Abstract
Pandemic 2009 influenza A virus (A/H1N1/2009) has emerged globally. In this study, we performed a comprehensive detection of potential pathogens by de novo sequencing using a next-generation DNA sequencer on total RNAs extracted from an autopsy lung of a patient who died of viral pneumonia with A/H1N1/2009. Among a total of 9.4×106 40-mer short reads, more than 98% appeared to be human, while 0.85% were identified as A/H1N1/2009 (A/Nagano/RC1-L/2009(H1N1)). Suspected bacterial reads such as Streptococcus pneumoniae and other oral bacteria flora were very low at 0.005%, and a significant bacterial infection was not histologically observed. De novo assembly and read mapping analysis of A/Nagano/RC1-L/2009(H1N1) showed more than ×200 coverage on average, and revealed nucleotide heterogeneity on hemagglutinin as quasispecies, specifically at two amino acids (Gly172Glu and Gly239Asn of HA) located on the Sa and Ca2 antigenic sites, respectively. Gly239 and Asn239 on antigenic site Ca2 appeared to be minor amino acids compared with the highly distributed Asp239 in H1N1 HAs. This study demonstrated that de novo sequencing can comprehensively detect pathogens, and such in-depth investigation facilitates the identification of influenza A viral heterogeneity. To better characterize the A/H1N1/2009 virus, unbiased comprehensive techniques will be indispensable for the primary investigations of emerging infectious diseases.
Introduction
In April 2009, an H1N1 triple-reassortant swine influenza virus (A/H1N1/2009) was detected in humans with febrile respiratory illness in North America [1], and the virus has rapidly spread worldwide by human-to-human transmission. According to the disease outbreak news from the World Health Organization, at least 14,711 people died from A/H1N1/2009 between April 2009 and January 2010 (http://www.who.int/csr/don/en/). Fatal cases from A/H1N1/2009 viral infection were summarized in a report by Gill et al. [2].
The genome of influenza A virus (family Orthomyxoviridae) consists of 8 single-stranded negative sense RNA molecules spanning approximately 13.5 kb. The segments range in length from 890 to 2341 nucleotides (nt) and encode a total of 11 proteins [3]. Genetic diversity in influenza virus results from a high mutation rate associated with replication using a low-fidelity RNA polymerase and the reshuffling (reassortment) of segments among coinfecting strains. Multiple-reassortant influenza viruses from avian, human, and swine origins emerged as major pandemic influenza viruses (i.e., 1918 H1N1, 1957 H2N2, and 1968 H3N2) causing significant mortality in humans in the 20th century [4]. Such an “antigenic shift” by multiple reassortant drives the emergence of pandemic influenza viruses, with their severity and clinical outcome always unpredictable [5].
Influenza A virus can evade antibodies specific to its attachment protein, hemagglutinin (HA), by the accumulation of amino acid substitutions in HA epitopes [6]. This “antigenic drift” in HA epitopes [7] affects recognition by antibodies that neutralize viral infectivity by blocking the interaction of HA with sialic acid residues on host-cell membranes. The H1 subtype HA has four antigenic sites recognized by monoclonal antibodies with high neutralizing activity, designated Sa, Sb, Ca, and Cb [8]. In addition, 8 continuous B cell/antibody epitopes for human H1N1 HA proteins have been experimentally defined by the Immune Epitope Database and Analysis Resource (IEDB: http://www.immuneepitope.org/) [9]. Immune epitope analysis of HA epitopes in A/H1N1/2009 is also summarized in the Influenza Research Database (http://www.fludb.org/brc/homeExtraPage.do?decorator=influenza&extraPage=separate) [10].
To better predict a future pandemic of influenza A virus, the characterization of possible antigenic drift will be indispensable. Igarashi et al. and Shen et al. reported that a structural comparison of HAs could predict probable future antigenic changes during the evolution of A/H1N1/2009 in the human population [11], [12].
In addition to this prediction, extensive investigations on viral quasispecies will be required to reveal the actual appearance of those antigenic changes. Nakamura et al. demonstrated the direct detection of potential pathogens, including influenza virus, using de novo pyrosequencing [13], but the detection appeared to have insufficient redundant sequencing reads to reveal the genetic variation of the viruses. Ramakrishnan et al. demonstrated the discrimination of quasispecies in mixed HA subtype infections of influenza A virus using the same pyrosequencing approach [14]. However, it was shown that the influenza viral RNA sample should be enriched through sequence-specific oligonucleotide capturing prior to pyrosequencing, indicating that such enrichment might represent a biased result.
Here, we performed de novo sequencing using total RNAs extracted from an autopsy lung of a patient infected with A/H1N1/2009, and detected potential pathogens such as bacteria in addition to A/H1N1/2009. Extensive DNA sequencing using the Illumina Genome Analyzer II (GA II) revealed that quasispecies for the HA sequence were generated in single patient. Such heterogeneity demonstrated the antigenic drift of HA, implying the existence of a mechanism to escape pre-existing human immunity to the virus.
Results
Summary of sequencing reads and detection of potential pathogens
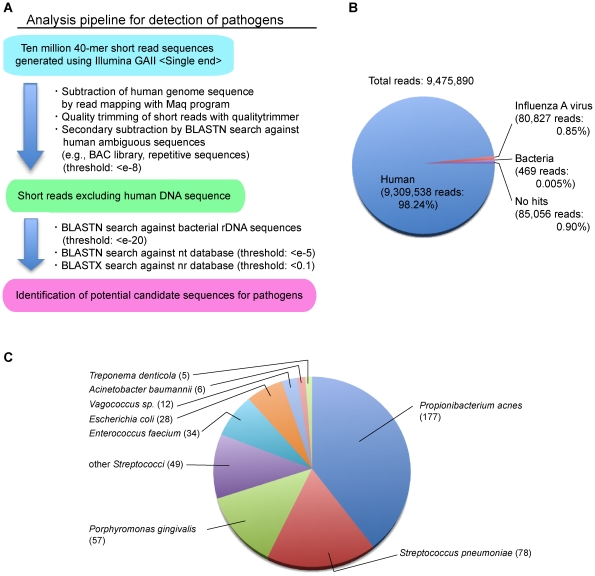
To determine the influenza A virus sequence and other potential pathogens, we performed de novo sequencing of double-stranded cDNA from total RNA extracted from the autopsy lung of a patient infected with the A/H1N1/2009 virus (A/Nagano/RC1/2009(H1N1)) in August 2009 in Japan. The patient was found to be positive for A/H1N1/2009 by real-time reverse transcriptase-polymerase chain reaction (RT-PCR); histopathological findings were also reported [15]. GA II produced 9.4×106 40-mer reads from the cDNA library (Fig. 1B). To exclude the human-derived read sequences, we performed an analysis pipeline as follows (Fig. 1A). Initially, all 9,475,890 reads were aligned to a reference sequence of human genomic DNA, followed by quality trimming to remove low-quality reads and excluding reads with similarities to ambiguous human sequences, resulting in 9,309,538 reads (98.24%) with a possible human source (Fig. 1B). The remaining 166,352 reads (1.75%) were further analyzed using a BLAST search against non-redundant databases, revealing 80,827 (0.85%), 469 (0.005%), and 85,056 (0.90%) reads as influenza A virus, bacteria, and no hits, respectively (Fig. 1B).
Figure 1. Detection of potential pathogens by comprehensive de novo sequencing.
(A) Schematic representation of the analysis pipeline for the detection of pathogens from comprehensive sequencing of human clinical specimens. After excluding human-derived DNA sequences using Maq software and a BLAST homology search against human genomic DNA and human ambiguous sequences extracted from the nt database, the remaining short reads were subjected to a BLAST search to detect potential pathogens. (B) Pie chart of the homology search results for the 40-mer short reads. Read numbers and their percentage to the total reads are shown in parenthesis. (C) Pie chart of identified bacterial hits. Number of hit reads is shown in parenthesis. Bacteria with less than 5 hit reads were excluded.
Regarding the bacterial hits, species classification was determined based on the results of a BLASTN search against the nt database (Fig. 1C). The most abundant bacterium was Propionibacterium acnes, but our other sequencing trials for clinical specimens suggest that this species is always detected (data not shown). Therefore, the presence of P. acnes could be the result of contamination at some point from the autopsy to the preparation of the cDNA library. In addition to P. acnes, Escherichia coli and Acinetobacter baumannii were frequently detected as possible contaminants. Suspected bacterial pathogens were identified as Streptococcus pneumoniae and Porphyromonas gingivalis. Specific PCR using 16S-rDNA and the lytA gene was performed for further verification of the presence of S. pneumoniae (data not shown). Although S. pneumoniae was not sufficiently abundant to conclude a coinfection with A/H1N1/2009, the severity of the A/H1N1/2009 infection could be correlated with S. pneumoniae, as reported by Palacios et al [16]. The other detected bacteria, such as Streptococcus sp., generally constitute the normal human oral flora.
de novo assembly of the A/H1N1/2009 virus
Whole 40-mer short reads, including human-derived reads, were assembled using Euler-SR or the Velvet de novo assembler. The resultant contigs generated using Euler-SR had longer extended sequences than those generated using Velvet (data not shown); thus, all further analyses were performed using the contigs generated using Euler-SR (Texts S1 and S2). All contigs showed high similarity to the sequences of A/H1N1/2009 (Table 1). Among the 8 segments, almost the whole lengths of segments 2 (2321 nt), 3 (2231 nt), 4 (1765 nt), 5 (1562 nt), 7 (1026 nt), and 8 (892 nt) were correctly assembled as single contigs of 2204, 2198, 1761, 1514, 1019, and 834 nt, respectively (Table 1 and Fig. 2). Segments 1 and 6 were divided into several contigs, but were correctly aligned to the known sequences (Table 1 and Fig. 2).
Table 1. BLASTN search results of de novo assembly contigs against database of Influenza virus sequences.
| Euler-SR_contigs | Contig length (bp) | Virus segment | Top hit of accession number using BLASTN search against databse of Influenza virus sequences | Length of subject (bp) | Identities | Contig location for A/Toronto/T0106/2009(H1N1) |
| >826 183 2835 | 183 | 1 | gb|GQ328865|INFLUENZA A virus (A/Finland/553/2009(H1N1)) segment 1 polymerase PB2 (PB2) | 2345 | 167/168 (99%) | 5–168 |
| >324 1558 136 | 1558 | 1 | gb|GQ365425|INFLUENZA A virus (A/Fukushima/1/2009(H1N1)) segment 1 polymerase PB2 (PB2) | 2280 | 1556/1558 (99%) | 201–1758 |
| >1194 239 112 | 239 | 1 | gb|GQ894926|INFLUENZA A virus (A/Delaware/03/2009(H1N1)) segment 1 polymerase PB2 (PB2) | 2280 | 214/214 (100%) | 1894–2107 |
| >887 174 3294 | 174 | 1 | gb|GQ894833|INFLUENZA A virus (A/Rhode Island/08/2009(H1N1)) segment 1 polymerase PB2 (PB2) | 2280 | 156/156 (100%) | 2145–2300 |
| >890 2204 4651 | 2204 | 2 | gb|GQ894924|INFLUENZA A virus (A/New Mexico/04/2009(H1N1)) segment 2 polymerase PB1 (PB1) | 2274 | 2200/2204 (99%) | 41–2244 |
| >696 2198 3968 | 2198 | 3 | gb|GQ866924|INFLUENZA A virus (A/Thailand/CU-H106/2009(H1N1)) segment 3 polymerase PA (PA) | 2238 | 2152/2155 (99%) | 54–2208 |
| >868 1761 3831 | 1761 | 4 | gb|CY045503|INFLUENZA A virus (A/Bayern/66/2009(H1N1)) segment 4 sequence | 1754 | 1750/1754 (99%) | 1–1741 |
| >897 1514 1710 | 1514 | 5 | gb|GQ502907|INFLUENZA A virus (A/Toronto/R8557/2009(H1N1)) segment 5 nucleocapsid protein | 1558 | 1511/1514 (99%) | 36–1549 |
| >224 101 9 | 101 | 6 | gb|GQ502908|INFLUENZA A virus (A/Toronto/R8557/2009(H1N1)) segment 6 neuraminidase (NA) | 1458 | 101/101 (100%) | 3–103 |
| >1206 1302 2468 | 1302 | 6 | gb|GQ906584|INFLUENZA A virus (A/Stockholm/49/2009(H1N1)) segment 6 neuraminidase (NA) | 1447 | 1299/1300 (99%) | 124–1423 |
| >750 1019 1128 | 1019 | 7 | gb|CY045957|INFLUENZA A virus (A/Toronto/T0106/2009(H1N1)) segment 7 sequence | 1026 | 1017/1017 (100%) | 9–1025 |
| >809 834 4399 | 834 | 8 | gb|GQ485660|INFLUENZA A virus (A/Ekaterinburg/01/2009(H1N1)) segment 8 nuclear export | 877 | 828/830 (99%) | 52–881 |
Schematic representation of contigs is shown in Fig. 2.
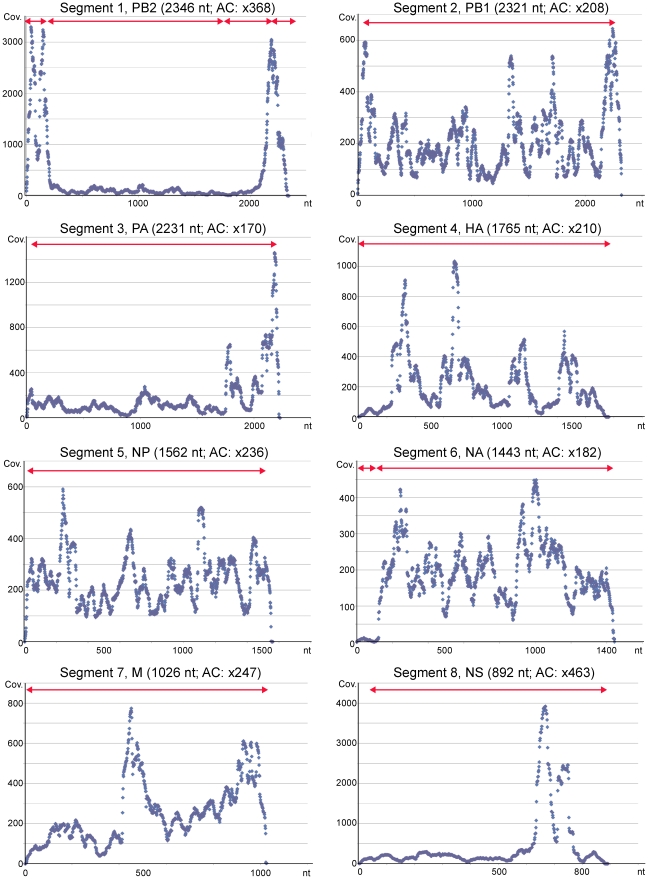
Figure 2. Dot plot of short read coverage (Cov.) at every nucleotide for the 8 segments of A/Nagano/RC1-L/2009(H1N1).
To obtain the consensus sequences for the respective 8 segments, 40-mer short reads were aligned to the complete segment sequences of A/Tronto/T0106/2009(H1N1) (gb|CY045951.1 – .8). Short read sequencing was performed using total RNA including human RNA, and also vRNA, cRNA, and mRNA from influenza A virus; thus, coverage bias was detected throughout the segments, but the average coverage (AC) is likely to be similar at approximately ×200 or more. The horizontal red arrows show the location of the contigs obtained by de novo assembly, as shown in Table 1.
Read mapping analysis of the A/H1N1/2009 virus
To obtain whole sequences and identify single nucleotide polymorphisms (SNPs) for the 8 segments, the 40-mer short reads were aligned to the sequence of A/Tronto/T0106/2009(H1N1), which was found to be the most similar to the A/H1N1/2009 virus using a BLASTN search. Figure 2 shows dot plot images of the coverage at every nucleotide of the segments. Read coverage was observed at ∼×200 on average for all segments, indicating a sufficient redundancy to identify the viral sequences and SNPs. The obtained viral sequences, designated as A/Nagano/RC1-L/2009(H1N1), were consistent with those from A/Nagano/RC1/2009(H1N1) passaged in the Madin-Darby canine kidney (MDCK) cell line, except for 3 possible heterogeneous nucleotides in HA.
The coverage plot curves were not flat throughout the segments. Intriguingly, both ends of segment 1 (encoding PB2), the 3′–end of segment 3 (encoding PA), and approximately 700 nt of segment 8 (encoding NS) showed significant abundant coverage greater than ×1000.
Genetic population analysis of the A/H1N1/2009 virus
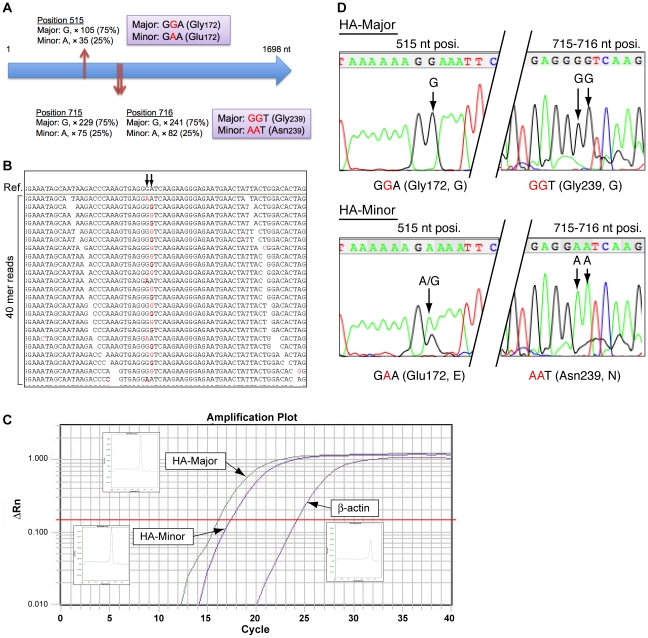
To identify heterogeneous populations, alignment results were screened using MapView software (Fig. 3B). Three potential heterogeneous genetic populations were found in segment 4 (encoding HA) at the 515, 715, and 716 nt positions (Fig. 3A), but not in other segments. The read alignments shown in Fig. 3B indicate that either the GGT or AAT sequence appeared at the 715–717 nt position, but not the GAT or AGT sequence. In addition, the read coverage implied that the major (GG; HA-Major) or minor (AA; HA-Minor) nucleotides were detected at the frequencies of 75% and 25%, respectively. To validate these variations, HA-Major- or HA-Minor-specific quantitative RT-PCR (qRT-PCR) was performed for the preparation of specific PCR products between the 434 and 738 nt in the HA coding sequence (Fig. 3C). qRT-PCR demonstrated that the expression of HA was ∼40,000-fold greater than that of human β-actin, and the expression ratio of HA-Major/HA-Minor was 4.05, suggesting that it corresponds to the read mapping shown in Fig. 3A. Furthermore, HA-Major and HA-Minor sequences were verified by Sanger DNA sequencing of the specific PCR products (Fig. 3D). Taken together, these results suggest the following amino acid substitutions of HA: one nucleotide alteration causes Gly172Glu and the other two alterations cause Gly239Asn (Fig. 3D).
Figure 3. Genetic variations of the HA nucleotide sequence.
(A) Schematic representation of 3 nucleotide variations (positions 515, 715, and 716 nt) in the HA coding nucleotide sequence. Three variations were classified as Major (75% appearance) or Minor (25% appearance) by read coverage (×), and the coding amino acids are also shown. (B) Arrows indicate positions 715 and 716 nt of the HA sequence, and the alignment image of the 40-mer reads. Nucleotides shown in red are the mismatches to the reference sequence of A/Tronto/T0106/2009(H1N1). Every read suggested that either the GGT or AAT sequence was present, but not the GAT or AGT sequence. (C) An amplification plot for HA-specific qRT-PCR. (D) Validation of genetic variation by Sanger capillary sequencing. HA-Major or HA-Minor PCR products were obtained by qRT-PCR using HA-Major- or HA-Minor-specific PCR primers. HA-Major PCR product shows G nucleotides at positions 515, 715, and 716 nt, while HA-Minor shows A nucleotides.
Epitope analysis of heterogeneous HA
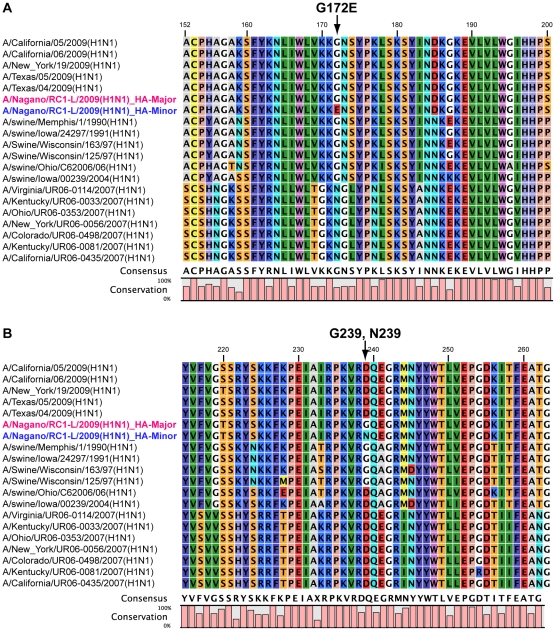
To elucidate whether the Gly172Glu and Gly239Asn amino acid substitutions in the HA sequence could be associated with antigenic drift, they were compared to known potential epitopes [8], [9]. Representative HA amino acid sequences of the H1N1 influenza A virus were aligned with the heterogeneous HA-Major and HA-Minor sequences. The Gly172Glu substitution (corresponding to Gly158 in the mature HA lacking a signal peptide) was located on the Sa antigenic site (Fig. 4A).
Figure 4. Alignment of HA amino acid sequences in influenza A virus around the identified mutations in A/Nagano/RC1-L/2009(H1N1).
(A) Genetic variation at position 515 nt causes the amino acid substitution Gly172Glu; HA-Major: Gly172, HA-Minor: Glu172. (B) Genetic variation at position 715 and 716 nt causes the amino acid substitution Gly239Asn; HA-Major: Gly239, HA-Minor: Asn239.
The HA Gly172Glu substitution is likely to be rare thus far because a BLASTP search against the non-redundant nr database revealed only two identical hits, A/Bayern/62/2009(H1N1) in Germany and A/Catalonia/S1207/2009(H1N1) in Spain (data not shown). One intriguing hit was to A/Pennsylvania/14/2009(H1N1) isolated in the US, whose HA sequence has an Xaa amino acid at position 172 due to the presence of the heterogeneous nucleotide R (A or G) (Fig. 5A). This deposited sequence completely coincides with our observation, suggesting that two variants of HAs are likely to coexist in the human lung, further implying that such heterogeneous populations might frequently be generated during infection.
Figure 5. BLAST homology search of the HA sequences against the nr or nt databases.
(A) Glu172 of HA-Minor. (B) Gly239 of HA-Major. R: A or G.
Furthermore, HA Gly239Asn was located on the Ca2 antigenic site that contributes to binding with the host's sialic acid receptor [17]. Asp239 (corresponding to Asp225 in the mature HA lacking a signal peptide) was frequently distributed in H1N1 HAs (Fig. 4B), but Gly239 and Asn239 were found to be minor amino acids among HAs; a BLASTN search found 18 and 5 hit entries on the nt database, respectively. As was observed for Gly172, Xaa239 was found in the nt database (Fig. 5B), suggesting that HA heterogeneity of both minor amino acids may affect its binding affinity to the sialic acid receptor.
Discussion
In this study, we demonstrated the detection of potential pathogens using a next-generation DNA sequencer. We speculated that, in addition to influenza A virus, additional potential pathogens such as S. pneumoniae could contribute to the severity and fatality of infection with the A/H1N1/2009 virus [2], [16]. In this case, the amount of bacteria detected was small (Fig. 1B and 1C), and they were considered to be the result of contamination during the course of the experiment, from autopsy to short read sequencing. The clinical outcome of the patient and histopathological examination suggest that this was a case of influenza viral pneumonia rather than bacterial infection [15], although S. pneumoniae coinfection has been reported to play a crucial role in the severity of influenza virus infection in some cases [16], [18].
In the present autopsy lung sample, very significant viral copies (∼40,000-fold greater than β-actin) were determined using qRT-PCR, but this was not always observed in autopsy samples from other patients (less than β-actin) (data not shown). Such abundant read sequencing enabled us to obtain almost full-coverage contig sequences for the viral segments using de novo assembly, suggesting the importance of this result in terms of being able to evaluate uncharacterized emerging infectious agents using an unbiased sequencing technique at the outbreak of a pandemic. Indeed, this study demonstrated that whole contigs can be identified as A/H1N1/2009, but not seasonal H1N1 or other subtypes (Table 1).
The read coverage profile generated by mapping was very indicative for segment 1 encoding PB2 (Fig. 2). Both ends were highly redundant with up to ×3000 coverage. The coverage is reflected by the amounts of vRNA, cRNA, and mRNA of influenza A virus, implying that the coverage bias may detect a more stable region as it is dependent on the expression level or manner of replication.
Contrary to the viral sequences obtained for A/Nagano/RC1/2009(H1N1) isolated from passaging in MDCK cells, de novo sequencing revealed the presence of A/Nagano/RC1-L/2009(H1N1) HA quasispecies in the autopsy sample (Fig. 3). Despite the fact that immunity to A/H1N1/2009 viruses is supposed to be limited among the general human population [19], we detected the amino acid substitution Gly172Glu in the HA Sa antigenic site in A/Nagano/RC1-L/2009(H1N1), and this appears to be a very rare event among A/H1N1/2009 viruses to date.
We also observed another substitution of Gly239Asn in the HA Ca2 antigenic site of A/Nagano/RC1-L/2009(H1N1). This antigenic site plays a crucial role in conferring specificity to galactose of the human α2-6 sialylated glycan receptor [20]. Interestingly, Asp239 (corresponding to Asp225 in the mature HA that lacks a signal peptide) is highly prevalent in known H1N1 HAs, indicating that both Gly239 and Asn239 appear to be very minor amino acids among all HA sequences.
Thus far, amino acid substitutions in the HAs of A/H1N1/2009 have been identified compared with seasonal H1N1 HAs. Homology-based structural investigations [17], [21] suggest that A/H1N1/2009 HA has the necessary residues to provide optimal contacts for high affinity binding to α2-6 sialylated glycans present in the human upper airway [22], while it apparently shows minimal contact with α2-3 sialylated glycans present in the human lower respiratory tract [23]. Indeed, the recombinant A/H1N1/2009 HA has been characterized to exhibit lower binding to the alveolar tissue of the lower respiratory tract [17]. However, we previously detected abundant viral nucleoprotein of A/Nagano/RC1-L/2009(H1N1) in pneumocytes expressing α2-3 sialylated glycans in autopsy lung tissue sections [15], suggesting that the above substitutions could affect the binding affinity to both types of sialylated glycans.
Very suggestive reports predicted the possible future antigenic drift of A/H1N1/2009 viruses from viral sequence and structural comparative analyses [11], [12]. Prior to the initiation of the current study (September 2009), Igarashi et al. predicted possible substitutions and these included the two amino acid substitutions presented here (Gly172Glu and Asp239Gly) [11]. Furthermore, Shen et al. suggested that host-driven antigenic drift based on evolutionary trends appeared to favor Asp239 (corresponding to Asp225 in the mature HA) in swine HAs and the 1918 pandemic, while Asp204 (corresponding to Asp190 in the mature HA) was favored in seasonal H1N1 HAs [12]. These predictions are very attractive and our experiments demonstrated one of them a posteriori. Furthermore, recent study has shown that receptor-binding avidity can influence antigenic drift [24]. HA antigenic sites Sa is the membrane proximal region, therefore, the identified variations on both Sa and Ca2 might contribute to the alteration of antigenicity and receptor-binding avidity by synergistic effect. The newly identified Asn239 substitution could be a probable candidate for further investigation of the manner of viral binding to sialic acid on the host receptors.
In conclusion, this study demonstrated that de novo sequencing can comprehensively detect pathogens, and such in-depth investigation facilitates the identification of influenza A viral heterogeneity during infection. The possibility of mixed infections with variants remains to be elucidated in this case, but worldwide sequencing efforts suggest that quasispecies of the A/H1N1/2009 virus evidently appear and are observed. To better characterize the currently emerging A/H1N1/2009 virus and prevent worse pandemics in the near future, unbiased de novo sequencing techniques will be indispensable for the primary investigations of emerging infectious diseases.
Materials and Methods
Ethics Statement
The study protocol was approved by the institutional medical ethical committee, National Institute of Infectious Diseases, Japan (Approval No.236), and the study was conducted according to the Declaration of Helsinki Principles. In the autopsy case, a written consent for autopsy was obtained from relatives.
Total RNA and cDNA preparation from autopsy human lung
Information for the patient was previously reported [15]. Briefly, in August 2009, a 33-year-old male patient with chronic heart failure due to dilated cardiomyopathy, mild diabetes mellitus, atopic dermatitis, asthma, and obesity (BMI: 38) died from respiratory failure and multiple organ dysfunction syndrome. A diagnosis of pandemic influenza A virus (A/H1N1/2009) infection was determined using RT-PCR testing in a clinical laboratory. Total RNA was prepared from a 5-mm cube of the autopsy lung tissue using ISOGEN (NipponGene, Japan), followed by Ambion TurboDNase treatment (Ambion, Austin, TX USA). Double-stranded cDNA was prepared from 5 µg of total RNA using the random priming method with SuperScript Choice System for cDNA synthesis (Invitrogen, Carlsbad, CA, USA). cDNA was purified using a QIAquick PCR Purification kit (QIAGEN, Hilden, Germany).
Short-read DNA sequencing using the Illumina Genome Analyzer II
An approximately 300-bp length cDNA library was prepared using a genomic DNA sample prep kit (Illumina, San Diego, CA, USA), and DNA clusters were generated on a slide using a Cluster Generation kit (ver. 2) on an Illumina cluster station (Illumina), according to the manufacturer's instructions. To obtain ∼1.0×107 clusters for one lane, the general procedure as described in the standard recipe (Illumina) was performed as follows: template hybridization, isothermal amplification, linearization, blocking, denaturation, and hybridization of the sequencing primer (Illumina). All sequencing runs for 40 mers were performed with GA II using the Illumina Sequencing kit (ver. 3). Fluorescent images were analyzed using the Illumina base-calling pipeline 1.4.0 to obtain FASTQ formatted sequence data.
Homology search analysis
The obtained DNA sequences were investigated using a BLAST search as shown in Fig. 1A. The results of the BLASTN search were analyzed and visualized using MEGAN v3.7.4 [25] with the following parameters: minimum support, 5; minimum score, 35.0.
de novo assembly of short reads
Prior to de novo assembly, all obtained 40-mer reads were trimmed based on the phred quality value obtained using the Euler-SR qualitytrimmer command [26]. Such trimmed read sequences were assembled using Velvet v0.7.55 [27] or Euler-SR v1.0 [26] with the default parameters (Velvet: hash length, 25; Euler-SR: vertex size, 25).
Read mapping
To obtain consensus sequences for the respective 8 segments of influenza A virus, 40-mer short reads were aligned to A/Tronto/T0106/2009(H1N1) sequences (gb|CY045951.1 – .8) as reference sequences with Maq software (ver. 0.7.1) [28] using the easyrun Perl-command. The consensus sequences were extracted as a “cns.fq” file for each segment, and deposited in the DNA Data Bank of Japan (DDBJ; accession numbers: AB538386 to AB538393 for the 8 segments of A/Nagano/RC1-L/2009(H1N1), and AB538394 for segment 4 encoding the HA-Minor sequence). Read coverage at every nucleotide was obtained using Maq software (ver. 0.7.1) with the pileup command. Read alignment for the validation of SNPs was performed using the MapView graphical alignment viewer [29].
qRT-PCR analysis
qRT-PCR was performed using 100 ng of total RNA, HA variant-specific primers (forward common primer: pdmFlu09-HA-F, 5′–CGAACAAAGGTGTAACGGCAGCAT–3′; HA-Major-specific reverse primer: pdmFlu-HA-R_Major, 5′–ATAGTTCATTCTCCCTTCTTGACC–3′; HA-Minor-specific reverse primer: 5′–ATAGTTCATTCTCCCTTCTTGATT–3′), and the SuperScript III Platinum SYBR Green One-Step qRT-PCR kit with ROX (Invitrogen), and analyzed using the ABI PRISM 7900HT Real-time PCR System (Applied Biosystems, Foster City, CA, USA). The following qRT-PCR program was used: RT reaction, 50°C for 3 min; initial denaturation, 95°C for 5 min; 2 steps of amplification (×40 cycles), 95°C for 15 sec and 60°C for 30 sec. The human β-actin gene was used as the internal control. PCR products were resolved by 5% polyacrylamide gel electrophoresis, followed by Sanger sequencing using the BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems).
Virus isolation
The A/H1N1/2009 virus was isolated from MDCK cells passaged once with trypsin.
Supporting Information
Fastq file of the 40-mer short reads with similarity to influenza A virus extracted from whole obtained short reads.
(11.19 MB PDF)
De novo assembly of the influenza A virus using Euler-SR v1.0 [26] with the default parameters (vertex size, 25).
(0.04 MB RTF)
Acknowledgments
We thank Tadahito Kanda for valuable suggestions.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: This work was supported by grants for Research on Emerging and Re-emerging Infectious Diseases (H20 and 21 Shinko-Ippan-6), from the Ministry of Health, Labour, and Welfare, Japan. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, et al. Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med. 2009;360:2605–2615. doi: 10.1056/NEJMoa0903810. [DOI] [PubMed] [Google Scholar]
- 2.Gill JR, Sheng ZM, Ely SF, Guinee DG, Beasley MB, et al. Pulmonary pathologic findings of fatal 2009 pandemic influenza A/H1N1 viral infections. Arch Pathol Lab Med. 2010;134:235–243. doi: 10.5858/134.2.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ghedin E, Sengamalay NA, Shumway M, Zaborsky J, Feldblyum T, et al. Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution. Nature. 2005;437:1162–1166. doi: 10.1038/nature04239. [DOI] [PubMed] [Google Scholar]
- 4.Kilbourne ED. Influenza pandemics of the 20th century. Emerg Infect Dis. 2006;12:9–14. doi: 10.3201/eid1201.051254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Nelson MI, Viboud C, Simonsen L, Bennett RT, Griesemer SB, et al. Multiple reassortment events in the evolutionary history of H1N1 influenza A virus since 1918. PLoS Pathog. 2008;4:e1000012. doi: 10.1371/journal.ppat.1000012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Knossow M, Skehel JJ. Variation and infectivity neutralization in influenza. Immunology. 2006;119:1–7. doi: 10.1111/j.1365-2567.2006.02421.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Air GM, Laver WG, Webster RG. Antigenic variation in influenza viruses. Contrib Microbiol Immunol. 1987;8:20–59. [PubMed] [Google Scholar]
- 8.Caton AJ, Brownlee GG, Yewdell JW, Gerhard W. The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell. 1982;31:417–427. doi: 10.1016/0092-8674(82)90135-0. [DOI] [PubMed] [Google Scholar]
- 9.Bui HH, Peters B, Assarsson E, Mbawuike I, Sette A. Ab and T cell epitopes of influenza A virus, knowledge and opportunities. Proc Natl Acad Sci U S A. 2007;104:246–251. doi: 10.1073/pnas.0609330104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Squires B, Macken C, Garcia-Sastre A, Godbole S, Noronha J, et al. BioHealthBase: informatics support in the elucidation of influenza virus host pathogen interactions and virulence. Nucleic Acids Res. 2008;36:D497–503. doi: 10.1093/nar/gkm905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Igarashi M, Ito K, Yoshida R, Tomabechi D, Kida H, et al. Predicting the antigenic structure of the pandemic (H1N1) 2009 influenza virus hemagglutinin. PLoS One. 2010;5:e8553. doi: 10.1371/journal.pone.0008553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Shen J, Ma J, Wang Q. Evolutionary trends of A(H1N1) influenza virus hemagglutinin since 1918. PLoS One. 2009;4:e7789. doi: 10.1371/journal.pone.0007789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nakamura S, Yang CS, Sakon N, Ueda M, Tougan T, et al. Direct metagenomic detection of viral pathogens in nasal and fecal specimens using an unbiased high-throughput sequencing approach. PLoS One. 2009;4:e4219. doi: 10.1371/journal.pone.0004219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ramakrishnan MA, Tu ZJ, Singh S, Chockalingam AK, Gramer MR, et al. The feasibility of using high resolution genome sequencing of influenza a viruses to detect mixed infections and quasispecies. PLoS One. 2009;4:e7105. doi: 10.1371/journal.pone.0007105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Nakajima N, Hata S, Sato Y, Tobiume M, Katano H, et al. The first autopsy case of pandemic influenza (A/H1N1pdm) virus infection in Japan: Detection of high copy number of the virus in type II alveolar epithelial cells by pathological and virological examination. Jpn J Infect Dis. 2010;63:67–71. [PubMed] [Google Scholar]
- 16.Palacios G, Hornig M, Cisterna D, Savji N, Bussetti AV, et al. Streptococcus pneumoniae coinfection is correlated with the severity of H1N1 pandemic influenza. PLoS One. 2009;4:e8540. doi: 10.1371/journal.pone.0008540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Maines TR, Jayaraman A, Belser JA, Wadford DA, Pappas C, et al. Transmission and pathogenesis of swine-origin 2009 A(H1N1) influenza viruses in ferrets and mice. Science. 2009;325:484–487. doi: 10.1126/science.1177238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Louie J, Jean C, Chen T-H, Park S, Ueki R, et al. Bacterial coinfections in lung tissue specimens from fatal cases of 2009 pandemic influenza A (H1N1) - United States, May-August 2009. MMWR Morb Mortal Wkly Rep. 2009;58:1071–1074. [PubMed] [Google Scholar]
- 19.Greenbaum JA, Kotturi MF, Kim Y, Oseroff C, Vaughan K, et al. Pre-existing immunity against swine-origin H1N1 influenza viruses in the general human population. Proc Natl Acad Sci U S A. 2009;106:20365–20370. doi: 10.1073/pnas.0911580106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gamblin SJ, Haire LF, Russell RJ, Stevens DJ, Xiao B, et al. The structure and receptor binding properties of the 1918 influenza hemagglutinin. Science. 2004;303:1838–1842. doi: 10.1126/science.1093155. [DOI] [PubMed] [Google Scholar]
- 21.Soundararajan V, Tharakaraman K, Raman R, Raguram S, Shriver Z, et al. Extrapolating from sequence–the 2009 H1N1 ‘swine’ influenza virus. Nat Biotechnol. 2009;27:510–513. doi: 10.1038/nbt0609-510. [DOI] [PubMed] [Google Scholar]
- 22.Chandrasekaran A, Srinivasan A, Raman R, Viswanathan K, Raguram S, et al. Glycan topology determines human adaptation of avian H5N1 virus hemagglutinin. Nat Biotechnol. 2008;26:107–113. doi: 10.1038/nbt1375. [DOI] [PubMed] [Google Scholar]
- 23.Shinya K, Ebina M, Yamada S, Ono M, Kasai N, et al. Avian flu: influenza virus receptors in the human airway. Nature. 2006;440:435–436. doi: 10.1038/440435a. [DOI] [PubMed] [Google Scholar]
- 24.Hensley SE, Das SR, Bailey AL, Schmidt LM, Hickman HD, et al. Hemagglutinin receptor binding avidity drives influenza A virus antigenic drift. Science. 2009;326:734–736. doi: 10.1126/science.1178258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mitra S, Klar B, Huson DH. Visual and statistical comparison of metagenomes. Bioinformatics. 2009;25:1849–1855. doi: 10.1093/bioinformatics/btp341. [DOI] [PubMed] [Google Scholar]
- 26.Chaisson MJ, Pevzner PA. Short read fragment assembly of bacterial genomes. Genome Res. 2008;18:324–330. doi: 10.1101/gr.7088808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zerbino DR, Birney E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008;18:821–829. doi: 10.1101/gr.074492.107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li H, Ruan J, Durbin R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 2008;18:1851–1858. doi: 10.1101/gr.078212.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bao H, Guo H, Wang J, Zhou R, Lu X, et al. MapView: visualization of short reads alignment on a desktop computer. Bioinformatics. 2009;25:1554–1555. doi: 10.1093/bioinformatics/btp255. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Fastq file of the 40-mer short reads with similarity to influenza A virus extracted from whole obtained short reads.
(11.19 MB PDF)
De novo assembly of the influenza A virus using Euler-SR v1.0 [26] with the default parameters (vertex size, 25).
(0.04 MB RTF)