasudevan S, Tong Y, Steitz JA.Switching from repression to activation: microRNAs can up-regulate translation. Science 2007; 318: 1931–1934. [DOI] [PubMed] [Google Scholar]