Abstract
The type 5 G protein β subunit (Gβ5) can form complexes with members of the Regulator of G protein Signaling 7 (RGS7) family, but the relevance to neuronal G protein signaling is unclear. We report that mouse RGS7/Gβ5 complexes bind to G protein–gated potassium channels and facilitate their functional coupling to GABAB receptors in neurons. These findings identify a novel compartmentalization mechanism critical for ensuring high temporal resolution of neuronal G protein signaling.
Gβ5 is a divergent member of the Gβ family that does not engage in signaling from G protein–coupled receptors to effectors but rather binds to the Gγ–like domain present in the R7 group of Regulator of G protein Signaling (R7 RGS) proteins 1. In mammals, Gβ5–R7 RGS complexes critically shape vision, nociception, and reward behavior by ensuring timely inactivation of G protein responses following termination of receptor activation 2. Association with Gβ5 is required for proper folding and proteolytic stability of R7 RGS proteins and ablation of the Gβ5 eliminates all R7 RGS protein expression 3. However, despite the obligate nature of their association, the functional role of Gβ5 in the context of RGS proteins is unknown.
The interface found in conventional Gβ subunits (Gβ1–4) that mediates interaction with Gα subunits and effectors is conserved in Gβ5 4. Thus, we probed for association between Gβ5 and three Gβγ–regulated effectors: G protein–gated inwardly–rectifying K+ (GIRK/Kir3) channels, phosphoinositide 3–kinase gamma (PI3Kγ), and β2 type phospholipase C (PLCβ2) (Fig. 1a,b,c; see Supplementary Material for a full description of methods). In transfected cells, Gβ5 co–immunoprecipitated with the GIRK channel subunit GIRK1 and the P101 subunit of PI3Kγ, but not with PLCβ2. However, among the three effectors examined, only GIRK1 could co–precipitate with the physiologically–relevant complex formed by Gβ5 and the prototypical R7 RGS protein RGS7.
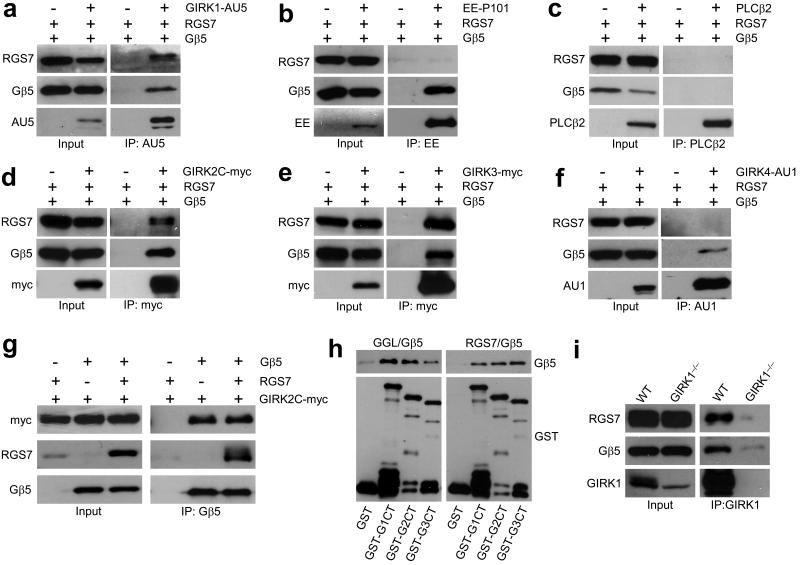
Figure 1. The Gβ5–RGS7 complex binds specifically to GIRK channels.
a–c) Co–immunoprecipitation of RGS7 and Gβ5 with GIRK1 (a) but not with P101 subunit of PI3Kγ (b) or PLCβ2 (c) from transfected 293T cells. ‘+’ and ‘−’ denote the presence and absence, respectively, of the pertinent expression construct in the transfection mixture. Immunoprecipitations (IP) were conducted with antibodies against epitope–tagged effectors (3 μg each) and resultant immunocomplexes were probed for the presence of RGS7, Gβ5, and effector by immunoblotting. Transfected cells without effector construct served as controls for non–specific binding. d–f) RGS7 and Gβ5 co–immunoprecipitate with GIRK2 and GIRK3 but not GIRK4. Tagged GIRK subunits were immunoprecipitated from cells co–transfected with RGS7 and/or Gβ5, and proteins in the eluates were detected by immunoblotting. g) GIRK2 co–immunoprecipitated with Gβ5 in the absence and presence of RGS7. h) Gβ5–RGS complexes bind to GIRK subunits via direct protein–protein interactions. GST–tagged C–terminal (ct) cytoplasmic domains of GIRK1 (GST–G1ct), GIRK2 (GST–G2ct), and GIRK3 (GST–G3ct) subunits were immobilized on beads and incubated with either purified recombinant full–length RGS7/Gβ5 (right) or Gβ5 bound to the Gγ–like (GGL) domain of RGS9 (left). Proteins retained on the beads after washing were detected by immunoblotting with anti–Gβ5 and anti–GST antibodies. i) Gβ5 and RGS7 associate with GIRK1 in the mouse hippocampus. Hippocampal membrane samples were prepared from wild–type and Girk1–/– mice and used in co–immunoprecipitation studies. Both Gβ5 and RGS7, co–immunoprecipitated with GIRK1 in wild–type but not Girk1–/– samples. Immunoblots were cropped for space reduction. Please refer to Supplemental Fig. S4 for full-length blots.
GIRK channels mediate the postsynaptic inhibitory effect of many neurotransmitters including GABA, and they play key roles in synaptic plasticity and behavior 5,6,7. Neuronal GIRK channels are formed primarily by GIRK1 (Kcnj3), GIRK2 (Kcnj6), and GIRK3 (Kcnj9) subunits 8. Interestingly, Gβ5–RGS7 also co–immunoprecipitated with GIRK2 and GIRK3, but not with the cardiac GIRK subunit GIRK4 (Kcnj5) (Fig. 1d,e,f). Further forward and reciprocal immunoprecipitation experiments revealed that Gβ5 could bind GIRK subunits in the absence of RGS7, suggesting that Gβ5 mediates formation of the GIRK–RGS complex (Fig. 1g, Supplemental Fig. S1). Indeed, C–terminal cytoplasmic domains of GIRK subunits bound to complexes consisting of Gβ5 and full–length RGS7 or a minimal binding fragment consisting of the RGS9 Gγ–like (GGL) domain in a pull–down assay (Fig. 1h). Importantly, anti–GIRK1 antibodies co–immunoprecipitated Gβ5 and RGS7 from wild–type but not Girk1–/– hippocampus, indicating that this complex exists in vivo (Fig. 1i).
We next probed the impact of Gβ5 (Gnb5) ablation on GIRK signaling in neurons. Consistent with published studies 9, the GABAB receptor agonist baclofen (100 μM) evoked a robust inward, barium–sensitive K+ current that was absent in cultured hippocampal neurons from mice lacking the Girk2 gene (Supplemental Fig. S2). As shown in Fig. 1i, Gβ5 and RGS7 are expressed and interact with GIRK1 in the mouse hippocampus; the complete absence of RGS7 protein in hippocampal samples from Gβ5–/– mice underscores the obligate nature of Gβ5/R7 RGS protein complexes (Supplemental Fig. S3). Gβ5 ablation did not alter the resting membrane potential (WT: −59±3 mV, n=9 vs. Gβ5–/–: −59±2 mV, n=13; P=0.92), input resistance (WT: 149±22 MΩ vs. Gβ5–/–: 178±24 MΩ; P=0.39), or baclofen–induced current density (WT: −6.9±1.1 pA/pF vs. Gβ5–/–: −7.3±0.7 pA/pF P=0.73) of cultured hippocampal neurons. Current deactivation kinetics, and to a lesser extent activation kinetics, however, were significantly slower in neurons from Gβ5–/– mice (τact=304±38 ms, P<0.05; τdeact=3281±211 ms; P<0.001) as compared to controls (τact=181±26 ms; τdeact=1522±134 ms) (Fig. 2a,b). Moreover, baclofen was more potent with respect to GIRK channel activation in neurons from Gβ5–/– mice (EC50 = 0.4–0.7 μM; 95% CI) as compared to controls (EC50 = 1.6–2.5 μM; 95% CI) (Fig. 2c,d). Collectively, these data argue that Gβ5, in complex with RGS7 and/or another R7 RGS protein, influences both the timing and sensitivity of the GABAB receptor–GIRK signaling pathway in hippocampal neurons.
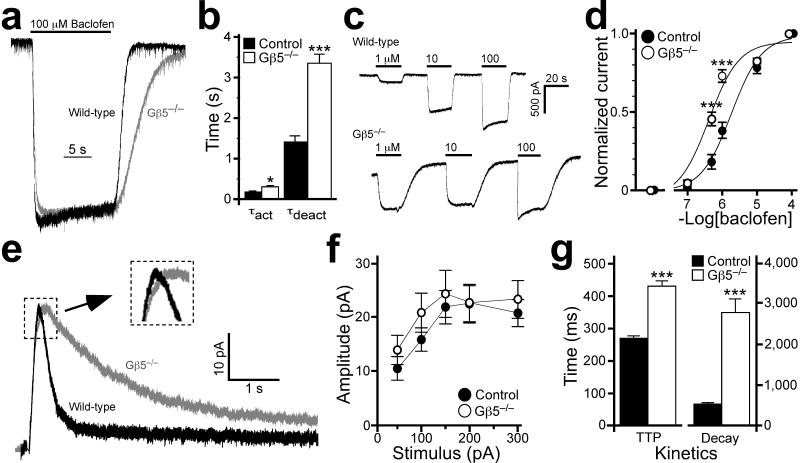
Figure 2. Gβ5 ablation delays GABAB–GIRK signaling kinetics.
a) Representative currents induced by 100 μM baclofen in a wild–type (WT, black) and Gβ5–/– (gray) neuron are normalized to emphasize the genotype–dependent differences in current kinetics. b) Activation (τact) and deactivation (τdeact) kinetics for the baclofen–induced current in control and Gβ5–/– neurons (n = 9–13 per group). Both activation and deactivation kinetics were delayed in neurons from Gβ5–/– mice. c) Representative inward currents measured in neurons from control (upper trace) and Gβ5–/– (lower trace) mice following application of 1, 10, and 100 μM baclofen. Note both the delayed current deactivation kinetics and the enhanced sensitivity to 1 μM baclofen in the Gβ5–/– neuron. d) Concentration–response relationship for baclofen–activation of GIRK current in wild–type (black circles) and Gβ5–/– (open circles) neurons. Steady–state inward currents evoked sequentially by either 0.1 or 0.5 μM baclofen, and then 1 and 10 μM baclofen, were normalized to the response to 100 μM baclofen applied at the end of the experiment (n=10–12 experiments per genotype). Main effects of genotype (F1,73=23.6, P<0.001) and concentration (F5,73=220.4, P<0.001), as well as an interaction between genotype and concentration (F5,73=11.2, P<0.001), were observed. e) Representative slow evoked IPSCs in CA1 neurons from wild–type (black) and Gβ5–/– (gray) mice; amplitudes are normalized to emphasize genotype–dependent kinetic differences. Each trace is an average of 15 sweeps and stimulus artifacts have been removed. Inset, magnified view showing the slower rise of the IPSC in Gβ5–/– mice. f) IPSC amplitude vs. stimulus intensity in control and Gβ5–/– mice (n = 14–18 per genotype). The stimulus response curves were indistinguishable (F1,23=1.0, P>0.05; interaction genotype × input, F3,69=0.6, P>0.05). g) The time–to–peak (TTP) and decay (90–37%) of the evoked IPSCs in wild–type and Gβ5–/– mice. Symbols: *,*** P<0.05 and 0.001 respectively, vs. control. Error bars are s.e.m.
We next measured evoked inhibitory postsynaptic currents (IPSCs) in hippocampal CA1 pyramidal neurons in acutely–isolated slices to determine whether Gβ5 ablation altered synaptically–driven responses. While the amplitude of the slow IPSC did not differ between genotypes (Fig. 2e,f), both the rise time (time–to–peak, TTP) and decay of the IPSC were significantly slower in slices from Gβ5–/– mice (Fig. 2g). Integration (area under the curve) revealed a ~2.5–fold increase (t(30)=3.3, P<0.01) in the synaptically–evoked slow IPSC in CA1 neurons from Gβ5–/– mice. To our knowledge, these data for the first time implicate RGS protein activity in shaping IPSC responses, a hallmark of inhibitory signaling in the nervous system.
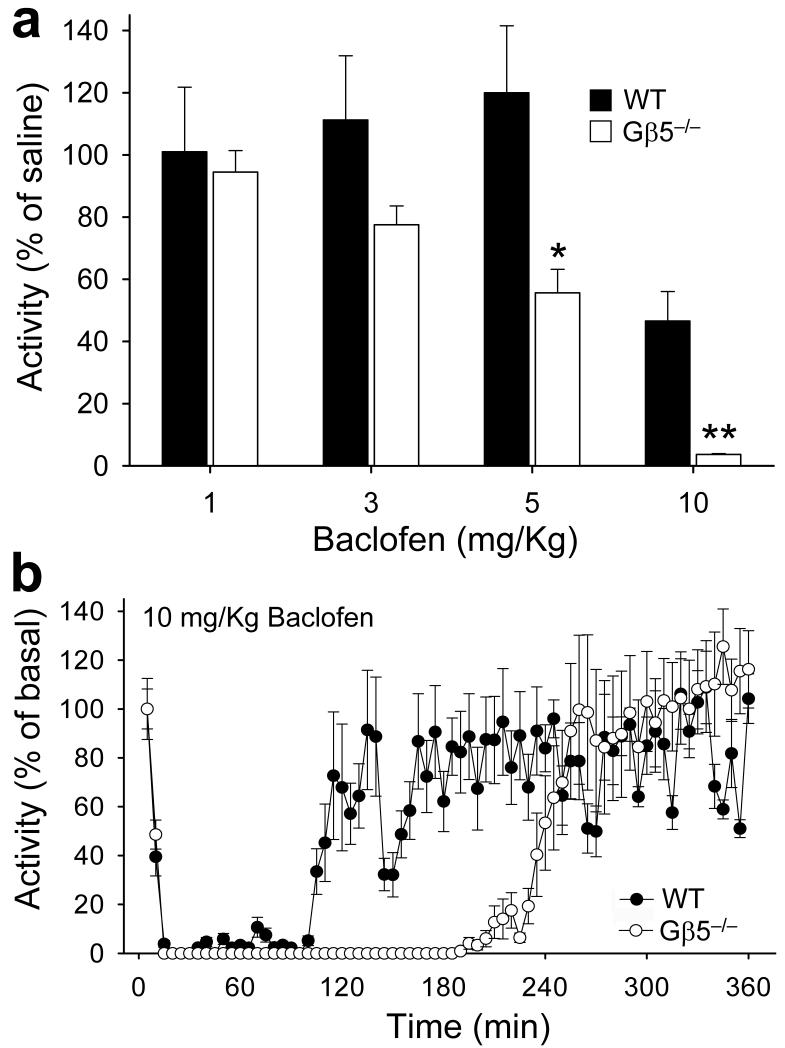
Baclofen evokes muscle–relaxation, sedation, and hypothermia effects 10 that are blunted in Girk–/– mice 6,11. Delayed GABAB–GIRK signaling kinetics and enhanced GABAB–dependent postsynaptic responses predict that these mice would exhibit enhanced behavioral responses to baclofen. To test this possibility, we monitored the effect of baclofen on Gβ5–/– mice and wild–type siblings in an open–field locomotor activity assay. Consistent with our predictions, baclofen at 5 mg/kg caused a pronounced reduction in locomotor activity in Gβ5–/– mice, while having no effect on wild–type siblings (Fig. 3a). Moreover, at 10 mg/kg baclofen, Gβ5–/– mice exhibited a prolonged (3 hr) immobilization, while wild–type littermates were markedly less affected (Fig. 3b). Thus, Gβ5 ablation in mice yielded enhanced sensitivity to a key behavioral effect of GABAB receptor stimulation.
Figure 3. Gβ5 ablation potentiates GABAB–dependent behavior.
a) Open–field locomotor activity of wild–type (WT, n=7) and Gβ5–/– (n = 5) mice following administration of increasing baclofen doses (1, 3, 5, 10 mg/kg, s.c.). Activity was monitored for 3 hr after baclofen injection and then normalized to the activity of saline–treated controls of the same genotype. Basal locomotor activities of saline–treated controls were: 109±11 and 980±53 meters traveled per 3 hr session for wild–type and Gβ5–/– mice, respectively. b) The effect of 10 mg/kg baclofen on motor activity as a function of time. Symbols: *, P<0.01 vs. control. Error bars are s.e.m. Mice were used in accordance with the protocols approved by the Animal Care and Use Committee at the University of Minnesota.
In summary, we present evidence that Gβ5 recruits R7 RGS proteins to neuronal GIRK channels via direct interaction with the intracellular C–terminal domain of GIRK channel subunits. This finding helps explain high potency of R7 RGS–Gβ5 complexes seen in reconstituted systems 12,13,14 and provides a mechanism for the stabilization–independent enhancement of RGS7– and RGS9–mediated acceleration of GIRK channel kinetics by Gβ5 12,15. Moreover, we show that in the absence of Gβ5, the high temporal fidelity that characterizes the GIRK channel response to GABAB receptor activation in neurons is disrupted and that a GIRK–dependent motor–inhibitory effect linked to GABAB receptor activation is enhanced. Thus, this new example of RGS–effector association provides important insight into the compartmentalization of GABAB–GIRK signaling in neurons while revealing a mechanism for controlling temporal and spatial characteristics of inhibitory signaling in neuronal circuitry. Finally, while specificity for GIRK channels was observed with the subset of effectors examined herein, it is possible that Gβ5 recruits RGS proteins to other effectors as well, making the identification of additional effector targets an important and exciting future direction of research.
Supplementary Material
Acknowledgements
We thank Dr. Jason Chen (Virginia Commonwealth University) for providing the line of Gβ5–/– mice, Dr. William Simonds for the gift of anti–Gβ5 and anti–RGS7 antibodies and Marc Parent for assistance with the hippocampal slicing technique. This work was supported by NIH grants DA021743 (K.A.M.), DA026405 (K.A.M.), EY018139 (K.A.M.), MH061933 (K.W.), DA011806 (K.W.), DA019666 (M.J.T.), and MH078291 (J.C.S.), a McKnight Land–Grant Award (K.A.M), and an award from the Whitehall Foundation (M.J.T.).
REFERENCES
- 1.Sondek J, Siderovski DP. Biochem. Pharmacol. 2001;61:1329–1337. doi: 10.1016/s0006-2952(01)00633-5. [DOI] [PubMed] [Google Scholar]
- 2.Anderson GR, Posokhova E, Martemyanov KA. Cell. Biochem. Biophys. 2009;54:33–46. doi: 10.1007/s12013-009-9052-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chen CK, et al. Proc. Natl. Acad. Sci. U.S.A. 2003;100:6604–6609. doi: 10.1073/pnas.0631825100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cheever ML, et al. Nat. Struct. Mol. Biol. 2008;15:155–162. doi: 10.1038/nsmb.1377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Luscher C, Jan LY, Stoffel M, Malenka RC, Nicoll RA. Neuron. 1997;19:687–695. doi: 10.1016/s0896-6273(00)80381-5. [DOI] [PubMed] [Google Scholar]
- 6.Pravetoni M, Wickman K. Genes Brain Behav. 2008;7:523–531. doi: 10.1111/j.1601-183X.2008.00388.x. [DOI] [PubMed] [Google Scholar]
- 7.Chung HJ, et al. Proc. Natl. Acad. Sci. U.S.A. 2009;106:635–640. doi: 10.1073/pnas.0811685106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Karschin C, Dissmann E, Stuhmer W, Karschin AJ. Neurosci. 1996;16:3559–3570. doi: 10.1523/JNEUROSCI.16-11-03559.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chung HJ, Qian X, Ehlers M, Jan YN, Jan LY. Proc. Natl. Acad. Sci. U.S.A. 2009;106:629–634. doi: 10.1073/pnas.0811615106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bettler C.V.a.B. Curr. Drug Targets CNS Neurol. Disord. 2003;2:248–259. doi: 10.2174/1568007033482814. [DOI] [PubMed] [Google Scholar]
- 11.Costa AC, Stasko MR, Stoffel M, Scott–McKean JJ. J. Neurosci. 2005;25:7801–7804. doi: 10.1523/JNEUROSCI.1699-05.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kovoor A, et al. J. Biol. Chem. 2000;275:3397–3402. doi: 10.1074/jbc.275.5.3397. [DOI] [PubMed] [Google Scholar]
- 13.Rahman Z, et al. Neuron. 2003;38:941–952. doi: 10.1016/s0896-6273(03)00321-0. [DOI] [PubMed] [Google Scholar]
- 14.Drenan RM, et al. J. Biol. Chem. 2006;281:28222–28231. doi: 10.1074/jbc.M604428200. [DOI] [PubMed] [Google Scholar]
- 15.Keren–Raifman T, et al. FEBS Lett. 2001;492:20–28. doi: 10.1016/s0014-5793(01)02220-7. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.