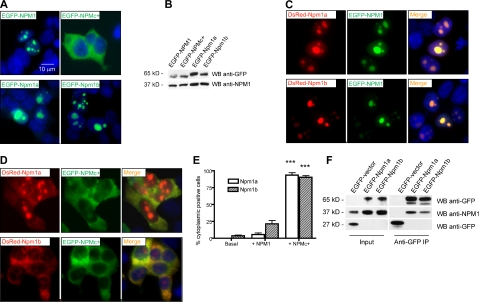
Figure 4.
Zebrafish npm1 orthologs interact with human NPM1 protein. (A) Epifluorescent images of 293T cells transfected with pEGFP-C1 expression vector encoding NPM1 (top left), NPMc+ (top right), Npm1a (bottom left), or Npm1b (bottom right). Expressed proteins are shown in green; nuclei are counterstained in blue with DAPI. (B) Anti-GFP WB analysis of protein lysates from transfected cells shown in Figure 2A. Levels of transfected proteins (top row) are shown along with levels of endogenous NPM1 (bottom row). (C) Epifluorescent images of 293T cells cotransfected with pEGFP-C1-NPM1 and pDsRed-monomer-C1 encoding zebrafish Npm1a (top row) or Npm1b (bottom row). Colocalization areas between Npm1a or Npm1b (in red, left column) and NPM1 (in green, middle column) are shown in yellow in the merged images (right column). Nuclei are counterstained in blue with DAPI. (D) Epifluorescent images of 293T cells cotransfected with pEGFP-C1-NPM1c+ and pDsRed-monomer-C1 encoding Npm1a (top row) or Npm1b (bottom row). Colocalization areas between Npm1a or Npm1b (in red, left column) and NPMc+ (in green, middle column) are shown in yellow in the merged images (right column). Nuclei are counterstained in blue with DAPI. All fluorescent images were acquired with a Zeiss Axio imager Z1 microscope using a Zeiss 63×/1.4 NA Apochromat oil lens (Carl Zeiss) and Openlab software (Perkin Elmer). (E) Quantification of the Npm1a and Npm1b delocalization by NPMc+. Bars indicate the percentage of cells with aberrant Npm1a (□) or Npm1b (▧) cytoplasmic expression (with or without residual nucleolar positivity). Error bars indicate SD. *Statistically significant differences between the NPMc+-injected and both NPM1-injected and uninjected embryos (***P < .001; Student t test). More than 200 cells were counted in 3 different microscopic fields for this analysis. (F) Anti-GFP coimmunoprecipitation assays of 293T cells transfected with the empty pEGFP-C1 vector, pEGFP-C1-Npm1a, or pEGFP-C1-Npm1b. Left panels, input; right panels, anti-GFP coimmunoprecipitation lanes. Top row: anti-GFP WB, detecting EGFP-Npm1a and EGFP-Npm1b fusion proteins. Middle row: anti-NPM1 WB, detecting endogenous NPM1. Bottom row: anti-GFP WB, detecting EGFP expressed by the empty vector. In the anti-GFP immunoprecipitation lanes, a dimmer band under the EGFP-Npm1a and EGFP-Npm1b bands probably represents a degradation product of the fusion protein, as it is also visible in the input lanes at longer exposures.