Abstract
In preadipocytes, retinoic acid (RA) regulates gene expression by activating the nuclear RA receptor (RAR) and its cognate intracellular lipid-binding protein CRABP-II. It was previously reported that RA inhibits adipocyte differentiation but only when administered early during the differentiation program. The data presented here indicate that the diminished ability of RA to activate RAR following induction of differentiation stems from down-regulation of CRABP-II. The observations show that expression of CRABP-II in preadipocytes is repressed by all three components of the classical hormonal mixture that induces adipocyte differentiation, i.e. isobutylmethylxanthine, insulin, and dexamethasone. Isobutylmethylxanthine-dependent activation of protein kinase A triggered the phosphorylation of the transcription factor cAMP-response element-binding protein, which induced the expression of the cAMP-response element-binding protein family repressor cAMP-response element modulator. In turn, cAMP-response element modulator was found to associate with a cognate response element in the CRABP-II promoter and to repress CRABP-II expression. The data further show that CRABP-II is a direct target gene for the glucocorticoid receptor and that it is subjected to dexamethasone-induced glucocorticoid receptor-mediated repression during adipogenesis. Finally, the observations demonstrate that permanent repression of CRABP-II in mature adipocytes is exerted by the master regulator of adipocyte differentiation CCAAT/enhancer-binding protein α and is directly mediated through CCAAT/enhancer-binding protein α-response elements in the CRABP-II promoter. Taken together, the observations emphasize the important role of CRABP-II in regulating the transcriptional activity of RA through RAR, and they demonstrate that repression of this gene is critical for allowing adipogenesis to proceed.
Keywords: Adipocyte, Adipose Tissue, Lipid-binding Protein, Nuclear Receptors, Retinoid, Retinoid-binding Protein, Transcription Factors, Transcription Regulation
Introduction
Adipogenesis entails a timed progression of tightly controlled changes in gene expression, culminating in terminal differentiation. Studies using cultured preadipocytes demonstrated that adipocyte differentiation, which begins with cell-cell contact inhibition, can be induced by a hormonal mixture containing insulin, isobutylmethylxanthine (IBMX),3 and a glucocorticoid such as dexamethasone (1, 2). This differentiation mixture triggers an initial growth arrest followed by one or two cell divisions known as clonal expansion. Cells then exit the cell cycle and proceed to terminal differentiation driven by the “master regulators” of adipocyte differentiation, the transcription factors peroxisome proliferator-activated receptor γ (PPARγ) and CCAAT/enhancer-binding protein α (C/EBPα). Mature adipocytes are characterized by their ability to accumulate lipids and by the appearance of adipocyte markers such as the fatty acid-binding protein FABP4 and the lipid transporter CD36 (3, 4).
The individual components of the mixture that trigger differentiation facilitate different aspects of adipogenesis. IBMX, a phosphodiesterase inhibitor, elevates cellular cAMP levels leading to activation of protein kinase A (PKA). One of the PKA substrates in adipocyte is the transcription factor cAMP-response element-binding protein (CREB). Activated CREB positively regulates the expression of multiple genes, including the transcription factor C/EBPβ, which promotes cell cycle arrest and clonal expansion (3, 5–7). CREB also induces the expression of another CREB family member, the cAMP-response element modulator gene (CREM). In contrast with CREB, CREM lacks the ability to recruit coactivators, and it functions as a cAMP-responsive negative regulator of gene expression (8–10).
Glucocorticoids are involved in adipocyte differentiation by activating glucocorticoid receptor (GR) whose targets include C/EBPδ, a transcription factor that induces the expression of PPARγ (11, 12). GR also represses the expression of the negative regulator of adipocyte differentiation Pref-1 (13). The downstream effectors that mediate the involvement of insulin in adipocyte differentiation are poorly understood, and the effects may stem from cross-activation of the insulin-like growth factor-1 receptor (3).
Studies using the well established adipogenesis model NIH3T3-L1 cells revealed that the vitamin A metabolite retinoic acid (RA) effectively inhibits adipocyte differentiation (14). The biological activities of RA are mediated by two nuclear receptors, RAR (15) and PPARβ/δ (17–19), and by two intracellular lipid-binding proteins (iLBP) that selectively deliver RA from its sites of synthesis in the cytosol to specific RA receptors in the nucleus. It has been demonstrated in regard to this that RA is delivered to RAR by the iLBP cellular retinoic acid-binding protein II (CRABP-II) and that it is transported to PPARβ/δ by the iLBP termed fatty acid-binding protein 5 (FABP5) (18, 20–24). Hence, the partitioning of RA between its two receptors and the biological activities of the hormone are determined by the relative expression levels of these iLBPs. RA signals through RAR in cells that highly express CRABP-II, and it activates PPARβ/δ in cells that display a high level of FABP5 (18–21, 25).
It was previously reported in preadipocytes that CRABP-II is highly expressed and consequently that RA functions through RAR in these cells. Interestingly, it was shown that adipocyte differentiation is accompanied by down-regulation of CRABP-II and RAR and up-regulation of FABP5, resulting in a marked shift of RA signaling toward the FABP5/PPARβ/δ pathway (14, 25). Available information thus indicates that the ability of RA to inhibit adipogenesis is exerted by activities of the hormone that are mediated by CRABP-II and its cognate receptor RAR in preadipocytes and thus suggests that down-regulation of CRABP-II is a critical component of the differentiation process. The observations reported here support this notion, and they delineate the pathways by which the expression of CRABP-II is repressed during adipogenesis and is maintained at a low level in mature adipocytes.
EXPERIMENTAL PROCEDURES
Reagents
Antibodies against CREM, phospho-CREB, and GR were from Santa Cruz Biotechnology. C/EBPα and CREB antibodies were from Cell Signaling and Millipore, respectively. Antibodies against CRABP-II were generously provided by Cecile Rochette-Egly (Institut Génétique Biologie Moléculaire Cellulaire). Adenovirus expressing CRABP-II was generated at the Gene Transfer Vector Core, University of Iowa. Wortmannin and RA were purchased from Calbiochem. Actinomycin D, cycloheximide, forskolin, IBMX, insulin, dexamethasone, RU486, H89, and antibodies against β-tubulin were from Sigma. Lentiviruses harboring shRNAs were purchased from Open Biosystems. The triglyceride assay kit was obtained from Zen-Bio.
Cells
3T3-L1s were obtained from ATCC and cultured in Dulbecco's modified Eagle's medium supplemented with 10% calf serum. Cells were grown 2 days postconfluence and then induced to differentiate with Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum, 10 μg/ml insulin, 0.5 mm IBMX, and 0.25 mm dexamethasone. Adipocyte differentiation was confirmed by Oil Red O staining and by immunostaining for the presence of FABP4.
Chromatin Immunoprecipitation (ChIP) Assays
Preadipocytes were treated with IBMX (0.5 mm, 4 h) or dexamethasone (250 nm, 2 h) prior to cross-linking with formaldehyde. 1.5 m glycine was added, and lysates were sonicated. Appropriate antibodies were added, and proteins were immunoprecipitated (12 h) and attached to protein A/G beads (2 h). Cross-link was reversed and protein degraded using protease K. DNA was precipitated using ethanol. PCR was performed using the following primer sets: CRE response element, sense 5′-CGTTGTGCCTACCTA-3′ and antisense 5′-CAGAGCGATTTTAGGGCTTG-3′; GR half-site response element, sense 5′-AGCTGTCCAGAGCACCAAAT-3′ and antisense 5′-GAGAAGCGTCAGAAGGCAAG-3′; C/EBPα-response element, sense 5′-GAGCATCAGGCTCCTGTTTT-3′ and antisense 5′-CTCCGAGGCTGATTTTTACG-3′.
Quantitative Real Time PCR (Q-PCR)
RNA was extracted using TRIzol (Molecular Research Center). cDNA was generated using GeneAmp RNA PCR (Applied Biosystems). Q-PCR was carried out using TaqMan chemistry and Assays-on-Demand probes (Applied Biosystems) for CRABP-II Mm00801691_m1, CREM Mm00516346_m1, and CREB Mm00501607_m1.
Isolation of Human Preadipocytes
Adipose tissue was collected from patients undergoing abdominoplasty, and primary preadipocytes were isolated as described previously (26). Briefly, 20 g of tissue was cut into fine pieces, placed in 10 ml of phosphate-buffered saline containing collagenase (10 mg) and 1% bovine serum albumin, and incubated at 37 °C for 90 min. Material was centrifuged (500 × g, 10 min, 4 °C). The pellet was suspended in erythrocyte buffer (1.54 mm NH4Cl, 5.2 mm K2HPO4, 0.1 mm EDTA) and placed on ice for 10 min. Cells were centrifuged (500 × g, 10 min, 4 °C), resuspended in Dulbecco's modified Eagle's/F-12 media containing 10% calf serum, and cultured (37 °C, 5% CO2). Cells were recultured three times prior to experimentation.
RESULTS
CRABP-II Enhances the Ability of RA to Inhibit Adipogenesis
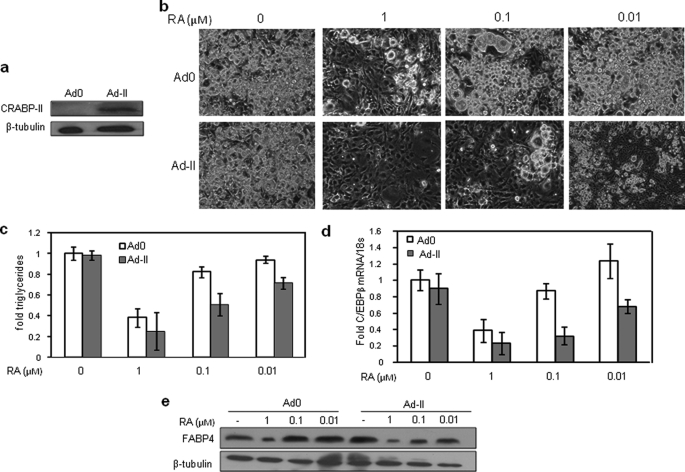
It has been reported that RA inhibits the differentiation of NIH3T3-L1 cells, a well established cultured cell model of adipogenesis (14). In this model, preadipose cells are grown 2 days post-confluence and induced to differentiate using a hormone mixture containing insulin (10 μg/ml), IBMX (0.5 mm), and dexamethasone (0.25 mm). Following 3 days of treatment, media were replaced, and cells were grown for 4 days to obtain mature adipocytes. To examine the involvement of CRABP-II in RA-induced inhibition of adipogenesis, NIH3T3-L1 cells were grown to ∼80% confluence and then infected with an adenovirus harboring cDNA for CRABP-II (Ad-II, Fig. 1a). Cells were induced to differentiate in the presence of varying concentrations of RA, and adipogenesis was examined by monitoring lipid accumulation and by examining the expression of the early and late adipocyte markers C/EBPβ and FABP4, respectively. RA had little effect at low concentrations. However, treatment with 1 μm of the hormone markedly inhibited adipogenesis as reflected by decreased lipid accumulation (Fig. 1, b and c) and lower expression of the adipocyte markers C/EBPβ (Fig. 1d) and FABP4 (Fig. 1e). Strikingly, in cells that ectopically overexpressed CRABP-II, RA inhibited differentiation at concentrations as low as 10 nm (Fig. 1, b–e), demonstrating that the protein markedly enhances the ability of RA to inhibit adipogenesis.
FIGURE 1.
CRABP-II sensitizes NIH3T3-L1 cells to RA-induced inhibition of adipogenesis. a, NIH3T3-L1 preadipocytes were infected with an empty adenovirus (Ad0) or adenovirus harboring CRABP-II cDNA (Ad-II) at 80% confluence. Two days later, cells were induced to differentiate. CRABP-II level in resulting mature adipocytes was assessed by immunoblotting. b, preadipocytes were infected with Ad0 or Ad-II as described in a and induced to differentiate in the absence or presence of denoted concentrations of RA. Micrographs show lipid accumulation at the end of differentiation. c, cells were treated as described in b, and triglyceride content at the end of differentiation was measured (Zen-Bio, triglyceride assay kit). d, cells were treated as described in b, and levels of C/EBP-β mRNA were measured by Q-PCR for 24 h following induction. Error bars indicate S.D. e, cells were treated as described in b, and expression of FABP4 in mature adipocytes was measured by immunoblots.
Expression of CRABP-II Is Down-regulated upon Induction of Adipocyte Differentiation
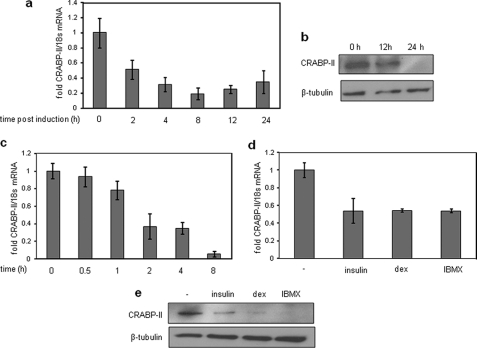
Examination of the expression of CRABP-II in NIH3T3-L1 cells showed that both CRABP-II mRNA (Fig. 2a) and protein (Fig. 2b) are rapidly down-regulated following induction of differentiation by the standard differentiation mixture consisting of insulin, dexamethasone, and IBMX. The rate of decrease of CRABP-II mRNA levels following treatment with the transcription inhibitor actinomycin D (Fig. 2c) was similar to that observed upon treatment with the differentiation mixture (Fig. 2a), suggesting that the expression of the gene during adipogenesis may be regulated at the level of transcription.
FIGURE 2.
Adipocyte differentiation is accompanied by down-regulation of CRABP-II expression. a, preadipocytes were induced to differentiate, and CRABP-II expression at denoted time points was analyzed by Q-PCR. b, immunoblots of CRABP-II expression in preadipocytes at the denoted time points following differentiation induction. c, preadipocytes were treated with actinomycin D (5 μg/ml) for denoted time points, and levels of CRABP-II mRNA were analyzed by Q-PCR. d and e, preadipocytes were treated with insulin (10 μg/ml), dexamethasone (dex) (0.25 mm), or IBMX (0.5 mm) for 4 h. Levels of CRABP-II mRNA were analyzed by Q-PCR (d) and CRABP-II protein levels assessed by immunoblots (e). Error bars indicate S.D.
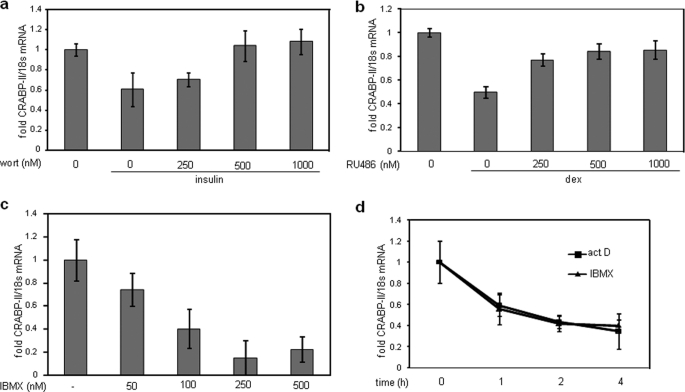
To examine which of the individual components of the differentiation mixture controls CRABP-II expression, cells were treated separately with insulin, dexamethasone, or IBMX, and the expression of CRABP-II mRNA (Fig. 2d) and protein (Fig. 2e) was measured. The data show that all three components down-regulated CRABP-II expression. Treatment with wortmannin, an inhibitor of phosphatidylinositol 3-kinase, abolished the ability of insulin to repress CRABP-II (Fig. 3a), and treatment with RU486, a glucocorticoid receptor antagonist, inhibited down-regulation of CRABP-II by dexamethasone (Fig. 3b). CRABP-II expression was down-regulated in a dose-dependent manner following a 4-h treatment of IBMX (Fig. 3c). The rate of decrease in CRABP-II mRNA following IBMX treatment was similar to that observed upon treatment with actinomycin D (Fig. 3d), suggesting that IBMX regulates the expression of CRABP-II by inhibiting its transcription.
FIGURE 3.
Insulin, dexamethasone, and IBMX down-regulate CRABP-II expression. a and b, preadipocytes were pretreated with denoted concentrations of wortmannin (wort) (a) or with RU486 (b) and then treated with 10 μg/ml insulin (a) or 250 nm dexamethasone (b). CRABP-II mRNA expression was analyzed by Q-PCR. c, preadipocytes were treated with denoted concentrations of IBMX, and CRABP-II mRNA expression was analyzed by Q-PCR. d, preadipocytes were treated with either actinomycin D (act D) (5 μg/ml) or IBMX (0.5 mm) for the denoted times, and CRABP-II mRNA expression was analyzed by Q-PCR. Error bars indicate S.D.
PKA Represses CRABP-II Expression by Activating a CREB/CREM Pathway
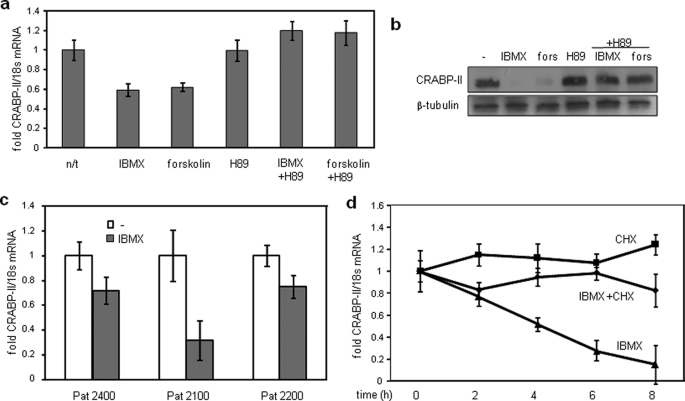
The pathways by which IBMX down-regulates CRABP-II were then investigated. Cells were treated with the adenylate cyclase activator forskolin, a treatment which, like IBMX, increases intracellular cAMP levels. Similarly to IBMX, forskolin decreased the levels of CRABP-II (Fig. 4, a and b). The PKA inhibitor H89 alleviated the repression induced by either IBMX or forskolin (Fig. 4, a and b). Hence, activation of PKA by cAMP is closely involved in regulating CRABP-II expression.
FIGURE 4.
Repression of CRABP-II by IBMX is mediated by PKA. a and b, preadipocytes were treated with IBMX (0.5 mm), forskolin (1 μm), or the PKA inhibitor H89 (1 μm) for 4 h or pretreated with H89 prior to treatment with IBMX or forskolin (fors). n/t, not treated. Levels of CRABP-II mRNA (a) and protein (b) were measured by Q-PCR and immunoblots, respectively. c, primary human preadipocytes isolated from three individual patients were treated with IBMX (0.5 mm, 4 h), and CRABP-II expression was analyzed by Q-PCR. d, preadipocytes were pretreated with cycloheximide (CHX) (10 μg/ml, 15 min) and then treated with IBMX (0.5 mm, 4 h). CRABP-II mRNA expression was analyzed by Q-PCR. Error bars indicate S.D.
The effect of PKA activation on CRABP-II expression was also examined using primary preadipocytes isolated from human adipose tissue. Tissue was dissociated in phosphate-buffered saline containing collagenase (1 mg/ml) and bovine serum albumin (1%). Cells were collected, resuspended in a buffer containing 1.54 mm NH4Cl, 5.2 mm K2HPO4, and 0.1 mm EDTA, centrifuged, suspended in Dulbecco's modified Eagle's/F-12 media containing 10% calf serum, and plated. Cells were recultured three times prior to experimentation. Similarly to the response observed in NIH3T3-L1 cells, treatment of primary human preadipocytes with IBMX decreased the level of expression of CRABP-II (Fig. 4c).
To determine whether PKA-mediated down-regulation of CRABP-II expression is direct or whether it reflects secondary responses to PKA activation, NIH3T3-L1 preadipocytes were pretreated with the protein synthesis inhibitor cycloheximide (10 μg/ml, 15 min) prior to treatment with IBMX. Cycloheximide treatment abolished the ability of IBMX to decrease CRABP-II expression levels (Fig. 4d), demonstrating that the effect required de novo protein synthesis.
A well established downstream effector of PKA is the transcription factor CREB. Activation of CREB by PKA-mediated phosphorylation allows this factor to bind to cAMP-response elements (CRE) in promoter regions of target genes and to positively regulate gene expression (3, 4, 8). Indeed, treatment of NIH3T3-L1 cells with IBMX enhanced the phosphorylation level of CREB (Fig. 5a). To examine a possible involvement of CREB in regulating CRABP-II expression, the expression of this factor in NIH3T3-L1 preadipocytes was decreased using lentiviruses harboring CREB shRNA (Fig. 5b). Cells were then treated with IBMX, and CRABP-II expression was measured. Decreased expression of CREB abrogated the ability of IBMX to down-regulate CRABP-II mRNA (Fig. 5c) and protein (Fig. 5d). In adipocytes, activated CREB is known to induce early responsive genes, such as CREM (27). Unlike CREB, which induces gene expression, CREM does not associate with coactivators. Instead, this factor recruits transcriptional corepressors, and upon binding to CRE, it represses the expression of its target genes (3, 4). Considering the observations that CREB negatively regulates CRABP-II expression (Fig. 5, c and d) and that the effect is indirect (Fig. 4d), possible involvement of CREM was examined. Induction of adipocyte differentiation was accompanied by a transient up-regulation of CREM (Fig. 5e), and interestingly, the maximal level of CREM expression at 4 h post-induction coincided with 50% reduction in CRABP-II mRNA (Fig. 5e). Decreasing the expression level of CREM using a lentivirus harboring CREM shRNA (Fig. 5g) hampered the ability of IBMX to suppress CRABP-II attesting to the involvement of the factor in mediating the repression of CRABP-II (Fig. 5h).
FIGURE 5.
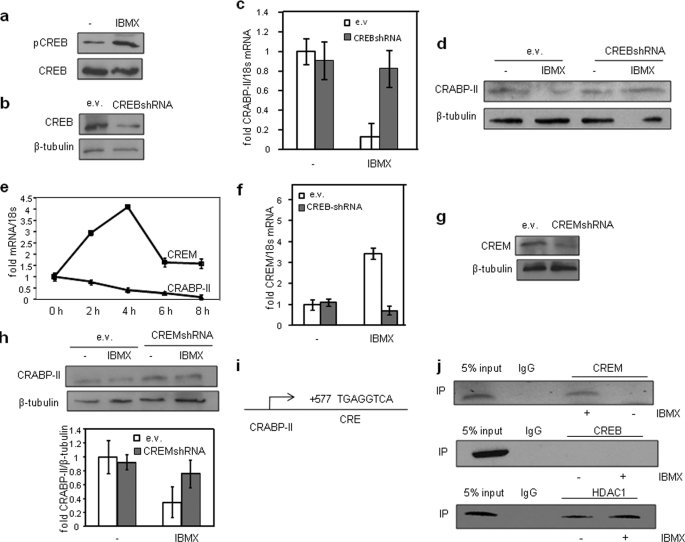
Repression of CRABP-II by PKA is mediated by CREB and its target gene CREM. a, preadipocytes were treated with IBMX (0.5 mm) for 15 min, and phosphorylation of CREB was assessed by immunoblots. b, preadipocytes were infected with empty lentivirus (ev) or lentivirus harboring CREB shRNA. Decreased expression of CREB was assessed by immunoblots. c and d, preadipocytes infected with lentivirus as denoted were then treated with IBMX (0.5 mm) for 4 (c) or 12 h (d), and levels of CRABP-II mRNA (a) and protein (d) were assessed by Q-PCR and immunoblots, respectively. e, preadipocytes were treated with IBMX (0.5 mm) for the denoted times. Levels of mRNA for CREM and CRABP-II were measured by Q-PCR. f, preadipocytes were infected with denoted lentiviruses and treated with IBMX for 4 h. Levels of mRNA for CREM were measured by Q-PCR. g, immunoblots of CREM in preadipocytes were infected with empty lentivirus or with lentivirus harboring CREM shRNA. h, top, immunoblots of CRABP-II in preadipocytes were infected with denoted lentiviruses and treated with IBMX for 12 h. Bottom, quantification of immunoblots from three independent experiments. Data are mean ± S.E. i, location of the Cre element in the first intron of the CRABP-II gene. j, ChIP assays indicating recruitment of CREM and HDAC1 to the Cre element of the CRABP-II gene following treatment with IBMX (0.5 mm, 4 h.) (see under “Experimental Procedures” for assay details). IP, immunoprecipitation.
The Alibaba 2 program of the Transfac site was used to identify response elements that may mediate CREM-dependent repression of CRABP-II. Two putative CREs were found, a site at 649 bp upstream from the gene start site, and an additional element located in the first intron of CRABP-II (Fig. 5i). ChiP assays were carried out to examine whether either CREB or CREM is associated with these elements in NIH3T3-L1 cells. Neither factor associated with the upstream element either (data not shown). In contrast, IBMX treatment efficiently induced recruitment of CREM (but not CREB) to the CRE located in the first intron of CRABP-II (Fig. 5j). IBMX treatment also recruited the transcriptional corepressor HDAC1 to this element. (Fig. 5j). The data thus indicate that CREM and its associated corepressors directly repress CRABP-II expression.
GR Negatively Regulates CRABP-II Expression
In addition to the repression of CRABP-II by cAMP signaling, the expression of this gene in preadipocytes was also found to be down-regulated upon treatment with the GR agonist dexamethasone (Fig. 2, d and e). To examine whether CRABP-II is a direct target gene for GR, cells were pretreated with cycloheximide prior to treatment with dexamethasone. The observations that the ligand decreased CRABP-II expression in the presence of the protein synthesis inhibitor (Fig. 6a) suggest that CRABP-II is a direct target for GR-mediated repression. In accordance, down-regulation of GR expression using a lentivirus harboring GR shRNA (Fig. 6b) inhibited dexamethasone-induced down-regulation of CRABP-II (Fig. 6, c and d). Examination of the CRABP-II promoter revealed the presence of a putative GR-response element at 247 bp upstream from the gene start site (Fig. 6e). ChiP assays demonstrated that GR is recruited to this element following treatment with dexamethasone (Fig. 6f). The observations thus establish that GR directly suppresses CRABP-II during adipocyte differentiation.
FIGURE 6.
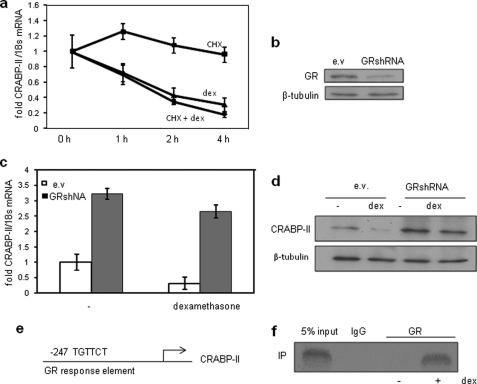
CRABP-II is directly repressed by GR. a, preadipocytes were pretreated with cycloheximide (CHX) (10 μg/ml, 15 min) and then treated with dexamethasone (dex) for 4 h. CRABP-II expression at the denoted time was assessed by Q-PCR. b, immunoblot of GR in preadipocytes infected with empty lentivirus (ev) or with lentivirus harboring GR shRNA. c and d, preadipocytes were infected with lentoviruses and treated with dexamethasone for 4 h (c) or 12 h (d). Expression levels of CRABP-II mRNA (c) and protein (d) were assessed by Q-PCR and immunoblots, respectively. e, GR half-site in the CRABP-II promoter. Error bars indicate S.D. f, ChIP assays indicating recruitment of GR to the GRE of the CRABP-II gene following treatment with dexamethasone (see “Experimental Procedures” for assay details). IP, immunoprecipitation.
In Mature Adipocytes, CRABP-II Expression Is Repressed by C/EBPα
Following its down-regulation during adipogenesis, CRABP-II expression remains low in mature adipocytes (25). These observations raise the question of the mechanisms that sustain CRABP-II repression following differentiation. The question is especially pertinent considering that although the major CRABP-II repressor CREM is induced upon triggering differentiation, the effect is transient, and CREM expression returns to a basal level as early as 8 h following induction (Fig. 5e). In accordance with these observations, ChIP assays indicated that although CREM was associated with CRE of the CRABP-II gene 4 h following IBMX treatment (Fig. 5j), the factor was not bound to the element in mature adipocytes (Fig. 7b).
FIGURE 7.
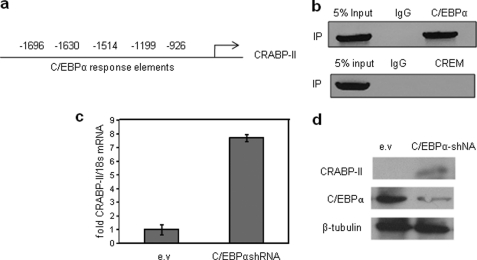
C/EBPα represses CRABP-II in mature adipocytes. a, five C/EBPα sites are located in the CRABP-II promoter. b, top, ChIP assays indicate recruitment of C/EBPα to the promoter region containing the putative C/EBPα sites in mature adipocytes. Bottom, ChIP assays indicating that in mature adipocytes CREM does not occupy the Cre in the CRABP-II promoter (compare with Fig. 5j). c and d, expression of C/EBPα in mature adipocytes was decreased using a lentivirus harboring C/EBPα shRNA. CRABP-II expression was analyzed by Q-PCR (b) or immunoblot (c). IP, immunoprecipitation. Error bars indicate S.D.
One possibility is that sustained repression of CRABP-II in mature adipocytes is exerted by C/EBPα, a master regulator of adipogenesis that is highly expressed in mature adipocytes and can function either as an activator or as a repressor of gene expression (3, 4, 28). Inspection of the CRABP-II promoter revealed that a region, located at about 1,000 bp upstream from the gene start site, contains five putative C/EBPα-response elements (Fig. 7a). ChiP assays indicated that C/EBPα is indeed associated with this region in mature adipocytes (Fig. 7b). The involvement of C/EBPα in regulating CRABP-II expression was further examined by down-regulating C/EBPα expression in mature adipocytes (Fig. 7d). Decreasing the expression level of C/EBPα resulted in marked up-regulation of CRABP-II mRNA (Fig. 7c) and protein (Fig. 7d). Taken together, these data demonstrate that the low level of expression of CRABP-II in mature adipocytes is maintained by C/EBPα.
DISCUSSION
RA regulates gene transcription by activating the nuclear receptor RAR and its cognate iLBP CRABP-II and by an alternative pathway mediated by PPARβ/δ and FABP5 (15–17, 20, 21, 24, 29–31). We previously reported that although RA predominantly signals through CRABP-II/RAR in preadipocytes, adipocyte differentiation is accompanied by a shift in the balance between the two pathways. This shift is accomplished by down-regulation of CRABP-II, RARα, and RARγ and by up-regulation of FABP5 during adipogenesis, enabling activation of the FABP5/PPARβ/δ pathway in the mature adipocyte (25). Consequently, in mature adipocytes, RA regulates lipid homeostasis and insulin responses by controlling the expression of target genes for both RAR and PPARβ/δ (25). The data presented here demonstrate that CRABP-II markedly sensitizes preadipocytes to RA-induced inhibition of differentiation (Fig. 1). The observations thus reveal that down-regulation of CRABP-II, which sets the stage for proper RA signaling in mature adipocytes, is also critical for bypassing RA-induced, RAR-mediated inhibition of differentiation and allowing adipogenesis to proceed. Interestingly, it was reported that RA inhibits adipocyte differentiation if administered within a short time frame following its induction but that it fails to do so when administered late in the program even in cells that ectopically overexpress RARs (32). The observations that suppression of CRABP-II is an early adipogenic event (Fig. 2a) emphasize the importance of CRABP-II in activation of RAR and provide a rationale for understanding the failure of RA to inhibit adipogenesis late in the program.
Expression of CRABP-II was found to be repressed by all three components of the classical adipocyte differentiation mixture (Fig. 8). Although the pathway(s) that mediate the effect of insulin remain to be explored, the observations delineate the mechanisms by which dexamethasone and IBMX control CRABP-II expression. The data demonstrate that treatment of preadipocytes with dexamethasone results in recruitment of GR to a GR element located 247 bp upstream from the CRABP-II gene start site (Fig. 6e) and that repression of CRABP-II by dexamethasone is a GR-mediated response that does not require de novo protein synthesis (Fig. 6, a–d). Hence, CRABP-II is a direct target gene for dexamethasone-induced, GR-mediated repression.
FIGURE 8.
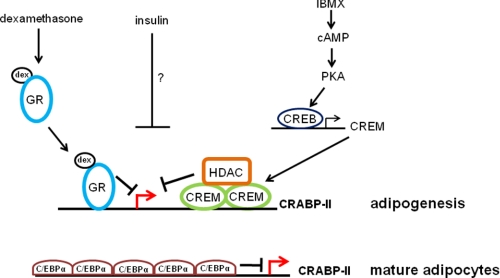
Model for the regulation of CRABP-II expression during adipogenesis and in mature adipocytes. The classical adipocyte differentiation inducers, insulin, IBMX, and dexamethasone (dex), all repress the expression of CRABP-II. IBMX increases cellular levels of cAMP leading to activation of PKA. Activated PKA phosphorylates CREB, which positively regulates the expression of its direct target gene CREM. In turn, CREM associates with a CRE in the first intron of CRABP-II and recruits HDAC1 to repress CRABP-II transcription. Dexamethasone activates GR. In turn, GR binds to a GRE in the CRABP-II promoter and negatively regulates CRABP-II expression. The details of the mechanism through which insulin represses CRABP-II expression remain to be explored. In mature adipocytes, low levels of expression of CRABP-II are maintained by direct repression by C/EBPα.
Treatment with IBMX or the adenylate cyclase activator forskolin decreased the expression of CRABP-II in NIH3T3-L1 cells as well as in primary human preadipocytes, and inhibition of PKA activity abolished the effect (Fig. 4, a–c). The data thus establish that cAMP and PKA signaling down-regulates the expression of CRABP-II. Suppression of CRABP-II by IBMX was indeed found to be mediated by the transcription factor CREB, a PKA substrate in preadipocytes (Fig. 5, b and c). However, the observations that down-regulation of CRABP-II by IBMX required de novo protein synthesis (Fig. 4d) and the known activity of CREB as a positive regulator of transcription (3) suggested that repression of CRABP-II by CREB is a secondary response. The data indicate that CRABP-II expression is controlled by CREB indirectly through induction of the direct CREB target gene, the transcriptional repressor CREM (Fig. 8). In support of this conclusion, IBMX-induced repression of CRABP-II was found to require the expression of CREM (Fig. 5h), and treatment with IBMX resulted in recruitment of CREM to a cognate CRE in the first intron of CRABP-II (Fig. 5, i and j). Association of CREM with this CRE was accompanied by recruitment of HDAC1 (Fig. 5j), revealing that CREM functions as a direct repressor of CRABP-II.
CREM is up-regulated upon induction of adipocyte differentiation, but its expression is transient and returns to basal level 8 h later (Fig. 5e). Accordingly, CREM was found to be associated with the CRE of the CRABP-II gene in early stages of adipogenesis (Fig. 5j) but not in mature adipocytes (Fig. 7b). These observations indicate that another mechanism must be in place to maintain the low level of CRABP-II expression observed in mature adipocytes (25). The data show that, in mature adipocytes, CRABP-II expression is suppressed by C/EBPα, a critical regulator of adipogenesis whose expression is induced late during differentiation and remains high in mature adipose cells (4). Inspection of the CRABP-II promoter revealed the presence of five consensus C/EBPα-response elements, and ChiP assays indicated that C/EBPα is associated with this region in mature adipocytes (Fig. 7, a and b). The observations that C/EBPα effectively down-regulates the expression of CRABP-II (Fig. 7, c and d) are in agreement with the reports that this transcription factor can recruit transcriptional repressors and function as a negative regulator of gene expression (28, 33).
The data thus demonstrate that down-regulation of CRABP-II is critical for allowing adipocyte differentiation, and they elucidate the mechanisms by which the expression of this protein is suppressed during adipogenesis and remains low in mature adipocytes. The mechanisms through which RA, functioning by activating RAR and CRABP-II, inhibits adipogenesis remain to be clarified.
It has been reported that CRABP-II inhibits the growth of cultured mammary carcinoma cells (29, 30) and suppresses tumor development in mouse models of breast cancer (18, 19, 34). It was further shown that, in the murine mammary tumor virus-neu mouse model of breast cancer, tumorigenesis is accompanied by down-regulation of CRABP-II expression (19). However, the molecular events that underlie the loss of this protein during tumor initiation and progression are unknown. The observations that the expression of CRABP-II is suppressed by cAMP signaling may provide insight into such mechanisms. It is interesting to note in regard to this that cAMP activities have been shown to promote carcinoma cell growth (35, 36) and were implicated in the initiation and progression of multiple types of tumors (37). cAMP-dependent enhancement of carcinoma cell growth may be mediated, at least in part, by suppression of CRABP-II.
This work was supported, in whole or in part, by National Institutes of Health Grant DK060684 (to N. N.).
Supported by National Institutes of Health Grant DK073195T32.
- IBMX
- isobutylmethylxanthine
- C/EBPα
- CCAAT/enhancer-binding protein α
- RA
- retinoic acid
- RAR
- retinoic acid receptor
- CREB
- cAMP-response element-binding protein
- CREM
- cAMP-response element modulator
- GR
- glucocorticoid receptor
- PKA
- protein kinase A
- PPARγ
- peroxisome proliferator-activated receptor γ
- ChIP
- chromatin immunoprecipitation
- Q-PCR
- quantitative real time PCR
- iLBP
- intracellular lipid-binding protein
- shRNA
- small hairpin RNA
- CRE
- cAMP-response element.
REFERENCES
- 1.Green H., Meuth M. (1974) Cell 3, 127–133 [DOI] [PubMed] [Google Scholar]
- 2.Green H., Kehinde O. (1975) Cell 5, 19–27 [DOI] [PubMed] [Google Scholar]
- 3.Rosen E. D., Spiegelman B. M. (2000) Annu. Rev. Cell Dev. Biol. 16, 145–171 [DOI] [PubMed] [Google Scholar]
- 4.Rosen E. D., Walkey C. J., Puigserver P., Spiegelman B. M. (2000) Genes Dev. 14, 1293–1307 [PubMed] [Google Scholar]
- 5.Zhang J. W., Klemm D. J., Vinson C., Lane M. D. (2004) J. Biol. Chem. 279, 4471–4478 [DOI] [PubMed] [Google Scholar]
- 6.Petersen R. K., Madsen L., Pedersen L. M., Hallenborg P., Hagland H., Viste K., Døskeland S. O., Kristiansen K. (2008) Mol. Cell. Biol. 28, 3804–3816 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Reusch J. E., Colton L. A., Klemm D. J. (2000) Mol. Cell. Biol. 20, 1008–1020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Foulkes N. S., Borrelli E., Sassone-Corsi P. (1991) Cell 64, 739–749 [DOI] [PubMed] [Google Scholar]
- 9.Don J., Stelzer G. (2002) Mol. Cell. Endocrinol. 187, 115–124 [DOI] [PubMed] [Google Scholar]
- 10.Sassone-Corsi P. (1995) Annu. Rev. Cell Dev. Biol. 11, 355–377 [DOI] [PubMed] [Google Scholar]
- 11.Shi X. M., Blair H. C., Yang X., McDonald J. M., Cao X. (2000) J. Cell. Biochem. 76, 518–527 [DOI] [PubMed] [Google Scholar]
- 12.Pantoja C., Huff J. T., Yamamoto K. R. (2008) Mol. Biol. Cell 19, 4032–4041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Smas C. M., Chen L., Zhao L., Latasa M. J., Sul H. S. (1999) J. Biol. Chem. 274, 12632–12641 [DOI] [PubMed] [Google Scholar]
- 14.Schwarz E. J., Reginato M. J., Shao D., Krakow S. L., Lazar M. A. (1997) Mol. Cell. Biol. 17, 1552–1561 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chambon P. (1996) FASEB J. 10, 940–954 [PubMed] [Google Scholar]
- 16.Deleted in proof
- 17.Shaw N., Elholm M., Noy N. (2003) J. Biol. Chem. 278, 41589–41592 [DOI] [PubMed] [Google Scholar]
- 18.Schug T. T., Berry D. C., Shaw N. S., Travis S. N., Noy N. (2007) Cell 129, 723–733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Schug T. T., Berry D. C., Toshkov I. A., Cheng L., Nikitin A. Y., Noy N. (2008) Proc. Natl. Acad. Sci. U.S.A. 105, 7546–7551 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Budhu A., Gillilan R., Noy N. (2001) J. Mol. Biol. 305, 939–949 [DOI] [PubMed] [Google Scholar]
- 21.Budhu A. S., Noy N. (2002) Mol. Cell. Biol. 22, 2632–2641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Dong D., Ruuska S. E., Levinthal D. J., Noy N. (1999) J. Biol. Chem. 274, 23695–23698 [DOI] [PubMed] [Google Scholar]
- 23.Sessler R. J., Noy N. (2005) Mol. Cell 18, 343–353 [DOI] [PubMed] [Google Scholar]
- 24.Tan N. S., Shaw N. S., Vinckenbosch N., Liu P., Yasmin R., Desvergne B., Wahli W., Noy N. (2002) Mol. Cell. Biol. 22, 5114–5127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Berry D. C., Noy N. (2009) Mol. Cell. Biol. 29, 3286–3296 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bakker A. H., Van Dielen F. M., Greve J. W., Adam J. A., Buurman W. A. (2004) Obes. Res. 12, 488–498 [DOI] [PubMed] [Google Scholar]
- 27.Tenbrock K., Juang Y. T., Leukert N., Roth J., Tsokos G. C. (2006) J. Immunol. 177, 6159–6164 [DOI] [PubMed] [Google Scholar]
- 28.Vernochet C., Peres S. B., Davis K. E., McDonald M. E., Qiang L., Wang H., Scherer P. E., Farmer S. R. (2009) Mol. Cell. Biol. 29, 4714–4728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Donato L. J., Noy N. (2005) Cancer Res. 65, 8193–8199 [DOI] [PubMed] [Google Scholar]
- 30.Donato L. J., Suh J. H., Noy N. (2007) Cancer Res. 67, 609–615 [DOI] [PubMed] [Google Scholar]
- 31.Berry D. C., Noy N. (2007) PPAR Res. 2007, 73256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Xue J. C., Schwarz E. J., Chawla A., Lazar M. A. (1996) Mol. Cell. Biol. 16, 1567–1575 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Soliera A. R., Lidonnici M. R., Ferrari-Amorotti G., Prisco M., Zhang Y., Martinez R. V., Donato N. J., Calabretta B. (2008) Blood 112, 1942–1950 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Manor D., Shmidt E. N., Budhu A., Flesken-Nikitin A., Zgola M., Page R., Nikitin A. Y., Noy N. (2003) Cancer Res. 63, 4426–4433 [PubMed] [Google Scholar]
- 35.de Rooij J., Rehmann H., van Triest M., Cool R. H., Wittinghofer A., Bos J. L. (2000) J. Biol. Chem. 275, 20829–20836 [DOI] [PubMed] [Google Scholar]
- 36.Detry C., Lamour V., Castronovo V., Bellahcène A. (2008) Bone 42, 422–431 [DOI] [PubMed] [Google Scholar]
- 37.Stork P. J., Schmitt J. M. (2002) Trends Cell Biol. 12, 258–266 [DOI] [PubMed] [Google Scholar]