Abstract
The motor protein, non-muscle myosin II (NMII), must undergo dynamic oligomerization into filaments to participate in cellular processes such as cell migration and cytokinesis. A small non-helical region at the tail of the long coiled-coil region (tailpiece) is a common feature of all dynamically assembling myosin II proteins. In this study, we investigated the role of the tailpiece in NMII-C self-assembly. We show that the tailpiece is natively unfolded, as seen by circular dichroism and NMR experiments, and is divided into two regions of opposite charge. The positively charged region (Tailpiece1946–1967) starts at residue 1946 and is extended by seven amino acids at its N terminus from the traditional coiled-coil ending proline (Tailpiece1953–1967). Pull-down and sedimentation assays showed that the positive Tailpiece1946–1967 binds to assembly incompetent NMII-C fragments inducing filament assembly. The negative region, residues 1968–2000, is responsible for NMII paracrystal morphology as determined by chimeras in which the negative region was swapped between the NMII isoforms. Mixing the positive and negative peptides had no effect on the ability of the positive peptide to bind and induce filament assembly. This study provides molecular insight into the role of the structurally disordered tailpiece of NMII-C in shifting the oligomeric equilibrium of NMII-C toward filament assembly and determining its morphology.
Keywords: Molecular Motors/Myosin, Peptides/Interactions, Protein/Protein-Protein Interactions, Protein/Assembly, Protein Structure, Myosin II Filament Assembly, Myosin II Non-helical Tailpiece
Introduction
Myosin II is a hexameric protein (Fig. 1A) of a family of actin-based molecular motors that are involved in cellular activities such as muscle contraction, cell migration, and cytokinesis (1–4). Each hexamer comprises two identical heavy chains and two sets of light chains. The N-terminal region of myosin II heavy chain is a globular head containing the actin binding and ATPase domains, followed by a large coiled-coil rod, which enables independent myosin II hexamers to assemble into large filaments (1). Non-muscle myosin II (NMII)4 is a ubiquitous protein expressed by all cell types that mediates cellular processes that require contractility (2–4). In contrast to skeletal myosin II, NMII undergoes dynamic filament assembly-disassembly cycles to facilitate its diverse range of activities (5). NMII can only function when in filament form. Therefore, the process of filament assembly is an important regulatory step controlling NMII action (1). Several crucial factors for NMII filament assembly have been identified, including charge periodicity in the amino acid sequence along the coiled-coil rod and the presence of two small assembly competence domains located at the C terminus of the coiled-coil (6–11). Additionally myosin II light chain phosphorylation activates the motor domain and promotes filament assembly (12–14).
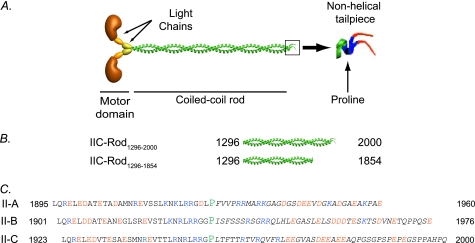
FIGURE 1.
A, schematic presentation of NMII functional domains. B, schematic presentation of IIC-Rod fragments. C, sequence alignment of the C-terminal region of NMII isoforms. Italic characters, tailpiece sequences; blue characters, positively charged residues; red characters, negatively charged residues; P in green, proline ending the traditional coiled-coil. The numbers represent amino acid positions in the full-length protein.
Three isoforms of NMII (NMII-A, NMII-B, and NMII-C) have been identified in mammals (15–17), each with specific tissue distributions and functions (18–23). The C terminus of NMII contains a non-helical tailpiece (tailpiece) of 33–47 amino acids depending on the isoform (Fig. 1A). All myosin II molecules that undergo dynamic assembly and disassembly have non-helical tailpieces. This tailpiece has been shown to be an important regulatory domain, affecting NMII filament assembly and cellular functions (24–28). Removing the tailpiece from chicken smooth muscle myosin II and NMII changed their solubility and impaired the ability of NMII to form large filamentous structures (24, 28). Furthermore, smooth muscle myosin II has a splice variant lacking the tailpiece, which exhibited significantly different properties of filament assembly that were reflected by different molecular packing inside the filament (26). Swapping the tailpieces among the different NMII isoforms showed that the tailpiece affects NMII function in vivo and determines the specific morphology of NMII filaments (28). Additionally, phosphorylating the tailpiece by protein kinase C and casein kinase II resulted in decreased NMII assembly as determined by sedimentation and critical concentration experiments (29–34). Mutations creating a stop codon that prevents the NMII tailpiece from being expressed, resulted in platelet disorders and hearing problems (35). Despite its apparent importance no structural data explaining the tailpiece function in filament assembly are available.
The distribution of hydrophobic, basic, and acidic amino acids among the different NMII isoform tailpieces is preserved although the lengths and sequences vary (Fig. 1C). This distribution divides the tailpiece into a positively charged region at the N terminus followed by a negatively charged region at the C terminus (Fig. 1C). These two conserved charged regions may be an important feature for the function of the tailpiece in filament assembly.
In this study NMII-C served as a model to explore the structure and mechanism of action of the NMII tailpiece. Using a combination of structural, biophysical, and biochemical methods we show that the NMII tailpiece is unstructured and that the newly defined, positively charged region of the tailpiece promotes filament assembly.
EXPERIMENTAL PROCEDURES
Peptide Synthesis, Labeling, and Purification
Tailpiece peptides and a control peptide KKLANAPRRLKKNSS with a positive charge of +6 were synthesized using an Applied Biosystems (ABI) 433A peptide synthesizer. Tailpiece1955–2000 and Tailpiece1946–2000 were synthesized using a Liberty microwave-assisted peptide synthesizer (CEM). Labeled Tailpiece1946–2000 was ordered from GL Biochem (Shanghai). The peptides were labeled with Trp at their N terminus for UV spectroscopy. For pull-down experiments, the N termini of the peptides were labeled with fluorescein (50). Peptide purification was performed with a Gilson HPLC using a reverse-phase C8 semi-preparative column (ACE, Advanced Chromatography Technologies) with a gradient of 5–60% acetonitrile in water (both containing 0.1% (v/v) trifluoroacetic acid). Peptide purity was confirmed by MALDI-TOF mass spectrometry and analytical HPLC (supplemental Fig. S1). The peptide concentration was determined using a UV spectrophotometer (Shimadzu Kyoto, Japan) as described previously (51).
Proteins Used in This Study
NMII-A, GenBankTM accession number NP_002464, NMII-B, GenBankTM accession number A59252, and NMII-C, GenBankTM accession number AY363100.
Construction of NMII Mutants
The IIC-Rod1296–1854 construct was created by introducing a stop codon at position 1855 on the NMII rods in pET21 as previously described (28) using QuikChangeTM site-directed mutagenesis kit (Stratagene, La Jolla, CA) with the following primers 5′-GCC CAG GCA GAG GAG CAG TAG TAA GCA GGA GAG CAG GGA GCG CAT C-3′. Negatively charged tail swapping mutants were created by adding a KpnI site after the positively charged region of each tailpiece isoform by site-directed mutagenesis with the following primers: NMII-A KpnI 5′-CGG AAA GGC GCC GGG GGT ACC TCC GAC GAC GAA GAG GTA GAT-3′. NMII-B KpnI 5′-GCG CCA GCT GCA CCT TGG TAC CGC TTC CCT GGA GCT CTC-3′. NMII-C KpnI 5′-CGT CAG AAG CCA CGC CGG TAC CCA GCC GGA ACA CCT GG-3′. Fragments from the tailpieces containing the negatively charged region were removed by digestion with KpnI and DraIII followed by ligation using Mighty-Mix ligation kit (Takara Bio, Shiga, Japan) into the different NMII isoforms in pET21, digested with the same enzymes. Finally, the KpnI site was restored to the original sequence using site-directed mutagenesis with the following primers: NMII-A negative B KpnI fix: 5′-GGA AAG GCG CCG GGG AAG GAG CTT CCC TGG AGC TCT CCG-3′. NMII-A negative C KpnI fix: 5′-GGA AAG GCG CCG GGG AAG AGG GCG TGG CTT CTG ACG AG-3′. NMII-B negative A KpnI fix: 5′-GCG CCA GCT GCA CCT TGA TGG CTC CGA CGA AGA GGT AGA TGG-3′. NMII-B negative C KpnI fix: 5′-GCG CCA GCT GCA CCT TGS SGS GGG CGT GGC TTC TGS CGS G-3′. NMII-C negative A KpnI fix: 5′-CAG GTG TTC CGG CTG GAT GGC TCC GAC GAA GAG GTA GAT GGC-3′. NMII-C negative B KpnI fix: 5′-GGT GTT CCG GCT GGA AGG AGC TTC CCT GGA GCT CTC CG-3′. A 6× His tag IIC-Rod1296–1854 construct was created by annealing a 6× His tag primer with NdeI sites and ligating with NMII in pET21 constructs described above. FWD 6× HIS primer: 5′-TAT GCA TCA CCA TCA CCA TCA CCA-3′. REV 6× HIS primer: 5′-TAT GGT GAT GGT GAT GCA-3′. The sequences of all constructs were confirmed by DNA sequencing (Center for Genomic Analysis, The Hebrew University of Jerusalem).
Purification of NMII Fragments from E. coli, Sedimentation Assays, and Negative Staining for Electron Microscopy
NMII fragment purification was performed as described (36). Assays were performed as described previously (10, 36) with the following modification: Filament formation for electron microscopy was performed in a buffer containing 25 mm phosphate buffer, pH 7.5, 12 mm CaCl2, and 18 mm MgCl2. An additional wash step was added after protein deposition on the grid for the electron microscopy experiments. Filaments were measured using Image-Pro plus software (Media Cybernetics) on calibrated electron micrographs at ×88000 magnification. Measurements were consequently averaged and subjected to a two-tailed, two-sampled unequal variance Student's t test using at least 15 measurements.
Pull-down Assay
His6-tagged IIC-Rod1296–1854, was expressed and purified as described above. The purified protein was incubated with Ni-nitrilotriacetic acid beads (Qiagen Gmbh, Hilden, Germany) for 2 h in buffer A (25 mm phosphate buffer, pH 7.5, 800 mm NaCl and 10 mm imidazole) at 4 °C followed by three washes in buffer A. The Ni-bound His6-tagged IIC-Rod1296–1854 was quantified by SDS-PAGE and Coomassie Blue staining, and the amounts of protein were normalized. His6-tagged IIC-Rod1296–1854-bound Ni-beads were incubated for 2 h at 4 °C in 300 μl of binding buffer with an ionic strength of 100 mm (25 mm phosphate buffer, pH 7.5, 42 mm NaCl) with 0.5 μm fluorescein-labeled peptides. After binding, the beads were washed three times followed by eluting the bound peptides with 200 μl of elution buffer (25 mm phosphate buffer, pH 7.5, and 800 mm NaCl). The amount of peptide bound to the Ni-bead-IIC-Rod1296–1854 was measured with a Galaxy fluorescent spectrophotometer using 485/520 nm filters. The amount of bound peptide was calculated by subtracting the background values obtained for peptide bound to Ni-beads only.
NMR Measurements
Samples of each peptide (Table 1) were prepared by dissolving the lyophilized peptide in 25 mm sodium phosphate buffer, pH 7.2, 100 mm NaCl, and D2O was added to give a final 10% solution (v/v). In assays in which the peptides were mixed with IIC-Rod1296–1854, the peptides were dissolved in IIC-Rod1296–1854 solution (25 mm sodium phosphate buffer, pH 7.2, 100 mm NaCl). pH values were adjusted with 0.1 m NaOH to give pH 6.80 and a final concentration of 0.5 mm. The final IIC-Rod1296–1854 concentration was 0.25 mm. NMR measurements were performed on a Bruker Avance 600 MHz DMX spectrometer (Bruker, Germany) operating at the proton frequency of 600.13 MHz. TOSCY (37, 38), and NOESY (39) experiments were acquired under identical conditions. All spectra were acquired at 10 °C. Spectra were processed and analyzed with the XWINNMR software package (Bruker Analytische Messtechnik GmbH) and SPARKY (provided by Goddard T. D. and Kneller D. G., SPARKY 3, University of California, San Francisco).
TABLE 1.
NMII-C tailpiece peptides used in this study
a Red characters, positively charged amino acids; blue characters, negatively charged amino acids.
CD Measurements
Samples of each peptide (Table 1) were prepared by dissolving lyophilized peptide in 25 mm sodium phosphate buffer, pH 7.2, 100 mm NaCl. Solutions of IIC-Rod1296–1854 in the presence and absence of each peptide were prepared by mixing stock solutions of each peptide and protein. The final concentration of IIC-Rod1296–1854 was 0.01 mm. CD spectra were recorded using a J-810 spectropolarimeter (Jasco) in a 0.1-cm quartz cuvette for far-UV CD spectroscopy. Far-UV CD spectra were collected over 190–260 nm at room temperature.
RESULTS
The NMII-C Tailpiece Is Natively Unfolded
Traditionally, the NMII tailpiece has been defined as the C-terminal portion of the NMII rod downstream to the proline residue that presumably breaks the α-helix structure of the coiled-coil rod (40). According to this definition, the NMII-C tailpiece is between amino acids 1954 and 2000 (Fig. 1C). We tested the structure of the NMII-C tailpiece using a chemically synthesized peptide corresponding to the 47-amino acid NMII-C tailpiece (Tailpiece1954–2000, Table 1). The circular dichroism (CD) spectrum of Tailpiece1954–2000 shows a minimum absorption around 200 nm, which is characteristic of an unfolded protein (Fig. 2A). NMR TOCSY experiments showed a fingerprint region of the spectrum with low chemical shift dispersion (∼0.8 ppm) and high overlap, further indicating that the tailpiece has an unstructured character (Fig. 2B).
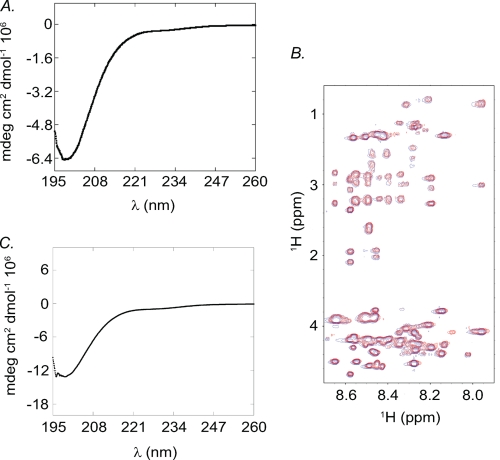
FIGURE 2.
Secondary structural analysis of NMII-C tailpieces. A, CD spectra of 0.01 mm Tailpiece1954–2000. B, overlaid fingerprint region of the NMR TOCSY spectra of the Tailpiece1954–2000 in the presence (red) and absence (blue) of IIC-Rod1296–1854. C, CD spectra of 0.03 mm Tailpiece1946–2000.
The Traditional NMII-C Tailpiece Does Not Interact with IIC-Rod1296–1854
Because the tailpiece is important for filament assembly, its interaction with the coiled-coil region of NMII-C during the assembly process was studied using a fragment of NMII-C coiled-coil lacking the C-terminal 146 amino acids (IIC-Rod1296–1854, Fig. 1B). This fragment does not create filaments even at low salt concentrations (supplemental Fig. S2) making it suitable for studying the effect of the tailpiece on filament assembly. The effect of the tailpiece on IIC-Rod1296–1854 filament assembly was studied by sedimentation assays, which are used to determine the ability of NMII to form filaments (24, 29, 41, 42). In this assay, filamentous NMII is insoluble and remains in the pellet upon high speed centrifugation, whereas non-filamentous NMII appears in the supernatant. IIC-Rod1296–1854 alone is completely soluble (Fig. 3), indicating that it is unable to assemble into filaments. Incubating IIC-Rod1296–1854 with Tailpiece1954–2000 had no effect on IIC-Rod1296–1854 sedimentation (Fig. 3). This was confirmed by CD analysis of IIC-Rod1296–1854 in the presence of increasing concentrations of Tailpiece1954–2000. The resulting CD spectra were the sum of IIC- Rod1296–1854 and Tailpiece1954–2000 spectra, indicating that no structural changes occurred (supplemental Fig. S4). NMR TOCSY spectra (Fig. 2B) of Tailpiece1954–2000 alone and in the presence of IIC-Rod1296–1854 (at a molar ratio of 2:1 peptide:IIC-Rod1296–1854) showed no deviations in the chemical shift, further indicating that the traditional NMII-C tailpiece does not interact with IIC-Rod1296–1854.
FIGURE 3.
Sedimentation of IIC-Rod1296–1854 by tailpiece peptides. 5 μm IIC-Rod1296–1854 were incubated in the presence of 20 μm tailpiece peptides at an ionic strength of 100 mm for 4 h at 4 °C. Samples were centrifuged for 1 h at 100,000 × g. Equal amounts of supernatant and pellet were separated on SDS-PAGE and stained with Coomassie Blue. Control peptide: KKLANAPRRLKKNSS. S, supernatant; P, pellet; M, molecular weight marker.
The Positively Charged Part of Tailpiece1946–1967 Induces IIC-Rod1296–1854 Filament Assembly
The NMII-C tailpiece has been traditionally defined as amino acids 1954–2000. However, the margins of the non-helical portion have not been exactly defined biochemically, and no structural data exist for the tailpiece. All NMII isoforms have a short positively charged region N-terminal to the proline, which is preceded by a negatively charged region (Fig. 1C). We propose that the positively charged region upstream to the proline is an integral part of the tailpiece forming one continuous positively charged region. According to this hypothesis, the positive residues N-terminal to the proline (1946–1953) forms a continuous domain with the positively charged region C-terminal to the proline (1954–1967) of NMII-C. The properties of the extended non-helical tailpiece region (Fig. 1) encompassing amino acids 1946–2000 were investigated. CD spectra of Tailpiece1946–2000, a peptide corresponding to this region, showed it to be natively unfolded, similar to Tailpiece1954–2000 (Fig. 2C). Because all NMII tailpieces are composed of a positive region followed by a negatively charged region (Fig. 1C), the distinct role of each charged region in NMII filament assembly was tested. Tailpiece1946–2000 was divided into two peptides, one derived from the N-terminal part having a net positive charge of +7 (Tailpiece1946–1967 and Table 1), and the second from the C-terminal region having a net negative charge of −9 (Tailpiece1968–2000, Table 1). Sedimentation experiments of IIC-Rod1296–1854 with the positively charged Tailpiece1946–1967 showed that IIC-Rod1296–1854 formed insoluble structures that appeared only in the pellet (Fig. 3). In contrast, IIC-Rod1296–1854 showed no evidence of assembly when incubated with the negatively charged Tailpiece1968–2000 as it appeared only in the supernatant (Fig. 3). Incubating IIC-Rod1296–1854 with a positively charged control peptide with a net charge of +6 had no affect on its sedimentation properties (Fig. 3).
CD spectra of IIC-Rod1296–1854 showed changes in secondary structure upon adding the positively charged tailpiece peptide Tailpiece1946–1967 (Fig. 4A). IIC-Rod1296–1854 alone adopted an expected α-helical conformation. Adding the positively charged Tailpiece1946–1967 caused a decrease in α-helical content, which was dependent on the concentration of the positively charged Tailpiece1946–1967 (Fig. 4A). This decrease in the absorption amplitude indicates a decrease in the concentration of IIC-Rod1296–1854 in solution, possibly because of filament assembly. Similar experiments using IIC-Rod1296–1854 with the negatively charged Tailpiece1968–2000 showed a typical CD spectrum representing the sum of IIC-Rod1296–1854 and Tailpiece1968–2000 spectra indicating no structural change (Fig. 4B). NMR TOCSY peaks of the positively charged Tailpiece1946–1967 with a 2:1 molar ratio of IIC-Rod1296–1854, disappeared relative to the unreacted peptide, which may be caused by increased correlation times due to binding to the large IIC-Rod1296–1854 (supplemental Fig. S3A), whereas similar NMR experiments using the negatively charged Tailpiece1968–2000 showed no change in the peak intensities or chemical shift (supplemental Fig. S3B). This supports the sedimentation and CD results, indicating that the positively charged Tailpiece1946–1967 induces IIC-Rod1296–1854 filament assembly into insoluble structures, but the negatively charged Tailpiece1968–2000 does not.
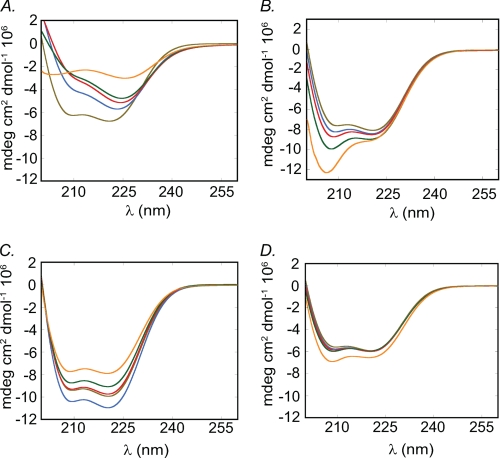
FIGURE 4.
Secondary structural analysis of IIC-Rod1296–1854 in the presence of tailpiece peptides. A, CD spectra of 0.01 mm IIC-Rod1296–1854 incubated with increasing concentrations of the positively charged Tailpiece1946–1967. B, CD spectra of 0.01 mm IIC-Rod1296–1854 incubated with increasing concentrations of the negatively charged Tailpiece1968–2000. C, CD spectra of 0.01 mm IIC-Rod1296–1854 in the presence of increasing concentrations of the positively charged Tailpiece 1946–1953. D, CD spectra of 0.01 mm IIC-Rod1296–1854 in the presence of increasing concentrations of the positively charged Tailpiece1953–1967. IIC-Rod1296–1854 alone is represented by a brown line. Blue, IIC-Rod1296–1854 and appropriate tailpiece peptide at 1:1 ratio; red, 1:2 ratio; green, 1:4 ratio; and orange, 1:10 ratio.
The Entire Positively Charged Region of the Tailpiece Is Required for Inducing IIC-Rod1296–1855 Filament Assembly
The role of the seven amino acids located N-terminal to the proline (Tailpiece1946–2000, Fig. 1C and Table 1) on IIC-Rod1296–1854 filament assembly was studied using two additional peptides: one corresponding to the seven extended residues (Tailpiece1946–1952, Table 1) and the other corresponding to the positively charged part of the traditional Tailpiece1953–1967 (Table 1). Sedimentation experiments of IIC-Rod1296–1854 in the presence of Tailpiece1946–1952 or Tailpiece1953–1967 showed only a slight increase in IIC-Rod1296–1854 filament assembly (Fig. 3). CD experiments using the positively charged Tailpiece1946–1952 and Tailpiece1953–1967 showed only a small conformational change upon incubation with IIC-Rod1296–1854 (Fig. 4, C and D). This minimal change in conformation was stable over time, indicating the absence of subsequently formed higher order assemblies (supplemental Fig. S5). The positively charged region of the tailpiece binds the NMII-C coiled-coil rod and induces NMII-C paracrystal formation. Sedimentation and CD experiments indicated that the positively charged Tailpiece1946–1967 may interact with IIC-Rod1296–1854 and that this interaction may induce its assembly. Binding of the different tailpiece peptides to the coiled-coil rod of NMII-C was studied by pull-down assays using His-tagged-IIC-Rod1296–1854 immobilized on nickel beads and fluorescein-labeled peptides. This assay tests whether the peptides bind to the coiled-coil rod in an assembly-independent manner. Peptides containing amino acids 1953–1967 bound IIC-Rod1296–1854 with Tailpiece1946–1967 having the strongest binding (Fig. 5). This is consistent with Tailpiece1946–1967 results from sedimentation and CD assays. No significant binding of Tailpiece1946–1952 to IIC-Rod1296–1854 was detected. The negative region, Tailpiece1968–2000, and control peptide also did not bind IIC-Rod1296–1854 (Fig. 5). These experiments confirm the importance of the positively charged region for filament assembly.
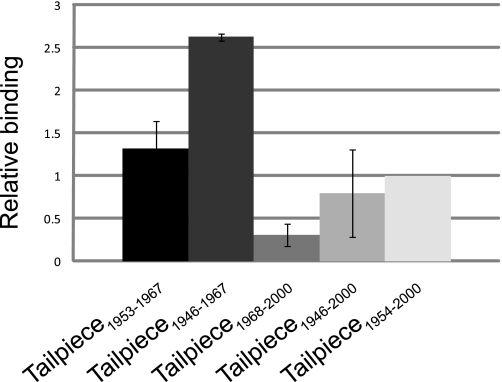
FIGURE 5.
The interaction between IIC-Rod1296–1854 and tailpiece peptides. Fluorescein-labeled tailpiece peptides were incubated for 2 h at 4 °C with Ni-beads-immobilized His-tagged IIC-Rod1296–1854. Bound peptides were eluted with 800 mm NaCl, 25 mm phosphate buffer, pH 7.5, and the fluorescence was detected by fluorescence spectrophotometer as described under “Experimental Procedures.” The extent of peptide bound to His-tagged-IIC-Rod1296–1854 was calculated by subtracting the background value of peptide bound to Ni-beads only from the value of peptide bound to His-tagged-IIC-Rod1296–1854. Values are the average of 3–4 independent experiments ± S.D. and are normalized to Tailpiece1954–2000 binding. Tailpiece1947–1952 and control peptide had binding values below background.
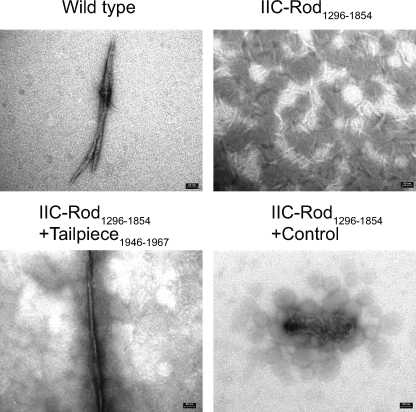
NMII forms large paracrystal filaments at low salt concentrations (43). These structures have been used to examine the capability of NMII to assemble into large filamentous structures (10, 24, 26, 35, 44). The effect of the tailpiece on NMII-C filament assembly was studied by inducing IIC-Rod1296–1854 to form paracrystals in the presence of the different tailpiece peptides. IIC-Rod1296–1854 alone is not capable of creating the distinctive long paracrystals of wild-type NMII-C (28), and only extremely small, needle-like structures were observed (Fig. 6). Adding Tailpiece1946–1967, corresponding to the full positively charged region of the tailpiece, to IIC-Rod1296–1854 resulted in the formation of filamentous paracrystals similar to those formed by wild-type NMII-C. In accordance with the CD and sedimentation experiments, no large filaments were observed upon incubating IIC-Rod1296–1854 with each of the truncated positively charged tailpieces (Tailpiece1946–1952 or Tailpiece1953–1967). Furthermore, the traditional Tailpiece1954–2000 or the peptide corresponding to the negatively charged Tailpiece1968–2000 had no effect on IIC-Rod1296–1854 paracrystal formation (data not shown). These data are consistent with sedimentation experiments.
FIGURE 6.
Tailpiece peptides effect on IIC-Rod1296–1854 paracrystal morphology. IIC-Rod1296–1854 was mixed with the different peptides at a 1:4 ratio in filament buffer (25 mm phosphate buffer, pH 7.5, 12 mm CaCl2, and 18 mm MgCl2). Filaments were negatively stained with uranyl acetate prior to viewing by electron microscope at ×88000 as described under “Experimental Procedures.”
The Negatively Charged Region of NMII-C Tailpiece Regulates the Positively Charged Region and Determines Filament Morphology
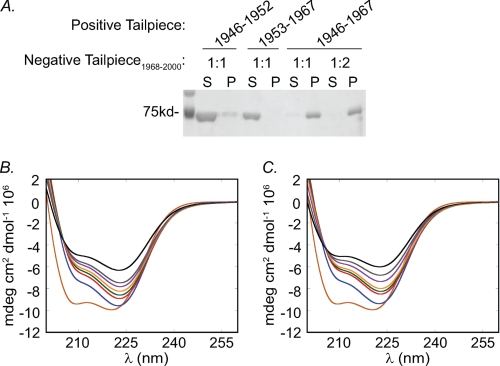
The role of the negatively charged region of the tailpiece on IIC-Rod1296–1854 filament assembly was studied by sedimentation and CD experiments in which the full positively charged Tailpiece1946–1967 was mixed with the negatively charged Tailpiece1968–2000 and added to IIC-Rod1296–1854. Sedimentation results showed that IIC-Rod1296–1854 was completely insoluble upon adding the mixture of positively and negatively charged tailpieces (Fig. 7A). A similar effect on IIC-Rod solubility was seen using the positively charged Tailpiece1946–1967 alone (Fig. 3). The change in CD spectra of IIC-Rod1296–1854 after adding an equimolar mixture of positively and negatively charged peptides was similar to the change seen when the positively charged Tailpiece1946–1967 was added alone (Fig. 7, B and C). As expected similar results were obtained when mixing the two short positive peptides (Fig. 7A). Thus in contrast to the complete tailpiece containing both the positively and negatively charged regions, the negatively charged Tailpiece1968–2000 by itself does not alter the effect of the positively charged Tailpiece1946–1967 on IIC-Rod1296–1854 filament assembly.
FIGURE 7.
The effect of mixing the negatively and positively charged tailpiece regions on IIC-Rod1296–1854 filament assembly. A, sedimentation assay of IIC-Rod1296–1854 in the presence of either of the positively charged Tailpiece1946–1952, Tailpiece1953–1967 or Tailpiece1946–1967 and the negatively charged Tailpiece1968–2000 were performed as described in Fig. 3. B, CD spectra of 0.01 mm IIC-Rod1296–1854 incubated with 0.01 mm Tailpiece1946–1967 alone. C, CD spectra of 0.01 mm IIC-Rod1296–1854 in the presence of both 0.01 mm Tailpiece1946–1967 and 0.01 mm Tailpiece1968–2000. Brown line, IIC-Rod1296–1854 alone; Black line, IIC-Rod1296–1854 and Tailpiece(s) that were collected after 0 min; Red line, after 1 min; blue line, after 5 min; green line, after 10 min; magenta line after 15 min; light blue line after 30 min; light green line, after 1 h, and orange line, after 2 h.
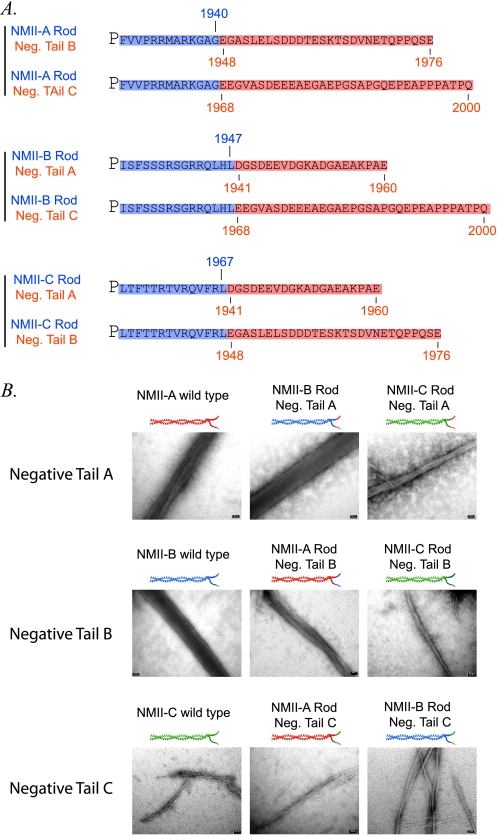
The tailpiece has been shown to be important for determining the isoform specific morphology of NMII paracrystals (28). As the negatively charged region does not promote IIC-Rod1296–1854 filament assembly it may be important in determining NMII paracrystal morphology. NMII fragment chimeras were created in which the negatively charged region was swapped among the NMII-A, NMII-B, and NMII-C isoforms. We have previously shown that rod fragments of the three NMII isoforms create different paracrystal morphology: NMII-A and NMII-B form large wide filaments (width measuring 1.21 ± 0.4 nm and 1.22 ± 0.38 nm, respectively) while NMII-C forms delicate thin filaments (0.32 ± 0.07 nm) (28). Swapping the negatively charged region of either NMII-A or NMII-B with the negatively charged region of NMII-C resulted in thin filaments similar to NMII-C (0.269 nm ±0.06 and 0.239 nm ±0.04, respectively) (Fig. 8 and Table 2). Accordingly, swapping the negatively charged region of NMII-C with the negatively charged region of NMII-A resulted in large filaments (0.534 ± 0.075 nm). Surprisingly swapping the negative tailpiece of NMII-C with the negative of NMII-B increased the filament width minimally (0.348 ± 0.075 nm) (Fig. 8 and Table 2). These results indicate that the negative region of the tailpiece plays a role in organizing NMII in the growing filament.
FIGURE 8.
The effect of the tailpiece negatively charged region on NMII paracrystal morphology. A, amino acids sequence of the negatively charged tailpiece swapped chimeras. Blue highlight, positively charged region, red highlight, negatively charged region. Numbers indicate amino acid position of the respective region. Only amino acids C-terminal to the proline are presented. B, NMII isoforms A, B, and C rod fragments with the negative region swapped among the isoforms were dialyzed against filament buffer and stained with uranyl acetate prior to viewing by electron microscope at ×88000 as described under “Experimental Procedures.” Blue represents NMII-A, red represents NMII-B, and green represents NMII-C originating amino acid sequence. Scale bars, 50 nm.
TABLE 2.
Width measurements of NMII negatively charged tailpiece chimeras
Paracrystal width was measured using ImagePro software from negatively stained electron micrographs as described in the legend to Fig. 8. p values are Student's t test comparing to wild-type measurement population.
| Name | Morphology | Width ± S.D. | p value (compared to wild type) |
|---|---|---|---|
| nm | |||
| NMII-A wild typea | Wide | 1.21 ± 0.4 | N/Ab |
| NMII-B wild typea | Wide | 1.22 ± 0.4 | N/A |
| NMII-C wild type | Thin | 0.32 ± 0.07 | N/A |
| NMII-A Neg. Tail B | Wide | 0.782 ± 0.21 | 7.34e-5 |
| NMII-A Neg. Tail C | Thin | 0.269 ± 0.06 | 1.64e-11 |
| NMII-B Neg. Tail A | Wide | 1.39 ± 0.3 | 0.1 |
| NMII-B Neg. Tail C | Thin | 0.239 ± 0.04 | 1.98e-17 |
| NMII-C Neg. Tail A | Wide | 0.534 ± 0.075 | 3.6e-6 |
| NMII-C Neg. tail B | Thin | 0.348 ± 0.075 | 0.33 |
a Data taken from Ref. 28.
b N/A, not applicable.
DISCUSSION
Extending the Tailpiece Boundary
As the C-terminal region of myosin II has not been extensively analyzed, its exact definition is unknown. Our results show that the actual tailpiece domain extends further toward the N terminus of NMII from the putative coiled-coil breaking proline. This region of seven amino acids is also positively charged, suggesting that it is an integral part of the positive region together with the traditional tailpiece. Data from other NMII isoforms have also shown that the small region upstream to the proline is important for assembly (28, 45). Both the traditional Tailpiece1954–2000 and the newly defined Tailpiece1946–2000 were found to be unfolded by CD (Fig. 2). The entire positively charged Tailpiece1946–1967 binds IIC-Rod1296–1854, and induces it to form structures similar to wild-type NMII-C as seen by CD and EM experiments. Dividing this region into what was traditionally thought to be the tailpiece of NMII-C (Tailpiece1953–1967) and a peptide representing only the seven amino acids upstream of the proline (Tailpiece1946–1952), showed that Tailpiece1953–1967 may be responsible for initial binding to IIC-Rod1296–1854. However this binding induced only minimal assembly of IIC-Rod1296–1854 as seen by CD and sedimentation experiments. This indicates that the entire positively charged region is a single domain responsible for promoting filament assembly.
The Negatively Charged C Terminus of the Tailpiece Has a Role Distinct from the Positive Region
Peptides corresponding to the negatively charged region of the Tailpiece1968–2000 were found to be unfolded and did not bind IIC-Rod1296–1854 as seen in pull-down assays. In addition, this peptide did not affect IIC-Rod1296–1854 assembly as it remained completely soluble in the presence of the peptide. Furthermore, CD experiments showed no conformational changes of IIC-Rod1296–1854 after incubation with the negatively charged Tailpiece1968–2000 (Fig. 4B). This is in agreement with previous experiments in which removing the negatively charged region of the tailpiece from smooth muscle myosin II had no effect on filament assembly properties (24, 46). Nevertheless the negatively charged Tailpiece1968–2000 was sufficient to determine NMII paracrystal morphology as seen in chimeric NMII isoforms in which only the negatively charged region was swapped among the isoforms (Fig. 8). Each region of the tailpiece seems to have a unique role in NMII filament assembly. Experiments exploring the individual roles of each region showed that although each region has a distinct role, the negatively charged region may regulate the positively charged region during filament assembly process. Neither the newly defined tailpiece, Tailpiece1946–2000, nor the traditional tailpiece, Tailpiece1954–2000, had any effect on IIC-Rod1296–1854 filament assembly even though these peptides contain the positively charged region (Fig. 3). Furthermore, binding experiments showed that extending either positively charged peptide to include the negatively charged region greatly reduced binding to IIC-Rod1296–1854 (Fig. 5). However, mixing experiments showed that adding the negatively charged Tailpiece1968–2000 did not affect the ability of the positively charged Tailpiece1947–1968 to induce IIC-Rod1296–1854 filament assembly (Fig. 7). This indicates that when in integral form, the negatively charged region may act as a regulator of the positively charged region.
A Model for the Role of the Tailpiece in Regulating Myosin Assembly
The proposed model for the role of the tailpiece in the assembly of NMII coiled-coil rod includes a positively charged region spanning both sides of the proline (amino acids 1946–1967) that is responsible for binding the coiled-coil rod and inducing it to form high oligomeric structures. Once the positively charged region has bound to the NMII rod, the negatively charged region can exert its effect on determining the morphology of the growing filament. This negatively charged region is also capable of masking the positively charged region, thereby hindering its ability to bind and hence modulating the assembly process.
The Tailpiece Is a Shiftide: A Peptide That Shifts the Oligomerization Equilibrium of Proteins
NMII is in equilibrium among individual hexamers and high order oligomers in filamentous form. In cases where proteins are in equilibrium among several oligomeric states, peptides or small molecules can bind specifically to one of these oligomeric species, stabilize the state, and thus shift the oligomerization equilibrium toward that specific state, according to the law of mass action (47, 48). We have termed such peptides “shiftides.” The positively charged Tailpiece1946–1967 is an example for such a shiftide, because it shows an ability to shift IIC-Rod1296–1854 from an individual hexameric state to a high order oligomeric structure. The mechanism by which the positively charged tailpiece peptide acts still needs to be investigated. Because NMII can only perform its functions when in filaments, peptides modulating filament assembly may shift oligomerization equilibrium toward the functional form of NMII. This peptide may have therapeutic potential by shifting the equilibrium toward filament assembly in diseases caused by defects in NMII assembly (49).
Supplementary Material
Acknowledgment
We thank Dr. Robert S. Adelstein for the NMII-C construct.
This work was supported in part by Israel Ministry of Health Grant 3114 (to S. R.) and a starting grant from the European Research Council (to A. F.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S5.
- NMII
- non-muscle myosin II
- tailpiece
- non-helical tailpiece.
REFERENCES
- 1.Sellers J. R. (1999) Myosins, 2nd Ed., Oxford University Press, Oxford, UK [Google Scholar]
- 2.Conti M. A., Adelstein R. S. (2008) J. Cell Sci. 121, 11–18 [DOI] [PubMed] [Google Scholar]
- 3.Matsumura F. (2005) Trends Cell Biol. 15, 371–377 [DOI] [PubMed] [Google Scholar]
- 4.Lauffenburger D. A., Horwitz A. F. (1996) Cell 84, 359–369 [DOI] [PubMed] [Google Scholar]
- 5.Sellers J. R. (2000) Biochim. Biophys. Acta 1496, 3–22 [DOI] [PubMed] [Google Scholar]
- 6.Sohn R. L., Vikstrom K. L., Strauss M., Cohen C., Szent-Gyorgyi A. G., Leinwand L. A. (1997) J. Mol. Biol. 266, 317–330 [DOI] [PubMed] [Google Scholar]
- 7.Atkinson S. J., Stewart M. (1992) J. Mol. Biol. 226, 7–13 [DOI] [PubMed] [Google Scholar]
- 8.McLachlan A. D., Karn J. (1982) Nature 299, 226–231 [DOI] [PubMed] [Google Scholar]
- 9.Nakasawa T., Takahashi M., Matsuzawa F., Aikawa S., Togashi Y., Saitoh T., Yamagishi A., Yazawa M. (2005) Biochemistry 44, 174–183 [DOI] [PubMed] [Google Scholar]
- 10.Rosenberg M., Straussman R., Ben-Ya'acov A., Ronen D., Ravid S. (2008) PLoS ONE 3, e1496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Straussman R., Squire J. M., Ben-Ya'acov A., Ravid S. (2005) J. Mol. Biol. 353, 613–628 [DOI] [PubMed] [Google Scholar]
- 12.Bresnick A. R. (1999) Curr. Opin. Cell Biol. 11, 26–33 [DOI] [PubMed] [Google Scholar]
- 13.Kolega J., Kumar S. (1999) Cell Motil. Cytoskel. 43, 255–268 [DOI] [PubMed] [Google Scholar]
- 14.Tan J. L., Ravid S., Spudich J. A. (1992) Annu. Rev. Biochem. 61, 721–759 [DOI] [PubMed] [Google Scholar]
- 15.Golomb E., Ma X., Jana S. S., Preston Y. A., Kawamoto S., Shoham N. G., Goldin E., Conti M. A., Sellers J. R., Adelstein R. S. (2004) J. Biol. Chem. 279, 2800–2808 [DOI] [PubMed] [Google Scholar]
- 16.Shohet R. V., Conti M. A., Kawamoto S., Preston Y. A., Brill D. A., Adelstein R. S. (1989) Proc. Natl. Acad. Sci. U.S.A. 86, 7726–7730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Simons M., Wang M., McBride O. W., Kawamoto S., Yamakawa K., Gdula D., Adelstein R. S., Weir L. (1991) Circ. Res. 69, 530–539 [DOI] [PubMed] [Google Scholar]
- 18.Bao J., Jana S. S., Adelstein R. S. (2005) J. Biol. Chem. 280, 19594–19599 [DOI] [PubMed] [Google Scholar]
- 19.Bao J., Ma X., Liu C., Adelstein R. S. (2007) J. Biol. Chem. 282, 22102–22111 [DOI] [PubMed] [Google Scholar]
- 20.Even-Ram S., Doyle A. D., Conti M. A., Matsumoto K., Adelstein R. S., Yamada K. M. (2007) Nat. Cell Biol. 9, 299–309 [DOI] [PubMed] [Google Scholar]
- 21.Jana S. S., Kawamoto S., Adelstein R. S. (2006) J. Biol. Chem. 281, 24662–24670 [DOI] [PubMed] [Google Scholar]
- 22.Sandquist J. C., Swenson K. I., Demali K. A., Burridge K., Means A. R. (2006) J. Biol. Chem. 281, 35873–35883 [DOI] [PubMed] [Google Scholar]
- 23.Wylie S. R., Chantler P. D. (2008) Mol. Biol. Cell 19, 3956–3968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hodge T. P., Cross R., Kendrick-Jones J. (1992) J. Cell Biol. 118, 1085–1095 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sato M. K., Takahashi M., Yazawa M. (2007) Mol. Biol. Cell 18, 1009–1017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rovner A. S., Fagnant P. M., Lowey S., Trybus K. M. (2002) J. Cell Biol. 156, 113–123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sinard J. H., Rimm D. L., Pollard T. D. (1990) J. Cell Biol. 111, 2417–2426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ronen D., Ravid S. (2009) J. Biol. Chem. 284, 24948–24957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Dulyaninova N. G., Malashkevich V. N., Almo S. C., Bresnick A. R. (2005) Biochemistry 44, 6867–6876 [DOI] [PubMed] [Google Scholar]
- 30.Even-Faitelson L., Ravid S. (2006) Mol. Biol. Cell 17, 2869–2881 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kelley C. A., Adelstein R. S. (1990) J. Biol. Chem. 265, 17876–17882 [PubMed] [Google Scholar]
- 32.Murakami N., Chauhan V. P., Elzinga M. (1998) Biochemistry 37, 1989–2003 [DOI] [PubMed] [Google Scholar]
- 33.Rosenberg M., Ravid S. (2006) Mol. Biol. Cell 17, 1364–1374 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Straussman R., Even L., Ravid S. (2001) J. Cell Sci. 114, 3047–3057 [DOI] [PubMed] [Google Scholar]
- 35.Franke J. D., Dong F., Rickoll W. L., Kelley M. J., Kiehart D. P. (2005) Blood 105, 161–169 [DOI] [PubMed] [Google Scholar]
- 36.Straussman R., Ben-Ya'acov A., Woolfson D. N., Ravid S. (2007) J. Mol. Biol. 366, 1232–1242 [DOI] [PubMed] [Google Scholar]
- 37.Piotto M., Saudek V., Sklenár V. (1992) J. Biomol. NMR 2, 661–665 [DOI] [PubMed] [Google Scholar]
- 38.Sklenar V., Piotto M., Leppik R., Saudek V. (1993) J. Magn. Resonance A 102, 241–245 [Google Scholar]
- 39.Jeener J., Meier B. H., Bachmann P., Ernst R. R. (1979) J. Chem. Physics 71, 4546–4553 [Google Scholar]
- 40.Chou P. Y., Fasman G. D. (1974) Biochemistry 13, 222–245 [DOI] [PubMed] [Google Scholar]
- 41.Hostetter D., Rice S., Dean S., Altman D., McMahon P. M., Sutton S., Tripathy A., Spudich J. A. (2004) PLoS Biol. 2, e356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Murakami N., Kotula L., Hwang Y. W. (2000) Biochemistry 39, 11441–11451 [DOI] [PubMed] [Google Scholar]
- 43.Kendrick-Jones J., Szent-Gyorgyi A. S., Cohen C. (1971) J. Mol. Biol. 59, 527–529 [DOI] [PubMed] [Google Scholar]
- 44.Atkinson S. J., Stewart M. (1991) J. Cell Sci. 99, 823–836 [DOI] [PubMed] [Google Scholar]
- 45.Ikebe M., Komatsu S., Woodhead J. L., Mabuchi K., Ikebe R., Saito J., Craig R., Higashihara M. (2001) J. Biol. Chem. 276, 30293–30300 [DOI] [PubMed] [Google Scholar]
- 46.Turbedsky K., Pollard T. D. (2005) J. Mol. Biol. 345, 351–361 [DOI] [PubMed] [Google Scholar]
- 47.Hayouka Z., Rosenbluh J., Levin A., Loya S., Lebendiker M., Veprintsev D., Kotler M., Hizi A., Loyter A., Friedler A. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 8316–8321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Jaffe E. K. (2005) Trends Biochem. Sci. 30, 490–497 [DOI] [PubMed] [Google Scholar]
- 49.Vicente-Manzanares M., Ma X., Adelstein R. S., Horwitz A. R. (2009) Nat. Rev. 10, 778–790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Weber P. J., Bader J. E., Folkers G., Beck-Sickinger A. G. (1998) Bioorg. Med. Chem. Lett. 8, 597–600 [DOI] [PubMed] [Google Scholar]
- 51.Gill S. C., von Hippel P. H. (1989) Anal. Biochem. 182, 319–326 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.