Abstract
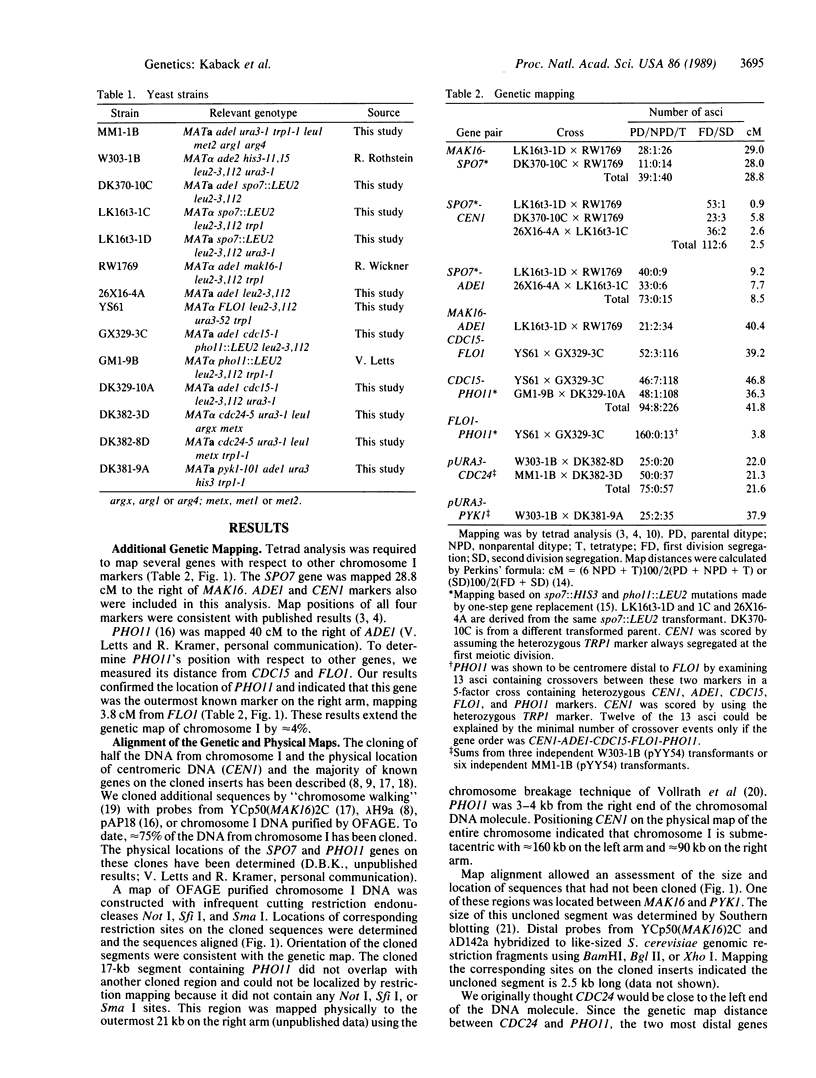
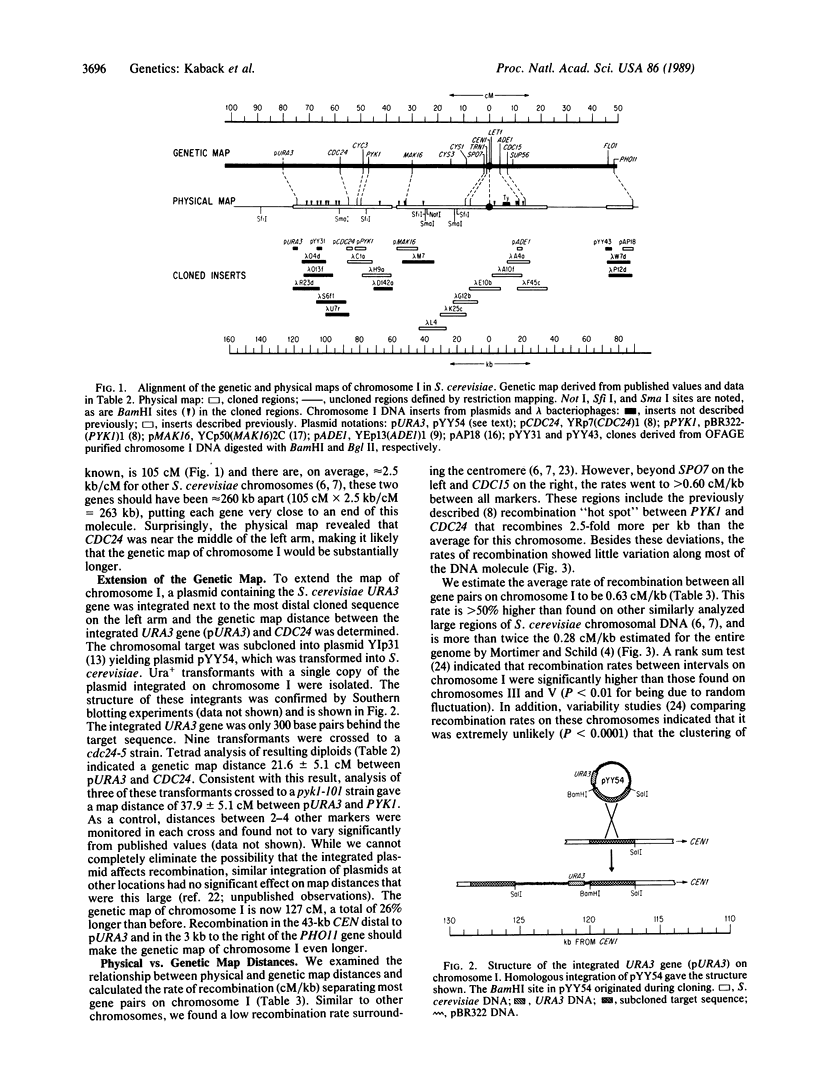
Chromosome I is the smallest chromosome in Saccharomyces cerevisiae and contains a DNA molecule that is only 250 kilobases (kb). Approximately 75% of this DNA molecule has been cloned. A restriction map for the entire DNA molecule from chromosome I was determined and most of its genetically mapped genes were located on this physical map. Based on the average rate of recombination (centimorgans/kb) found for other S. cerevisiae chromosomes, the outermost markers on the genetic map of chromosome I were expected to be close to the ends of the DNA molecule. While the rightmost genetic marker was 3 kb from the end, the leftmost marker, CDC24, was located near the middle of the left arm, suggesting that the genetic map would be much longer. To extend the genetic map, a copy of the S. cerevisiae URA3 gene was integrated in the outermost cloned region located 32 kb centromere distal to CDC24, and the genetic map distance between these two genes was determined. The new marker substantially increased the genetic map length of chromosome I. In addition, we determined the relationship between physical and genetic map distance along most of the length of the chromosome. Consistent with the longer genetic map, the average rate of recombination between markers on chromosome I was greater than 50% higher than the average found on other yeast chromosomes. Owing to its small size, it had been estimated that approximately 5% of the chromosome I homologues failed to undergo meiotic recombination. New measurements of the zero-crossover class indicated that the enhanced rate of recombination ensures at least one genetic exchange between virtually every pair of chromosome I homologues.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andersen N., Thill G. P., Kramer R. A. RNA and homology mapping of two DNA fragments with repressible acid phosphatase genes from Saccharomyces cerevisiae. Mol Cell Biol. 1983 Apr;3(4):562–569. doi: 10.1128/mcb.3.4.562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker B. S., Carpenter A. T., Esposito M. S., Esposito R. E., Sandler L. The genetic control of meiosis. Annu Rev Genet. 1976;10:53–134. doi: 10.1146/annurev.ge.10.120176.000413. [DOI] [PubMed] [Google Scholar]
- Bender W., Spierer P., Hogness D. S. Chromosomal walking and jumping to isolate DNA from the Ace and rosy loci and the bithorax complex in Drosophila melanogaster. J Mol Biol. 1983 Jul 25;168(1):17–33. doi: 10.1016/s0022-2836(83)80320-9. [DOI] [PubMed] [Google Scholar]
- Carle G. F., Olson M. V. An electrophoretic karyotype for yeast. Proc Natl Acad Sci U S A. 1985 Jun;82(11):3756–3760. doi: 10.1073/pnas.82.11.3756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coleman K. G., Steensma H. Y., Kaback D. B., Pringle J. R. Molecular cloning of chromosome I DNA from Saccharomyces cerevisiae: isolation and characterization of the CDC24 gene and adjacent regions of the chromosome. Mol Cell Biol. 1986 Dec;6(12):4516–4525. doi: 10.1128/mcb.6.12.4516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooke H. J., Brown W. R., Rappold G. A. Hypervariable telomeric sequences from the human sex chromosomes are pseudoautosomal. Nature. 1985 Oct 24;317(6039):687–692. doi: 10.1038/317687a0. [DOI] [PubMed] [Google Scholar]
- Cummins C. M., Culbertson M. R., Knapp G. Frameshift suppressor mutations outside the anticodon in yeast proline tRNAs containing an intervening sequence. Mol Cell Biol. 1985 Jul;5(7):1760–1771. doi: 10.1128/mcb.5.7.1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dawson D. S., Murray A. W., Szostak J. W. An alternative pathway for meiotic chromosome segregation in yeast. Science. 1986 Nov 7;234(4777):713–717. doi: 10.1126/science.3535068. [DOI] [PubMed] [Google Scholar]
- Jackson J. A., Fink G. R. Meiotic recombination between duplicated genetic elements in Saccharomyces cerevisiae. Genetics. 1985 Feb;109(2):303–332. doi: 10.1093/genetics/109.2.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lambie E. J., Roeder G. S. Repression of meiotic crossing over by a centromere (CEN3) in Saccharomyces cerevisiae. Genetics. 1986 Nov;114(3):769–789. doi: 10.1093/genetics/114.3.769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mann C., Davis R. W. Meiotic disjunction of circular minichromosomes in yeast does not require DNA homology. Proc Natl Acad Sci U S A. 1986 Aug;83(16):6017–6019. doi: 10.1073/pnas.83.16.6017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mortimer R. K., Schild D. Genetic map of Saccharomyces cerevisiae, edition 9. Microbiol Rev. 1985 Sep;49(3):181–213. doi: 10.1128/mr.49.3.181-213.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mortimer R. K., Schild D. Genetic map of Saccharomyces cerevisiae. Microbiol Rev. 1980 Dec;44(4):519–571. doi: 10.1128/mr.44.4.519-571.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olson M. V., Dutchik J. E., Graham M. Y., Brodeur G. M., Helms C., Frank M., MacCollin M., Scheinman R., Frank T. Random-clone strategy for genomic restriction mapping in yeast. Proc Natl Acad Sci U S A. 1986 Oct;83(20):7826–7830. doi: 10.1073/pnas.83.20.7826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parent S. A., Fenimore C. M., Bostian K. A. Vector systems for the expression, analysis and cloning of DNA sequences in S. cerevisiae. Yeast. 1985 Dec;1(2):83–138. doi: 10.1002/yea.320010202. [DOI] [PubMed] [Google Scholar]
- Perkins D. D. Biochemical Mutants in the Smut Fungus Ustilago Maydis. Genetics. 1949 Sep;34(5):607–626. doi: 10.1093/genetics/34.5.607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Steensma H. Y., Crowley J. C., Kaback D. B. Molecular cloning of chromosome I DNA from Saccharomyces cerevisiae: isolation and analysis of the CEN1-ADE1-CDC15 region. Mol Cell Biol. 1987 Jan;7(1):410–419. doi: 10.1128/mcb.7.1.410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strathern J. N., Newlon C. S., Herskowitz I., Hicks J. B. Isolation of a circular derivative of yeast chromosome III: implications for the mechanism of mating type interconversion. Cell. 1979 Oct;18(2):309–319. doi: 10.1016/0092-8674(79)90050-3. [DOI] [PubMed] [Google Scholar]
- Vollrath D., Davis R. W., Connelly C., Hieter P. Physical mapping of large DNA by chromosome fragmentation. Proc Natl Acad Sci U S A. 1988 Aug;85(16):6027–6031. doi: 10.1073/pnas.85.16.6027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickner R. B., Koh T. J., Crowley J. C., O'Neil J., Kaback D. B. Molecular cloning of chromosome I DNA from Saccharomyces cerevisiae: isolation of the MAK16 gene and analysis of an adjacent gene essential for growth at low temperatures. Yeast. 1987 Mar;3(1):51–57. doi: 10.1002/yea.320030108. [DOI] [PubMed] [Google Scholar]
- de Jonge P., de Jongh F. C., Meijers R., Steensma H. Y., Scheffers W. A. Orthogonal-field-alternation gel electrophoresis banding patterns of DNA from yeasts. Yeast. 1986 Sep;2(3):193–204. doi: 10.1002/yea.320020307. [DOI] [PubMed] [Google Scholar]



