Abstract
Bacteria and chloroplasts require the ring-forming cytoskeletal protein FtsZ for division. Although bacteria accomplish division with a single FtsZ, plant chloroplasts require two FtsZ types for division, FtsZ1 and FtsZ2. These proteins colocalize to a mid-plastid Z ring, but their biochemical relationship is poorly understood. We investigated the in vitro behavior of recombinant FtsZ1 and FtsZ2 separately and together. Both proteins bind and hydrolyze GTP, although GTPase activities are low compared with the activity of Escherichia coli FtsZ. Each protein undergoes GTP-dependent assembly into thin protofilaments in the presence of calcium as a stabilizing agent, similar to bacterial FtsZ. In contrast, when mixed without calcium, FtsZ1 and FtsZ2 exhibit slightly elevated GTPase activity and coassembly into extensively bundled protofilaments. Coassembly is enhanced by FtsZ1, suggesting that it promotes lateral interactions between protofilaments. Experiments with GTPase-deficient mutants reveal that FtsZ1 and FtsZ2 form heteropolymers. Maximum coassembly occurs in reactions containing equimolar FtsZ1 and FtsZ2, but significant coassembly occurs at other stoichiometries. The FtsZ1:FtsZ2 ratio in coassembled structures mirrors their input ratio, suggesting plasticity in protofilament and/or bundle composition. This behavior contrasts with that of α- and β-tubulin and the bacterial tubulin-like proteins BtubA and BtubB, which coassemble in a strict 1:1 stoichiometry. Our findings raise the possibility that plasticity in FtsZ filament composition and heteropolymerization-induced bundling could have been a driving force for the coevolution of FtsZ1 and FtsZ2 in the green lineage, perhaps arising from an enhanced capacity for the regulation of Z ring composition and activity in vivo.
Keywords: Arabidopsis, Chloroplast, Cytoskeleton, Protein Self-assembly, Tubulin, Bacterial Cell Division, Chloroplast Division, FtsZ, Plastid Division
Introduction
Cell division in bacteria and chloroplast division in plants both require FtsZ, a tubulin-like GTPase that functions as a contractile ring, the Z ring, inside the cell or chloroplast. Mutations in bacterial FtsZ genes disrupt cell division, resulting in long filamented cells (reviewed in Refs. 1–3). Purified bacterial FtsZ undergoes GTP-dependent, hydrolysis-independent polymerization primarily into single protofilaments (4, 5), but protofilament sheets and bundles form in the presence of stabilizing agents that promote lateral interactions (6–8). Polymerization stimulates FtsZ GTPase activity because the catalytic site for GTP hydrolysis lies in the longitudinal interface between adjacent monomers within the polymer (9). GTP hydrolysis destabilizes protofilaments, leading to disassembly (10). The in vivo structure of the bacterial Z ring is not yet clear, but recent models suggest it may be built from short overlapping protofilaments that are stabilized at the division site by accessory factors (11, 12). The Z ring functions partly as a scaffold for other cell division proteins and recently has also been shown to provide contractile force for membrane constriction (13, 14).
In most bacteria, including the cyanobacterial relatives of chloroplasts, a single FtsZ accomplishes cell division. In contrast, chloroplast division in plants and green algae involves two forms of FtsZ, FtsZ1 and FtsZ2, that colocalize to a mid-plastid Z ring (reviewed in Refs. 15 and 16). Depletion or overexpression of either protein impairs division and results in enlarged chloroplasts, which are the phenotypic equivalent of the bacterial filamentation phenotype (17–19). These findings, along with data showing that the molar ratio between Arabidopsis FtsZ1 and FtsZ2 remains constant throughout leaf development (20), suggest that their stoichiometry is important for chloroplast division activity. In vitro, FtsZ1 and FtsZ2 have been reported to exhibit GTPase activity and to undergo GTP-dependent assembly into protofilaments (21–23). However, the previous analyses have been limited in scope, and whether FtsZ1 and FtsZ2 form heteropolymers has not been investigated. Heteropolymerization is suggested by the observation that FtsZ1 and FtsZ2 are always tightly colocalized in vivo even when FtsZ filament morphology is perturbed (24, 25) and by studies showing that FtsZ1 and FtsZ2 interact in two-hybrid assays (26) and in a native complex (20).
Here, we present a detailed study of the in vitro biochemical behavior of FtsZ1 and FtsZ2 individually and in combination. We show that, similar to bacterial FtsZ, FtsZ1 and FtsZ2 both bind and hydrolyze GTP and individually undergo GTP-dependent assembly into thin protofilaments, although only when stabilized by calcium. However, when mixed, FtsZ1 and FtsZ2 exhibit enhanced GTP-dependent protofilament formation and extensive protofilament bundling in the absence of calcium. Experiments with mutant proteins lacking GTPase activity show that FtsZ1 and FtsZ2 form heteropolymers. Our findings yield new insight into the assembly properties of chloroplast FtsZ1 and FtsZ2 and provide clues to understanding the mechanistic driving force for the coevolution of two FtsZ families in green algae and plants.
EXPERIMENTAL PROCEDURES
Production of Recombinant FtsZ Proteins
FtsZ1 (AtFtsZ1-1, At5g55280) and FtsZ2-1 (AtFtsZ2-1, At2g36250) lacking their transit peptides (57 and 48 N-terminal amino acids, respectively; see supplemental Methods) were expressed as either C-terminally His6-tagged or untagged fusion proteins. All of the fragments were PCR-amplified from AtFtsZ1-1 (GenBankTM accession number AY113896) or AtFtsZ2-1 (27) cDNAs. Primers used for the tagged proteins were 5′-AGTGGTCCATGGCCAGGTCTAAGTCGATGCGATTG-3′ (forward) and 5′-TGCACCCTCGAGCTAATGATGATGATGATGATGGAAGAAAAGTCTACGGGGA-3′ (reverse) for FtsZ1 and 5′-AGTGGTCCATGGCCGCCGCTCAGAAATCTGAATC-3′ (forward) and 5′-TGCACCCTCGAGTTAATGATGATGATGATGATGGACTCGGGGATAACGAGAGCTG-3′ (reverse) for FtsZ2. The same forward primers were used for the untagged proteins; reverse primers were 5′-TGCACCCTCGAGCTAATGGAAGAAAAGTCTACGGGGA-3′for FtsZ1 and 5′-TGCACCCTCGAGTTAGACTCGGGGATAACGAGAGCTG-3′ for FtsZ2. PCR fragments were subcloned into the NcoI and XhoI sites of the expression vector pDB328 (28).
His-tagged and untagged FtsZ1 and FtsZ2 were expressed in Rosetta (DE3) cells (Novagen) overexpressing the Escherichia coli ftsQAZ operon (29). Overnight cultures were grown at 37 °C and diluted 1:1000 (culture volume) into fresh LB medium. When A600 reached ∼0.6, 0.6 mm isopropyl β-d-thiogalactopyranoside was added, and the culture was grown for 3 h at 37 °C. More than 80% of expressed plant FtsZ was found in inclusion bodies irrespective of expression conditions. The cells were harvested at 9,000 × g for 10 min at 4 °C and stored at −80 °C.
Prior to purification, frozen cell pellets were resuspended in 0.025 culture volume in extraction buffer (25 mm Tris-Cl, pH 8.0, 500 mm NaCl, 10 mm imidazole with Roche Complete EDTA-free protease inhibitor mixture), and the cells were sonicated six times for 30 s at full tip power with 5 min between sonications using a Bronson microtip sonicator. The lysates were centrifuged at 18,000 × g for 20 min at 4 °C, and the supernatant was discarded. Inclusion bodies in the pellet were resuspended in 0.0125 culture volume in 2× extraction buffer, sonicated again, and solubulized by adding urea and water to a final volume of 0.025 culture volume in 6 m urea. Insoluble material was removed by centrifugation at 18,000 × g for 20 min at 4 °C.
Purification and Refolding of His-tagged FtsZ Proteins
Urea-solubilized lysates were applied to a 15-ml nickel-Sepharose column (GE Healthcare) at 1 ml min−1 and washed with 5 column volumes of buffer A (25 mm Tris-Cl, pH 8.0, 500 mm NaCl, 10 mm imidazole, 6 m urea). FtsZ was eluted with a linear gradient of 0.01–1 m imidazole in buffer A at 5 ml min−1; fractions containing FtsZ proteins, which typically eluted between ∼100 and 300 mm imidazole, were pooled and stored at −80 °C for up to 3 months prior to refolding.
His-tagged FtsZ1 and FtsZ2 were refolded by dialysis immediately before use in biochemical assays. Imidazole was first removed by dialysis against 25 mm Tris-Cl, pH 8.0, 500 mm NaCl, 3 m urea. GDP was then added directly to the dialysis bag at a 5-fold molar excess relative to the total protein concentration, and the contents were dialyzed against four changes of TMK (25 mm Tris-Cl, pH 7.0, 5 mm MgCl2, 100 mm KCl). Aggregated protein was removed by centrifugation at 18,000 × g for 20 min at 4 °C. Protein in the supernatant was concentrated by ultrafiltration (Amicon Centricon; molecular mass cut-off, 20 kDa) to ∼15 μm. Aggregated FtsZ and minor contaminants were removed by size exclusion chromatography on a HiPrep 26/10 column (GE Healthcare) with TMK at 10 ml min−1, and 5-ml fractions were collected. Peak protein-containing fractions were pooled and concentrated to 15–20 μm by ultrafiltration. The proteins were stored at 0 °C for up to 1 week.
SDS-PAGE and quantitative amino acid analysis were used to assess the purity of recombinant FtsZs. A typical preparation was >95% pure. Quantitative amino acid analysis also demonstrated that the BCA assay underestimates recombinant FtsZ1 and FtsZ2 by ∼20%, similar to EcFtsZ (30) and consistent with our previous quantitative analyses (20).
Purification of Untagged FtsZ Proteins
Untagged FtsZ1 and FtsZ2 were purified by assembly-based purification. Urea-solubilized proteins were dialyzed against HMK (50 mm HEPES-KOH, pH 7.0, 100 mm KCl, and 5 mm MgSO4), and CaCl2 was then added to a final concentration of 10 mm. Assembly of FtsZ1 or FtsZ2 was triggered by adding GTP to a final concentration of 2 mm at room temperature (23–25 °C). After 30 min, assembled protein was centrifuged at 20,000 × g for 20 min at 4 °C and resuspended in 6 m urea in HMK. Untagged FtsZ1 and FtsZ2 were subsequently refolded and subjected to size exclusion chromatography as described above for the His-tagged proteins. E. coli FtsZ (EcFtsZ) was prepared by assembly as described (31) for use in control experiments.
Site-directed Mutagenesis
Plasmids encoding the T7 loop mutants FtsZ1D275A and FtsZ2D322A were produced by site-directed mutagenesis (32) of the His-tagged FtsZ1 and FtsZ2 expression plasmids using the primers 5′-GTCAATGTGGATTTTGCAGCTGTGAAGGCAGTCATGAAA-3′ and 5′-TTTCATGACTGCCTTCACAGCTGCAAAATCCACATTGAC-3′ for FtsZ1 and 5′-GTGAATGTGGATTTTGCTGCTGTGAGAGCTATAATGGCA-3′ and 5′-TGCCATTATAGCTCTCACAGCAGCAAAATCCACATTCAC-3′ for FtsZ2. Expression and purification were performed as described for the His-tagged unmutagenized proteins.
GTP Binding and Hydrolysis Assays
GTP binding to FtsZ1, FsZ2, FtsZ1D275A, FtsZ2D322A, and EcFtsZ was assayed as described (33) in HMK containing 1 mm GTP and 5 μm total protein. GTPase activity was measured with a Pi release assay (34) using 5 μm total protein (except in supplemental Fig. S7) in HMK. Nucleotide and temperature were varied as described under “Results” and in the figure legends. For all of the GTPase and assembly assays, guanine nucleotide concentration in stock solutions was verified spectrophotometrically prior to use. GTPase activities were compared using the Student's t test.
Sedimentation Assays
Plant FtsZs (5 μm total protein) were incubated at 25 °C in HMK containing 1 or 3 mm GTP or GDP as indicated. Other assay conditions were varied as described under “Results” and in the figure legends. The aliquots were removed after 3 and 30 min and centrifuged at 51,000 × g for 30 min at 4 °C. Supernatant and pellet fractions were analyzed by SDS-PAGE and Coomassie staining. The proteins in the pellet fractions were quantified by densitometry.
Light Scattering Assays
90° light scattering assays were conducted at room temperature as described for EcFtsZ (35) using a Photon Technology International fluorescence spectrophotometer equipped with a model 814 photomultiplier operated in digital mode at 1000 V. The assays were performed with 0.5-nm excitation and 1-nm emission slit widths and 350-nm excitation and emission wavelengths. Assembly was carried out in HMK at room temperature. The buffers were filtered through a 0.22-μm filter prior to use, and the protein preparations were centrifuged for 30 min at 51,000 × g at 25 °C immediately before assay. The maximum assembly rates were derived by linear regression fit of the maximum linear increase in light scattering (35).
Electron Microscopy
FtsZ assembly reactions prepared as described for light scattering assays were performed at room temperature with 5 μm total protein. After 10 min, 2 μl from each reaction was pipetted onto a carbon-coated 400-mesh nickel grid and stained with 2% aqueous uranyl acetate. The assembled structures were visualized with a JEOL100 CXII (Japan Electron Optics Laboratories) transmission electron microscope at an accelerating voltage of 100 kV at 10,000–450,000× magnification.
RESULTS
Production of Recombinant Arabidopsis FtsZ1 and FtsZ2
FtsZ1 and FtsZ2 are encoded in the nucleus as preproteins and targeted to the chloroplast by transit peptides, which are cleaved upon import (36, 37). To produce recombinant Arabidopsis thaliana FtsZ1 (AtFtsZ1-1, At5g55280) and FtsZ2 (AtFtsZ2-1, At2g36250) resembling the mature in vivo forms, we expressed constructs lacking transit peptides as His-tagged fusion proteins in E. coli. A second FsZ2 isoform in Arabidopsis, AtFtsZ2-2, is functionally redundant with AtFtsZ2-1 (19); therefore, we used only AtFtsZ2-1 (henceforth FtsZ2) in our experiments. Because recombinant proteins were mostly in inclusion bodies, they were solublized and subjected to nickel affinity chromatography in urea, then refolded by dialysis, and further purified by size exclusion chromatography. Purity of both FtsZ1 and FtsZ2 was typically >95%. Most of the assays were carried out with these His-tagged proteins; however, untagged proteins behaved similarly to the tagged forms (described below), indicating that the tags did not interfere with or significantly alter FtsZ1 or FtsZ2 behavior.
FtsZ1 and FtsZ2 Bind and Hydrolyze GTP Individually and When Mixed
FtsZ1 and FtsZ2 contain all features important for GTP binding and hydrolysis in bacterial FtsZ (15, 25). When assayed separately for GTP binding in 1 mm GTP (33), FtsZ1 and FtsZ2 bound 1.05 ± 0.03 and 0.98 ± 0.22 mol of GTP/mol of protein, respectively. In control experiments, neither FtsZ1 nor FtsZ2 bound detectable levels of ATP, indicating specificity for guanine nucleotides. E. coli FtsZ (EcFtsZ) used as a positive control bound 1.09 ± 0.17 mol of GTP/FtsZ but did not bind ATP, consistent with published results (33, 38, 39).
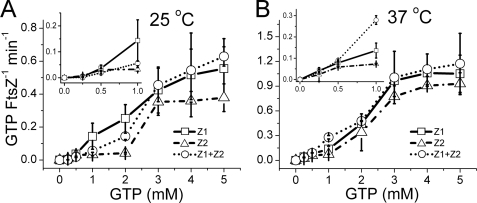
GTP hydrolysis rates for FtsZ1 and FtsZ2 (5 μm) were assayed for 30 min at various GTP concentrations at 25 and 37 °C. Little activity was observed below 0.5 mm GTP (Fig. 1, insets), but the rates increased above 0.5 mm GTP and began to plateau at 3 mm GTP. The maximum activities at 25 °C were 0.56 and 0.38 GTP FtsZ−1 min−1 for FtsZ1 and FtsZ2, respectively (Fig. 1A). The corresponding maximum activities at 37 °C were 1.1 and 0.93 GTP FtsZ−1 min−1 (Fig. 1B). Mutation of the predicted catalytic T7 loops (9) in both proteins abolished their GTPase activities (described below). In control assays, maximum EcFtsZ activities at 25 and 37 °C were 2.1 and 5.0 GTP FtsZ−1 min−1, respectively (supplemental Fig. S1, A and B), consistent with reported data (30, 33). The ability of FtsZ1 and FtsZ2 to each hydrolyze GTP indicates that their active sites can be completed by interaction between identical monomers, as occurs for bacterial FtsZ (9).
FIGURE 1.
GTP hydrolysis rates of FtsZ1 and FtsZ2 assayed separately and together. GTPase activities of 5 μm FtsZ1 (□, solid lines), 5 μm FtsZ2 (▵, dashed and dotted lines), and 2.5 μm each FtsZ1 plus FtsZ2 (○, dotted lines) were assayed at the indicated GTP concentrations. A, 25 °C. B, 37 °C.
We also investigated the effects of mixing FtsZ1 and FtsZ2 (2.5 μm each) on GTPase activity. Instead of showing an average of their individual activities as would be expected if FtsZ1 and FtsZ2 did not interact, GTP hydrolysis was stimulated slightly at 3 mm GTP and above at 25 °C (Fig. 1A; also see Fig. 7B) and above 0.5 mm GTP at 37 °C (Fig. 1B). Maximum activities in the mixed reactions were 0.63 and 1.17 GTP FtsZ−1 min−1 at 25 and 37 °C, respectively. Although the stimulation in activity of the mixture was small, at 3 mm GTP it was statistically significant (see Fig. 7B, left three sets of bars). These results suggest that FtsZ1 and FsZ2 interact and that interaction between FtsZ1 and FtsZ2 subunits creates a slightly more active GTPase than interaction between identical subunits, particularly when compared with FtsZ2 (Fig. 1). Above 0.5 mm GTP, the maximum activities for the individual and mixed proteins (Fig. 1) were considerably lower than those of EcFtsZ (supplemental Fig. S1, A and B), indicating that the plant proteins hydrolyze GTP more slowly. GTP hydrolysis was not detected above the background control (heated proteins) in the presence of GDP, GTP plus EDTA, or ATP (supplemental Fig. S1C).
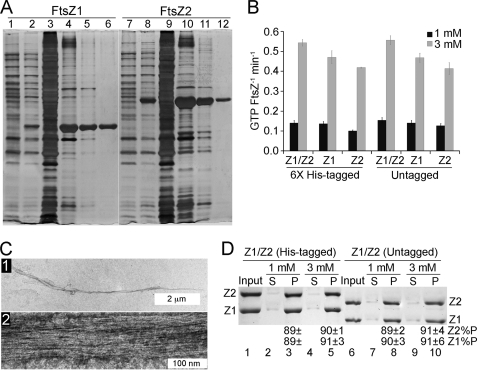
FIGURE 7.
Comparison of untagged and His-tagged FtsZ1 and FtsZ2. A, SDS-PAGE and silver staining of assembly-purified untagged FtsZ1 (lanes 1–6) and FtsZ2 (lanes 7–12). Lanes 1 and 7, uninduced crude cell extracts; lanes 2 and 8, crude cell extracts from isopropyl β-d-thiogalactopyranoside-induced cultures; lanes 3 and 9, soluble fractions from isopropyl β-d-thiogalactopyranoside-induced cultures; lanes 4 and 10, insoluble fractions from isopropyl β-d-thiogalactopyranoside-induced cultures (inclusion bodies); lanes 5 and 6, FtsZ1 before and after assembly purification, respectively; lanes 11 and 12, FtsZ2 before and after assembly purification, respectively. B, GTP hydrolysis rates of His-tagged and untagged FtsZ1 (Z1) and FtsZ2 (Z2) assayed individually (5 μm) or mixed 1:1 (2.5 μm each; Z1/Z2) with 1 mm (black bars) or 3 mm (gray bars) GTP in HMK at 25 °C. At 3 mm GTP, the activity of equally mixed His-tagged FtsZ1 + FtsZ2 is significantly higher than that of FtsZ1 (p = 0.05, n = 6) and FtsZ2 (p = 0.01, n = 6), and the activity of equally mixed untagged FtsZ1 + FtsZ2 is significantly higher than that of FtsZ1 (p = 0.01, n = 3) and FtsZ2 (p = 0.01, n = 3). C, electron micrographs of coassembled untagged FtsZ1 and FtsZ2 (2.5 μm each) in HMK with 1 mm GTP. Scale bars, 2 μm and 100 nm in panels 1 and 2, respectively. D, coassembly of His-tagged (lanes 2–5) and untagged (lanes 7–10) FtsZ1 and FtsZ2 (2.5 μm each) in HMK with 1 mm (lanes 2, 3, 7, and 8) or 3 mm (lanes 4, 5, 9, and 10) GTP monitored by sedimentation as in Fig. 2. Input, FtsZ1 (Z1) and FtsZ2 (Z2) in the starting reactions (lanes 1 and 6); S, supernatant; P, pellet. Percentages of input FtsZ1 (Z1%P) and FtsZ2 (Z2%P) in the pellet fraction ± S.E. (n = 2) are shown below the relevant lanes.
FtsZ1 and FtsZ2 Each Exhibit GTP-dependent Assembly in the Presence of Calcium
To quantify the assembly competence of recombinant FtsZ1 and FtsZ2, we performed sedimentation assays (10). The reactions were carried out at room temperature (23–25 °C), similar to the temperature at which Arabidopsis is typically grown. FtsZ1 and FtsZ2 (5 μm) were incubated separately with 1 mm GTP or GDP in either HMK (see “Experimental Procedures”) or HMK containing 5 mm CaCl2 (HMKCa). After 3 and 30 min, the reactions were centrifuged, and pellet and supernatant fractions were analyzed by SDS-PAGE and Coomassie staining. Assembled structures in some reactions were visualized by negative stain transmission electron microscopy (EM)4 (8).
In HMK lacking nucleotide, nearly all of the FtsZ1 and FtsZ2 remained in the supernatant after 30 min (Fig. 2A). The addition of GTP (HMK-GTP) slightly increased protein in the pellet fractions to ∼3 and 7% of the total FtsZ1 and FtsZ2, respectively (Fig. 2B). Assembled structures in these reactions were not detected by EM, presumably because of their low abundance. In contrast, when incubated in HMKCa with GTP (HMKCa-GTP), protein in the pellet fractions after 30 min increased to 83 and 79% of the total for FtsZ1 and FtsZ2, respectively (Fig. 2D, lanes 5 and 10), and filaments could be observed in these reactions by EM (Fig. 3, A and B; described below). These findings indicate that Ca2+ stabilizes FtsZ1 and FtsZ2 filaments formed in the presence of GTP. However, the amount of protein in the pellet fractions in Fig. 2D probably overestimates the proportion undergoing GTP-dependent assembly because a significant, although smaller, proportion of FtsZ1 and FsZ2 also pelleted in HMKCa lacking nucleotide (supplemental Fig. S2, A and B, lanes 3 and 5) or containing GDP (HMKCa-GDP) (Fig. 2C and supplemental Fig. S2, A and B, lanes 7 and 9), suggesting the possibility of CaCl2-induced aggregation. EM confirmed that protein aggregates but not filaments formed in both HMKCa and HMKCa-GDP (supplemental Fig. S2, C–F). However, fewer and smaller aggregates were visible in pellets from the HMKCa-GTP treatments (Fig. 3, A and B, asterisks), suggesting that protofilament assembly may compete with aggregation when GTP is present and making quantitative estimates of GTP-dependent assembly in the presence of Ca2+ unreliable.
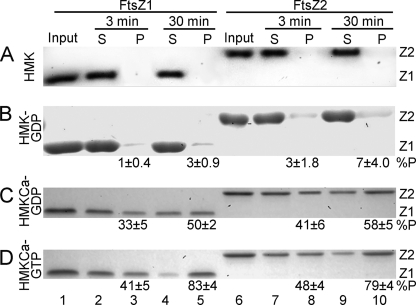
FIGURE 2.
GTP-dependent assembly of FtsZ1 and FtsZ2 monitored by sedimentation. 5 μm FtsZ1 (lanes 2–5) or FtsZ2 (lanes 7–10) was incubated at room temperature (25 °C) for 3 or 30 min in the indicated buffer. Assembly was initiated by the addition of nucleotide. Supernatant (S) and pellet (P) fractions were obtained by centrifugation and analyzed by SDS-PAGE, Coomassie staining, and densitometry. Input, FtsZ1 (Z1, lanes 1) or FtsZ2 (Z2, lanes 6) in the starting reactions. The percentage of input protein in the pellet fraction ± S.E. (n = 4) (%P) is shown below the relevant lane. The buffers used were HMK (A), HMK-GTP (B), HMKCa-GDP (C), and HMKCa-GTP (D). FtsZ1 and FtsZ2 migrate at ∼40 and 45–46 kDa, respectively (20).
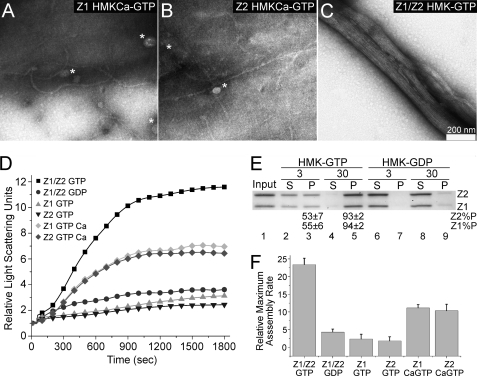
FIGURE 3.
Assembly of FtsZ1 and FtsZ2 separately and together. The reactions were performed at room temperature with 5 μm total protein. Assembly was initiated by the addition of nucleotide. A–C, electron micrographs of reaction aliquots taken after 10 min. A, FtsZ1 in HMKCa-GTP. B, FtsZ2 in HMKCa-GTP. C, FtsZ1 plus FtsZ2 (2.5 μm each) in HMK-GTP. D, time course of assembly of FtsZ1 in HMK-GTP (gray triangle) and HMKCa-GTP (gray diamond), FtsZ2 in HMK-GTP (inverted black triangle), and HMKCa-GTP (black diamond), and FtsZ1 plus FtsZ2 (2.5 μm each) in HMK-GTP (black square) or HMK-GDP (black circle) monitored by light scattering. E, coassembly of FtsZ1 and FtsZ2 (2.5 μm each) in HMK-GTP (lanes 2–5) or HMK-GDP (lanes 6–9) monitored by sedimentation as in Fig. 2. Input, FtsZ1 (Z1) or FtsZ2 (Z2) in the starting reactions (lane 1). The percentages of input FtsZ1 (Z1%P) and FtsZ2 (Z2%P) in the pellet fraction ± S.E. (n = 4) are shown below the relevant lanes. S, supernatant; P, pellet. F, relative maximum assembly rates for reactions in D estimated by taking the average of the maximum slopes from three independent light scattering traces.
The FtsZ1 and FtsZ2 filaments formed in HMKCa-GTP had similar morphologies (Fig. 3, A and B). Thin filaments ∼5–8 nm thick were usually observed, suggesting that they are single or double protofilaments based on the dimensions of the bacterial proteins and their similarity to the plant proteins (20, 25). Filaments were typically 600–800 nm in length, although filaments up to 1.5 μm were occasionally observed. Many filaments exhibited some curvature. The increased amount of protein in the pellet fractions in HMKCa-GTP and observation of filaments in these reactions showed that FtsZ1 and FtsZ2 are individually capable of protofilament formation when stabilized by Ca2+.
FtsZ1 and FtsZ2 Exhibit GTP-dependent Coassembly in the Absence of Calcium
In chloroplasts, FtsZ1 and FtsZ2 always colocalize, suggesting that they coassemble (24, 40). To address this possibility, we carried out sedimentation assays in which FtsZ1 and FtsZ2 were mixed 1:1 (2.5 μm each). In HMK containing 1 mm GDP (HMK-GDP), little FtsZ pelleted (Fig. 3E, lanes 6–9). In contrast, in 1 mm GTP (HMK-GTP) 53 and 94% of the total protein pelleted after 3 and 30 min, respectively (Fig, 3E, lanes 2–5). A similar degree of sedimentation occurred at 3 mm GTP (described below). EM revealed that the structures formed in HMK-GTP consisted mainly of large bundles of protofilaments (Fig. 3C and supplemental Fig. S3, A–D). These bundles were often ∼5 μm in length, sometimes longer. Less frequently bundles <3 μm in length were observed. Bundles varied in thickness from ∼30 to ∼140 nm (supplemental Fig. S3, A and B) and appeared to be composed largely of straight or slightly curved protofilaments. In some places thick bundles appeared to consist of several thinner bundles. The results described below show that the bundling behavior was not a consequence of the His tags. Notably, the formation of protofilaments and bundles in the mixed reactions occurred in the absence of Ca2+ as a stabilizing agent. These findings indicate that coassembly (polymerization and bundling) of FtsZ1 and FtsZ2 is strongly favored over assembly of the individual proteins.
To further investigate the time course of FtsZ1 and FtsZ2 coassembly, we monitored assembly continuously by 90° light scattering (LS) (35). Consistent with sedimentation assays (Figs. 2B and 3E), only a small increase in LS occurred when FtsZ1 and FtsZ2 were incubated either separately in HMK-GTP (Fig. 3D, triangles) or together in HMK-GDP (Fig. 3D, circles). The increase in LS was somewhat greater for the mixed proteins with GDP (Fig. 3D, circles) than for the single proteins with GTP (Fig. 3D, triangles), suggesting a minor degree of coassembly even in GDP. The addition of Ca2+ to the individual FtsZs resulted in an increased LS signal (Fig. 3D, diamonds), mirroring the increased sedimentation observed under these conditions (Fig. 2D). The largest increase in LS occurred for the mixed proteins in HMK-GTP (Fig. 3D, squares), consistent with the increased sedimentation and formation of protofilaments and bundles in these reactions (Fig. 3, C and E). The signal increase showed a slight initial lag, suggestive of cooperative assembly. Consistent with this observation, equimolar FtsZ1 and FtsZ2 exhibited a critical concentration (Cc) for coassembly of ∼0.75 μm (Fig. 4). The LS signal for the mixed proteins in HMK-GTP increased for ∼800 s and then began to plateau (Fig. 3D, squares, and supplemental Fig. S4, black triangles). Some variability between experiments was noted (Fig. 5, A and B, black squares). The maximum rate of assembly was higher for the coassembled proteins in GTP than in the other reactions (Fig. 3F). No decrease in LS was observed for the coassembled proteins over 2 h of monitoring. This behavior contrasts with that of EcFtsZ, which showed an increase in LS for ∼10 min in 1 mm GTP and ∼25 min in 5 mm GTP and then a slow decrease (supplemental Fig. S4), consistent with protofilament disassembly caused by GTP hydrolysis and depletion (35). These results indicate that coassembled FtsZ1 and FtsZ2 polymers are more stable than EcFtsZ polymers in vitro, possibly as a result of both their intrinsically lower GTP hydrolysis rates and their bundling activity.
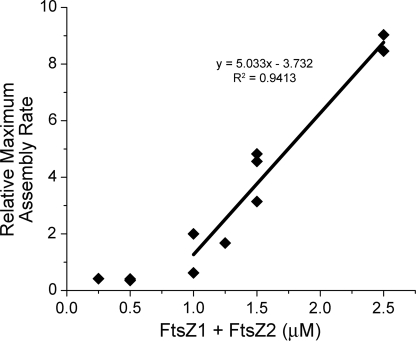
FIGURE 4.
Critical concentration for FtsZ1 and FtsZ2 coassembly. Coassembly of equimolar FtsZ1 and FtsZ2 in HMK-GTP at room temperature was monitored by light scattering at the indicated total FtsZ concentration. The relative maximum assembly rate for each reaction, estimated by taking the maximum slope of the corresponding light scattering trace, was plotted against protein concentration (35). All of the data points except those below 1.0 μm FtsZ, where assembly was not observed, were fit with the linear regression equation shown. The x intercept indicated a Cc of ∼0.75 μm for FtsZ1/FtsZ2 coassembly, similar to the Cc of ∼1 μm reported for EcFtsZ (10, 64).
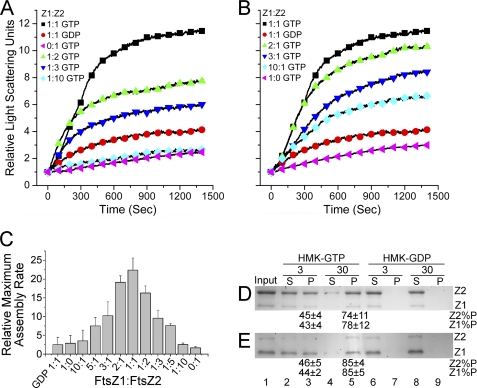
FIGURE 5.
Coassembly of FtsZ1 and FtsZ2 at various stoichiometries. All of the reactions were performed at room temperature with 5 μm total protein. Assembly was initiated by the addition of nucleotide. A and B, time course of coassembly of FtsZ1 (Z1) and FtsZ2 (Z2) at the indicated molar ratio monitored by light scattering. The experiment was repeated three times; data from a single experiment are shown. The control reactions (red circles) were in HMK-GDP; all others were in HMK-GTP. A, reactions with excess FtsZ2. B, reactions with excess FtsZ1. C, relative maximum assembly rates ± S.E. (n = 3) derived by averaging the maximum slopes of the light scattering traces from three experiments. D and E, coassembly of 1:5 (D) or 5:1 (E) FtsZ1:FtsZ2 in HMK-GTP (lanes 2–5) or HMK-GDP (lanes 6–9) monitored by sedimentation as described in the legend for Fig. 2. Input, FtsZ1 (Z1) and FtsZ2 (Z2) in the starting reactions (lanes 1). Percentages of input FtsZ1 (Z1 %P) and FtsZ2 (Z2 %P) in the pellet fraction ± S.E. (n = 4) are shown below the relevant lanes. S, supernatant; P, pellet.
Maximum FtsZ1 and FtsZ2 Coassembly Occurs at a 1:1 Stoichiometry
We also investigated Ca2+-independent FtsZ assembly at different FtsZ1:FtsZ2 ratios (5 μm total protein). In LS assays, the extent of assembly was highest when the proteins were mixed 1:1 in HMK-GTP (Fig. 5, A and B, black squares). Excess FtsZ2 decreased the LS signals (Fig. 5A), indicating reduced polymerization and/or bundling. Similar results were obtained for excess FtsZ1 (Fig. 5B). Maximum rates of assembly followed the same pattern (Fig. 5C), with the maximum rates observed at equimolar FtsZ1:FtsZ2.
We further quantified the effects of unequal molar ratios on coassembly using sedimentation assays. In reactions containing 1:5 FtsZ1:FtsZ2 (5 μm total FtsZ), ∼44 and 76% of the protein sedimented after 3 and 30 min, respectively (Fig. 5D, lanes 3 and 5). When the ratio was reversed, the percentages were ∼45 and 85% (Fig. 5E). In both cases, coassembly was GTP-dependent (Fig. 5, D and E, lanes 6–9). Although the extent of coassembly was somewhat less than in equimolar reactions (Fig. 3E, lanes 3 and 5), significant protofilament formation and bundling were still observed (supplemental Fig. S3, E–G). Interestingly, in all of the mixing experiments, the FtsZ1:FtsZ2 ratio in the pellet fractions was approximately the same as the input ratio (e.g. compare lane 1 to lane 5 in Figs. 3E and 5, D and E), suggesting plasticity in the composition of coassembled structures. In addition, in nonequimolar reactions, the extent of assembly was consistently higher in reactions containing more FtsZ1 than FtsZ2 at a given ratio (e.g. Fig. 5, A and B, 2:1 versus 1:2 FtsZ1:FtsZ2; Fig. 5, D and E, lanes 5), suggesting that FtsZ1 enhances coassembly to a greater extent than FtsZ2.
FtsZ1 and FtsZ2 Copolymerize
In bacterial FtsZ, interaction of the T7 loop of one subunit with GTP bound to an adjacent subunit completes the GTPase active site and catalyzes hydrolysis (9, 33, 41). T7 loop mutants lack GTPase activity but still undergo GTP-dependent assembly (9, 33, 42). We introduced mutations into the predicted T7 loop regions of FtsZ1 and FtsZ2 (15, 25) to create FtsZ1D275A and FtsZ2D322A, respectively, which correspond to the EcFtsZ mutants D212A and D212G (9, 33, 42), and investigated their biochemical properties. GTPase activities of FtsZ1D275A and FtsZ2D322A were <10% of the activities of the corresponding nonmutant (hereafter called WT) proteins (Fig. 6B, compare last to first set of bars), similar to the background control (supplemental Fig. S1C), showing that GTP hydrolysis by FtsZ1 and FtsZ2 depends on an intact T7 loop. We recently identified missense alleles of Arabidopsis FtsZ1 (25) and FtsZ2 (supplemental Fig. S5) that bear mutations in their predicted T7 loops and cause dominant-negative defects in chloroplast division, highlighting the importance of these regions for in vivo function. However, although FtsZ1D275A and FtsZ2D322A were GTPase-deficient, they bound 1.15 ± 0.25 and 0.95 ± 0.12 GTP/FtsZ, respectively, indicating that GTP binding was unaffected by the mutations. Consistent with this finding, the individual mutant proteins formed Ca2+-stabilized protofilaments (supplemental Fig. S6, A and B) similar to those formed by the WT proteins (Fig. 3, A and B). Further, when mixed 1:1, the mutant proteins coassembled into protofilament bundles in the absence of Ca2+ (Fig. 6A). These data show that, similar to bacterial FtsZ (9, 10, 42), assembly of the chloroplast FtsZs does not require GTP hydrolysis.
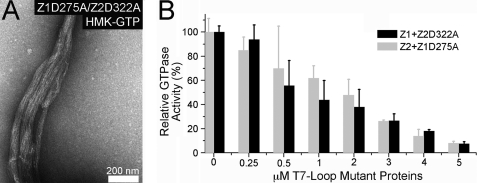
FIGURE 6.
Coassembly of FtsZ1 and FtsZ2 T7 loop mutants and effect on GTPase activity of the wild type proteins. A, electron micrograph of FtsZ1D275A and FtsZ2D322A (2.5 μm each) coassembled HMK-GTP at room temperature. B, GTPase activities of FtsZ1 assayed at 25 °C in the presence of increasing FtsZ2D322A (Z1+Z2D322A, black bars) and FtsZ2 assayed in the presence of increasing FtsZ1D275A (Z2+Z1D275A, gray bars). The total protein concentration was kept at 5 μm, and each assay was performed three times.
The slight stimulation in GTPase activity when FtsZ1 and FtsZ2 are mixed 1:1 at 3 mm GTP or above (Figs. 1 and 7B) suggests they associate to form an active site. To address this possibility more directly, we asked whether FtsZ1D275A and FtsZ2D322A would act as competitive inhibitors of the WT proteins in GTPase assays containing different stoichiometric ratios of WT and mutant FtsZ (5 μm total FtsZ). Experiments were performed in 3 mm GTP where the WT proteins showed near maximum activity (Fig. 1). When FtsZ1 was incubated with increasing amounts of FtsZ2D322A, its GTPase activity decreased proportionately (Fig. 6B, black bars). Similar results were obtained when FtsZ2 was incubated with increasing amounts of FtsZ1D275A (Fig. 6B, gray bars). In the reactions containing 1:1 FtsZ1D275A:FtsZ2 and 1:1 FtsZ2D322A:FtsZ1, protofilament bundles were observed (supplemental Fig. S6, C and D). The decreased GTPase activity in mutant-containing reactions was not due to the reduced concentration of WT protein because, in control assays containing only WT protein in 3 mm GTP, GTPase activities did not vary over the range of WT protein concentrations used in the mixing experiments (supplemental Fig. S7). These findings indicate that GTPase-deficient FtsZ1 substoichiometrically inhibits the GTPase activity of FtsZ2 and vice versa, providing strong evidence that the GTPase active site in the mixed reactions is completed by interaction between FtsZ1 and FtsZ2 subunits. Because the active site in FtsZ proteins is oriented along the longitudinal axis of a protofilament (9, 41), these results also indicate that FtsZ1 and FtsZ2 form heteropolymers.
His Tags Do Not Affect the in Vitro Behavior of FtsZ1 and FtsZ2
To ensure that the enzymatic and assembly properties of FtsZ1 and FtsZ2 described above are not artifacts of their His tags, we expressed untagged forms of the proteins in E. coli and purified them from inclusion bodies by assembly-based purification (Fig. 7A, lanes 6 and 12). Assembly-purified proteins were solubilized in urea and then refolded and further purified by size exclusion chromatography, after which they were >95% pure.
GTPase activities of the individual and mixed untagged and His-tagged proteins (5 μm total FtsZ) were compared at 1 and 3 mm GTP. In all reactions, activities of the tagged and untagged proteins were similar (Fig. 7B). Further, at 3 mm GTP, equimolar mixtures of untagged FtsZ1 and FtsZ2 exhibited slightly enhanced GTPase activity, similar to the tagged proteins (Fig. 7B), consistent with their coassembly.
For bacterial FtsZ, His tags have been reported to induce formation of bundles and sheets under some conditions (33, 43). To test whether the bundling behavior of coassembled FtsZ1 and FtsZ2 was due to their His tags, we used sedimentation assays and EM to compare the coassembly properties of the untagged and His-tagged proteins. Similar to the tagged proteins, ∼90% of the untagged FtsZ1 and FtsZ2 (2.5 μm each) pelleted after 30 min in both 1 and 3 mm GTP, and pelleting was GTP-dependent (Fig. 7D). EM showed the coassembled untagged proteins in large protofilament bundles (Fig. 7C) resembling those formed by the coassembled His-tagged proteins (Fig. 3C and supplemental Fig. S3, A and B). We conclude that the His tags do not alter the in vitro behavior of FtsZ1 and FtsZ2 under the conditions reported here and that bundling is an intrinsic property of FtsZ1/FtsZ2 coassembly.
DISCUSSION
FtsZ1 and FtsZ2 arose via post-endosymbiotic gene duplication in a common ancestor of green algae and land plants and remain as distinct families throughout these lineages (44, 45). FtsZ2 is more ancestral based on phylogenetic analysis and retention of the C-terminal core motif common to most bacterial FtsZs, and FtsZ1 and FtsZ2 interact with different partner proteins (26, 44–46). However, both proteins share >45% amino acid identity with many cyanobacterial FtsZs and >35% identity with FtsZs in other bacteria (18). Consistent with the degree of conservation, FtsZ1 and FtsZ2 are typical prokaryotic FtsZs in many respects. They each bind one GTP/monomer, undergo GTP-dependent assembly into thin filaments in the presence of Ca2+, and exhibit GTPase activities that are dependent on GTP concentration. Although their maximum GTPase activities are lower than those reported for E. coli and other well studied bacterial proteins, they are in the range of activities reported for Mycobacterium tuberculosis and Haloferax volcanii FtsZ (47, 48).
Although FtsZ1 and FtsZ2 are capable of assembling individually, in vivo they always colocalize and function together in chloroplast division (24, 40). Whether their colocalization reflects copolymerization or close proximity of separately assembled protofilaments has not been clear (15). Our in vitro analyses show that FtsZ1 and FtsZ2 do coassemble and that coassembly is strongly favored over self-assembly even when their ratios in assembly reactions are highly skewed (Fig. 5). Coassembly involves two distinct activities. One is the formation of heteropolymers. This was revealed by our finding that FtsZ1 T7 loop mutants act as substoichiometric inhibitors of WT FtsZ2 GTPase activity and vice versa (Fig. 6), indicating that FtsZ1 and FtsZ2 are capable of completing their active sites in trans (9). This is the first time that heteropolymerization of FtsZ1 and FtsZ2 has been demonstrated. The other component of coassembly is protofilament bundling. Bacterial FtsZ protofilaments also bundle in vitro, but usually only in the presence of cationic stabilizers (7, 8, 30, 49–51). In contrast, the extensive bundling observed upon FtsZ1/FtsZ2 coassembly occurs in the absence of such stabilizers and appears to be stimulated by heteropolymerization. Although EM suggests that bundling may be most pronounced in equimolar reactions, consistent with the maximum degree of sedimentation and light scattering observed under these conditions, significant bundling was also observed at different stoichiometric ratios, at lower FtsZ concentrations (1.25 μm each FtsZ1 and FtsZ2) (supplemental Fig. S8A), at the physiological pH of the stroma in the light (pH 8) (supplemental Fig. S8, B and C), and in reactions containing both WT and T7 loop mutant proteins (supplemental Fig. S6). Thus FtsZ1/FtsZ2 heteropolymers have a strong tendency to bundle under a variety of conditions, indicating that this is an intrinsic property of their coassembly.
Because bundling stabilizes bacterial FtsZ protofilaments (35, 49–51), bundling of coassembled plant FtsZ could explain their enhanced stability relative to that of EcFtsZ (supplemental Fig. S4). In addition, the slow GTPase activity of the plant proteins may also contribute to polymer stability because GTP hydrolysis is thought to destabilize FtsZ protofilaments (10, 47, 52), although the exact relationship between GTP hydrolysis and FtsZ dynamics is not entirely clear (12, 14, 53). However, because GTPase activity was not reduced, and in fact was slightly increased, in reactions in which coassembly and bundling were observed (Figs. 1A and 7, B–D), the enhanced assembly in the mixed reactions may not be due to a bundling-associated decrease in GTPase activity, as has been proposed for bundling of EcFtsZ (35, 49, 51). The primary effect of bundling could rather be to reduce the rate of monomer dissociation. Interestingly, the greater stability of the assembled plant proteins is consistent with the presence of mid-plastid Z rings in nearly all plastids, including those in mature leaf cells in which chloroplast division has slowed or ceased (24). This suggests that plastid FtsZ filaments may be inherently more stable than bacterial filaments in vivo as well as in vitro. This would further imply that chloroplast Z ring dynamics should be regulated in part by factors that inhibit protofilament bundling. One candidate for such a factor is the chloroplast division protein ARC3, a proposed functional replacement for bacterial MinC (46, 54), which was recently shown to inhibit lateral interactions between EcFtsZ protofilaments assembled in vitro (55).
GTP-dependent protofilament formation has been demonstrated previously for Medicago truncatula and Nicotiana tabacum FtsZ1 (21, 22), but Nicotiana FtsZ2 did not assemble (21), possibly because the recombinant FtsZ2 protein contained its transit peptide. A more recent study by Smith et al. (23) using Arabidopsis FtsZ1 and FtsZ2 bearing both c-Myc and His tags on their C termini indicated that both FtsZ1 and FtsZ2 are capable of GTP-dependent protofilament formation. However, the results of the latter study differed in many respects from those reported here. GTPase activities in the two studies are difficult to compare directly because Smith et al. did not report the GTP concentration used in their assays, and the protein concentrations given in their figure legends and supplemental methods are inconsistent. However, they reported that mixing FtsZ1 and FtsZ2 in equimolar reactions inhibited GTPase activity, whereas we observed a slight stimulation at 3 mm GTP and above. In assembly assays, Smith et al. observed single protofilaments for individual and mixed FtsZ1 and FtsZ2, as reported for bacterial FtsZ (4, 5), but indicated that the filament morphologies were unlike any observed for the bacterial proteins (23). In contrast, the morphologies of our assemblies appear more like those seen for bacterial FtsZs (e.g. Refs. 30, 35, and 47). Further, Smith et al. did not observe protofilament bundling when FtsZ1 and FtsZ2 were mixed, whereas we observed extensive bundling under a variety of conditions. Finally, Smith et al. concluded that polymerization of FtsZ1 and FtsZ2 requires GTP hydrolysis based on a lack of assembly in the presence a nonhydrolozyable GTP analog, but we found that FtsZ1 and FtsZ2 mutants lacking GTPase activity still exhibit GTP-dependent assembly, consistent with data showing that assembly of bacterial FtsZ, although GTP-dependent, does not require GTP hydrolysis (9, 33, 43, 56). These differences may be due in part to differences in assay conditions or C-terminal tags, but another explanation could be dissimilarities in the lengths of the recombinant proteins used in the two studies, which were truncated to remove the predicted chloroplast transit peptides. We estimated the FtsZ1 and FtsZ2 transit peptide lengths at 57 and 48 amino acids, respectively, based on a combination of sequence alignments, targeting predictions, and molecular mass data (supplemental Methods). According to the primer sequences reported by Smith et al. in their supplemental methods, their proteins were truncated after amino acids 39 and 73 for FtsZ1 and FtsZ2, respectively, which would have left a portion of the predicted transit peptide on FtsZ1 and removed conserved residues from the probable mature form of FtsZ2. Further experiments will be required to understand why our findings differ from those of Smith et al. (23).
FtsZ proteins are widely believed to be the evolutionary progenitors of eukaryotic α- and β-tubulin (7, 8, 57), and the discovery of two FtsZ types in chloroplasts raised the question of whether they might have a similar functional relationship (15, 25). The demonstration that FtsZ1 and FtsZ2 are capable of polymerizing independently of one another (Refs. 21–23 and this study) indicated that this was probably not the case. Nevertheless, we have shown that FtsZ1 and FtsZ2 form heteropolymers and that their coassembly is favored at a 1:1 stoichiometry. However, we also found in sedimentation assays that the FtsZ1:FtsZ2 ratio in the pellet fraction reflects their input ratio. This behavior contrasts with that of BtubA and BtubB, two tubulin-like proteins recently identified in the bacterium Prosthecobacter dejongeii (58, 59). Unlike FtsZ1 and FtsZ2, in sedimentation assays BtubA and BtubB always pellet in a 1:1 stoichiometry regardless of input ratio and only polymerize when both proteins are present. Recent data have shown that BtubA/B polymers assemble from heterodimers, similar to αβ-tubulin (60). Thus, although FtsZ1 and FtsZ2 copolymerize, they do not behave like BtubA/B or αβ-tubulin.
Although FtsZ1 and FtsZ2 coassemble with variable stoichiometry in mixed reactions, we do not yet know the molecular makeup of the assemblies that pellet at different input ratios. One possibility is that the bundles that form in skewed reaction mixtures (supplemental Fig. S3, E–G) are composed of protofilaments in which FtsZ1 and FtsZ2 heteropolymerize at the input stoichiometry. This would suggest that interaction between FtsZ1 and FtsZ2, although kinetically favored (Fig. 5C), is dynamic, and their dissociation could allow polymers of variable stoichiometry to form. Another possibility is that the FtsZ1:FtsZ2 stoichiometry in bundled protofilaments is actually 1:1 regardless of input ratio and that the excess FtsZ1 or FtsZ2 in the input reaction assembles into thin homopolymers that still come down in the pellets. However, in reactions containing only FtsZ1 or FtsZ2, thin protofilaments form and pellet only in the presence of Ca2+ (Figs. 2, B and D, and 3, A and B). Therefore, this latter scenario would imply that assembly of 1:1 heteropolymers in the skewed reaction mixtures, which lack Ca2+, could facilitate or nucleate the assembly of excess FtsZ into thin homopolymers. Defining the composition of protofilaments and bundles that form at various FtsZ1/FtsZ2 input ratios will be important for understanding their coassembly properties.
Interestingly, the apparent plasticity of FtsZ coassembly in vitro is consistent with genetic studies showing that chloroplast FtsZ filaments assemble under a wide range of in vivo stoichiometries, although significant deviations from the wild type stoichiometry alters FtsZ filament morphology and perturbs plastid division (17, 19, 20, 24, 61). Nevertheless, the ability of FtsZ1 and FtsZ2 to coassemble at different stoichiometries raises the intriguing possibility that the degree of heteropolymerization and/or bundling may be regulated in vivo as a means of regulating FtsZ filament morphology and hence Z ring constriction and dynamics. Such regulation may be necessary because plants have multiple plastid types that vary greatly in size and division activity; perhaps the relative levels of FtsZ1 and FtsZ2 vary in different plastid types or at different stages of development (61). In this context, it is interesting to note that complete FtsZ2 rings are occasionally observed in small plastids of ftsZ1 null mutants (25), but the converse is not true (19). This suggests that even though both proteins are capable of protofilament formation in vitro, FtsZ1 may have evolved as a bundling factor rather than as a distinct ring-forming polymer.
The in vitro behavior of FtsZ1 and FtsZ2 begins to suggest potential mechanistic explanations for the evolution of two FtsZ families in plants. One possibility is that longer or stronger protofilaments resulting from heteropolymer-induced bundling provide additional force for the division of organelles that are generally larger in diameter than bacteria, although the role of FtsZ bundling in force generation remains unclear (11–13, 62, 63). Another is that two FtsZ types allow for dynamic regulation of protofilament composition and degree of bundling during constriction, perhaps permitting the significant change in membrane curvature involved in constricting a large organelle. Further biochemical studies of plant FtsZ1 and FtsZ2 will shed light on these issues and may yield new insights into the mechanisms of FtsZ function in bacteria as well.
Supplementary Material
Acknowledgments
We thank Alicia Pastor for EM support; Jonathan Glynn, Aaron Schmitz, and Yue Yang for helpful discussions and comments on the manuscript; Jonathan Glynn and John Sherbeck for identification of the CAN3 allele of AtFtsZ2-1; and Harold Erickson for the gift of the EcFtsZ expression plasmid and invaluable suggestions.
This work was supported by Michigan State University Plant Sciences and N. E. Tolbert Memorial Fellowships (to B. J. S. C. O.) and by National Science Foundation Grant 0544676 (to K. W. O.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S8 and Methods.
- EM
- electron microscopy
- LS
- light scattering
- WT
- wild type.
REFERENCES
- 1.Romberg L., Levin P. A. (2003) Annu. Rev. Microbiol. 57, 125–154 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Michie K. A., Löwe J. (2006) Annu. Rev. Biochem. 75, 467–492 [DOI] [PubMed] [Google Scholar]
- 3.Adams D. W., Errington J. (2009) Nat. Rev. Microbiol. 7, 642–653 [DOI] [PubMed] [Google Scholar]
- 4.Mukherjee A., Lutkenhaus J. (1994) J. Bacteriol. 176, 2754–2758 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Huecas S., Llorca O., Boskovic J., Martín-Benito J., Valpuesta J. M., Andreu J. M. (2008) Biophys. J. 94, 1796–1806 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Popp D., Iwasa M., Narita A., Erickson H. P., Maéda Y. (2009) Biopolymers 91, 340–350 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Löwe J., Amos L. A. (1999) EMBO J. 18, 2364–2371 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Erickson H. P., Taylor D. W., Taylor K. A., Bramhill D. (1996) Proc. Natl. Acad. Sci. U.S.A. 93, 519–523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Scheffers D. J., de Wit J. G., den Blaauwen T., Driessen A. J. (2002) Biochemistry 41, 521–529 [DOI] [PubMed] [Google Scholar]
- 10.Mukherjee A., Lutkenhaus J. (1998) EMBO J. 17, 462–469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Li Z., Trimble M. J., Brun Y. V., Jensen G. J. (2007) EMBO J. 26, 4694–4708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Erickson H. P. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 9238–9243 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Osawa M., Anderson D. E., Erickson H. P. (2008) Science 320, 792–794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Osawa M., Anderson D. E., Erickson H. P. (2009) EMBO J. 28, 3476–3484 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Osteryoung K. W., McAndrew R. S. (2001) Annu. Rev. Plant Physiol. Plant Mol. Biol. 52, 315–333 [DOI] [PubMed] [Google Scholar]
- 16.Yang Y., Glynn J. M., Olson B. J., Schmitz A. J., Osteryoung K. W. (2008) Curr. Opin. Plant Biol. 11, 577–584 [DOI] [PubMed] [Google Scholar]
- 17.Strepp R., Scholz S., Kruse S., Speth V., Reski R. (1998) Proc. Natl. Acad. Sci. U.S.A. 95, 4368–4373 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Osteryoung K. W., Stokes K. D., Rutherford S. M., Percival A. L., Lee W. Y. (1998) Plant Cell 10, 1991–2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Schmitz A. J., Glynn J. M., Olson B. J., Stokes K. D., Osteryoung K. W. (2009) Mol. Plant 2, 1211–1222 [DOI] [PubMed] [Google Scholar]
- 20.McAndrew R. S., Olson B. J., Kadirjan-Kalbach D. K., Chi-Ham C. L., Vitha S., Froehlich J. E., Osteryoung K. W. (2008) Biochem. J. 412, 367–378 [DOI] [PubMed] [Google Scholar]
- 21.El-Kafafi el-S., Mukherjee S., El-Shami M., Putaux J. L., Block M. A., Pignot-Paintrand I., Lerbs-Mache S., Falconet D. (2005) Biochem. J. 387, 669–676 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lohse S., Hause B., Hause G., Fester T. (2006) Plant Cell Physiol 47, 1124–1134 [DOI] [PubMed] [Google Scholar]
- 23.Smith A. G., Johnson C. B., Vitha S., Holzenburg A. (2010) FEBS Lett. 584, 166–172 [DOI] [PubMed] [Google Scholar]
- 24.Vitha S., McAndrew R. S., Osteryoung K. W. (2001) J. Cell Biol. 153, 111–120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yoder D. W., Kadirjan-Kalbach D., Olson B. J., Miyagishima S. Y., Deblasio S. L., Hangarter R. P., Osteryoung K. W. (2007) Plant Cell Physiol. 48, 775–791 [DOI] [PubMed] [Google Scholar]
- 26.Maple J., Aldridge C., Møller S. G. (2005) Plant J. 43, 811–823 [DOI] [PubMed] [Google Scholar]
- 27.Stokes K. D., McAndrew R. S., Figueroa R., Vitha S., Osteryoung K. W. (2000) Plant Physiol. 124, 1668–1677 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bendezú F. O., Hale C. A., Bernhardt T. G., de Boer P. A. (2009) EMBO J. 28, 193–204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Jeong K. J., Lee S. Y. (2003) Appl. Environ. Microbiol. 69, 1295–1298 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lu C., Stricker J., Erickson H. P. (1998) Cell Motil. Cytoskeleton 40, 71–86 [DOI] [PubMed] [Google Scholar]
- 31.Lu C., Erickson H. P. (1998) Methods Enzymol. 298, 305–313 [DOI] [PubMed] [Google Scholar]
- 32.Fisher C. L., Pei G. K. (1997) BioTechniques 23, 570–574 [DOI] [PubMed] [Google Scholar]
- 33.Redick S. D., Stricker J., Briscoe G., Erickson H. P. (2005) J. Bacteriol. 187, 2727–2736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Geladopoulos T. P., Sotiroudis T. G., Evangelopoulos A. E. (1991) Anal. Biochem. 192, 112–116 [DOI] [PubMed] [Google Scholar]
- 35.Mukherjee A., Lutkenhaus J. (1999) J. Bacteriol. 181, 823–832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.McAndrew R. S., Froehlich J. E., Vitha S., Stokes K. D., Osteryoung K. W. (2001) Plant Physiol. 127, 1656–1666 [PMC free article] [PubMed] [Google Scholar]
- 37.Osteryoung K. W., Vierling E. (1995) Nature 376, 473–474 [DOI] [PubMed] [Google Scholar]
- 38.de Boer P., Crossley R., Rothfield L. (1992) Nature 359, 254–256 [DOI] [PubMed] [Google Scholar]
- 39.RayChaudhuri D., Park J. T. (1992) Nature 359, 251–254 [DOI] [PubMed] [Google Scholar]
- 40.Vitha S., Froehlich J. E., Koksharova O., Pyke K. A., van Erp H., Osteryoung K. W. (2003) Plant Cell 15, 1918–1933 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Löwe J., Amos L. A. (1998) Nature 391, 203–206 [DOI] [PubMed] [Google Scholar]
- 42.Mukherjee A., Saez C., Lutkenhaus J. (2001) J. Bacteriol. 183, 7190–7197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Oliva M. A., Huecas S., Palacios J. M., Martín-Benito J., Valpuesta J. M., Andreu J. M. (2003) J. Biol. Chem. 278, 33562–33570 [DOI] [PubMed] [Google Scholar]
- 44.Stokes K. D., Osteryoung K. W. (2003) Gene 320, 97–108 [DOI] [PubMed] [Google Scholar]
- 45.Rensing S. A., Kiessling J., Reski R., Decker E. L. (2004) J. Mol. Evol. 58, 154–162 [DOI] [PubMed] [Google Scholar]
- 46.Maple J., Vojta L., Soll J., Møller S. G. (2007) EMBO Rep. 8, 293–299 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Chen Y., Anderson D. E., Rajagopalan M., Erickson H. P. (2007) J. Biol. Chem. 282, 27736–27743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang X., Lutkenhaus J. (1996) Mol. Microbiol. 21, 313–319 [DOI] [PubMed] [Google Scholar]
- 49.Yu X. C., Margolin W. (1997) EMBO J. 16, 5455–5463 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lu C., Reedy M., Erickson H. P. (2000) J. Bacteriol. 182, 164–170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Marrington R., Small E., Rodger A., Dafforn T. R., Addinall S. G. (2004) J. Biol. Chem. 279, 48821–48829 [DOI] [PubMed] [Google Scholar]
- 52.Huecas S., Andreu J. M. (2004) FEBS Lett. 569, 43–48 [DOI] [PubMed] [Google Scholar]
- 53.Scheffers D. J., den Blaauwen T., Driessen A. J. (2000) Mol. Microbiol. 35, 1211–1219 [DOI] [PubMed] [Google Scholar]
- 54.Glynn J. M., Miyagishima S. Y., Yoder D. W., Osteryoung K. W., Vitha S. (2007) Traffic 8, 451–461 [DOI] [PubMed] [Google Scholar]
- 55.Dajkovic A., Lan G., Sun S. X., Wirtz D., Lutkenhaus J. (2008) Curr. Biol. 18, 235–244 [DOI] [PubMed] [Google Scholar]
- 56.Mohammadi T., Ploeger G. E., Verheul J., Comvalius A. D., Martos A., Alfonso C., van Marle J., Rivas G., den Blaauwen T. (2009) Biochemistry 48, 11056–11066 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Erickson H. P. (2007) Bioessays 29, 668–677 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Schlieper D., Oliva M. A., Andreu J. M., Löwe J. (2005) Proc. Natl. Acad. Sci. U.S.A. 102, 9170–9175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Sontag C. A., Staley J. T., Erickson H. P. (2005) J. Cell Biol. 169, 233–238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Sontag C. A., Sage H., Erickson H. P. (2009) PLoS ONE 4, e7253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.El-Kafafi el-S., Karamoko M., Pignot-Paintrand I., Grunwald D., Mandaron P., Lerbs-Mache S., Falconet D. (2008) Biochem. J. 409, 87–94 [DOI] [PubMed] [Google Scholar]
- 62.Allard J. F., Cytrynbaum E. N. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 145–150 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Lan G., Daniels B. R., Dobrowsky T. M., Wirtz D., Sun S. X. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 121–126 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Chen Y., Bjornson K., Redick S. D., Erickson H. P. (2005) Biophys. J. 88, 505–514 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.