Abstract
ATP-dependent proteases are currently emerging as key regulators of mitochondrial functions. Among these proteolytic systems, Pim1, a Lon-like serine protease in Saccharomyces cerevisiae, is involved in the control of selective protein turnover in the mitochondrial matrix. In the absence of Pim1, yeast cells have been shown to accumulate electron-dense inclusion bodies in the matrix space, to lose integrity of mitochondrial genome, and to be respiration-deficient. Because of the severity of phenotypes associated with the depletion of Pim1, this protease appears to be an essential component of the protein quality control machinery in mitochondria and to exert crucial functions during the biogenesis of this organelle. Nevertheless, its physiological substrates and partners are not fully characterized. Therefore, we used the combination of different proteomic techniques to assess the nature of oxidized protein substrates and physiological partners of Pim1 protease under non-repressing growth conditions. The results presented here supply evidence that Pim1-mediated proteolysis is required for elimination of oxidatively damaged proteins in mitochondria.
Keywords: Oxygen/Reactive, Protease/ATP-dependent, Protein/Protein-protein Interactions, Protein/Degradation, Proteomics, Subcellular Organelles/Mitochondria
Introduction
Proteolysis by AAA+ proteases (ATPases associated with various cellular activities) (1) plays a fundamental role in the maintenance of mitochondrial functions. These energy-dependent proteases are present in various subcompartments of mitochondria. Two major proteases, Pim1/Lon and ClpP, are present in the mitochondrial matrix, and two membrane-embedded proteases, Yme1 (i-AAA) and Yta10/Yta12 (m-AAA), with their proteolytic domains facing the matrix and the intermembrane space, respectively, are known to degrade membrane proteins (2–4). These ATP-dependent proteases are believed to fulfill a dual function in the organelle. First, they carry out the elimination of non-assembled, damaged, or misfolded proteins, thus ensuring the proper stoichiometry of multienzyme complexes and preventing the accumulation of harmful aggregated polypeptides in mitochondria. Additionally, these proteases display regulatory functions by selectively degrading some mitochondrial protein targets, which appears to be crucial for mitochondrial biogenesis.
Among these proteolytic systems, Pim1 (proteolysis into mitochondria) is a highly conserved ATP-dependent Lon-like protease in S. cerevisiae (5–7). No homologue of ClpP was found in yeast and Pim1 is therefore the only ATP-dependent protease in the mitochondrial matrix of this organism. Unlike many AAA+ proteases, Pim1 contains three domains (N domain, AAA+ module, and proteolytic domain) that are encoded in the same polypeptide chain and forms homoheptameric complexes (8). Under cellular stress conditions, induction of the Pim1/Lon transcript has been observed, suggesting a role for the protease in the elimination of misfolded or damaged proteins (6, 9, 10). Pim1 cooperates with chaperone proteins from the Hsp70 and the Hsp100 family, which allow stabilization of substrate polypeptides in a soluble conformation susceptible to degradation (11–14). There is no evidence for the existence of a proteolytic targeting signal in mitochondria, such as polyubiquitination addressing degradation substrates to the 26 S proteasome in the cytosol or SsrA that targets proteins for proteolysis in bacteria (15, 16). Hence, the non-native conformation of substrate polypeptides appears to be a signal for proteolysis by Pim1/Lon (11, 13, 17–19).
Disruption of PIM1 causes severe phenotypes. Cells lacking PIM1 accumulate electron-dense inclusion bodies in the matrix space, presumably corresponding to aggregated proteins, and are respiration-deficient. This has been attributed to the loss of mtDNA5 integrity in Pim1-deficient cells, resulting in a ρ− phenotype (6, 7). The molecular mechanisms responsible for the loss of mtDNA integrity in Pim1 mutants are still elusive, but the proteolytic function of Pim1 protease has been implicated (20). Moreover, impaired respiratory competence in Pim1 mutants can also be assigned to the role of the protease in the splicing of introns encoding maturases in COB and COX1 transcripts (21).
Pim1 appears to be an essential component of the protein quality control system in mitochondria, also harboring crucial functions in the biogenesis of the organelle. Nevertheless, its physiological substrates and partners are not fully characterized. Major et al. (10) were able to identify a novel subset of Pim1 substrates by a proteomic approach using glucose as a carbon source. In the present work, we investigated the substrate specificity of Pim1 protease by comparing mitochondrial oxidized proteomes between wild-type (WT) and Pim1-depleted cells (Δpim1) grown under non-repressing conditions, using two-dimensional gel electrophoresis (GE) analyses. Because mitochondrial protein composition depends on protein degradation mediated by proteases present in the organelle, we hypothesized that the absence of Pim1 would result in the accumulation of its specific degradation substrates. Using the combination of different proteomic approaches, 19 proteins were found to be more abundant in Δpim1 mitochondria, accumulating or not as preferentially oxidized products. We also examined the influence of repressing/non-repressing growth conditions on the steady state level of Pim1 substrates, and we assessed the range of Pim1 partners using tandem affinity purification (TAP). According to their nature and function, proteins of interest were classified into five categories: 1) mitochondrial stress proteins, 2) mitochondrial metabolic enzymes, 3) respiratory chain subunits, 4) mitochondrial ribosomal proteins, and 5) mtDNA nucleoid proteins. Our results provide new insights into regulatory functions of Pim1 in mitochondria and illustrate the role performed by mitochondrial ATP-dependent proteases in the protein quality control and the biogenesis of this organelle.
EXPERIMENTAL PROCEDURES
Yeast Strains and Growth Conditions
The strains used in this study were SC0437 (MATa; ade2; arg4; leu2-3, 112; trp1-289; ura3-52; lon::TAP-K.I.URA3) and SC0000 (MATa; ade2; arg4; leu2-3, 112; trp1-289; ura3-52), ordered from Euroscarf (Frankfurt, Germany). The Δlon (Δpim1) and isogenic wild-type WT strains referred to throughout this study had been described previously (22): JK93dα wild-type WT ρ0 (EtBr-treated, leu2-3, 112, ura3-52, trp1, his4, rme1, HMLa); JK93dα Δlon (Δpim1) ρ0 (EtBr treated, leu2-3,112, ura3-52, trp1, his4, rme1, lon::KAN, HMLa). Non-respiring mutant strains were tested for the presence of wild-type mtDNA (ρ+) by scoring growth on a non-fermentable carbon source. SC0000 and SC0437 strains were grown at 30 °C in YPGR (1% yeast extract, 2% peptone, 0.1% glucose, 2% raffinose). Δlon (Δpim1) and isogenic WT strains were grown in yeast nitrogen base lacking uracil containing 0.1% dextrose and 2% raffinose (raffinose medium) or 2% dextrose (dextrose medium).
Isolation of Mitochondria and Preparation of Protein Extracts
Cells were harvested by centrifugation at 3000 × g for 20 min at 4 °C, washed with 50 ml of ice-cold water, and resuspended in MST buffer (210 mm mannitol, 70 mm sucrose, 5 mm Tris-HCl, pH 7.5, with 1 tablet/50 ml of complete protease inhibitor mixture (Roche Applied Science)), at a protein concentration of 25 mg/ml. About 4 ml of 425–600-μm acid-washed glass beads (Sigma) were added to each tube, and the cells were lysed by vortexing three times for 2 min with 2-min intervals on ice. All of the subsequent steps were carried out at 4 °C. Glass beads and unbroken cells were removed by centrifugation at 1,000 × g for 5 min, and supernatants were used to prepare total cell protein extracts. Alternatively, supernatants were centrifuged (14,000 × g, 10 min, 4 °C); pellets were resuspended with MST buffer to prepare mitochondrial protein extracts, whereas supernatants were used to prepare cytosolic extracts. The purity of mitochondrial extracts was evaluated using specific antibodies (supplemental Fig. S1). Mitochondrial membrane fractions were prepared from isolated mitochondria swollen for 10 min at 4 °C in 50 mm sucrose, 1 mm EDTA, 10 mm Mops/KOH, pH 7.2, and subjected to an ultracentrifugation step (100,000 × g, 1 h at 4 °C). Protein content was determined using the Bradford method (Bio-Rad). Samples were then prepared in Laemmli sample buffer and incubated for 5 min at 100 °C.
Immunochemical Reagents
Rabbit polyclonal antibodies against Om45, Phb1, and Phb2 were a gift from Dr. M. Yaffe (University of California, San Diego). Rabbit polyclonal antibodies specific to the yeast Cox4 and Rip1 proteins were a gift from Dr. G. Isaya (Mayo Clinic and Foundation, Rochester, MI). Rabbit polyclonal antibodies raised against Atp2 and Atp7 were a gift from Dr. J. Velours (IBGC, Bordeaux, France). Polyclonal antibody against lipoic acid was purchased from Calbiochem. Anti-porin antibody was a mouse monoclonal antibody obtained from Molecular Probes/Invitrogen. Anti-TAP polyclonal antibody was obtained from Biovalley (Marne la Vallée, France). Anti-Hsp60 was a mouse monoclonal antibody obtained from StressMarq (Victoria, Canada), and anti-Hsp104 was a rabbit polyclonal antibody from MBL (Woburn, MA). Monoclonal antibody against Mrp20 was a gift from Dr. C. Suzuki (University of Medicine and Dentistry of New Jersey, Newark, NJ). Rabbit polyclonal antibodies against Ilv5 and Hsp78 were provided by Dr. J. Marszalek (University of Gdansk). A polyclonal antibody against yeast aconitase targeted to residues 495–511 of S. cerevisiae mitochondrial aconitase and a polyclonal antibody against Pim1 targeted to residues 337–353 were described previously (23).
Western Blot Analysis
Samples were separated by SDS-PAGE in a 12% (w/v) polyacrylamide gel, and proteins were electrotransferred onto a Hybond nitrocellulose membrane (GE Healthcare). Primary antibody binding was detected by incubation with a peroxidase-conjugated secondary antibody and chemiluminescent substrate ECL Plus (GE Healthcare).
Detection of Carbonylated Proteins
Carbonylated proteins were detected and analyzed following derivatization of protein carbonyl groups with 2,4-dinitrophenylhydrazine, using the OxyBlot kit (Millipore, Billerica, MA). Immunodetection was performed utilizing 10 μg of mitochondrial protein per lane with a primary antibody directed against dinitrophenylhydrazone.
Measurement of Mitochondrial ATP-stimulated Proteolytic Activity
Mitochondrial proteolytic activity was assayed by determining the rate of fluorescein isothiocyanate-casein (Sigma) degradation, as described previously (23).
Purification of Tagged Complexes
The tagged complexes were purified from 8 mg of mitochondria isolated from two different yeast strains (from Euroscarf), SC0000 (control strain expressing the Pim1 protein) and SC0437 (strain expressing the Pim1-TAP fusion protein). Mitochondria were isolated as described under “Experimental Procedures.” The TAP purification was performed by sequential IgG affinity and calmodulin affinity columns. Briefly, mitochondria were lysed with 0.25% Nonidet P-40 and cleared by low speed centrifugation. The supernatant was further lysed with 1.25% Nonidet P-40, cleared by high speed centrifugation, and loaded on IgG affinity chromatography by binding to Protein A tag. The bound complexes were eluted by tobacco etch virus protease cleavage (N terminus to Protein A tag). Fractions were pooled and purified by calmodulin affinity chromatography by binding to calmodulin-binding peptide tag and eluted with EGTA.
RNA Isolation and Real-time Quantitative PCR Analysis
Total RNA was extracted from WT and Δpim1 yeasts cultured in YNB medium lacking uracil containing 0.1% dextrose and 2% raffinose to an A600 nm of ∼0.7. We followed the Gary Kobs protocol (75), using the SV Total RNA Isolation System (Promega Corp., Madison, WI). cDNA was synthesized from 3 μg of RNA, using the SuperScript III first strand synthesis system for reverse transcription-PCR (Invitrogen). PCRs were performed on a LightCycler®480 system (Roche Applied Science) at the technical platform of the IFR 83 at Université Pierre et Marie Curie-Paris 6. Each reaction was carried out in 10 μl with 2 μl of cDNA (equivalent to 10 ng of reverse transcribed RNA), 500 nm primer concentration, and 1× LightCycler® 480 SYBR Green I master mix (Roche Applied Science). Serial dilutions of cDNA (37–3.7 ng) were used to generate a quantitative PCR standard curve. The LightCycler protocol was as follows: 5 min of 95 °C hot start enzyme activation; 40 cycles of 95 °C denaturation for 10 s, 60 °C annealing for 15 s, and 72 °C elongation for 15 s; and melting at 95 °C for 5 s, 65 °C for 60 s, and then heating to 97 °C. Water was used as the template for negative control amplifications included with each PCR run. All reactions were performed in triplicate. Data were analyzed using Roche LightCycler® 480 Software, and CP was calculated by the second derivative maximum method. The amount of the target mRNA was examined and normalized to the RPO21 gene mRNA. The relative expression ratio of a target gene was calculated as described by Pfaffl (24), based on real-time PCR efficiencies. Primer pairs are listed in Table 2 and were designed using Probe Finder software (available on-line from Roche Applied Science). The results presented are from three independent experiments (n = 3).
TABLE 2.
Analysis of the expression level of genes of interest
The expression level of genes of interest was determined in WT and Δpim1 cells by quantitative PCR experiments, as described under “Experimental Procedures.” For each gene, relative expression (Δpim1/WT) was determined by calculating the ratio between the expression level from Δpim1 and WT. Sequences of primers used are depicted: the first is the forward primer, and the second is the reverse primer.
| Genes | Primers (5′–3′) | Expression level in WT | Expression level in Δpim1 | Relative expression Δpim1/WT |
|---|---|---|---|---|
| HSP78 | CGACGGTAAATTAGACCCTGTC | 0.84 ± 0.07 | 1.54 ± 0.22 | 1.83 |
| ATTTGAATAGCTCTTGCGATTTCT | ||||
| HSP60 | GAAGTTGCCTCCAAAACCAA | 0.89 ± 0.05 | 2.15 ± 0.18 | 2.41 |
| TGGCTCTACCTAAGACAGTAGCAG | ||||
| SOD2 | AAACCACTGTCTATTCTGGGAAA | 0.90 ± 0.19 | 1.34 ± 0.06 | 1.49 |
| CTCGTCGATTGCCTTTGC | ||||
| PDB1 | ATACGGGTTCACAGGTTTGG | 0.88 ± 0.05 | 0.74 ± 0.04 | 0.84 |
| GCTTGCATAGAGAAATTGAACG | ||||
| LAT1 | AAAGACCAGTACGAAGGCTCAG | 0.88 ± 0.05 | 0.76 ± 0.08 | 0.86 |
| GCTGGCGCTTCTTGTTTCT | ||||
| LPD1 | TTCGTTAAAGGAAATTCCCAAA | 0.86 ± 0.53 | 0.68 ± 0.05 | 0.79 |
| ACCCATTTCCAATCCGATG | ||||
| COQ5 | TTACGCAAGCTCACAGAGCA | 1.39 ± 0.244 | 1.17 ± 0.17 | 0.84 |
| GCTGCGGATGATAAAGGACT | ||||
| DLD1 | TATTTGAGCGACCACGGTTT | 0.73 ± 0.04 | 0.48 ± 0.05 | 0.66 |
| AACCACCAATCTGTGCACCT | ||||
| ALD4 | CGGGTACGGTCTGGATAAAC | 1.36 ± 0.29 | 1.74 ± 0.22 | 1.28 |
| AACCCACCGAAAGGAACTG | ||||
| ILV5 | GCCATTGTTGACCAAGGGTA | 0.73 ± 0.16 | 0.64 ± 0.10 | 0.88 |
| TCCTTGAAGACTGGGGAGAA | ||||
| QCR2 | GCGGCCTATTTACTCTGTTTGT | 0.97 ± 0.17 | 1.17 ± 0.13 | 1.21 |
| CCGCAACAATTTTCTTGATGT | ||||
| RIP1 | TGTGTTCCAATTGGTGAAGC | 0.85 ± 0.31 | 1.35 ± 0.30 | 1.59 |
| TGAACCATGGCAAGGACAG | ||||
| ATP1 | CTCCTGGTCGTGAAGCCTAC | 1.17 ± 0.25 | 1.17 ± 0.04 | 1.00 |
| GCGGCTCTTTCTAGCAATCTT | ||||
| ATP2 | TCCGTGGTGAAAAGGTTCTT | 0.93 ± 0.05 | 0.90 ± 0.02 | 0.97 |
| TCCCTAAAGTTTCTCTCCCAACT | ||||
| MRP20 | GCCAGAGCAGGTTCCAATAA | 1.25 ± 0.27 | 0.56 ± 0.43 | 0.45 |
| GCCGCTTTTGTCGAGTATTT |
Two-dimensional Electrophoresis
Mitochondrial fractions were isolated from wild-type and Δpim1 yeast cells cultured in YNB medium lacking uracil containing 0.1% dextrose and 2% raffinose. For isoelectric focusing, mitochondrial protein extracts (300 μg) were resuspended in sample buffer (7 m urea, 2 m thiourea, 4% (w/v) CHAPS, 1% (w/v) dithiothreitol, 2% (v/v) ampholites (pH 3–10, NL), and 0.001% (w/v) bromphenol blue). Aliquots (250 μl) were applied on Immobiline Drystrips (pH 3–10 NL, 13 cm; GE Healthcare), which were then incubated overnight at 20 °C in a reswelling tray (GE Healthcare). Reswelling was followed by protein electrofocusing for 4 h at 300 V, 30 min at 750 V, 30 min at 1500 V, 16 h at 2500 V, and then 2 h at 3500 V, using the Multiphor II system (GE Healthcare). The strips were then equilibrated by incubation for 10 min in equilibration buffer (50 mm Tris-HCl, pH 8.8, 6 m urea, 30% (v/v) glycerol, 1% (w/v) SDS) supplemented with 1% (w/v) dithiothreitol and for 10 min in equilibration buffer containing 3% (w/v) iodoacetamide and 0.001% (w/v) bromphenol blue. The second dimension consisted of SDS-PAGE in a 12% (w/v) polyacrylamide gel, using the Protean II system (Bio-Rad). Gels were running for 16 h at 96 V. Proteins were detected by colloidal blue staining according to the manufacturer (Sigma). Gels were scanned using ImageScanner (GE Healthcare), and protein spots were quantified using ImageMaster 2D Platinum version 7.0 software (GE Healthcare). To account for experimental variations, four independent experiments were performed with independent samples.
Two-dimensional OxyBlots
Isoelectric focusing was performed using 250 μg of mitochondrial protein extracts from wild-type and Δpim1 cells as described above. After the first dimension, strips were frozen at −80 °C for 2 h, and protein carbonyl groups were derivatized; strips were incubated in a solution containing 2 m HCl and 10 mm 2,4-dinitrophenylhydrazine (Sigma) for 20 min at 20 °C. The strips were then incubated twice in a solution containing 2 m Tris base and 30% glycerol (v/v) for 20 min each, at 20 °C, and were equilibrated as described above. After the second dimension (as described above), proteins were electrotransferred onto a Hybond nitrocellulose membrane (GE Healthcare). The primary antibody (OxyBlot kit, Millipore) used for immunodetection was raised against dinitrophenylhydrazone. Films were scanned using ImageScanner (GE Healthcare), and protein spots were quantified using ImageMaster 2D Platinum version 7.0 software (GE Healthcare). To account for experimental variations, four independent experiments were performed with independent biological samples.
Quantification of Two-dimensional Gels and Two-dimensional OxyBlots
Protein spots were matched on two-dimensional gels and two-dimensional OxyBlots, and quantification was performed using ImageMaster 2D Platinum version 7.0 software (GE Healthcare). For the different experimental conditions (WT or Δpim1), the volume of each spot was quantified and normalized to the total spot volume per gel or per OxyBlot. Thus, we obtained the percentage of volume for each spot. Then we determined protein folds or oxidation folds by calculating the ratio (change) between percentage of volume of spots in Δpim1 compared with WT (Δpim1/WT), respectively, for two-dimensional gels or two-dimensional OxyBlots. The S.D. value was calculated, and statistical significance was assessed using Student's t test among four independent experiments (n = 4). Oxidation indexes were calculated as the ratio between oxidation folds and protein folds, for each spot of interest.
Mass Spectrometry
Bands or spots of interest were manually excised from one-dimensional gels or two-dimensional gels, washed with deionized water and then twice with 100% acetonitrile for 15 min and 2 min. The gel plugs were dried in a vacuum centrifuge and rehydrated by incubation in 2% trypsin (Promega Corp.) in 50 mm NH4HCO3 at 4 °C. After 20 min of incubation, the excess trypsin solution was removed, 30 μl of 50 mm NH4HCO3 was added, and digestion was allowed to proceed at 37 °C overnight. Samples were stored at −20 °C until use. Peptides were desalted on custom-made reverse-phase microcolumns prepared with R2 resin (Perseptive Biosystems Inc., Framingham, MA), as described previously (25). A peptide mixture, obtained by trypsin digestions of each spot, was loaded onto the microcolumn, which was then washed with 10 μl of 1% trifluoroacetic acid. Bound peptides were eluted with 0.8 μl of matrix solution (5 μg/μl α-cyano-4-hydrocynnamic acid in 70% acetonitrile and 0.1% trifluoroacetic acid) directly onto the matrix-assisted laser desorption ionization (MALDI) target plate. Peptide mass spectra were acquired in positive reflector mode on a 4700 Proteomics Analyzer MALDI time-of-flight (TOF)/TOF apparatus (Applied Biosystems, Foster City, CA), using an acceleration voltage of 20 kV. Each spectrum was obtained with a total of 1000–1200 laser shots and was externally calibrated using peptides derived by digesting β-lactoglobulin with trypsin. Tandem mass spectra were acquired using the same instrument in MS/MS-positive mode. Peak lists were generated from the raw data output by Data Explorer version 4.6 software (Applied Biosystems). MS and MS/MS peak lists were combined into a single search file and used to search the Swiss-Prot data base (release of May 18, 2009) with the Mascot search engine, version 2.2 (Matrix Science, Ltd., London, UK; release of May 18, 2009). The search parameters were as follows: data base, Swiss-Prot; taxonomy, S. cerevisiae; enzyme, trypsin; number of missed cleavages tolerated, 1; fixed modifications, 0; variable modifications, methionine oxidation; peptide mass tolerance, 70 ppm; fragment mass tolerance, 500 ppm. Protein hits with a significant Mascot score (>65) were considered as positively identified.
RESULTS
Accumulation of Oxidatively Damaged Proteins in Δpim1 Cells
WT and Δpim1 cells were converted to ρ0 mutants by ethidium bromide treatment (26) in order to reduce genotypic variability so that WT and Δpim1 cells absolutely display the same defect in mtDNA and that only the presence or absence of Pim1 protease differs between these cells. As a result, WT and Δpim1 cells used in this study are unable to grow on a non-fermentable carbon source, such as glycerol/ethanol (data not shown). Moreover, because one of our goals was to characterize Pim1 functions in mitochondrial biogenesis, cells were grown in raffinose medium.
Pim1 processing requires two independent cleavages after mitochondrial import of the precursor to generate a pro-form and a mature form of Pim1, (20). To assess that Pim1 protein is correctly imported and processed into a mature form in ρ0 cell mitochondria, we performed immunoblots of WT and Δpim1 mitochondrial fractions (Fig. 1A). Two distinct bands around 150 kDa were detected from the wild type. Both the pro-form and the mature form are shown to co-purify with mitochondrial membranes (Fig. 1A), suggesting that Pim1 could be recruited to mitochondrial inner membrane. Furthermore, we did not observe any change in the mitochondrial content between WT and Δpim1 cells, as shown by immunoblots of total cell extracts using antibodies against Por1 (mitochondrial porin) or Om45 (a protein of the outer membrane of mitochondria).
FIGURE 1.
Processing of Pim1 preprotein, ATP-stimulated proteolytic activity, and oxidized protein content in WT and Δpim1 mitochondria. WT and Δpim1 cells were grown at 30 °C in Raffinose medium. A, immunoblot analysis (50 μg of protein/lane) of mitochondrial proteins (M), mitochondrial membrane fractions (Mb), and total fractions (T) isolated from WT and Δpim1 cells, using antibodies directed against Pim1 (p, pro-form; m, mature form), Por1, or Om45. B, mitochondrial ATP-stimulated proteolytic activity was assessed in WT and Δpim1 mitochondria by analysis of the proteolytic breakdown of casein-fluorescein isothiocyanate (for details, see “Experimental Procedures”). Fluorescence released into supernatant fractions was quantified, and Vi values, corresponding to casein degradation-hydrolyzing activity, were calculated for WT (6 ± 0.24 pmol/min/mg) and Δpim1 (3.9 ± 0.17 pmol/min/mg). Vi was taken as 100% for the wild type (n = 3). The data in the bar graph represent means ± S.E. The asterisk indicates the level of statistical significance: *, p < 0.03 according to Student's t test. C, representative OxyBlot showing oxidized protein content in WT and Δpim1 mitochondria. 10 μg of mitochondrial proteins were loaded/lane (see details under “Experimental Procedures”).
In order to evaluate the implications of Pim1 in mitochondrial ATP-dependant proteolysis, we analyzed the proteolytic breakdown of casein-fluorescein isothiocyanate by WT or Δpim1 isolated mitochondrial extracts (Fig. 1B). Mitochondrial ATP-dependent proteolytic activity was reduced by 35% in Δpim1 cells, thus representing the contribution of Pim1 protease in energy-dependent protein degradation in this organelle. Because the human Lon protease has been shown to degrade oxidized proteins (27, 28), we investigated the possibility of a functional conservation in yeast. To determine the level of oxidative damage to proteins, the amount of carbonyl groups was monitored in isolated mitochondria from WT and Δpim1 cells by OxyBlot analysis. Oxidized proteins were found to accumulate at elevated levels in Δpim1 mitochondria (Fig. 1C).
Identification of Oxidized Protein Substrates of Pim1 Protease
Proteomic approaches are well adapted to monitor remodeling of the organellar protein content in response to changes in metabolism or genetic environment. Comparative analyses of mitochondrial oxidized proteomes were performed between WT and Δpim1 cells grown under non-repressing conditions using raffinose as a carbon source to obtain a comprehensive overview of Pim1-mediated protein turnover reactions in mitochondria.
Two-dimensional-GE analyses are presented in Fig. 2. Two-dimensional gels were either stained using colloidal Coomassie Brilliant Blue G to detect proteins (Fig. 2, A and B) or were electrotransferred onto nitrocellulose membranes to subsequently detect carbonylated proteins using the OxyBlot technique (Fig. 2, C and D). 14 protein spots are more abundant in Δpim1 mitochondria (Fig. 2, A versus B), all of which accumulate as carbonylated proteins (Fig. 2, C versus D).
FIGURE 2.
Comparative analysis of mitochondrial oxidized proteomes between WT and Δpim1 cells. Mitochondrial fractions were isolated from WT and Δpim1 cells grown at 30 °C in raffinose medium and were subjected to two-dimensional GE analysis. Following two-dimensional GE, gels were either stained with colloidal Coomassie Brilliant Blue G to detect proteins (A and B) or were electrotransferred onto nitrocellulose membranes to subsequently detect carbonylated proteins using the OxyBlot technique (C and D), as described under “Experimental Procedures.” A and C, WT mitochondrial extracts; B and D, Δpim1 mitochondrial extracts. The arrows and numbers refer to spots accumulating in Δpim1 mitochondria, which are identified in Fig. 3 and listed in supplemental Table S1 and Table 1. Presented results are from one representative experiment of four independent experiments.
A wide range of proteins were identified by mass spectrometry (Fig. 3 and supplemental Table S1). Hsp60 (heat shock protein 60) and Sod2 (mitochondrial superoxide dismutase) are stress proteins known for their refolding or antioxidant properties, respectively. Pdb1 (pyruvate dehydrogenase E1 component subunit β), Lat1 (dihydrolipoamide acetyltransferase component (E2) of pyruvate dehydrogenase complex), Lpd1 (dihydrolipoamide dehydrogenase (E3) component of pyruvate dehydrogenase complex), CoQ5 (ubiquinone biosynthesis methyltransferase COQ5), Dld1 (d-lactate dehydrogenase), Ald4 (potassium-activated aldehyde dehydrogenase), Aim17 (uncharacterized oxidoreductase YHL021C), and Ilv5 (ketol acid reductoisomerase) are all metabolic enzymes, most of which are involved in mitochondrial energetic pathways and metabolic adaptation. Notably, Pdb1, Lat1, and Lpd1 correspond to three subunits of the pyruvate dehydrogenase complex. Atp1 (F1F0-ATP synthase subunit α), Atp2 (F1F0-ATP synthase subunit β), Qcr2 (subunit 2 of the ubiquinol cytochrome c reductase complex), and Rip1 (Rieske iron-sulfur protein of the ubiquinol-cytochrome c reductase complex) are subunits of respiratory chain complexes. Interestingly, some of the proteins of interest (Ilv5, Pdb1, Lpd1, Ald4, Atp1, and Hsp60) correspond to bifunctional proteins that, in addition to their known primary function in mitochondria, have been identified as components of mtDNA nucleoids in S. cerevisiae (29–31). To differentiate proteins that are preferentially oxidized in Δpim1 mitochondria, we performed quantification of protein spots. Results are presented in Table 1. After normalization of oxidation folds by the corresponding protein folds, we obtained oxidation indexes. Seven proteins (Hsp60, Atp2, Pdb1, Ald4, Lpd1, Ilv5, and Sod2) display an oxidation index greater than 1 and most likely correspond to Pim1-specific oxidized protein substrates.
FIGURE 3.
Identification of spots of interest. Spots of interest were identified using mass spectrometry as described under “Experimental Procedures” (see supplemental Table S1 and “Supplemental Proteomic Data” for details). Enlarged areas of two-dimensional gels (colloidal Coomassie Brilliant Blue G) and two-dimensional OxyBlot regions containing spots of interest are presented for WT and Δpim1 mitochondria. Spots of interest are indicated with an arrow. The corresponding identified proteins are shown on two-dimensional gels, whereas spots are numbered on two-dimensional OxyBlots (as in Fig. 2). Presented results are from one representative experiment of four independent experiments.
TABLE 1.
Quantification data and associated statistics from two-dimensional analyses
Spot quantification (percentage volume of each spot) was performed on two-dimensional gels and two-dimensional OxyBlots from WT and Δpim1 mitochondria using ImageMaster 2D Platinum version 7.0 software (GE Healthcare). Protein fold (Δpim1/WT) and oxidation fold (Δpim1/WT) were calculated (n = 4). For each protein of interest, the oxidation fold was normalized by the corresponding protein fold to assess the oxidation index. Proteins with an oxidation index greater than 1 (indicated in boldface type) correspond to preferentially accumulating oxidized proteins in Δpim1 mitochondria (only proteins with an oxidation index of ≥1,15 were considered). Statistical significance for differential expression between WT and Δpim1 (p values from two-dimensional gels and two-dimensional OxyBlots) are indicated. According to Student's t test, p values were calculated from quantification data obtained from four independent experiments.
| Genesa | Spotsb | Protein foldsc (Δpim1/WT) | Oxidation foldsd (Δpim1/WT) | Oxidation indexese (Δpim1/WT) |
p valuesf |
|
|---|---|---|---|---|---|---|
| Two-dimensional gels | Two-dimensional OxyBlots | |||||
| HSP60 | 1 | 3.26 ± 0.71 | 6.82 ± 0,74 | 2.09 | <0.01 | <0.01 |
| ATP2 | 2 | 2.85 ± 0.67 | 3.69 ± 1,16 | 1.29 | <0.05 | <0.05 |
| PDB1 | 3 | 3.18 ± 0.95 | 6.33 ± 2,93 | 1.99 | <0.05 | <0.01 |
| COQ5 | 4 | 3.73 ± 0.80 | 3.64 ± 1,08 | 0.98 | <0.05 | <0.05 |
| LAT1 | 5 | 2.33 ± 0.42 | 1.79 ± 0,87 | 0.77 | <0.05 | <0.05 |
| DLD1 | 6 | 4.16 ± 1.27 | 3.08 ± 0,38 | 0.74 | <0.01 | <0.005 |
| ALD4 | 7 | 3.85 ± 0.60 | 4.44 ± 1,63 | 1.15 | <0.005 | <0.05 |
| QCR2 | 8 | 2.28 ± 0.53 | 2.19 ± 1,09 | 0.96 | <0.05 | <0.05 |
| AIM17 | 9 | 3.07 ± 0.92 | 3.17 ± 1,21 | 1.03 | <0.05 | <0.05 |
| LPD1 | 10 | 4.98 ± 1.73 | 7.68 ± 3,69 | 1.54 | <0.05 | <0.01 |
| ATP1 | 11 | 3.56 ± 0.68 | 2.93 ± 1,23 | 0.82 | <0.01 | <0.05 |
| ILV5 | 12 | 2.37 ± 0.27 | 12.36 ± 7,51 | 5.22 | <0.005 | <0.01 |
| RIP1 | 13 | 3.21 ± 0.93 | 3.37 ± 1,31 | 1.05 | <0.05 | <0.05 |
| SOD2 | 14 | 2.51 ± 0.29 | 2.98 ± 0,83 | 1.19 | <0.05 | <0.05 |
a Genes correspond to gene names found in the Saccharomyces Genome Database.
b Spots refer to the numbered spots on Figs. 2 and 3 (as in supplemental Table S1).
c Protein folds are calculated from quantification of two-dimensional gels.
d Oxidation folds are calculated from quantification of two-dimensional OxyBlots.
e Oxidation indexes correspond to oxidation folds normalized by the corresponding protein folds.
f p values obtained using Student's t test (n = 4).
To further characterize the involvement of Pim1 in the degradation of the identified protein substrates, quantitative PCR analysis was performed from WT and Δpim1 cells grown under non-repressing conditions using raffinose as a carbon source. 15 genes encoding the identified proteins were analyzed, and the results are described in Table 2. These results show that HSP60, SOD2, and HSP78 genes are increased transcriptionally in Δpim1 cells compared with WT cells, which is consistent with an overexpression of these genes in response to mitochondrial stress due to the unfolded and oxidized proteins that accumulate in Δpim1 cells. In contrast, PDB1, LAT1, LPD1, COQ5, DLD1, ALD4, QCR2, RIP1, ATP1, and ATP2 transcription levels did not show any increase in mRNA expression in Δpim1 cells (Table 2). Thus, these results do not correlate with the elevated protein levels observed by two-dimensional GE (Table 1), indicating that the corresponding proteins accumulate in the mitochondria from Δpim1 cells because they are not degraded. In addition, Ilv5 and Mrp20 proteins, which are known to be specific Pim1 substrates (14, 22), also show no change in mRNA expression but a significant increase in protein amount in Δpim1 mitochondria (Table 1 and Fig. 4).
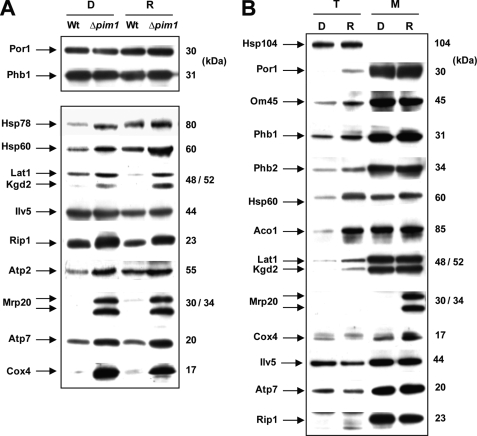
FIGURE 4.
Influence of dextrose versus raffinose medium on the steady state level of Pim1 degradation substrates. A, mitochondria were isolated from WT and Δpim1 cells grown either in dextrose (D) or in raffinose (R) medium at 30 °C. Immunodetection of Pim1 substrates was performed using specific antibodies (25 μg of mitochondrial protein/lane). Immunodetection of Por1 (mitochondrial porin) and Phb1 (prohibitin 1) was used as loading control. B, immunoblot analysis of total cell extracts (T; 50 μg of protein/lane) or mitochondrial extracts (M; 20 μg of protein/lane) from WT cells grown either in dextrose or in raffinose medium was performed using specific antibodies directed against various mitochondrial proteins, especially Pim1 substrates (Hsp60, Lat1, Kgd2, Mrp20, Cox4, Ilv5, Atp7, and Rip1). Immunoblot of Hsp104 (cytosolic heat shock protein 104) was used as a control.
Influence of Dextrose Versus Raffinose Growth on the Steady State Level of Pim1 Substrates in WT and Δpim1 Mitochondria
Many studies have examined transcriptional and metabolic patterns and controls operating during growth of S. cerevisiae in various carbon sources (reviewed in Ref. 32). In glucose-rich medium, this organism represses the synthesis of functional components of several pathways that are not required for glycolysis (reviewed in Refs. 33 and 34). Numerous mitochondrial functions and most of the nuclear genes encoding certain mitochondrial components are repressed, including components of the tricarboxylic acid cycle, respiratory chain subunits, and mitochondrial ribosomes (35).
To obtain more indications concerning Pim1 substrate specificity, the influence of dextrose versus raffinose cell growth on the steady state level of Pim1 substrates was compared in isolated mitochondria. Five proteins already identified using two-dimensional GE analyses (Hsp60, Lat1, Ilv5, Rip1, and Atp2) (Fig. 3, Table 1, and supplemental Table S1) were confirmed to accumulate in Δpim1 mitochondria (Fig. 4A and Table 2). Mrp20, a mitochondrial subunit of the large ribosomal particle that was previously shown to be degraded by Pim1 (22), was also found. Furthermore, to obtain additional substrates of Pim1, we also performed immunoblots using specific antibodies directed against Hsp78 (heat shock protein 78), Cox4 (subunit 4 of the cytochrome oxidase complex), Atp7 (subunit d of ATP-synthase), and lipoic acid (a prosthetic group present in E2 components of pyruvate dehydrogenase complex and keto acid dehydrogenase complex, referring to Lat1 and Kgd2, respectively (36)). As shown in Fig. 4A, most Pim1 substrates, such as Lat1, Kgd2 (dihydrolipoyl transsuccinylase, E2 component of the mitochondrial α-ketoglutarate dehydrogenase complex), Rip1, Mrp20, Atp7, and Cox4, accumulate at nearly identical levels in Δpim1 mitochondria from cells grown either in dextrose or in raffinose. In contrast, Ilv5 accumulates only in Δpim1 derepressed mitochondria (Figs. 3 and 4A). This result is supported by a previous study showing that a mutant form of Ilv5 with a tendency to aggregate is degraded by Pim1 when raffinose but not glucose is used as a carbon source (14). Moreover, the overexpression level of this protein was considerably decreased when analysis was performed by Western blot (Fig. 4A) compared with when using two-dimensional GE analysis (Fig. 3). This could be attributed to a better solubilization of protein aggregates in urea/thiourea when using two-dimensional GE analysis than when using one-dimensional GE analysis.
To further characterize catabolite repression/derepression effects on the steady state level of mitochondrial proteins, especially Pim1 substrates, we performed immunoblots of total cell extracts and mitochondrial extracts of WT cells grown either in dextrose or in raffinose using specific antibodies (Fig. 4B). Most of the mitochondrial proteins analyzed (see Por1, Om45, Phb1, Phb2, Hsp60, Aco1, Lat1, and Kgd2) are present at increased amounts in total extracts of cells grown in raffinose as a carbon source compared with dextrose-grown cells, without changes in the expression of the same proteins in isolated mitochondria (Fig. 4B) (also confirmed by comparing mitochondrial contents of Ilv5, Atp7, and Rip1 in dextrose versus raffinose growing conditions). This demonstrates that dextrose/raffinose growth conditions have an influence on nucleus-encoded mitochondrial protein expression and may modulate mitochondrial mass according to catabolite repression/derepression. Hsp60 and Hsp78 were found to be overexpressed in WT mitochondria from cells grown in raffinose medium compared with dextrose medium, which suggests the need for refolding machineries due to massive protein import during biogenesis of mitochondria (Fig. 4, A and B; see also Refs. 37 and 38). Accordingly, a general stress response in terms of heat shock and oxidative stress responses has been shown to occur transcriptionally in S. cerevisiae in response to catabolite derepression (35, 39). In contrast, Mrp20 or Cox4 are barely detectable in total cell extracts and are present at considerably decreased amounts in WT mitochondria upon dextrose repression (Fig. 4B). In the absence of Pim1, these two proteins accumulate at nearly similar levels either under repressing or non-repressing conditions (Fig. 4A), which exclude transcriptional, posttranscriptional, or translational regulations. These results reflect specific post-translational regulation functions of Pim1 protease that could selectively modulate the degradation of key proteins, such as Mrp20 and Cox4, involved in the biogenesis process of the organelle. Notably, Atp2 is also affected by this post-translational regulation (Fig. 4A). Furthermore, Ilv5 and Atp7 are not overexpressed in cells upon raffinose derepression, and these two proteins are even present at decreased amounts in total extract from cells grown under raffinose conditions compared with dextrose-grown cells (Fig. 4B). Accordingly, the ILV5 gene is known to specifically respond to amino acid starvation because the encoded product is involved in branched-chain amino acid synthesis (40), and our results show that Ilv5 protein is specifically degraded by Pim1 in response to catabolite derepression (Figs. 3 and 4A; see also Ref. 14).
Identification of Pim1 Partners Using TAP
To further investigate mitochondrial proteins that stably interact with Pim1, we used a strain of S. cerevisiae expressing a fusion of Pim1 protein to TAP tag (SC0437), which allows purifying Pim1 and its interacting partners using TAP. As expected, this fusion protein has a slower mobility than Pim1 protein from the control SC0000 strain (Fig. 5A). After TAP purification, no protein was observed in the elution fraction using mitochondrial extracts from the control SC0000 strain (Fig. 5B). This demonstrates the specificity of the purification process. In contrast, several proteins interacting with Pim1-TAP were recovered after TAP purification using the SC0437 strain (Fig. 5B). Protein identification by mass spectrometry revealed that Phb1, Phb2, and Atp7 interact with Pim1-TAP (supplemental Table S2). The identity of these interacting partners was confirmed by Western blot using specific antibodies (Fig. 5C).
FIGURE 5.
Identification of Pim1 interacting partners using TAP. Cells from SC0000 and SC0437 strains were grown in YPGR at 30 °C. A, mitochondrial protein extracts (50 μg of protein/lane) from SC0000 and SC0437 strains were electrotransferred onto nitrocellulose membrane and incubated with anti-Pim1 or anti-TAP tag antibodies. Immunodetection of Por1 (porin) was used as loading control. B, TAP purification was performed using SC0000 and SC0437 mitochondrial fractions as described under “Experimental Procedures.” Elution fractions were analyzed on one-dimensional gel using silver staining. Protein bands, obtained from the SC0437 strain, were identified by mass spectrometry (for more details, see “Experimental Procedures” and see supplemental Table S2 and “Supplemental Proteomic Data”). Immunoblot analysis of mitochondrial fractions from the SC0437 strain (M; 25 μg of protein) and elution fractions from SC0000 or SC0437 mitochondria (same volume of elution fraction for each) was performed using an antibody directed against the TAP tag. The presented results are from one representative experiment of three independent experiments. C, Pim1 and its interacting partners (Phb1, Phb2, and Atp7) were confirmed by immunoblotting of the elution fraction from the TAP purification performed with SC0437 mitochondria using specific antibodies. WB, Western blot.
In mitochondria, Phb1 and Phb2 assemble into a high molecular weight complex in the inner membrane (41, 42). This large complex interacts with the m-AAA protease and has been proposed as a membrane-bound chaperone for the stabilization of newly imported respiratory chain subunits (43–45). In addition, Atp7 is found to interact with Pim1 (Fig. 5, B and C). This predominantly hydrophilic membrane protein exposes a large domain in the matrix space. Atp7 was also found to be a substrate of Pim1 (Fig. 4A), thus suggesting specific tasks of Pim1 regarding this protein, because ATP-dependent proteases are usually interacting weakly and transiently with their degradation substrates.
DISCUSSION
We have examined the role of Pim1 protease in mitochondrial proteolysis using WT(ρ0) and Δpim1(ρ0) cells in order to focus on the proteolytic function of Pim1 independently of its impact on mtDNA integrity. In ρ0 cells, many nucleus-encoded subunits of respiratory chain complexes as well as nucleus-encoded mitochondrial ribosomal proteins are degraded rapidly after import due to defective assembly in mitochondria (reviewed in Ref. 46). This model is therefore particularly adapted to study mitochondrial proteolytic processes. By analyzing purified mitochondria using two-dimensional GE, OxyBlots, and immunoblots, 19 protein substrates of Pim1 were identified, and three interacting partners of this protease were evidenced by TAP purification. According to their nature and function, proteins of interest found using the different approaches were classified into five functional categories: 1) mitochondrial stress proteins, 2) mitochondrial metabolic enzymes, 3) respiratory chain subunits, 4) mitochondrial ribosomal proteins, and 5) mtDNA nucleoid proteins (Table 3). Among the substrates that were found, Atp1, Atp2, and a mutant form of Ilv5 have been previously shown to be degraded by Pim1 protease (7, 14, 47) as well as Mrp20 (22). Furthermore, in a recent proteomics-based study, Major et al. (10) have identified a subset of mitochondrial proteins all composed of metabolic enzymes (Ilv1, Ilv2, Lsc1, Lys4, and Yjl200c) as substrates of Pim1. However, none of these identified proteins matches with metabolic enzymes that we found. A probable reason is that the authors performed analysis under glucose-repressing growth conditions, which could imply remodeling in metabolism-related mitochondrial protein expression and modulation of Pim1 substrate specificity compared with raffinose-non-repressing growth conditions. In addition, they used WT(ρ−) and Δpim1(ρ−) cells. This could explain the general difference between the cluster of mitochondrial metabolic enzymes identified by Major et al. (10) and the different protein groups identified in our study (Table 3 and Fig. 6). Taken together, our findings highlight the dual function of Pim1 in mitochondrial protein quality control and biogenesis.
TABLE 3.
Classification of proteins of interest into functional categories
Proteins of interest were classified into five distinct functional categories: mitochondrial stress proteins, mitochondrial metabolic enzymes, respiratory chain subunits, mitochondrial ribosomal proteins, and mtDNA nucleoid proteins (marked with asterisks). Submitochondrial localization of proteins is indicated (M, matrix; IM, inner mitochondrial membrane) as well as their significance in the present study. Indicated protein names are from the Saccharomyces Genome Database.
| Functional categories and proteins of interest | Mitochondrial localization | mtDNA nucleoid proteins | Significance in this study |
|---|---|---|---|
| Mitochondrial stress proteins | |||
| Hsp78 | M | Mitochondrial stress response in Δpim1 mitochondria | |
| Hsp60 | M | * (29) | Mitochondrial stress response in Δpim1 mitochondria |
| Sod2 | M | Mitochondrial stress response in Δpim1 mitochondria | |
| Phb1 | IM | Interacting partner of Pim1 (also interacts with the m-AAA protease (43)) | |
| Phb2 | IM | Interacting partner of Pim1 (also interacts with the m-AAA protease (43)) | |
| Mitochondrial metabolic enzymes | |||
| Pdb1 (PDH complex) | M | * (30) | Degradation substrate of Pim1 protease |
| Lat1 (PDH complex) | M | Degradation substrate of Pim1 protease | |
| Lpd1 (PDH complex) | M | * (29) | Degradation substrate of Pim1 protease |
| Kgd2 (KGDH complex) | M | * (29) | Degradation substrate of Pim1 protease |
| CoQ5 | M | Degradation substrate of Pim1 protease | |
| Dld1 | IM | Degradation substrate of Pim1 protease | |
| Ald4 | M | * (29) | Degradation substrate of Pim1 protease |
| Aim17 | M | Degradation substrate of Pim1 protease | |
| Ilv5 | M | * (29) | Degradation substrate of Pim1 protease (already identified (14)) |
| Respiratory chain subunits | |||
| Qcr2 (complex III) | IM | Degradation substrate of Pim1 protease | |
| Rip1 (complex III) | IM | Degradation substrate of Pim1 protease | |
| Cox4 (complex IV) | IM | Degradation substrate of Pim1 protease | |
| Atp1 (complex V) | IM | * (29) | Degradation substrate of Pim1 protease (already identified (47)) |
| Atp2 (complex V) | IM | Degradation substrate of Pim1 protease (already identified (7, 47)) | |
| Atp7 (complex V) | IM | Degradation substrate of Pim1 protease/Interacting partner of Pim1 (also partially degraded by the m-AAA protease (74)) | |
| Mitochondrial ribosomal protein | |||
| Mrp20 | M | Degradation substrate of Pim1 protease (already identified (22)) | |
FIGURE 6.
Pim1 protease is involved in mitochondrial biogenesis and integrity. Pim1 degrades various mitochondrial protein substrates. The turnover of unassembled respiratory chain subunits by Pim1 protease could be mediated by the prohibitin complex (Phb1/2) in the inner membrane. Many substrates of Pim1 protease are part of mtDNA nucleoid proteins, underscoring the role of this protease in mtDNA metabolism. In addition, Pim1-mediated proteolysis is responsible for elimination of oxidatively damaged proteins. Accordingly, depletion of Pim1 protease leads to a mitochondrial stress response (overexpression of Hsp78, Hsp60, and Sod2). PDH, pyruvate dehydrogenase complex (E1, E2, and E3 subunits); M, matrix; IM, inner mitochondrial membrane; IMS, intermembrane space; OM, outer mitochondrial membrane. Proteins of interest identified in this study are shown in red.
Pim1 Plays a Prevalent Role in Mitochondrial Protein Quality Control
Our results show that Pim1 protease is a key component of the proteolytic defense system against oxidative damages in mitochondria by degrading preferentially several carbonylated proteins (Table 1 and Fig. 6). Thus, carbonylation may act as an unfolding signal, ensuring that mitochondrial damaged proteins enter the degradation pathway by being targeted to Pim1, rather than the chaperone/refolding pathway because carbonylation is an irreversible modification (48). In the absence of Pim1 protease, oxidized Atp1 and Atp2 (α and β subunits of ATP synthase) accumulate at elevated levels (Table 1). By degrading these two proteins, Pim1 would ensure their stoichiometric concentration in mitochondria and prevent their aggregation when not incorporated into functional F1 particles (49, 50). We also identified Ilv5 as a Pim1 substrate, with an oxidation index of 5.22 (Table 1). This protein was reported to be highly susceptible to aggregation following a single point mutation (14). Thus, these specific substrates of Pim1 are good candidates to be responsible for initial events promoting formation of aggregate in Δpim1 mitochondria (7). As a result, a mitochondrion-specific stress response was observed in PIM1-depleted cells because three stress proteins, Hsp78, Hsp60, and Sod2, were overexpressed in Δpim1 mitochondria (Table 2 and Fig. 6). Accordingly, the absence of the mammalian Lon protease leads to the accumulation of aggregated, oxidatively damaged proteins in mitochondria (28, 51), and Pim1/Lon-deficient cells have been shown to accumulate mitochondrial aggregated proteins from bacteria to humans (7, 51, 52). All together, these findings underscore the role of Pim/Lon protease in the maintenance of mitochondrial integrity during the aging process (Table 3) (27, 48, 53, 54).
The turnover of mitochondrial energy-transducing complex subunits is considered necessary to ensure stoichiometric amounts of different subunits essential for the proper assembly of the respiratory chain and to prevent the possibly deleterious accumulation of nonnative polypeptides in the inner membrane. Mitochondrial membrane-integrated proteases, m-AAA and i-AAA proteases, are known to carry out this membrane protein degradation process (55, 56). The Phb1-Phb2 complex is linked to proteolysis in the inner membrane and was functionally associated with the F1F0-ATP synthase (43–45, 57). Our results show that Pim1 could be recruited to the inner membrane of mitochondria (Fig. 1A) and is involved in the turnover of unassembled matrix-exposed subunits of the respiratory chain and the inner membrane protein Dld1 (Table 3). These findings indicate that prohibitins, that were found to interact with Pim1, could serve as a recruiter complex in the inner membrane to assist the quality control of these membrane proteins by Pim1 protease. Interestingly, Atp7 was found to be in the same time substrate and partner of Pim1. This subunit corresponds to the matrix-exposed subunit d of ATP synthase that is required for the correct assembly of the catalytic particle F1 with the F0 membranous particle (58). This could suggest that Pim1 is able to interact and stabilize the unassembled Atp7 subunit during biogenesis of the ATP synthase complex (47, 59). Alternatively, the interaction could be mediated by prohibitins and Pim1 may play a role similar to Atp23, which mediates maturation of Atp6 and chaperones its assembly into F1F0-ATP synthase complexes (57).
Pim1 Protease Is Involved in the Biogenesis of Mitochondria
In addition to its role in posttranscriptional control of mitochondrial genes COB and COX1 (21), the Pim1 protease has been shown to degrade a broad range of protein substrates involved in various mitochondrial functions (Table 3 and Fig. 6). Our findings add new dimensions to the role of Pim1 in mitochondrial biogenesis that include a significant degree of control over the substrate specificity and activity of Pim1 protease according to metabolic growth conditions. Our results tend to show that Pim1-mediated proteolysis of metabolic enzymes indirectly controls mitochondrial concentrations of specific proteins (quinone pool) and metabolites (NADH and FADH2) that are basic elements for energy production in the organelle (see Fig. 6). Notably, conversion of lactate to acetyl-CoA by the sequential action of Dld1 and pyruvate dehydrogenase complex and subsequent activation of the citric acid cycle are necessary for cell adaptation accompanying the diauxic shift. In ρ0 cells, the biogenesis of respiratory chain complexes is obviously defective, but the increase of mitochondrial mass occurs all the same in response to catabolite derepression (Fig. 4B). Here, we show that Pim1 is not required for the increase of mitochondrial mass (Fig. 1A) but exerts some posttranslational regulations by selectively modulating the degradation of specific proteins, such as Cox4, Mrp20, and Atp2, according to metabolic growth conditions (Fig. 4, A and B).
Importantly, our results also assign novel functions of Pim1 in mtDNA metabolism (Fig. 6). Indeed, a large part of Pim1 substrates (Ilv5, Pdb1, Lpd1, Kgd2, Ald4, Atp1, and Hsp60) correspond to bifunctional proteins that associate with mtDNA to form nucleoids (Table 3 and Fig. 6) (29–31). These mtDNA nucleoid proteins are involved in mtDNA maintenance, and mutations in either of the corresponding genes generate different levels of mitochondrial genome instability (60, 61). Recently, mtDNA nucleoids in yeast were identified as dynamic structures that are remodeled in response to metabolic cues (62). Hsp60 and Ilv5 are recruited to mtDNA nucleoids during glucose repression and amino acid starvation, respectively (62, 63). By selectively degrading some mitochondrial nucleoid proteins, such as Ilv5, under non-repressing growth conditions, Pim1 could participate to this remodeling process and therefore influence the modulation of mtDNA dynamic changes with respect to mitochondrial gene expression. Moreover, DNA binding by the ATP-dependent Lon protease was demonstrated in bacteria, mouse, and human but not for the yeast Pim1 ortholog (64–67). Interactions of mtDNA with mitochondrial enzymes seemingly unrelated to mtDNA transactions as well as with protein folding machinery are conserved in higher eukaryotes (68–72). Interestingly, most of the mitochondrial nucleoid proteins found in our study correspond to Pim1 substrates that are preferentially degraded when oxidized (see Pdb1, Ald4, Lpd1, Ilv5, and Hsp60 in Table 1, Table 2, and Fig. 6). This raises the possibility that these mtDNA nucleoid proteins exert protection of mtDNA against oxidative damages by trapping reactive oxygen species. Thereby, in the absence of Pim1/Lon protease, it is conceivable that mitochondrial nucleoid proteins would be stabilized on mtDNA, which could possibly increase the protection of mtDNA against oxidative damage (73). Although further experiments are required to clearly demonstrate the role of Pim1-mediated proteolysis in remodeling mtDNA nucleoids, our results shed new light on the role of Pim1 in mitochondrial biogenesis and provide new insights into the level of regulation exerted by Pim1 protease during this process (Fig. 6 and Table 3).
Supplementary Material
Acknowledgments
We thank Dr. M. Yaffe for providing antibodies directed against Phb1, Phb2, and Om45. We thank Dr. G. Isaya for anti-Cox 4 and anti-Rip1 antibodies. We thank Dr. J. Velours for anti-Atp2 and anti-Atp7 antibodies. We thank Dr. C. Suzuki for anti-Mrp20 antibody and for providing the JK93dα strains. We thank Dr. J. Marszalek for antibodies directed against Ilv5 and Hsp78. We thank S. Bornens for help and advice concerning quantitative PCR experiments.
This work was supported by funds from the Ministere de l'Education Nationale, de la Recherche, et de la Technologie (MENRT), and Proteomage Grant LSHM-CT-518230.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Fig. S1 and Tables S1 and S2.
- mtDNA
- mitochondrial DNA
- GE
- gel electrophoresis
- TAP
- tandem affinity purification
- CHAPS
- 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonic acid
- Mops
- 4-morpholinepropanesulfonic acid
- MALDI
- matrix-assisted laser desorption ionization
- TOF
- time-of-flight
- MS
- mass spectrometry
- MS/MS
- tandem MS.
REFERENCES
- 1.Mogk A., Haslberger T., Tessarz P., Bukau B. (2008) Biochem. Soc. Trans. 36, 120–125 [DOI] [PubMed] [Google Scholar]
- 2.Leidhold C., Voos W. (2007) Ann. N.Y. Acad. Sci. 1113, 72–86 [DOI] [PubMed] [Google Scholar]
- 3.Koppen M., Langer T. (2007) Crit. Rev. Biochem. Mol. Biol. 42, 221–242 [DOI] [PubMed] [Google Scholar]
- 4.Tatsuta T., Langer T. (2008) EMBO J. 27, 306–314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Wang N., Gottesman S., Willingham M. C., Gottesman M. M., Maurizi M. R. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 11247–11251 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Van Dyck L., Pearce D. A., Sherman F. (1994) J. Biol. Chem. 269, 238–242 [PubMed] [Google Scholar]
- 7.Suzuki C. K., Suda K., Wang N., Schatz G. (1994) Science 264, 273–276 [DOI] [PubMed] [Google Scholar]
- 8.Stahlberg H., Kutejová E., Suda K., Wolpensinger B., Lustig A., Schatz G., Engel A., Suzuki C. K. (1999) Proc. Natl. Acad. Sci. U.S.A. 96, 6787–6790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hori O., Ichinoda F., Tamatani T., Yamaguchi A., Sato N., Ozawa K., Kitao Y., Miyazaki M., Harding H. P., Ron D., Tohyama M., Stern D. M., Ogawa S. (2002) J. Cell Biol. 157, 1151–1160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Major T., von Janowsky B., Ruppert T., Mogk A., Voos W. (2006) Mol. Cell. Biol. 26, 762–776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wagner I., Arlt H., van Dyck L., Langer T., Neupert W. (1994) EMBO J. 13, 5135–5145 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Savel'ev A. S., Novikova L. A., Kovaleva I. E., Luzikov V. N., Neupert W., Langer T. (1998) J. Biol. Chem. 273, 20596–20602 [DOI] [PubMed] [Google Scholar]
- 13.Rottgers K., Zufall N., Guiard B., Voos W. (2002) J. Biol. Chem. 277, 45829–45837 [DOI] [PubMed] [Google Scholar]
- 14.Bateman J. M., Iacovino M., Perlman P. S., Butow R. A. (2002) J. Biol. Chem. 277, 47946–47953 [DOI] [PubMed] [Google Scholar]
- 15.Schwartz A. L., Ciechanover A. (2009) Annu. Rev. Pharmacol. Toxicol. 49, 73–96 [DOI] [PubMed] [Google Scholar]
- 16.Karzai A. W., Roche E. D., Sauer R. T. (2000) Nat. Struct. Biol. 7, 449–455 [DOI] [PubMed] [Google Scholar]
- 17.von Janowsky B., Knapp K., Major T., Krayl M., Guiard B., Voos W. (2005) Biol. Chem. 386, 1307–1317 [DOI] [PubMed] [Google Scholar]
- 18.Ondrovicová G., Liu T., Singh K., Tian B., Li H., Gakh O., Perecko D., Janata J., Granot Z., Orly J., Kutejová E., Suzuki C. K. (2005) J. Biol. Chem. 280, 25103–25110 [DOI] [PubMed] [Google Scholar]
- 19.Gur E., Sauer R. T. (2008) Genes Dev. 22, 2267–2277 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wagner I., van Dyck L., Savel'ev A. S., Neupert W., Langer T. (1997) EMBO J. 16, 7317–7325 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.van Dyck L., Neupert W., Langer T. (1998) Genes Dev. 12, 1515–1524 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.van Dijl J. M., Kutejová E., Suda K., Perecko D., Schatz G., Suzuki C. K. (1998) Proc. Natl. Acad. Sci. U.S.A. 95, 10584–10589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bulteau A. L., Dancis A., Gareil M., Montagne J. J., Camadro J. M., Lesuisse E. (2007) Free Radic. Biol. Med. 42, 1561–1570 [DOI] [PubMed] [Google Scholar]
- 24.Pfaffl M. W. (2001) Nucleic Acids Res. 29, e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gobom J., Nordhoff E., Mirgorodskaya E., Ekman R., Roepstorff P. (1999) J. Mass Spectrom. 34, 105–116 [DOI] [PubMed] [Google Scholar]
- 26.Fox T. D., Folley L. S., Mulero J. J., McMullin T. W., Thorsness P. E., Hedin L. O., Costanzo M. C. (1991) Methods Enzymol. 194, 149–165 [DOI] [PubMed] [Google Scholar]
- 27.Bota D. A., Van Remmen H., Davies K. J. (2002) FEBS Lett. 532, 103–106 [DOI] [PubMed] [Google Scholar]
- 28.Bota D. A., Davies K. J. (2002) Nat. Cell Biol. 4, 674–680 [DOI] [PubMed] [Google Scholar]
- 29.Kaufman B. A., Newman S. M., Hallberg R. L., Slaughter C. A., Perlman P. S., Butow R. A. (2000) Proc. Natl. Acad. Sci. U.S.A. 97, 7772–7777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chen X. J., Wang X., Kaufman B. A., Butow R. A. (2005) Science 307, 714–717 [DOI] [PubMed] [Google Scholar]
- 31.Kucej M., Butow R. A. (2007) Trends Cell Biol. 17, 586–592 [DOI] [PubMed] [Google Scholar]
- 32.Schüller H. J. (2003) Curr. Genet. 43, 139–160 [DOI] [PubMed] [Google Scholar]
- 33.Gancedo J. M. (1998) Microbiol. Mol. Biol. Rev. 62, 334–361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Carlson M. (1999) Curr. Opin. Microbiol. 2, 202–207 [DOI] [PubMed] [Google Scholar]
- 35.DeRisi J. L., Iyer V. R., Brown P. O. (1997) Science 278, 680–686 [DOI] [PubMed] [Google Scholar]
- 36.Onder O., Yoon H., Naumann B., Hippler M., Dancis A., Daldal F. (2006) Mol. Cell Proteomics 5, 1426–1436 [DOI] [PubMed] [Google Scholar]
- 37.Craig E. A., Gambill B. D., Nelson R. J. (1993) Microbiol. Rev. 57, 402–414 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Voos W., Röttgers K. (2002) Biochim. Biophys. Acta 1592, 51–62 [DOI] [PubMed] [Google Scholar]
- 39.Roberts G. G., Hudson A. P. (2006) Mol. Genet. Genomics 276, 170–186 [DOI] [PubMed] [Google Scholar]
- 40.Petersen J. G., Holmberg S. (1986) Nucleic Acids Res. 14, 9631–9651 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Back J. W., Sanz M. A., De Jong L., De Koning L. J., Nijtmans L. G., De Koster C. G., Grivell L. A., Van Der Spek H., Muijsers A. O. (2002) Protein Sci. 11, 2471–2478 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tatsuta T., Model K., Langer T. (2005) Mol. Biol. Cell 16, 248–259 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Steglich G., Neupert W., Langer T. (1999) Mol. Cell. Biol. 19, 3435–3442 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Nijtmans L. G., de Jong L., Artal Sanz M., Coates P. J., Berden J. A., Back J. W., Muijsers A. O., van der Spek H., Grivell L. A. (2000) EMBO J. 19, 2444–2451 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Nijtmans L. G., Artal S. M., Grivell L. A., Coates P. J. (2002) Cell Mol. Life Sci. 59, 143–155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Rep M., Grivell L. A. (1996) Curr. Genet. 30, 367–380 [DOI] [PubMed] [Google Scholar]
- 47.Rep M., van Dijl J. M., Suda K., Schatz G., Grivell L. A., Suzuki C. K. (1996) Science 274, 103–106 [DOI] [PubMed] [Google Scholar]
- 48.Nyström T. (2005) EMBO J. 24, 1311–1317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lefebvre-Legendre L., Vaillier J., Benabdelhak H., Velours J., Slonimski P. P., di Rago J. P. (2001) J. Biol. Chem. 276, 6789–6796 [DOI] [PubMed] [Google Scholar]
- 50.Lefebvre-Legendre L., Salin B., Schaëffer J., Brèthes D., Dautant A., Ackerman S. H., di Rago J. P. (2005) J. Biol. Chem. 280, 18386–18392 [DOI] [PubMed] [Google Scholar]
- 51.Bota D. A., Ngo J. K., Davies K. J. (2005) Free Radic. Biol. Med. 38, 665–677 [DOI] [PubMed] [Google Scholar]
- 52.Rosen R., Biran D., Gur E., Becher D., Hecker M., Ron E. Z. (2002) FEMS Microbiol. Lett. 207, 9–12 [DOI] [PubMed] [Google Scholar]
- 53.Ngo J. K., Davies K. J. (2007) Ann. N.Y. Acad. Sci. 1119, 78–87 [DOI] [PubMed] [Google Scholar]
- 54.Squier T. C. (2001) Exp. Gerontol. 36, 1539–1550 [DOI] [PubMed] [Google Scholar]
- 55.Leonhard K., Herrmann J. M., Stuart R. A., Mannhaupt G., Neupert W., Langer T. (1996) EMBO J. 15, 4218–4229 [PMC free article] [PubMed] [Google Scholar]
- 56.Arnold I., Langer T. (2002) Biochim. Biophys. Acta 1592, 89–96 [DOI] [PubMed] [Google Scholar]
- 57.Osman C., Wilmes C., Tatsuta T., Langer T. (2007) Mol. Biol. Cell 18, 627–635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Norais N., Promé D., Velours J. (1991) J. Biol. Chem. 266, 16541–16549 [PubMed] [Google Scholar]
- 59.Suzuki C. K., Rep M., van Dijl J. M., Suda K., Grivell L. A., Schatz G. (1997) Trends Biochem. Sci. 22, 118–123 [DOI] [PubMed] [Google Scholar]
- 60.Chen X. J., Butow R. A. (2005) Nat. Rev. Genet. 6, 815–825 [DOI] [PubMed] [Google Scholar]
- 61.Contamine V., Picard M. (2000) Microbiol. Mol. Biol. Rev. 64, 281–315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Kucej M., Kucejova B., Subramanian R., Chen X. J., Butow R. A. (2008) J. Cell Sci. 121, 1861–1868 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.MacAlpine D. M., Perlman P. S., Butow R. A. (2000) EMBO J. 19, 767–775 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Nomura K., Kato J., Takiguchi N., Ohtake H., Kuroda A. (2004) J. Biol. Chem. 279, 34406–34410 [DOI] [PubMed] [Google Scholar]
- 65.Lu B., Liu T., Crosby J. A., Thomas-Wohlever J., Lee I., Suzuki C. K. (2003) Gene 306, 45–55 [DOI] [PubMed] [Google Scholar]
- 66.Liu T., Lu B., Lee I., Ondrovicová G., Kutejová E., Suzuki C. K. (2004) J. Biol. Chem. 279, 13902–13910 [DOI] [PubMed] [Google Scholar]
- 67.Fu G. K., Markovitz D. M. (1998) Biochemistry 37, 1905–1909 [DOI] [PubMed] [Google Scholar]
- 68.Bogenhagen D. F., Wang Y., Shen E. L., Kobayashi R. (2003) Mol. Cell Proteomics 2, 1205–1216 [DOI] [PubMed] [Google Scholar]
- 69.Wang Y., Bogenhagen D. F. (2006) J. Biol. Chem. 281, 25791–25802 [DOI] [PubMed] [Google Scholar]
- 70.Malka F., Lombès A., Rojo M. (2006) Biochim. Biophys. Acta 1763, 463–472 [DOI] [PubMed] [Google Scholar]
- 71.Bogenhagen D. F., Rousseau D., Burke S. (2008) J. Biol. Chem. 283, 3665–3675 [DOI] [PubMed] [Google Scholar]
- 72.Chen S. H., Suzuki C. K., Wu S. H. (2008) Nucleic Acids Res. 36, 1273–1287 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Lu B., Yadav S., Shah P. G., Liu T., Tian B., Pukszta S., Villaluna N., Kutejová E., Newlon C. S., Santos J. H., Suzuki C. K. (2007) J. Biol. Chem. 282, 17363–17374 [DOI] [PubMed] [Google Scholar]
- 74.Korbel D., Wurth S., Käser M., Langer T. (2004) EMBO Rep. 5, 698–703 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Kobs G. (2001) Promega Notes, Vol. 68, pp. 28–29, Promega Corp., Madison, WI [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.