Abstract
Expression of the five β-like globin genes (ϵ, Gγ, Aγ, δ, β) in the human β-globin locus depends on enhancement by the locus control region, which consists of five DNase I hypersensitive sites (5′HS1 through 5′HS5). We report here a novel enhancer activity in 5′HS1 that appears to be potent in transfected K562 cells. Deletion analyses identified a core activating element that bound to GATA-1, and a two-nucleotide mutation that disrupted GATA-1 binding in vitro abrogated 5′HS1 enhancer activity in transfection experiments. To determine the in vivo role of this GATA site, we generated multiple lines of human β-globin YAC transgenic mice bearing the same two-nucleotide mutation. In the mutant mice, ϵ-, but not γ-globin, gene expression in primitive erythroid cells was severely attenuated, while adult β-globin gene expression in definitive erythroid cells was unaffected. Interestingly, DNaseI hypersensitivity near the 5′HS1 mutant sequence was eliminated in definitive erythroid cells, whereas it was only mildly affected in primitive erythroid cells. We therefore conclude that, although the GATA site in 5′HS1 is critical for efficient ϵ-globin gene expression, hypersensitive site formation per se is independent of 5′HS1 function, if any, in definitive erythroid cells.
Keywords: Chromatin Immunoprecipitation (ChiP), Hemoglobin, Mouse, Transcription Enhancers, Transgenic, DNaseI Hypersensitive Site, GATA, YAC, β-Globin
Introduction
The human β-globin locus consists of five β-like globin genes (ϵ, Gγ, Aγ, δ, β; Fig. 1A) whose temporal and spatial expression pattern switches twice during development (1). While the embryonic ϵ gene is expressed in the yolk sac, the two γ genes are expressed in the fetal liver. In contrast, the δ and β genes are expressed in the adult spleen and bone marrow. High level expression of all the β-like globin genes depends on the locus control region (LCR)2 enhancer activity (2, 3), which is located 6–22 kbp upstream of the ϵ-globin gene and consists of one ubiquitous and four erythroid-specific DNase I hypersensitive sites (5′HS5 and 5′HS4-1, respectively, Fig. 1A) (4).
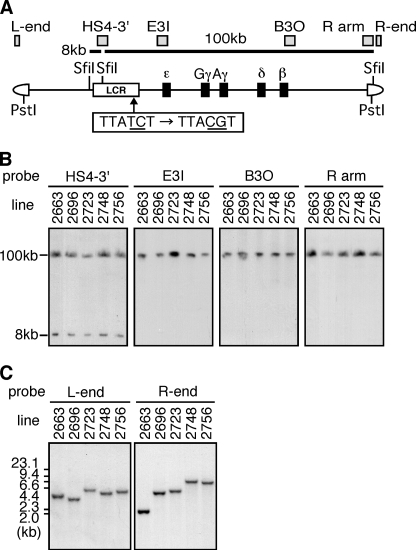
FIGURE 1.
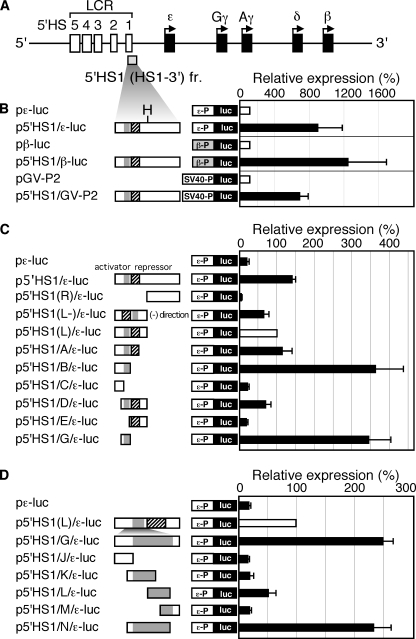
Identification of core sequences of the 5′HS1 enhancer. A, structure of the human β-globin locus. The 5′HS1 region (formerly termed HS1–3′, (24) is located from −6189 to −5248 nucleotides upstream of the ϵ-globin transcription start site. H: HindIII site. B–D, 5′HS1 fragment and its deletion mutants were linked to the luciferase (luc) reporter gene under the control of the ϵ-globin (ϵ-P; −174 to +38, relative to transcription start site), β-globin (β-P), or SV40 (SV40-P) promoters. These reporter constructs, as well as the control plasmid CMV-β-gal, were cotransfected into K562 cells. Luciferase activities were normalized to β-Gal activities to control for transfection efficiencies. The luciferase activities are expressed relative to those (open bars, set at 100) in the constructs without a linked enhancer fragment in B or in the p5′HS1(L)/ϵ-luc in C and D. Each value represents the mean ± S.D. for at least three independent experiments. Gray and hatched boxes represent putative enhancer and repressor sequences, respectively.
The mechanism leading to the highly reproducible pattern of globin gene switching has been intensively studied by introducing mutations into the endogenous mouse locus or into the human locus followed by subsequent analysis in transgenic mice (TgM). In the latter cases, the whole human locus has been introduced into the mouse genome via cosmid or yeast artificial chromosome (YAC) vectors, in which gene switching was reasonably faithfully recapitulated (5–7). Because the mouse exhibits a single switch, however, the human ϵ and γ (high level) genes are expressed in the yolk sac (in primitive erythroid cells), whereas the γ (low level), δ, and β genes are expressed in the fetal liver and spleen/bone marrow (definitive erythroid cells) of TgM.
Transient cell transfection assays in K562 and MEL cells have reproduced partial LCR enhancer activity in 5′HS2, but not in 5′HS3 or 5′HS4 (8, 9), while stable transfection and TgM assays revealed significant enhancer activity in all these hypersensitive sites (9–11). It is not fully understood whether individual HSs independently or cooperatively contribute to gene switching in the β-globin locus. In the mouse locus, individual HSs appeared to be structurally and functionally independent and each additively contributes to LCR enhancer activity (12, 13). Although human HSs have distinct preferences for gene activation when they are separately analyzed (14, 15), other in vivo evidence demonstrated that the HSs form a single functional unit (the LCR holocomplex) (16), in which 5′HS2, 3, and 4 are structurally and functionally involved (17–20). Although 5′HS1 is erythroid-specific and stable throughout development (2), it has not been established whether or not 5′HS1 is a component of the holocomplex.
5′HS1 was capable of activating a γ-globin promoter by severalfold in stable, but not in transient, transfection assays in K562 cells (representing primitive erythrocytes) (9), suggesting that 5′HS1 requires integration into chromatin for its enhancing activity to be detected. In stably transfected MEL cells (definitive erythrocytes), however, 5′HS1 was barely capable of activating the adult β-globin promoter (10, 21). In TgM, 5′HS1 activated the γ- and β-globin genes (the ϵ gene was not examined) to quite low levels at all developmental stages (14). In accord with these results, an intact 5′HS1 in a patient bearing the Hispanic thalassemia deletion, in which 5′HS2 through HS5 as well as additional upstream sequences are removed, is unable to significantly activate the adult β-globin gene (22). Furthermore, a small deletion including 5′HS1 in an Italian thalassemia patient, which was in trans to the thalassemic allele, does not cause reduced adult β-globin gene expression in cis (23). These results therefore suggest that 5′HS1 may not play an important role, at least in adult β-globin gene activation, in the context of the entire locus. However, it is not known whether embryonic ϵ-globin gene expression may be activated in the Hispanic deletion allele and/or attenuated in the 5′HS1 deletion allele at an earlier developmental stage.
In our previous work (24), we were able to identify an enhancer activity lying within a 1 kb fragment around human 5′HS1, unexpectedly in transiently transfected K562 cells. We termed this activity an HS1–3′ enhancer, because it contained the 3′ portion of 5′HS1 and further downstream region (Fig. 1A).
In this current work, we further explored this observation and identified the core enhancer sequences to which transcription factor GATA-1 bound in vitro and in vivo. Because core enhancer activity was eventually localized within the 5′HS1 portion, we renamed and called this activity a “5′HS1” enhancer, hereafter (Fig. 1A). A two-nucleotide mutation, which disrupted in vitro GATA-1 binding, completely abolished the enhancer activity of 5′HS1 in the transfection assay. To determine the in vivo role of this GATA site, we generated five lines of intact, single copy human β-globin YAC TgM bearing the same two-nucleotide mutation in 5′HS1. Analysis of the mutant TgM revealed that GATA site in the 5′HS1 was crucial for ϵ-, but not γ-globin gene expression in the yolk sac, while it was unnecessary for β-globin gene expression in the adult spleen. ChIP analysis revealed significantly reduced occupancy of GATA-1 at the mutant 5′HS1 site, and consequently, DNaseI hypersensitivity at 5′HS1 disappeared in definitive erythroid cells. In contrast, 5′HS1 formation was only marginally affected in primitive erythroid cells, in which ϵ-globin gene expression was severely attenuated. Thus, although the lone GATA binding site in 5′HS1 of the LCR is indispensable for efficient ϵ-globin gene expression, the 5′HS1 structure anticorrelates with its transcriptional enhancing ability on target β-like globin genes.
EXPERIMENTAL PROCEDURES
Reporter Plasmid Construction
To generate pβ-luc, human β-globin promoter fragment (HpaI-NcoI, from nucleotides (nt) 61320 to 62190; GenBankTM, HUMHBB) was subcloned into SmaI/NcoI-digested pGV-B2 (TOYO INK). The 5′HS1 (formerly termed HS1–3′) fragment was excised as a KpnI-XbaI fragment from the pHS1–3′/ϵ-luc (24) and subcloned into KpnI/NheI-digested pβ-luc and pGV-P2 (TOYO INK) to generate p5′HS1/β-luc and p5′HS1/GV-P2, respectively.
The 5′HS1(R) fragment was excised as a BamHI-SacII fragment (from nt 13769 to 14240) from the plasmid pHS1–3′/ϵ-luc and subcloned into BglII/SacII-digested pϵ-luc (24) to generate p5′HS1(R)/ϵ-luc. The 5′HS1(L) fragment (from nt 13305 to 13765) was PCR-amplified with the primers HS1–5S and HS1–3A2, digested with XhoI enzyme and subcloned into XhoI-digested pϵ-luc in sense or antisense orientations to generate p5′HS1(L)/ϵ-luc and p5′HS1(l-)/ϵ-luc, respectively. The truncated 5′HS1(L) fragments, 5′HS1/A-E, G, J-O, V, L(m2), and V(m1, m2), were PCR-amplified with their respective primers, digested with XhoI and SacII enzymes, and subcloned into the XhoI/SacII-digested pϵ-luc. To generate p5′HS1/T/ϵ-luc, the oligonucleotides, HS1–5S/d9 and HS1–3A/d8 were annealed (from nt 13481 to 13504) and subcloned into the XhoI/SacII-digested pϵ-luc. To generate p5′HS1/Z/ϵ-luc, the oligonucleotides, HS1–5S/d10 and HS1–3A/d9 were annealed (from nt 13443 to 13467) and subcloned into the SmaI-digested pϵ-luc. Oligo sequences are listed in the supplemental Tables S1 and S2.
To generate fragments with the GATA site mutation (5′HS1(L)(m4′) and N(m4′)), artificially introduced SnaBI restriction enzyme site around the m4′ sequences was utilized. Briefly, PCR-amplified 5′ and 3′ part of the fragments were digested with an XhoI/SnaBI and SnaBI/SacII, respectively. These and XhoI/SacII-digested reporter plasmids (pϵ-luc or pGV-B2) were subjected to three-fragment ligation. Primer sequences are listed in the supplemental Tables S1 and S2. All the vector sequences were verified by DNA sequencing.
Transfection Assay
For DNA transfection into human erythroleukemia K562 cells, 950 ng of test constructs and 50 ng of pCMV-β-gal in 10 μl of H2O were mixed with 25 μl of Opti-MEM (Invitrogen), then with 4 μl of PLUS reagent (Invitrogen), followed by incubation for 15 min at room temperature. Separately, 2 μl of Lipofectamine (Invitrogen) and 25 μl of Opti-MEM reagents were mixed. Two mixtures were combined, vortexed, and incubated for 15 min at room temperature. Cells (4 × 105/200 μl of Opti-MEM/24-well dish) were exposed to the DNA/reagent mix for 4h at 37 °C. RPMI 1640 medium with 10% fetal bovine serum (1 ml) was added to each dish and incubated for another 48 h at 37 °C. Luciferase activity in the cell extracts was measured by an ARVOTMSX (Wallac Berthold) and normalized by β-galactosidase activity in the same sample. Assays were performed at least in triplicate, and average values with S.D. were calculated.
EMSA
Double-stranded L DNA fragment was end-labeled by using T4 polynucleotide kinase (TOYOBO) and [γ-32P]ATP. Nuclear extracts (3–7 μg), prepared by using Nuclear Extract Kit (Active Motif), were preincubated for 4 h at 4 °C in the reaction mixture (5 mm Tris-Cl, pH 8.0, 0.5 mm dithiothreitol, 0.5 mm EDTA, 25 mm NaCl, and 1% Ficoll) in the presence or absence of 10–100-fold molar excess of specific double-stranded competitor DNA or 2 μg of antibody. 0.5 ng (10,000 cpm) of a radiolabeled DNA probe was added, and the incubation was continued for 15–30 min at 4 °C. The incubation mixture was loaded on a 5% non-denaturing polyacrylamide gel in 0.5× TBE buffer, and electrophoresed at 4 °C with buffer circulation. The gels were dried and exposed to x-ray film. The competitor oligonucleotides used are listed in the supplemental Table S3. Anti-GATA1 (sc-1233) and anti-GATA2 (sc-9008) antibodies were obtained from Santa Cruz Biotechnology.
Chromatin Immunoprecipitation
Definitive erythroid cells were collected from phenylhydrazine-treated anemic adult spleen and made single cell suspension. K562 or adult spleen cells were fixed in PBS(-) with 1% formaldehyde for 10 min at room temperature, and the reaction was terminated by the addition of glycine to 0.125 m for 5 min at room temperature. Nuclei (from 1.5 × 107 cells) were prepared and digested with 0.01 units/μl of MNase at 37 °C for 10 min. DNA was prepared from a chromatin and used as the “input” sample. Chromatin was incubated with anti-GATA-1, anti-GATA-2 antibodies (above-described), or normal goat (sc-2028) or rabbit (sc-2027) IgGs overnight at 4 °C and precipitated with pre-blocked Dynabeads protein G magnetic beads (Invitrogen). Immunoprecipitated materials were then washed extensively and reverse cross-linked. DNA from input and bound samples was purified by QIAquick PCR purification kit (Qiagen) and subjected to quantitative real-time PCR (qPCR) analysis. Primer sequences used in the qPCR are listed in supplemental Table S4 (25). The qPCR was performed in the Thermal Cycler Dice (TAKARA) with cycling as follows: an initial cycle for 10 s at 95 °C, followed by 40 cycles of 5 s at 95 °C and 30 s at 60 °C.
YAC Mutagenesis and Transgenic Mice
The XhoI-SacII fragment from the p5′HS1(L)(m4′)/ϵ-luc was subcloned into XhoI/SacII-digested pRS306 to generate pRS306/5′HS1(L)mGATA. This plasmid DNA was linearized with SpeI and used for mutagenizing the human β-globin YAC (A201F4.3) (26, 27), to make A201F4.3/mutGATA/5′HS1. Successful homologous recombination in yeast was confirmed by Southern blot analyses with several combinations of restriction enzymes and probes. The SnaBI restriction enzyme site, which was artificially introduced by the 2-bp GATA site mutation, was utilized to distinguish mutated from wild-type YAC. The vector and YAC sequences were verified by DNA sequencing.
Generation and structural analysis of human β-globin YAC TgM has been described elsewhere (28). Animal experiments were carried out in a humane manner and were approved by the Institutional Animal Experiment Committee of the University of Tsukuba. Experiments were conducted in accordance with the Regulation of Animal Experiments of the University of Tsukuba and the Fundamental Guidelines for Proper Conduct of Animal Experiments and Related Activities in Academic Research Institutions under the jurisdiction of the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Semi-quantitative RT-PCR
To generate acute anemia and obtain abundant nucleated erythroid progenitor cells, adult mice were injected with N-acetyl-phenylhydrazine. Total RNA was extracted from yolk sacs (9.5 days postcoitum), fetal livers (14.5 days postcoitum), or anemic adult spleens (1–2 month old) by using ISOGEN (Nippon Gene). First strand cDNA was synthesized from 2.5 μg of RNA with ReverTra Ace (Toyobo). A twentieth of the reaction was used for PCR amplification using the following parameters: 94 °C for 30 s, 58 °C for 1 min, and 72 °C for 1 min. Cycle numbers used for PCR analyses were as follows: 16 for δ and 12 for both the β and α cDNAs in spleen; 18 and 12 for the γ and β/α cDNAs, respectively, in the fetal liver; and 18 and 12 for the ϵ and γ/α cDNAs, respectively, in the yolk sac. An aliquot of each PCR reaction was electrophoresed on 8% polyacrylamide gels, dried, and subjected to x-ray autoradiography and quantitative analysis with PhosphorImager (GE Healthcare) and ImageQuant software. The PCR primers for amplifying the δ-globin cDNA were: 5′-CTCATGGCAAGAAGGTGCTA-3′ and 5′-TTCTGATAGGCAGCCTGCAT-3′. The sequences of other primers used have been described elsewhere (28).
DNaseI Hypersensitive (HS) Site Mapping
DNaseI HS site mapping was essentially carried out as described elsewhere with minor modifications (27). Blood cells were collected from embryos and yolk sacs (10.5 days postcoitum) or phenylhydrazine-induced anemic adult spleens (1–2-month-old). Nuclei were prepared and digested with increasing concentrations of DNaseI (either 0, 2.5, 10, 20, 40, 80 units/ml final concentration) at 37 °C for 5 min. Purified DNA was cut to completion with restriction enzymes and subjected to gel electrophoresis and Southern blotting. For mapping DNaseI HS sites 2–4 and 1–2, probes corresponding to the 3′ region of HS4 (XbaI-HindIII, nt 2352–3266) and the 3′ region of HS1 (HindIII-XbaI, nt 13769–14235) were used, respectively.
RESULTS
Identification of the Core Sequences of the 5′HS1 Enhancer
We have previously reported an enhancer activity corresponding to the human β-globin LCR-5′HS1 (formerly termed HS1–3′ enhancer), which is located between −6.2 and −5.2 kb 5′ to the ϵ-globin gene transcriptional start site (Fig. 1A) (24). To test for promoter specificity of this enhancer, we linked the 5′HS1 fragment to the ϵ-globin, β-globin, and SV40 promoters and measured its activity by transient transfection into K562 cells (representing primitive erythroid cells; Fig. 1B). The fragment activated all of the promoters to a similar extent, suggesting that the 5′HS1 enhancer does not possess intrinsic promoter preferences. Curiously, however, the fragment failed to activate the same three promoters in MEL (definitive erythrocytes), HepG2, or HeLa cells (data not shown), suggesting that the 5′HS1 enhancer possesses cell type and developmental stage specificity. This activity was not caused by any theoretical cryptic promoter activity in 5′HS1 because the inverted fragment also exhibited significant enhancer activity (p5′HS1(L-)/ϵ-luc; Fig. 1C).
To identify the core sequences of the enhancer, we generated 3′- and 5′-deletions of the 5′HS1 enhancer fragment and tested their activities after transfection into K562 cells (Fig. 1C). Although the 5′HS1 fragment appeared to bear both activator and repressor activities, a single fragment “G” exhibited the strongest enhancer activity and was functional in both orientations (supplemental Fig. S1). We then divided the G fragment into four pieces (J-M) and tested their individual activities (Fig. 1D). Among these, only one of the fragments, L, exhibited any enhancer activity above background, while combining the K and L fragments (corresponding to fragment “N”) restored full enhancer activity as in fragment G.
Identification of Transcription Factor Binding Sites in the 5′HS1 Enhancer
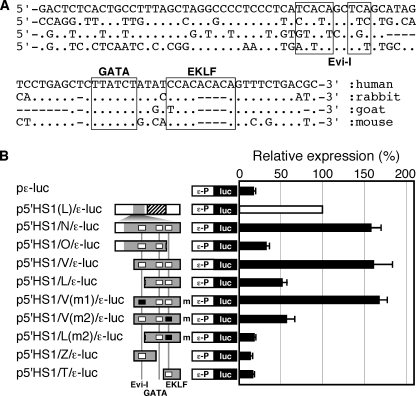
We next searched for putative transcription factor binding motifs in the N fragment (Fig. 2A). Although putative Evi-I, GATA, and EKLF binding motifs could be identified in the human sequence, only the GATA sites were conserved between rabbit, goat, and mouse. Based on this sequence information, we introduced smaller deletion and/or point mutations into the “N” fragment and tested their activities by transfection of the mutants into K562 cells (Fig. 2B). Whereas deletion of the 5′-part of the fragment significantly attenuated the enhancer activity (“L”), a point mutation in the 5′ Evi-I binding motif (“V(m1)”) had no effect, suggesting that other transcription factors were responsible for the enhancer activity. Deletion of part of the 3′-end of the fragment (“O”) or introduction of a point mutation into the 3′ EKLF binding motif (“V(m2)”) severely attenuated enhancer activity. Therefore, both ends of the “V” fragment appear to play significant quantitative roles in 5′HS1 enhancer activity. However, the 5′ (“Z”), 3′ (“T”), or middle (“L(m2)”) fragments individually displayed no enhancer activity, suggesting either that these elements are functionally interdependent or that one large enhancer element spans multiple segments of the V region of the 5′HS1 fragment.
FIGURE 2.
Identification of transcription factor binding sites in the 5′HS1 enhancer. A, cross-species comparison between the “N” enhancer fragment sequences. Putative transcription factor binding motifs found in the human sequences are boxed. (.) : identical nucleotides; (−): missing nucleotides. B, deletions and/or point mutations (shown in Fig. 3A) were introduced into the HS1/N fragment to systematically disrupt putative transcription factor binding motifs; their effects on enhancer activity were tested in K562 cells as described in the legend to Fig. 1, C and D. Open boxes, intact Evi-I, GATA, and EKLF motifs shown in A. Solid boxes (m), mutated factor binding motifs.
In Vitro and in Vivo Binding of GATA Factors to the Enhancer Core Element of 5′HS1
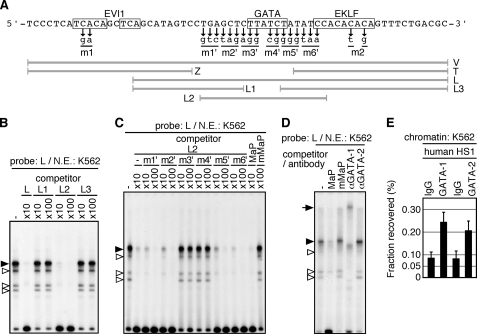
Because we failed to detect factor binding to either fragments Z or T in initial gel-shift assays (data not shown), we focused on identifying factor binding to L (Fig. 3A), which displays moderate enhancer activity alone (Figs. 1D and 2B). In the gel-shift assays employing K562 nuclear extracts, four shifted bands were observed with probe L, all of which were competed efficiently with only the middle part of the unlabeled L fragment, “L2” (Fig. 3B). We then introduced clustered two or three nucleotide mutations into the L2 fragment (m1′-m6′) and used them as competitors (Fig. 3, A and C), and found that mutations in the GATA motif (m3′ and m4′) disrupted its function as a competitor. In addition, the mouse α-globin promoter GATA site (MaP), but not its mutant version (mMaP), competed with the factor binding (Fig. 3C). We next conducted a super-shift assay with K562 nuclear extract and found that an anti-GATA-1, but not anti-GATA-2, antibody supershifted all the bands, demonstrating that these complexes with the L fragment include GATA-1 (Fig. 3D).
FIGURE 3.
In vitro and in vivo binding of GATA-1 to the 5′HS1 enhancer core sequences. A, sequence of the enhancer core element and portions of probe or competitor DNA fragments used in the EMSA (gray thick lines). Two- or three-base pair substitutions (m1, m2, and m1′–m6′) were introduced to mutate the factor binding motifs (boxed). B–D, EMSA. K562 nuclear extract was incubated with unlabeled 10- or 100-fold molar excess of competitor DNA fragments or specific antibodies, followed by the addition of radiolabeled L fragment probe; complex formation was analyzed by neutral polyacrylamide gel electrophoresis. Shifted bands are indicated by solid (strong binding) or open triangles. Mouse α-globin promoter sequences containing intact (MaP) or mutated (mMaP) GATA motifs were also included as competitors. E, GATA-1 and GATA-2 ChIP of the human 5′HS1 in K562 cells. Chromatin was immunoprecipitated with anti-GATA-1, anti-GATA-2 antibodies or control IgG. Immunoprecipitated DNA was quantified by real-time quantitative PCR (qPCR) using specific primers and normalized to input DNA. ChIP was repeated twice for each chromatin, and qPCR was repeated at least three times for each immunoprecipitated DNA. Data represent the averages with S.D.
We finally examined the in vivo binding of GATA factors to the human 5′HS1 region by ChIP assay and found that both GATA-1 and GATA-2 were recruited in this vicinity in K562 cells (Fig. 3E). However, because there is more than a single GATA motif in the 5′HS1 region that was amplified for the ChIP experiment, it is not known which site is in fact occupied by which GATA factors. Based on the gel-shift result in Fig. 3D, however, we surmise that the GATA-2 binds to sites in vivo that lie outside the L region.
The GATA Site Is Critical for 5′HS1 Enhancer Function
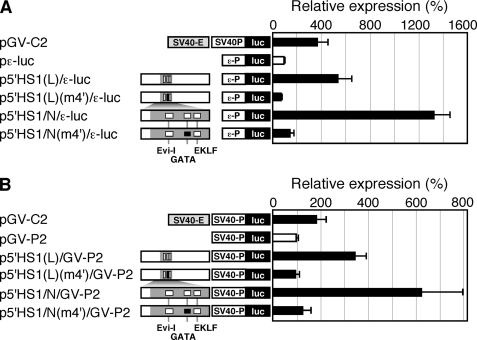
To test for effects of mutating the GATA motif on enhancer activity, we introduced a 2-bp mutation (m4′) into the N and 5′HS1(L) enhancer fragments and tested their transcriptional activity in K562 cells. As clearly shown in Fig. 4, A and B, the GATA site mutation completely abolished the enhancer activity of 5′HS1, demonstrating that the GATA site is indispensable for 5′HS1 enhancer activity in K562 cells.
FIGURE 4.
The GATA motif is responsible for the majority of 5′HS1 enhancer activity. The 2-bp GATA site mutation (m4′ in Fig. 3A) was introduced into the N, and 5′HS1(L) fragments. Wild-type and mutant enhancer fragments were linked to the ϵ-globin (A) or the SV40 (B) promoters, and their transcriptional activities were tested in K562 cells. Luciferase activities are expressed relative to those in the pϵ-luc or pGV-P2 (set at 100) in A and B, respectively. Solid box: mutated GATA motif.
Human β-Globin YAC TgM Bearing a 5′HS1 GATA Site Mutation
Because the data to this point suggested a vital role for the GATA site in 5′HS1 enhancer activity, we generated a 2-bp GATA site mutation (m4′) into a 150-kbp human β-globin YAC (Fig. 5A) to evaluate its in vivo function. We then generated 5 lines of intact, single copy YAC TgM (Fig. 5, B and C) and termed them mutGATA/5′HS1 TgM.
FIGURE 5.
Generation and structural analysis of mutGATA/5′HS1 TgM. A, structure of the 150-kb human β-globin YAC. The positions of the β-like globin genes (solid rectangles) are shown relative to the LCR. SfiI restriction enzyme sites are located 5′ to 5′HS5, between 5′HS4 and 5′HS3, and in the right arm of the YAC. Probes (gray rectangles) used for long-range (B) and end fragment (C) analyses, and expected restriction enzyme fragments with their sizes are shown. The GATA motif within the 5′HS1 region was mutated by substituting two nucleotides (underlined in the box) in the YAC. B, long-range structural analyses of the human β-globin YAC in TgM. The whole β-globin locus is contained within two SfiI fragments (8 and 100 kb, as above). DNAs from thymus cells were digested with SfiI in agarose plugs, separated by pulsed-field gel electrophoresis, and hybridized separately to the probes shown in A. C, tail DNA was digested with PstI, fractionated on agarose gels, and hybridized to L-end and R-end probes (shown in A).
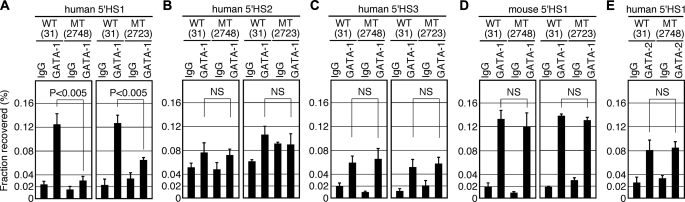
To examine the effects of GATA motif mutation in 5′HS1 on GATA factor binding in vivo, we first conducted a ChIP analysis using nucleated erythroid cells recovered from the spleens of adult TgM (Fig. 6). Three regions within the human LCR-5′HS1, HS2, and HS3, as well as the endogenous mouse 5′HS1 were analyzed by different PCR primer sets. ChIP analyses of two sets of TgM pairs (wild-type line 31 and mutant lines 2748 or 2723) revealed that human HS2 was not significantly occupied by GATA-1 in the WT or mutant TgM (Fig. 6B). Human HS3 (Fig. 6C) and mouse 5′HS1 (Fig. 6D) were both bound by GATA-1, regardless of whether they carry an intact or mutant 5′HS1 transgene. In striking contrast, GATA-1 was preferentially recruited to human 5′HS1 in the WT, but clearly not in the MT, TgM (Fig. 6A). The difference in enrichment between WT and MT alleles was 2–4-fold. Considering that there are other GATA motifs in the vicinity of human LCR-HS1 (not shown), this difference is quite significant. In contrast, GATA-2 recruitment to the human 5′HS1 did not differ significantly between WT and MT TgM (Fig. 6E), supporting our previous contention that GATA-2 binds to sites lying outside of the L region in 5′HS1. Thus, GATA-1 in fact binds to the human and mouse 5′HS1 in vivo and mutation of the GATA site severely attenuated GATA-1 binding to the human 5′HS1 in definitive erythroid progenitor cells.
FIGURE 6.
In vivo analysis of GATA factor recruitment to the LCR-5′HSs. 1–2-month-old animals carrying either WT or mutant (MT) YAC transgenes were made anemic by phenylhydrazine treatment, and spleens were collected. Chromatin was subjected to ChIP reactions using antibodies against GATA-1, GATA-2, or control IgG. Immunoprecipitated DNA was quantified by real-time qPCR using specific primers recognizing the human (A–C and E) or mouse (D) HS sites, and normalized to input DNA. Sequences of primers are listed in supplemental Table S4. ChIP was repeated twice for each chromatin and qPCR was repeated at least three times for each immunoprecipitated DNA. Data represent the averages with S.D. TgM lines are shown in parentheses. NS, not significant.
ϵ-Globin Gene Expression Is Attenuated in mutGATA/5′HS1 TgM
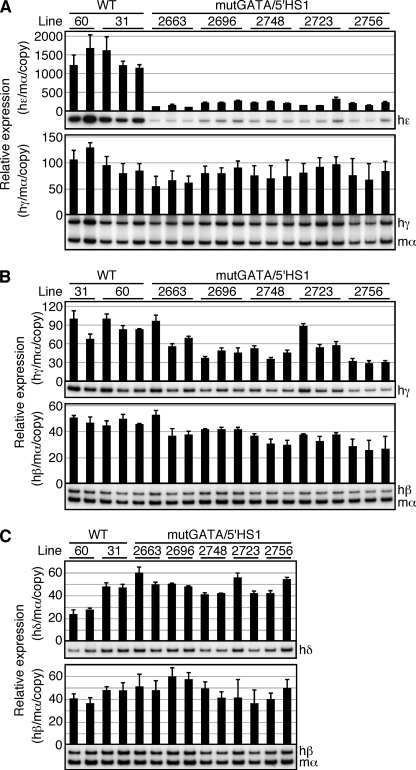
We next analyzed human ϵ- and γ-globin gene expression in primitive erythroid cells of the mutGATA/HS1 TgM (Fig. 7A). Total RNA was recovered from the yolk sacs (9.5 dpc) and expression of globin mRNAs was measured by semi-quantitative RT-PCR using endogenous mouse α-globin gene expression as an internal control. The data show that ϵ gene expression fell by 80–90% (Fig. 7A, top) in the mutant lines (mutGATA/5′HS1) in comparison to wild-type (WT; 29, 30) TgM. This phenotype was gene-specific because γ-globin gene expression was not significantly affected by the same mutation (Fig. 7A, bottom).
FIGURE 7.
Expression of the human β-like globin genes in the TgM. A, total RNA was prepared from the yolk sacs of more than two embryos (9.5 dpc) derived from the intercross of male transgenic and female non-Tg animals. Expression of human ϵ (hϵ)- and human γ (hγ)-globin compared with endogenous mouse α (mα)-globin genes was separately analyzed by semi-quantitative RT-PCR. The signals for hϵ-globin at 18 cycles and hγ/mα-globin at 12 cycles were quantified by phosphorimager, and the ratios of hϵ/mα (top) and hγ/mα (bottom) were calculated (the mα signal at 12 cycles was set at 100%). B, total RNA was prepared from the fetal livers of more than two fetuses (14.5 dpc). Expression of hγ- and human β (hβ)-globin compared with endogenous mα-globin genes was analyzed. The signals for hγ-globin at 18 cycles and hβ/mα-globin at 12 cycles were quantified, and the ratios of hγ/mα (top) and hβ/mα (bottom) were calculated (the mα signal at 12 cycles was set at 100%). C, total RNA was prepared from the anemic spleens of 1-month-old mice. Samples were collected from two individuals from each line of TgM. Expression of human δ (hδ) and hβ-globin compared with endogenous mα-globin genes was analyzed. The signals for hδ-globin at 16 cycles and hβ/mα-globin at 12 cycles were quantified, and the ratio of hδ/mα (top) and hβ/mα (bottom) were calculated (the mα signal at 12 cycles was set at 100%). The average ± S.D. from at least three independent experiments was calculated and graphically depicted. Representative results are shown below each panel.
We next collected fetal livers (14.5 dpc) from TgM and analyzed human γ- and β-globin gene expression in these definitive erythroid progenitor cells (Fig. 7B). Although RT-PCR analysis showed that both γ- and β-globin gene expression levels were slightly reduced in the mutant TgM, statistically significant differences were found only in a subset of the lines.
After 5 days of phenylhydrazine treatment, anemic spleens were collected from two animals representing each line for RNA extraction and RT-PCR analysis (Fig. 7C). No significant difference was observed in δ- or β-globin gene expression between the mutant and wild-type TgM.
DNaseI Hypersensitive Site Mapping of Wild-type and Mutant β-Globin LCRs
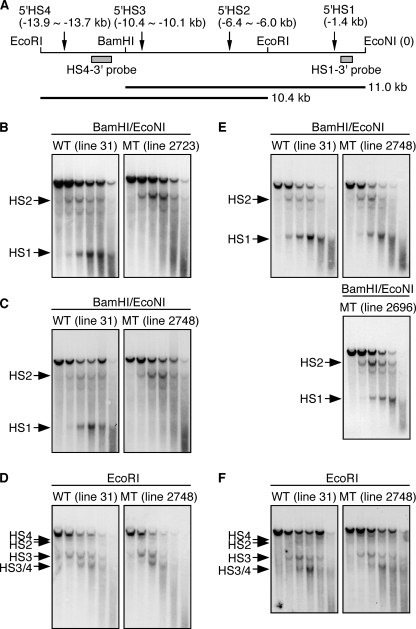
To examine possible chromatin effects of mutating the GATA motif in 5′HS1, we performed DNaseI HS site mapping experiments on definitive erythroid cells from wild-type (line 31) and mutant (lines 2723 and 2748) human β-globin YAC TgM lines (Fig. 8, B–D). Fig. 8A shows the structure of the human LCR and the experimental strategy we used to detect the formation of 5′HS4–2 and 5′HS1, using the probes “HS4–3′” and “HS1–3′”, respectively. We first tested if the GATA motif mutation disrupted the formation of 5′HS1 in the definitive erythroid chromatin of the YAC TgM. As shown in the data presented in Fig. 8, B and C, 5′HS1 disappeared in the TgM lines harboring the 2-bp GATA motif mutation, whereas it was easily detectable in the LCR of wild-type TgM.
FIGURE 8.
DNaseI hypersensitive site mapping of the human LCR chromatin in wild-type and mutant YAC transgenic mice. A, schematic representation of the human LCR. Fragments generated by digestion with BamHI/EcoNI (11.0 kb) or EcoRI (10.4 kb) are shown as horizontal thick lines. The location of the core elements of 5′HS1 (located at −1.4 kb relative to the EcoNI site at the 3′-end of the fragment), 5′HS2 (between −6.4 and −6.0 kb), 5′HS3 (−10.4 and −10.1 kb), and 5′HS4 (−13.9 and −13.7 kb) and the probes used for detecting the fragments (HS4–3′ and HS1–3′) are indicated by vertical arrows or gray boxes, respectively. Nuclei were isolated from blood cells of anemic spleens (B–D) or 10.5 dpc embryos (E and F) of TgM (line numbers in parentheses) carrying either WT or mutant (MT) human β-globin YACs, and treated with increasing concentrations of DNaseI. DNA was then isolated and digested with BamHI/EcoNI (B, C, and E) or EcoRI (D and F). After agarose gel electrophoresis, the DNA was transferred to nylon membranes and hybridized to either HS1–3′ (B, C, and E) or HS4–3′ (D) probes. The positions of the HS sites formed are indicated by arrows.
We have previously generated a TgM carrying human β-globin YAC with the 5′HS2 to 4 sequences all replaced with the polyoma virus enhancer sequences. In this mutant mouse, 5′HS1 and 5′HS5 formation, in addition to hypersensitive sites 2–4, were lost, suggesting their hierarchically lower positions in the process of HS site formation (27). We therefore analyzed the formation of 5′HS2 to 4 in the definitive erythroid cells of mutGATA/5′HS1 TgM. As expected (Fig. 8D), 5′HS2, 5′HS3 and 5′HS4 were clearly formed in both wild-type and mutant YAC TgM. Thus, mutation of the GATA motif in the core enhancer of 5′HS1 caused specific loss of 5′HS1 formation in the LCR. The presence of the 5′HS2 to 4 in the mutGATA/5′HS1 TgM, together with the observation that β-globin gene expression in the definitive erythroid cells was not affected by the GATA motif mutation in the 5′HS1 (Fig. 7C), clearly demonstrated that 5′HS1 is dispensable for LCR function at this developmental stage.
Next, we examined the consequences of mutating the 5′HS1 GATA motif in primitive erythroid cells (E10.5). 5′HS1 formation was only moderately affected by the mutation (Fig. 8E), while those in the 5′HS2–4 regions were not significantly affected (Fig. 8F). Thus, the clear presence of 5′HS1 to 4, together with the observation that ϵ-globin gene expression was severely attenuated in the mutGATA/HS1 TgM (Fig. 7A), allows the conclusion that this specific GATA site function, rather than its contribution to hypersensitive site formation, is critically involved in ϵ-globin gene activation in primitive erythroid cells.
DISCUSSION
Previous studies by Fedosyuk et al. (31) reported a 16- and 5-fold reduction in the expression of ϵ-globin in the yolk sac and γ-globin in the fetal liver, respectively, in the 213-kb human β-globin YAC TgM when examining a 0.8 kb 5′HS1 deletion that includes the GATA site that was intensively analyzed in the present work. Although the phenotypes we reported here are quantitatively less severe, the data are qualitatively consistent. Therefore, we conclude that a role for the GATA site examined in this study dominates 5′HS1 enhancer activity. Thus according to the current results, the activity of human 5′HS1 is predicted to be gene- and developmental stage-specific.
In contrast, however, Bender et al. (13) tested the effects of deleting 2.3 kb surrounding the 5′HS1 region from the endogenous mouse β-globin locus and found only moderate reduction (by up to 24%) of all the β-like globin genes, demonstrating that the activity of 5′HS1 exhibits neither gene- nor developmental stage-specificity. Although somewhat distinct phenotypes among these reports may be simply attributable to different experimental design, such as size of the deleted region, we propose that some species-specific differences in the context of the 5′HS1 enhancer may additionally be part of the basis for the observed discrepancy. There are several other (than GATA-1) transcription factor binding motifs in human 5′HS1 (Fig. 2A and Ref. 32). In the cell transfection assay, we showed that these cis sequences, including EKLF/Sp1 binding motifs, contributed to the 5′HS1 enhancer activity (Fig. 2B) and importantly, introduction of a 2 bp GATA motif mutation in the 5′HS1 fragment completely disrupted its enhancer function, although the other cis sequences were intact (Fig. 4). Both GATA and CAC motifs are frequently found in close proximity to, or within, enhancers and promoters of numerous erythroid specific genes and appear to synergistically cooperate in directing their expression (33–35). Perhaps curiously, however, EKLF/Sp1 binding motifs were not found within the homologous region in mouse 5′HS1 (Fig. 2A). We therefore assume that, in human 5′HS1, multiple elements cooperate synergistically to generate a single powerful enhancer activity, in which the GATA site plays a dominant role, and that this activity locally and specifically potentiates proximal ϵ-globin gene transcription. In the mouse, the GATA site in 5′HS1 moderately and only additively contributes to LCR enhancer activity that broadly potentiates all the β-like globin genes in the locus.
In proper development of the primitive as well as the definitive erythroid cell lineages, both GATA-1 and GATA-2 play important roles (36, 37). In the mouse, abundant GATA-1 (but little GATA-2) (38, 39) is expressed in mature erythrocytes, while GATA-2 plays a role in the earliest phases of erythroid development (37). We therefore attempted to clarify which factor contributed the 5′HS1 enhancer activity in vivo. In K562 cell chromatin, both GATA-1 and GATA-2 were recruited to the 5′HS1 region, which carries several GATA motifs (Fig. 3E and data not shown). In the gel-shift assay, however, only GATA-1 bound to the GATA sequence in the L enhancer fragment (Fig. 3D). In addition, we showed that recruitment of GATA-1, but not GATA-2, to the 5′HS1 region in vivo was significantly affected by GATA motif mutation in mutGATA/5′HS1 TgM. We therefore assume that GATA-1, but not GATA-2, is responsible for the observed phenotypes in the mutGATA/5′HS1 TgM. Unfortunately, however, since our attempts to demonstrate differential GATA factor recruitment to the wild-type versus mutant human 5′HS1 by ChIP analyses of 10.5 dpc primitive erythroid cells was unsuccessful (data not shown), probably because of the quite limited number of primitive erythrocytes we could recover, it is formally possible that GATA-2 is involved in GATA site-mediated ϵ-globin gene activation in primitive erythroid cells.
Although 5′HS1 is apparently required, it appears not sufficient to confer high level ϵ-globin gene expression. We have previously generated transgenic mice carrying the human β-globin YAC with LCR-HS2, -3, and -4 all replaced with the polyoma virus enhancer (27). In definitive erythroid cells of the PyE-substituted mutant mouse, 5′HS1 was not formed in the absense of other three HSs even though the 5′HS1 sequence was intact. Furthermore, ϵ-globin gene expression was completely silent in the yolk sac of this mutant mouse. Therefore, at least in the transgenic β-globin locus, LCR-HS2–4 may be required for opening chromatin before 5′HS1 can activate ϵ-globin transcription as an enhancer in primitive erythroid cells. In this sense, 5′HS1 may serve as a local enhancer that preferentially stimulates ϵ-globin, rather than functioning as a part of an LCR holocomplex. We have previously reported that ϵ-globin gene expression in primitive erythroid cells of YAC TgM is sensitive to distance from the LCR enhancer (24). Now, we showed here that efficient ϵ-globin gene transcription is critically dependent on 5′HS1 enhancer activity. Collectively, it is possible that ϵ-globin gene transcription is sensitive to distance from the 5′HS1 GATA site.
In summary, we found that the GATA site in the human LCR-HS1 region is required for conferring DNaseI hypersensitivity on 5′HS1, but that the same site is dispensable for adult β-globin gene expression in definitive erythroid cells. In contrast, the 5′HS1-GATA site is critically required for efficient expression of the ϵ-globin gene, while it is dispensable for formation of 5′HS1 in primitive erythroid cells. Therefore, the GATA site, rather than DNase I hypersensitivity within the 5′HS1 region, is involved in the developmental stage-specific gene activation mechanism in the human β-globin gene locus.
Supplementary Material
Acknowledgment
We thank Y. Tanimoto for technical assistance in generating TgM.
This work was supported in part by research grants from the Ministry of Education, Culture, Sports, Science, and Technology of Japan, Grant-in-Aid for Scientific Research (A) and for Young Scientists (S).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Fig. S1 and Tables S1–S4.
- LCR
- locus control region
- YAC
- yeast artificial chromosome
- TgM
- transgenic mice
- ChIP
- chromatin immunoprecipitation assay
- HS
- hypersensitive
- nt
- nucleotide
- EMSA
- electrophoretic mobility shift assay
- WT
- wild type
- qPCR
- real-time quantitative PCR.
REFERENCES
- 1.Stamatoyannopoulos G., Neinhuis A. W. (1994) in The Molecular Basis of Blood Diseases (Stamatoyannopoulos G., Neinhuis A. W., Majerus P., Varmus H. ed), pp. 107–155, 2 Ed., W. B. Saunders Co., New York [Google Scholar]
- 2.Forrester W. C., Thompson C., Elder J. T., Groudine M. (1986) Proc. Natl. Acad. Sci. U.S.A. 83, 1359–1363 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Grosveld F., Greaves D., Philipsen S., Talbot D., Pruzina S., deBoer E., Hanscombe O., Belhumeur P., Hurst J., Fraser P., et al. (1990) Ann. N.Y. Acad. Sci. 612, 152–159 [DOI] [PubMed] [Google Scholar]
- 4.Tuan D., London I. M. (1984) Proc. Natl. Acad. Sci. U.S.A. 81, 2718–2722 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Strouboulis J., Dillon N., Grosveld F. (1992) Genes Dev. 6, 1857–1864 [DOI] [PubMed] [Google Scholar]
- 6.Gaensler K. M., Kitamura M., Kan Y. W. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 11381–11385 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Peterson K. R., Clegg C. H., Huxley C., Josephson B. M., Haugen H. S., Furukawa T., Stamatoyannopoulos G. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 7593–7597 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Tuan D. Y., Solomon W. B., London I. M., Lee D. P. (1989) Proc. Natl. Acad. Sci. U.S.A. 86, 2554–2558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Moon A. M., Ley T. J. (1991) Blood 77, 2272–2284 [PubMed] [Google Scholar]
- 10.Collis P., Antoniou M., Grosveld F. (1990) EMBO J. 9, 233–240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fraser P., Hurst J., Collis P., Grosveld F. (1990) Nucleic Acids Res. 18, 3503–3508 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bender M. A., Mehaffey M. G., Telling A., Hug B., Ley T. J., Groudine M., Fiering S. (2000) Blood 95, 3600–3604 [PubMed] [Google Scholar]
- 13.Bender M. A., Roach J. N., Halow J., Close J., Alami R., Bouhassira E. E., Groudine M., Fiering S. N. (2001) Blood 98, 2022–2027 [DOI] [PubMed] [Google Scholar]
- 14.Fraser P., Pruzina S., Antoniou M., Grosveld F. (1993) Genes Dev. 7, 106–113 [DOI] [PubMed] [Google Scholar]
- 15.Engel J. D. (1993) Trends Genet. 9, 304–309 [DOI] [PubMed] [Google Scholar]
- 16.Palstra R. J., Tolhuis B., Splinter E., Nijmeijer R., Grosveld F., de Laat W. (2003) Nat. Genet. 35, 190–194 [DOI] [PubMed] [Google Scholar]
- 17.Milot E., Strouboulis J., Trimborn T., Wijgerde M., de Boer E., Langeveld A., Tan-Un K., Vergeer W., Yannoutsos N., Grosveld F., Fraser P. (1996) Cell 87, 105–114 [DOI] [PubMed] [Google Scholar]
- 18.Bungert J., Davé U., Lim K. C., Lieuw K. H., Shavit J. A., Liu Q., Engel J. D. (1995) Genes Dev. 9, 3083–3096 [DOI] [PubMed] [Google Scholar]
- 19.Bungert J., Tanimoto K., Patel S., Liu Q., Fear M., Engel J. D. (1999) Mol. Cell Biol. 19, 3062–3072 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fang X., Sun J., Xiang P., Yu M., Navas P. A., Peterson K. R., Stamatoyannopoulos G., Li Q. (2005) Mol. Cell Biol. 25, 7033–7041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Emery D. W., Chen H., Li Q., Stamatoyannopoulos G. (1998) Blood Cells Mol. Dis. 24, 322–339 [DOI] [PubMed] [Google Scholar]
- 22.Driscoll M. C., Dobkin C. S., Alter B. P. (1989) Proc. Natl. Acad. Sci. U.S.A. 86, 7470–7474 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kulozik A. E., Bail S., Bellan-Koch A., Bartram C. R., Kohne E., Kleihauer E. (1991) J. Clin. Invest. 87, 2142–2146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Shimotsuma M., Matsuzaki H., Tanabe O., Campbell A. D., Engel J. D., Fukamizu A., Tanimoto K. (2007) Mol. Cell Biol. 27, 5664–5672 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hou C., Zhao H., Tanimoto K., Dean A. (2008) Proc. Natl. Acad. Sci. U.S.A. 105, 20398–20403 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gaensler K. M., Burmeister M., Brownstein B. H., Taillon-Miller P., Myers R. M. (1991) Genomics 10, 976–984 [DOI] [PubMed] [Google Scholar]
- 27.Tanimoto K., Liu Q., Bungert J., Engel J. D. (1999) Nucleic Acids Res. 27, 3130–3137 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tanimoto K., Liu Q., Grosveld F., Bungert J., Engel J. D. (2000) Genes Dev. 14, 2778–2794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tanimoto K., Liu Q., Bungert J., Engel J. D. (1999) Nature 398, 344–348 [DOI] [PubMed] [Google Scholar]
- 30.Tanimoto K., Sugiura A., Omori A., Felsenfeld G., Engel J. D., Fukamizu A. (2003) Mol. Cell Biol. 23, 8946–8952 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fedosyuk H., Peterson K. R. (2007) Blood Cells Mol. Dis. 39, 44–55 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hardison R., Slightom J. L., Gumucio D. L., Goodman M., Stojanovic N., Miller W. (1997) Gene 205, 73–94 [DOI] [PubMed] [Google Scholar]
- 33.Gregory R. C., Taxman D. J., Seshasayee D., Kensinger M. H., Bieker J. J., Wojchowski D. M. (1996) Blood 87, 1793–1801 [PubMed] [Google Scholar]
- 34.Ren S., Li J., Atweh G. F. (1996) Nucleic Acids Res. 24, 342–347 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Merika M., Orkin S. H. (1995) Mol. Cell Biol. 15, 2437–2447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Pevny L., Simon M. C., Robertson E., Klein W. H., Tsai S. F., D'Agati V., Orkin S. H., Costantini F. (1991) Nature 349, 257–260 [DOI] [PubMed] [Google Scholar]
- 37.Tsai F. Y., Keller G., Kuo F. C., Weiss M., Chen J., Rosenblatt M., Alt F. W., Orkin S. H. (1994) Nature 371, 221–226 [DOI] [PubMed] [Google Scholar]
- 38.Leonard M., Brice M., Engel J. D., Papayannopoulou T. (1993) Blood 82, 1071–1079 [PubMed] [Google Scholar]
- 39.Minegishi N., Ohta J., Suwabe N., Nakauchi H., Ishihara H., Hayashi N., Yamamoto M. (1998) J. Biol. Chem. 273, 3625–3634 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.