Abstract
Spider venoms provide a highly valuable source of peptide toxins that act on a wide diversity of membrane-bound receptors and ion channels. In this work, we report isolation, biochemical analysis, and pharmacological characterization of a novel family of spider peptide toxins, designated β/δ-agatoxins. These toxins consist of 36–38 amino acid residues and originate from the venom of the agelenid funnel-web spider Agelena orientalis. The presented toxins show considerable amino acid sequence similarity to other known toxins such as μ-agatoxins, curtatoxins, and δ-palutoxins-IT from the related spiders Agelenopsis aperta, Hololena curta, and Paracoelotes luctuosus. β/δ-Agatoxins modulate the insect NaV channel (DmNaV1/tipE) in a unique manner, with both the activation and inactivation processes being affected. The voltage dependence of activation is shifted toward more hyperpolarized potentials (analogous to site 4 toxins) and a non-inactivating persistent Na+ current is induced (site 3-like action). Interestingly, both effects take place in a voltage-dependent manner, producing a bell-shaped curve between −80 and 0 mV, and they are absent in mammalian NaV channels. To the best of our knowledge, this is the first detailed report of peptide toxins with such a peculiar pharmacological behavior, clearly indicating that traditional classification of toxins according to their binding sites may not be as exclusive as previously assumed.
Keywords: Insect, Ion Channels, Membrane Biophysics, Membrane Proteins, Neurobiology, Neurotoxin, Peptides, Protein-Protein Interactions, Sodium Channels, Spider Venom
Introduction
Spider venom is a rich mixture with a complex biochemical content, including proteins, peptides, and low molecular mass molecules (1–4). To attain high efficiency of the venom to paralyze or kill prey, many of the bioactive components specifically act on targets in the nervous system of the recipient, such as ion channels, and voltage-gated sodium (NaV)4 channels in particular. NaV channels are the trademark of electro-excitable cells, carrying the fast transient inward Na+ current during the depolarization phase of an action potential. They consist of a ∼260-kDa pore-forming α-subunit associated with auxiliary β-subunits of ∼30 kDa (β1–β4 for mammalian and tipE for insect NaV channels) (5–7). The α-subunit contains four homologous but non-identical repeats (DI–DIV), enclosing the ion conduction pore in a clockwise orientation. Each of these four repeats consists of six membrane-spanning segments (S1–S6). The first four segments (S1–S4) of each repeat comprise the voltage sensor domain with highly conserved positively charged amino acid residues (Arg or Lys) in the S4 segment serving as gating charges (5). The four positively charged S4 segments are thought to move outward through the membrane in response to depolarization, changing the channel conformation and thereby opening the ion conduction pathway to generate a transient Na+ current. Still, the exact mechanism underlying the gating process is poorly understood (8, 9). Segments S5 and S6 from all four repeats surround the pore of the channel, with the extracellular linker between S5 and S6 dipping back into the membrane to form the ion selectivity filter (5). The NaV channel is a common target for numerous xenobiotics, including animal toxins, therapeutic drugs, and insecticides (10, 11). These molecular probes can bind to at least seven different binding sites (so-called receptor sites) located on the α-subunit and thereby strongly alter normal channel functioning. Peptide toxins from spiders and other venomous animals can bind to at least four of these receptor sites (12, 13). Site 1 is located on the extracellular mouth of the channel and is targeted by a few peptide toxins, including μ-conotoxins and probably some spider toxins such as hainantoxins I and III–V and huwentoxin IV (14–17). Like the milestone tetrodotoxin and saxitoxin, these peptides are thought to act as a plug, physically blocking the pore of the NaV channel. Receptor sites 3, 4, and 6 are targeted by peptide toxins that modulate channel gating rather than block the pore of the channel. Scorpion α-toxins typically target site 3, together with some sea anemone and spider toxins, including Magi 4 and δ-atracotoxins (18–22). These site 3 toxins slow or inhibit the inactivation process of NaV channels. Scorpion β-toxins are known as the classic site 4 toxins and cause a hyperpolarizing shift of the activation of the NaV channel (46). Some spider toxins like Magi 5, δ-palutoxins-IT, μ-agatoxins, and curtatoxins are also reported to bind to site 4, although this is only speculative for the latter two groups (21, 23–25). Finally, site 6 is targeted by δ-conotoxins that slow the inactivation of NaV channels upon binding similar to site 3 toxins (26, 27). Site 6 has not been studied in detail and probably overlaps at least partially with site 3. Moreover, some peptide toxins, such as μO-conotoxins, may target receptor sites distinct from those described (28).
Because most spiders prey on insects, it does not seem to be very surprising that their venom contains toxins specifically designed to target insect NaV channels. To date, a number of spider toxins have been reported to modulate the functioning of insect NaV channels with a high selectivity (13, 29, 30). The insect NaV channels share a high primary structure similarity with their mammalian counterparts (31). Yet, insect NaV channels are much more conserved. Until today, only one gene (para) encoding NaV channels has been identified in insects and successfully expressed in a heterologous system, in contrast to mammalian NaV channels where nine distinct genes were found to encode for functionally distinct isoforms NaV1.1–1.9 (isoforms 1.1–1.8 successfully expressed heterologously) (32). In insect NaV channels, however, functional diversity is thought to be achieved mainly by alternative splicing and RNA editing (33, 34).
Agelena orientalis is an “Old World” spider from the family Agelenidae. These araneomorph spiders are sometimes referred to as “funnel-web” spiders but should not be confused with the Australian mygalomorph funnel-web spiders from the Hexathelidae family, which includes the infamous Atrax and Hadronyche genera (22). Certain functionally diverse groups of toxins have been isolated and characterized from the venom of agelenid spiders. Thus, agelenin isolated from Agelena opulenta is presumed to act on insect Ca2+ channels (35). Curtatoxins are insecticidal toxins from Hololena curta (25, 36). From Agelenopsis aperta, three groups of toxins were isolated and characterized: α-, μ-, and ω-agatoxins (24). The α- and ω-agatoxins are known to target glutamate receptors and voltage-gated Ca2+ channels, respectively, whereas the μ-agatoxins act on insect NaV channels. This latter group of toxins is known to shift the voltage dependence of sodium channel activation toward more hyperpolarized potentials (24, 37–40). Still, very few electrophysiological data showing the effects of μ-agatoxins on NaV channels are available today. In this study, we present isolation, biochemical analysis, and pharmacological characterization of β/δ-agatoxins, a novel group of toxins isolated from the venom of the agelenid spider A. orientalis. They are unique in their narrow voltage-dependent effects on both activation and inactivation of insect NaV channels.
EXPERIMENTAL PROCEDURES
Our generalized scheme for spider toxin purification, sequencing, and analysis may be found elsewhere (41).
Toxin Purification
Crude venom of the spider A. orientalis (Agelenidae) was purchased from Fauna Laboratories, Ltd. (Republic of Kazakhstan). Lyophilized venom was dissolved in distilled water (10 mg in 1 ml) and separated by reversed-phase high-performance liquid chromatography (RP-HPLC) on a Jupiter C5 column (4.6 × 150 mm, Phenomenex) using a 45-min linear gradient of acetonitrile concentration (0–20% in 5 min, 20–80% in 40 min) in 0.1% triethylamine (titrated with acetic acid to pH 10) at a flow rate of 1 ml/min. All fractions obtained were exposed to a second round RP-HPLC on the same column under acidic conditions using a 60-min linear gradient of acetonitrile concentration (0–60%) in 0.1% aqueous trifluoroacetic acid at a flow rate of 1 ml/min. Eluate absorbance was always monitored at 280 nm.
Mass Spectrometry
Molecular mass measurements were carried out using the method of matrix-assisted laser desorption-ionization (MALDI) on a M@LDI LR mass spectrometer (Micromass, UK) with identification of positive ions in linear mode. The 2,5-dihydroxy benzoic acid (10 mg/ml in 50% acetonitrile, 0.1% trifluoroacetic acid) matrix was used. Samples were prepared using the dried droplet method: equal volumes (0.5 μl) of sample and matrix were mixed on the target plate and left to dry in the air. Calibration was performed using the ProteoMass calibration kit with a molecular mass range of 700–66000 Da (Sigma).
Reduction of Disulfide Bonds and Modification of Thiol Groups
Dried samples were dissolved in 40 μl of solution containing 6 m guanidine hydrochloride, 3 mm EDTA, 0.5 m Tris-HCl (pH 8.5). Then 2 μl of 1.4 m 1,4-dithiothreitol was added, and the sample was incubated overnight at 30 °C. Four microliters of 50% 4-vinylpyridine in isopropanol was added to the samples, followed by incubation for 15–20 min at room temperature in the dark. Modified polypeptides were separated by RP-HPLC on a Jupiter C5 column (2 × 150 mm, Phenomenex) using a 60-min linear gradient of acetonitrile concentration (15–60%) in 0.1% trifluoroacetic acid at a flow rate of 0.3 ml/min.
Protein Sequencing
The N-terminal amino acid sequences of the alkylated peptides were determined on a Procise Model 492 protein/peptide sequencer (Applied Biosystems) according to the manufacturer's protocol.
Insect Toxicity Assay
Chromatographic fractions as well as purified toxins were assayed by injection into flesh fly Sarcophaga carnaria maggots (weighing ∼50–60 mg). Samples were dissolved in physiological saline: 140 mm NaCl, 5 mm KCl, 5 mm CaCl2, 1 mm MgCl2, 4 mm NaHCO3, and 5 mm HEPES (pH 7.2), and a volume of 2 μl was injected into the fourth segment of fly larva. Groups of 3–5 individuals were used for every tested concentration. Controls received pure saline. Paralytic and lethal effects were monitored for up to 24 h after the injection. Sample toxicities were expressed as median lethal doses (LD50) causing 50% mortality.
Expression of NaV Channels in X. laevis Oocytes
For expression in Xenopus laevis oocytes, cDNAs encoding rNaV1.2, DmNaV1, and tipE subunits were subcloned into vectors pLCT1 (42), pGH19-13-5 (7), and pGH19 (6), respectively. For in vitro transcription, these plasmids were linearized with NotI. Subsequently, capped cRNAs were synthesized from the linearized plasmid using the T7 mMESSAGE mMACHINE transcription kit (Ambion). The rβ1/pSP64T (43) plasmid was linearized with EcoRI and transcribed with the SP6 mMESSAGE mMACHINE transcription kit (Ambion).
Stage V–VI oocytes were harvested from the ovarian lobes of anesthetized female X. laevis frogs as described previously (44). The oocytes were injected with 50 nl of cRNA at a concentration of 1 ng/nl using a micro-injector (Drummond). The injected oocytes were incubated in a solution containing 96 mm NaCl, 2 mm KCl, 1.8 mm CaCl2, 2 mm MgCl2, and 5 mm HEPES (pH 7.4), supplemented with 50 mg/liter gentamycin sulfate.
Electrophysiological Experiments
Two-electrode voltage clamp recordings were performed at room temperature (18–22 °C) using a GeneClamp 500 amplifier (Molecular Devices) controlled by a pClamp data acquisition system (Molecular Devices). Whole cell currents from X. laevis oocytes were recorded 2–5 days after injection. Voltage and current electrodes were filled with 3 m KCl. Resistance of both electrodes was kept as low as possible (<1 MΩ). The bath solution composition was: 96 mm NaCl, 2 mm KCl, 1.8 mm CaCl2, 2 mm MgCl2, and 5 mm HEPES (pH 7.4). The evoked currents were sampled at 20 kHz and filtered at 1 kHz using a four-pole low-pass Bessel filter. Linear leak currents were corrected with a P/4 protocol. To avoid overestimation of a potential toxin-induced shift in the current-voltage relationships as a result of inadequate voltage control when measuring large sodium currents in oocytes, only data obtained from cells exhibiting currents with peak amplitude below 2 μA were considered for analysis.
For the electrophysiological characterization of the toxins, a number of voltage protocols were applied from a holding potential of −90 mV with a start-to-start pulse frequency of 0.2 Hz. (i) Current traces were evoked by 100-ms depolarizations to Vmax (i.e. the voltage corresponding to maximal INa in control). (ii) The current-voltage relationships were determined by 50-ms step depolarizations between −90 and 60 mV, using 5-mV increments. The sodium conductance (gNa) was calculated from the currents according to Ohm's law (Equation 1): gNa = INa/(V − Vrev), where INa represents the Na+ current peak amplitude at a given test potential V, and Vrev is the reversal potential. The values of gNa were plotted as a function of voltage and fitted using the Boltzmann equation (Equation 2): gNa/gmax = [1 + (exp(Vg − V)/k)]−1, where gmax represents maximal gNa, Vg is the voltage corresponding to half-maximal conductance, and k is the slope factor. (iii) To assess the toxin-induced effects on the steady-state inactivation, a standard two-step protocol was applied. In this protocol, 100-ms conditioning, 5-mV step prepulses ranging from −90 to 60 mV were followed by a 50-ms test pulse to −30 or −10 mV. Data were normalized to the maximal Na+ current amplitude, plotted against prepulse potential, and fitted using the Boltzmann equation (Equation 3): INa/Imax = [(1 − C)/(1 + exp((V − Vh)/k))] + C, where Imax is the maximal INa, Vh is the voltage corresponding to half-maximal inactivation, V is the test voltage, k is the slope factor, and C is a constant representing a non-inactivating persistent fraction (close to zero in control). (iv) The recovery from inactivation was assayed with a double-pulse protocol, where a 100-ms conditioning pulse to −30 or −10 mV was followed by a 50-ms test pulse to the same voltage. Both pulses were interspersed by a repolarization to −90 mV for a gradually increasing time interval (1–40 ms). The INa obtained in the test pulse was normalized to the INa obtained in the conditioning pulse, plotted against the corresponding time interval, and fitted with the following exponential equation (Equation 4): f(t) = Ae−t/τ + C, where t represents the time, A is the amplitude of the current, τ is the time constant for the fast inactivation, and C is a constant representing a non-inactivating persistent fraction (close to zero in control). The time constants (τ) of the NaV channel fast inactivation were measured directly from the decay phase of the recorded Na+ current using a single exponential fit (Equation 4). To assess the dose-response relationships, data were fitted with the Hill equation (Equation 5): y = 100/[1 + (EC50/[toxin])h], where y is the amplitude of the toxin-induced effect, EC50 is the toxin concentration at half-maximal efficacy, [toxin] is the toxin concentration, and h is the Hill coefficient.
Data were analyzed with pClamp Clampfit 8.0 (Molecular Devices) and Origin 6.1 software (OriginLab). Comparison of two sample means was made using a paired Student's t test (p < 0.05). All data are presented as means ± S.E. of at least three independent experiments (n ≥ 3).
Molecular Modeling
The model of the β/δ-agatoxin-1 spatial structure was built using the program MODELLER 9v7 (available on-line) on the basis of the known structure of the homologous δ-palutoxin-IT2 (PDB code 1V91). Structure visualization was performed using the program PyMOL (available on-line).
RESULTS
Toxin Isolation and Purification
Venom of the spider A. orientalis showed high insect toxicity. In flesh fly larvae, for instance, its LD50 was ∼3 μg/g. At 0.1 mg/ml, the crude venom showed a pronounced effect on the insect DmNaV1/tipE channel (activation facilitation and inactivation inhibition) and virtually no activity on mammalian NaV1.2, 1.5, or 1.7 channels in an X. laevis oocyte expression system (45). This venom was therefore utilized to identify novel selective molecules affecting insect NaV channels.
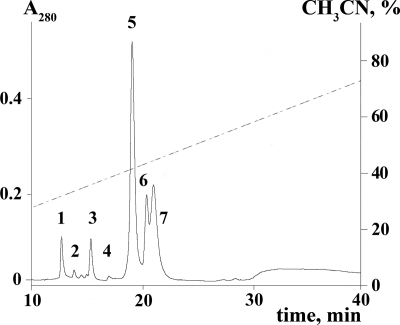
The new toxins were isolated using a bioassay-guided two-step RP-HPLC procedure. Crude A. orientalis venom was first separated on an RP column under basic conditions (Fig. 1), and the fractions active on DmNaV1/tipE channels were then again separated on the same column under acidic conditions (see “Experimental Procedures”). For proteomic analysis of the venom, see Ref. 60. In total, seven novel polypeptides were purified and named β/δ-agatoxins (see Table 1 for sequence data). Their homogeneity was verified by analytical HPLC and matrix-assisted laser desorption ionization mass spectrometry (the measured molecular masses are listed in Table 1). The major venom constituent corresponding to at least 30% of total polypeptide content (fraction 5 in Fig. 1) retained much of the crude venom toxicity (LD50 = 7 μg/g in flesh fly larvae) and was named β/δ-agatoxin-1 (β/δ-Aga-1). Other β/δ-agatoxins were numbered in accordance with the degree of their similarity to the major peptide (see sequence data below).
FIGURE 1.
Purification of β/δ-agatoxins. RP-HPLC of A. orientalis crude venom on a Jupiter C5 column (4.6 × 150 mm) using a linear gradient of acetonitrile concentration in 0.1% triethylamine (shown with a line). Numbered fractions contain toxins active on DmNaV1/tipE channels: 1, β/δ-Aga-5; 2, β/δ-Aga-4; 3, β/δ-Aga-6; 4, β/δ-Aga-7; 5, β/δ-Aga-1; 6, β/δ-Aga-2; and 7, β/δ-Aga-3.
TABLE 1.
Full-length sequences, average molecular masses, and accession numbers of β/δ-agatoxins
All peptides are C-terminally amidated (-NH2). Breaks are introduced to allow sequence comparison, cysteine residues are printed in bold, and residues that differ from those in β/δ-Aga-1 are marked gray.
Primary Structure
Alkylation of non-reduced molecules with 4-vinylpyridine revealed no free thiol groups in the native toxins. Full amino acid sequences of the new sodium channel modulators were established by automated Edman degradation. Direct N-terminal sequencing of reduced and alkylated peptides provided 36 amino acid residues for β/δ-Aga-5 and -6, 37 residues for β/δ-Aga-3, -4, and -7, and 38 residues for β/δ-Aga-1 and -2 (see Table 1 for sequence data). Each peptide contains eight cysteine residues forming four intramolecular disulfide bridges. Because of the 1-Da difference between the measured molecular masses and the corresponding calculated values based on the sequences, it was concluded that all seven peptides are C-terminally amidated.
Electrophysiological Study
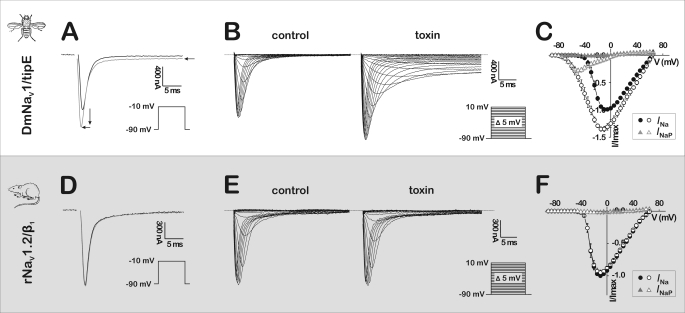
Because the spider A. orientalis is known to prey on small insects, we firstly investigated the effects of β/δ-Aga-1, the most abundant compound in the venom, on the cloned insect NaV channel DmNaV1/tipE from the fruit fly Drosophila melanogaster, expressed in X. laevis oocytes. On currents elicited by depolarizations to −10 mV (i.e. Vmax, the test potential corresponding to maximal INa), addition of 1 μm β/δ-Aga-1 resulted in three significant effects (Fig. 2A): (i) the peak of the transient sodium current (INa) increased by 35.2 ± 4.8% (n = 6), (ii) the time-to-peak of INa decreased from 1.07 ± 0.12 to 0.86 ± 0.11 ms (n = 6), and (iii) a persistent current (INaP) of 5.5 ± 2.2% (n = 6) of the peak was induced. Consequently, the toxin-induced effects on the current-voltage (I-V) relationships were assayed at 1 μm (Fig. 2B) and analyzed for peak currents (Fig. 2C, INa) and persistent currents (Fig. 2C, INaP). Clearly, β/δ-Aga-1 caused a profound hyperpolarizing shift in the INa-V relationship, without changing the reversal potential indicating that the ion selectivity of the DmNaV1/tipE channels was not altered. The toxin-induced persistent currents (INaP) could be observed only in the voltage range between −80 and 0 mV, following a bell-shaped INaP-V relationship with a maximal amplitude of 20.7 ± 2.4% of INa peak amplitude at −45 mV (n = 6). In contrast, β/δ-Aga-1 failed to affect the mammalian rNaV1.2/β1 channel from Rattus norvegicus (see Fig. 2, D–F) at concentrations up to 1 μm.
FIGURE 2.
Activity of β/δ-Aga-1 on NaV channels. Typical effects induced by β/δ-Aga-1 on currents from the cloned insect Na+ channel DmNaV1/tipE from D. melanogaster (A–C) and the cloned mammalian Na+ channel rNaV1.2/β1 from R. norvegicus (D–F), expressed in X. laevis oocytes. A and D, representative whole cell current traces in control (black trace) and in the presence of 1 μm β/δ-Aga-1 (gray trace). Arrows indicate the toxin-induced effect on the current of DmNaV1/tipE: an increase in peak amplitude, a decrease in time-to-peak, and the generation of a persistent current. B and E, superimposed representative whole cell current traces elicited by 5-mV step depolarizations between −90 and 10 mV in control (left) and in the presence of 1 μm β/δ-Aga-1 (right). The dotted line indicates the zero current level. C and F, normalized current-voltage relationships (n = 6 for DmNaV1/tipE and n = 3 for rNaV1.2/β1) of transient peak currents (INa, black circles) and persistent currents measured after 50 ms (INaP, gray triangles) in control (closed symbols) and in the presence of 1 μm β/δ-Aga-1 (open symbols).
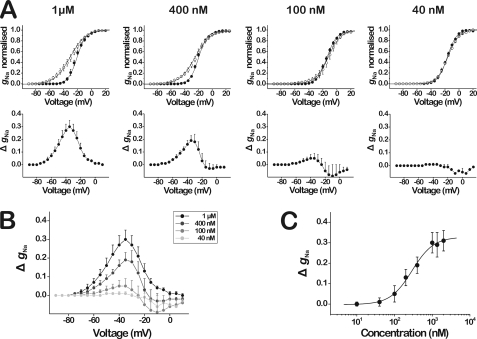
To further investigate the effects of β/δ-Aga-1 on the activation of the insect DmNaV1/tipE channel, the Na+ conductance (gNa) was calculated from INa peak amplitude according to Ohm's law (Equation 1), normalized, plotted against voltage, and fitted with the Boltzmann equation (Equation 2), yielding the steady-state activation curve (Fig. 3A, top panels). At a concentration of 1 μm, the toxin induced a large hyperpolarizing shift marked by a prominent change in the slope of the curve from 6.3 ± 0.3 to 11.3 ± 0.6 ms (n = 6). The threshold potential for activation was shifted approximately from −50 to −80 mV, whereas the potential corresponding to maximal activation was not changed. The potential corresponding to half-maximal activation (Vg) was shifted from −20.8 ± 0.5 to −30.4 ± 1.0 mV (n = 6). To display the voltage range wherein the action of toxin on DmNaV1/tipE channel activation takes place, the normalized gNa values in control were subtracted from the normalized gNa values in the presence of toxin. The resulting ΔgNa was then plotted against voltage (see Fig. 3A, bottom panels). The voltage dependence thus found revealed a bell-shaped relationship in the voltage range between −80 and 0 mV reaching a maximal amplitude around −35 mV. When measured at different toxin concentrations, a concentration- and voltage-dependent bell-shaped relationship was observed (see compilation, Fig. 3B). To assess the dose-response relationship (Fig. 3C), the ΔgNa (subtracted conductances) measured at −35 mV were plotted against toxin concentration and fitted with the Hill equation (Equation 5) yielding an EC50 value of 288.2 ± 116.4 nm (n = 3–6, Hill coefficient = 1.5 ± 0.7).
FIGURE 3.
Bell-shaped voltage dependence of the effects of β/δ-Aga-1 on the activation of the insect Na+ channel DmNaV1/tipE. A, upper panels show the steady-state activation curves (n = 3–6) in control (closed circles) and in the presence of varying concentrations of β/δ-Aga-1 (open circles). All steady-state activation curves are fit with the Boltzmann equation. The bottom panels display the subtraction (toxin minus control) of the normalized gNa values (ΔgNa), revealing the voltage range wherein the toxin-induced effects on the activation occur. B, superimposed curves of the ΔgNa at varying concentrations of β/δ-Aga-1 (see inset at upper right), showing the concentration dependence of the toxin-induced effects on the activation. C, dose-response curve of the ΔgNa measured at −35 mV (i.e. the voltage at which the effect of toxin on gNa is maximal). The data are fit with the Hill equation, yielding an EC50 value of 288.2 ± 116.4 nm (n = 3–6; Hill coefficient is 1.5 ± 0.7).
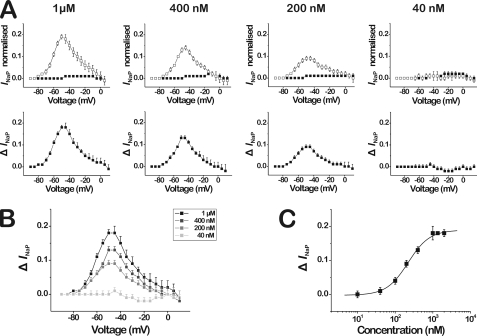
The toxin-induced persistent currents (INaP) were analyzed in an analogous manner as the effect of toxin on the activation of DmNaV1/tipE. The INaP was quantified by measuring the current amplitude after 50 ms and normalizing to the maximal current amplitude in control (not necessarily at the same voltage). The thus obtained normalized INaP values were then plotted against voltage (Fig. 4A, top panels). To display the voltage range wherein the toxin-induced persistent currents take place, control values for normalized INaP were subtracted from normalized INaP in the presence of the toxin (see Fig. 4B, bottom panels). The resulting ΔINaP followed a bell-shaped voltage dependence, reaching a maximum around −45 mV. When tested at different toxin concentrations, a concentration- and voltage-dependent bell-shaped relationship was observed (see compilation, Fig. 4B). To obtain the dose-response relationship (Fig. 4C), the ΔINaP (subtracted persistent currents) measured at −45 mV were plotted against toxin concentration and fitted with the Hill equation (Equation 5) yielding an EC50 value of 220.2 ± 32.9 nm (n = 3–6, Hill coefficient = 1.5 ± 0.4).
FIGURE 4.
Bell-shaped voltage dependence of the persistent currents (INaP) induced by β/δ-Aga-1 on the insect Na+ channel DmNaV1/tipE. To quantify INaP, the current amplitude after 50 ms was normalized to peak current amplitude. A, upper panels show the normalized INaP (n = 3–6) in control (closed circles) and in the presence of varying concentrations of β/δ-Aga-1 (open circles). The bottom panels display the subtraction (toxin minus control) of the normalized INaP values (ΔINaP), revealing the voltage range wherein the toxin-induced persistent currents occur. B, superimposed curves of the ΔINaP at varying concentrations of β/δ-Aga-1 (see inset at upper right), showing the concentration dependence of the toxin-induced persistent currents. C, dose-response curve of the ΔINaP at −45 mV (i.e. the voltage at which the toxin-induced persistent currents are maximal). The data are fit with the Hill equation, yielding an EC50 value of 220.2 ± 32.9 nm (n = 3–6, Hill coefficient 1.5 ± 0.4).
For a better understanding of the mechanism behind the persistent currents induced by β/δ-Aga-1, we continued by investigating the toxin effects on the DmNaV1/tipE inactivation process into more detail. Firstly, the steady-state inactivation was assayed with a typical double-pulse protocol (Fig. 5A). When the test pulse voltage was set at −10 mV (i.e. Vmax, the voltage corresponding to maximal current in control), 1 μm β/δ-Aga-1 did not change Vh (i.e. the midpoint of the steady-state inactivation curve). Yet, a small but significant non-inactivating component (5.3 ± 0.6%; n = 6) was induced at prepulse voltages more depolarized than −40 mV. Subsequently, the test potential was chosen more centrally in the voltage range where the persistent currents occur. At a test potential of −30 mV, Vh was again not significantly shifted (from −46.7 ± 0.1 to −50.0 ± 0.3 mV), but the non-inactivating component was now increased up to 15.8 ± 0.6% (n = 3). The effects of β/δ-Aga-1 on the recovery from inactivation of DmNaV1/tipE were examined next (see Fig. 5B). At a test potential of −10 mV, β/δ-Aga-1 decreased the time needed to recover from inactivation. The time constant for recovery from inactivation significantly decreased from 2.56 ± 0.11 ms in control to 1.65 ± 0.13 ms in the presence of 1 μm β/δ-Aga-1 (n = 4). When the test potential was set at −30 mV, the time constant for recovery from inactivation decreased significantly from 2.86 ± 0.08 to 1.77 ± 0.06 ms (n = 4). Interestingly, the channels not only recovered faster from inactivation in the presence of the toxin, they also yielded significantly higher currents (up to 114.2 ± 1.0%) in the test pulse compared with the conditioning pulse when short interpulse intervals (4–28 ms) were applied. These unusually high normalized currents (>1) decreased with increasing interpulse time interval, to return to unity when interpulse intervals were 28 ms or longer. To test whether the toxin altered the real-time kinetics of fast inactivation of DmNaV1/tipE, the time constants of inactivation were measured directly from the decay phase of currents with a mono-exponential fit as described by Equation 4. The use of a multiexponential equation did not yield beneficial fitting results over the mono-exponential equation. At voltages more depolarized than −25 mV, 1 μm β/δ-Aga-1 did not significantly change the time constant of inactivation (see Fig. 5, n = 4). At more negative potentials than −25 mV, however, the time constants of inactivation significantly decreased in the presence of 1 μm β/δ-Aga-1 (n = 4).
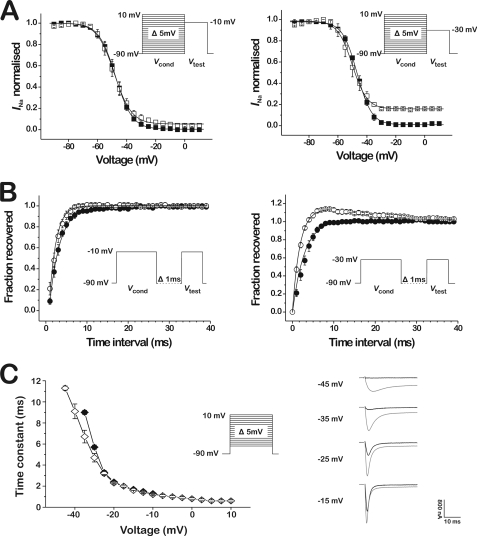
FIGURE 5.
Effect of β/δ-Aga-1 on the inactivation process of the insect Na+ channel DmNaV1/tipE. A, steady-state inactivation curves (n = 3–6) in control (closed squares) and in the presence of 1 μm β/δ-Aga-1 (open squares). Test voltage is set at −10 mV in the left graph and −30 mV in the right graph. B, recovery from inactivation (n = 4) in control (closed circles) and in the presence of 1 μm β/δ-Aga-1 (open circles). Test voltage is set at −10 mV in the left graph and −30 mV in the right graph. C, voltage dependence of the time constants of fast inactivation of DmNaV1/tipE (n = 4) in control (closed diamonds) and in the presence of 1 μm β/δ-Aga-1 (open diamonds). On the right, superimposed representative current traces in control (black traces) and in the presence of 1 μm β/δ-Aga-1 (gray traces) at different test potentials.
Six other peptides (designated β/δ-agatoxins-2–7) that are highly homologous to β/δ-Aga-1 were isolated from the venom of A. orientalis (see Table 1). These naturally occurring homologues displayed a similar dual electrophysiological effect on the insect channel DmNaV1/tipE as β/δ-Aga-1, causing a hyperpolarizing shift in the channel activation and inducing persistent currents. The ΔgNa and ΔINaP values were plotted against voltage for each homologue in an analogous manner as in Figs. 2 and 3. Similar to the observations with β/δ-Aga-1, these curves describe a strong bell-shaped voltage dependence (Fig. 6A). To facilitate a quantitative comparison, all homologues (β/δ-Aga-1–7) were tested at a concentration of 1 μm (n = 3–6). The amplitudes of ΔgNa and ΔINaP, taken at −35 and −45 mV, respectively, were normalized by setting the β/δ-agatoxin homologue with the highest amplitude as 100% (Fig. 6B). Interestingly, β/δ-agatoxins 4 (2.4 ± 2.4% for ΔgNa and 9.1 ± 4.5% for ΔINaP) and 7 (26.2 ± 11.9% for ΔgNa and 18.2 ± 9.1% for ΔINaP) appeared to affect the insect channel only weakly, compared with β/δ-agatoxins 2 (66.7 ± 11.9% for ΔgNa and 100.0 ± 18.2% for ΔINaP) and 3 (100.0 ± 11.9% for ΔgNa and 90.9 ± 13.6% for ΔINaP), which are comparable in amplitude to β/δ-Aga-1 (71.4 ± 11.9% for ΔgNa and 81.8 ± 9.1% for ΔINaP). Two other homologues, β/δ-agatoxins 5 and 6, shifted the activation of DmNaV1/tipE with a similar amplitude as β/δ-Aga-1 (ΔgNa of 92.9 ± 26.2% and 78.6 ± 21.7%, respectively), but induced persistent currents to a significantly lesser extent than β/δ-Aga-1 (ΔINaP of 36.4 ± 4.5% and 36.4 ± 13.6%, respectively).
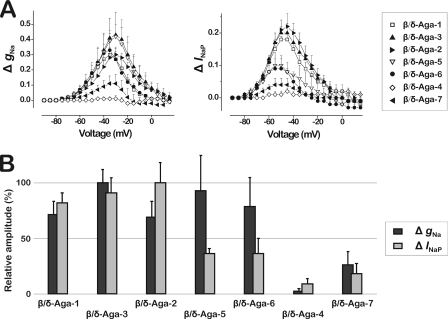
FIGURE 6.
Comparison of the effects of the β/δ-agatoxin homologues on the insect Na+ channel DmNaV1/tipE. To facilitate a proper comparison, all data represent experiments conducted at a toxin concentration of 1 μm (n = 3–6). A, superimposed bell-shaped curves showing the amplitude of the effects of β/δ-agatoxin homologues on the activation (left graph) and the persistent currents (right graph). Analysis of the toxin-induced effects on activation (ΔgNa) and persistent currents (ΔINaP) were done in the same manner as in Figs. 3 and 4, respectively. B, bar diagrams showing the relative amplitudes of the effects on the activation (represented by ΔgNa measured at −35 mV) and the persistent currents (represented by ΔINaP measured at −45 mV). Amplitudes were normalized by setting the β/δ-agatoxin homologue with the highest effect as 100%.
DISCUSSION
In this work, we describe β/δ-agatoxins, a new group of peptides found in A. orientalis spider venom, specifically targeting insect NaV channels. The electrophysiological profile of β/δ-agatoxins was assayed on cloned insect and mammalian NaV channels expressed in X. laevis oocytes, revealing, to the best of our knowledge, a novel and unique behavior. The studied toxins cause a 3-fold effect on the currents mediated by the insect DmNaV1/tipE channel: (i) an increase of peak sodium current, (ii) a decrease in time-to-peak, and (iii) generation of a non-inactivating persistent current. In contrast, the mammalian rNaV1.2/β1 channel was not affected by β/δ-Aga-1, which was somewhat expected, as the natural prey of A. orientalis does not include mammals.
β/δ-Agatoxins shift the current-voltage relationships of insect DmNaV1/tipE channels toward more hyperpolarizing potentials, comparable to typical site 4 toxins such as scorpion β-toxins, which could explain the increase in peak current amplitude at low depolarizations. However, β/δ-agatoxins differ in their actions from scorpion β-toxins in that the latter generally cause a small current inhibition (<15%) at the voltage of maximal activation (46–49). Moreover, the shift in the voltage dependence of activation observed with β/δ-agatoxins is marked by a significant increase in the slope factor of the activation curve, indicating that the cooperativity between the four S4 segments is changed, which is not the case with the classic β-toxins.
Importantly, the foremost prominent difference with typical site 4 toxins is the persistent current generated by β/δ-agatoxins. In fact, the increase in peak current amplitude and the appearance of persistent currents macroscopically resemble the effects of typical site 3 toxins, such as scorpion α-toxins, which are known to slow the fast inactivation of NaV channels. However, the observed actions of β/δ-agatoxins differ thoroughly from the actions of typical site 3 toxins. The β/δ-agatoxin-induced persistent currents only took place in the voltage range between −80 and 0 mV, following a bell-shaped voltage dependence with a maximal amplitude around −45 mV. In addition, β/δ-Aga-1 did not change the time constants of inactivation (τ) at voltages more positive than −25 mV and even caused a decrease in τ at potentials below −25 mV rather than an increase, which would have been expected if the toxin actually slowed the channel inactivation. The interpretation of this unexpected observation is difficult because of the dual effect of the toxin. Besides the apparent slowing of the inactivation that could result in the persistent currents, β/δ-Aga-1 also inflicts a hyperpolarizing shift in the voltage dependence of activation, causing the channels to open at lower depolarizations compared with control conditions. Therefore, currents in the presence of β/δ-Aga-1 between −80 and −25 mV have an increased peak amplitude and a faster inactivation than currents elicited at the same voltage in control situation (see current traces in Fig. 5C). β/δ-Aga-1 did not shift Vh nor alter the slope of the steady-state inactivation curve. Yet, a small but significant component of non-inactivating channels was observed at prepulse potentials more depolarized than −40 mV, a phenomenon previously observed with site 3 toxins (20, 50, 51). This small non-inactivating component was observed using a classic steady-state inactivation voltage protocol with the test potential set at Vmax (−10 mV). When the test potential was chosen at −30 mV, a potential more centrally located in the voltage range where the toxin-induced effects take place, the non-inactivating component was increased significantly. If one assumes a bimodal gating scheme in the modulated receptor mechanism as proposed by Hondeghem and Katzung (52), it could be deduced that the toxin promotes the inactivated-to-open state transition (as indicated by the non-inactivating component in the steady-state inactivation; see Fig. 5A) and the closed-to-open state transition (as observed in real-time by the decreased time-to-peak and in steady-state by the hyperpolarizing shift in activation; see Fig. 2, A–C). In addition, β/δ-Aga-1 increased the rate of recovery from inactivation of DmNaV1/tipE (see Fig. 5B), which may indicate that the toxin promotes the transition from the inactivated to the closed state of the toxin-bound channel. A remarkable observation from the recovery protocol is that the normalized currents in the presence of β/δ-Aga-1 exceed unity with short interpulse intervals (4–28 ms). This “additional” normalized current (Itest > Icond) gradually decreases toward unity as interpulse intervals increase. Apparently, when using such a double-pulse protocol, the duration of repolarization to −90 mV between the two pulses is crucial for unbinding of the toxin from the channel. The “additional” normalized current (Itest > Icond) observed in the recovery protocol with short interpulse intervals could be explained by the promotion of the inactivated-to-open state transition by the toxin, which leads to a higher fraction of channels that return to the open state in the test pulse compared with the conditioning pulse. As the repolarization interval increases, the fraction of unbound channels increases, and this effect decreases. All together, these observations show that binding of β/δ-Aga-1 relatively favors the open state of the DmNaV1/tipE channel, giving rise to larger sodium influx in the narrow bell-shaped voltage range, with important consequences in vivo.
Amino acid sequences of β/δ-agatoxins 1–7 show considerable similarity to some other known spider toxins, including μ-agatoxins 1–6 from A. aperta (from 45.9% between β/δ-Aga-1 or -2 and μ-Aga-2 up to 91.9% between β/δ-Aga-7 and μ-Aga-2), curtatoxins 1–3 from H. curta (from 38.9% between β/δ-Aga-7 and curtatoxin-1 up to 75% between β/δ-Aga-5 or 6 and curtatoxin-1) (both spiders are from the same family Agelenidae; μ-Aga-3 and curtatoxin-2 have identical sequences), and δ-palutoxins-IT from Paracoelotes luctuosus (family Amaurobiidae, related to Agelenidae) (from 30.6% between β/δ-Aga-6 and δ-Palu-IT3 up to 81.1% between β/δ-Aga-7 and δ-Palu-IT4) (see Fig. 7). In effect, all of these toxins form a large family of related peptides affecting insect NaV channels. Apart for overall sequence similarity, all of these toxins (except for δ-Palu-IT3) are C-terminally amidated and contain eight strictly conserved half-cystine residues that most probably account for a presumably common molecular fold. To date, spatial structures of μ-Aga-1 and δ-palutoxins-IT1 and -2 are known, all showing a high degree of similarity (53, 54) and conforming to the inhibitor cystine knot (ICK) motif most common for spider venom neurotoxins (4, 55). Even in the absence of experimental data proving the exact disulfide connectivity, it is therefore not unreasonable to surmise the typical ICK motif for the β/δ-agatoxins. Moreover, all of these toxins seem to exhibit high taxon specificity, effectively discriminating between insect and mammalian NaV channels (being inactive against the latter).
FIGURE 7.
Comparison of the amino acid sequences of β/δ-agatoxins and similar spider toxins. Sequence identities are shown relative to β/δ-Aga-1, β/δ-Aga-6, and β/δ-Aga-7. Amino acid residues identical with β/δ-Aga-1 are shown on a black background, conservative substitutions are shown on a gray background, and cysteine residues are printed in bold. Breaks (dashes) have been introduced to maximize alignment. All the aligned toxins except for δ-Palu-IT3 are C-terminally amidated, as indicated by asterisks.
For β/δ-agatoxins, the corresponding mRNA sequences from venom glands were established, and the precursor prepropeptide structures were thus deduced (56). The sequences are organized in the conventional manner, with N-terminal signal peptides, intervening profragments and C-terminal mature peptides (57, 58). Limited proteolysis occurs at the processing quadruplet motifs: XXXR, where any X = E. All precursors bear C-terminal extensions -G or -GK serving signals of C-terminal amidation and confirming our protein sequencing results.
Historically the first discovered μ-agatoxins from A. aperta venom (37, 40) were later shown to act specifically on insect NaV channels (24, 38, 39). Most notably, the toxins μ-Aga-1 and -4 studied in detail were shown to shift the voltage-activation curve to more negative potentials similar to scorpion β-toxins. However, they also slowed NaV channel inactivation and produced a non-inactivating persistent current. These effects have not been explored in detail but at least for the best studied μ-Aga-1 and -4 qualitatively resemble those described by us for the β/δ-agatoxins. Thus, we believe that the “μ-” prefix is misleading, because it has a more widespread pore-blocking mechanism connotation and propose that these toxins be renamed accordingly. Curtatoxins from H. curta venom (25) have not been studied in detail but are believed to act similarly to μ-agatoxins, δ-palutoxins-IT (δ-Palu-IT), and the newly discovered β/δ-agatoxins based on their sequence similarity. To date, δ-palutoxins-IT from P. luctuosus venom (59) are probably the best studied toxins from the family. Unlike the homologous μ-agatoxin-1 and -4 and β/δ-agatoxins, δ-palutoxins-IT1 and -2 do not affect NaV channel activation but instead induce effects resembling scorpion α-toxins, i.e. antagonize channel inactivation. However, profound differences between δ-Palu-IT1 and -2 and scorpion α-toxins were noted, the former competing with scorpion β-toxins for binding to receptor site 4 and not influencing the kinetics of channel inactivation (23, 53). It was therefore suggested that δ-Palu-IT1 and -2 only macroscopically resemble typical scorpion α-toxins but, unlike them, influence the inactivated-to-closed state transition process (53).
Although numerous exceptions exist, high polypeptide sequence similarity generally suggests common spatial structure organization and mechanism of action. We may therefore speculate that, albeit producing diverse effects on their target, β/δ- and μ-agatoxins, curtatoxins, and δ-Palu-IT all share a common fold and bind to the same receptor site 4 of NaV channels, as shown for δ-Palu-IT2 (23). In Fig. 8, a spatial structure model for β/δ-Aga-1 is shown, based on sequence similarity to δ-Palu-IT2 (62.2% identical residues). Residues critical for channel binding have been determined in δ-Palu-IT2 by alanine scanning mutagenesis: Arg-8, Trp-12, Tyr-22, Ser-24, Arg-26, Met-28, Tyr-30, Arg-32, and Arg-34 (colored red in Fig. 8 at the proper positions) (23). Of these, Trp-12 and Tyr-22 are almost 100% conserved across all family members (see Fig. 7) with the only W12A mutation in curtatoxin-1 (a neighboring Tyr-13 in this toxin may take over the functions) and Y22W substitution in δ-Palu-IT3; a hydroxy amino acid residue (Ser or Thr) is found at position equivalent to Ser-24 with Asn only in curtatoxin-1; Arg-26 is substituted for Met in β/δ-Aga-6, Ser in curtatoxin-1, and Pro in δ-Palu-IT3; a hydrophobic residue is present at position corresponding to Met-28; either positively charged or bulky hydrophobic residues are found at the positions of Arg-32 and Arg-34; and only Arg-8 and Tyr-30 are quite variable, still with preference for Arg/Lys, His, or Gln at position 8 and Lys or Tyr at position 30 (see Fig. 7). Therefore, the general features of the δ-Palu-IT2 pharmacophore are well conserved among all members of the family.
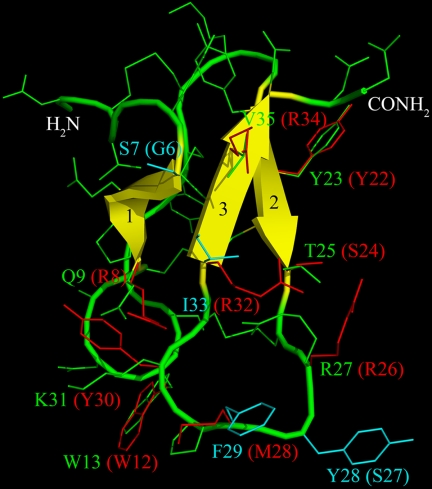
FIGURE 8.
Spatial structure model of β/δ-Aga-1. Model was built on the basis of the structure of the homologous δ-palutoxin-IT2 (PDB accession code: 1V91). Main-chain atoms are hidden, side chains are shown as lines, N and C termini are labeled, and β-strands are colored yellow and numbered. Presumed functionally important residues are indicated, the corresponding residues of δ-Palu-IT2 are given in parenthesis; the putative pharmacophore of δ-Palu-IT2 (23) is colored red; and residues believed to conform the scorpion β-toxin-like activity to β/δ-Aga-1 are marked cyan.
The decreased potency of β/δ-Aga-4 and -7 compared with β/δ-Aga-1, -2, and -3 (Fig. 6) may be attributed to P16L and K31G mutations, respectively. In fact, the least active homologue β/δ-Aga-4 differs from the most active homologue β/δ-Aga-3 by one P16L mutation solely. Proline might be important structurally, and this residue was shown to form an unusual cis-peptide bond in δ-Palu-IT1 and -2 (53), whereas the K31G substitution hits the δ-Palu-IT2 pharmacophore. The relative “uncoupling” of the two effects seen in β/δ-Aga-5 and -6 could be explained by the structure difference of the Cys-2 to Cys-3 loop, which is shorter in these two peptides.
In fact, β/δ- and μ-agatoxins (the best studied μ-Aga-1 and -4) may be considered as having an “additional” scorpion β-toxin-like activity as compared with the α-toxin-like δ-palutoxins-IT (δ-Palu-IT1 and -2). Following the same reasoning as above, residues that account for this additional feature may be located. In β/δ-Aga-1, these are Ser-7, Tyr-28, Phe-29, and Ile-33 (cyan in Fig. 8), the corresponding residues in δ-Palu-IT2 being Gly-6, Ser-27, Met-28, and Arg-32. All other substitutions in δ-Palu-IT2 as compared with β/δ-Aga-1 can also be found in other toxins exhibiting the β-toxin-like activity and are therefore considered neutral. Interestingly, all of the presumed residues assuring the β-toxin-like activity are located close in space (see Fig. 8) with two of the four coinciding with the δ-Palu-IT2 pharmacophore. The envisaged future mutagenesis studies will shed more light onto these issues.
In conclusion, we argue that the “classic” concept of receptor sites 3 and 4 of NaV channels binding toxins that exhibit the α- and β-effects (site 3- and site 4-like activities), respectively, becomes rather “blurred” as new data are being accumulated. Indeed, it seems that no strict correspondence exists between the toxin binding site and its effect on the channel function.
Acknowledgments
We thank A. L. Goldin (University of California, Irvine) for sharing rNaV1.2, S. H. Heinemann (Friedrich-Schiller-Universität, Jena, Germany) for sharing rβ1, and M. S. Williamson (Institute of Arable Crops Research Rothamsted, Harpenden, UK) for sharing DmNaV1 and tipE. We are also very grateful to Sergey Kozlov, Steve Peigneur, Marijke Stevens, and Frank Bosmans for their helpful discussions and support in molecular biology.
This work was supported by the Federal Agency for Education of the Russian Federation (State Contract P1388), the Program of Cell and Molecular Biology of the Russian Academy of Sciences, the Russian Foundation for Basic Research (Grant 05-04-50828), Grant BIL05/50 (bilateral collaboration between Flanders and the Russian Federation), Grants G.0257.08 and G.0330.06 (F.W.O. Vlaanderen), Grant OT-05–64 (K.U. Leuven), and Grant UA P6/31 (Interuniversity Attraction Poles Program, Belgian State, Belgian Science Policy).
- NaV channel
- voltage-gated sodium channel
- gNa
- sodium conductance
- INaP
- persistent sodium current
- INa
- transient sodium current
- LD50
- median lethal dose
- RP-HPLC
- reversed-phase high-performance liquid chromatography
- δ-Palu-IT
- δ-palutoxin-IT
- β/δ-Aga
- β/δ-agatoxin
- μ-Aga
- μ-agatoxin
- EC50
- toxin concentration causing a half-maximal effect
- ICK
- inhibitor cystine knot.
REFERENCES
- 1.Estrada G., Villegas E., Corzo G. (2007) Nat. Prod. Rep. 24, 145–161 [DOI] [PubMed] [Google Scholar]
- 2.Grishin E. (1999) Eur. J. Biochem. 264, 276–280 [DOI] [PubMed] [Google Scholar]
- 3.Rash L. D., Hodgson W. C. (2002) Toxicon 40, 225–254 [DOI] [PubMed] [Google Scholar]
- 4.Vassilevski A. A., Kozlov S. A., Grishin E. V. (2009) Biochemistry 74, 1505–1534 [DOI] [PubMed] [Google Scholar]
- 5.Catterall W. A. (2000) Neuron 26, 13–25 [DOI] [PubMed] [Google Scholar]
- 6.Feng G., Deák P., Chopra M., Hall L. M. (1995) Cell 82, 1001–1011 [DOI] [PubMed] [Google Scholar]
- 7.Warmke J. W., Reenan R. A., Wang P., Qian S., Arena J. P., Wang J., Wunderler D., Liu K., Kaczorowski G. J., Van der Ploeg L. H., Ganetzky B., Cohen C. J. (1997) J. Gen. Physiol. 110, 119–133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Armstrong C. M. (1981) Physiol. Rev. 61, 644–683 [DOI] [PubMed] [Google Scholar]
- 9.Elinder F., Nilsson J., Arhem P. (2007) Physiol. Behav. 92, 1–7 [DOI] [PubMed] [Google Scholar]
- 10.Billen B., Bosmans F., Tytgat J. (2008) Curr. Pharm. Des. 14, 2492–2502 [DOI] [PubMed] [Google Scholar]
- 11.Wang S. Y., Wang G. K. (2003) Cell Signal 15, 151–159 [DOI] [PubMed] [Google Scholar]
- 12.Cestèle S., Catterall W. A. (2000) Biochimie 82, 883–892 [DOI] [PubMed] [Google Scholar]
- 13.Nicholson G. M. (2007) Toxicon. 49, 490–512 [DOI] [PubMed] [Google Scholar]
- 14.Li D., Xiao Y., Hu W., Xie J., Bosmans F., Tytgat J., Liang S. (2003) FEBS Lett. 555, 616–622 [DOI] [PubMed] [Google Scholar]
- 15.Xiao Y., Liang S. (2003) Eur. J. Pharmacol. 477, 1–7 [DOI] [PubMed] [Google Scholar]
- 16.Xiao Y. C., Liang S. P. (2003) Toxicon 41, 643–650 [DOI] [PubMed] [Google Scholar]
- 17.Liang S. (2004) Toxicon 43, 575–585 [DOI] [PubMed] [Google Scholar]
- 18.Rogers J. C., Qu Y., Tanada T. N., Scheuer T., Catterall W. A. (1996) J. Biol. Chem. 271, 15950–15962 [DOI] [PubMed] [Google Scholar]
- 19.Bosmans F., Tytgat J. (2007) Toxicon 49, 550–560 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yamaji N., Little M. J., Nishio H., Billen B., Villegas E., Nishiuchi Y., Tytgat J., Nicholson G. M., Corzo G. (2009) J. Biol. Chem. 284, 24568–24582 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Corzo G., Gilles N., Satake H., Villegas E., Dai L., Nakajima T., Haupt J. (2003) FEBS Lett. 547, 43–50 [DOI] [PubMed] [Google Scholar]
- 22.Nicholson G. M., Little M. J., Birinyi-Strachan L. C. (2004) Toxicon 43, 587–599 [DOI] [PubMed] [Google Scholar]
- 23.Corzo G., Escoubas P., Villegas E., Karbat I., Gordon D., Gurevitz M., Nakajima T., Gilles N. (2005) Biochemistry 44, 1542–1549 [DOI] [PubMed] [Google Scholar]
- 24.Adams M. E. (2004) Toxicon 43, 509–525 [DOI] [PubMed] [Google Scholar]
- 25.Stapleton A., Blankenship D. T., Ackermann B. L., Chen T. M., Gorder G. W., Manley G. D., Palfreyman M. G., Coutant J. E., Cardin A. D. (1990) J. Biol. Chem. 265, 2054–2059 [PubMed] [Google Scholar]
- 26.Fainzilber M., Kofman O., Zlotkin E., Gordon D. (1994) J. Biol. Chem. 269, 2574–2580 [PubMed] [Google Scholar]
- 27.Leipold E., Hansel A., Olivera B. M., Terlau H., Heinemann S. H. (2005) FEBS Lett. 579, 3881–3884 [DOI] [PubMed] [Google Scholar]
- 28.Ekberg J., Craik D. J., Adams D. J. (2008) Int J Biochem. Cell Biol. 40, 2363–2368 [DOI] [PubMed] [Google Scholar]
- 29.De Lima M. E., Figueiredo S. G., Pimenta A. M., Santos D. M., Borges M. H., Cordeiro M. N., Richardson M., Oliveira L. C., Stankiewicz M., Pelhate M. (2007) Comp Biochem. Physiol C. Toxicol. Pharmacol. 146, 264–279 [DOI] [PubMed] [Google Scholar]
- 30.King G. F., Escoubas P., Nicholson G. M. (2008) Channels (Austin) 2, 100–116 [DOI] [PubMed] [Google Scholar]
- 31.Zlotkin E. (1999) Annu. Rev. Entomol. 44, 429–455 [DOI] [PubMed] [Google Scholar]
- 32.Catterall W. A., Goldin A. L., Waxman S. G. (2005) Pharmacol. Rev. 57, 397–409 [DOI] [PubMed] [Google Scholar]
- 33.Dong K. (2007) Invert. Neurosci. 7, 17–30 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Tan J., Liu Z., Nomura Y., Goldin A. L., Dong K. (2002) J. Neurosci. 22, 5300–5309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yamaji N., Sugase K., Nakajima T., Miki T., Wakamori M., Mori Y., Iwashita T. (2007) FEBS Lett. 581, 3789–3794 [DOI] [PubMed] [Google Scholar]
- 36.Quistad G. B., Reuter C. C., Skinner W. S., Dennis P. A., Suwanrumpha S., Fu E. W. (1991) Toxicon 29, 329–336 [DOI] [PubMed] [Google Scholar]
- 37.Adams M. E., Herold E. E., Venema V. J. (1989) J. Comp. Physiol. A 164, 333–342 [DOI] [PubMed] [Google Scholar]
- 38.Cohen C. J., Bale T. A., Ertel E. A., Warren V. A., Smith M. M. (1993) Biophys. J. 64, A4 [Google Scholar]
- 39.Norris T. M., Lee D., Adams M. E. (1995) Soc. Neurosci. Abstr. 21, 1820 [Google Scholar]
- 40.Skinner W. S., Adams M. E., Quistad G. B., Kataoka H., Cesarin B. J., Enderlin F. E., Schooley D. A. (1989) J. Biol. Chem. 264, 2150–2155 [PubMed] [Google Scholar]
- 41.Vassilevski A. A., Kozlov S. A., Egorov T. A., Grishin E. V. (2010) Methods Mol. Biol. 615, 87–100 [DOI] [PubMed] [Google Scholar]
- 42.Noda M., Ikeda T., Suzuki H., Takeshima H., Takahashi H., Kuno M., Numa S. (1986) Nature 322, 826–828 [DOI] [PubMed] [Google Scholar]
- 43.Gellens M. E., George A. L., Jr., Chen L. Q., Chahine M., Horn R., Barchi R. L., Kallen R. G. (1992) Proc. Natl. Acad. Sci. U.S.A. 89, 554–558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Billen B., Vassilevski A., Nikolsky A., Tytgat J., Grishin E. (2008) Toxicon 52, 309–317 [DOI] [PubMed] [Google Scholar]
- 45.Nikolsky A. S., Billen B., Vassilevski A. A., Filkin S. Y., Tytgat J., Grishin E. V. (2009) Biochemistry (Moscow) Supplement Series A: Membrane and Cell Biology 3, 245–253 [Google Scholar]
- 46.Cestèle S., Qu Y., Rogers J. C., Rochat H., Scheuer T., Catterall W. A. (1998) Neuron 21, 919–931 [DOI] [PubMed] [Google Scholar]
- 47.Cohen L., Karbat I., Gilles N., Ilan N., Benveniste M., Gordon D., Gurevitz M. (2005) J. Biol. Chem. 280, 5045–5053 [DOI] [PubMed] [Google Scholar]
- 48.Bosmans F., Martin-Eauclaire M. F., Tytgat J. (2007) Toxicol. Appl. Pharmacol. 218, 45–51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Corzo G., Sabo J. K., Bosmans F., Billen B., Villegas E., Tytgat J., Norton R. S. (2007) J. Biol. Chem. 282, 4643–4652 [DOI] [PubMed] [Google Scholar]
- 50.Nicholson G. M., Walsh R., Little M. J., Tyler M. I. (1998) Pflugers Arch. 436, 117–126 [DOI] [PubMed] [Google Scholar]
- 51.Nicholson G. M., Willow M., Howden M. E., Narahashi T. (1994) Pflugers Arch. 428, 400–409 [DOI] [PubMed] [Google Scholar]
- 52.Hondeghem L. M., Katzung B. G. (1984) Annu. Rev. Pharmacol. Toxicol. 24, 387–423 [DOI] [PubMed] [Google Scholar]
- 53.Ferrat G., Bosmans F., Tytgat J., Pimentel C., Chagot B., Gilles N., Nakajima T., Darbon H., Corzo G. (2005) Proteins 59, 368–379 [DOI] [PubMed] [Google Scholar]
- 54.Omecinsky D. O., Holub K. E., Adams M. E., Reily M. D. (1996) Biochemistry 35, 2836–2844 [DOI] [PubMed] [Google Scholar]
- 55.Mouhat S., Jouirou B., Mosbah A., De Waard M., Sabatier J. M. (2004) Biochem. J. 378, 717–726 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kozlov S., Malyavka A., McCutchen B., Lu A., Schepers E., Herrmann R., Grishin E. (2005) Proteins 59, 131–140 [DOI] [PubMed] [Google Scholar]
- 57.Kozlov S. A., Grishin E. V. (2007) Toxicon 49, 721–726 [DOI] [PubMed] [Google Scholar]
- 58.Kozlov S. A., Vassilevski A. A., Grishin E. V. (2009) in Protein Biosynthesis, (Esterhouse T. E., Petrinos L. B. eds) pp. 225–248, Nova Biomedical Books, New York [Google Scholar]
- 59.Corzo G., Escoubas P., Stankiewicz M., Pelhate M., Kristensen C. P., Nakajima T. (2000) Eur. J. Biochem. 267, 5783–5795 [DOI] [PubMed] [Google Scholar]
- 60.Shlyapnikov Y. M., Kozlov S. A., Feodorov A. A., Grishin E. V. (2010) Bioorg. Khim. 36, 81–88 [PubMed] [Google Scholar]