Abstract
Sickle cell disease (SCD) is characterized by a prothrombotic state. Plasminogen activator inhibitor-1 (PAI-1) is known to modulate fibrinolysis, lung injury/fibrosis, and angiogenesis. However, its role in SCD is less understood, and the molecular mechanisms underlying increased PAI-1 are unknown. Herein, we show a novel link between PAI-1 and sickle erythropoiesis. Plasma PAI-1 levels were high in SCD patients at steady state and in two humanized sickle mouse models, with increased PAI-1 immunolabeling in sickle mouse lung, bronchial epithelial cells, alveolar macrophages, and pulmonary microvascular endothelial cells. Placenta growth factor (PlGF), released at high levels by sickle erythroblasts, induced PAI-1 expression in primary human pulmonary microvascular endothelial cells and monocytes through activation of c-Jun N-terminal kinase (JNK), NADPH oxidase, and hypoxia-inducible factor-1α (HIF-1α). Analysis of the human PAI-1 promoter revealed this induction was mediated by hypoxia-response element (HRE)-1, HRE-2, and distal activator protein (AP-1) sites. We also identify the involvement of c-Jun, c-Jun/c-Fos, and JunD, but not JunB, in binding with AP-1 sites of the PAI-1 promoter upon PlGF induction. Consistent with these findings, levels of PAI-1 were low in PlGF knock-out mice and sickle-PlGF knock-out mice; overexpression of PlGF in normal mice increased circulating PAI-1. In conclusion, we identify a novel mechanism of PAI-1 elevation in SCD.
Keywords: Endothelium, Erythropoeisis, Jun Transcription Factor, Plasma, Plasminogen Inhibitor, Human Pulmonary Microvascular Endothelial Cells, Hypoxia-inducible Factor-1 alpha, Placenta Growth Factor, Plasminogen Activator Inhibitor-1, Sickle Cell Disease
Introduction
SCD5 is caused by a mutation in the β-globin chain of hemoglobin that results in sickling of red blood cells upon deoxygenation. Although vaso-occlusive phenomena and hemolysis are its clinical hallmarks, other clinical manifestations are also seen (1–3), such as hypercoagulation (4–7). SCD patients have increased levels of activated coagulation factors and decreased levels of anticoagulant proteins, including an abnormal fibrinolytic system (8, 9). Elevated plasma levels of circulating tissue factor (4) and PAI-1 have been reported at steady state and are further increased during sickle vaso-occlusive crises (10, 11). A prothrombotic state predisposes patients to pulmonary hypertension and stroke, two devastating complications of SCD (12).
In addition to its role in coagulation and fibrinolysis, PAI-1 functions in development of lung injury/fibrosis, tumor progression, angiogenesis, atherosclerosis, and wound healing by modulating the process of fibrinolysis or cell adhesion (13–15). Befitting its broad functional profile, PAI-1 is synthesized by a variety of cell types, such as endothelial cells, hepatocytes, platelets, fibroblasts, mononuclear cells, and smooth muscle cells (16–19). Its expression is modulated by a number of factors: hypoxia (17, 20), interleukin-1 (IL-1)(21), lipopolysaccharide(22), transforming growth factor-β (TGF-β) (23), D-dimer (18), and insulin (24). Induction of PAI-1 expression by hypoxia involves the transcription factors, HIF-1α, early growth response factor 1 (Egr-1), and CCAAT/enhancer-binding protein α (C/EBPα) (20). Oxidative stress regulates PAI-1 expression through activator protein (AP-1) (25). Little is known regarding plasma factors that lead to the elevated expression of PAI-1 in SCD.
We previously reported high circulating levels of PlGF derived from erythroblasts in SCD that correlate with increased propensity of vaso-occlusive crises (26). PlGF is an angiogenic growth factor that, in the fetus, is made by placental syncytiotrophoblasts and, postnatally, by erythroblasts. We showed that PlGF activates downstream target genes in endothelial cells via HIF-1α in a hypoxia-independent manner (27). Because PAI-1 expression is induced by HIF-1α (in a hypoxia-dependent fashion) and PlGF activates HIF-1α (in a hypoxia-independent fashion), we hypothesized that the PAI-1 elevation in SCD is mediated via PlGF acting through HIF-1α. We found high levels of PAI-1 in sickle mouse pulmonary endothelial cells, alveolar macrophages, bronchial epithelial cells, and plasma. In addition, we show that PlGF increased levels of PAI-1 through HIF-1α and the AP-1 pathway. Finally, we confirmed these results in vivo using genetic models to knock down or overexpress PlGF.
EXPERIMENTAL PROCEDURES
Cells and Reagents
Primary human pulmonary microvascular endothelial cells (HPMVEC) were obtained from Lonza Group Ltd. (Basel, Switzerland) and grown as described previously (27). Cultured HPMVEC were confirmed to express CD31 and von Willebrand factor (data not shown). Unless otherwise specified, HPMVEC were incubated overnight in EBM-2 medium containing 2% serum followed by serum deprivation for 3 h prior to PlGF treatment or other experimental condition.
Human recombinant PlGF, mouse PlGF-2, and human PAI-1 immunoassay kits were obtained from R&D Systems (Minneapolis, MN). Pharmacological inhibitors of cell signaling (supplemental Table S1) were obtained from Tocris Bioscience (Ellisville, MO), and [32P]UTP was obtained from MP Biomedicals (Solon, OH). Antibodies against vascular endothelial growth factor receptor 1 (VEGFR1), vascular endothelial growth factor receptor 2 (VEGFR2), HIF-1α, phospho-c-Jun, c-Jun, phospho-JNK, and JNK, as well as horseradish peroxidase-conjugated secondary antibodies, were obtained from Santa Cruz Biotechnology (Santa Cruz, CA).
HIF-1α siRNA and scrambled short hairpin RNA were synthesized at the Microchemical Core facility of the University of Southern California-Norris Comprehensive Cancer Center (USC-NCCC). TranSilent siRNA vectors for Egr-1, c-Jun, and cAMP-response element-binding protein (CREB1) were from Panomics Inc. (Fremont, CA). Oligonucleotides used for electrophoretic mobility shift assay (EMSA) were obtained from Integrated DNA Technologies (Coralville, IA). All other reagents were purchased from Sigma.
Mice and Human Subjects
C57BL/6 and Berkeley sickle mice (28) were purchased from The Jackson Laboratory (Bar Harbor, ME). Berkeley sickle mice carry two copies of a transgene carrying the human α- and sickle β-globin genes (SS) and are knock-outs for the mouse α- and β-globin genes. The genetic background of this strain is a mixture of FVB/N, 129, DBA/2, Black Swiss, and >50% C57BL/6 genomes. It was then backcrossed to C57BL/6J one generation to allow a predominant C57BL/6J background. Backcrossing into a homogenous C57BL/6J background results in an extremely severe sickle phenotype (with hemoglobin levels of 2–3 gm/dl), which is hard to propagate (29). Knock-in sickle (SS) and normal (AA) mice, where the human α- and β-globin loci are knocked in by homologous recombination in place of the respective mouse loci, were kindly provided by Dr. Timothy Townes (University of Alabama, Birmingham, AL) (30). PlGF knock-out mice (in the C57BL/6J strain), were provided by Dr. Peter Carmeliet (Vesalius Research Center, VIB-Vlaams Instituut voor Biotechnologie, Leuven, Belgium) (31). Mice were housed in the animal facility of the Cincinnati Children's Research Foundation using protocols approved by the Institutional Animal Care and Use Committee. Berkeley AA mice were provided by Dr. Cheryl Hillary, University of Wisconsin, Madison, WI. These mice carry two copies of the human α- and β-globin transgenes, are in the same mixed background as the Berkeley SS mice, and have been interbred with the Berkeley SS mice for six generations. Sex-matched males and females, between 8 and 24 weeks of age, were used for experiments; experimental arms had similar aged mice. Mice were bled via the retro-orbital sinus after isoflurane anesthesia in EDTA-containing tubes. Plasma was isolated and stored at −80 °C until assayed.
Human samples were obtained by routine clinical blood draws from children with homozygous SCD at steady state during their elective clinic appointment. We obtained the samples through the Hematology Repository, using an Institutional Review Board-approved protocol at the Cincinnati Children's Hospital Medical Center. The plasma and mononuclear cells (MNC) samples were derived from the same subjects as described previously (32); MNC were isolated as described previously (26).
Quantitative Real-time Reverse Transcription-PCR (qRT-PCR)
Total RNA from MNC was isolated using RNA Stat-60 (IsoTex Diagnostics Inc., Friendswood, TX), and qRT-PCR analysis was performed using iScript one-step RT-PCR kit with SYBR Green (Bio-Rad) and an ABI PRISM 7900 HT sequence detection system (Applied Biosystems, Foster City, CA), according to the manufacturer's instructions. Relative quantification values of PAI-1 mRNA expression were calculated as described previously (32).
Estimation of PAI-1 and Tissue Plasminogen Activator in Plasma or Bronchoalveolar Lavage (BAL)
PAI-1 levels in human plasma and cell culture supernatants were determined by ELISA. The MADPK-71K PAI-1 single plex kit (Millipore Corp., Billerica, MA) was used to assay mouse PAI-1 and read on a Bioplex analyzer (Bio-Rad). Tissue plasminogen activator (tPA) levels in plasma were determined using ELISA as per the manufacturer's instructions (Innovative Research, Novi, MI). Data are expressed as mean ± S.E. of three independent experiments with duplicate samples.
RNase Protection Assay (RPA)
RPA was performed using custom-made RiboQuant multi-probe templates comprised of PAI-1, TGF-β, HIF-1α, and glyceraldehyde-3-phosphate dehydrogenase (BD Biosciences), as described previously (33). Band intensity was analyzed using Spot-Denso software on an AlphaImager 2000 gel documentation system (Alpha Innotech Corp., San Leandro, CA). The mRNA levels were normalized to glyceraldehyde-3-phosphate dehydrogenase mRNA.
Western Blot Analysis
Cytosolic and nuclear extracts were prepared as described previously (34) and subjected to Western blot analysis. Protein bands were detected with Immobilon Western blot reagents (Millipore Corp.).
Mutagenesis of PAI-1 Promoter
Mutations in the WT pGL3-PAI-12345 promoter construct were performed by QuikChange site-directed mutagenesis kit according to the manufacturer's instructions (Stratagene, Cedar Creek, TX), using the primers listed in Table 1, and verified by automated DNA sequencing (Microchemical Core Facility, USC-NCCC).
TABLE 1.
Oligonucleotide primers used in this study
The abbreviations are: GAPDH, glyceraldehyde-3-phosphate dehydrogenase; mut, mutant; SDM, site-directed mutagenesis; oligo, oligonucleotide. Bold and underlined sequences highlight specific mutations.
| Gene/Fragment location | Method | Forward sequence (5′ to 3′) | Reverse sequence (5′ to 3′) |
|---|---|---|---|
| PAI-1 | qRT-PCR | gctgaattcctggagctcag | ctgcgccacctgctgaaaca |
| GAPDH | qRT-PCR | aacctgccaagtacgatgacatc | gtagcccaggatgcccttga |
| PAI-1 promoter AP-1 (−154/−147 bp)mut | SDM | cacacacacacatgccttgccaagtcccagagaggg | Ccctctctgggacttggcaaggcatgtgtgtgtgtg |
| PAI-1 promoter HRE-1 (−158/−151 bp)mut | SDM | cacacacatgtatgcctcagcaagtcccagagaggg | Ccctctctgggacttgctgaggcatacatgtgtgtg |
| PAI-1 promoter AP-1/HRE1 (−154/−151 bp)mut | SDM | cacacacacacaatgctcagcaagtcccagagaggg | Ccctctctgggacttgctgagcattgtgtgtgtgtg |
| PAI-1:WT oligo (−163/−143 bp) | EMSA | acacacacatgcctcagcaag | cttgctgaggcatgtgtgtgt |
| PAI-1:HRE-1/AP-1mut | EMSA | acacacaaaagccttggcaag | cttgccaaggcttttgtgtgt |
| PAI-1:AP1mut | EMSA | acacacacaagccttggcaag | cttgccaaggcttgtgtgtgt |
| PAI-1 promoter (−366/−82 bp) | ChIP | ctccaagctgaacactagggg | ctggctctgggagtccgtctg |
| PAI-1 promoter (−114/+98 bp) | ChIP | ccgctggctgttcagacggac | ctggcggcggctgctgagctg |
Transient Transfection of HPMVEC
HPMVEC (1 × 106 cells) were resuspended in 100 μl of Nucleofector solution containing β-galactosidase reporter plasmid and 1 μg of expression plasmid, 1 μg of promoter luciferase constructs, or 50 nm siRNA constructs followed by nucleofection using HMVEC-L Nucleofection kit (Lonza Group Ltd.). After transfection, cells were kept in complete medium overnight and serum-free medium for 3 h and then treated with PlGF for the indicated time. Cells were then lysed and analyzed for luciferase activity using a luminometer (Lumat LB 9501, Berthold, Bad Wildbad, Germany) and β-galactosidase activity using kits (Promega, Madison, WI). Luciferase values were normalized to β-galactosidase values. Data are expressed relative to the activity of the promoter-less pGL3 basic vector.
EMSA
Nuclear extracts from untreated and PlGF-treated HPMVEC were subjected to EMSA analysis using WT and mutant oligonucleotide probes (Table 1), as described previously (27).
Chromatin Immunoprecipitation (ChIP) Assay
HPMVEC (5 × 106 cells) were treated with PlGF in serum-free medium, in the presence or absence of inhibitors, for the indicated time. ChIP analysis was carried out by using either HIF-1α antibody or c-Jun antibody as described previously (35). PCR was set for 30 cycles under the following conditions: 95 °C for 30 s, 56 °C for 60 s, and 72 °C for 120 s, using the primers shown in Table 1. ChIP products were analyzed by agarose gel electrophoresis.
Immunohistochemistry
Lungs were fixed in 10% formalin, embedded in paraffin, and stained as described by Zhou et al. (36). Briefly, 5-μm paraffin sections were deparaffinized and then microwaved in buffer containing 0.1 m citric acid and 0.1 m sodium citrate (pH 6.0) at high power for 8 min and at mid-low power for 15 min (37). Sections were treated with 3% hydrogen peroxide for 30 min, blocked with 5% normal goat serum at room temperature for 2 h, and incubated overnight at 4 °C with anti-PAI-1 antibody (Molecular Innovations) or control rabbit IgG (Invitrogen). Slides were developed using the Vector Elite ABC method (Vector Laboratories; Burlingame, CA). Antigen localization was enhanced with nickel-diaminobenzidine, followed by incubation with Tris-cobalt (10 mm Tris, 0.5% cobalt chloride) for 4 min at room temperature, and counterstained with 0.1% Nuclear Fast Red for 2 min.
Statistical Analysis
Results are expressed as mean ± S.E. The significance of differences in mean values between untreated and PlGF-treated samples was determined by Student's t test. Parametric one-way analysis of variance followed by a Turkey-Kramer test was used to determine the significance of differences in mean values between multiple groups using Instat 2 software (GraphPad Software, San Diego, CA). ***, p < 0.001, **, p < 0.01, not significant or p > 0.05 (indicated by ns in Figs. 1–4).
FIGURE 1.
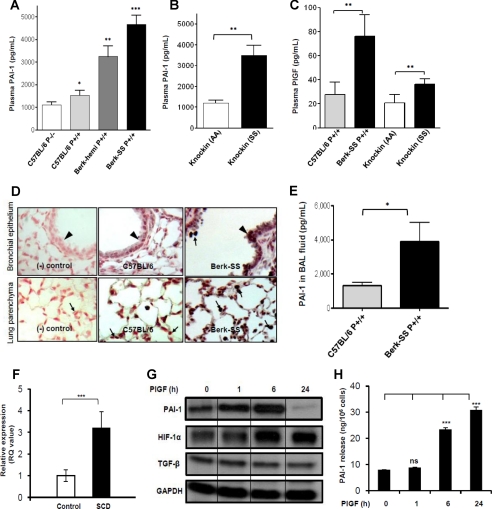
PAI-1 expression in mice and human subjects. A and B, plasma levels of PAI-1 in mice as measured by ELISA. Berk-hemi, hemizygous Berkeley mice; Berk-SS, Berkeley SS mice. C, plasma PlGF levels in mice by ELISA. The respective strain is indicated at the bottom of the figure. D, immunohistochemistry for PAI-1 expression in C57BL/6 (normal) and Berkeley SS mouse lungs (shown in brown; nuclei are stained red). The top panel shows: (i) antibody control and PAI-1 expression in bronchial epithelial cells of normal and (ii) Berkeley SS mice; the bottom panel shows corresponding staining for PAI-1 in their lung parenchyma. Arrows indicate alveolar macrophages, and arrowheads indicate bronchial epithelial layer. E, the levels of PAI-1 were quantified in BAL of indicated strains of mice by ELISA. F, qRT-PCR analysis of PAI-1 mRNA in MNC from SCD subjects (n = 9) and normal controls (n = 9). RQ, relative quantification. G and H, HPMVEC were treated with human recombinant PlGF (250 ng/ml) for the indicated time periods. G, total RNA was subjected to RPA for the expression of indicated genes. H, the culture supernatants were assayed for PAI-1 by ELISA. RPA data are representative of three independent experiments. GAPDH, glyceraldehyde-3-phosphate dehydrogenase. Where indicated, the vertical lines show repositioned gel lanes. *, p < 0.05, **, p < 0.01, ***, p < 0.001. ns, not significant.
FIGURE 2.
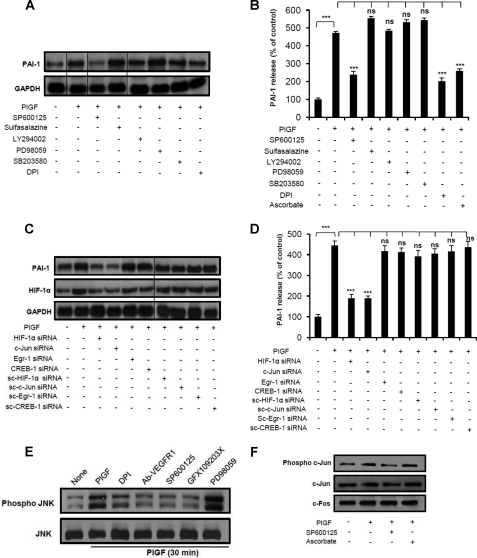
Cellular signaling pathway of PlGF-induced PAI-1 expression. A and B, HPMVEC were pretreated for 30 min with pharmacological inhibitors of PI3K, LY294002 (10 μm); Erk, PD98059 (10 μm); P38α, SB203580 (1 μm); JNK, SP600125 (0.1 μm); NADPH oxidase, DPI (10 μm); HIF-1α, ascorbate (25 μm), and NF-κB, sulfasalazine (2 mm) followed by treatment with PlGF for 6 h. GAPDH, glyceraldehyde-3-phosphate dehydrogenase. C and D, HPMVEC were transfected with indicated siRNA or scrambled (sc) siRNA constructs followed by PlGF treatment for 6 h. A and C, RPA analysis of the indicated genes. Where indicated, the vertical lines show repositioned gel lanes. Data are representative of three independent experiments. B and D, PAI-1 release by ELISA. The data are representative of three independent experiments. ***, p < 0.001. ns, not significant. E and F, HPMVEC were treated with PlGF either in the presence or in the absence of pharmacological inhibitors for 30 min. E, the cytosolic extracts were prepared and subjected to Western blotting for phospho-JNK, and the same blot was reprobed for total JNK. Ab, antibody. F, the nuclear extracts were analyzed for phospho-c-Jun, and the same blot was reprobed for total c-Jun and c-Fos. Data are representative of three independent experiments.
FIGURE 3.
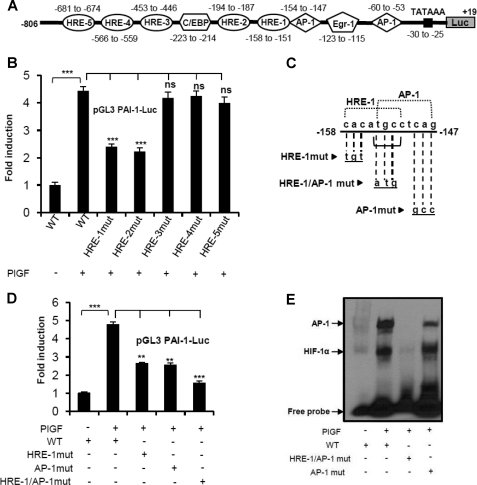
Identification of functional cis-acting elements in PlGF-induced PAI-1 promoter activation. A, schematics of PAI-1 promoter (−806/+19 bp, relative to transcription start site), indicating the location of HRE-1 to -5, proximal and distal AP-1, C/EBP, and Egr-1 binding sites. Luc, luciferase. B, HPMVEC were co-transfected with either WT or individual HRE mutant (mut) constructs of PAI-1 promoter and β-galactosidase plasmid prior to PlGF treatment for 6 h. C, illustrations of an overlapping region (−158/−147 bp) containing the HRE-1 and AP-1 binding sites in the PAI-1 promoter. The mutated oligonucleotides corresponding to HRE-1, AP-1, and the overlapping region of HRE-1 and AP-1 are underlined. D, HPMVEC were co-transfected with either WT or HRE-1 mutant or AP-1 mutant or HRE-1/AP-1 mutant of PAI-1 promoter and β-galactosidase plasmid prior to PlGF treatment for 6 h. The luciferase and β-galactosidase activities were estimated. E, nuclear extracts from untreated and PlGF-treated HPMVEC (10 μg) were incubated either with a biotinylated double-stranded oligonucleotide probe corresponding to an overlapping region containing HRE-1 and AP-1 sites or with a probe containing mutations in an overlapping four shared nucleotides or mutation only in the AP-1 site. Data are representative of three independent experiments. ns, not significant.
FIGURE 4.
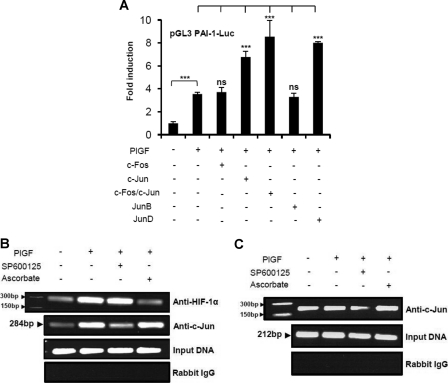
Role of individual components of AP-1 in PlGF-induced PAI-1 promoter activity. A, HPMVEC were co-transfected with expression plasmid for either c-Fos or c-Jun or both c-Fos + c-Jun or JunB or JunD along with the WT PAI-1 promoter and β-galactosidase plasmid prior to PlGF treatment for 6 h. Luciferase and β-galactosidase activities were estimated. PlGF promotes HIF-1α and AP-1 binding to PAI-1 promoter in vivo. ns, not significant. B and C, HPMVEC were pretreated with pharmacological inhibitors of either JNK (SP600125) or HIF-1α (ascorbate) prior to PlGF stimulation for 4 h. The soluble chromatin was isolated and immunoprecipitated with indicated antibody or control rabbit IgG. DNA was amplified with the primers (−366 to −82 bp) corresponding to HRE-1 and -2 and a distal AP-1 site (B) and the primers (−114 to +98 bp) corresponding to a proximal AP-1 binding site (C). The primer sequences are listed in Table 1. The input DNA panel represents the amplification of samples before immunoprecipitation. Data are representative of three independent experiments.
RESULTS
PAI-1 Expression in Sickle Cell Anemia Mice and Human Subjects
We analyzed plasma levels of PAI-1 in two different models of sickle mice with strain-appropriate normal controls: (i) with Berkeley SS mice with severe SCD (where mouse α- and β-globins are knocked out and human α- and β-sickle globins are present as a transgene), hemizygous Berkeley mice, Berkeley AA mice (humanized mice that carry the human α- and β-globin transgenes and express normal human hemoglobin A), and C57BL/6 mice serving as controls (Fig. 1A and supplemental Fig. S1A) (28) and (ii) knock-in SS mice with moderate SCD, where human βS- and α-globin genes replace the mouse globin genes at the endogenous loci, with their humanized counterparts (knock-in AA mice) serving as controls (Fig. 1B) (30). Berkeley SS mice and knock-in SS mice had 3-fold higher PAI-1 in plasma as compared with their corresponding C57BL/6 and AA controls. The Berkeley SS mice have more severe anemia and higher PlGF levels than the knock-in SS mice (Fig. 1C). Furthermore, PAI-1 plasma levels were significantly higher in Berkeley hemizygous mice as compared with normal Berkeley AA or C57BL/6 controls (supplemental Fig. S1A and Fig. 1A), the effect of the sickle gene dose. A temporal analysis of PAI-1 expression performed in the Berkeley SS (supplemental Fig. S1B) and knock-in SS mice (supplemental Fig. S1C) showed a linear increase in PAI-1 levels with increasing age, in contrast to the corresponding control mice. More importantly, plasma PAI-1 levels were lower in C57BL/6 PlGF−/− mice than in C57BL/6 PlGF+/+ controls. The immunohistochemical data showed significantly higher PAI-1 staining in endothelial cells, bronchial epithelial cells, and alveolar macrophages of Berkeley lungs as compared with C57BL/6 controls (Fig. 1D). Because PAI-1 can bind vitronectin and urokinase-type plasminogen activator, resulting in increased immunolabeling of sickle lungs, higher expression of PAI-1 in sickle tissue was confirmed by BAL fluid of Berkeley SS and C57BL/6 control mice (Fig. 1E) and PAI-1 mRNA expression in the cells obtained by BAL (alveolar macrophages and granulocytes) by real-time RT-PCR. There was a 2-fold higher PAI-1 mRNA expression in BAL cells. Circulating levels of tPA in Berkeley SS and knock-in SS mice were significantly lower than corresponding controls (supplemental Fig. S1D), consistent with reports of lower tPA plasma levels in patients with SCD (10, 11). To assess whether PlGF affected tPA levels, we stimulated primary HPMVEC with PlGF and measured release of tPA in the medium. A small but statistically insignificant increase in tPA was observed (supplemental Fig. S1E).
In humans, we confirmed earlier reports (10, 11) of high circulating levels of PAI-1 in plasma of SCD patients at steady state (17 ± 3.2 ng/ml) as compared with normal subjects (3.7 ± 1.5 ng/ml, p < 0.001; supplemental Fig. S2A). Plasma PlGF levels were also elevated in the same SCD subjects (24.7 ± 1.6 pg/ml versus 13.7 ± 0.4 pg/ml, p < 0.001; supplemental Fig. S2B). We also confirmed lower levels of tPA in plasma of SCD patients (2.74 ± 0.83 ng/ml) as compared with controls (1.26 ± 0.97 ng/ml, p < 0.001; supplemental Fig. S2C). Because PAI-1 immunolabeling was strong in alveolar macrophages and endothelial cells of sickle mice, we used real-time qRT-PCR to determine whether MNC from SCD subjects at steady state had increased expression of PAI-1. In fact, PAI-1 mRNA was 3-fold higher in MNC from SCD subjects as compared with controls (p < 0.001; Fig. 1F). We next exposed primary HPMVEC to PlGF and found that it induced PAI-1 mRNA in a time-dependent manner (Fig. 1G). TGF-β is a known autocrine inducer of PAI-1 mRNA (23). However, TGF-β mRNA expression remained unchanged in HPMVEC exposed to PlGF (Fig. 1G). Consistent with our prior observations, HIF-1α mRNA increased in response to PlGF stimulation (Fig. 1G). In addition, PAI-1 secretion from HPMVEC increased in response to PlGF stimulation (Fig. 1H).
PlGF-induced PAI-1 Expression Involves JNK and NADPH Oxidase
To determine the cell signaling mechanism of PlGF-induced PAI-1 expression in HPMVEC, we utilized pharmacological kinase inhibitors (supplemental Table S1). PlGF-induced PAI-1 mRNA expression was significantly inhibited by SP600125 but not by PD98059, SB203580, or sulfasalazine (Fig. 2A). We recently showed that PlGF increases endothelin-1 (ET-1) mRNA expression in HPMVEC by activation of phosphoinositide 3-kinase (PI3K), NADPH oxidase, and HIF-1α (27). We therefore examined the role of these pathways in regulation of PAI-1 expression. PlGF-induced PAI-1 mRNA expression in HPMVEC was inhibited by DPI but not by LY294002. Consistent with these data, we found that PlGF-induced PAI-1 secretion was significantly diminished by inhibitors of JNK, NADPH oxidase, and HIF-1α but not by PI3K inhibitor. Furthermore, transfection with either dominant negative PI3K (DnPI3K) or phosphatase and tensin homolog (negative regulator of PI3K/Akt) did not affect PlGF-induced PAI-1 mRNA expression or protein release (supplemental Fig. S3, A and B). Taken together, these results indicate that PlGF-mediated PAI-1 expression in HPMVEC involves activation of JNK and NADPH oxidase, but not PI3K, a mechanism distinct from ET-1 induction by PlGF.
PlGF-induced PAI-1 Expression Requires HIF-1α and c-Jun
The transcription factors Egr-1, HIF-1α, and C/EBPα have been shown to play an important role in regulation of the rat PAI-1 gene by hypoxia (20). AP-1 is another transcription factor, which has been implicated in the regulation of PAI-1 expression by oxidative stress (25) and by a fibrin fragment, D-dimer (18). Recently, angiotensin II-mediated PAI-1 expression has been shown to be regulated by CREB1 in rat cerebrovascular smooth muscle cells (38). We examined the role of these transcription factors in PlGF-mediated PAI-1 expression by transfecting HPMVEC with siRNA for each. Although PlGF-mediated expression of PAI-1 mRNA and protein was completely abrogated by HIF-1α and c-Jun siRNA, Egr-1, and CREB1 siRNA had no effect (Fig. 2, C and D).
PlGF Mediates Phosphorylation of Cytosolic JNK and Nuclear c-Jun
Because PlGF-induced PAI-1 expression was inhibited by SP600125, we wanted to determine whether PlGF activated JNK in HPMVEC. Interestingly, PlGF induced phosphorylation of both 46-kDa and 54-kDa isoforms of JNK, with maximal phosphorylation at 30 min followed by dramatic reduction at 60 min, likely due to its translocation into the nucleus (supplemental Fig. S4A). This effect was reduced by antibody to the cognate receptor of PlGF, VEGFR1, showing that the effect was specifically induced by PlGF. JNK phosphorylation was also reduced by DPI, SP600125, and GFX109203X; however, PD98059 had no effect (Fig. 2E). PlGF also increased the levels of phospho-c-Jun, a downstream target of JNK, in nuclear extracts; this increase was inhibited by SP600125 but not by ascorbate (Fig. 2F). However, levels of total c-Jun and c-Fos proteins remain unchanged. Moreover, PlGF caused a time-dependent increase in HIF-1α protein levels in cytosolic extracts of HPMVEC, an effect that was specifically attenuated by DPI and ascorbate but not by SP600125 (supplemental Fig. S4B). Taken together, our data indicate that PlGF-mediated activation of the JNK pathway leads to phosphorylation of nuclear c-Jun and activation of NADPH oxidase, resulting in increased HIF-1α.
PlGF-mediated PAI-1 Promoter Activation Requires HRE and AP-1 Binding Elements
An in silico analysis of the human PAI-1 promoter revealed five HREs and two AP-1, Egr-1, and C/EBP binding sites in the proximal promoter (GenBankTM accession number X13323; Fig. 3A). Additionally, we identified a 12-bp region from −158/−147 bp containing putative binding elements for both HIF-1α (HRE-1, −158/−151 bp) and AP-1 (−154/−147 bp), with an overlap of 4 bp. Previous studies suggested that the PAI-1 promoter responds to hypoxia and D-dimer via HRE-2 (17) and proximal AP-1 (−60 to −53 bp) (18) motifs, respectively. To identify the functional HIF-1α binding sites that respond to PlGF, HPMVEC were transfected with either WT PAI-1 promoter or individual HRE mutant PAI-1 promoter constructs followed by treatment with PlGF. A 4.5-fold increase in PAI-1 promoter activity was observed (Fig. 3B). This effect was reduced 50% by mutations in either HRE-1 or HRE-2 but not by mutations in HRE-3, -4, or -5. We also performed mutations of the binding sites for HRE-1, AP-1, and overlapping four shared nucleotides between them in the 12-bp region (Fig. 3C). PlGF-mediated PAI-1 promoter activity was attenuated by ∼50% in either the HRE-1 or the AP-1 binding site mutant (Fig. 3D), but the inhibition was significantly greater in the mutant for overlapping region (85%). The roles of HRE-1 and AP-1 in activating PAI-1 were further substantiated by their increased DNA binding activity in an EMSA. There were two bands of DNA-protein complexes. Mutation in the overlapping region dramatically decreased both complexes to nearly undetectable levels (Fig. 3E, third lane). When an oligonucleotide containing a mutation in the AP-1 site was used, there was reduced intensity in the upper band, but the lower band remained unchanged (Fig. 3E, fourth lane), suggesting that the upper and lower bands correspond to AP-1-DNA and HIF-1α-DNA binding complexes, respectively. PlGF-induced PAI-1 promoter activity was inhibited by VEGFR1 antibody, SP600125, GF109203X, and ascorbate but not by VEGFR2 antibody and PD98059 (supplemental Fig. S5). These results suggest that an overlapping region comprised of HRE-1 and AP-1 binding sites, as well as the HRE-2 binding site, are the functional elements required for PlGF-mediated PAI-1 promoter activity.
The Role of Individual Components of the AP-1 Transcription Complex
Because the AP-1 binding site (−154/−147 bp) was identified as one of the functional elements involved in PlGF-mediated PAI-1 promoter activity, we studied individual components of the AP-1 family. PlGF treatment of HPMVEC transfected with WT PAI-1 promoter resulted in a 3.6-fold increase in luciferase activity (Fig. 4A), whereas co-transfection with a c-Jun expression plasmid resulted in a 6.8-fold increase. Interestingly, although transfection with a c-Fos expression plasmid had no effect, co-transfection of both c-Fos and c-Jun expression plasmids had a synergistic effect, an 8.8-fold increase. Overexpression of JunD, but not JunB, increased PlGF-mediated PAI-1 promoter activity by 8-fold. Taken together, our data indicate that c-Jun homodimers or c-Fos/C-Jun heterodimers or JunD homodimers interact with the AP-1 binding site to regulate PAI-1 promoter activity.
ChIP analysis of PlGF-treated HPMVEC with HIF-1α antibody showed a 3-fold enrichment of HIF-1α binding at the PAI-1 promoter region (−366/−82 bp) containing the HRE-2 and the HRE-1/AP-1 overlapping binding sites (Fig. 4B, upper). No such enrichment of HIF-1α binding was seen at the HRE-3 to -5 binding sites (supplemental Fig. S6A). Pretreatment with ascorbate, but not with SP600125, reduced the expected PCR product by ∼80%. To determine whether AP-1 also binds to PAI-1 promoter in native chromatin in response to PlGF, chromatin from PlGF-treated HPMVEC was immunoprecipitated with c-Jun antibody (Fig. 4B, lower). PlGF caused a 3-fold enrichment of c-Jun at −366/−82 bp of PAI-1 promoter, which was reduced by SP600125 but not by ascorbate. ChIP with c-Jun antibody followed by amplification of the −114/+98-bp promoter region showed no enrichment of c-Jun (Fig. 4C, upper), suggesting that the proximal AP-1 site (−60/−53 bp) was not involved in PlGF-mediated PAI-1 promoter activation. As a negative control, the binding of c-Jun with PAI-1 promoter region (−704/−404 bp) that lacks the AP-1 binding site was performed, and no binding was observed (supplemental Fig. S6B). These results show that PlGF regulates PAI-1 promoter activity via HRE-2 and an overlapping region comprised of HRE-1 and AP-1 functional elements.
PlGF Contributes to PAI-1 Levels in Vivo
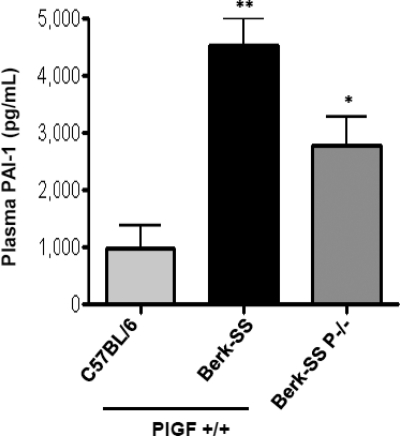
To conclusively implicate PlGF in PAI-1 induction, we interbred PlGF−/− mice and Berkeley SS mice. The former have very small litters, and the latter are difficult to breed due to the SCD. Indeed, very few Berkeley SS/PlGF−/− mice were generated. Nevertheless, plasma PAI-1 levels were significantly higher in Berkeley SS PlGF+/+ mice as compared with Berkeley SS PlGF−/− mice (Fig. 5). In both cases, PAI-1 levels were significantly higher than in control mice, suggesting that PlGF is one of the factors that contributes to increased PAI-1 in SCD. Next, we injected an adenovirus vector encoding PlGF or GFP cDNA into C57BL/6 mice. Published reports show that this method results in liver gene transfer in the short term, for 5–6 weeks. Ad-PlGF-injected mice had increased circulating PAI-1 levels, as compared with control Ad-GFP mice (supplemental Fig. S7).
FIGURE 5.
Role of PlGF in PAI-1 expression in sickle mice. Berkeley SS mice (Berk-SS) were bred with PlGF−/− mice to obtain Berkeley SS/PlGF−/− mice. The plasma levels of PAI-1 are measured by ELISA in the indicated strains of mice (n = 6).
DISCUSSION
PAI-1 is the primary physiologic inhibitor of both tPA and urokinase-type plasminogen activator and hence inhibits fibrinolysis. The factors that elevate PAI-1 in SCD (10, 11) are unknown. In this study, we observed increased PAI-1 expression in pulmonary endothelial cells and macrophages of sickle mice and identified the molecular mechanisms of PlGF-mediated PAI-1 induction in primary HPMVEC. Our results show that PlGF is a novel inducer of PAI-1 expression in HPMVEC. We further show that PlGF acts by activating JNK and NADPH oxidase but not PI3K, p38 mitogen-activated protein kinase (MAPK), or MAPK.
Previous reports have shown PAI-1 up-regulation in endothelial cells by several factors, including lipopolysaccharide (22), thrombin (39), TGF-β (40), IL-1 (41), insulin (24), and tumor necrosis factor-α (43), many of which are elevated in SCD. Lipopolysaccharide-mediated PAI-1 expression in macrophages involved activation of the JNK pathway (22). In contrast, insulin-mediated PAI-1 expression in hepatoma cells was shown to involve activation of PI3K (24) and MAPK (44) In addition, we reported previously that PlGF-mediated up-regulation of ET-1 in HPMVEC was mediated by activation of PI3K (27). In the present study, PlGF-mediated activation of JNK led to increased phosphorylation of nuclear c-Jun protein, whereas the total amount of nuclear c-Jun and c-Fos proteins remained unaltered. On the other hand, a previous study showed PlGF-mediated induction in c-Fos and FosB expression in human umbilical vein endothelial cells (45). The difference in the studies is perhaps the result of differences in cell type, dose, and/or duration of PlGF stimulation.
Our work also identifies a role for the transcription factors HIF-1α and c-Jun, but not Egr-1 and CREB1, in the PlGF-mediated PAI-1 expression. We observed an increase in HIF-1α mRNA and protein in HPMVEC in response to PlGF stimulation. Notably, HIF-1α induction occurred independent of hypoxia. Previous studies have shown that hypoxia-mediated PAI-1 gene expression in rats involves an interplay among Egr-1, HIF-1α, and C/EBPα transcription factors (20), and D-dimer-mediated PAI-1 expression in humans involves AP-1-response element (18). In contrast, angiotensin II has been shown to regulate PAI-1 expression through CREB1 (38). Additionally, studies have shown that there are at least five putative HRE-1 to -5 in the human PAI-1 promoter and that HIF-1α interacts only with HRE-2 to regulate PAI-1 transcription in response to hypoxia (17). Our study shows that PlGF induces activity of the human PAI-1 promoter via HRE-1 and -2 (but not HRE-3, -4, or -5), as well as the distal AP-1 site, and that the HRE-1 and AP-1 sites overlap. AP-1 is a family of dimeric transcription factors consisting of either Jun homodimers or Jun heterodimers of certain Jun/Fos members (46). We show here that PlGF-mediated PAI-1 promoter activation involved formation of c-Jun/c-Fos heterodimers, c-Jun homodimers, and JunD homodimers but not c-Fos and JunB homodimers. In the absence of PlGF stimulation, overexpression of c-Jun and c-Fos proteins did not result in the activation of PAI-1 promoter, indicating that phosphorylation of c-Jun/c-Fos is required for PlGF-mediated PAI-1 gene expression. PAI-1 expression mediated by D-dimer (18), oxidative stress (25), and nickel (47) has been shown to involve c-Fos/JunD binding to a proximal AP-1-response element in the PAI-1 promoter. Previous studies have shown that JunB and JunD display opposite effects, with JunD repressing and JunB activating heme oxygenase-1 expression in human renal epithelial cells (48). In contrast, we show that PlGF-mediated PAI-1 transcription is activated by JunD, and overexpression of JunB has no effect. Our results show that PlGF activates JNK, which causes downstream phosphorylation of c-Jun, resulting in formation of AP-1 complex, comprised minimally of c-Jun and JunD proteins.
The present study shows that PlGF-induced PAI-1 expression involves HIF-1α and AP-1. Indeed, humans with Chuvash polycythemia have constitutively elevated HIF-1α in the absence of hypoxia and have been reported to have elevations in plasma PAI-1 (49).
Thus, we show that PlGF increases PAI-1 expression through a unique signaling pathway. This is in contrast to our previous studies showing PlGF-mediated increased expression of inflammatory cytochemokines (26), ET-1 (27), and five lipoxygenase-activating proteins (32). Our in vitro finding that PlGF increases PAI-1 expression received added support from our animal data showing that PAI-1 levels were lowest in PlGF knock-out mice and highest in plasma of Berkeley SS mice and knock-in SS mice. Indeed, hemizygous Berkeley SS mice had PAI-1 levels midway between Berkeley SS and normal mice, and Berkeley SS mice had higher PAI-1 than knock-in SS mice. Moreover, the inverse was true for functional tPA. These data suggest that PAI-1 levels correlated to sickle gene dosage or severity. Our data showing that MNC of SCD patients at steady state had increased levels of PAI-1 and that PAI-1 was induced by PlGF in HPMVEC are novel; our observation that PlGF is also elevated in SCD patients is consistent with previous studies (10, 11).
The clinical consequences of elevation of PAI-1 can result from hypercoagulation, due to decreased fibrinolysis, or from the non-hemostatic effects of PAI-1, which could result in increased fibrosis. PAI-1, D-dimers, and tPA levels have been measured in patients with SCD at steady state and then following vaso-occlusion by Nsiri et al. (10). These investigators found that PAI-1 and D-dimer levels were elevated nearly 2.5–3-fold at steady state as compared with controls. However, no further increases in PAI-1 levels were seen during vaso-occlusive crises, although D-dimers were elevated further during vaso-occlusive events. In a separate study, the same group (11) showed that two-thirds of sickle cell disease patients showed a poor tPA response to vaso-occlusion. Indeed, our data in two strains of sickle mice are consistent with their results of lower tPA levels in SCD. Of note, their study showed that complications of SCD occurred predominantly in patients who showed a poor fibrinolytic response (defined as elevated PAI-1 or no increase in tPA with vaso-occlusion). Although these studies are performed on a small number of patients, they provide important leads for larger studies, necessary to address the antifibrinolytic and non-hemostatic effects of elevated PAI-1 in SCD.
We have previously shown that PlGF levels are also elevated in patients with other chronic hemolytic anemia, such as β-thalassemia (26). Although plasma PAI-1 levels have not been reported in thalassemia patients, it is interesting to note that a hypercoagulable and prothrombotic state, with elevated D-dimers, is present in this disease (42). Up-regulation of PAI-1 could be a contributory factor to the hypercoagulable state observed in β-thalassemia. Our PAI-1 and tPA data in two strains of sickle mice and in human subjects with SCD certainly show a state of reduced fibrinolysis, which would result in a prothrombotic phenotype. In conclusion, our studies show that PlGF, which is released from erythroid cells and, at greatly elevated levels, from sickle erythroid cells, leads to the expression and release of PAI-1 from HPMVEC. In addition to revealing a novel activator of PAI-1, this study provides the first demonstration that PlGF-induced PAI-1 expression involves the hypoxia-independent activation of HIF-1α and AP-1. Whether PlGF and high PAI-1 play an important role in development of a prothrombotic state or pulmonary fibrosis in SCD remains to be determined.
Acknowledgments
We thank Dr. Peter Carmeliet (Vesalius Research Center, VIB-Vlaams Instituut voor Biotechnologie, Leuven, Belgium) for kindly providing PlGF knock-out mice. We also thank Dr. Timothy Townes (University of Alabama, Birmingham, AL) for providing knock-in sickle mice and AA control mice, Dr. Trine Fink (University of Aalborg, Aalborg, Denmark) for kindly providing PAI-1 WT and individual HRE mutant promoter constructs, Dr. Michael Karin (University of California San Diego, La Jolla, CA) for providing c-Fos, c-Jun, and JunD expression plasmids, Dr. Anupam Agarwal (University of Alabama, Birmingham, AL) for JunB expression plasmid, and Dr. Debbie Johnson (USC Keck School of Medicine, Los Angeles, CA) for providing DnPI3K and phosphatase and tensin homolog plasmids. We thank Dr. Jeffrey Molkentin, Division of Cardiology, Heart Institute, Cincinnati Children's Hospital Medical Center, for providing us with the Ad-PlGF and Ad-GFP vector constructs. We are grateful to Jeff Bailey and Victoria Summey of the Comprehensive Mouse Core, Cincinnati Children's Hospital Medical Center for their assistance with murine injections and the Hematology Repository and the Translational Core Laboratory at the Cincinnati Children's Hospital Medical Center for processing and providing human samples. We also thank the Institutional Core of USC Research Center for Liver Disease for the use of a sequence detection instrument (supported by National Institutes of Health Grant P30-DK 048522) and Dr. Susan Wert, Perinatal Institute, CCHMC, for assistance with immunohistochemistry.
This work was supported, in whole or in part, by National Institutes of Health Grants R01-HL-079916 and HL-070595 (to P. M.).
This article was selected as a Paper of the Week.

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S7 and Tables S1 and S2.
- SCD
- sickle cell disease
- HIF-1α
- hypoxia inducible factor-1α
- HPMVEC
- human pulmonary microvascular endothelial cells
- HRE
- hypoxia-response elements
- PAI-1
- plasminogen activator inhibitor-1
- tPA
- tissue plasminogen activator
- PlGF
- placenta growth factor
- JNK
- c-Jun N-terminal kinase
- MAPK
- mitogen-activated protein kinase
- TGF
- transforming growth factor
- C/EBPα
- CCAAT/enhancer-binding protein α
- CREB1
- cAMP-response element-binding protein
- PI3K
- phosphoinositide 3-kinase
- MNC
- mononuclear cells
- WT
- wild type
- GFP
- green fluorescent protein
- BAL
- bronchoalveolar lavage
- RPA
- RNase protection assay
- qRT-PCR
- quantitative real-time reverse transcription-PCR
- ELISA
- enzyme-linked immunosorbent assay
- EMSA
- electrophoretic mobility shift assay
- ChIP
- chromatin immunoprecipitation
- siRNA
- small interfering RNA
- DPI
- diphenylene iodonium
- AA
- normal-homozygous adult globin
- SS
- sickle cell disease-homozygous sickle globin.
REFERENCES
- 1.Francis R. B., Jr., Johnson C. S. (1991) Blood 77, 1405–1414 [PubMed] [Google Scholar]
- 2.Gladwin M. T., Sachdev V., Jison M. L., Shizukuda Y., Plehn J. F., Minter K., Brown B., Coles W. A., Nichols J. S., Ernst I., Hunter L. A., Blackwelder W. C., Schechter A. N., Rodgers G. P., Castro O., Ognibene F. P. (2004) N. Engl. J. Med. 350, 886–895 [DOI] [PubMed] [Google Scholar]
- 3.Bunn H. F. (1997) N. Engl. J. Med. 337, 762–769 [DOI] [PubMed] [Google Scholar]
- 4.Key N. S., Slungaard A., Dandelet L., Nelson S. C., Moertel C., Styles L. A., Kuypers F. A., Bach R. R. (1998) Blood 91, 4216–4223 [PubMed] [Google Scholar]
- 5.Ataga K. I., Moore C. G., Hillery C. A., Jones S., Whinna H. C., Strayhorn D., Sohier C., Hinderliter A., Parise L. V., Orringer E. P. (2008) Haematologica. 93, 20–26 [DOI] [PubMed] [Google Scholar]
- 6.Berney S. I., Ridler C. D., Stephens A. D., Thomas A. E., Kovacs I. B. (1992) Am. J. Hematol. 40, 290–294 [DOI] [PubMed] [Google Scholar]
- 7.Ataga K. I., Key N. S. (2007) Hematology Am. Soc. Hematol. Educ. Program 91–96 [DOI] [PubMed] [Google Scholar]
- 8.Tomer A., Harker L. A., Kasey S., Eckman J. R. (2001) J. Lab. Clin. Med. 137, 398–407 [DOI] [PubMed] [Google Scholar]
- 9.Westerman M. P., Green D., Gilman-Sachs A., Beaman K., Freels S., Boggio L., Allen S., Zuckerman L., Schlegel R., Williamson P. (1999) J. Lab. Clin. Med. 134, 352–362 [DOI] [PubMed] [Google Scholar]
- 10.Nsiri B., Gritli N., Bayoudh F., Messaoud T., Fattoum S., Machghoul S. (1996) Hematol. Cell Ther. 38, 279–284 [DOI] [PubMed] [Google Scholar]
- 11.Nsiri B., Gritli N., Mazigh C., Ghazouani E., Fattoum S., Machghoul S. (1997) Hematol. Cell Ther. 39, 229–232 [DOI] [PubMed] [Google Scholar]
- 12.Hillery C. A., Panepinto J. A. (2004) Microcirculation 11, 195–208 [DOI] [PubMed] [Google Scholar]
- 13.Carmeliet P., Stassen J. M., Schoonjans L., Ream B., van den Oord J. J., De Mol M., Mulligan R. C., Collen D. (1993) J. Clin. Invest. 92, 2756–2760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bajou K., Noël A., Gerard R. D., Masson V., Brunner N., Holst-Hansen C., Skobe M., Fusenig N. E., Carmeliet P., Collen D., Foidart J. M. (1998) Nat. Med. 4, 923–928 [DOI] [PubMed] [Google Scholar]
- 15.Idell S. (2003) Crit. Care Med. 31, S213–S220 [DOI] [PubMed] [Google Scholar]
- 16.Sprengers E. D., Kluft C. (1987) Blood 69, 381–387 [PubMed] [Google Scholar]
- 17.Fink T., Kazlauskas A., Poellinger L., Ebbesen P., Zachar V. (2002) Blood 99, 2077–2083 [DOI] [PubMed] [Google Scholar]
- 18.Olman M. A., Hagood J. S., Simmons W. L., Fuller G. M., Vinson C., White K. E. (1999) Blood 94, 2029–2038 [PubMed] [Google Scholar]
- 19.Simpson A. J., Booth N. A., Moore N. R., Bennett B. (1991) J. Clin. Pathol. 44, 139–143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liao H., Hyman M. C., Lawrence D. A., Pinsky D. J. (2007) FASEB J. 21, 935–949 [DOI] [PubMed] [Google Scholar]
- 21.Nachman R. L., Hajjar K. A., Silverstein R. L., Dinarello C. A. (1986) J. Exp. Med. 163, 1595–1600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Arndt P. G., Young S. K., Worthen G. S. (2005) J. Immunol. 175, 4049–4059 [DOI] [PubMed] [Google Scholar]
- 23.Keeton M. R., Curriden S. A., van Zonneveld A. J., Loskutoff D. J. (1991) J. Biol. Chem. 266, 23048–23052 [PubMed] [Google Scholar]
- 24.Kietzmann T., Samoylenko A., Roth U., Jungermann K. (2003) Blood 101, 907–914 [DOI] [PubMed] [Google Scholar]
- 25.Vulin A. I., Stanley F. M. (2004) J. Biol. Chem. 279, 25172–25178 [DOI] [PubMed] [Google Scholar]
- 26.Perelman N., Selvaraj S. K., Batra S., Luck L. R., Erdreich-Epstein A., Coates T. D., Kalra V. K., Malik P. (2003) Blood 102, 1506–1514 [DOI] [PubMed] [Google Scholar]
- 27.Patel N., Gonsalves C. S., Malik P., Kalra V. K. (2008) Blood 112, 856–865 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Pászty C., Brion C. M., Manci E., Witkowska H. E., Stevens M. E., Mohandas N., Rubin E. M. (1997) Science 278, 876–878 [DOI] [PubMed] [Google Scholar]
- 29.Levasseur D. N., Ryan T. M., Pawlik K. M., Townes T. M. (2003) Blood 102, 4312–4319 [DOI] [PubMed] [Google Scholar]
- 30.Wu L. C., Sun C. W., Ryan T. M., Pawlik K. M., Ren J., Townes T. M. (2006) Blood 108, 1183–1188 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Carmeliet P., Moons L., Luttun A., Vincenti V., Compernolle V., De Mol M., Wu Y., Bono F., Devy L., Beck H., Scholz D., Acker T., DiPalma T., Dewerchin M., Noel A., Stalmans I., Barra A., Blacher S., Vandendriessche T., Ponten A., Eriksson U., Plate K. H., Foidart J. M., Schaper W., Charnock-Jones D. S., Hicklin D. J., Herbert J. M., Collen D., Persico M. G. (2001) Nat. Med. 7, 575–583 [DOI] [PubMed] [Google Scholar]
- 32.Patel N., Gonsalves C. S., Yang M., Malik P., Kalra V. K. (2009) Blood 113, 1129–1138 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Giri R. K., Selvaraj S. K., Kalra V. K. (2003) J. Immunol. 170, 5281–5294 [DOI] [PubMed] [Google Scholar]
- 34.Dignam J. D., Lebovitz R. M., Roeder R. G. (1983) Nucleic Acids Res. 11, 1475–1489 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kim K. S., Rajagopal V., Gonsalves C., Johnson C., Kalra V. K. (2006) J. Immunol. 177, 7211–7224 [DOI] [PubMed] [Google Scholar]
- 36.Zhou L., Lim L., Costa R. H., Whitsett J. A. (1996) J. Histochem. Cytochem. 44, 1183–1193 [DOI] [PubMed] [Google Scholar]
- 37.Shi S. R., Key M. E., Kalra K. L. (1991) J. Histochem. Cytochem. 39, 741–748 [DOI] [PubMed] [Google Scholar]
- 38.Rose P., Bond J., Tighe S., Toth M. J., Wellman T. L., Briso de Montiano E. M., Lewinter M. M., Lounsbury K. M. (2008) Am. J. Physiol. Heart. Circ. Physiol. 294, H1075–H1085 [DOI] [PubMed] [Google Scholar]
- 39.Gelehrter T. D., Sznycer-Laszuk R. (1986) J. Clin. Invest. 77, 165–169 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Petzelbauer E., Springhorn J. P., Tucker A. M., Madri J. A. (1996) Am. J. Pathol. 149, 923–931 [PMC free article] [PubMed] [Google Scholar]
- 41.Bevilacqua M. P., Schleef R. R., Gimbrone M. A., Jr., Loskutoff D. J. (1986) J. Clin. Invest. 78, 587–591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Eldor A., Rachmilewitz E. A. (2002) Blood 99, 36–43 [DOI] [PubMed] [Google Scholar]
- 43.Soeda S., Tsunoda T., Kurokawa Y., Shimeno H. (1998) Biochim. Biophys. Acta 1448, 37–45 [DOI] [PubMed] [Google Scholar]
- 44.Dimova E. Y., Kietzmann T. (2006) Ann. N. Y. Acad. Sci. 1090, 355–367 [DOI] [PubMed] [Google Scholar]
- 45.Holmes D. I., Zachary I. (2004) FEBS. Letters 557, 93–98 [DOI] [PubMed] [Google Scholar]
- 46.Shaulian E., Karin M. (2002) Nat. Cell Biol. 4, E131–E136 [DOI] [PubMed] [Google Scholar]
- 47.Andrew A. S., Klei L. R., Barchowsky A. (2001) Am. J. Physiol. Lung Cell. Mol. Physiol. 281, L616–L623 [DOI] [PubMed] [Google Scholar]
- 48.Hock T. D., Liby K., Wright M. M., McConnell S., Schorpp-Kistner M., Ryan T. M., Agarwal A. (2007) J. Biol. Chem. 282, 6875–6886 [DOI] [PubMed] [Google Scholar]
- 49.Gordeuk V. R., Prchal J. T. (2006) Semin. Thromb. Hemost. 32, 289–294 [DOI] [PubMed] [Google Scholar]