Abstract
The molecular mechanisms regulating organ growth and size remain unclear. Sonic hedgehog (SHH) signaling is a major player in the regulation of cerebellar development: SHH is secreted by Purkinje neurons and acts on the proliferation of granule cell precursors (GCPs) in the external germinal layer. These then become postmitotic and form the internal granular layer but do so in the presence of SHH ligand, begging the question of how the proliferative response to SHH signaling is downregulated in differentiating GCPs. Here, we have determined the precise cellular localization of the expression of insulin-like growth factor (IGF) network components in the developing mouse cerebellum and show that this network modulates the proliferative effects of SHH signaling on GCPs. IGF1 and IGF2 are potent mitogens for GCPs and both synergize with SHH in inducing GCP proliferation. Whereas the proliferative activity of IGF1 or IGF2 on GCPs does not require intact SHH signaling, aspects of SHH activity on GCP proliferation require signaling through the IGF receptor 1. Moreover, we find that 3 of the IGF-binding proteins, IGFBP2, IGFBP3 and IGFBP5, inhibit IGF1/2-induced cell proliferation, whereas IGFBP5 also inhibits SHH-induced GCPs proliferation. This novel function of IGFBP5 that we have uncovered demonstrates the exquisite regulation of SHH signaling by different components of the IGF network.
Key Words: Cerebellum, Proliferation, Granule cell precursor, Sonic hedgehog, Insulin-like growth factor, IGF-binding protein
Introduction
The size and shape of the cerebellum depend largely on the size of the multilayered cerebellar cortex. It is thought that the regulation of the granule cell precursor (GCP) cell number is critical to the definition of cerebellar size. Moreover, medulloblastomas, one of the most devastating pediatric brain tumors, are thought to arise from GCPs, indicating that the regulation of the GCP number is also critical to prevent tumorigenesis.
GCPs are born in the rhombic lip at the level of the fourth ventricle and migrate tangentially to form the external germinal layer (EGL) [1,2,3,4]. In mice, GCPs then undergo a highly controlled proliferation program in the EGL during the first 2 weeks of the postnatal life. This proliferation is controlled in part by Purkinje neurons (PNs), located in the Purkinje layer (PL) internal to the EGL [5].
Sonic hedgehog (SHH) signaling is responsible for the growth and patterning of the cerebellum: SHH is secreted by PNs and regulates the proliferation of GCPs [6,7,8,9]. Deregulation or inappropriate activation of the SHH signaling pathway has been proposed to contribute to medulloblastoma [10,11,12]; reviewed in [13]. How SHH signaling is tightly regulated during GCP proliferation is thus a key issue to understand normal development and tumorigenesis.
Recent studies have provided insights into the interactions of the SHH pathway with other molecules during the development of GCPs. These regulators include vitronectin [14], bone morphogenetic proteins [14,15], fibroblast growth factor [9], Notch [16], stromal-derived factor-1 [17], proteoglycans [18], β-1 integrins [19], pituitary adenylate cyclase-activating polypeptide-1 [20] as well as the insulin-like growth factor (IGF) system [21].
The IGF signaling network is composed of 2 ligands (IGF1 and IGF2), 2 receptors (IGFR1 and IGFR2) and 6 IGF-binding proteins (IGFBPs) known to modulate the activity of the ligands [22,23]. IGFR1, a tyrosine kinase receptor, is thought to transduce most of the biological activities on growth and differentiation of the 2 ligands (reviewed in Werner and Le Roith [24]). IGF1 has been shown to act on the survival and proliferation of GCPs [e.g. [25,26,27]]. Members of this network are expressed in the cerebellum and interactions between the SHH and the IGF pathways have been postulated. For instance, mice lacking Ptch1 require Igf2 function to develop rhabdomyosarcoma [21,28]. Igf2 has also been proposed to be a target of SHH signaling, suggesting that it acts as a critical mediator of SHH function [21]. Moreover, synergism between SHH and IGF2 on GCPs has been suggested [29]. However, this derives from data using whole cerebellar cell cultures, making it impossible to know which cells signal and which respond. Indeed, GCPs are not the only cell type that responds to SHH from PNs [6]. However, more recently, Parathath et al. [30] have shown that the insulin receptor substrate I (IRS-I) is an effector of the SHH signaling in the GCPs. Nevertheless, apart from this last study and the wealth of information on the role of IGF signaling in other systems [31,32,33,34,35] little is known about how the IGF network, including the IGFBPs, interacts with SHH signaling in cerebellar GCPs and about the precise cellular bases of such interactions in the developing cerebellum.
Here we have investigated these interactions during GCP proliferation. We show that the mitogenic effect of IGF1/2 on GCPs is inhibited by IGFBPs and is not dependent on SHH signaling. On the contrary, we demonstrate that SHH and IGF signaling synergize and that aspects of SHH signaling in GCPs require functional IGFR1. Finally, we show that only IGFBP5 is able to efficiently inhibit the SHH-induced proliferation of GCPs. We propose that the regulation of GCP proliferation by SHH signaling, and thus cerebellar growth, may therefore depend on the modulatory effect of the IGF signaling network.
Materials and Methods
Animals and Tissue Preparation for in situ Hybridization
CD-1 embryos and pups were staged counting the morning after conception as embryonic day (E) 0.5 and the morning after birth as postnatal day (P) 0. Embryos were dissected from uteri in ice-cold PBS and cold-anesthetized pups were intracardially perfused with 4% paraformaldehyde (PFA) in phosphate buffer.
Riboprobe Synthesis
Total RNA was extracted from fresh murine tissues using Trizol (Invitrogen), treated with DNase I (Promega) then reverse transcribed using Superscript II reverse transcriptase according to the manufacturer's instructions (Invitrogen). cDNA fragments corresponding to each member of the mouse IGF system were cloned by PCR amplification using the following primers:
Igf1 5′-GGACCTACCAAAATGACCGC-3′ and 5′-GGTTGCTCAAGCAGCAAAGG-3′,
Igf2 5′-CATCAATCTGTGACCTCCTC-3′ and 5′-GAAAGACAGAACTAGCAGCC-3′,
Igfr1 5′-GACGAGTGGAGAAATCTGTG-3′ and 5′-CTTGGGCACATTTTCTGGCA-3′,
Igfbp1 5′-CTGTTGTTTCTTGGCCGTTC-3′ and 3′-GGCTCCTTCCATTTCTTGAG-5′,
Igfbp3 5′-GAAACATCAGTGAGTCCGAG-3′ and 3′-CAATGTACGTCGTCTTTCCC-5′,
Igfbp4 5′-CTTGCTCCGAGGAGAAGCTG-3′ and 3′-CTTCCGATCCACACACCAGC-5′,
Igfbp6 5′-CTACAAAGGAGAGCAAACCC-3′ and 3′-AACTAGCTGTGTAAAGGCCC-5′.
cDNA fragments were amplified from the following tissues: whole E11.5 embryo (Igf1, Igfr1, Igfbp3), whole E12.5 embryo (Igfbp1), E18.5 cerebellum (Igfbp4), adult olfactory bulb (Igfbp6), E12.5 liver (Igf2). The amplified fragments were cloned into the pGEMT easy vector (Promega) and their identity was confirmed by direct sequencing. The Igfbp5 mouse cDNA fragment (located in the 3′UTR) was isolated from an embryonic neocortex cDNA library. The IMAGE clone 2654907 was used as a template to synthesize the mouse Igfbp2 RNA riboprobes. Digoxigenin-labeled sense and antisense riboprobes were obtained by in vitro transcription using T7 and SP6 transcriptase (Promega) and digoxigenin-UTP (Roche). They were stored in hybridization buffer (see below) at the concentration of 10 μg/ml.
In situ Hybridization of Frozen Sections on Glass Slides
The brains were removed, fixed in 4% PFA for 2–24 h at 4°C, cryoprotected in 30% sucrose/PBS, embedded in OCT and cryosectioned at 14 μm. In situ hybridization was performed as previously described [36]. The sections were washed in PBS, then RIPA (150 mM of NaCl, 1% NP-40, 0.5% Na deoxycholate, 0.1% SDS, 1 mM of EDTA, pH 8, 50 mM of Tris, pH 8), post-fixed in 4% PFA for 5 min and treated with 0.25 acetic anhydre in 0.1 M of triethanolamine. Hybridization was performed with digoxigenin-labeled sense or antisense riboprobes (1 μg/ml) overnight at 70°C in hybridization buffer (50% formamide, 5X SSC, 5X Denhardt's, 250 μg/ml yeast tRNA, 125 mg/l salmon sperm DNA). The sections were washed with 50% formamide/2X SSC/0.1%, Tween 20 for 2 h at 70°C then MABT (100 mM of maleic acid and 150 mM of NaCl at pH 7.5, 0.1% Tween 20) at room temperature. After blocking with 10% goat serum/MABT, hybridization was revealed by incubation with an alkaline phosphatase (AP) coupled antibody (Roche, dilution 1:2,500 in MABT, 10% heat-inactivated goat serum blocking buffer) overnight at 4°C. The slides were washed with MABT and with AP buffer (100 mM of Tris, pH 9, 100 mM of NaCl, 50 mM of MgCl2, 0.24 mg/ml levamisole/0.1%, Tween 20). A 5-bromo-4-chloro-3-indolyl phosphate/nitroblue tetrazolium (Sigma) solution was then layered on top of the sections on the slides and covered with parafilm.
Primary Cerebellar Granule Cell Cultures
Granule cell culture was established using Percoll gradient fractionation as previously described [6,26,37] with minor modifications. Cerebella from postnatal day 4–5 (P4–5) CD-1 mice were dissected in HBSS and the meninges were carefully removed. The cerebella were treated with trypsin (Sigma) and DNase I (Serlabo) for 20 min at 37°C and then triturated using fire-polished glass Pasteur pipettes. After centrifugation the cell suspensions were resuspended in DMEM-F12 supplemented with glutamine (2 mM), penicillin (50 units/ml), streptomycin (50 μg/ml) and B27 (1×; Gibco) and passed through a cell strainer (Falcon). The GCPs were then purified by depleting adherent cells with 2 rounds of 30 min plating on plastic dishes coated with poly-D-lysine (MP Biomedicals). The resulting cultures contained >95% of granule cells. The cells were then pelleted and resuspended in DMEM-F12 media supplemented as described above and plated on glass coverslips coated with poly-D-lysine in 24-well plates at the density of 0.8.106 cells/well. For reaggregate cultures, cells were plated at 1.5.106 cells/well in uncoated 24-well plates. The day after, the GCPs were treated with the different growth factors.
Cerebellar Explants
Cerebella from P0 or P1 CD-1 mice were removed, embedded in 2.5% agarose and cut into 250-μm-thick sagittal sections using a vibratome. Meninges were carefully removed and slices were grown on Milicel culture inserts (Millipore) in 6-well plates containing serum-free defined medium as described above.
IGF, SHH and Other Treatments
IGF1 was purchased from Roche, IGF2 and SHH from R&D Systems, IGFBP1, IGFBP3 and IGFBP5 from Upstate and IGFBP2 from Gropep. Cyclopamine was purchased from Toronto Research Chemicals. IGF1, IGF2, IGFBPs, SHH and cyclopamine were added at the time of plating for explant cultures, after cell adhesion for dissociated cells and after aggregate formation in reaggregate cultures. The IGFR1 blocking antibody [38], as well as the control antibody (nonrelated antibody of the same subtype) was a gift from Dr. D. Hicklin and Dr. D. Ludwig (ImClone Systems Inc.) and used at 20 μg/ml. Each treatment was performed with serum-free defined medium consisting of DMEM-F12 supplemented with penicillin (50 units/ml), streptomycin (50 μg/ml), transferrin (100 μg/ml), putrescine (100 μM), progesterone (20 nM), selenium (30 nM), glutamine (2 mM), glucose (6 mg/ml), guanosine (200 μM) and bovine serum albumin (10 mg/ml). The cultures were maintained for 48 h after factors had been added.
Bromodeoxyuridine Incorporation and Detection
Bromodeoxyuridine (BrdU) (12 μg/ml) was added to the culture medium 18 h (2 h for explants) before fixation with 4% PFA for 1 h. Then granule cells and explant cultures were washed in 0.1% Triton X-100/PBS, treated for 30 min with 2 N HCl at 37°C, neutralized with 0.1 M of sodium borate, pH 8.5, for 20 min at room temperature, blocked with 10% goat serum/0.1% Triton X-100/PBS at room temperature and incubated overnight at 4°C with a mouse anti-BrdU antibody (Becton Dickinson, dilution 1:400). The samples were washed and stained with a goat anti-mouse IgG conjugated to FITC (Jackson, dilution 1:500) for 45 min at room temperature. The nuclei were counterstained with Hoechst (Sigma) and the samples were mounted on slides using Mowiol/Dabko. BrdU-positive cells were counted with a fluorescent microscope using a ×20 objective (Axioskop; Zeiss). At least 10 independent fields for each culture condition were counted per culture condition. Statistical analyses were performed with the Student t test.
Results
Expression Pattern of Members of the IGF Signaling Network in the Late Embryonic and Postnatal Mouse Cerebellum
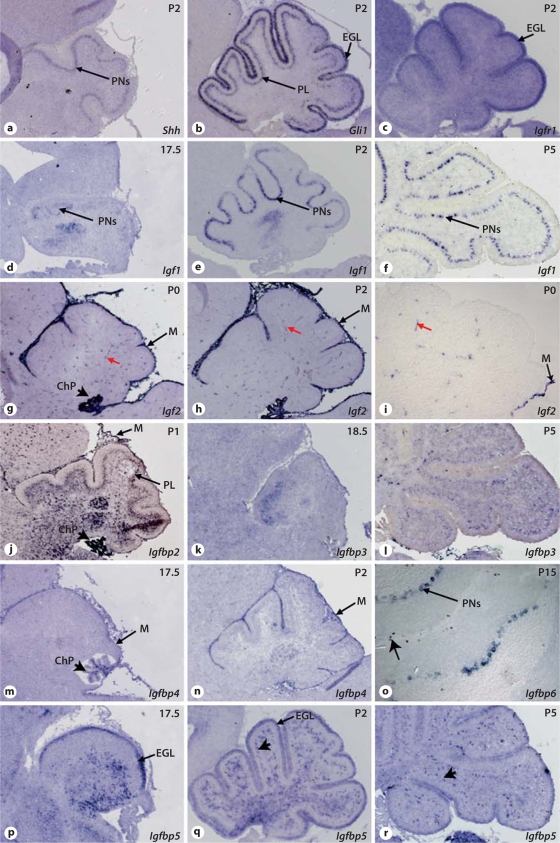
In order to assess the role of the IGF signaling network in SHH-driven proliferation of GCPs, we examined the expression patterns of members of this signaling pathway in late embryonic and postnatal cerebellar development by in situhybridization. These patterns of expression were compared with those of Shh, encoding the secreted ligand, and Gli1, encoding the final mediator of SHH signaling, the expression of which is also a reliable target of the SHH pathway [e.g. [39,40]]. As previously described [e.g. [6,41]] Shh was expressed by PNs in the cerebella of P1–P5 mice (fig. 1a), and Gli1 was expressed both in GCPs in the EGL and in Bergmann glial cells in the PL (fig. 1b [41]). Igfr1, the main IGF receptor, was expressed ubiquitously in the cerebellum at all stages examined with a high level of expression in the EGL (fig. 1c). Consistent with previous data [42], we found that Igf1 was expressed in PNs as early as E17.5 (fig. 1d). Igf1 expression in PNs was not uniform as it was found only in a subset of PNs at P2 (fig. 1e), becoming more widely expressed around P5 (fig.1f). Igf2 was expressed in the meninges and in blood vessels within the cerebellum (fig. 1g–i). It was also highly expressed by the choroid plexus (fig. 1g–h). We observed the same expression pattern for Igf2 in a cerebellum in which meninges were removed (fig. 1i). Thus, neither Igf1 nor Igf2 were found by in situ hybridization to be expressed in mouse GCPs (fig. 1d–i).
Fig. 1.
Expression analysis of members of the IGF signaling network in the developing embryonic and postnatal cerebellum. Sagittal sections of mouse embryonic (E17.5) and postnatal (P0, P1, P2 or P5) cerebellum were hybridized with RNA probes corresponding to mouse Shh, Gli1 and members of the IGF signaling network. Shh is expressed in the PNs (a) and Gli1 expression in the EGL and in cells in the PL (b). Igfr1 expression is ubiquitous in the cerebellum, with stronger expression in EGL cells (c, arrow). Igf1 expression in the PNs is apparent as early as E17.5 (d, arrow), becoming stronger during the early postnatal stages (e, f). Igf2 is detected in meninges (g–i, arrows), choroid plexus (arrowhead in g) and in blood vessels (red arrows in g–i) in the cerebellar tissue. Igfbp2 is expressed in the PL, choroids plexus and meninges (j). Igfbp3 is expressed in the PL (k, l), while Igfbp4 expression is observed in the meninges (arrow) and choroid plexus (arrowhead in m) (m, n). Igfbp6 is expressed in a subset of PNs (o); note that proliferating phosphoshistone H3 positive cells are detected in the EGL of P15 cerebellum (arrow in o). Igfbp5 expression is detected at all stages in the EGL and cells within the PL (arrows in p, r). M = Meninges; ChP = choroid plexus.
The Igfbps genes exhibited different patterns of expression during the development of the cerebellum (fig. 1j–r). Igfbp2 was found in the meninges, the PL, the internal granular layer, the plexus choroid (fig. 1j, see also Bondy and Lee [42]) as well as in the EGL (online suppl. fig. 1, www.karger.com/doi/10.1159/000274458). Igfbp3 mRNA was found in the PL (fig. 1k, l). Igfbp4 mRNA was observed in the meninges and choroid plexus, with a similar pattern to Igf2, although at lower levels (fig. 1m, n). Igfbp5 expression was found in the EGL cells, mostly in proliferating outer EGL (fig. 1p–r; see also Bondy and Lee [42]). Igfbp5 was also strongly expressed in cells within the PL (fig. 1q, r) that do not correspond to calbindin-positive PNs (online suppl. fig. 2). Igfbp6 was not detected in the cerebellum until P8, when it was seen in a subset of PNs and was still observed at P15, the latest stage examined (fig. 1o). We did not observe any expression of Igfbp1 in the cerebellum at any stage examined.
Both IGF1 and IGF2 Are Mitogenic for GCPs
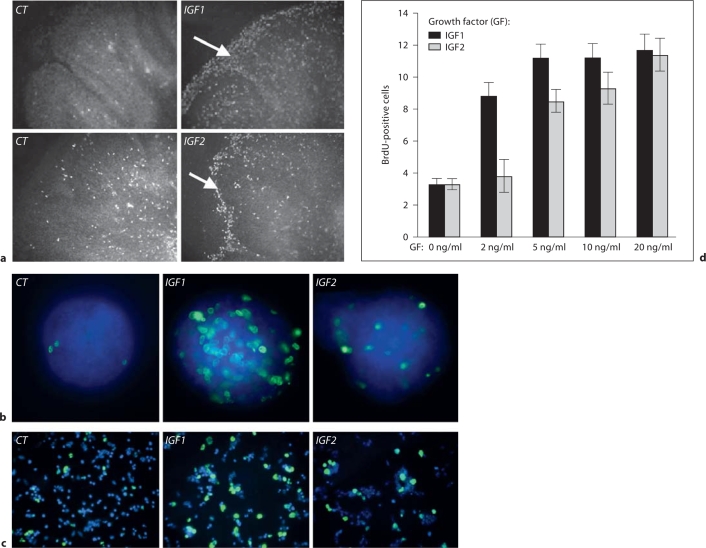
In order to analyze the functional relationship between the IGF signaling network and the SHH signaling pathway, we first examined the role of recombinant IGF1 and IGF2 on GCPs. We used 3 systems to investigate their activities: cerebellar explants, purified GCPs cultured as aggregates [6,26] and isolated purified GCPs cultured on polylysine-coated dishes [8,9]. We analyzed cell proliferation using a pulse of BrdU incorporation to mark cells that have newly synthetized DNA. In all systems, we found that IGF1 and IGF2 are potent mitogens for GCPs (fig. 2a–c). Quantification of these effects was done on isolated cultures of purified GCPs and showed a clear dose-dependent effect of these 2 growth factors on the number of BrdU-positive cells (fig. 2d).
Fig. 2.
IGF1 and IGF2 act as mitogens on GCP proliferation. The effects of IGFs on GCP proliferation were tested using 250-μm postnatal cerebellar explants (a), GCP aggregates (b) or isolated, purified GCPs (c). After 48 h in culture, IGF1 (20 ng/ml) and IGF2 (40 ng/ml) induce an increase in cell proliferation in the outer side of the explants containing the GCPs (arrows in a), in the GCP aggregates (b) or in isolated cells (c) as visualized by BrdU incorporation. b, c All nuclei are stained with Hoechst in blue and BrdU is in green. The quantification of the IGF1 and IGF2 proliferative effect on GCPs in isolated cultures shows a dose effect of these growth factors (d).
SHH Acts Synergistically with IGF1 and IGF2
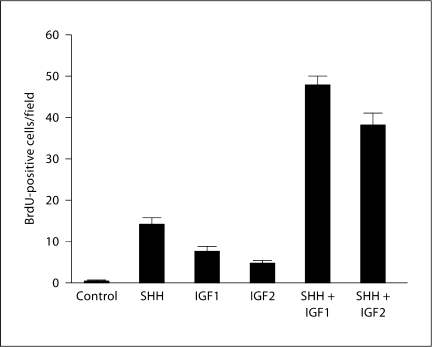
Since these effects of the IGF network parallel the mitogenic effects of SHH signaling on GCPs, we assayed for the ability of IGF1 or IGF2 to modify the proliferative response of purified GCPs to SHH. As expected, treatment with SHH (recombinant N-SHH) alone enhanced the proliferation of GCPs (fig. 3) in a dose-dependent manner (not shown). Treatment of GCPs with IGF1 and IGF2 also resulted in an increase in BrdU incorporation, although it was smaller than that observed with SHH alone (fig. 3). However, simultaneous treatment with SHH plus IGF1 or IGF2 resulted in a strong synergistic effect: BrdU incorporation in GCPs was enhanced approximately 3-fold in response to combined SHH and IGF signaling as compared with SHH alone and about 6- or 7-fold as compared with IGF1 or IGF2 alone, respectively (fig. 3). These results strongly suggest that IGF and SHH signaling interact to increase the proliferation of GCPs.
Fig. 3.
SHH acts synergistically with IGF1 and IGF2. Purified GCPs were either not treated (control) or treated with SHH or SHH + IGF1 (n = 4 for each condition). GCPs were also treated with SHH or SHH + IGF2 (n = 5 for each condition). Cell proliferation is significantly higher when GCPs are treated with both SHH + IGF1 (p < 0.001) and SHH + IGF2 (p < 0.001) than when they are treated with SHH alone. The concentrations of growth factors were as follows: IGF1: 20 ng/ml; IGF2: 40 ng/ml; SHH was used at 400 or 600 ng/ml. Similar results were obtained with these 2 concentrations of SHH in all experiments.
IGF-1 and IGF2-Induced Proliferation Is Not Dependent on the Function of Smoothened
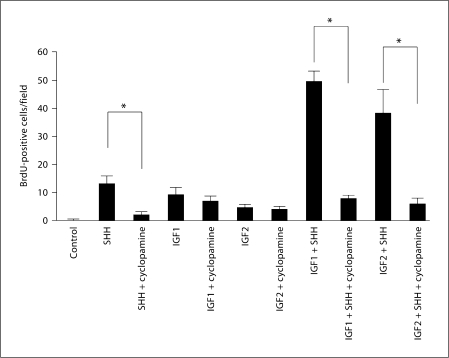
The combined action of IGFs and SHH signaling could be explained by the action of IGF signaling downstream of the SHH pathway as Igf2, for instance, is a required element in tumors induced by activation of SHH function [21]. In this case, exogenous IGF would simply be enhancing the outcome of SHH signaling on GCPs. However, neither Igf1 nor Igf2 are detected in GCPs in vivo (fig. 1). Thus, IGFs from surrounding cells may affect GCPs in the EGL. This raises the possibility that the synergistic effect we document could derive from a combined action of IGFs and SHH on the SHH receptor complex. To test if IGF signaling requires the function of Smoothened, which is the critical transmembrane signal transducer of the SHH pathway [43], we treated purified GCPs with SHH, IGF1 or IGF2 alone or each plus cyclopamine, a specific inhibitor of Smoothened [44] (fig. 4). Cyclopamine (5 μM) drastically diminished the effect of SHH. In contrast, it had no significant effect on the level of BrdU incorporation induced by IGF1 or IGF2 (fig. 4). Thus, IGFs do not appear to require active Smoothened signaling to enhance GCP proliferation.
Fig. 4.
Cyclopamine does not affect IGF1- or IGF2-induced proliferation of GCPs. Purified GCPs were treated with SHH (400 ng/ml; n = 3), IGF1 (20 ng/ml) or IGF2 (40 ng/ml) alone or in combination with cyclopamine (5 μM). All growth factors alone enhance cell proliferation. Addition of cyclopamine significantly decreases cell proliferation for cells treated with SHH (p < 0.0001; n = 4), but it does not induce any significant decrease in the number of BrdU-positive cells when GCPs are treated with either IGF1 (p = 0.19; n = 4) or IGF2 (p = 0.15; n = 4). ∗ p < 0.05.
SHH-Induced Proliferation Is Dependent on IGF Receptor Function
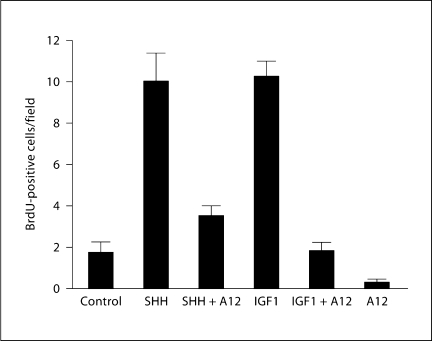
To directly test for the involvement of IGFR1, the main transmembrane tyrosine kinase IGF receptor involved in transducing mitogenic signals (reviewed in [24]), purified GCPs were treated with IGF1 and the monoclonal antibody A12, a specific antibody to IGFR1 that blocks its function [45]. The proliferative response of GCPs treated with IGF1 was antagonized by addition of the anti-IGFR1 blocking antibody (fig. 5). We then tested if blocking IGFR1 signaling had any effect on SHH-induced proliferation. In this assay we used a lower dose of SHH than the one in the previous experiments as we wanted to be in the range of the minimal doses that can induce an effect. Treatment of purified GCPs with SHH (200 ng/ml) produced a marked enhancement of BrdU incorporation (fig. 5). Surprisingly, SHH-induced proliferation was inhibited by >50% by treatment with anti-IGFR1 blocking antibody (fig. 5) but not by treatment with a nonrelated antibody of the same subtype (ImClone; not shown).
Fig. 5.
IGF signaling is required for SHH-induced proliferation of GCPs. Purified GCPs were treated with SHH (200 ng/ml, n = 7 independent tests) or IGF1 (20 ng/ml, n = 6) alone or in combination with the monoclonal antibody A12, an anti-IGFR1 blocking antibody (20 μg/ml). Blockade of IGFR1 induces a decrease in proliferation on GCPs following treatment with IGF1 (p < 0.0001) or SHH (200 ng/ml; p < 0.0001).
IGFBPs Inhibit the Activity of IGFs on GCPs Proliferation
Igfbps were also found to be highly expressed in the cerebellum (fig. 1, [46]), but their functions in this brain region are not known. As they are able to bind IGFs [23], one hypothesis is that they modulate the activity of IGF1 and IGF2 during proliferation of GCPs. Culture of GCPs in the presence of IGF1 or IGF2 together with recombinant IGFBPs showed that all IGFBPs tested strongly reduced the number of BrdU-positive cells induced in response to IGF1 or IGF2 (fig. 6; not shown).
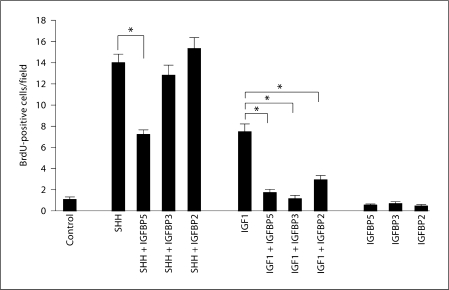
Fig. 6.
SHH-induced proliferation is specifically inhibited by IGFBP5 but not by IGFBP2 or IGFBP3. Purified GCPs were either left untreated (control, n = 14 independent tests) or treated with the following binary combinations of factors: SHH or SHH + IGFBP2 (n = 5 for each condition); SHH or SHH + IGFBP3 (n = 5 for each condition); SHH or SHH + IGFBP5 (n = 9 for each condition), IGF1 or IGF1 + IGFBP2 (n = 5 for each condition), IGF1 or IGF1 + IGFBP3 (n = 4 for each condition), IGF1 or IGF1 + IGFBP5 (n = 5 for each condition). IGFBPs were used at 400 ng/ml, IGF1 at 20 ng/ml and IGF2 at 40 ng/ml. Similar results were obtained with SHH concentrations of 400 or 600 ng/ml. IGF1-induced proliferation on GCPs is reduced when cells are co-treated with IGFBP2 (p < 0.0003), IGFBP3 (p < 0.0001) or IGFBP5 (p < 0.0001). However, only IGFBP5 causes a significant decrease in SHH-induced GCP proliferation (p < 0.0001), while IGFBP2 (p = 0.4) or IGFBP3 (p = 0.8) have no significant effect on SHH activity. ∗ = Significant differences.
IGFBP5 Selectively Diminishes SHH-Induced Proliferation of Purified GCPs and Cerebellar Explants
We therefore tested for the ability of IGFBPs to inhibit the effect of SHH on GCPs and found that while all the ones we tested (IGFBP2, −3 and −5) have the ability to inhibit IGF1 activity (fig. 6), only IGFBP5 also inhibited by half the effects of SHH on the proliferation of purified GCPs (fig. 6). IGFBP2 or IGFBP3 were unable to alter the response of GCPs to treatment with SHH, and IGFBPs on their own had no effect on GCP proliferation (fig. 6).
IGFBP5 was able to inhibit SHH-induced proliferation of GCPs even when high doses of SHH were used (1 μg/ml, data not shown) or when IGFBP5 was used at a lower dose (200 ng/ml, not shown). These results suggest that the mode of action of IGFBP5 is distinct from that of other IGFBPs as it appears to inhibit IGF signaling on one hand and SHH signaling on the other.
Discussion
IGF signaling has been implicated in many processes [47] during development and homeostasis. A number of the components of the IGF signaling network have also been previously shown to be expressed in the cerebellum [42,48,49,50]. However, the role of members of this signaling network in the cerebellum has remained unclear, in part due to the lack of knowledge of the precise sources and targets of signaling and inhibitory molecules, as well as of any potential interaction with SHH, which is the main regulator of cerebellar growth.
Two recent studies have brought important insights into the understanding of the collaboration between the SHH and IGF signaling pathways. The first one suggests that SHH and IGF2 act together in vitro [29]. In this study, however, whole cerebellar cultures were used, thus precluding the identification of the direct cellular targets affected. For instance, both GCPs in the EGL and Bergmann glia in the PL respond to SHH secreted from PNs [6]. The second study demonstrates that IRS-I is an effector of SHH signaling in GCPs [30]. These and other works urge a better characterization of SHH and IGF functions and, importantly, of their interactions in the cerebellum.
We find that IGF1 and IGF2 on their own have moderate effects on the proliferation of purified GCPs, in agreement with the phenotype of deletion of Igf1, Igf2 or Igfr1 [31,32,33]. However, they directly enhance the proliferation of GCPs in combination with exogenous SHH. This synergism supports the idea that the full effect of SHH signaling involves cooperation with the IGF signaling network. IGF1 and IGF2 appear to derive from local sources: the meninges and blood vessels for IGF2 and PNs for IGF1 (this work and [42,51]). They are also found in the circulation, raising the possibility that systemic IGF signaling could also be a general positive parallel cofactor for the full SHH mitogenic response in organs other than the nervous system.
Our finding showing that blockade of IGFR-I highly decreases SHH-induced GCP proliferation is consistent with recent data showing that IRS-I is an effector of SHH signaling [30]. However, in these studies GCPs were grown in the presence of N2 supplement [30], which contains insulin at levels sufficient to activate IGF-1R, thus making it impossible to sort out if IGF-1R (and IRS-1) function is solely dependent on SHH signaling. Instead, since we have grown GCPs in the absence of insulin sources, our data suggest an activation of IGF-1R by SHH signaling. How this happens remains unclear. While we cannot rule out the possibility that there is an SHH-independent tonic low level of cell-autonomous IGF-1R activity in GCPs in culture, one possibility is that the interactions between the SHH and IGF pathways parallel the interaction between the estrogen receptor and IGF-1R through an adaptor protein [52]. Other modes of interaction include the activation of PI3K and AKT by IGF-IGFR1 [29,53], since PI3K-AKT signaling has been proposed to enhance the outcome of SHH signaling by inhibiting the degradation of N-Myc and cyclin D1 through the inhibition of GSK3β [53]. As N-Myc is a critical mediator of aspects of SHH signaling in the cerebellum [53,54], IGF signaling may synergize with the SHH pathway through enhancement of the action of such targets of the SHH pathway. These possibilities are not exclusive and there may be multiple sites or modes of pathway interaction. Indeed, IGF2 enhances the level of Gli1 expression in cerebellar cultures [29], and since Gli1 is the last mediator of SHH signaling [40] and is itself a regulator of Nmyc and cyclin D1, an interaction between IGF-PI3K and SHH could also occur at the level of Gli1 regulation, upstream of the function of SHH-Gli targets such as N-Myc and D-type cyclins.
One of the most intriguing aspects of our data is the finding that while all IGFBPs tested diminish IGF1 and IGF2 function, only IGFBP5 also negatively modulates significantly the proliferative effect of SHH signaling on GCPs and cerebellar explants. IGF signaling does not require the SHH pathway, as cyclopamine does not inhibit the proliferative effects of IGF1 or IGF2, and SHH signaling does not activate AKT function [29,53]. IGFBP5 has been proposed to be able to either inhibit or activate IGF signaling in a context-dependent manner [23] and here we show that it acts to inhibit IGF function in cerebellar GCPs. Thus, it is possible that IGFBP5 has a selective dual role, acting as a common negative regulator of both the IGF signaling network and the SHH pathway.
The modulation of SHH signaling by IGFBP5 that we have uncovered may be general. This idea is supported by the striking conservation of the overlap of the Shh and Igfbp5 expression domains throughout the developing brain and in other organs (N.D., unpublished).
Thus, from the same or adjacent sources, and as secreted molecules, SHH and IGFBP5 are likely to affect the same or overlapping responding cell populations. We note that Igfbp5 has neither been described nor found in microarray analyses as an SHH-regulated gene, although in the chick neural tube, Igfbp5 is regulated by cyclopamine treatment [55], raising the possibility that SHH may regulate its expression and that IGFBP5 may act as a feedback inhibitor to limit the duration or strength of SHH signaling. In addition, IGFBP5 and other IGFBPs may also act locally to negatively modulate the effects of IGFs. However, it is not known how IGFBP5, but not the other IGFBPs, negatively modulates SHH signaling. It remains possible that IGFBP5 could also act through a receptor other than IGFR1, much like IGFBP3 has been shown to act through the TGFβ receptor [56] or it may also act in the nucleus as IGFBP5 has been shown to be nuclear and to interact with transcription factors [57].
Our data support a model in which the modulation of SHH effects by the IGF signaling network is important for the biological action of SHH on distinct cerebellar cell types. Collectively, our data show that the IGF signaling network and the SHH pathway interact in both positive and negative ways in cerebellar GCPs, suggesting a tug of war between different IGF signaling network components that leads to the refined modulation of the effect of SHH signaling on the regulation of GCP proliferation in the developing cerebellum. This includes both positive synergistic effects by IGFs, negative regulation of IGF function by IGFBPs, and negative modulation of both IGF and SHH signaling by IGFBP5. In this sense, IGF and insulin signaling have been known for many years to control organismal size and cell number [31,32,33,58], and here we link this important function to SHH signaling, which is similarly critical to the control of body size [59], cell number and possibly organ shape [60,61].
These findings have an important corollary for disease. Medulloblastomas are cerebellar tumors that are thought to derive, at least in part, from GCPs through the misregulation of the SHH pathway [6,8,9,10,28,62,63,64,65]. Our work suggests that the IGF signaling network may often be deregulated towards a positive, synergistic mode in relation with SHH in medulloblastoma. This is consistent with the requirement for IGF2 function in medulloblastoma and other tumors [21,29]. In this sense, IGFs in the circulation could play a critical role in sustaining SHH-dependent tumor growth by securing an overall positive input of the IGF signaling network on the proliferative role of the SHH pathway.
Supplementary Material
Igfbp2 expression in postnatal cerebellum. Sagittal sections of mouse postnatal P1 cerebellum were hybridized with RNA probe corresponding to full-length mouse Igfbp2. Igfbp2 expression is detected in the EGL, in cells within the PL, in cells in the IGL and in meninges.
Igfbp5 expression does not colocalize with Calbindin expression. Sagittal sections of mouse postnatal P1 cerebellum were hybridized with an antisense RNA probe corresponding to full-length mouse Igfbp5 and incubated with the anti-Calbindin antibody. Igfbp5 expression (purple) is detected in the EGL and in scattered cells within or around the Purkinje cell layer. The igfbp5 expression in the isolated cells does not colocalize with Calbindin expressing cells (brown) which correspond to Purkinje neurons (PN).
Acknowledgements
We thank Christophe Bail for the excellent technical assistance, and Drs. D. Hickling and D. Ludwig (ImClone Systems Inc.) for providing us with A12, the blocking IGFR1 monoclonal antibody as well as the control antibody. This work was supported by postdoctoral fellowships from la Fondation pour la Recherche Médicale and the Philippe Fondation to V.T. The work in the N.D. laboratory was supported by a CNRS ATIPE grant, l’Association pour la Recherche contre le Cancer, la Fondation pour la Recherche Médicale while at the Developmental Biology Institute of Marseille and currently by grants from the Albert R. Taxin Brain Tumor Research Center at the Wistar Institute, the V foundation, the WW Smith Charitable Trust. Support from the Wistar Institute Cancer Center grant NIH P30 CA010815 to N.D. laboratory is also acknowledged.
Footnotes
C.F. and V.T. have equally contributed to the work.
References
- 1.Goldowitz D, Hamre K. The cells and molecules that make a cerebellum. Trends Neurosci. 1998;21:375–382. doi: 10.1016/s0166-2236(98)01313-7. [DOI] [PubMed] [Google Scholar]
- 2.Hatten ME, Heintz N. Mechanisms of neural patterning and specification in the developing cerebellum. Annu Rev Neurosci. 1995;18:385–408. doi: 10.1146/annurev.ne.18.030195.002125. [DOI] [PubMed] [Google Scholar]
- 3.Sotelo C. Cellular and genetic regulation of the development of the cerebellar system. Prog Neurobiol. 2004;72:295–339. doi: 10.1016/j.pneurobio.2004.03.004. [DOI] [PubMed] [Google Scholar]
- 4.Wang VY, Zoghbi HY. Genetic regulation of cerebellar development. Nat Rev Neurosci. 2001;2:484–491. doi: 10.1038/35081558. [DOI] [PubMed] [Google Scholar]
- 5.Smeyne RJ, Chu T, Lewin A, Bian F, Sanlioglu SC, Kunsch C, Lira SA, Oberdick J. Local control of granule cell generation by cerebellar Purkinje cells. Mol Cell Neurosci. 1995;6:230–251. doi: 10.1006/mcne.1995.1019. [DOI] [PubMed] [Google Scholar]
- 6.Dahmane N, Ruiz i Altaba A. Sonic hedgehog regulates the growth and patterning of the cerebellum. Development. 1999;126:3089–3100. doi: 10.1242/dev.126.14.3089. [DOI] [PubMed] [Google Scholar]
- 7.Lewis PM, Gritli-Linde A, Smeyne R, Kottmann A, McMahon AP. Sonic hedgehog signaling is required for expansion of granule neuron precursors and patterning of the mouse cerebellum. Dev Biol. 2004;270:393–410. doi: 10.1016/j.ydbio.2004.03.007. [DOI] [PubMed] [Google Scholar]
- 8.Wallace VA. Purkinje-cell-derived sonic hedgehog regulates granule neuron precursor cell proliferation in the developing mouse cerebellum. Curr Biol. 1999;9:445–448. doi: 10.1016/s0960-9822(99)80195-x. [DOI] [PubMed] [Google Scholar]
- 9.Wechsler-Reya RJ, Scott MP. Control of neuronal precursor proliferation in the cerebellum by sonic hedgehog. Neuron. 1999;22:103–114. doi: 10.1016/s0896-6273(00)80682-0. [DOI] [PubMed] [Google Scholar]
- 10.Berman DM, Karhadkar SS, Hallahan AR, Pritchard JI, Eberhart CG, Watkins DN, Chen JK, Cooper MK, Taipale J, Olson JM, Beachy PA. Medulloblastoma growth inhibition by hedgehog pathway blockade. Science. 2002;297:1559–1561. doi: 10.1126/science.1073733. [DOI] [PubMed] [Google Scholar]
- 11.Dahmane N, Sanchez P, Gitton Y, Palma V, Sun T, Beyna M, Weiner H, Ruiz i Altaba A. The sonic hedgehog-gli pathway regulates dorsal brain growth and tumorigenesis. Development. 2001;128:5201–5212. doi: 10.1242/dev.128.24.5201. [DOI] [PubMed] [Google Scholar]
- 12.Pomeroy SL, Tamayo P, Gaasenbeek M, Sturla LM, Angelo M, McLaughlin ME, Kim JY, Goumnerova LC, Black PM, Lau C, Allen JC, Zagzag D, Olson JM, Curran T, Wetmore C, Biegel JA, Poggio T, Mukherjee S, Rifkin R, Califano A, Stolovitzky G, Louis DN, Mesirov JP, Lander ES, Golub TR. Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature. 2002;415:436–442. doi: 10.1038/415436a. [DOI] [PubMed] [Google Scholar]
- 13.Ruiz i Altaba A, Sanchez P, Dahmane N. Gli and hedgehog in cancer: tumours, embryos and stem cells. Nat Rev Cancer. 2002;2:361–372. doi: 10.1038/nrc796. [DOI] [PubMed] [Google Scholar]
- 14.Pons S, Trejo JL, Martinez-Morales JR, Marti E. Vitronectin regulates sonic hedgehog activity during cerebellum development through CREB phosphorylation. Development. 2001;128:1481–1492. doi: 10.1242/dev.128.9.1481. [DOI] [PubMed] [Google Scholar]
- 15.Rios I, Alvarez-Rodriguez R, Marti E, Pons S. Bmp2 antagonizes sonic hedgehog-mediated proliferation of cerebellar granule neurones through Smad5 signalling. Development. 2004;131:3159–3168. doi: 10.1242/dev.01188. [DOI] [PubMed] [Google Scholar]
- 16.Solecki DJ, Liu XL, Tomoda T, Fang Y, Hatten ME. Activated Notch2 signaling inhibits differentiation of cerebellar granule neuron precursors by maintaining proliferation. Neuron. 2001;31:557–568. doi: 10.1016/s0896-6273(01)00395-6. [DOI] [PubMed] [Google Scholar]
- 17.Klein RS, Rubin JB, Gibson HD, DeHaan EN, Alvarez-Hernandez X, Segal RA, Luster AD. SDF-1 α induces chemotaxis and enhances sonic hedgehog-induced proliferation of cerebellar granule cells. Development. 2001;128:1971–1981. doi: 10.1242/dev.128.11.1971. [DOI] [PubMed] [Google Scholar]
- 18.Rubin JB, Choi Y, Segal RA. Cerebellar proteoglycans regulate sonic hedgehog responses during development. Development. 2002;129:2223–2232. doi: 10.1242/dev.129.9.2223. [DOI] [PubMed] [Google Scholar]
- 19.Blaess S, Graus-Porta D, Belvindrah R, Radakovits R, Pons S, Littlewood-Evans A, Senften M, Guo H, Li Y, Miner JH, Reichardt LF, Muller U. Beta1-integrins are critical for cerebellar granule cell precursor proliferation. J Neurosci. 2004;24:3402–3412. doi: 10.1523/JNEUROSCI.5241-03.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nicot A, Lelievre V, Tam J, Waschek JA, DiCicco-Bloom E. Pituitary adenylate cyclase-activating polypeptide and sonic hedgehog interact to control cerebellar granule precursor cell proliferation. J Neurosci. 2002;22:9244–9254. doi: 10.1523/JNEUROSCI.22-21-09244.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hahn H, Wojnowski L, Specht K, Kappler R, Calzada-Wack J, Potter D, Zimmer A, Muller U, Samson E, Quintanilla-Martinez L, Zimmer A. Patched target IGF2 is indispensable for the formation of medulloblastoma and rhabdomyosarcoma. J Biol Chem. 2000;275:28341–28344. doi: 10.1074/jbc.C000352200. [DOI] [PubMed] [Google Scholar]
- 22.D'Ercole AJ, Ye P, O'Kusky JR. Mutant mouse models of insulin-like growth factor actions in the central nervous system. Neuropeptides. 2002;36:209–220. doi: 10.1054/npep.2002.0893. [DOI] [PubMed] [Google Scholar]
- 23.Firth SM, Baxter RC. Cellular actions of the insulin-like growth factor binding proteins. Endocr Rev. 2002;23:824–854. doi: 10.1210/er.2001-0033. [DOI] [PubMed] [Google Scholar]
- 24.Werner H, Le Roith D. New concepts in regulation and function of the insulin-like growth factors: implications for understanding normal growth and neoplasia. Cell Mol Life Sci. 2000;57:932–942. doi: 10.1007/PL00000735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Dudek H, Datta SR, Franke TF, Birnbaum MJ, Yao R, Cooper GM, Segal RA, Kaplan DR, Greenberg ME. Regulation of neuronal survival by the serine-threonine protein kinase Akt. Science. 1997;275:661–665. doi: 10.1126/science.275.5300.661. [DOI] [PubMed] [Google Scholar]
- 26.Gao WO, Heintz N, Hatten ME. Cerebellar granule cell neurogenesis is regulated by cell-cell interactions in vitro. Neuron. 1991;6:705–715. doi: 10.1016/0896-6273(91)90168-y. [DOI] [PubMed] [Google Scholar]
- 27.Ye P, Xing Y, Dai Z, D'Ercole AJ. In vivo actions of insulin-like growth factor-I (IGF-I) on cerebellum development in transgenic mice: evidence that IGF-I increases proliferation of granule cell progenitors. Brain Res Dev Brain Res. 1996;95:44–54. doi: 10.1016/0165-3806(96)00492-0. [DOI] [PubMed] [Google Scholar]
- 28.Hahn H, Wojnowski L, Zimmer AM, Hall J, Miller G, Zimmer A. Rhabdomyosarcomas and radiation hypersensitivity in a mouse model of Gorlin syndrome. Nat Med. 1998;4:619–622. doi: 10.1038/nm0598-619. [DOI] [PubMed] [Google Scholar]
- 29.Hartmann W, Koch A, Brune H, Waha A, Schuller U, Dani I, Denkhaus D, Langmann W, Bode U, Wiestler OD, Schilling K, Pietsch T. Insulin-like growth factor II is involved in the proliferation control of medulloblastoma and its cerebellar precursor cells. Am J Pathol. 2005;166:1153–1162. doi: 10.1016/S0002-9440(10)62335-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Parathath SR, Mainwaring LA, Fernandez LA, Campbell DO, Kenney AM. Insulin receptor substrate 1 is an effector of sonic hedgehog mitogenic signaling in cerebellar neural precursors. Development. 2008;135:3291–3300. doi: 10.1242/dev.022871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Baker J, Liu JP, Robertson EJ, Efstratiadis A. Role of insulin-like growth factors in embryonic and postnatal growth. Cell. 1993;75:73–82. [PubMed] [Google Scholar]
- 32.DeChiara TM, Efstratiadis A, Robertson EJ. A growth-deficiency phenotype in heterozygous mice carrying an insulin-like growth factor II gene disrupted by targeting. Nature. 1990;345:78–80. doi: 10.1038/345078a0. [DOI] [PubMed] [Google Scholar]
- 33.Liu JP, Baker J, Perkins AS, Robertson EJ, Efstratiadis A. Mice carrying null mutations of the genes encoding insulin-like growth factor I (IGF-1) and type 1 IGF receptor (IGF1R) Cell. 1993;75:59–72. [PubMed] [Google Scholar]
- 34.Pintar JE, Schuller A, Cerro JA, Czick M, Grewal A, Green B. Genetic ablation of IGFBP-2 suggests functional redundancy in the IGFBP family. Prog Growth Factor Res. 1995;6:437–445. doi: 10.1016/0955-2235(95)00029-1. [DOI] [PubMed] [Google Scholar]
- 35.Wood TL, Rogler L, Streck RD, Cerro J, Green B, Grewal A, Pintar JE. Targeted disruption of IGFBP-2 gene. Growth Regul. 1993;3:5–8. [PubMed] [Google Scholar]
- 36.Tiveron MC, Hirsch MR, Brunet JF. The expression pattern of the transcription factor Phox2 delineates synaptic pathways of the autonomic nervous system. J Neurosci. 1996;16:7649–7660. doi: 10.1523/JNEUROSCI.16-23-07649.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hatten ME. Neuronal regulation of astroglial morphology and proliferation in vitro. J Cell Biol. 1985;100:384–396. doi: 10.1083/jcb.100.2.384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Burtrum D, Zhu Z, Lu D, Anderson DM, Prewett M, Pereira DS, Bassi R, Abdullah R, Hooper AT, Koo H, Jimenez X, Johnson D, Apblett R, Kussie P, Bohlen P, Witte L, Hicklin DJ, Ludwig DL. A fully human monoclonal antibody to the insulin-like growth factor I receptor blocks ligand-dependent signaling and inhibits human tumor growth in vivo. Cancer Res. 2003;63:8912–8921. [PubMed] [Google Scholar]
- 39.Romer JT, Kimura H, Magdaleno S, Sasai K, Fuller C, Baines H, Connelly M, Stewart CF, Gould S, Rubin LL, Curran T. Suppression of the Shh pathway using a small molecule inhibitor eliminates medulloblastoma in ptc1(+/–)p53(–/–) mice. Cancer Cell. 2004;6:229–240. doi: 10.1016/j.ccr.2004.08.019. [DOI] [PubMed] [Google Scholar]
- 40.Lee J, Platt KA, Censullo P, Ruiz i Altaba A. Gli1 is a target of sonic hedgehog that induces ventral neural tube development. Development. 1997;124:2537–2552. doi: 10.1242/dev.124.13.2537. [DOI] [PubMed] [Google Scholar]
- 41.Traiffort E, Charytoniuk DA, Faure H, Ruat M. Regional distribution of sonic hedgehog, patched, and smoothened mRNA in the adult rat brain. J Neurochem. 1998;70:1327–1330. doi: 10.1046/j.1471-4159.1998.70031327.x. [DOI] [PubMed] [Google Scholar]
- 42.Bondy C, Lee WH. Correlation between insulin-like growth factor (IGF)-binding protein 5 and IGF-I gene expression during brain development. J Neurosci. 1993;13:5092–5104. doi: 10.1523/JNEUROSCI.13-12-05092.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ingham PW, McMahon AP. Hedgehog signaling in animal development: paradigms and principles. Genes Dev. 2001;15:3059–3087. doi: 10.1101/gad.938601. [DOI] [PubMed] [Google Scholar]
- 44.Chen JK, Taipale J, Cooper MK, Beachy PA. Inhibition of Hedgehog signaling by direct binding of cyclopamine to Smoothened. Genes Dev. 2002;16:2743–2748. doi: 10.1101/gad.1025302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Burtrum D, Zhu Z, Lu D, Anderson DM, Prewett M, Pereira DS, Bassi R, Abdullah R, Hooper AT, Koo H, Jimenez X, Johnson D, Apblett R, Kussie P, Bohlen P, Witte L, Hicklin DJ, Ludwig DL. A fully human monoclonal antibody to the insulin-like growth factor I receptor blocks ligand-dependent signaling and inhibits human tumor growth in vivo. Cancer Res. 2003;63:8912–8921. [PubMed] [Google Scholar]
- 46.Russo VC, Gluckman PD, Feldman EL, Werther GA. The insulin-like growth factor system and its pleiotropic functions in brain. Endocr Rev. 2005;26:916–943. doi: 10.1210/er.2004-0024. [DOI] [PubMed] [Google Scholar]
- 47.D'Ercole AJ, Ye P, Calikoglu AS, Gutierrez-Ospina G. The role of the insulin-like growth factors in the central nervous system. Mol Neurobiol. 1996;13:227–255. doi: 10.1007/BF02740625. [DOI] [PubMed] [Google Scholar]
- 48.Lee WH, Michels KM, Bondy CA. Localization of insulin-like growth factor binding protein-2 messenger RNA during postnatal brain development: correlation with insulin-like growth factors I and II. Neuroscience. 1993;53:251–265. doi: 10.1016/0306-4522(93)90303-w. [DOI] [PubMed] [Google Scholar]
- 49.Naeve GS, Vana AM, Eggold JR, Verge G, Ling N, Foster AC. Expression of rat insulin-like growth factor binding protein-6 in the brain, spinal cord, and sensory ganglia. Brain Res Mol Brain Res. 2000;75:185–197. doi: 10.1016/s0169-328x(99)00262-4. [DOI] [PubMed] [Google Scholar]
- 50.Werther GA, Abate M, Hogg A, Cheesman H, Oldfield B, Hards D, Hudson P, Power B, Freed K, Herington AC. Localization of insulin-like growth factor-I mRNA in rat brain by in situ hybridization – relationship to IGF-I receptors. Mol Endocrinol. 1990;4:773–778. doi: 10.1210/mend-4-5-773. [DOI] [PubMed] [Google Scholar]
- 51.Stylianopoulou F, Herbert J, Soares MB, Efstratiadis A. Expression of the insulin-like growth factor II gene in the choroid plexus and the leptomeninges of the adult rat central nervous system. Proc Natl Acad Sci USA. 1988;85:141–145. doi: 10.1073/pnas.85.1.141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Song RX, Barnes CJ, Zhang Z, Bao Y, Kumar R, Santen RJ. The role of SHC and insulin-like growth factor 1 receptor in mediating the translocation of estrogen receptor α to the plasma membrane. Proc Natl Acad Sci USA. 2004;101:2076–2081. doi: 10.1073/pnas.0308334100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kenney AM, Widlund HR, Rowitch DH. Hedgehog and PI-3 kinase signaling converge on NMYC1 to promote cell cycle progression in cerebellar neuronal precursors. Development. 2004;131:217–228. doi: 10.1242/dev.00891. [DOI] [PubMed] [Google Scholar]
- 54.Kenney AM, Cole MD, Rowitch DH. NMYC upregulation by sonic hedgehog signaling promotes proliferation in developing cerebellar granule neuron precursors. Development. 2003;130:15–28. doi: 10.1242/dev.00182. [DOI] [PubMed] [Google Scholar]
- 55.Allan GJ, Zannoni A, McKinnell I, Otto WR, Holzenberger M, Flint DJ, Patel K. Major components of the insulin-like growth factor axis are expressed early in chicken embryogenesis, with IGF-binding protein (IGFBP)-5 expression subject to regulation by sonic hedgehog. Anat Embryol (Berl) 2003;207:73–84. doi: 10.1007/s00429-003-0321-x. [DOI] [PubMed] [Google Scholar]
- 56.Leal SM, Liu Q, Huang SS, Huang JS. The type V transforming growth factor β receptor is the putative insulin-like growth factor-binding protein 3 receptor. J Biol Chem. 1997;272:20572–20576. doi: 10.1074/jbc.272.33.20572. [DOI] [PubMed] [Google Scholar]
- 57.Amaar YG, Thompson GR, Linkhart TA, Chen ST, Baylink DJ, Mohan S. Insulin-like growth factor-binding protein 5 (IGFBP-5) interacts with a 4 and a half LIM protein 2 (FHL2) J Biol Chem. 2002;277:12053–12060. doi: 10.1074/jbc.M110872200. [DOI] [PubMed] [Google Scholar]
- 58.Oldham S, Hafen E. Insulin/IGF and target of rapamycin signaling: a TOR de force in growth control. Trends Cell Biol. 2003;13:79–85. doi: 10.1016/s0962-8924(02)00042-9. [DOI] [PubMed] [Google Scholar]
- 59.Milenkovic L, Goodrich LV, Higgins KM, Scott MP. Mouse patched1 controls body size determination and limb patterning. Development. 1999;126:4431–4440. doi: 10.1242/dev.126.20.4431. [DOI] [PubMed] [Google Scholar]
- 60.Ruiz i Altaba A, Palma V, Dahmane N. Hedgehog-Gli signalling and the growth of the brain. Nat Rev Neurosci. 2002;3:24–33. doi: 10.1038/nrn704. [DOI] [PubMed] [Google Scholar]
- 61.Corrales JD, Blaess S, Mahoney EM, Joyner AL. The level of sonic hedgehog signaling regulates the complexity of cerebellar foliation. Development. 2006;133:1811–1821. doi: 10.1242/dev.02351. [DOI] [PubMed] [Google Scholar]
- 62.Goodrich LV, Milenkovic L, Higgins KM, Scott MP. Altered neural cell fates and medulloblastoma in mouse patched mutants. Science. 1997;277:1109–1113. doi: 10.1126/science.277.5329.1109. [DOI] [PubMed] [Google Scholar]
- 63.Hahn H, Wicking C, Zaphiropoulous PG, Gailani MR, Shanley S, Chidambaram A, Vorechovsky I, Holmberg E, Unden AB, Gillies S, Negus K, Smyth I, Pressman C, Leffell DJ, Gerrard B, Goldstein AM, Dean M, Toftgard R, Chenevix-Trench G, Wainwright B, Bale AE. Mutations of the human homolog of drosophila patched in the nevoid basal cell carcinoma syndrome. Cell. 1996;85:841–851. doi: 10.1016/s0092-8674(00)81268-4. [DOI] [PubMed] [Google Scholar]
- 64.Johnson RL, Rothman AL, Xie J, Goodrich LV, Bare JW, Bonifas JM, Quinn AG, Myers RM, Cox DR, Epstein EH, Jr, Scott MP. Human homolog of patched, a candidate gene for the basal cell nevus syndrome. Science. 1996;272:1668–1671. doi: 10.1126/science.272.5268.1668. [DOI] [PubMed] [Google Scholar]
- 65.Oliver TG, Read TA, Kessler JD, Mehmeti A, Wells JF, Huynh TT, Lin SM, Wechsler-Reya RJ. Loss of patched and disruption of granule cell development in a pre-neoplastic stage of medulloblastoma. Development. 2005;132:2425–2439. doi: 10.1242/dev.01793. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Igfbp2 expression in postnatal cerebellum. Sagittal sections of mouse postnatal P1 cerebellum were hybridized with RNA probe corresponding to full-length mouse Igfbp2. Igfbp2 expression is detected in the EGL, in cells within the PL, in cells in the IGL and in meninges.
Igfbp5 expression does not colocalize with Calbindin expression. Sagittal sections of mouse postnatal P1 cerebellum were hybridized with an antisense RNA probe corresponding to full-length mouse Igfbp5 and incubated with the anti-Calbindin antibody. Igfbp5 expression (purple) is detected in the EGL and in scattered cells within or around the Purkinje cell layer. The igfbp5 expression in the isolated cells does not colocalize with Calbindin expressing cells (brown) which correspond to Purkinje neurons (PN).