Abstract
A purification protocol that comprised ion exchange chromatography on DEAE-cellulose, affinity chromatography on Affi-gel blue gel, ion exchange chromatography on SP-Sepharose, and gel filtration by FPLC on Superdex 75 was complied to isolate two trypsin inhibitors from Phaseolus vulgaris cv “White Cloud Bean”. Both trypsin inhibitors exhibited a molecular mass of 16 kDa and reduced the activity of trypsin with an IC50 value of about 0.6 μM. Dithiothreitol attenuated the trypsin inhibitory activity, signifying that an intact disulfide bond is indispensable to the activity. [Methyl-3H] thymidine incorporation by leukemia L1210 cells was inhibited with an IC50 value of 28.8 μM and 21.5 μM, respectively. They were lacking in activity toward lymphoma MBL2 cells and inhibitory effect on HIV-1 reverse transcriptase and fungal growth when tested up to 100 μM.
1. Introduction
Protease inhibitors have been purified from an array of leguminous and nonleguminous species encompassing Torresea cearensis [1], Erythrina caffra [2], Dolichos lablab [3], Crotalaria paulina [4], Medicago scutellata [5, 6], Canavalia gladiata [7], Pisum sativum [8], Dimorphandra mollis [9], Swartzia pickellii [10], Psophocarpus tetragonolobus [11], Delonix regina [12], Poecilanthe parviflora [13], Adenanthera pavonina [14], Cajanus cajan [15], Dolichos biflorus [16], Phaseolus acutifolius [17], Arachis hypogaea [18], Leucaena leucocephala [19], Bauhinia bauhinioides [20], Bauhinia variegata [21], Bauhinia ungulata [22], Vigna unguiculata [23], Lens culinaris [24], Glycine max [25], Peltophorum dubium [26], Pithecellobium dulce [27], Glycine soja [28], and barley [29].
Protease inhibitors impair the activity of insect midgut proteases and thus adversely affect protein digestion and health in insects. They represent one of the multitude of entomotoxic proteins [30, 31]. In addition to insecticidal activity [32–35], protease inhibitors demonstrate antiproliferative and antitumor activities [36–45]. The objective of the present study was to isolate and characterize proteins with protease inhibitory activity from white cloud beans.
2. Material and Methods
2.1. Isolation of Trypsin Inhibitor
An aqueous extract of the beans (250 g) was produced by blending in distilled water (3 ml/g) followed by centrifugation (14000 g for 25 minutes at 4°C). The resulting supernatant was applied to a 5 × 20 cm column of DEAE-cellulose (Sigma) in 10 mM Tris-HCl buffer (pH 7.4). After elution of unadsorbed proteins (fraction D1), the column was eluted successively with 0.2 M NaCl and 1 M NaCl in the Tris-HCl buffer. Fraction D2 eluted with 0.2 M NaCl was dialyzed to remove NaCl and then subjected to affinity chromatography on a 5 × 15 cm of Affi-gel blue gel (Bio-Rad) in 10 mM Tris HCl buffer (pH 7.4). The unadsorbed fraction (B1) was dialyzed against 10 mM NH4OAc buffer (pH 5) and then applied to a 2.5 × 20 cm column of SP-Sepharose (GE Healthcare). After elution of unadsorbed proteins (fraction S1), the column was eluted with a 0-1 M NaCl concentration gradient in the NH4OAc buffer. The first and second adsorbed fractions (SP2 and SP3) were then further purified by gel filtration on a Superdex 75 HR 10/30 column (GE Healthcare) in 0.2 M NH4HCO3 buffer (pH 8.5) using an AKTA Purifier (GE Healthcare). The second absorbance peak represented purified trypsin inhibitor.
2.2. Electrophoresis, Molecular Mass Determination, and N-Terminal Sequence Analysis
The molecular mass of the isolated proteins was determined by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) following the procedure of Laemmli and Favre [46]. Gel filtration on an FPLC-Superdex 75 column, previously calibrated with molecular mass marker proteins (GE Healthcare), was employed to determine the molecular mass of the protein. The N-terminal sequence of the protein was analyzed by using a Hewlett-Packard HP G1000A Edman degradation unit and an HP 1000 HPLC System [47].
2.3. Assay for Trypsin Inhibitory Activity
The test sample (20 μl) was added to 160 μl of a 1% casein solution in 0.1 M Tris-HCl buffer (pH 7.4). Trypsin (20 μl of a 0.5 mg/ml solution) was then added and the mixture was incubated at 37°C for 15 minutes followed by addition of Trichloroacetic acid (0.4 ml, 5%) that was added to bring the reaction to an end. After centrifugation the absorbance of the resulting supernatant, which indicates the amount of casein fragments produced by trypsin, was read at 280 nm. The % inhibition of trypsin activity is equal to the % decrease in absorbance of the supernatant [48].
2.4. Effect of Dithiothreitol (DTT) on Trypsin Inhibitory Activity
The isolated trypsin inhibitor (2.5 μM) was treated with dithiothreitol (DTT) at the final concentration 2.5, 10 and 40 mM for 5, 20, and 80 minutes at 37°C. Soybean trypsin inhibitor from Sigma (2.5 μM) was similarly treated and used as a positive control. The reaction was terminated by adding iodoacetamide at twice the amount of thiol functions at each DTT concentration. Residual trypsin inhibitor activity was measured at pH 7.4 as described above in assay for trypsin inhibitory activity. The highest iodoacetamide concentration used in the test was devoid of any effect on trypsin activity and the trypsin inhibitory activity of isolated trypsin inhibitor and soybean trypsin inhibitor [47].
2.5. Assay for Antiproliferative Activity Toward Tumor Cells
The antiproliferative activity of the purified trypsin inhibitor was assayed as described below. The cell lines L1210 (human leukemia) and MBL2 (murine lymphoma) were obtained from American Type Culture Collection. The cell line was maintained in Dulbecco Modified Eagles' Medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 100 mg/l streptomycin, and 100 IU/ml penicillin, at 37°C in a humidified atmosphere of 5% CO2. Cells (1 × 104) in their exponential growth phase were seeded into each well of a 96-well culture plate (Nunc, Denmark) and incubated for 3 hours prior to addition of the trypsin inhibitor. Incubation was performed for an additional 48 hours. Radioactive precursor, 1 μCi ([3H-methyl]-thymidine, from GE Healthcare), was then introduced to each well and the incubation continued for 6 hours. The cultures were then harvested using a cell harvester. The radioactivity incorporated was measured in a liquid scintillation counter [49].
2.6. Assay for HIV-1 Reverse Transcriptase Inhibitory Activity
The assay for HIV reverse transcriptase inhibitory activity was carried out in view of the report that trypsin inhibitors manifest this activity [50, 51]. It was conducted according to instructions supplied with the assay kit from Boehringer Mannheim (Germany). The assay makes following use of the ability of reverse transcriptase to synthesize DNA, commencing from the template/primer hybrid poly (A) oligo (dT) 15. The digoxigenin- and biotin-labeled nucleotides in an optimized ratio are incorporated into one of the same DNA molecule, which is freshly synthesized by the reverse transcriptase (RT). The detection and quantification of synthesized DNA as a parameter for RT activity are based on a sandwich ELISA protocol. Biotin-labeled DNA binds to the surface of microtiter plate modules that have been precoated with strepatavidin. In the following step, an antibody to digoxigenin, conjugated to peroxidase, binds to the digoxigenin-labeled DNA. In the last step, the peroxidase substrate is added. The peroxidase enzyme affects the cleavage of the substrate, yielding a colored reaction product. The absorbance of the sample at 405 nm which is directly correlated to the level of RT activity can be measured using a microtiter plate (ELISA) reader. A fixed amount (4–6 ng) of recombinant HIV-1 reverse transcriptase was used. The inhibitory activity of the trypsin inhibitor was calculated as percent inhibition compared to a control without the trypsin inhibitor [49].
2.7. Assay of Ability to Inhibit HIV-1 Integrase
2.7.1. Expression and Purification of Recombinant HiV-1 Integrase
The plasmid that expressed His-tagged wild-type HIV-1 integrase, pT7-7-His (Y | TX)-HIV-1-IN, was a generous gift from Professor S.A. Chow (School of Medicine, UCLA). To express the enzyme, a 1-liter culture of E. coli BL21 (DE3) cells containing the expressing plasmid was grown at 37°C until it reached OD600 0.7-0.8. Cells were induced by addition of 0.8 mM IPTG (isopropyl-β-d-thiogalactopyranoside) and harvested, after 4 hours of incubation, by centrifugation at 6000 g for 10 minutes at 4°C. Cells were suspended at a concentration of 10 ml/g wet cell paste in 20 mM Tris-HCl buffer (pH 8.0) containing 0.1 mM EDTA, 2 mM β-mercaptoethanol, 0.5 M NaCl, and 5 mM imidazole. Lysozyme was added to a concentration of 0.2 mg/ml. After incubation at 4°C for 1 hour, the lysate was sonicated and centrifuged at 40 000 g at 4°C for 20 minutes. The pellet was homogenized in 50 ml buffer A (20 mM Tris-HCl, pH 8.0, 2 M NaCl, 2 mM β-mercaptoethanol) which contained 5 mM imidazole. The suspension was rotated at 4°C for 1 hour and cleared by centrifugation at 40 000 g at 4°C for 20 minutes. The supernatant was applied to a 1 ml chelating Sepharose (GE Healthcare) column charged with 50 mM imidazole. The column was eluted with five column volumes of buffer A containing 5 mM imidazole, and the protein was eluted with three column volumes of buffer A containing 200 and 400 mM imidazole, respectively. Protein containing fractions were pooled, and EDTA was added to a final concentration of 5 mM, followed by dialysis against buffer B (20 mM HEPES, pH 7.5, 1 mM EDTA, 1 M NaCl, 20% glycerol) containing 2 mM β-mercaptoethanol and then against buffer B containing 1 mM dithiothreitol. Aliquots of the protein were stored at −70°C [49].
2.7.2. HIV-1 Integrase Assay
A nonradioactive ELISA-based HIV-1 integrase assay was carried out according to the DNA-coated plate method. In this study, 1 μg of Smal-linearized pBluescript SK was coated onto each well in the presence of 2 M NaCl as target DNA. The donor DNA was prepared by annealing VU5BR (5′-biotin-GTGTGGAAAATCTCTA- GCAGT-3′) and VU5 (5′-ACTGCTAGAGATTTTCCACAC-3′) in 10 mM Tris-HC1, pH 8.0, 1 mM EDTA, and 0.1 M NaCl at 80°C, followed by 30 minutes at room temperature. Integrase reaction was conducted in 20 mM HEPES (pH 7.5) containing 10 mM MnCl2, 30 mM NaCl, 10 mM dithiothreitol, and 0.05% Nonidet-P40 (Sigma). After the reaction, biotinylated DNA immobilized on the wells was incubated with streptavidin-conjugated alkaline phosphatase (Boehringer-Mannheim, Mannheim, Germany), followed by colorimetric detection with 1 mg/ml p-nitrophenyl phosphate in 10% diethanolamine buffer (pH 9.8) containing 0.5 mM MgCl2. The absorbance was read at 415 nm. The ribosome inactivating protein trichosanthin was employed as a positive control [49].
2.8. Screening for Inhibitory Effect on SARS Coronavirus Protease
The activity of SARS coronavirus (CoV) protease was reflected by a cleavage of designed substrate which is composed of two proteins linked by a cleavage site for SARS CoV protease. The reaction was carried out in a mixture containing 5 μM SARS CoV protease, 5 μM sample, and 20 μM substrate and buffer [20 mM Tris-HCl (pH 7.5), 20 mM NaCl and 10 mM beta-mercaptoethanol] for 40 minutes at 37°C. The reaction was then terminated by heating at 100°C for 2 minutes. Then the reaction mixture was analysed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). If SARS CoV protease is inhibited by the test sample, there is only one band, which is the intact substrate, shown in SDS-PAGE [52].
2.9. Assay for Antifungal Activity
This assay was performed since some trypsin inhibitors demonstrate antifungal activity [8]. The assay for antifungal activity toward Botrytis cinerea and Fuserium oxysporum was executed in 100 mM × 15 mM petri plates containing 10 ml of potato dextrose agar. After the mycelial colony had grown to a sufficiently large size, sterile blank paper disks (0.625 cm in diameter) were laid at a distance of 0.5 cm away from the edge of the mycelial colony. An aliquot (15 μl) of the trypsin inhibitor was added to a disk. The plates were exposed at 23°C for 72 hours until mycelial growth had enveloped the disks containing the control and had produced crescents of inhibition around disks containing samples with antifungal activity [49].
3. Results
3.1. Isolation of Trypsin Inhibitor
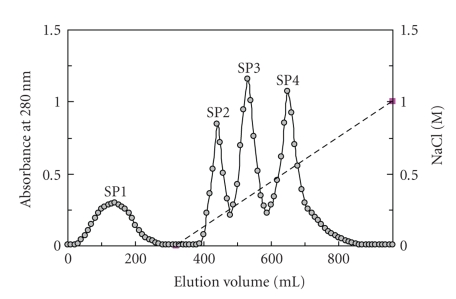
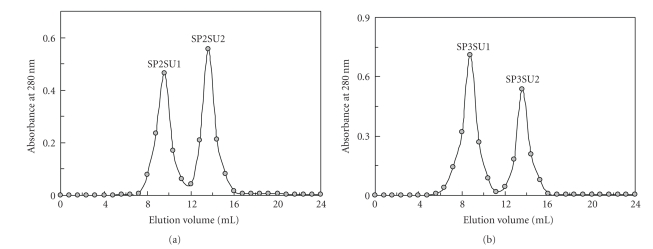
Anion exchange chromatography of the bean extract on DEAE-cellulose resolved it into three fractions, an unadsorbed fraction D1 together with two adsorbed fractions D2 and D3. Trypsin inhibitory activity was confined to fraction D2. After affinity chromatography of fraction D2 on Affi-gel blue gel, the activity appeared in the unadsorbed fraction B1 (data not shown). Ion exchange chromatography of fraction B1 on SP-Sepharose resolved it into a small unadsorbed fraction SP1 and three large adsorbed fractions (SP2, SP3, and SP4) of about the same size (Figure 1). Trypsin inhibitory activity was detected in fractions SP2 and SP3. Final purification of SP2 on Superdex 75 produced two fractions, SP2SU1 and SP2SU2 (Figure 2(a)). Gel filtration of SP3 on Superdex 75 yielded two fractions SP3SU1 and SP3SU2 (Figure 2(b)). Fractions SP2SU2 and SP3SU2, both with a molecular mass of 16 kDa as determined by gel filtration on Superdex 75, were the only fractions with trypsin inhibitory activity. The remaining fractions SP2SU1 and SP3SU1 were inactive. The yields of the various chromatographic fractions are presented in Table 1. Both SP2SU2 and SP3SU2 displayed a molecular mass of 16 kDa in SDS-PAGE (Figure 3) and gel filtration (Figure 2). Their N-terminal sequences are shown in Table 2. These two white cloud bean trypsin inhibitors inhibited trypsin with an IC50 of about 0.6 μM (Figure 4).
Figure 1.
Results of ion exchange chromatography on SP-Sepharose column (2.5 × 20 cm). The sample is fraction of white cloud bean extract previously adsorbed on DEAE-cellulose and eluted with 0.2 M NaCl added to the buffer, and subsequently unadsorbed on Affi-gel blue gel in the Tris-HCl buffer. Starting buffer for SP-Sepharose chromatography is 10 mM NH4OAc buffer (pH 5). Broken line across the right half of the chromatography represents the linear 0-1 M NaCl gradient in 10 mM NH4OAc buffer (pH 5) employed to elute adsorbed proteins. Trypsin inhibitory activity was found in both fractions SP2 and SP3.
Figure 2.
(a) Fast protein liquid chromatography gel filtration of fraction SP2 from SP-Sepharose column on a Superdex 75 HR10/30 column in 0.2 M NH4HCO3 buffer (pH 8.5) at a flow rate of 0.4 ml/min. Fraction size is 0.8 ml. Trypsin inhibitory activity was restricted to the second fraction (SP2SU2). (b) Fast protein liquid chromatography gel filtration of fraction SP3 from SP-Sepharose column on a Superdex 75 HR10/30 column in 0.2 M NH4HCO3 buffer (pH 8.5) at a flow rate of 0.4 ml/min. Fraction size is 0.8 ml. Trypsin inhibitory activity was restricted to the second fraction (SP3SU2).
Table 1.
Yields and trypsin inhibitory activities of various chromatographic fractions (from 250 g dry white cloud beans).
| Fraction | Yield (mg) | IC50 (μg/ml) | Fraction | Yield (mg) | IC50 (μg/ml) |
|---|---|---|---|---|---|
| Extract | 4150 | 236.8 | SP2 | 82.1 | 17.3 |
| D1 | 1079 | — | SP3 | 102.6 | 23.5 |
| D2 | 904 | 72.4 | SP4 | 109.4 | — |
| D3 | 958 | — | SP2SU1 | 33.2 | — |
| B1 | 469.8 | 45.1 | SP2SU2 | 39.9 | 9.6 |
| B2 | 179.1 | — | SP3SU1 | 41.8 | — |
| SP1 | 47.9 | — | SP3SU2 | 31.7 | 8.8 |
Trypsin inhibitor-enriched fractions are highlighted in boldface.
Figure 3.
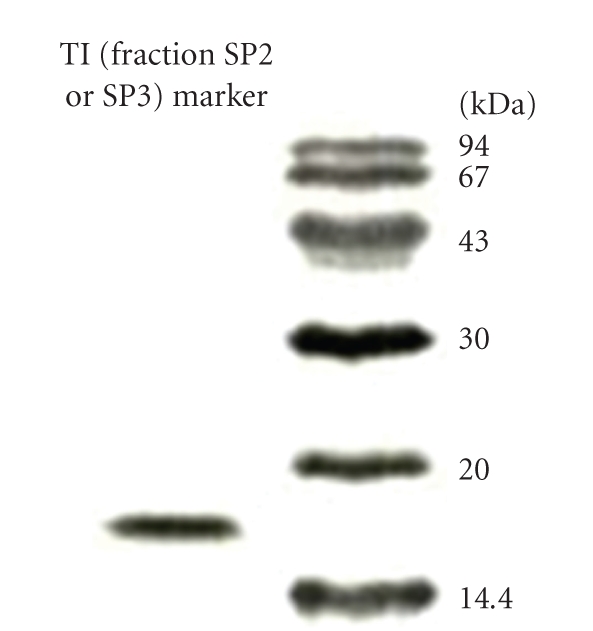
Results of SDS-PAGE of the two white cloud bean trypsin inhibitors (TI, fractions SU2 and SU3) and molecular mass marker. Molecular mass of both trypsin inhibitors was calculated to be 16 kDa.
Table 2.
N-terminal sequence comparison of white cloud bean trypsin inhibitors SP3SU2 and SP2SU2 with other leguminous trypsin inhibitors (Results of BLAST search).
| Trypsin inhibitor | Amino acid sequences |
|---|---|
| SP3SU2 | GSGHRHESTDEPSSSKAACCD |
| SP2SU2 | GH -HRHESTDEPSESKKACCDHCACTKKIPPQCRRDLLLL |
| PVTI (34–62) | –HRHESTDEPSESSKACCDHCACTKSIPPQ |
| VUTI (80–118) | –NHHDSDSSDEPSESSEPCCDSCICSKSIPPQCHHTDIRL |
| VMTI (32–51) | –SGRHHESTDEPSESSKPCCD |
| PLTI (32–51) | –SGHHHESTDEPSESSKPCCD |
Identical corresponding residues are underlined.
VUTI: Vigna unguiculata trypsin inhibitor.
PVTI: double-headed trypsin inhibitor from Phaseolus vulgaris.
PLTI: double-headed trypsin inhibitor from Phaseolus lunatus.
VMTI: double-headed trypsin inhibitor from Vugna mungo.
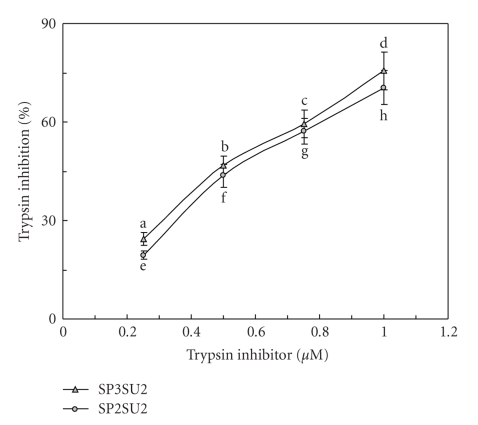
Figure 4.
Inhibitory effects by the two white cloud bean trypsin inhibitors on trypsin. IC50 = circa 0.6 μM. IC50 for soybean trypsin inhibitor = 1.5 μM. Results are presented as mean ± SD (n = 3). Results are presented as mean ± SD (n = 3). IC50 = 0.25 μM. Different letters (a, b, c, d) next to the data points indicate statistically significant difference (P < .05) when the data are analyzed by analysis of variance followed by Duncan's multiple range test.
3.2. Characterization of Isolated Trypsin Inhibitor
DTT treatment curtailed the trypsin inhibiting activity in a dose- and time-dependent manner (Table 3). The two trypsin inhibitors did not inhibit HIV-1 reverse transcriptase when tested at various concentrations up to 100 μM (Table 4). They lacked antifungal activity when tested up to 24 μg/disk (100 μM, 15 μl) (data not shown). The IC50 values of the inhibitory effects of the trypsin inhibitors on L1210 cells were, respectively, 21.5 μM and 28.8 μM (Table 4). However, there was no activity toward MBL2 cells when tested up to 100 μM. They showed marked homology to partial sequences of other leguminous trypsin inhibitors.
Table 3.
Inhibition rate (%) of dithiothreitol (DTT) on the trypsin inhibitory activity of white cloud bean trypsin inhibitor and soybean trypsin inhibitor after incubation at 37°C for different durations.
| Incubation time (min) | Isolated trypsin inhibitor (SP3SU2) | Isolated trypsin inhibitor (SP2SU2) | Soybean trypsin inhibitor | ||||
|---|---|---|---|---|---|---|---|
| 2.5 mM DTT | 10 mM DTT | 40 mM DTT | 2.5 mM DTT | 10 mM DTT | 40 mM DTT | 2.5 mM DTT | |
| 5 | 9.7 ± 1.1a | 24.7 ± 2.5b | 28.8 ± 2.4c | 9.0 ± 1.0a | 22.8 ± 1.9b | 27.0 ± 2.2c | 78.2 ± 5.7a |
| 20 | 30.6 ± 2.9d | 55.0 ± 4.1e | 67.4 ± 4.1f | 28.3 ± 3.0d | 54.1 ± 3.6e | 65.5 ± 4.3f | 93.7 ± 5.0b |
| 80 | 61.7 ± 6.5g | 83.2 ± 5.9h | 94.5 ± 5.6i | 59.6 ± 4.7g | 80.6 ± 6.5h | 91.9 ± 6.6i | 96.4 ± 4.3c |
Results are presented as mean ± SD (n = 3). Different alphabets (e.g., a, b, and c) indicate statistically significant difference (P < .05) when (I) data at same time point and different DTT concentrations or (II) data at same DTT concentration but different time point were analyzed by analysis of variance followed by Duncan's multiple range test.
Table 4.
Inhibition rate (%) of white cloud bean trypsin inhibitors on L1210 cells, MBL2 cells, and HIV-1 reverse transcriptase (RT). Results are presented as mean ± SD (n = 3).
| Dose (μM) | Trypsin inhibitor (SP3SU2) | trypsin inhibitor (SP2SU2) | ||||
|---|---|---|---|---|---|---|
| L1210 cells | MBL2 cells | HIV-1 RT | L1210 cells | MBL2 cells | HIV-1 RT | |
| 100 | 93.2 ± 5.7a | 4.8 ± 1.5 | 3.8 ± 0.6 | 88.1 ± 7.2a | 5.0 ± 1.4 | 5.0 ± 0.8 |
| 50 | 79.3 ± 5.0b | 2.7 ± 0.7 | 5.3 ± 1.1 | 68.6 ± 5.3b | 4.1 ± 0.8 | 4.7 ± 1.0 |
| 25 | 56.0 ± 3.8c | 3.4 ± 1.2 | 3.4 ± 1.3 | 44.9 ± 3.4c | 3.5 ± 1.5 | 4.5 ± 1.3 |
| 12.5 | 24.7 ± 3.1d | 4.2 ± 0.9 | 4.0 ± 1.8 | 21.3 ± 2.2d | 4.3 ± 1.0 | 3.1 ± 0.6 |
Different alphabets (e.g., a, b, c, and d) indicate statistically significant difference (P < .05) when data were analyzed by analysis of variance followed by Duncan's multiple range test.
4. Discussion and Conclusions
The present study disclosed the production of two trypsin inhibitors with closely related N-terminal sequences chromatographic behavior and bioactivities by the white cloud bean variety of Phaseolus vulgaris. The trypsin inhibitors demonstrate the same molecular mass; both are adsorbed on DEAE-cellulose and unadsorbed on Affi-gel blue gel but can be separated by ion exchange chromatography on SP-Sepharose. They have approximately the same trypsin inhibitory potency, and neither of them inhibits HIV-1 reverse transcriptase inhibitory activity, HIV-1 integrase inhibitory activity, SARS coronavirus proteinase inhibitory activity, or fungal growth. Both inhibitors exhibit an antiproliferative activity toward L1210 cells albeit with a small difference in potency, while there is little activity toward MBL2 cells. The difference in yields of the two trypsin inhibitors from the white cloud beans is only slight.
The two trypsin inhibitors exhibit N-terminal sequence homology with those of other leguminous trypsin inhibitors such as inhibitors mungbean (Vigna mungo), cowpea (Vigna unguiculata), and lima bean (Phaseolus lunatus). Whereas leguminous antifungal proteins [48, 49], like nonleguminous antifungal proteins [53], are unadsorbed on DEAE-cellulose and adsorbed on Affi-gel blue gel, white cloud bean trypsin inhibitors are adsorbed on DEAE-cellulose and unadsorbed on Affi-gel blue gel. Thus the purification procedure adopted in the present investigation can be conveniently used to separate trypsin inhibitors from antifungal proteins. The antiproliferative activity of white cloud bean trypsin inhibitors is consistent with similar observations on field bean trypsin inhibitor [39, 40, 42, 43, 54]. It is noteworthy that white cloud bean trypsin inhibitors do not exert a similar action on lymphoma MBL2 cells. Thus the action of white cloud bean trypsin inhibitors is tumour cell-specific. The ribosome inactivating protein trichosanthin exerts different antiproliferative potencies toward different tumor cells [55]. In contrast to broad bean trypsin inhibitor [48, 56], those from white cloud bean do not inhibit HIV-1 reverse transcriptase. A variety of trypsin inhibitors exhibit antifungal activity [48, 56]. However, white cloud bean trypsin isoinhibitors lack such activity. This is reminiscent of the finding that lentil and Vigna mungo inhibitors have little or no HIV-1 reverse transcriptase inhibitory and antifungal activities [57, 58].
In summary, the isolation of two trypsin inhibitors with very similar biochemical and biological characteristics from white cloud beans was achieved in the present investigation. The presence of multiple trypsin inhibitors has previously been reported in Momordica cochinchinensis seeds [59]. White cloud bean trypsin inhibitors demonstrate antiproliferative activity against tumor cells but do no inhibit mycelial growth or HIV-1 reverse transcriptase. Previously isolated P. vulgaris trypsin inhibitors have not been so tested [60–64]. They reportedly have a molecular mass of about 9 kDa [63, 65] or 13 kDa [61], smaller than the value of 16 kDa obtained for white cloud bean trypsin inhibitors.
Acknowledgment
This work was financially supported by National Grants of China (nyhyzx07-008 and 2007BAD89B00).
References
- 1.Tanaka AS, Sampaio MU, Marangoni S, et al. Purification and primary structure determination of a Bowman-Birk trypsin inhibitor from Torresea cearensis seeds. Biological Chemistry. 1997;378(3-4):273–281. doi: 10.1515/bchm.1997.378.3-4.273. [DOI] [PubMed] [Google Scholar]
- 2.Minuth T, Krämer B, Lehle K, Jaenicke R, Kohnert U. The spectroscopic analysis, inhibition and binding studies demonstrate the equivalence of Erythrina caffra trypsin inhibitor and the recombinant substitution variant recSerETI. Journal of Biotechnology. 1998;62(3):231–239. doi: 10.1016/s0168-1656(98)00066-2. [DOI] [PubMed] [Google Scholar]
- 3.Devaraj VR, Manjunatha NH. Purification and characterization of a proteinase inhibitor from field bean, Dolichos lablab perpureus L. Journal of Protein Chemistry. 1999;18(1):47–54. doi: 10.1023/a:1020695315964. [DOI] [PubMed] [Google Scholar]
- 4.Pando LA, Di Ciero L, Novello JC, Oliveira B, Weder JKP, Marangoni S. Isolation and characterization of a new trypsin inhibitor from Crotalaria paulina seeds. IUBMB Life. 1999;48(5):519–523. doi: 10.1080/713803553. [DOI] [PubMed] [Google Scholar]
- 5.Ceciliani F, Tava A, Iori R, Mortarino M, Odoardi M, Ronchi S. A trypsin inhibitor from snail medic seeds active against pest proteases. Phytochemistry. 1997;44(3):393–398. doi: 10.1016/s0031-9422(96)00483-9. [DOI] [PubMed] [Google Scholar]
- 6.Lanza A, Tava A, Catalano M, et al. Effects of the Medicago scutellata trypsin inhibitor (MsTI) on cisplatin-induced cytotoxicity in human breast and cervical cancer cells. Anticancer Research. 2004;24(1):227–233. [PubMed] [Google Scholar]
- 7.Park S-S, Sumi T, Ohba H, Nakamura O, Kimura M. Complete amino acid sequences of three proteinase inhibitors from white sword bean (Canavalia gladiata) Bioscience, Biotechnology and Biochemistry. 2000;64(10):2272–2275. doi: 10.1271/bbb.64.2272. [DOI] [PubMed] [Google Scholar]
- 8.Jouvensal L, Quillien L, Ferrasson E, Rahbé Y, Guéguen J, Vovelle F. PA1b, an insecticidal protein extracted from pea seeds (Pisum sativum): 1H-2-D NMR study and molecular modeling. Biochemistry. 2003;42(41):11915–11923. doi: 10.1021/bi034803l. [DOI] [PubMed] [Google Scholar]
- 9.Mello GC, Oliva MLV, Sumikawa JT, et al. Purification and characterization of a new trypsin inhibitor from Dimorphandra mollis seeds. Journal of Protein Chemistry. 2001;20(8):625–632. doi: 10.1023/a:1013764118579. [DOI] [PubMed] [Google Scholar]
- 10.Cavalcanti MDSM, Oliva MLV, Fritz H, et al. Characterization of a Kunitz trypsin inhibitor with one disulfide bridge purified from Swartzia pickellii. Biochemical and Biophysical Research Communications. 2002;291(3):635–639. doi: 10.1006/bbrc.2002.6436. [DOI] [PubMed] [Google Scholar]
- 11.Dattagupta JK, Podder A, Chakrabarti C, Sen U, Dutta SK, Singh M. Structure of a Kunitz-type chymotrypsin inhibitor from winged bean seeds at 2.95 Å resolution. Acta Crystallographica Section D. 1996;52(3):521–528. doi: 10.1107/S0907444996000224. [DOI] [PubMed] [Google Scholar]
- 12.Pando SC, Oliva MLV, Sampaio CAM, Di Ciero L, Novello JC, Marangoni S. Primary sequence determination of a Kunitz inhibitor isolated from Delonix regia seeds. Phytochemistry. 2001;57(5):625–631. doi: 10.1016/s0031-9422(01)00080-2. [DOI] [PubMed] [Google Scholar]
- 13.Garcia VA, Freire MDGM, Novello JC, Marangoni S, Macedo MLR. Trypsin inhibitor from Poecilanthe parviflora seeds: purification, characterization, and activity against pest proteases. Protein Journal. 2004;23(5):343–350. doi: 10.1023/b:jopc.0000032654.67733.d5. [DOI] [PubMed] [Google Scholar]
- 14.Macedo MLR, De Sá CM, Freire MDGM, Parra JRP. A Kunitz-type inhibitor of coleopteran proteases, isolated from adenanthera pavonina L. seeds and its effect on Callosobruchus maculatus. Journal of Agricultural and Food Chemistry. 2004;52(9):2533–2540. doi: 10.1021/jf035389z. [DOI] [PubMed] [Google Scholar]
- 15.Haq SK, Khan RH. Characterization of a Proteinase Inhibitor from Cajanus cajan (L.) Journal of Protein Chemistry. 2003;22(6):543–554. doi: 10.1023/b:jopc.0000005504.57372.5b. [DOI] [PubMed] [Google Scholar]
- 16.Kumar P, Sreerama YN, Gowda LR. Formation of Bowman-Birk inhibitors during the germination of horsegram (Dolichos biflorus) Phytochemistry. 2002;60(6):581–588. doi: 10.1016/s0031-9422(02)00178-4. [DOI] [PubMed] [Google Scholar]
- 17.Campos JE, Whitaker JR, Yip T-T, Hutchens TW, Blanco-Labra A. Unusual structural characteristics and complete amino acid sequence of a protease inhibitor from Phaseolus acutifolius seeds. Plant Physiology and Biochemistry. 2004;42(3):209–214. doi: 10.1016/j.plaphy.2003.12.002. [DOI] [PubMed] [Google Scholar]
- 18.Dodo HW, Viquez OM, Maleki SJ, Konan KN. cDNA clone of a putative peanut (Arachis hypogaea L.) trypsin inhibitor has homology with peanut allergens Ara h 3 and Ara h 4. Journal of Agricultural and Food Chemistry. 2004;52(5):1404–1409. doi: 10.1021/jf034765c. [DOI] [PubMed] [Google Scholar]
- 19.Sattar R, Ali SA, Kamal M, Khan AA, Abbasi A. Molecular mechanism of enzyme inhibition: prediction of the three-dimensional structure of the dimeric trypsin inhibitor from Leucaena leucocephala by homology modelling. Biochemical and Biophysical Research Communications. 2004;314(3):755–765. doi: 10.1016/j.bbrc.2003.12.177. [DOI] [PubMed] [Google Scholar]
- 20.Oliva MLV, R. Mendes C, Juliano MA, et al. Characterization of a tissue kallikrein inhibitor isolated from Bauhinia bauhinioides seeds: inhibition of the hydrolysis of kininogen related substrates. Immunopharmacology. 1999;45(1–3):163–169. doi: 10.1016/s0162-3109(99)00075-2. [DOI] [PubMed] [Google Scholar]
- 21.Di Ciero L, Oliva MLV, Torquato R, et al. The complete amino acid sequence of a trypsin inhibitor from Bauhinia variegata var. candida seeds. Journal of Protein Chemistry. 1998;17(8):827–834. doi: 10.1023/a:1020734519908. [DOI] [PubMed] [Google Scholar]
- 22.Oliva MLV, Andrade S, Batista IFC, et al. Human plasma kallikrein and tissue kallikrein binding to a substrate based on the reactive site of a factor Xa inhibitor isolated from Bauhinia ungulata seeds. Immunopharmacology. 1999;45(1–3):145–149. doi: 10.1016/s0162-3109(99)00146-0. [DOI] [PubMed] [Google Scholar]
- 23.Rao KN, Hegde SS, Lewis RJ, Suresh CG. Crystallization and preliminary X-ray diffraction studies of a Bowman-Birk inhibitor from Vigna unguiculata seeds. Acta Crystallographica Section D. 1999;55(11):1920–1922. doi: 10.1107/s0907444999010021. [DOI] [PubMed] [Google Scholar]
- 24.Weder JKP, Hinkers SC. Complete amino acid sequence of the lentil trypsin-chymotrypsin inhibitor LCI-1.7 and a discussion of atypical binding sites of Bowman-Birk inhibitors. Journal of Agricultural and Food Chemistry. 2004;52(13):4219–4226. doi: 10.1021/jf030768d. [DOI] [PubMed] [Google Scholar]
- 25.Koiwa H, Shade RE, Zhu-Salzman K, et al. Phage display selection can differentiate insecticidal activity of soybean cystatins. Plant Journal. 1998;14(3):371–379. doi: 10.1046/j.1365-313x.1998.00119.x. [DOI] [PubMed] [Google Scholar]
- 26.Troncoso MF, Zolezzi PC, Hellman U, Wolfenstein-Todel C. A novel trypsin inhibitor from Peltophorum dubium seeds, with lectin-like properties, triggers rat lymphoma cell apoptosis. Archives of Biochemistry and Biophysics. 2003;411(1):93–104. doi: 10.1016/s0003-9861(02)00726-9. [DOI] [PubMed] [Google Scholar]
- 27.Delgado-Vargas F, López-Valdés HE, Valdés-Rodríguez S, Blanco-Labra A, Chagolla-López A, López-Valenzuela EDJ. Isolation and properties of a Kunitz-type protein inhibitor obtained from Pithecellobium dulce seeds. Journal of Agricultural and Food Chemistry. 2004;52(20):6115–6121. doi: 10.1021/jf049694b. [DOI] [PubMed] [Google Scholar]
- 28.Deshimaru M, Hanamoto R, Kusano C, Yoshimi S, Terada S. Purification and characterization of proteinase inhibitors from wild soja (Glycine soja) seeds. Bioscience, Biotechnology and Biochemistry. 2002;66(9):1897–1903. doi: 10.1271/bbb.66.1897. [DOI] [PubMed] [Google Scholar]
- 29.McPhalen CA, James MNG. Crystal and molecular structure of the serine proteinase inhibitor CI-2 from barley seeds. Biochemistry. 1987;26(1):261–269. doi: 10.1021/bi00375a036. [DOI] [PubMed] [Google Scholar]
- 30.Gerhardt IR, Paes NS, Bloch C, Jr., et al. Molecular characterization of a new arcelin-5 gene. Biochimica et Biophysica Acta—Gene Structure and Expression. 2000;1490(1-2):87–98. doi: 10.1016/s0167-4781(99)00219-5. [DOI] [PubMed] [Google Scholar]
- 31.Chen J-J, Chen G-H, Hsu H-C, Li S-S, Chen C-S. Cloning and functional expression of a mungbean defensin VrD1 in Pichia pastoris. Journal of Agricultural and Food Chemistry. 2004;52(8):2256–2261. doi: 10.1021/jf030662i. [DOI] [PubMed] [Google Scholar]
- 32.Haq SK, Atif SM, Khan RH. Protein proteinase inhibitor genes in combat against insects, pests, and pathogens: natural and engineered phytoprotection. Archives of Biochemistry and Biophysics. 2004;431(1):145–159. doi: 10.1016/j.abb.2004.07.022. [DOI] [PubMed] [Google Scholar]
- 33.Bell HA, Fitches EC, Down RE, et al. Effect of dietary cowpea trypsin inhibitor (CpTI) on the growth and development of the tomato moth Lacanobia oleracea (Lepidoptera: Noctuidae) and on the success of the gregarious ectoparasitoid Eulophus pennicornis (Hymenoptera: Eulophidae) Pest Management Science. 2001;57(1):57–65. doi: 10.1002/1526-4998(200101)57:1<57::AID-PS273>3.0.CO;2-4. [DOI] [PubMed] [Google Scholar]
- 34.Foissac X, Edwards MG, Du JP, Gatehouse AMR, Gatehouse JA. Putative protein digestion in a sap-sucking homopteran plant pest (rice brown plant hopper; Nilaparvata lugens: Delphacidae)—identification of trypsin-like and cathepsin B-like proteases. Insect Biochemistry and Molecular Biology. 2002;32(9):967–978. doi: 10.1016/s0965-1748(02)00033-4. [DOI] [PubMed] [Google Scholar]
- 35.Giri AP, Harsulkar AM, Ku MSB, et al. Identification of potent inhibitors of Helicoverpa armigera gut proteinases from winged bean seeds. Phytochemistry. 2003;63(5):523–532. doi: 10.1016/s0031-9422(03)00181-x. [DOI] [PubMed] [Google Scholar]
- 36.Goetz IE, Weinstein C, Roberts E. Effects of protease inhibitors on growth of hamster tumor cells in culture. Cancer Research. 1972;32(11):2469–2474. [PubMed] [Google Scholar]
- 37.von Hofe E, Newberne PM, Kennedy AR. Inhibition of N-nitrosomethylbenzylamine-induced esophageal neoplasms by the Bowman-Birk protease inhibitor. Carcinogenesis. 1991;12(11):2147–2150. doi: 10.1093/carcin/12.11.2147. [DOI] [PubMed] [Google Scholar]
- 38.Kennedy AR, Billings PC, Maki PA, Newberne P. Effects of various preparations of dietary protease inhibitors on oral carcinogenesis in hamsters induced by DMBA. Nutrition and Cancer. 1993;19(2):191–200. doi: 10.1080/01635589309514249. [DOI] [PubMed] [Google Scholar]
- 39.Banerji AP, Fernandes AO. Field bean protease inhibitor preparations, unlike methotrexate, can completely suppress Yoshida sarcoma tumor in rats. Cell Biology International. 1994;18(11):1025–1034. doi: 10.1006/cbir.1994.1026. [DOI] [PubMed] [Google Scholar]
- 40.Fernandes AO, Banerji AP. Inhibition of benzopyrene-induced forestomach tumors by field bean protease inhibitor(s) Carcinogenesis. 1995;16(8):1843–1846. doi: 10.1093/carcin/16.8.1843. [DOI] [PubMed] [Google Scholar]
- 41.Birk Y. Protein proteinase inhibitors in legume seeds–overview. Archivos Latinoamericanos de Nutrición. 1996;44(4, supplement 1):26S–30S. [PubMed] [Google Scholar]
- 42.Fernandes AO, Banerji AP. Erratum: the field bean protease inhibitor can effectively suppress 72-dimethylbenz[a]anthracene-induced skin tumorigenesis in mice. Cancer Letters. 1996;106(1):p. 145. doi: 10.1016/0304-3835(96)04253-x. [DOI] [PubMed] [Google Scholar]
- 43.Banerji A, Fernandes A, Bane S, Ahire S. The field bean protease inhibitor has the potential to suppress B16F10 melanoma cell lung metastasis in mice. Cancer Letters. 1998;129(1):15–20. doi: 10.1016/s0304-3835(98)00090-1. [DOI] [PubMed] [Google Scholar]
- 44.Sammon AM. Protease inhibitors and carcinoma of the esophagus. Cancer. 1998;83(3):405–408. [PubMed] [Google Scholar]
- 45.Kobayashi H, Suzuki M, Kanayama N, Terao T. A soybean Kunitz trypsin inhibitor suppresses ovarian cancer cell invasion by blocking urokinase upregulation. Clinical and Experimental Metastasis. 2004;21(2):159–166. doi: 10.1023/b:clin.0000024751.73174.c2. [DOI] [PubMed] [Google Scholar]
- 46.Laemmli UK, Favre M. Maturation of the head of bacteriophage T4. I. DNA packaging events. Journal of Molecular Biology. 1973;80(4):575–599. doi: 10.1016/0022-2836(73)90198-8. [DOI] [PubMed] [Google Scholar]
- 47.Azarkan M, Dibiani R, Goormaghtigh E, Raussens V, Baeyens-Volant D. The papaya Kunitz-type trypsin inhibitor is a highly stable β-sheet glycoprotein. Biochimica et Biophysica Acta. 2006;1764(6):1063–1072. doi: 10.1016/j.bbapap.2006.02.014. [DOI] [PubMed] [Google Scholar]
- 48.Ye XY, Ng TB, Rao PF. A bowman-birk-type trypsin-chymotrypsin inhibitor from broad beans. Biochemical and Biophysical Research Communications. 2001;289(1):91–96. doi: 10.1006/bbrc.2001.5965. [DOI] [PubMed] [Google Scholar]
- 49.Wong JH, Ng TB. Gymnin, a potent defensin-like antifungal peptide from the Yunnan bean (Gymnocladus chinensis Baill) Peptides. 2003;24(7):963–968. doi: 10.1016/s0196-9781(03)00192-x. [DOI] [PubMed] [Google Scholar]
- 50.Wang H, Ng TB. Pleureryn, a novel protease from fresh fruiting bodies of the edible mushroom Pleurotus eryngii. Biochemical and Biophysical Research Communications. 2001;289(3):750–755. doi: 10.1006/bbrc.2001.6037. [DOI] [PubMed] [Google Scholar]
- 51.Wang H, Ng TB. Isolation and characterization of velutin, a novel low-molecular-weight ribosome-inactivating protein from winter mushroom (Flammulina velutipes) fruiting bodies. Life Sciences. 2001;68(18):2151–2158. doi: 10.1016/s0024-3205(01)01023-2. [DOI] [PubMed] [Google Scholar]
- 52.Leung EHW, Wong JH, Ng TB. Concurrent purification of two defense proteins from French bean seeds: a defensin-like antifungal peptide and a hemagglutinin. Journal of Peptide Science. 2008;14(3):349–353. doi: 10.1002/psc.946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wang HX, Ng TB. Isolation of cucurmoschin, a novel antifungal peptide abundant in arginine, glutamate and glycine residues from black pumpkin seeds. Peptides. 2003;24(7):969–972. doi: 10.1016/s0196-9781(03)00191-8. [DOI] [PubMed] [Google Scholar]
- 54.Banerji A, Fernandes A, Bane S. Treatment with field bean protease inhibitor can effectively repress ethylnitrosourea (ENU)-induced neoplasms of the nervous system in Sprague-Dawley rats. Cancer Letters. 1998;130(1-2):161–167. doi: 10.1016/s0304-3835(98)00135-9. [DOI] [PubMed] [Google Scholar]
- 55.Puri M, Kaur I, Kanwar RK, Gupta RC, Chauhan A, Kanwar JR. Ribosome inactivating proteins (RIPs) from Momordica charantia for anti viral therapy. Current Molecular Medicine. 2009;9(9):1080–1094. doi: 10.2174/156652409789839071. [DOI] [PubMed] [Google Scholar]
- 56.Ye XY, Ng TB. A new peptidic protease inhibitor from Vicia faba seeds exhibits antifungal, HIV-1 reverse transcriptase inhibiting and mitogenic activities. Journal of Peptide Science. 2002;8(12):656–662. doi: 10.1002/psc.425. [DOI] [PubMed] [Google Scholar]
- 57.Cheung AHK, Ng TB. Isolation and characterization of a trypsin-chymotrypsin inhibitor from the seeds of green lentil (Lens culinaris) Protein and Peptide Letters. 2007;14(9):859–864. doi: 10.2174/092986607782110310. [DOI] [PubMed] [Google Scholar]
- 58.Cheung AHK, Wong JH, Ng TB. Trypsin-chymotrypsin inhibitors from Vigna mungo seeds. Protein and Peptide Letters. 2009;16(3):277–284. doi: 10.2174/092986609787601714. [DOI] [PubMed] [Google Scholar]
- 59.Wong RCH, Fong WP, Ng TB. Multiple trypsin inhibitors from Momordica cochinchinensis seeds, the Chinese drug mubiezhi. Peptides. 2004;25(2):163–169. doi: 10.1016/j.peptides.2004.01.002. [DOI] [PubMed] [Google Scholar]
- 60.Birk Y. Trypsin isoinhibitors from garden beans (Phaseolus vulgaris) Methods in Enzymology. 1976;45:710–716. doi: 10.1016/s0076-6879(76)45063-2. [DOI] [PubMed] [Google Scholar]
- 61.Miyoshi M, Hamaguchi Y, Matsumoto K, Mizuno I. The isolation and characterization of a trypsin inhibitor from Kintoki bean (Phaseolus vulgaris) Journal of Nutritional Science and Vitaminology. 1978;24(2):195–204. doi: 10.3177/jnsv.24.195. [DOI] [PubMed] [Google Scholar]
- 62.Gerstenberg H, Belitz H-D, Weder JKP. Isolation and characterization of some proteinase inhibitors from Phaseolus vulgaris var. nanus. Zeitschrift für Lebensmittel Untersuchung und Forschung. 1980;171(1):28–34. doi: 10.1007/BF01044414. [DOI] [PubMed] [Google Scholar]
- 63.De Carvalho PGB, Bloch C, Jr., Morhy L, Da Silva MCM, De Mello LV, Neshich G. Amino acid sequence of the Phaseolus vulgaris var. “Fogo na Serra” inhibitor and interactive surface modeling for the enzyme-inhibitor complex. Journal of Protein Chemistry. 1996;15(6):591–598. doi: 10.1007/BF01908541. [DOI] [PubMed] [Google Scholar]
- 64.Kasahara K, Hayashi K, Arakawa T, et al. Complete sequence, subunit structure, and complexes with pancreatic α-amylase of an α-amylase inhibitor from Phaseolus vulgaris white kidney beans. Journal of Biochemistry. 1996;120(1):177–183. doi: 10.1093/oxfordjournals.jbchem.a021381. [DOI] [PubMed] [Google Scholar]
- 65.Sattar AKMA, Yamamoto N, Yoshimoto T, Tsuru D. Purification and characterization of an extracellular prolyl endopeptidase from Agaricus bisporus. Journal of Biochemistry. 1990;107(2):256–261. doi: 10.1093/oxfordjournals.jbchem.a123035. [DOI] [PubMed] [Google Scholar]