Abstract
Humanized mice that express the human UDP-glucuronosyltransferase (UGT) 1 locus have been developed in a Ugt1-null background as a model to improve predictions of human UGT1A-dependent drug clearance. Enzyme kinetic parameters (Km and Vmax) and pharmacokinetic properties of three probe drugs were compared using wild-type and humanized UGT1 mice that express the Gilbert’s UGT1A1*28 allele [Tg(UGT1A1*28) Ugt1(−/−) mice]. The well characterized substrate for UGT1A1, 7-ethyl-10-hydroxy-camptothecin (SN-38), showed the greatest difference in parent drug exposure (∼3-fold increase) and clearance (∼3-fold decrease) in Tg(UGT1A1*28) Ugt1(−/−) mice after intravenous administration compared with wild-type and phenobarbital-treated animals. In contrast, the clearance of the UGT2B7 substrate (−)-17-allyl-4, 5α-epoxy-3, 14-dihydroxymorphinan-6-one (naloxone) was not altered in Tg(UGT1A1*28) Ugt1(−/−) mice. In addition, pharmacokinetic parameters with 1-(4-fluorophenyl)3(R)-[3-(4-fluorophenyl)-3(S)-hydroxypropyl]-4(S)-(4-hydroxyphenyl)-2-azetidinone (ezetimibe, Zetia; Merck & Co., Whitehouse Station, NJ), considered to be a major substrate for UGT1A1, showed small to no dependence on UGT1A1-directed glucuronidation. Enzyme kinetic parameters assessed for SN-38, ezetimibe, and naloxone using liver microsomes prepared from wild-type and Tg(UGT1A1*28) Ugt1(−/−) mice showed patterns consistent with the in vivo pharmacokinetic data. For SN-38 glucuronidation, Vmax decreased 5-fold in Tg(UGT1A1*28) Ugt1(−/−) mouse liver microsomes compared with microsomes prepared from wild-type mice, and decreased 10-fold compared with phenobarbital-treated Tg(UGT1A1*28) Ugt1(−/−) mice. These differences are consistent with SN-38 glucuronidation activities using HLMs isolated from individuals genotyped as UGT1A1*1 or UGT1A1*28. For ezetimibe and naloxone the differences in Vmax were minimal. Thus, Tg(UGT1A1*28) Ugt1(−/−) mice can serve as a pharmacokinetic model to further investigate the effects of UGT1A1 expression on drug metabolism.
UDP-glucuronosyltransferases (UGTs) exist as an enzyme “superfamily” and are quantitatively among the most important drug-metabolizing enzymes. Together with cytochrome P450 (P450), they are responsible for the elimination of more than 90% of hepatically cleared drugs (Miners et al., 2004). Within the UGT superfamily, the nine UGTs encoded by the UGT1 locus in humans are responsible for the conjugation of most exogenous compounds (such as drugs, environmental toxicants, and carcinogens) and endogenous substances (such as bile acids, fatty acids, steroids, hormones, and bilirubin) (Tukey and Strassburg, 2000; Miners et al., 2002). They exhibit distinct, but often overlapping, substrate and inhibitor selectivity and differ in the occurrence and frequency of genetic polymorphism and regulation (Tukey and Strassburg, 2000; Miners et al., 2002). Of particular interest among the UGT1A subfamily is UGT1A1 (Bosma, 2003). This enzyme is solely responsible for the glucuronidation of bilirubin and contributes to the conjugation of exogenous and endogenous substances, including SN-38, the active metabolite of irinotecan (CPT-11) (Iyer et al., 1998; Strassburg et al., 2000; Bosma, 2003).
Numerous polymorphisms in the UGT1A1 gene have been identified (Udomuksorn et al., 2007). A common variant with functional consequence is the UGT1A1*28 allele, resulting from a (TA) insertion into the UGT1A1 promoter region. The UGT1A1*28 genotype has been linked to Gilbert's syndrome, and these individuals have been reported to experience episodes of mild hyperbilirubinemia. In addition, this population may have altered pharmacokinetic drug profiles and, more importantly, are susceptible to the dose/exposure-limited toxicities exemplified by the anticancer drug CPT-11 (Iyer et al., 2002; Tukey et al., 2002; Nagar and Blanchard, 2006). Even though the underlying mechanism for CPT-11 toxicity is yet to be determined, it is linked to the decreased hepatic UGT1A1 activity observed in Gilbert's syndrome, leading to reduced systemic clearance and consequently sustained elevated levels of SN-38, the active form of CPT-11 (Iyer et al., 1998, 1999; Gagné et al., 2002). Human liver microsomes (HLMs), isolated from individuals with the UGT1A1*28 polymorphism, show a decrease in catalytic activity toward SN-38 and other UGT1A1 substrates. Thus, genotyping for the UGT1A1*28 allele is recommended along with appropriate dose adjustments for CPT-11 treatment (O'Dwyer and Catalano, 2006; Ando et al., 2007).
Despite the fact that the UGT1A1*28 allele represents a common polymorphism and has been shown to lead to a clinically relevant phenotype, there are few tools to assess and predict whether lower UGT1A1 expression will affect the overall clearance of a new chemical entity. This is in part because of the lack of tools for UGT reaction phenotyping, such as specific chemical substrates, chemical and antibody inhibitors, and well characterized expressed enzymes (Miners et al., 2010). Currently, the most widely used and effective tool for attributing substrate specificity to a specific UGT is expressed UGT enzymes (Ethell et al., 2001). However, the utility of an expressed enzyme can be complicated by several factors shown recently to alter the kinetics of glucuronidation. These include, but are not limited to, bovine serum albumin effects (Miners et al., 2006; Rowland et al., 2008), UGT binary complex formation (Fujiwara et al., 2007), coexpression of multiple UGTs (Fujiwara et al., 2007), and coexpression of UGTs with CYP3A4 (Takeda et al., 2005; Ishii et al., 2007). Alternatively, a more laborious approach, with limited utility because of the lack of specific substrates for a given UGT isozyme, is the correlation analysis of an activity known to be specific for a single UGT with the glucuronidation of an unknown compound across an array of individual HLMs (Court, 2005). However, caution should be taken when drawing any conclusions from these data without any confirmatory data obtained from other methods.
An alternative approach to assess the substrate potential of a new chemical entity for UGT1A1, and thus potentially assess the likelihood for altered pharmacokinetics in individuals with the UGT1A1*28 genotype, is the implementation of humanized mice. In this report, humanized mice that carry the UGT1A1*28 allele and have been shown to duplicate the hyperbilirubinemia condition present in Gilbert's syndrome were evaluated for the first time as a model for assessing UGT1A1-related clearance and metabolism. Thus, three literature compounds were chosen for this research: SN-38, an exclusive UGT1A1 substrate (Iyer et al., 1998); ezetimibe, a partial UGT1A1 substrate (Ghosal et al., 2004); and naloxone, a UGT2B7 substrate. Pharmacokinetic and enzyme kinetic parameters for SN-38, ezetimibe, and naloxone in wild-type and humanized UGT1 mice were compared. The data are discussed in the context of evaluating the suitability of the humanized mouse model for assessing UGT1A1-related clearance and metabolism for humans.
Materials and Methods
Materials.
Male C57BL/6 mice were purchased from The Jackson Laboratory (Bar Harbor, ME). HLMs were purchased from BD Gentest (Woburn, MA). SN-38 and ezetimibe were synthesized internally by Pfizer Global Research and Development laboratories. Naloxone was purchased from Sigma-Aldrich (St. Louis, MO). Phenobarbital was purchased from Henry Schein (Melville, NY).
Generation of Humanized UGT1 Mice.
Transgenic mice that express the human UGT1 locus (Tg-UGT1) have been reported (Chen et al., 2005), and these studies have shown the expression of the UGT1A genes in humanized UGT1 mice are identical to those shown for expression of the UGT1 locus in Tg-UGT1 mice. In addition, these patterns closely resemble the expression patterns observed in human tissues. DNA sequence analysis of the UGT1A1 promoter that is expressed in Tg-UGT1 mice contains an extra (TA) insertion in the TATAA element [A(TA)7TAA] that identifies the gene as the human UGT1A1*28 allele. Breeding experiments with Tg-UGT1 mice and Ugt1(+/−) mice (Nguyen et al., 2008) led to a cross that expressed the UGT1 and Ugt1(+/−) alleles, which when backcrossed produced mice that were transgenic for the UGT1 locus in a Ugt1(−/−) background. Because the UGT1 locus encodes the nine UGT1A genes, including the UGT1A1*28 allele, these mice were identified as being humanized for the UGT1A1*28 gene and are designated as Tg(UGT1A1*28)Ugt1(−/−) mice.
Microsomal Enzyme Kinetics.
Liver microsomes from wild-type (C57BL/6) and Tg(UGT1A1*28)Ugt1(−/−) mice were prepared as described previously (Chen et al., 2005). Using either HLMs or mouse liver microsomes (MLMs), the following procedure was used. Incubations, in triplicate, containing 50 mM Tris, pH 7.4, 0.25 mg/ml microsomal protein, 5 mM MgCl2, 50 μg/ml alamethicin, and 5 mM UDP-glucuronic acid in a final concentration of 200 μl were conducted in 96-deep well plates at 37°C for 30 min. The reactions were stopped by the addition 100 μl of cold 0.1% formic acid in acetonitrile containing 0.25 μM carbamazepine as the internal standard. The samples were briefly centrifuged, and the supernatants were analyzed using liquid chromatography/tandem mass spectrometry (LC/MS/MS). For substrate concentration-dependent glucuronidation activities of selected HLMs (donor designations of HH9 and HH81, each genotyped as *28/*28; and donors HH83 and HH112, each genotyped as *1/*1) and MLMs, the substrate concentrations were 0.49 to 100 μM for ezetimibe, 0.23 to 30 μM for SN-38, and 7.8 to 1000 μM for naloxone.
LC/MS/MS Conditions and Data Analysis for Enzyme Kinetics.
No glucuronide standards were used for quantitation. Rather, quantitation was accomplished by using SN-38, ezetimibe, or naloxone as a standard and assuming similar ionization between parent and glucuronide. Samples were analyzed by LC/MS/MS in positive mode using a Thermo Fisher Scientific (Waltham, MA) AQUASIL C18 2.1 × 20-mm, 3-μm column at a flow rate of 0.5 ml/min (A, 0.1% formic acid in water; B, 0.1% formic acid in acetonitrile). Initial conditions were 5% B at 0.5 ml/min followed by a linear gradient from 5% to 95% B in 1 min, hold at 95% B for 1 min, and then a 1-min re-equilibration at 5% B. Mass transitions and retention times for the SN-38 glucuronide, ezetimibe glucuronide, naloxone glucuronide, and carbamazepine (internal standard) were 569→393 at 1.06 min, 586→410 at 1.24 min, 504→328 at 0.76 min, and 237→194 at 1.29 min, respectively. Standard curves were fitted to a 1× weighted linear regression model, and correlation coefficients were ≥0.99. The Km and Vmax values of each enzyme were estimated by fitting the glucuronidation activities to the Michaelis-Menten equation using a nonlinear regression analysis in GraphPad Software Inc. (San Diego, CA) Prism 5.00 for Windows (Microsoft, Redmond, WA).
Pharmacokinetic Studies in Mice.
Wild-type and Tg(UGT1A1*28)Ugt1(−/−) mice were single- or double-housed in polycarbonate microisolator cages with contact bedding on arrival. Animal room temperature and humidity were targeted to be between 70 ± 5°F and 30 to 70%, respectively. Lighting was set to maintain an approximate 12-h light/dark cycle. Food (Certified Rodent Diet 5002; Nestle Purina PetCare Co., St. Louis, MO) and water were available ad libitum. Animal housing and husbandry were performed in accordance with the provisions of the Guide for the Care and Use of Laboratory Animals (Institute of Laboratory Animal Resources, 1996). The Pfizer Institutional Animal Care and Use Committee reviewed and approved the animal use in these studies. The animal care and use program is fully accredited by the Association for Assessment and Accreditation of Laboratory Animal Care, International.
The mice were weighed the morning of dosing, and the dose volumes were calculated based on body weights (Table 1). Intravenous dose volumes for the mouse studies were either 4 or 8 ml/kg. Blood was collected into lithium heparin blood tubes at various time points via retro-orbital bleeding under general anesthesia (80% CO2/20% O2). A terminal blood sample was taken from the posterior vena cava after euthanasia using CO2. The blood was separated into plasma by centrifugation and then aliquoted and stored at −80°C until analyzed.
TABLE 1.
Intravenous formulation and time points for pharmacokinetic studies of SN-38, ezetimibe, and naloxone in wild-type and Tg(UGT1A1*28)Ugt1(−/−) mice (n = 2 animals/time point)
| Compounds | Mouse Type | Intravenous Dose | Formulation | Time Point Range |
|---|---|---|---|---|
| mg/kg | ||||
| SN-38 | WT | 1 | 10% Ethanol/70% PEG-400/20% PBS | 5 min–8 h |
| WT | 2 | 10% Ethanol/70% PEG-400/20% PBS | 5 min–3 h | |
| Tg(UGT1A1*28)Ugt1(−/−) | 1 | 10% Ethanol/70% PEG-400/20% PBS | 5–45 min | |
| Tg(UGT1A1*28)Ugt1(−/−) | 2 | 10% Ethanol/70% PEG-400/20% PBS | 2.5 min–3 h | |
| Tg(UGT1A1*28)Ugt1(−/−) with pb treatment | 1 | 10% Ethanol/70% PEG-400/20% water | 3–80 min | |
| Ezetimibe | WT | 1 | 10% Ethanol/50% PEG-400/40% PBS | 5 min–8 h |
| WT | 5 | 10% Ethanol/50% PEG-400/40% PBS | 5 min–3 h | |
| Tg(UGT1A1*28)Ugt(−/−) | 1 | 10% Ethanol/50% PEG-400/40% PBS | 5–45 min | |
| Tg(UGT1A1*28)Ugt1(−/−) | 5 | 10% ethanol/50% PEG-400/40% PBS | 2.5 min–3 h | |
| Naloxone | WT | 5 | 10% NMP/50% PEG-400/40% PBS | 5 min–8 h |
| Tg(UGT1A1*28)Ugt1(−/−) | 5 | 10% NMP/50% PEG-400/40% PBS | 5 min–8 h |
WT, C57BL/6 mouse; PEG, polyethylene glycol; PBS, phosphate-buffered saline; NMP, N-methylpyrrolidone; pb, phenobarbital.
In one study, male humanized mice were dosed via intraperitoneal injection for 4 days with 100 mg/kg/day sodium phenobarbital (12 mg/ml free base equivalent; 8 ml/kg) in sterile water. On the 5th day, the mice received SN-38 via intravenous administration, and blood was collected at various time points as described previously (n = 2 animals per time point for each study). After the terminal bleed (last time point), livers were removed, rinsed in saline, blotted dry, and frozen on dry ice. Livers were stored at −80°C until microsome preparation and Western blot analysis.
Sample Preparation, LC/MS/MS Conditions, and Data Analysis for Mouse Pharmacokinetic Studies.
Mouse plasma samples were prepared using a protein precipitation procedure with acetonitrile containing analytical internal standard (carbamazepine) and analyzed using an LC/MS/MS system composed of an SCL-10A VP Shimadzu (Kyoto, Japan) pump LC system controller, coupled with a triple quadrupole mass spectrometer using multiple reaction monitoring in the positive electrospray ionization mode (API4000; Applied Biosystems/MDS Sciex, Foster City, CA). The separation of the analytes was achieved by using an AQUASIL C18 20 × 2.1-mm (3 μm) (Thermo Fisher Scientific) column. A 3.5-min gradient at 0.5 ml/min flow rate was applied with the following mobile phases: A, 0.1% formic acid in water; B, acetonitrile with 0.1% formic acid. The same linear elution LC gradient was used as described for the in vitro enzyme kinetic experiments. Mass transitions and retention times for SN-38, ezetimibe, naloxone, and internal standard (carbamazepine) were 393.0→249.2 at 1.08 min, 410.1→132.9 at 1.4 min, 328.25→253.1 at 0.95 min, and 237.0→194.0 at 1.28 min, respectively. Quantitation of the analytes was performed using Analyst 1.2 software (Applied Biosystems/MDS Sciex).
Pharmacokinetic calculations were performed using the noncompartmental approach with the aid of Watson LIMS (version 7.2.0.03; Thermo Fisher Scientific). The initial plasma concentration (C0) was determined by linear regression extrapolation from the first two data points from mean animals. The area under the plasma concentration-time curve [AUC(0-t)] was determined using the linear trapezoidal method.
Immunoblot Analysis of Humanized UGT1 Mouse and HLMs.
All the Western blots were performed using NuPAGE BisTris polyacrylamide gels as outlined by the supplier (Invitrogen, Carlsbad, CA). Protein was heated at 70°C for 10 min in loading buffer and resolved in 4 to 12% BisTris gels under denaturing conditions (50 mM MOPS, 50 mM Tris base, pH 7.7, 0.1% SDS, 1 mM EDTA) before transferring the proteins to a polyvinylidene difluoride membrane using a semidry transfer system (Invitrogen). The membrane was blocked with 5% nonfat dry milk in 10 mM Tris-HCl, pH 7.4, containing 0.15 M NaCl and 0.05% Tween 20 (Tris-buffered saline) for 1 h at room temperature, followed by incubation with primary antibodies (mouse anti-human UGT1A1, a gift from Professor Joseph K. Ritter, Department of Pharmacology and Toxicology, Virginia Commonwealth University, Richmond, VA) in Tris-buffered saline overnight at 4°C. Membranes were washed and exposed to horseradish peroxidase-conjugated secondary antibodies for 1 h at room temperature. Each membrane was washed again, and the conjugated horseradish peroxidase was detected using the enhanced chemiluminescence plus Western blotting detection system (GE Healthcare, Waukesha, WI), and the proteins were detected after exposure to X-ray film.
RNA Preparation and Reverse Transcription-Polymerase Chain Reaction Analysis.
Pooled liver tissues were pulverized in liquid nitrogen, and approximately 100 mg of tissue was homogenized into 1 ml of TRIzol (Invitrogen). Two hundred microliters of chloroform was added with vigorous shaking. The solution was centrifuged, and the aqueous phase was transferred to a new tube. The RNA was precipitated down by adding 500 μl of isopropanol, collected by centrifugation, and followed by 75% ethyl alcohol wash. The RNA was dissolved into diethyl pyrocarbonate water, and the concentration was determined. Using Iscript Reverse Transcriptase (Bio-Rad Laboratories, Hercules, CA), 1 μg of total RNA in a volume of 20 μl was used for the generation of cDNA as outlined by the manufacturer. Thereafter, synthesized cDNA was used for amplifications of human UGT1A1, UGT1A3, UGT1A4, UGT1A6, and UGT1A9, as described previously (Chen et al., 2005). In brief, the polymerase chain reaction with 1 μl of cDNA and 0.25 μM of each of the UGT1A-specific oligonucleotide primer pairs was carried out in a total volume of 20 μl by polymerase master mix (Denville Scientific, South Plainfield, NJ). The polymerase was activated at 95°C for 10 min followed by 30 cycles of 95°C for 30 s, 63°C for 30 s, and 72°C for 45 s, and a final extension at 72°C for 7 min. After the reaction, 15 μl of the polymerase chain reaction products was subjected to electrophoresis in 1.5% agarose gels and documented by ChemiDoc XRS gel imaging system (Bio-Rad Laboratories).
Results
Mouse Pharmacokinetic Studies.
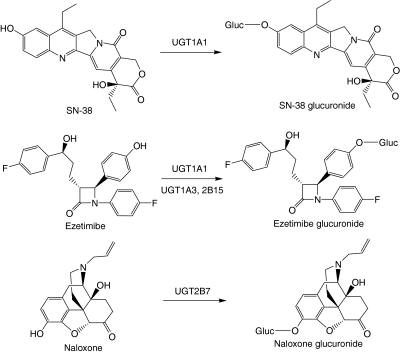
Pharmacokinetic parameters were determined in mice (intravenous route) for three established UGT substrates: SN-38, the active metabolite of CPT-11 that is cleared predominantly by UGT1A1; ezetimibe (Zetia), which is believed to form the phenolic glucuronide by UGT1A1, UGT1A3, and UGT2B15 (ezetimibe also forms a very minor benzylic glucuronide by UGT2B7, which is not monitored in this report); and naloxone, which is a UGT2B7 substrate (Fig. 1). In the case of SN-38, at the doses tested, no adverse effect was seen with either wild-type C57BL/6 or Tg(UGT11A*28)Ugt1(−/−) mice. The pharmacokinetic results are summarized in Table 2 and displayed in Fig. 2.
Fig. 1.
Glucuronidation reactions investigated in this study.
TABLE 2.
Pharmacokinetic parameters of SN-38, ezetimibe, and naloxone in wild-type C57BL/6 and Tg(UGT1A1*28)Ugt1(−/−) mice (dosed intravenously, n = 2 mice/time points; values in parentheses represent individual determinations)
| Compounds | Mouse Type | Intravenous Dose | AUC | CL |
|---|---|---|---|---|
| mg/kg | μM · h | ml/min/kg | ||
| SN-38 | WT | 1 | 0.14 (0.15, 0.12) | 320 (276, 363) |
| WT | 2 | 0.37 (0.34, 0.39) | 230 (241, 219) | |
| Humanized | 1 | 0.54 (0.57, 0.50) | 80 (74, 86) | |
| Humanized | 2 | 0.87 (0.79, 0.95) | 96 (100, 92) | |
| Humanized with pb treatment | 1 | 0.15 (0.18, 0.11) | 285 (240, 330) | |
| Ezetimibe | WT | 1 | 0.30 (0.31, 0.29) | 136 (131, 141) |
| WT | 5 | 1.7 (1.67, 1.74) | 120 (122, 117) | |
| Humanized | 1 | 0.31 (0.28, 0.35) | 132 (145, 118) | |
| Humanized | 5 | 3.3 (3.60, 3.09) | 63 (59, 66) | |
| Naloxone | WT | 5 | 1.3 (1.2, 1.4) | 194 (197, 191) |
| Humanized | 5 | 1.2 (1.3, 1.1) | 210 (199, 221) |
CL, clearance; WT, C57BL/6 mouse; pb, phenobarbital.
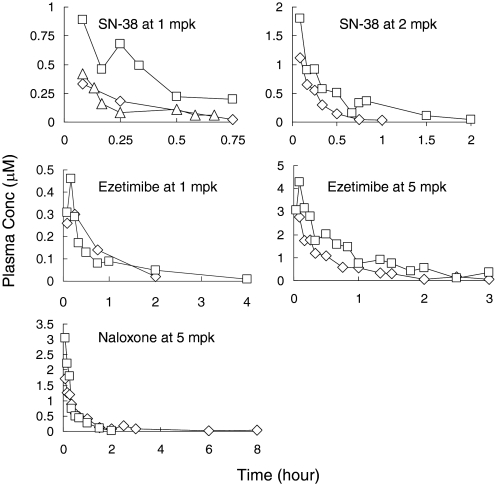
Fig. 2.
Time courses of SN-38, ezetimibe, and naloxone in mice dosed intravenously (n of 2 mice/time point, and each data point shown represents the average of two determinations). Diamonds, wild-type C57BL/6 mice; squares, humanized UGT1 mice; triangles, humanized UGT1 mice with phenobarbital treatment.
Wild-Type C57BL/6 Mouse Studies.
All three compounds were rapidly cleared in wild-type mice at the dose levels evaluated. For the 1 mg/kg i.v. dose, SN-38 exhibited a clearance of 320 ml/min/kg and an AUC of 0.14 μM·h, whereas at the 2 mg/kg i.v. dose, the clearance was lower and the exposure was higher (clearance of 230 ml/min/kg and an AUC of 0.37 μM·h, respectively). For ezetimibe, 1 mg/kg i.v. yielded a clearance value of 136 ml/min/kg and an AUC of 0.30 μM·h. At higher doses (5 mg/kg), clearance was constant, whereas the AUC increased approximately 5-fold. Naloxone, at 5 mg/kg i.v., had a high clearance of 194 ml/min/kg and an AUC of 1.3 μM·h.
Humanized UGT1 Mouse Studies.
The development of Tg(UGT1A1*28)Ugt1(−/−) mice leads to minimal expression of hepatic UGT1A1 (Fig. 3). In 8-week-old mice, the serum bilirubin levels are mildly elevated compared with same-aged wild-type mice, a finding that links the TATAA promoter polymorphism in the UGT1A1*28 gene to the observed hyperbilirubinemia. Thus, drugs that enter the circulation and become targeted for UGT1A1-dependent glucuronidation would be expected to show differences in clearance and AUC. For SN-38, the AUC increased from 0.54 to 0.87 μM·h as the dose increased from 1 to 2 mg/kg, whereas clearance was fairly consistent (80 versus 96 ml/min/kg). When the mice were treated with phenobarbital before administration of SN-38, hepatic UGT1A1 expression was greatly induced (Fig. 3). In addition, at the gene level, all the UGT1A genes expressed in human liver (except for UGT1A6, which was only mildly induced) were significantly induced in these mice after phenobarbital treatment (Fig. 4). Consistent with the induction of UGT1A1, clearance of SN-38 in phenobarbital-treated mice increased (96–285 ml/min/kg), an observation that is concordant with a reduction in AUC (0.87 to 0.15 μM·h).
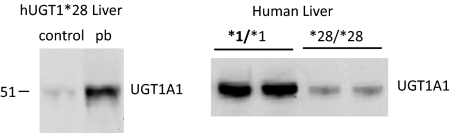
Fig. 3.
Protein expression of UGT1A1 in Tg(UGT1A1*28)Ugt1(−/−) male MLMs and HLMs. Mice were treated with phenobarbital at 100 mg/kg/day for 4 days. Left, control = Tg(UGT1A1*28)Ugt1(−/−) mice without phenobarbital treatment; pb, Tg(UGT1A1*28)Ugt1(−/−) mice with phenobarbital treatment. HLMs have been genotyped for expression of the UGT1A1*28 or UGT1A1*1 alleles.
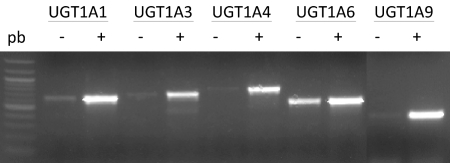
Fig. 4.
Human UGT1A gene expression in Tg(UGT1A1*28)Ugt1(−/−) mice treated with phenobarbital. Six age-matched humanized Tg(UGT1A1*28)Ugt1(−/−) mice were treated intraperitoneally with either dimethyl sulfoxide (control) or phenobarbital (100 mg/kg) for 4 consecutive days. The day after the last dose the mice were sacrificed, and liver tissues were collected and pooled (n = 3). Liver total RNA was prepared, followed by reverse transcription-polymerase chain reaction to determine the expression levels of human UGT1A1, UGT1A3, UGTA4, UGT1A6, and UGT1A9.
When we examined the pharmacokinetic parameters of ezetimibe at a 1-mg/kg dose, there was very little difference in the pharmacokinetic parameters when comparing Tg(UGT1A1*28)Ugt1(−/−) mice with wild-type mice. At a higher dose of ezetimibe (5 mg/kg), small differences (approximately 2-fold) in pharmacokinetic parameters were seen between Tg(UGT1A1*28)Ugt1(−/−) and wild-type mice. Finally, naloxone pharmacokinetics were evaluated, and the AUC and clearance values were nearly identical when comparing results between wild-type and Tg(UGT1A1*28)Ugt1(−/−) mice.
Enzyme Kinetics.
The kinetics of glucuronidation were evaluated for the three compounds by monitoring the formation of glucuronide metabolites using liver microsomes. Each compound was investigated using either HLMs genotyped for the normal (UGT1A1*1) and the Gilbert's (UGT1A1*28) allele, in addition to MLMs prepared from untreated wild-type mice, Tg(UGT1A1*28)Ugt1(−/−) mice, and phenobarbital-treated Tg(UGT1A1*28)Ugt1(−/−) mice. In all cases, parent (not glucuronide) standards were used for quantitation. The enzyme kinetic parameters with HLMs and the various MLMs are reported in Table 3. Individual data and data fitting are displayed in Fig. 5 (for HLM) and Fig. 6 (for MLM).
TABLE 3.
Kinetic parameters of glucuronidation from incubations with individual HLMs and pooled MLMs (minimal n = 3 animals and triplicate determinations per data point)
| Liver Microsomes | SN-38 |
Ezetimibe |
Naloxone |
|||
|---|---|---|---|---|---|---|
| Km | Vmax | Km | Vmax | Km | Vmax | |
| μM | pmol/min·mg | μM | pmol/min·mg | μM | pmol/min·mg | |
| Human liver microsomes | ||||||
| HH83 (WT) | 0.964 ± 0.274 | 162 ± 10.9 | 5.18 ± 2.28 | 781 ± 85.9 | 58.5 ± 6.85 | 1593 ± 50.1 |
| HH112 (WT) | 0.680 ± 0.217 | 88.2 ± 6.73 | 14.7 ± 4.43 | 1515 ± 146 | 54.6 ± 6.49 | 2059 ± 67.6 |
| HH9 (*28/*28) | 1.46 ± 0.637 | 21.0 ± 2.37 | 16.9 ± 3.62 | 1619 ± 116 | 34.5 ± 6.66 | 1882 ± 91.3 |
| HH81 (*28/*28) | 0.635 ± 0.253 | 17.6 ± 1.51 | 11.5 ± 3.47 | 1330 ± 120 | 63.1 ± 15.7 | 1986 ± 149 |
| Mouse liver microsomes | ||||||
| WT C57BL/6 mouse | 3.59 ± 1.21 | 201 ± 21.5 | 15.0 ± 3.11 | 859 ± 64.5 | 278 ± 23.6 | 1635 ± 55.7 |
| Tg(UGT1A1*28)Ugt1(−/−) | 2.42 ± 1.23 | 43.6 ± 6.36 | 35.6 ± 2.33 | 1093 ± 31.7 | 363 ± 39.3 | 1787 ± 84.1 |
| Tg(UGT1A1*28)Ugt1(−/−) with pb treatment | 1.44 ± 0.359 | 497 ± 28.8 | 20.0 ± 3.31 | 1025 ± 65.8 | 432 ± 31.6 | 1723 ± 58.1 |
WT, C57BL/6 mouse; pb, phenobarbital.
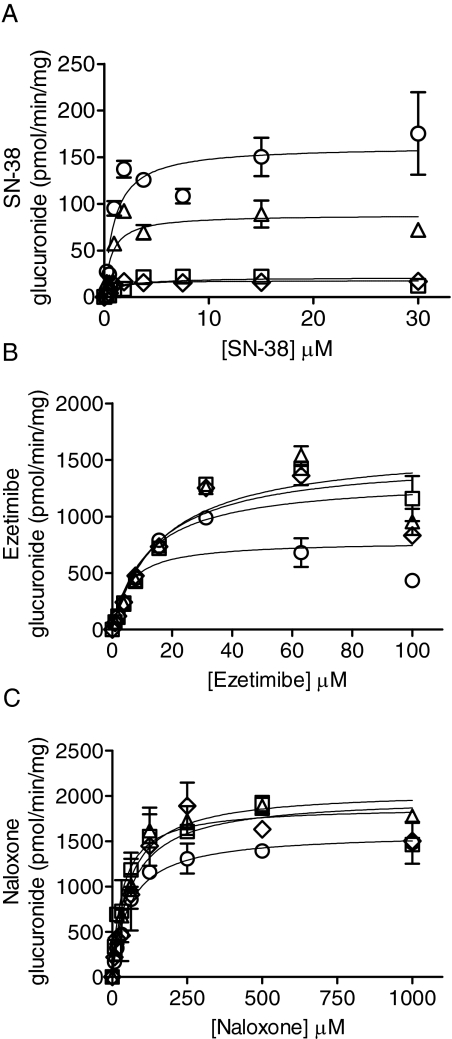
Fig. 5.
Substrate concentration-dependent glucuronidation of SN-38 (A), ezetimibe (B), and naloxone (C) by HLM genotyped as UGT1A1*28/*28 (HH9 and HH81) and UGT1A1*1/*1 (HH83 and HH112) (n of 3 replicates/data points). Open circles, HH83; open triangles, HH112; open squares, HH9; open diamonds, HH81.
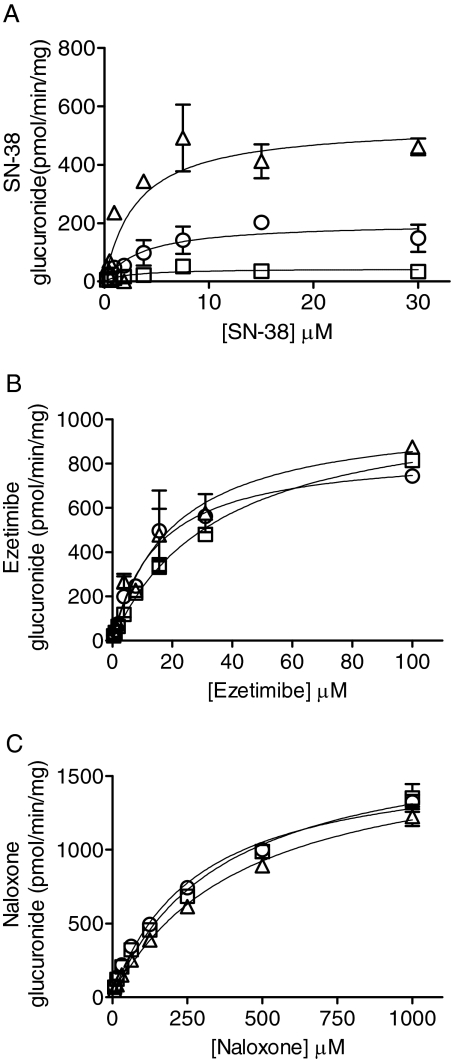
Fig. 6.
Substrate concentration-dependent glucuronidation of SN-38 (A), ezetimibe (B), and naloxone (C) by MLMs prepared from wild-type C57BL/6 mice (open circles), Tg(UGT1A1*28)Ugt1(−/−) mice without phenobarbital treatment (open squares), and with phenobarbital treatment (open triangles) (n of 3 replicates/data points using pooled MLMs prepared from a minimum of three animals).
Enzyme Kinetics with HLM.
Four individual lots of HLM were obtained commercially from BD Gentest. They were genotyped in-house, and Western blot analysis was also performed (Fig. 3) to ensure relative UGT1A1 protein content in individual HLM was consistent with genotype (Fang and Lazarus, 2004; Peterkin et al., 2007). This figure clearly shows a dramatic reduction of UGT1A1 protein in each of the microsomal preparations of UGT1A1*28 origin. For the kinetic analysis with SN-38, the Km ranged from 0.635 to 1.46 μM, whereas Vmax decreased 4- to 8-fold between HLM genotyped for the UGT1A1*1 and UGT1A1*28 alleles (Table 3).
For ezetimibe, Km ranged from 5.18 to 16.9 μM and Vmax from 781 to 1619 pmol/min·mg with UGT1A1*1 and UGT1A1*28 HLM, respectively. In the case of naloxone, the observed differences in Km (ranging 34.5–63.1 μM) or Vmax (ranging 1593–2059 pmol/min·mg) were minimal, as expected.
Enzyme Kinetics with MLM.
The enzyme kinetic results using MLMs prepared from wild-type and Tg(UGT1A1*28)Ugt1(−/−) mice in addition to Tg(UGT1A1*28)Ugt1(−/−) mice treated with phenobarbital are provided in Table 3. For SN-38, the Km values ranged from 1.44 μM in MLM prepared from phenobarbital-treated Tg(UGT1A1*28)Ugt1(−/−) mice to 3.59 μM in wild-type MLM. The Vmax increased from 43.6 pmol/min·mg in untreated Tg(UGT1A1*28)Ugt1(−/−) mice to 497 pmol/min·mg in MLM prepared from phenobarbital-treated Tg(UGT1A1*28)Ugt1 (−/−) mice (∼11-fold), a finding that supports the robust induction of UGT1A1 in microsomes from phenobarbital-treated Tg(UGT1A1*28)Ugt1 (−/−) mice. When ezetimibe and naloxone were evaluated, there were no differences in Km and Vmax values using microsomes from untreated and phenobarbital-treated Tg(UGT1A1*28)Ugt1(−/−) mice.
Discussion
Given the importance of P450 and UGTs in the metabolism and elimination of exogenous and endogenous xenobiotics, there is continuous interest in developing techniques and tools to identify the P450 and UGT enzyme(s) involved in the metabolism of a new drug and, additionally, in predicting human pharmacokinetic parameters to either guide compound selection in a discovery mode or select clinical doses in a development setting using in vitro kinetic parameters or in vivo preclinical pharmacokinetic data (Beaumont and Smith, 2009). Although qualitative and quantitative in vitro/in vivo correlation analysis based on data generated using human liver tissues and recombinant enzymes has been applied successfully to many drugs that are cleared by P450-dependent metabolism, these approaches have proven less definitive for glucuronidated compounds, mainly for the reasons alluded to earlier (Soars et al., 2003; Miners et al., 2004; Kilford et al., 2009). In addition, allometric scaling based on in vivo preclinical models has been successful for P450-eliminated drugs, provided that species-specific P450 enzymes have been well studied and understood (Caldwell et al., 2004; De Buck et al., 2007). On the other hand, given the current challenge of using preclinical species to predict human pharmacokinetics for predominantly glucuronidated compounds, more research is required to fully understand species differences and substrate specificity of UGT enzymes. It is clear that genetically modified mouse models, in which a specific gene is removed or replaced, have proven to be useful in studying drug-metabolizing enzymes (namely, P450 enzymes) and drug transporters even though quantitative prediction of the kinetic parameters of drugs is often limited by the complexity of gene transcription and translation and other factors, such as differences in levels and tissue expression patterns of enzymes in genetically modified mouse models (Lin, 2008). The recent generation of Tg(UGT1A1*28)Ugt1(−/−) mice has the potential to address issues of UGT-dependent drug clearance and toxicity because the humanization process does not alter expression of the constitutive mouse-specific UGT2 enzymes, as we have shown by Western blot analysis using UGT2B-specific antibodies (Nguyen et al., 2008). These humanized mice have been shown to be mildly hyperbilirubinemic, which serves as a biomarker of, and is fully consistent with, the expression of UGT1A1*28 and subsequent decrease in UGT1A1-specific glucuronidation (decreased Vmax).
To validate the utility of the humanized mouse model for studying hepatic glucuronidation, three literature compounds were chosen to cover a range of UGT1A1 involvement in the elimination of these compounds. The compounds evaluated were SN-38, the active metabolite of CPT-11 that is cleared predominantly by UGT1A1; ezetimibe (Zetia), which is reported to be eliminated partially by UGT1A1; and naloxone, which is not a UGT1A1 substrate (Fig. 1). CPT-11 and SN-38 have been well characterized because of their adverse effect (diarrhea) in some patients (Iyer et al., 1998, 1999; Onoue and Inui, 2008). After intravenous administration of a 1-mg/kg dose in mice, SN-38 showed approximately 4-fold differences in parent drug exposure (AUC) and clearance in the humanized mice compared with wild-type mice. The extremely high clearance values for SN-38 were likely caused by the extrahepatic elimination reported for the compound in rats (Chu et al., 1997). Similar differences in AUC and clearance were also seen between humanized and phenobarbital-treated Tg(UGT1A1*28)Ugt1(−/−) mice. Because UGT1A1 is efficiently induced by phenobarbital in Tg(UGT1A1*28)Ugt1(−/−) mice and others have shown that SN-38 is glucuronidated primarily by UGT1A1, these results are consistent with SN-38 glucuronidation by UGT1A1. In enzyme kinetic studies with HLM genotyped for normal UGT1A1*1 and variant UGT1A1*28, Vmax of SN-38 glucuronidation decreased 4- to 8-fold when comparing the *1/*1 and *28/*28 samples. These results are consistent with differences in protein concentration, with results reported by another laboratory where Vmax values showed a >10-fold difference using HLMs that were genotyped for the two UGT1A1 allelic variants (Zhang et al., 2007), and with the in vivo differences in pharmacokinetic parameters between phenobarbital-treated and untreated Tg(UGT1A1*28)Ugt1(−/−) mice. In addition, they follow the same rationale that the expression of UGT1A1 in liver microsomes obtained from UGT1A1*28 individuals would only affect the catalytic activity (with decreased catalytic activity, Vmax) but not the substrate affinity (Km) given that the only difference is reduced liver expression of UGT1A1 (Iyer et al., 1998; Gagné et al., 2002). Furthermore, the change in Vmax between HLMs genotyped as UGT1A1*1 compared with UGT1A1*28 supports the clinical finding that Gilbert's patients are more prone to adverse events with CPT-11 therapy (Iyer et al., 2002; Tukey et al., 2002). With MLM prepared from wild-type and Tg(UGT1A1*28)Ugt1(−/−) mice (with and without phenobarbital treatment), the Vmax for SN-38 glucuronidation showed similar trends as those determined in HLM. In particular, the increases in Vmax from humanized MLM to wild-type MLM and to phenobarbital-treated Tg(UGT1A1*28)Ugt1(−/−) microsomes were approximately 5- and 11-fold, respectively, which implies a direct translation of in vitro enzyme kinetic parameters from MLM to HLM.
In contrast to SN-38, the clearance of the UGT2B7 substrate, naloxone (Coffman et al., 1998), was not altered in the humanized mice compared with wild-type mice, given that it is not a UGT1A1 substrate. Consistent with this in vivo observation, any differences in the maximal velocity of naloxone glucuronidation were minimal between HLMs genotyped for normal UGT1A1*1 and UGT1A1*28 or between MLMs prepared from Tg(UGT1A1*28)Ugt1(−/−) mice with or without phenobarbital treatment.
Using recombinant UGT preparations, Ghosal et al. (2004) previously showed ezetimibe to be a substrate for multiple human UGT forms (UGT1A1, UGT1A3, UGT2B7, and UGT2B15). Our investigation began with mouse pharmacokinetic determinations and with ezetimibe administration to wild-type and Tg(UGT1A1*28)Ugt1(−/−) mice at 1 mg/kg i.v.; clearance of the drug was found to be very similar in both strains. At a higher dose (5 mg/kg), only a 2-fold increase in AUC and a 2-fold decrease in clearance were seen, which is likely the result of enzyme (UGT1A1) saturation in Tg(UGT1A1*28)Ugt1(−/−) mice (given the low protein levels; Fig. 3). Overall, this effect was a surprising observation, considering previous reports that ezetimibe served as a major substrate for recombinant UGT1A1. In addition, the expression of UGT1A1 in Tg(UGT1A1*28)Ugt1(−/−) hepatic tissue is minimal, as shown by Western blot analysis. To further evaluate this observation, ezetimibe glucuronidation was examined in microsomes from phenobarbital-treated Tg(UGT1A1*28)Ugt1(−/−) mice, where induction of human UGT1A1 was robust. The Km and Vmax values as determined in MLM from untreated and phenobarbital-treated mice were virtually identical, which is consistent with what was seen with HLM expressing either normal UGT1A1*1 or UGT1A1*28. More importantly, the rate of ezetimibe glucuronide formation was not reduced in HLM samples genotyped as UGT1A1*28 compared with normal UGT1A1*1 HLMs, indicating that glucuronidation by UGT1A1 may not play a major role in overall ezetimibe metabolism. Because phenobarbital leads to the induction of UGT1A4 (Argikar et al., 2009) and we have observed significant induction of UGT1A3 and UGT1A9 and mild induction of UGT1A6 in Tg-UGT1 mice (Fig. 4), it is instead likely that ezetimibe conjugation is catalyzed by UGT2B proteins. These results also indicate that the interpretation of enzymatic efficiency and substrate specificity using only expressed forms of the human UGTs must be interpreted with caution. The UGT functional specificity as they operate in vivo, which may depend on tissue-specific factors in addition to variations in heterodimer formation (Operaña and Tukey, 2007), may result in glucuronidation efficiencies that are much different from those observed in vitro using expressed forms of single UGTs.
In summary, the Tg(UGT1A1*28)Ugt1(−/−) mice have proven to be a useful tool to study UGT1A1-related drug clearance in vivo and in vitro and have the ability to at least qualitatively predict UGT1A1-involved clearance. Human liver UGT1A1 expression in individuals homozygous for the UGT1A1*1 allele can result in up to 10-fold greater protein concentration in liver compared with those that are homozygous for UGT1A1*28. These differences reflect similar patterns of human UGT1A1 expression in humanized mice when comparing their levels in untreated and phenobarbital-treated mice. These similarities validate the use of humanized UGT1 mice to examine pharmacokinetic parameters in vivo, especially when substrates are selectively metabolized by UGT1A1. Recent experiments using transgenic UGT1 mice have been used successfully to examine the pharmacokinetic parameters of lamotrigine, a popular antiepileptic drug that is metabolized by human UGT1A4 (Argikar et al., 2009). With current human safety assessments of new drugs relying primarily on standard animal models and in vitro experiments, coupled with arguments that intrinsic clearance values that rely on in vitro microsomal metabolism studies underpredict in vivo clearance values by an order of magnitude (Miners et al., 2006), the introduction of humanized animals modeled to express drug and xenobiotic-metabolizing enzymes should be perceived as a valued addition (Cheung and Gonzalez, 2008) toward providing accurate information in predicting human drug metabolism.
Acknowledgments.
We thank Steve Wene for important contributions to this research. We also thank Jeanne Rumsey, Dr. Larry Foresman, Debra Erbschloe, and Dr. Leonid Kirkovsky for assistance and/or helpful suggestions during various stages of this research.
This work was supported in part by the National Institutes of Health National Institute of Environmental Health Sciences [Grant ES10337]; Pfizer; and the Tobacco-Related Disease Research Program from the State of California [Grant 15RT-0251].
Article, publication date, and citation information can be found at http://dmd.aspetjournals.org.
doi:10.1124/dmd.109.030130.
- UGT
- UDP-glucuronosyltransferase
- P450
- cytochrome P450
- SN-38
- 7-ethyl-10-hydroxy-camptothecin
- CPT-11
- irinotecan
- HLM
- human liver microsome
- MLM
- mouse liver microsome
- LC/MS/MS
- liquid chromatography/tandem mass spectrometry
- AUC
- area under the curve.
References
- Ando Y, Fujita K, Sasaki Y, Hasegawa Y. (2007) UGT1AI*6 and UGT1A1*27 for individualized irinotecan chemotherapy. Curr Opin Mol Ther 9:258–262 [PubMed] [Google Scholar]
- Argikar UA, Senekeo-Effenberger K, Larson EE, Tukey RH, Remmel RP. (2009) Studies on induction of lamotrigine metabolism in transgenic UGT1 mice. Xenobiotica 39:826–835 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beaumont K, Smith DA. (2009) Does human pharmacokinetic prediction add significant value to compound selection in drug discovery research? Curr Opin Drug Discov Devel 12:61–71 [PubMed] [Google Scholar]
- Bosma PJ. (2003) Inherited disorders of bilirubin metabolism. J Hepatol 38:107–117 [DOI] [PubMed] [Google Scholar]
- Caldwell GW, Masucci JA, Yan Z, Hageman W. (2004) Allometric scaling of pharmacokinetic parameters in drug discovery: can human CL, Vss and t1/2 be predicted from in-vivo rat data? Eur J Drug Metab Pharmacokinet 29:133–143 [DOI] [PubMed] [Google Scholar]
- Chen S, Beaton D, Nguyen N, Senekeo-Effenberger K, Brace-Sinnokrak E, Argikar U, Remmel RP, Trottier J, Barbier O, Ritter JK, et al. (2005) Tissue-specific, inducible, and hormonal control of the human UDP-glucuronosyltransferase-1 (UGT1) locus. J Biol Chem 280:37547–37557 [DOI] [PubMed] [Google Scholar]
- Cheung C, Gonzalez FJ. (2008) Humanized mouse lines and their application for prediction of human drug metabolism and toxicological risk assessment. J Pharmacol Exp Ther 327:288–299 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu XY, Kato Y, Niinuma K, Sudo KI, Hakusui H, Sugiyama Y. (1997) Multispecific organic anion transporter is responsible for the biliary excretion of the camptothecin derivative irinotecan and its metabolites in rats. J Pharmacol Exp Ther 281:304–314 [PubMed] [Google Scholar]
- Coffman BL, King CD, Rios GR, Tephly TR. (1998) The glucuronidation of opioids, other xenobiotics, and androgens by human UGT2B7Y(268) and UGT2B7H(268). Drug Metab Dispos 26:73–77 [PubMed] [Google Scholar]
- Court MH. (2005) Isoform-selective probe substrates for in vitro studies of human UDP-glucuronosyltransferases. Methods Enzymol 400:104–116 [DOI] [PubMed] [Google Scholar]
- De Buck SS, Sinha VK, Fenu LA, Nijsen MJ, Mackie CE, Gilissen RA. (2007) Prediction of human pharmacokinetics using physiologically based modeling: a retrospective analysis of 26 clinically tested drugs. Drug Metab Dispos 35:1766–1780 [DOI] [PubMed] [Google Scholar]
- Ethell BT, Beaumont K, Rance DJ, Burchell B. (2001) Use of cloned and expressed human UDP-glucuronosyltransferases for the assessment of human drug conjugation and identification of potential drug interactions. Drug Metab Dispos 29:48–53 [PubMed] [Google Scholar]
- Fang JL, Lazarus P. (2004) Correlation between the UDP-glucuronosyltransferase (UGT1A1) TATAA box polymorphism and carcinogen detoxification phenotype: significantly decreased glucuronidating activity against benzo(a)pyrene-7,8-dihydrodiol(-) in liver microsomes from subjects with the UGT1A1*28 variant. Cancer Epidemiol Biomarkers Prev 13:102–109 [DOI] [PubMed] [Google Scholar]
- Fujiwara R, Nakajima M, Yamanaka H, Katoh M, Yokoi T. (2007) Interactions between human UGT1A1, UGT1A4, and UGT1A6 affect their enzymatic activities. Drug Metab Dispos 35:1781–1787 [DOI] [PubMed] [Google Scholar]
- Gagné JF, Montminy V, Belanger P, Journault K, Gaucher G, Guillemette C. (2002) Common human UGT1A polymorphisms and the altered metabolism of irinotecan active metabolite 7-ethyl-10-hydroxycamptothecin (SN-38). Mol Pharmacol 62:608–617 [DOI] [PubMed] [Google Scholar]
- Ghosal A, Hapangama N, Yuan Y, Achanfuo-Yeboah J, Iannucci R, Chowdhury S, Alton K, Patrick JE, Zbaida S. (2004) Identification of human UDP-glucuronosyltransferase enzyme(s) responsible for the glucuronidation of ezetimibe (Zetia). Drug Metab Dispos 32:314–320 [DOI] [PubMed] [Google Scholar]
- Institute of Laboratory Animal Resources (1996) Guide for the Care and Use of Laboratory Animals, 7th ed Institute of Laboratory Animal Resources, Commission on Life Sciences, National Research Council, Washington DC: [Google Scholar]
- Ishii Y, Iwanaga M, Nishimura Y, Takeda S, Ikushiro S, Nagata K, Yamazoe Y, Mackenzie PI, Yamada H. (2007) Protein-protein interactions between rat hepatic cytochromes P450 (P450s) and UDP-glucuronosyltransferases (UGTs): evidence for the functionally active UGT in P450-UGT complex. Drug Metab Pharmacokinet 22:367–376 [DOI] [PubMed] [Google Scholar]
- Iyer L, Das S, Janisch L, Wen M, Ramírez J, Karrison T, Fleming GF, Vokes EE, Schilsky RL, Ratain MJ. (2002) UGT1A1*28 polymorphism as a determinant of irinotecan disposition and toxicity. Pharmacogenomics J 2:43–47 [DOI] [PubMed] [Google Scholar]
- Iyer L, Hall D, Das S, Mortell MA, Ramírez J, Kim S, Di Rienzo A, Ratain MJ. (1999) Phenotype-genotype correlation of in vitro SN-38 (active metabolite of irinotecan) and bilirubin glucuronidation in human liver tissue with UGT1A1 promoter polymorphism. Clin Pharmacol Ther 65:576–582 [DOI] [PubMed] [Google Scholar]
- Iyer L, King CD, Whitington PF, Green MD, Roy SK, Tephly TR, Coffman BL, Ratain MJ. (1998) Genetic predisposition to the metabolism of irinotecan (CPT-11). Role of uridine diphosphate glucuronosyltransferase isoform 1A1 in the glucuronidation of its active metabolite (SN-38) in human liver microsomes. J Clin Invest 101:847–854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kilford PJ, Stringer R, Sohal B, Houston JB, Galetin A. (2009) Prediction of drug clearance by glucuronidation from in vitro data: use of combined cytochrome P450 and UDP-glucuronosyltransferase cofactors in alamethicin-activated human liver microsomes. Drug Metab Dispos 37:82–89 [DOI] [PubMed] [Google Scholar]
- Lin JH. (2008) Applications and limitations of genetically modified mouse models in drug discovery and development. Curr Drug Metab 9:419–438 [DOI] [PubMed] [Google Scholar]
- Miners JO, Knights KM, Houston JB, Mackenzie PI. (2006) In vitro-in vivo correlation for drugs and other compounds eliminated by glucuronidation in humans: pitfalls and promises. Biochem Pharmacol 71:1531–1539 [DOI] [PubMed] [Google Scholar]
- Miners JO, Mackenzie PI, Knights KM. (2010) The prediction of drug-glucuronidation parameters in humans: UDP-glucuronosyltransferase enzyme-selective substrate and inhibitor probes for reaction phenotyping and in vitro-in vivo extrapolation of drug clearance and drug-drug interaction potential. Drug Metab Rev 42:189–201 [DOI] [PubMed] [Google Scholar]
- Miners JO, McKinnon RA, Mackenzie PI. (2002) Genetic polymorphisms of UDP-glucuronosyltransferases and their functional significance. Toxicology 181–182:453–456 [DOI] [PubMed] [Google Scholar]
- Miners JO, Smith PA, Sorich MJ, McKinnon RA, Mackenzie PI. (2004) Predicting human drug glucuronidation parameters: application of in vitro and in silico modeling approaches. Annu Rev Pharmacol Toxicol 44:1–25 [DOI] [PubMed] [Google Scholar]
- Nagar S, Blanchard RL. (2006) Pharmacogenetics of uridine diphosphoglucuronosyltransferase (UGT) 1A family members and its role in patient response to irinotecan. Drug Metab Rev 38:393–409 [DOI] [PubMed] [Google Scholar]
- Nguyen N, Bonzo JA, Chen S, Chouinard S, Kelner MJ, Hardiman G, Bélanger A, Tukey RH. (2008) Disruption of the ugt1 locus in mice resembles human Crigler-Najjar type I disease. J Biol Chem 283:7901–7911 [DOI] [PubMed] [Google Scholar]
- O'Dwyer PJ, Catalano RB. (2006) Uridine diphosphate glucuronosyltransferase (UGT) 1A1 and irinotecan: practical pharmacogenomics arrives in cancer therapy. J Clin Oncol 24:4534–4538 [DOI] [PubMed] [Google Scholar]
- Onoue M, Inui K. (2008) [Role of UGT1A1*28 and UGT1A1*6 for irinotecan-induced adverse drug reaction]. Gan To Kagaku Ryoho 35:1080–1085 [PubMed] [Google Scholar]
- Operaña TN, Tukey RH. (2007) Oligomerization of the UDP-glucuronosyltransferase 1A proteins: homo- and heterodimerization analysis by fluorescence resonance energy transfer and co-immunoprecipitation. J Biol Chem 282:4821–4829 [DOI] [PubMed] [Google Scholar]
- Peterkin VC, Bauman JN, Goosen TC, Menning L, Man MZ, Paulauskis JD, Williams JA, Myrand SP. (2007) Limited influence of UGT1A1*28 and no effect of UGT2B7*2 polymorphisms on UGT1A1 or UGT2B7 activities and protein expression in human liver microsomes. Br J Clin Pharmacol 64:458–468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowland A, Knights KM, Mackenzie PI, Miners JO. (2008) The “albumin effect” and drug glucuronidation: bovine serum albumin and fatty acid-free human serum albumin enhance the glucuronidation of UDP-glucuronosyltransferase (UGT) 1A9 substrates but not UGT1A1 and UGT1A6 activities. Drug Metab Dispos 36:1056–1062 [DOI] [PubMed] [Google Scholar]
- Soars MG, Ring BJ, Wrighton SA. (2003) The effect of incubation conditions on the enzyme kinetics of UDP-glucuronosyltransferases. Drug Metab Dispos 31:762–767 [DOI] [PubMed] [Google Scholar]
- Strassburg CP, Kneip S, Topp J, Obermayer-Straub P, Barut A, Tukey RH, Manns MP. (2000) Polymorphic gene regulation and interindividual variation of UDP-glucuronosyltransferase activity in human small intestine. J Biol Chem 275:36164–36171 [DOI] [PubMed] [Google Scholar]
- Takeda S, Ishii Y, Mackenzie PI, Nagata K, Yamazoe Y, Oguri K, Yamada H. (2005) Modulation of UDP-glucuronosyltransferase 2B7 function by cytochrome P450s in vitro: differential effects of CYP1A2, CYP2C9 and CYP3A4. Biol Pharm Bull 28:2026–2027 [DOI] [PubMed] [Google Scholar]
- Tukey RH, Strassburg CP. (2000) Human UDP-glucuronosyltransferases: metabolism, expression, and disease. Annu Rev Pharmacol Toxicol 40:581–616 [DOI] [PubMed] [Google Scholar]
- Tukey RH, Strassburg CP, Mackenzie PI. (2002) Pharmacogenomics of human UDP-glucuronosyltransferases and irinotecan toxicity. Mol Pharmacol 62:446–450 [DOI] [PubMed] [Google Scholar]
- Udomuksorn W, Elliot DJ, Lewis BC, Mackenzie PI, Yoovathaworn K, Miners JO. (2007) Influence of mutations associated with Gilbert and Crigler-Najjar type II syndromes on the glucuronidation kinetics of bilirubin and other UDP-glucuronosyltransferase 1A substrates. Pharmacogenet Genomics 17:1017–1029 [DOI] [PubMed] [Google Scholar]
- Zhang D, Zhang D, Cui D, Gambardella J, Ma L, Barros A, Wang L, Fu Y, Rahematpura S, Nielsen J, et al. (2007) Characterization of the UDP glucuronosyltransferase activity of human liver microsomes genotyped for the UGT1A1*28 polymorphism. Drug Metab Dispos 35:2270–2280 [DOI] [PubMed] [Google Scholar]