Abstract
Spermatogenesis is a cyclic process in which diploid spermatogonia differentiate into haploid spermatozoa. This process is highly regulated, notably at the post-transcriptional level. MicroRNAs (miRNAs), single-stranded noncoding RNA molecules of about 20–25 nucleotides, are implicated in the regulation of many important biological pathways such as proliferation, apoptosis, and differentiation. We wondered whether miRNAs could play a role during spermatogenesis. The miRNA expression repertoire was tested in germ cells, and we present data showing that miR-34c was highly expressed only in these cells. Furthermore, our findings indicate that in male gonads, miR-34c expression is largely p53 independent in contrast to previous results showing a direct link in somatic cells between the miR-34 family and this tumor suppressor protein. In order to identify target genes involved in germinal lineage differentiation, we overexpressed miR-34c in HeLa cells, analyzed the transcriptome of these modified cells, and noticed a shift of the expression profile toward the germinal lineage. Recently, it has been shown that exogenous expression of Ddx4/Vasa in embryonic chicken stem cells (cESC) induces cESC reprogramming toward a germ cell fate. When we simultaneously expressed miR-34c in such cells, we could detect an up-regulation of germ cell-specific genes whereas the expression of other lineage specific markers remained unchanged. These data suggest that miR-34c could play a role by enhancing the germinal phenotype of cells already committed to this lineage.
Keywords: miR-34c, spermatogenesis, miRNA, DDX4/VASA
INTRODUCTION
Spermatogenesis is a complex multistep developmental process that leads undifferentiated diploid cells to eventually differentiate into haploid male gamete cells. Spermatogenesis is commonly divided into three main phases: (1) the formation of spermatogonia from germline stem cells followed by mitotic divisions to produce primary spermatocytes; (2) meiosis, by which diploid spermatocytes differentiate into haploid spermatids; and (3) spermiogenesis, which leads to the production of mature sperm from round spermatids. In mammals, the differentiation of germ cells into spermatozoa occurs in the seminiferous epithelium of the testis and is highly regulated by extrinsic and intrinsic factors. Male germ cell differentiation is achieved through the sequential expression of genes that can be used as markers of the different germ cell types, such as TP1 and TP2 for round spermatids and TH2B/P19 for pachytene spermatocytes (Hecht 1998; Marret et al. 1998; He et al. 2009). However, despite intensive investigations, the mechanisms regulating the cell-type specificity of their expression remain largely unknown. Furthermore, numerous mRNAs in the differentiating germ cells also undergo post-transcriptional and translational regulation.
MicroRNAs (miRNAs) are small, noncoding single-stranded RNA molecules of about 22 nucleotides (nt) that are potent negative regulators of mRNA transcription, stability, and translation. They recognize their mRNA target through base-pairing, generally in their 3′UTR (Ambros 2008; Bartel 2009; Younger et al. 2009). They are evolutionarily conserved and play critical roles in many biological processes in various species (e.g., neural, muscle, germline development, cell cycle control, apoptosis, cancer, etc.). A large number of them show a tissue-specific expression, and several experiments have confirmed their importance in regulating cellular differentiation (Plasterk 2006; Ambros 2008; Bartel 2009; He et al. 2009). Moreover, Lim et al. (2005) have shown that the overexpression of a tissue-specific miRNA in a recipient nonrelated cell shifted its transcriptome toward that of the lineage expressing the miRNA (miR-1 and muscle; miR-124 and brain).
All these observations lead the authors to consider that miRNAs could play an active and important role during spermatogenesis. Several lines of evidence suggest that this is indeed the case. Tnp2, a testis-specific gene involved in chromatin remodeling during spermatogenesis, is regulated by miR-122a (Yu et al. 2005). The microRNAs, miRNA-372 and miRNA-373, have been implicated as oncogenes in testicular germ cells, but their precise role during spermatogenesis remains elusive (Voorhoeve et al. 2006). miRNA expression profiles of various testicular cell populations have been described, and many testicular miRNAs have been isolated (Yu et al. 2005; Ro et al. 2007; Yan et al. 2007; Hayashi et al. 2008; Bannister et al. 2009; Yan et al. 2009). Furthermore, Dicer1, a protein necessary for miRNA processing, and Dnd1, a protein implicated in the protection of mRNAs from miRNAs, are essential for spermatogenesis completion (Kedde et al. 2007; Hayashi et al. 2008; Maatouk et al. 2008).
To explore the connection between miRNA and germ cells, we analyzed the miRNA repertoire expressed in pachytene spermatocytes and round spermatids. These cells were chosen because they could be considered as representative of two main phases of spermatogenesis, meiosis and the beginning of spermiogenesis, respectively. We then focused our attention on miR-34c, a particular miRNA emphasized by this study. We present data showing its germ cell-specific expression, and we demonstrate, through different overexpression experiments, that miR-34c could enhance spermatogenesis.
RESULTS
miR-34c is specifically expressed in germ cells
To test whether miRNA expression varies during the late stages of spermatogenesis, we determined the developmental expression profile of miRNAs in adult mouse pachytene spermatocytes and round spermatids; these germ cells are being considered as representative of two main phases of spermatogenesis: meiosis and the beginning of spermiogenesis. A miRNA array capable of monitoring the expression of murine miRNAs was used (mirVANA miRNA Probe Set, Ambion). Following hybridization and result analysis, five miRNAs were found to be expressed above the detection threshold in these cells: miR-17-5p, miR-191, miR-15b, miR-106a, and miR-34c. To confirm these results, we analyzed their expression by Northern blot. Only miR-34c showed a testis-specific expression pattern, the others being more or less ubiquitously expressed (Fig. 1A). Moreover, a high level of miR-34c was present in adult pachytenes spermatocytes and round spermatids. This microRNA, miR-34c, belongs to a family of evolutionarily conserved miRNAs (miR-34a, miR-34b, and miR-34c), whose expression is regulated by the tumor suppressor protein p53 and are implicated in the negative control of the cell cycle (Corney et al. 2007; Dutta et al. 2007; He et al. 2007; Raver-Shapira et al. 2007; Lujambio et al. 2008; Markey and Berberich 2008; Ji et al. 2009), cell cycle arrest (Paris et al. 2008; Sun et al. 2008), senescence (He et al. 2007; Tazawa et al. 2007; Kumamoto et al. 2008; Christoffersen et al. 2009), and apoptosis (Bommer et al. 2007; Chang et al. 2007; Raver-Shapira et al. 2007; Welch et al. 2007; Yamakuchi et al. 2008). However, the involvement of miR-34c in the differentiation process has not yet been reported.
FIGURE 1.
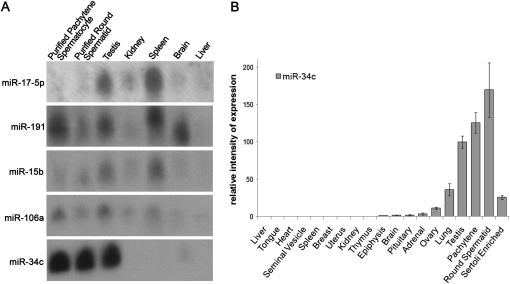
miR-34c expression profile in different mouse tissues. (A) Northern blot analysis of miR-34c, miR-106a, miR-15b, miR-191, and miR-17-5p expression in different tissues. Thirty micrograms of total RNA were loaded in each well and hybridized with the indicated probes. (B) RT-qPCR analysis of miR-34c expression in various tissues. A value of 100 was given to the miR-34c relative expression in testis. Two different RT reactions were carried out three times and used to calculate the mean and standard error (SEM).
We examined the expression of mature miR-34c in a larger panel of mouse tissues following the method described by Raymond et al. (2005). The highest expression was noticed in the testis, with a weaker one in the lungs (30% of testis expression) and almost no expression in other organs (Fig. 1B). These results are in agreement with previous reports. In these studies that used microarray profiling and RT-qPCR on pre-miRNA, miR-34a was shown to be ubiquitous and less expressed than miR-34c in testis and miR-34b to be similarly expressed to miR-34c, but to a lower level, except in fallopian tubes where miR-34b is more highly expressed (Baskerville and Bartel 2005; Dutta et al. 2007). We focused our attention on testis cells and we noticed a high miR-34c expression in purified pachytene spermatocytes and round spermatids and only a weak one in the enriched Sertoli cell fraction (contamination by germ cells was estimated at 30%) (Fig. 1B; see Materials and Methods). This suggested that miR-34c was germ cell specific or very weakly expressed in somatic testis cells.
To test this hypothesis, we examined different mouse models with various germ cell contents. Heterozygous c-Kit mice showed a reduced spermatogenesis proportional to the haplo deficiency of the receptor c-Kit. In this case, miR-34c expression in testis was also reduced when compared with the control (Fig. 2A). This suggested that miR-34c expression was closely correlated to the germ cell number. Moreover, wild-type (WT) mice were treated with busulfan, a cytotoxic drug known to deplete the germ cells in the testis. Busulfan treatment led to a miR-34c expression decrease in the same order of magnitude as the germ cell loss (95%) (Fig. 2B). Since germ cells in these two models were the direct target of either the genetic impairment or the chemical treatment, we therefore hypothesized that miR-34c is mainly expressed in germ cells.
FIGURE 2.
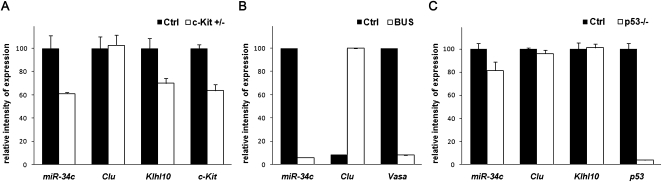
miR-34c is expressed preferentially in germ cells. (A) miR-34c expression in adult mouse c-Kit ± testis. miR-34c, Clusterin (Sertoli marker), Klhl10 (germ cell marker), and c-Kit RNA expressions were tested by RT-qPCR. (B) miR-34c expression in busulfan-treated adult mouse testis. miR-34c, Clusterin (Sertoli marker) and Vasa (germ cell marker) RNA expressions were tested by RT-qPCR. (C) miR-34c expression in p53-deficient mouse testis. miR-34c, Clusterin (Sertoli marker), Klhl10 (germ cell marker), and p53 RNA expressions were tested by RT-qPCR. For each experiment, a value of 100 was given to the relative gene expression in the wild-type mouse testis (Ctrl). Three different RT reactions were carried out three times and used to calculate the mean and standard error (SEM).
We then performed in situ hybridization on adult mouse testis sections with a locked nucleic acid (LNA) miR-34c-specific probe (Exiqon). In adult mouse seminiferous tubule sections, a significant signal was observed in all tubules and was more intense in late stages of the seminiferous epithelium (due to the presence of late pachytene spermatocytes) (Fig. 3, A1). No signal was noticed in interstitial tissues (testis somatic cells). At a higher magnification, we could see clearly that pachytene spermatocytes and round spermatids were stained (Fig. 3, A2). Moreover, the intensity of the staining was higher in late pachytene spermatocytes (belonging to stages IX to XI) than in young pachytene spermatocytes (present at stages I to VI) (Fig. 3, A1,A2). Furthermore, as illustrated in Figure 3, A3, Sertoli cells showed no staining; this latter observation confirmed that somatic cells of the testis did not express miR-34c in contrast to germ cells. Altogether, these results suggested that miR-34c is highly expressed in testis and is germ cell specific.
FIGURE 3.
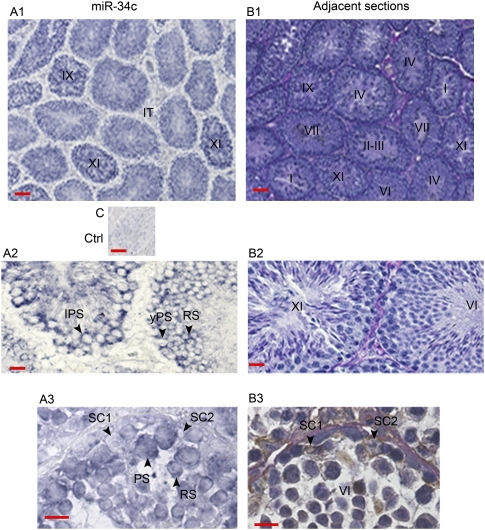
Localization of miR-34c by in situ hybridization of adult mouse seminiferous tubule sections. (Section A1) A significant signal was observed in all tubules and was more intense in late stages (IX and XI) of the seminiferous epithelium. No signal was noticed in interstitial tissue (IT). (Section A2) The expression of miR-34c was localized in meiotic cells: young pachytene spermatocytes (yPS), late pachytene spermatocytes (LPS), and in round spermatids (RS). (Section A3) No signal was observed in Sertoli cells (SC). Two Sertoli cells (SC1 and SC2) are shown on the section treated with miR-34c probe and on the control adjacent section. Note that when the nucleus of a SC is seen on a section, only a small part of it is present on the adjacent section. (Section C) Background in the absence of the miR-34c-specific probe. The stages of the seminiferous epithelium cycle (roman numerals) were identified from adjacent sections B1, B2, and B3. Adjacent sections B1 and B2) were stained with PAS and hematoxylin. Adjacent section B3 was incubated with an antivimentin antibody (a somatic cell marker) and stained with PAS and hematoxylin. Magnification: sections A1, B1, and C, X100; sections A2 and B2, ×400; and sections A3 and B3, ×1000. Bars in sections A1, B1, and C represent 50 μm. Bars in sections A2, B2, A3, and B3 represent 10 μm.
Since miR-34 family expression is regulated by p53 and its functions could be closely interwoven with those of the tumor suppressor, we wondered whether this was also the case in the testis. We compared miR-34c expression between p53-deficient mice testis and its normal counterpart. We observed only a slight miR-34c expression decrease (20%) (Fig. 2C), suggesting that miR-34c expression, at least in testis, is not only p53 dependent.
miR-34c overexpression induced a shift of the HeLa expression profile
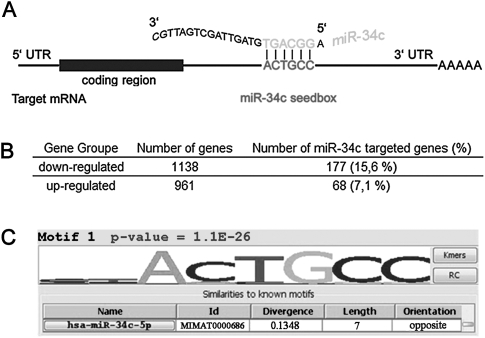
Lim et al. (2005) have shown that the overexpression of a cell type-specific microRNA could shift the expression profile of the recipient cells toward that of the cell originally expressing the microRNA. Since miR-34c appeared to be germ specific, to explore the biological effects of miR-34c we wondered whether miR-34c overexpressing HeLa cells could present a modified transcriptome. HeLa cells were therefore transfected with a miR-34c miRNA precursor (Ambion). Recent work has established that the miR-34 family negatively controls the cell cycle and induces apoptosis in p53-expressing cells (Bommer et al. 2007; Chang et al. 2007; Corney et al. 2007; He et al. 2007; Raver-Shapira et al. 2007; Tarasov et al. 2007; Kumamoto et al. 2008; Paris et al. 2008; Rokhlin et al. 2008). Two days after RNA transfection, we also noticed that miR-34c overexpression in HeLa cells led to a substantial inhibition of growth and to a mild apoptosis, as already reported for p53-deficient cells (data not shown). These positive preliminary controls being completed, DNA array gene profiling was performed to examine the transcriptomes of these cells. We analyzed their mRNA profiles (fold change of 1.5 between control and miR-34c-expressing cells) and noticed that 1138 genes showed a decreased expression and 961 an increased expression (see Supplemental Data Table S3). This list of genes (up and down) was then compared with the list of potential miR-34c target genes predicted by the miRGen target interface software. Significant enrichment of miR-34c target genes was noticed in the down-regulated genes (15.6%, P-value <10−4) when compared with the up-regulated genes (7.1%, P-value = 0.58) and to the unmodified genes (6.5%) (Fig. 4B). We also performed another analysis with the Allegro algorithm, which examines the frequency of hexamer nucleotides present in a list of genes. The hexamer ACTGCC, which is the target sequence recognized by miR-34c (Fig. 4A, miR-34c seedbox), was the most statistically enriched motif present in the down-regulated gene 3′UTRs (Fig. 4C); no significant motif was found in the up-regulated gene 3′UTRs. These data confirmed that miR-34c exerted its effects by down-regulating the expression of its target genes in our model. To gain further insight into the global effects of miR-34c ectopic expression in HeLa cells, we examined the Gene Ontology Classification of the up- and down-regulated genes with annotation, mapping, expression, and network (AMEN) software. When we tested Gene Ontology (GO) terms and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, we found an enrichment of “cell cycle” genes in the list of down-regulated genes, as already shown in cells overexpressing miR-34 microRNAs. The only significant pathway identified by KEGG was the “p53-signaling pathway,” as expected for a miRNA acting downstream from p53 (Bommer et al. 2007; Chang et al. 2007; He et al. 2007). In the list of genes implicated in cell division and down-regulated here, we noticed CCND3, CCNG1, CCNBB1, CCNC, CCNE1, CDK4, CDK6, E2F5, FOS, CDC2, AURKA, and AURKB; most of these are direct targets of miR-34c except AURKA and AURKB. Furthermore, in the up-regulated genes there existed a subset of known p53-target genes including BAX, SFN (14-3-3σ), PPM1D, PUMA, SESN1, and ZMAT3.
FIGURE 4.
Sites complementary to the miR-34c targeted sequence are enriched in the 3′UTRs of down-regulated transcripts. (A) Schematic representation of miR-34c base pairing with its targets. (B) miRGen results: miRGen (http://www.diana.pcbi.upenn.edu/cgi-bin/miRGen/v3/Targets.cgi) is a tool that provides a list of genes potentially recognized by a miRNA. We compared the miR-34c targeted gene list provided by miRGen with our experimental lists (DNA arrays on miR-34c overexpressing cells) of up- and down-regulated genes. The down-regulated gene list harbored more miR-34c regulated genes in comparison with up-regulated genes (15.6% versus 7.1%). (C) Allegro results: overrepresented motifs in the 3′UTRs of down-regulated genes. We submitted the miR-34c down- and up-regulated gene lists to the allegro software (http://acgt.cs.tau.ac.il/allegro/). Allegro is a software tool for discovering motifs statistically overrepresented in gene 3′UTR. The only noticeable motif highlighted by this analysis was a sequence corresponding to the miR-34c seedbox (ACTGCC) in the down-regulated gene list, whereas no particular motif was recognized in the up-regulated gene list.
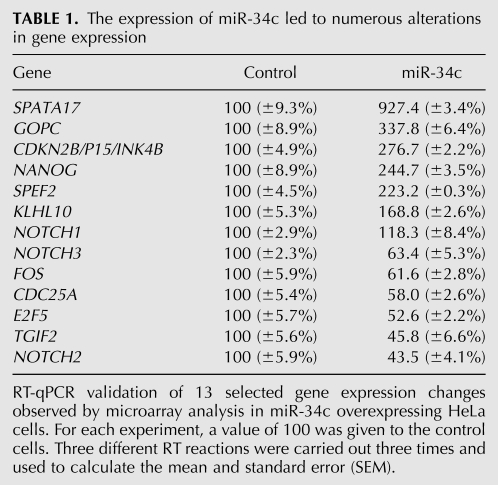
The AMEN suite of tools was then used to identify genes with a specific or preferential expression in a single tissue (Chalmel et al. 2007; Chalmel and Primig 2008). The most noticeable feature was that the percentage of genes preferentially expressed in testis diminished in the list of miR-34c down-regulated genes (2%; P-value <10−4) (Fig. 5; Supplemental Data Table S3) when compared with the unmodified ones (5.1%), whereas the percentage of genes preferentially expressed in testis remained unchanged in the list of miR-34c up-regulated genes (5%; P-value = 0.92). This meant that the list of genes down-regulated by miR-34c had proportionally less genes preferentially expressed in testis, and thus had a more distant relationship to the testis profile than the up-regulated genes or unmodified genes. In similar experiments, Lim et al. (2005) observed similar results for miR-1 and miR-124 and suggested that it could be envisaged as a transcriptome shift. In our case, we considered that the shift was weak but of interest in the context of germ cell differentiation. This is illustrated in Figure 5 by the heat map showing the expression of miR-34c up- and down-regulated genes in different organs and tissues. Although up-regulated genes were not different from the unmodified ones in terms of the percentages of those preferentially expressed in testis genes, we nevertheless noticed some interesting genes that we analyzed further by RT-qPCR. SPATA17, GOPC, SPEF2, and KLHL10 are germ-specific genes mostly expressed in the late stages of spermatogenesis, and were up-regulated in these miR-34c HeLa cells (Table 1). We did not find any direct association between them and miR-34c, Further studies are required to uncover the reality, if any, of this link or to determine if it was only due to a stochastic process.
FIGURE 5.
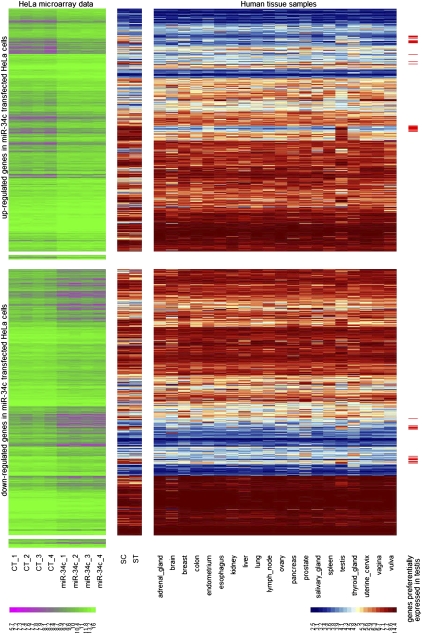
The expression of the up- and downregulated genes in purified germ cells and human tissue samples. A color-coded diagram (heatmap) of the up- and down-regulated genes is shown. Each column corresponds to a sample. Each line represents an Operon probe and its log2-transformed intensity signal: from pink (low signal) to green (high signal), as indicated in the bottom-left scale bar in the four control HeLa cells (CT_1–CT_4) and in the HeLa cells overexpressing miR-34c (miR-34c_1–miR-34c_4). Log2-transformed intensity signals of the affymetrix probe sets: from blue (low signal) to red (high signal), as indicated in the bottom-right scale bar, corresponding to the same gene were also displayed. These samples, hybridized on the Affymetrix GeneChip Human Genome U133 Plus 2.0 microarray, correspond to purified pachytene spermatocytes (SC) and round spermatids (ST), as well as 20 healthy human tissues including adrenal gland, brain, breast, colon, endometrium, esophagus, kidney, liver, lung, lymph node, ovary, pancreas, salivary gland, spleen, testis, thyroid gland, uterine cervix, vagina, and vulva. Probes showing a preferential expression in testis are shown with a red bar in the last column (right). Operon probes corresponding to genes not representated by affymetrix probe sets had no intensity signal using this technology (second and fourth sets of probes).
TABLE 1.
The expression of miR-34c led to numerous alterations in gene expression
TGIF2 and NOTCH2 are direct targets of miR-34c
Furthermore, among the down-regulated genes harboring a miR-34c recognition site in their 3′UTRs, we wondered whether some of these could be linked to pathways implicated in spermatogenesis control. Two down-regulated genes, TGIF2 and NOTCH2, appeared to be worthy of note. TGIF2 is an inhibitor of the TGFβ (TGFB1) pathway, and thus, its down-regulation enhances the signal elicited by this factor (Melhuish et al. 2001; Massague and Gomis 2006). Although the TGFβ signaling pathway was not highlighted by KEGG analysis, we nevertheless noticed that the expression of genes known to be targeted by TGFβ was modified by miR-34c ectopic expression in HeLa cells: an increased expression of CDKN2B/P15 and NANOG, with decreased expression of CDC25A (Table 1; Iavarone and Massague 1997; Massague and Gomis 2006; Xu et al. 2008). Combined with the fact that TGFβ plays a major role during spermatogenesis (Damestoy et al. 2005; Itman and Loveland 2008), we speculated that the potential role of miR-34c during this process could be closely associated with the TGFβ pathway, and so, we focused our attention on TGIF2. We confirmed by RT-qPCR, that TGIF2 gene expression was indeed down-regulated in miR-34c overexpressing cells (Fig. 6A), and we investigated its pattern of expression in testis. As illustrated in Figure 6B, Tgif2 was expressed in mouse testis, but principally in somatic cells. Tgif2 was virtually undetectable in round spermatids and pachytene spermatocytes, where conversely, miR-34c, as already mentioned, was highly expressed. We then explored whether miR-34c regulated TGIF2 directly. We fused a TGIF2 3′UTR to luciferase. Cotransfection with miR-34c (pre-miR-34c miRNA precursor) decreased luciferase activity in contrast to control miRNA (pre-miR miRNA precursor RNA negative control). Mutation in the miR-34c recognition site had a significant, although partial, effect (Fig. 6C). This could indicate the presence of other relevant cryptic miR-34c seed boxes or a combination of both direct and indirect effects, as already observed for other miR-34 targets such as CDK6 and CCNE2 (He et al. 2007; Lujambio et al. 2008; Sun et al. 2008). However, TGIF2 was clearly regulated by miR-34c.
FIGURE 6.
TGIF2 and NOTCH2 are direct targets of miR-34c. (A) We validated TGIF2 down-regulation by miR-34c in HeLa transfected cells by RT-qPCR. (B) Tgif2 has a low expression in mouse germ cells but is highly expressed in testis somatic cells, in contrast with miR-34c. Tgif2 expression was tested in bulsulfan-treated mouse testis (BUS), pachytene spermatocytes (PACH), round spermatids (RS), and in control testis (Ctrl). RT-qPCR showed that Tgif2 is highly expressed in bulsulfan-treated mouse testis (depleted in germ cells) when compared with the control untreated testis and to purified germ cells. For each experiment, a value of 100 was given to the control cells (testis). Three different RT reactions were carried out three times and used to calculate the mean and standard error (SEM). (C) Luciferase assays demonstrated that miR-34c directly down-regulates TGIF2 by targeting the TGIF2 3′UTR. Luciferase reporter constructs containing full-length (WT) and mutated miR-34c seedbox (MUT) 3′UTR of TGIF2 were cotransfected with pre-miR-34c miRNA precursor (miR-34c) or pre-miR miRNA precursor RNA negative control (Ctrl) into HeLa cells. Luciferase activity was measured 48 h after transfection. Mutation in the miR-34c binding site (MUT) resulted in a reduction of the inhibition induced by miR-34c. A value of 100 was given to the control cells. Three different transfections were carried out and luciferase activity was measured twice for each reaction. (D,E,F) The same experiments were undertaken with NOTCH2 and showed similar results.
The NOTCH signaling pathway controls numerous cell-fate specification events: cell differentiation inhibition, proliferation control, and stem cell number (Lai 2004; Fortini 2009). Furthermore, NOTCH genes have already been described as potential targets of miR-34 miRNAs (Lewis et al. 2003; Ji et al. 2008; Ji et al. 2009). The expression of NOTCH2 was greatly decreased in miR-34c transfected cells (Fig. 6D). Altogether, these results suggested that the potential regulation of NOTCH2 by miR-34c could be relevant during spermatogenesis; the down-regulation of such genes being necessary for the differentiation process to occur. RT-qPCR experiments confirmed the reduced expression of NOTCH2 in this model (Fig. 6D) and also showed a reduced Notch2 expression in mouse germ cells (Fig. 6E). We then performed luciferase assays with the NOTCH2 3′UTR fused to luciferase, following the same methodology as the one used for TGIF2, and we demonstrated that NOTCH2 was clearly a direct target of miR-34c (Fig. 6F).
Enhancement of germ cell marker expression by miR-34c
In order to investigate the role of miR-34c in the germinal lineage, we used the recently established cESC system modified by the chicken Vasa homolog (Cvh) gene (Lavial et al. 2009). These pluripotent chicken embryonic stem cells overexpressing the RNA binding protein Cvh differentiate into germ cells when cultured under appropriate conditions (Lavial et al. 2009). Since miR-34c could be involved in the control of the late steps of spermatogenesis, we wondered whether its overexpression could influence the fate of these Cvh overexpressing cells. miR-34c was transfected in the presence or absence of Cvh. We then analyzed by RT-qPCR the expression of a representative set of genes specific to different lineages in the selected cells: germ cells, Klhl10, Dnd1 (Deadend), Plzf, Piwi, Stra8, Sycp3, and Gcnf; germ and other lineages, Kit, Nanog, Pou5f1, Tudor, and Sdf1; and other lineages, Gata6, Sox7, Gsc, Cga, Hnf3, Lrh1, Sox4, Cxcr4, Fgf2, Gmnn, and Sox2.
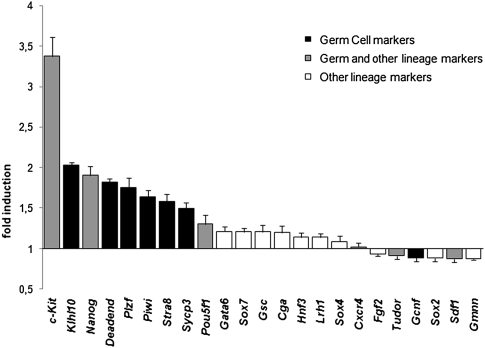
It appeared that miR-34c alone was unable to modify the expression level of these tested genes including germ cell markers (data not shown). However, when transfected with Cvh, miR-34c overexpression emphasized the germ cell nature of these cells (Fig. 7). Indeed, the expression levels of the somatic markers (Gata6, Sox7, Gsc, Cga, Hnf3, Lrh1, Sox4, Cxcr4, Fgf2, Gmnn, and Sox2) were not notably affected. In contrast, most germ cell markers were up-regulated: Kit, Klhl10, Deadend, Plzf, Piwi, Stra8, and Sycp3. Expression of the pluripotent cPouv and cNanog genes was also up-regulated as expected for genes that appear in numerous systems to be indispensable to germ cell commitment. In particular, the Nanog gene could be seen as playing an important role in germ cell formation, as already has been suggested (Chambers et al. 2007). Altogether, these results suggest that miR-34c alone is not able to induce the full germinal program, but rather plays a key role by reinforcing the germ cell phenotype.
FIGURE 7.
miR-34c overexpression enhanced the germ profile of cells already engaged in the first steps of the germ lineage. Chicken ES cells overexpressing chicken Vasa homolog (Cvh) and cultured under appropriate conditions differentiate toward the first steps of spermatogenesis. We cotransfected miR-34c in these Cvh overexpressing ES chicken cells cultured under the same conditions and analyzed the expression profile of selected differentiation markers. We analyzed the expression level of these markers by RT-qPCR in the miR-34c & Cvh co-transfected cells versus the Cvh control cells. A value of 1 was given to the expression level of these markers in the Cvh ES cells and the resulting up or down-regulation of these markers was noted on this graph.
DISCUSSION
The aim of this study was to identify new miRNAs that could be involved in spermatogenesis. We focused our attention on the expression profile of the miRNAs expressed in mouse pachytene spermatocytes and in round spermatids as these are two key steps in male germ cell differentiation.
By microarray experiments, we identified miR-34c as being highly expressed in germ cells and we showed that this expression was specific to these cells by Northern blotting, in situ hybridization, and the use of different mouse animal models. miR-34c belongs to a family of evolutionarily conserved miRNAs (miR-34a, miR-34b, and miR-34c) that have recently been shown to be involved in the negative control of the cell cycle (Corney et al. 2007; Dutta et al. 2007; He et al. 2007; Raver-Shapira et al. 2007; Lujambio et al. 2008; Markey and Berberich 2008; Ji et al. 2009), including cell cycle arrest (Paris et al. 2008; Sun et al. 2008), cellular senescence (He et al. 2007; Tazawa et al. 2007; Kumamoto et al. 2008; Christoffersen et al. 2009), and induction of apoptosis (Bommer et al. 2007; Chang et al. 2007; Raver-Shapira et al. 2007; Welch et al. 2007; Yamakuchi et al. 2008). In these systems, some targets directly down-regulated by miR-34s have also been experimentally validated, and it appears that most of them are involved in pathways related to these cellular processes: NOTCH1, MET, E2F1-3, MYC, MYCN, CDK4, CDK6, CCND1, and CCNE2, and cell cycle, SIRT1, BCL2, and apoptosis, and viral oncoprotein E6 (Lewis et al. 2003; Bommer et al. 2007; He et al. 2007; Tazawa et al. 2007; Welch et al. 2007; Kong et al. 2008; Leucci et al. 2008; Lujambio et al. 2008; Migliore et al. 2008; Sun et al. 2008; Wei et al. 2008; Yamakuchi et al. 2008; Li et al. 2009; Wang et al. 2009). Furthermore, the transcription of miR-34a, miR-34b, and miR-34c is regulated by p53, and these studies suggest that miR-34 miRNAs could constitute central mediators of p53 functions (Bommer et al. 2007; Chang et al. 2007; Corney et al. 2007; He et al. 2007; Raver-Shapira et al. 2007; Tarasov et al. 2007; Paris et al. 2008; Rokhlin et al. 2008). It is worthy of note that miR-34c has already been identified as a miRNA that is highly expressed in mature testis when compared with its immature counterpart (Yan et al. 2007, 2009). Pachytene spermatocytes and round spermatids are meiotic germ cells that are only produced in mature testis; thus, our results further support these observations. However, an interesting point raised by our observations is that p53 is not the sole transcriptional regulator of miR-34c in germ cells as miR-34c expression is only slightly reduced in p53-deficient mouse testis, and thus could be considered as p53-independent in this context. Unraveling the factors necessary to the transcriptional regulation of miR-34c in germ cells could reveal mechanisms used by p53 to become the guardian of the genome during evolution.
MicroRNAs play important gene-regulatory roles by targeting mRNAs to direct their post-transcriptional repression (degradation or translation inhibition) or by regulating transcription (Ambros 2008; Bartel 2009; Younger et al. 2009). Although in animals miRNAs are believed to act mainly through translational repression rather than mRNA degradation, microarray analysis of miRNA transfected cells has nevertheless revealed important clues to the understanding of miRNA functions and has shown that some miRNAs can down-regulate large amounts of mRNA. Lim et al. (2005) transfected HeLa cells with tissue-specific miRNAs and showed that this shifted the expression profile of these HeLa cells toward that of cells originally expressing the miRNA. Therefore, to gain insight into miR-34c function, we performed such experiments. We analyzed the expression profile of these miR-34c transfected HeLa cells. We noticed an overrepresentation of genes harboring a miR-34c seed box in the 3′UTRs of down-regulated genes, whereas no change was observed in the up-regulated ones. This result confirms that miR-34c is a negative regulator of gene expression and down-regulates the expression of many genes. Then, we tested the lists of genes for potential enrichment of genes implicated in particular pathways using the AMEN suite of tools. As already reported, we found several genes involved in cell cycle regulation (CCND3, CCNG1, CCNB1, CCNC, CCNE1, CDK4, CDK6, E2F5, FOS, CDC2, AURKA, and AURKB), and also genes belonging to the p53 pathway (BAX, SFN [14-3-3σ], PPM1D, PUMA, SESN1, and ZMAT3). No statistically original pathway that could be linked to spermatogenesis had been identified by these different analyses. To investigate the potential influence of miR-34c ectopic expression on HeLa cell fate, we also used AMEN to detect groups of significantly coexpressed genes, and thus similar patterns of expression through different lineages (Chalmel et al. 2007; Chalmel and Primig 2008). This analysis emphasized that the list of down-regulated genes identified in this study presented a lower percentage of genes preferentially expressed in testis (2%) than up-regulated and unchanged genes (5%). Lim et al. (2005), using the same cells (HeLa) with other miRNAs (miR-1 and mR-124), also observed a shift in the down-regulated gene expression profile (muscle and brain, respectively), and reached the conclusion that tissue-specific miRNA ectopic expression could influence the differentiation state of the recipient cells. We thus considered that miR-34c exerts its function through preferential down-regulation of genes less expressed in testis.
We then tried to decipher whether genes whose down-regulation by miR-34c could be relevant to spermatogenesis were present in the list of miR-34c down-regulated genes. Apart from genes directly involved in the cell control, we focused our attention on two genes: NOTCH2 and TGIF2. TGIF2 is a transcriptional repressor and is an inhibitor of the TGFβ pathway; its down-regulation enhances the signal elicited by this factor (Melhuish et al. 2001; Massague and Gomis 2006). Moreover, we noticed that the expression of genes known to be targeted by TGFβ was modified by miR-34c ectopic expression in HeLa cells: an increased expression of CDKN2B/P15 and NANOG, and a decreased expression of CDC25A (Iavarone and Massague 1997; Massague and Gomis 2006; Xu et al. 2008). Furthermore, TGFβ plays a major role during spermatogenesis, particularly at the end of the meiotic step (Damestoy et al. 2005; Itman and Loveland 2008). In this study, we showed that TGIF2 is down-regulated by miR-34c, is a direct target through a miR-34c seed box localized in its 3′UTR, and has a low expression in germ cells, as illustrated by its expression pattern in testis. We speculate that a possible role for miR-34c during spermatogenesis could be closely associated with the TGFβ pathway via the down-regulation of TGIF2.
The NOTCH signaling pathway controls numerous cell-fate specification events: cell differentiation inhibition, proliferation control, and stem cell number (Lai 2004; Fortini 2009). Furthermore, NOTCH genes have already been described as potential targets for miR-34 miRNAs (Lewis et al. 2003; Ji et al. 2008; Ji et al. 2009), and NOTCH2, which is hardly expressed in germ cells, could play an important role during testis somatic cell differentiation (Dirami et al. 2001; Mori et al. 2003; Tang et al. 2008). All these data led us to hypothesize a possible connection between miR-34c, NOTCH2 down-regulation, and germ cell differentiation. We demonstrated that NOTCH2 was a direct target of miR-34c and was expressed at a lower level than the other members of the family in this system. We postulated that the down-regulation of NOTCH2 by miR-34c could be necessary for the germ differentiation process to occur by blocking the capacity of germ cells to self-renew.
The conclusion we draw from the above results is that miR-34c seems to display antiproliferative properties by directly down-regulating genes involved in various pathways closely related to this process, and that miR-34c preferentially down-regulates genes less expressed in germ cells when compared with other organs. Altogether, this could constitute an explanation of its role during spermatogenesis.
As miR-34c is mainly expressed in the late stages of meiosis (pachytene spermatocytes and round spermatids), and is likely to influence germ cell fate during this period, we have also investigated its role during germ cell induction.
Recently, an in vitro cellular model reproducing the first steps of germ cell differentiation was generated by overexpressing the RNA binding protein Ddx4/Cvh (into chicken embryonic stem cells [cESC]) (Lavial et al. 2009). When coexpressed in the presence of the Cvh gene, miR-34c was able to enhance the germ cell character of the recipient cells by increasing the expression of germ cell markers. Thus, miR-34c would enhance the germ phenotype of cells already engaged in the germ cell lineage.
In the future, the generation of miR-34c invalidated mice should be of primary importance to ascertain the role of this miRNA during spermatogenesis and to explore the regulated biochemical pathways. Furthermore, in light of our results, miR-34c is a very promising candidate gene for testing in male germ cell cancer and male sterility.
MATERIALS AND METHODS
Elutriation
Pachytene spermatocytes, round spermatids, and Sertoli cells were prepared, separated, and analyzed as previously described by Weiss et al. (1997).
miRNA Array preparation and hybridization
Total RNA of spermatocytes and spermatids was extracted by TRIzol Reagent (Invitrogen). miRNA were purified from 100 μg of total RNA by the FlashPAGE method (Ambion). Cy5 or Cy3 labeling was performed according to the manufacturer's protocol (Ambion). Dual swap hybridization of the two samples was performed for 16 h at 45°C, on slides spotted by the DTAMB with the mirVana miRNA Probe Set (Ambion).
Genome microarray hybridizations
The Operon Human Genome Array (4.0) contained 35,035 oligonucleotide probes printed on epoxide glass microscope slides (www.operon.com). Total RNA was purified from the control and miR-34c transfected HeLa cells 48 h after transfection by using the TRIzol procedure (Invitrogen); RNA quality was checked by BioAnalyzer analysis (Agilent); and antisense RNA (aRNA) amplification (Van Gelder et al. 1990) was performed according to the Ambion protocol.
A total of 1.5 μg of combined dye-labeled aRNA (miR-34c versus control) was hybridized competitively to microarray slides for 16 h at 67°C in hybridization buffer (Agilent) under Agilent Gasket slide coverslips (Agilent hybridization chamber and Agilent hybridization oven). Each experiment was repeated twice with a switch of fluorescent Cy3/Cy5 labels to account for dye effects leading to four data points per hybridization spot.
miRNA and genome array scanning
After several post-hybridization washes, fluorescent microarray images were collected for both Cy3 and Cy5 by scanning the glass slides with a GenePix 4000B microarray scanner (Axon Instruments). Image intensity data were extracted and analyzed with GenePixPro6.0 microarray analysis software. After visual inspection, spots of insufficient quality were flagged and excluded from further analysis, and GenePix data files (gpr format) were generated.
Statistical analysis and interpretation of microarray data
The AMEN software was used for array data preprocessing, analysis, and visualization (Chalmel and Primig 2008). The “normexp” (Ritchie et al. 2007) and the “quantile–quantile” methods were used on the two-color array data for background correction and “between-array” normalization preprocesses, respectively. To identify differentially expressed genes, the empirical Bayes method (F-test), implemented in the LIMMA package and adjusted with the false discovery rate (FDR) method, was employed (fold change ≥1.5 and FDR ≤0.05). The differentially expressed genes were classified into two groups according to their expression profiles using the k-means clustering algorithm (with k = 2). Significantly enriched GO terms associated with the clusters were identified by using a P-value adjusted with an FDR ≤0.05. The minimal number of genes in a cluster associated with a given annotation term was set at ≥5 and the Gene Ontology information rate cutoff was set at ≥0.1.
In situ hybridization of mouse seminiferous tubules
Adult mouse testes were fixed either in PAF 4% or in AFA and then dehydrated and embedded in paraffin. Three sequential sections (5 μm) were prepared: the first one for hybridization with the miR-34c probe, the second for a histological study, and the third for hybridization with control. In situ hybridization was then performed according to the protocol supplied by Exiqon (nonradioactive in situ hybridization on paraffin sections using DIG-labeled mercury detection probes). miR-34c and a control probe (mercury LNA) were purchased from Exiqon. Some sections for histology were stained with periodic acid-Schiff and counterstained with hematoxylin. An immunocytochemical reaction against vimentin (solely expressed by somatic cells) was performed on other sections to discriminate between Sertoli cells and germ cells, as previously described (Perrard et al. 2007). 3,3′-Diaminobenzidine (DAB) was used as a chromogen, and then sections were stained with periodic acid-Schiff and counterstained with hematoxylin.
Northern blot analysis
A total of 30 μg of each total RNA sample was separated and hybridized as described by Kaddar et al. (2009). The probe sequences are provided in Supplemental Table S1.
Reverse transcription quantitative PCR
RT-qPCR was performed on the MXP-3000P QPCR-system (Stratagene/AGILENT) using Mix-Quantitect SYBR Green kit (Qiagen). All reactions were run in triplicate, and gene expression levels were calculated using the ΔΔCT method (http://www.gene-quantification.info) with the ribosomal gene Rs17 as a reference gene. For RT-qPCR on mature miRNA, we followed the protocol described by Raymond et al. (2005) (see Supplemental Table S1 for primer sequences).
miR-34c expression vector construction
Human DNA was used to amplify the pre-miR-34c (336 nt long) with the following primers (F: ACAGTCTCATGCCAGGAAAGC; R: GCACATTGATGATGCACAGGC). This pre-miR-34c region was then substituted for the luciferase gene in a pMIR-REPORT luciferase vector (Ambion).
Cell culture and RNA transfection
HeLa cells were cultured in Dulbecco's Modified Eagle Medium (DMEM) containing 10% heat-inactivated fetal bovine serum (Invitrogen). HeLa cells (3 × 105) were seeded in a six-well plate and transfected 1 d after plating with a pre-miR-34c miRNA precursor (100 nM) or a pre-miR miRNA precursor RNA negative control #1 (both from Ambion) using lipofectamine2000 reagent (Invitrogen). The transfection medium was replaced 5 h later by fresh growth medium, and the cells were harvested 48 h later.
cESC and GFP∷CVH modified cESC were maintained and transfected as previously described (Lavial et al. 2007, 2009) with either pMIR-REPORT-miR-34c expression vector or empty pMIR-REPORT as a control. The resulting clones obtained after neomycin (250 μg/mL) and puromycin (0.5 μg/mL) selection over 7 d were harvested, pooled, and grown in conditions inducing germ cell differentiation as previously reported (Lavial et al. 2007, 2009).
Luciferase reporter assay
The 3′UTRs of human TGIF2 (NM_021809.5, nucleotides 888–1426) or NOTCH2 (NM_024408.2, nucleotides 9444–10,522) encompassing the conserved miR-34c response elements (respectively, nucleotides 977–983 and 10,138–10,145) were cloned into the pMIR-REPORT vector (Ambion), between the SacI and HindIII sites, immediately downstream from the luciferase gene (WT). The miR-34c response element (CACTGCC), was mutated in both 3′UTRs using appropriate primers (MUT). The primers are described in Supplemental Table S1. A total of 400 ng of each pMIR-REPORT construct was cotransfected with pre-miR-34c miRNA precursor (miR-34c) or pre-miR miRNA precursor RNA negative control #1 (Ctrl) (50 nM) (Ambion) into HeLa cells in a 12-well plate using lipofectamine2000 (Invitrogen). After 48 h, firefly luciferase activity was measured on cell lysates with the dual-luciferase reporter system (Promega), according to the manufacturer's instructions.
Target prediction
The miR-34c target list was generated by miRGen (http://www.diana.pcbi.upenn.edu/cgi-bin/miRGen/v3/Targets.cgi), a target interface providing access to unions and intersections of four widely used target prediction programs and experimentally supported targets from TarBase (Megraw et al. 2007).
In addition, a motif analysis was performed with Allegro software, a tool for discovering motifs statistically overrepresented in gene 3′UTRs (http://acgt.cs.tau.ac.il/allegro/) (Linhart et al. 2008).
Animal models
To deplete germ cells from testis, Busulfan dissolved in DMSO (40 mg/kg) was injected intraperitoneally and the testes harvested 1 mo later. c-Kit mice have been described previously by Geissler et al. (1988). p53 deficient mice were generated following the method of Donehower et al. (1992).
SUPPLEMENTAL MATERIAL
Supplemental material can be found at http://www.rnajournal.org.
ACKNOWLEDGMENTS
We are grateful to M.J. Asensio, M.A. Chauvin, M. Billaud, Y. Mikaelian, T. Kaddar, R. Haber, B. Gibert, A. Rascalou, P. Monget, and the DTAMB for their help. Funding was provided in part by the Crédits incitatifs PHASE-INRA. F.B. was a recipient of a fellowship from the Organon-Schering-Plough PhD program and F.L. was a recipient of a fellowship from the PHASE INRA Department.
Footnotes
Article published online ahead of print. Article and publication date are at http://www.rnajournal.org/cgi/doi/10.1261/rna.1963810.
REFERENCES
- Ambros V. The evolution of our thinking about microRNAs. Nat Med. 2008;14:1036–1040. doi: 10.1038/nm1008-1036. [DOI] [PubMed] [Google Scholar]
- Bannister SC, Tizard ML, Doran TJ, Sinclair AH, Smith CA. Sexually dimorphic microRNA expression during chicken embryonic gonadal development. Biol Reprod. 2009;81:165–176. doi: 10.1095/biolreprod.108.074005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartel DP. MicroRNAs: Target recognition and regulatory functions. Cell. 2009;136:215–233. doi: 10.1016/j.cell.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baskerville S, Bartel DP. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA. 2005;11:241–247. doi: 10.1261/rna.7240905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, Zhai Y, Giordano TJ, Qin ZS, Moore BB, et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol. 2007;17:1298–1307. doi: 10.1016/j.cub.2007.06.068. [DOI] [PubMed] [Google Scholar]
- Chalmel F, Primig M. The annotation, mapping, expression, and network (AMEN) suite of tools for molecular systems biology. BMC Bioinformatics. 2008;9:86. doi: 10.1186/1471-2105-9-86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chalmel F, Rolland AD, Niederhauser-Wiederkehr C, Chung SS, Demougin P, Gattiker A, Moore J, Patard JJ, Wolgemuth DJ, Jegou B, et al. The conserved transcriptome in human and rodent male gametogenesis. Proc Natl Acad Sci. 2007;104:8346–8351. doi: 10.1073/pnas.0701883104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers I, Silva J, Colby D, Nichols J, Nijmeijer B, Robertson M, Vrana J, Jones K, Grotewold L, Smith A. Nanog safeguards pluripotency and mediates germline development. Nature. 2007;450:1230–1234. doi: 10.1038/nature06403. [DOI] [PubMed] [Google Scholar]
- Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ, et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007;26:745–752. doi: 10.1016/j.molcel.2007.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christoffersen NR, Shalgi R, Frankel LB, Leucci E, Lees M, Klausen M, Pilpel Y, Nielsen FC, Oren M, Lund AH. p53-independent upregulation of miR-34a during oncogene-induced senescence represses MYC. Cell Death Differ. 2009;17:236–245. doi: 10.1038/cdd.2009.109. [DOI] [PubMed] [Google Scholar]
- Corney DC, Flesken-Nikitin A, Godwin AK, Wang W, Nikitin AY. MicroRNA-34b and microRNA-34c are targets of p53 and cooperate in control of cell proliferation and adhesion-independent growth. Cancer Res. 2007;67:8433–8438. doi: 10.1158/0008-5472.CAN-07-1585. [DOI] [PubMed] [Google Scholar]
- Damestoy A, Perrard MH, Vigier M, Sabido O, Durand P. Transforming growth factor β-1 decreases the yield of the second meiotic division of rat pachytene spermatocytes in vitro. Reprod Biol Endocrinol. 2005;3:22. doi: 10.1186/1477-7827-3-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dirami G, Ravindranath N, Achi MV, Dym M. Expression of Notch pathway components in spermatogonia and Sertoli cells of neonatal mice. J Androl. 2001;22:944–952. doi: 10.1002/j.1939-4640.2001.tb03434.x. [DOI] [PubMed] [Google Scholar]
- Donehower LA, Harvey M, Slagle BL, McArthur MJ, Montgomery CA, Jr, Butel JS, Bradley A. Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature. 1992;356:215–221. doi: 10.1038/356215a0. [DOI] [PubMed] [Google Scholar]
- Dutta KK, Zhong Y, Liu YT, Yamada T, Akatsuka S, Hu Q, Yoshihara M, Ohara H, Takehashi M, Shinohara T, et al. Association of microRNA-34a overexpression with proliferation is cell type-dependent. Cancer Sci. 2007;98:1845–1852. doi: 10.1111/j.1349-7006.2007.00619.x. [DOI] [PubMed] [Google Scholar]
- Fortini ME. Notch signaling: The core pathway and its post-translational regulation. Dev Cell. 2009;16:633–647. doi: 10.1016/j.devcel.2009.03.010. [DOI] [PubMed] [Google Scholar]
- Geissler EN, Ryan MA, Housman DE. The dominant-white spotting (W) locus of the mouse encodes the c-kit proto-oncogene. Cell. 1988;55:185–192. doi: 10.1016/0092-8674(88)90020-7. [DOI] [PubMed] [Google Scholar]
- Hayashi K, Chuva de Sousa Lopes SM, Kaneda M, Tang F, Hajkova P, Lao K, O'Carroll D, Das PP, Tarakhovsky A, Miska EA, et al. MicroRNA biogenesis is required for mouse primordial germ cell development and spermatogenesis. PLoS One. 2008;3(3):e1738. doi: 10.1371/journal.pone.0001738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y, Xue W, Zender L, Magnus J, Ridzon D, et al. A microRNA component of the p53 tumor suppressor network. Nature. 2007;447:1130–1134. doi: 10.1038/nature05939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Z, Kokkinaki M, Pant D, Gallicano GI, Dym M. Small RNA molecules in the regulation of spermatogenesis. Reproduction. 2009;137:901–911. doi: 10.1530/REP-08-0494. [DOI] [PubMed] [Google Scholar]
- Hecht NB. Molecular mechanisms of male germ cell differentiation. Bioessays. 1998;20:555–561. doi: 10.1002/(SICI)1521-1878(199807)20:7<555::AID-BIES6>3.0.CO;2-J. [DOI] [PubMed] [Google Scholar]
- Iavarone A, Massague J. Repression of the CDK activator Cdc25A and cell-cycle arrest by cytokine TGF-β in cells lacking the CDK inhibitor p15. Nature. 1997;387:417–422. doi: 10.1038/387417a0. [DOI] [PubMed] [Google Scholar]
- Itman C, Loveland KL. SMAD expression in the testis: An insight into BMP regulation of spermatogenesis. Dev Dyn. 2008;237:97–111. doi: 10.1002/dvdy.21401. [DOI] [PubMed] [Google Scholar]
- Ji Q, Hao X, Meng Y, Zhang M, Desano J, Fan D, Xu L. Restoration of tumor suppressor miR-34 inhibits human p53-mutant gastric cancer tumorspheres. BMC Cancer. 2008;8:266. doi: 10.1186/1471-2407-8-266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ji Q, Hao X, Zhang M, Tang W, Yang M, Li L, Xiang D, Desano JT, Bommer GT, Fan D, et al. MicroRNA miR-34 inhibits human pancreatic cancer tumor-initiating cells. PLoS One. 2009;4:e6816. doi: 10.1371/journal.pone.0006816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaddar T, Rouault JP, Chien WW, Chebel A, Gadoux M, Salles G, Ffrench M, Magaud JP. Two new miR-16 targets: Caprin-1 and HMGA1, proteins implicated in cell proliferation. Biol Cell. 2009;101:511–524. doi: 10.1042/BC20080213. [DOI] [PubMed] [Google Scholar]
- Kedde M, Strasser MJ, Boldajipour B, Oude Vrielink JA, Slanchev K, le Sage C, Nagel R, Voorhoeve PM, van Duijse J, Orom UA, et al. RNA-binding protein Dnd1 inhibits microRNA access to target mRNA. Cell. 2007;131:1273–1286. doi: 10.1016/j.cell.2007.11.034. [DOI] [PubMed] [Google Scholar]
- Kong YW, Cannell IG, de Moor CH, Hill K, Garside PG, Hamilton TL, Meijer HA, Dobbyn HC, Stoneley M, Spriggs KA, et al. The mechanism of micro-RNA-mediated translation repression is determined by the promoter of the target gene. Proc Natl Acad Sci. 2008;105:8866–8871. doi: 10.1073/pnas.0800650105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumamoto K, Spillare EA, Fujita K, Horikawa I, Yamashita T, Appella E, Nagashima M, Takenoshita S, Yokota J, Harris CC. Nutlin-3a activates p53 to both down-regulate inhibitor of growth 2 and up-regulate mir-34a, mir-34b, and mir-34c expression, and induce senescence. Cancer Res. 2008;68:3193–3203. doi: 10.1158/0008-5472.CAN-07-2780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai EC. Notch signaling: control of cell communication and cell fate. Development. 2004;131:965–973. doi: 10.1242/dev.01074. [DOI] [PubMed] [Google Scholar]
- Lavial F, Acloque H, Bertocchini F, Macleod DJ, Boast S, Bachelard E, Montillet G, Thenot S, Sang HM, Stern CD, et al. The Oct4 homologue PouV and Nanog regulate pluripotency in chicken embryonic stem cells. Development. 2007;134:3549–3563. doi: 10.1242/dev.006569. [DOI] [PubMed] [Google Scholar]
- Lavial F, Acloque H, Bachelard E, Nieto MA, Samarut J, Pain B. Ectopic expression of Cvh (Chicken Vasa homolog) mediates the reprogramming of chicken embryonic stem cells to a germ cell fate. Dev Biol. 2009;330:73–82. doi: 10.1016/j.ydbio.2009.03.012. [DOI] [PubMed] [Google Scholar]
- Leucci E, Cocco M, Onnis A, De Falco G, van Cleef P, Bellan C, van Rijk A, Nyagol J, Byakika B, Lazzi S, et al. MYC translocation-negative classical Burkitt lymphoma cases: An alternative pathogenetic mechanism involving miRNA deregulation. J Pathol. 2008;216:440–450. doi: 10.1002/path.2410. [DOI] [PubMed] [Google Scholar]
- Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB. Prediction of mammalian microRNA targets. Cell. 2003;115:787–798. doi: 10.1016/s0092-8674(03)01018-3. [DOI] [PubMed] [Google Scholar]
- Li N, Fu H, Tie Y, Hu Z, Kong W, Wu Y, Zheng X. miR-34a inhibits migration and invasion by down-regulation of c-Met expression in human hepatocellular carcinoma cells. Cancer Lett. 2009;275:44–53. doi: 10.1016/j.canlet.2008.09.035. [DOI] [PubMed] [Google Scholar]
- Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM. Microarray analysis shows that some microRNAs down-regulate large numbers of target mRNAs. Nature. 2005;433:769–773. doi: 10.1038/nature03315. [DOI] [PubMed] [Google Scholar]
- Linhart C, Halperin Y, Shamir R. Transcription factor and microRNA motif discovery: The Amadeus platform and a compendium of metazoan target sets. Genome Res. 2008;18:1180–1189. doi: 10.1101/gr.076117.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lujambio A, Calin GA, Villanueva A, Ropero S, Sanchez-Cespedes M, Blanco D, Montuenga LM, Rossi S, Nicoloso MS, Faller WJ, et al. A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci. 2008;105:13556–13561. doi: 10.1073/pnas.0803055105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maatouk DM, Loveland KL, McManus MT, Moore K, Harfe BD. Dicer1 is required for differentiation of the mouse male germline. Biol Reprod. 2008;79:696–703. doi: 10.1095/biolreprod.108.067827. [DOI] [PubMed] [Google Scholar]
- Markey M, Berberich SJ. Full-length hdmX transcripts decrease following genotoxic stress. Oncogene. 2008;27:6657–6666. doi: 10.1038/onc.2008.266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marret C, Avallet O, Perrard-Sapori MH, Durand P. Localization and quantitative expression of mRNAs encoding the testis-specific histone TH2B, the phosphoprotein p19, the transition proteins 1 and 2 during pubertal development and throughout the spermatogenic cycle of the rat. Mol Reprod Dev. 1998;51:22–35. doi: 10.1002/(SICI)1098-2795(199809)51:1<22::AID-MRD3>3.0.CO;2-Y. [DOI] [PubMed] [Google Scholar]
- Massague J, Gomis RR. The logic of TGFβ signaling. FEBS Lett. 2006;580:2811–2820. doi: 10.1016/j.febslet.2006.04.033. [DOI] [PubMed] [Google Scholar]
- Megraw M, Sethupathy P, Corda B, Hatzigeorgiou AG. miRGen: A database for the study of animal microRNA genomic organization and function. Nucleic Acids Res. 2007;35:D149–D155. doi: 10.1093/nar/gkl904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melhuish TA, Gallo CM, Wotton D. TGIF2 interacts with histone deacetylase 1 and represses transcription. J Biol Chem. 2001;276:32109–32114. doi: 10.1074/jbc.M103377200. [DOI] [PubMed] [Google Scholar]
- Migliore C, Petrelli A, Ghiso E, Corso S, Capparuccia L, Eramo A, Comoglio PM, Giordano S. MicroRNAs impair MET-mediated invasive growth. Cancer Res. 2008;68:10128–10136. doi: 10.1158/0008-5472.CAN-08-2148. [DOI] [PubMed] [Google Scholar]
- Mori S, Kadokawa Y, Hoshinaga K, Marunouchi T. Sequential activation of Notch family receptors during mouse spermatogenesis. Dev Growth Differ. 2003;45:7–13. doi: 10.1046/j.1440-169x.2003.00670.x. [DOI] [PubMed] [Google Scholar]
- Paris R, Henry RE, Stephens SJ, McBryde M, Espinosa JM. Multiple p53-independent gene silencing mechanisms define the cellular response to p53 activation. Cell Cycle. 2008;7:2427–2433. doi: 10.4161/cc.6420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perrard MH, Vigier M, Damestoy A, Chapat C, Silandre D, Rudkin BB, Durand P. β-Nerve growth factor participates in an auto/paracrine pathway of regulation of the meiotic differentiation of rat spermatocytes. J Cell Physiol. 2007;210:51–62. doi: 10.1002/jcp.20805. [DOI] [PubMed] [Google Scholar]
- Plasterk RH. MicroRNAs in animal development. Cell. 2006;124:877–881. doi: 10.1016/j.cell.2006.02.030. [DOI] [PubMed] [Google Scholar]
- Raver-Shapira N, Marciano E, Meiri E, Spector Y, Rosenfeld N, Moskovits N, Bentwich Z, Oren M. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell. 2007;26:731–743. doi: 10.1016/j.molcel.2007.05.017. [DOI] [PubMed] [Google Scholar]
- Raymond CK, Roberts BS, Garrett-Engele P, Lim LP, Johnson JM. Simple, quantitative primer-extension PCR assay for direct monitoring of microRNAs and short-interfering RNAs. RNA. 2005;11:1737–1744. doi: 10.1261/rna.2148705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ritchie ME, Silver J, Oshlack A, Holmes M, Diyagama D, Holloway A, Smyth GK. A comparison of background correction methods for two-color microarrays. Bioinformatics. 2007;23:2700–2707. doi: 10.1093/bioinformatics/btm412. [DOI] [PubMed] [Google Scholar]
- Ro S, Park C, Sanders KM, McCarrey JR, Yan W. Cloning and expression profiling of testis-expressed microRNAs. Dev Biol. 2007;311:592–602. doi: 10.1016/j.ydbio.2007.09.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rokhlin OW, Scheinker VS, Taghiyev AF, Bumcrot D, Glover RA, Cohen MB. MicroRNA-34 mediates AR-dependent p53-induced apoptosis in prostate cancer. Cancer Biol Ther. 2008;7:1288–1296. doi: 10.4161/cbt.7.8.6284. [DOI] [PubMed] [Google Scholar]
- Sun F, Fu H, Liu Q, Tie Y, Zhu J, Xing R, Sun Z, Zheng X. Down-regulation of CCND1 and CDK6 by miR-34a induces cell cycle arrest. FEBS Lett. 2008;582:1564–1568. doi: 10.1016/j.febslet.2008.03.057. [DOI] [PubMed] [Google Scholar]
- Tang H, Brennan J, Karl J, Hamada Y, Raetzman L, Capel B. Notch signaling maintains Leydig progenitor cells in the mouse testis. Development. 2008;135:3745–3753. doi: 10.1242/dev.024786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tarasov V, Jung P, Verdoodt B, Lodygin D, Epanchintsev A, Menssen A, Meister G, Hermeking H. Differential regulation of microRNAs by p53 revealed by massively parallel sequencing: miR-34a is a p53 target that induces apoptosis and G1-arrest. Cell Cycle. 2007;6:1586–1593. doi: 10.4161/cc.6.13.4436. [DOI] [PubMed] [Google Scholar]
- Tazawa H, Tsuchiya N, Izumiya M, Nakagama H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc Natl Acad Sci. 2007;104:15472–15477. doi: 10.1073/pnas.0707351104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Gelder RN, von Zastrow ME, Yool A, Dement WC, Barchas JD, Eberwine JH. Amplified RNA synthesized from limited quantities of heterogeneous cDNA. Proc Natl Acad Sci. 1990;87:1663–1667. doi: 10.1073/pnas.87.5.1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R, Liu YP, van Duijse J, Drost J, Griekspoor A, et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell. 2006;124:1169–1181. doi: 10.1016/j.cell.2006.02.037. [DOI] [PubMed] [Google Scholar]
- Wang X, Wang HK, McCoy JP, Banerjee NS, Rader JS, Broker TR, Meyers C, Chow LT, Zheng ZM. Oncogenic HPV infection interrupts the expression of tumor-suppressive miR-34a through viral oncoprotein E6. RNA. 2009;15:637–647. doi: 10.1261/rna.1442309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei JS, Song YK, Durinck S, Chen QR, Cheuk AT, Tsang P, Zhang Q, Thiele CJ, Slack A, Shohet J, et al. The MYCN oncogene is a direct target of miR-34a. Oncogene. 2008;27:5204–5213. doi: 10.1038/onc.2008.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss M, Vigier M, Hue D, Perrard-Sapori MH, Marret C, Avallet O, Durand P. Pre- and postmeiotic expression of male germ cell-specific genes throughout 2-week cocultures of rat germinal and Sertoli cells. Biol Reprod. 1997;57:68–76. doi: 10.1095/biolreprod57.1.68. [DOI] [PubMed] [Google Scholar]
- Welch C, Chen Y, Stallings RL. MicroRNA-34a functions as a potential tumor suppressor by inducing apoptosis in neuroblastoma cells. Oncogene. 2007;26:5017–5022. doi: 10.1038/sj.onc.1210293. [DOI] [PubMed] [Google Scholar]
- Xu RH, Sampsell-Barron TL, Gu F, Root S, Peck RM, Pan G, Yu J, Antosiewicz-Bourget J, Tian S, Stewart R, et al. NANOG is a direct target of TGFβ/activin-mediated SMAD signaling in human ESCs. Cell Stem Cell. 2008;3:196–206. doi: 10.1016/j.stem.2008.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamakuchi M, Ferlito M, Lowenstein CJ. miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci. 2008;105:13421–13426. doi: 10.1073/pnas.0801613105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan N, Lu Y, Sun H, Tao D, Zhang S, Liu W, Ma Y. A microarray for microRNA profiling in mouse testis tissues. Reproduction. 2007;134:73–79. doi: 10.1530/REP-07-0056. [DOI] [PubMed] [Google Scholar]
- Yan N, Lu Y, Sun H, Qiu W, Tao D, Liu Y, Chen H, Yang Y, Zhang S, Li X, et al. Microarray profiling of microRNAs expressed in testis tissues of developing primates. J Assist Reprod Genet. 2009;26:179–186. doi: 10.1007/s10815-009-9305-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Younger ST, Pertsemlidis A, Corey DR. Predicting potential miRNA target sites within gene promoters. Bioorg Med Chem Lett. 2009;19:3791–3794. doi: 10.1016/j.bmcl.2009.04.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu Z, Raabe T, Hecht NB. MicroRNA Mirn122a reduces expression of the posttranscriptionally regulated germ cell transition protein 2 (Tnp2) messenger RNA (mRNA) by mRNA cleavage. Biol Reprod. 2005;73:427–433. doi: 10.1095/biolreprod.105.040998. [DOI] [PubMed] [Google Scholar]