Abstract
Summary: ViewDock TDW is a modification of the pre-existing ViewDock Chimera extension (http://www.cgl.ucsf.edu/chimera/) used to visualize results of virtual screening experiments. By combing TDW hardware and an enhanced ViewDock interface, dozens of ligand–protein complexes are rendered simultaneously to parallelize the analysis of candidate ligands. The ViewDock TDW GUI allows the user to easily and interactively manipulate the molecules on the TDW as an entire set, a selected subset or a single ligand–protein complex and preserves all Chimera functionality.
Availability and Implementation: ViewDock TDW is an open source software; freely available on the web at http://www.tdw-prime.webs.com. Chimera UCSF is also available, free of charge, at http://www.cgl.ucsf.edu/chimera/
Contact: jhaga@bioeng.ucsd.edu
1 INTRODUCTION
As an alternative to costly high-throughput empirical screening of millions of compounds, virtual screening has led to tremendous success in drug discovery (Kitchen et al., 2004) by computationally identifying compounds with higher probability of strong binding affinity to the target protein. These data are used to make ‘informed’ decisions in constructing a shorter list of compounds to purchase and test in the lab, significantly reducing the cost and time to discovery.
Growing databases of small molecules (Baykoucheva, 2007; Irwin et al., 2006) and rigorous virtual screening methods demand greater computational resources. High performance computing solutions, such as Grid computing (Levesque et al., 2008), have proved to be useful tools but visual analysis of resulting data remains as a bottleneck in the workflow. Visualizing hundreds to thousands of ligand–protein complexes one at a time, limits a researcher's ability to compare and contrast candidate compounds. Identifying details of chemical binding poses, recurring or unique chemical moieties, and false positives are more likely when multiple structures are analyzed simultaneously, whether this be two side-by-side or more than 20 structures.
A Tiled Display Wall (TDW) is a computing cluster with one or more displays (tiles) connected to each computer (node) of the cluster. They are typically dedicated for visualization of large datasets in the form of high-resolution images (Wei et al., 2000). Although the use of TDW clusters in drug discovery workflows is limited, their configuration can vary in size and can be as simple as two networked computers, which makes it an appealing technology for high-throughput virtual screening.
ViewDock TDW was designed for a hardware setup consisting of one node as the control node with its own display, keyboard and mouse, and the render nodes running instances of Chimera (Petterson et al., 2004) on the TDW. From the control node, files of ligand–protein structure data are selected and Chimera commands are sent to the render nodes. This setup allows the user to compare and visualize dozens of ligand–receptor interactions in parallel, thus providing a means to analyze virtual screening results more quickly, thoroughly, and effectively.
2 VIEWDOCK TDW DESCRIPTION
ViewDock TDW controls Chimera instances running on the render nodes by taking advantage of the pre-existing function of accepting information and instructions from hyperlinks in a web browser, referred to as Chimera web data. In ViewDock TDW, these web data are in the form of SSH (Secure SHell) remote calls sending Chimera commands that are interpreted by instances of Chimera on the render nodes. ‘Tile’ objects were designed in ViewDock TDW's python code to allow the control node to manage display tiles and their associated structures. A ‘tile’ object contains all the information regarding a particular instance of Chimera including the node running the instance and the port number used to send commands to specified molecules. The use of common scripting languages and command line tools driving the functionality of ViewDock TDW makes it compatible with most existing Unix-based TDWs.
ViewDock TDW scales to any number of tiles and allows the user to specify the position and number of instances of Chimera displayed on each tile. With resulting ligand binding conformation data from the molecular docking software DOCK (Lang et al., 2008), the user can compare the ligand–protein interactions, number of hydrogen bonds and calculated energy scores from DOCK all in parallel. Given that common TDW configurations contain 16–20 tiles, ViewDock TDW allows the user to view and control 32–40 molecules simultaneously. Surface generations or hydrogen-bond calculations can be time-consuming on a single computer for multiple structures. ViewDock TDW performs these operations in parallel on the render nodes significantly reducing required CPU time.
3 EXAMPLE USAGE
The visualization step of the virtual screening workflow is the final review of the in silico molecular docking and scoring results data. While scoring algorithms based on binding energy functions provide a method to rank the ligands from most to least likely hits, visually examining the predicted conformations of the ligands bound to the target protein is necessary for selecting a set of ligands to purchase and test empirically in the lab. After starting ViewDock TDW on the control node for the first time, the user is asked to define the TDW dimensions and hostnames of each tile, which are saved for future sessions. The user then selects the file containing multiple ligand structures, in their target protein-binding conformations, and Chimera instances open on the render nodes with the first set of ligands. Chimera automatically adjusts to full display resolution on the render nodes and splits for one or more instances per display.
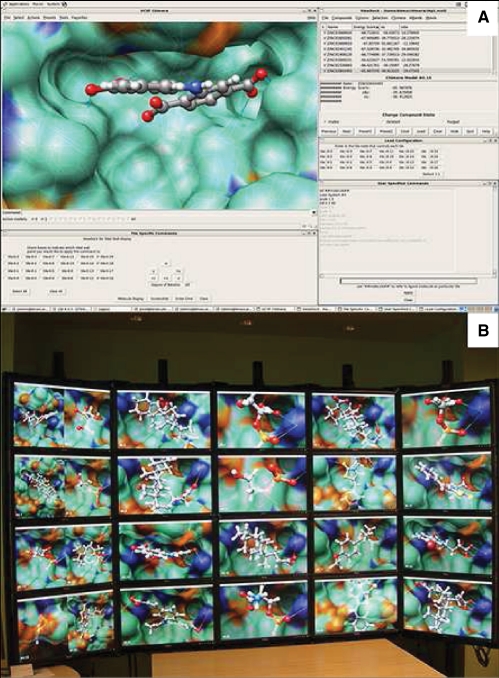
Figure 1A is a screenshot of the ViewDock TDW GUI on the control node. The candidate ligands' compound IDs, calculated binding scores and categories to group ligands are controlled in the main ViewDock TDW GUI. All communication to the TDW is facilitated through the ViewDock TDW GUI via keyboard commands and predefined buttons. In the ‘Tile Specific Commands’ window, the user selects structures of interest and subsequent commands are sent only to the selected molecules. Structure rotation, viewing angle manipulation and preset display operations were mapped to hotkeys and GUI button elements. For more advanced control, user-specified Chimera commands can be sent to render nodes using the ‘Cmd’ window and the selected structure shown on the control node display can be manipulated using the mouse just as with Chimera on its own.
Fig. 1.
(A) Screenshot of ViewDock TDW GUI. Specific molecules are selected using the checkboxes on the bottom left of screenshot. The bottom right window demonstrates the ability to send commands via command line to the molecules in Chimera. (B) Picture of a 4 × 5 Tiled Wall Display. Twenty-four different candidate ligand–protein complexes are rendered simultaneously. The leftmost column of the TDW has two Chimera instances per display.
Figure 1B shows a TDW running ViewDock TDW to review candidate ligands bound to the catalytic domain of the protein tyrosine phosphatase SHP-1 [PDB ID: 1FPR]. Using the ‘User Specified Commands’ window, surface transparency was changed and hydrogen bond thickness was increased. The ‘Next’ and ‘Prev’ buttons are used to navigate through the entire list of candidate compounds by loading the next or previous set of structures on the TDW. While proceeding through results, compounds can be tagged as ‘Viable’, ‘Deleted’ or ‘Purged’ and only compounds with the specified tag(s) appear on the TDW. Two instances of Chimera are shown per monitor on the left-most column of the TDW, demonstrating the scalability of ViewDock TDW. This scalability also allows for its use with a minimal setup of a two-computer network. We have built and tested this scenario where up to six instances of Chimera were run without noticeably impaired performance.
4 CONCLUSIONS
The manual, visual analysis step of the virtual screening-based, drug discovery workflow is enhanced by ViewDock TDW. ViewDock TDW scales to any display configuration, is easily installed, and gives the user global to fine-grained control over the rendered structures. Being able to compare, group, analyze and manipulate multiple candidate ligands simultaneously gives the researcher a better tool to effectively choose compounds to purchase for subsequent validation studies. This increases the efficacy and decreases the time involved in drug discovery.
ViewDock TDW is written in perl, python and shell scripts. It runs as a packaged, extension of UCSF Chimera, which can be downloaded free of charge at http://www.cgl.ucsf.edu/chimera/. ViewDock TDW has been created by a Licensee of Chimera and in accordance with the license agreement; our software is subject to same terms and conditions. Currently, ViewDock TDW only supports linux operating systems; downloads, instructions for example setup and usage as well as demo videos are available at http://tdw-prime.webs.com.
ACKNOWLEDGEMENTS
Molecular graphics images were produced using the UCSF Chimera package from the Resource for Biocomputing, Visualization and Informatics at the University of California, San Francisco supported by the National Institutes of Health [grant number P41 RR-01081].
Funding: National Science Foundation (OISE-0710726); the National Institutes of Health (HL085159); PRIUS (The Ministry of Education, Culture, Sports, Science and Technology funded University Education Internationalization Promotion Program); the Pacific Rim Applications and Grid Middleware Assembly (INT-0314015, OCI-0627026).
Conflict of Interest: none declared.
REFERENCES
- Baykoucheva S. A new era in chemical information: PubChem, Discovery-Gate, and Chemistry Central. ONLINE. 2007;31:17–20. [Google Scholar]
- Irwin JJ, et al. ZINC - a free database for virtual screening. J. Chem. Inf. Model. 2006;45:177–182. doi: 10.1021/ci049714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitchen DB, et al. Docking and scoring in virtual screening for drug discovery: methods and applications. Nat. Rev. Drug Discov. 2004;3:935–949. doi: 10.1038/nrd1549. [DOI] [PubMed] [Google Scholar]
- Lang PT, et al. DOCK, Version 6.2. 2008 Available at http://dock.compbio.ucsf.edu/ (last accessed date June 11, 2010) [Google Scholar]
- Levesque MJ, et al. Design of a grid service-based platform for in silico protein-ligand screenings. Comp. Meth. Prog. Biomed. 2008;93:73–82. doi: 10.1016/j.cmpb.2008.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pettersen EF, et al. UCSF Chimera – a visualization system for exploratory research and analysis. J. Comput. Chem. 2004;25:1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
- Wei B, et al. Visualization research with large displays. IEEE Comput. Graph. 2000;20:50–55. [Google Scholar]