Abstract
Background
The serum response element (SRE) in the c-fos promoter is a convergence point for several signaling pathways that regulate induction of the c-fos gene. Many transcription factors regulate the SRE, including serum response factor (SRF), ternary complex factor (TCF), and CCAAT/enhancer binding protein-beta (C/EBPβ). Independently, the TCFs and C/EBPβ have been shown to interact with SRF and to respond to Ras-dependent signaling pathways that result in transactivation of the SRE. Due to these common observations, we addressed the possibility that C/EBPβ and Elk-1 could both be necessary for Ras-stimulated transactivation of the SRE.
Results
In this report, we demonstrate that Elk-1 and C/EBPβ functionally synergize in transactivation of both a Gal4 reporter plasmid in concert with Gal4-SRF and in transactivation of the SRE. Interestingly, this synergy is only observed upon activation of Ras-dependent signaling pathways. Furthermore, we show that Elk-1 and C/EBPβ could interact both in an in vitro GST-pulldown assay and in an in vivo co-immunoprecipitation assay. The in vivo interaction between the two proteins is dependent on the presence of activated Ras. We have also shown that the C-terminal domain of C/EBPβ and the N-terminal domain of Elk-1 are necessary for the proteins to interact.
Conclusions
These data show that C/EBPβ and Elk-1 synergize in SRF dependent transcription of both a Gal-4 reporter and the SRE. This suggests that SRF, TCF, and C/EBPβ are all necessary for maximal induction of the c-fos SRE in response to mitogenic signaling by Ras.
Introduction
c-fos is a member of the family of immediate early genes, and its transcription is transiently induced in response to mitogenic signals [1]. The serum response element (SRE) is located approximately 300 bp upstream of the transcriptional start site in the c-fos promoter and is necessary for serum induction of c-fos [2]. The SRE binds a transcription factor named serum response factor (SRF) which was found to be necessary, but not sufficient, for serum induction of the SRE [3,4,5]. In vivo footprinting analysis shows that SRF is constitutively bound to the SRE in both quiescent and growth factor stimulated cells [6]. This suggests that it is the transcriptional activation of a complex of SRF and its accessory proteins that is regulated rather than regulation of SRF DNA binding.
The ternary complex factors (TCFs) are members of the ets family of transcription factors. The TCF family members Elk-1 [7], SAP-1 [8], and SAP-2/ERP/NET [9, 10] have been found to have a role in regulating the SRE. TCFs cannot bind the SRE autonomously, but require protein-protein interactions with SRF in order to bind the SRE [11,12,13,14,15]. The TCFs contain 3 conserved motifs termed the A, B, and C boxes [16]. The N-terminal A-box (amino acids (aa) 1-90 of Elk-1) is necessary to bind DNA, while the central B-box (aa 148-168) is the SRF interaction domain. The C-terminal C-box (aa 352-399), harboring the transactivation domain, contains several consensus mitogen activated protein kinase (MAPK) phosphorylation sites. Accordingly, the TCFs have been found to be targets of the Ras-Raf-MAPK signal transduction pathway [16]. In addition, the TCFs have been found to be targets for all three families of MAPKs: the extracellular signal-regulated kinases 1/2 (ERK1/2), the jun-N-terminal kinases/stress activated protein kinases (JNK/SAPK), and the p38 kinase [17, 18]. The transcriptional activity of the TCFs are stimulated by phosphorylation of the C-terminal MAPK sites [16, 17].
Another transcription factor that is involved in regulation of the c-fos SRE is CCAAT/Enhancer binding protein-beta (C/EBPβ). C/EBPβ (also known as NF-IL6, LAP, NF-M, AGP/EBP, and CRP2) is a member of the basic-leucine zipper family of transcription factors [19, 20]. The C/EBPβ mRNA contains three in-frame methionines which give rise to three different translation products: p38, p35, and p20-C/EBPβ [21]. p38 and p35-C/EBPβ both contain an N-terminal transactivation domain and a C-terminal DNA binding/dimerization domain. p20-C/EBPβ lacks the N-terminal transactivation domain and therefore acts as a repressor of transcription. Our lab has previously shown that p35-C/EBPβ activates an SRE-driven reporter construct while p20-C/EBPβ inhibits serum stimulation of the same reporter [22]. We have also shown that both p35-C/EBPβ and p20-C/EBPβ could interact with SRF in vivo and that the interaction between SRF and p35-C/EBPβ, but not between SRF and p20-C/EBPβ, is stimulated by activated Ras. The target for this Ras stimulation is Thr235 in a consensus MAPK site in C/EBPβ [23]. Therefore, C/EBPβ is a target of a Ras-dependent signaling pathway that regulates its interaction with SRF.
Based on the observations that TCF factors as well as p35-C/EBPβ: (1) interact with SRF (2) transactivate the SRE and (3) are responsive to Ras-dependent signaling pathways, we tested the possibility that both TCF and p35-C/EBPβ are necessary for maximal induction of the SRE in response to mitogenic stimulation. In this study, we show that p35-C/EBPβ and the TCF family member Elk-1 synergize in transactivation of SRF dependent transcription of both a Gal4 dependent reporter and an SRE-driven reporter construct, but only in response to mitogenic stimulation by Ras. We further show that Elk-1 and p35-C/EBPβ interact in vitro in a glutathione-S-transferase (GST)-pulldown assay as well as in an in vivo coimmunoprecipitation assay. The in vivo interaction is dependent on the presence of activated Ras. Finally, we show that the C-terminal domain of C/EBPβ is sufficient to interact with Elk-1 while the N-terminal A-box of Elk-1 is necessary to interact with C/EBPβ. These results suggest a cooperative role between the TCF and C/EBPβ transcription factors in regulation of the c-fos SRE in response to Ras-dependent signaling pathways.
Results
Elk-1 and p35-C/EBPβ synergize in transactivation of a GAL4 dependent reporter
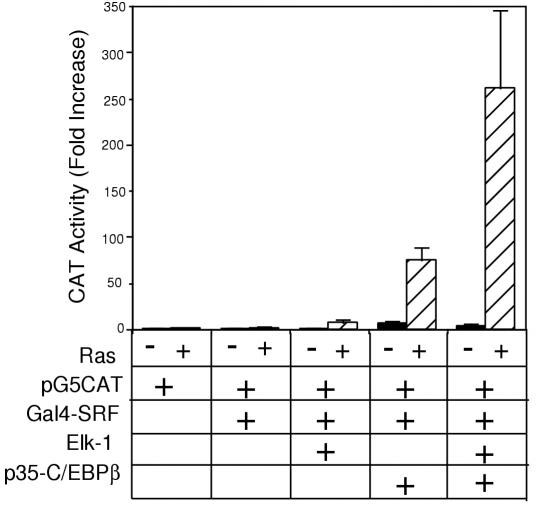
Due to the common observations that both Elk-1 and p35-C/EBPβ transactivate the SRE, interact with SRF, and are responsive to Ras-dependent signaling pathways, we hypothesized that Elk-1 and p35-C/EBPβ may both be necessary for maximal induction of SRF dependent transcription. To begin to test this hypothesis, we used a GAL4 dependent reporter construct, in which a CAT reporter gene is driven by five copies of a GAL4 binding site upstream of the adenovirus E1B minimal promoter (pG5CAT). We constructed a GAL4 DNA binding domain-SRF fusion protein, as has been described previously [23]. GAL4-SRF binds to the GAL4 reporter construct, thereby making transcription of the reporter gene dependent on the presence of SRF. As shown in Fig. 1, when a GAL4-SRF fusion construct is transfected with the reporter, there is no increase in CAT activity. The SRF transactivation domain is weak, as has been shown previously [24]. When CMV-Elk-1 is cotransfected with the reporter and GAL4-SRF, there is little increase in CAT activity in the absence of activated Ras (CMV-Ras.V12). This result is expected since the transactivation domain of Elk-1 is activated in response to Ras. Therefore, when activated Ras is transfected with Elk-1, GAL4-SRF, and the reporter, the CAT activity increases to 8-fold over basal levels. We cannot determine if the increase in CAT activity in the presence of Ras also reflects a stimulation of the interaction of the Elk-1 and SRF proteins.
Figure 1.
Elk-1 and p35-C/EBPβ synergize in SRF dependent transactivation of a GAL4 reporter construct. NIH 3T3 cells were transfected with 1 μg of pG5CAT alone, with 0.5 μg of pGAL4-SRF, or with 0.5 μg of pGAL4-SRF and 0.5 μg of pCMV-LAP (encoding p35-C/EBPβ) and/or 0.5 μg of pCMV-Elk-1 as indicated, in the absence and presence of 1 μg pCMV-Ras.V12. Total DNA in each transfection was adjusted to 3.5 μg with pUC19. Cells were serum deprived for 40 h prior to harvesting. CAT activity was measured as previously described [33]. Data are the average of 14 determinations, and error bars represent standard error.
When CMV-LAP (which encodes p35-C/EBPβ) is transfected with GAL4-SRF and the reporter construct, we see a 7-fold increase in CAT activity that is potentiated to 75-fold when activated Ras is cotransfected. Therefore, C/EBPβ results in a much larger increase in transcription than Elk-1. We have previously shown that Ras does not activate the transactivation domain of p35-C/EBPβ [23]. Thus, the increase in transcription in this assay is due to Ras stimulation of the SRF-p35-C/EBPβ interaction.
Interestingly, when all three constructs - SRF, Elk-1, and p35-C/EBPβ-are cotransfected with the reporter construct, there is an average 260-fold increase in CAT activity in the presence of activated Ras. The values of fold activation varied from as low as 60-fold to as high as 725-fold over basal levels, and we are unsure of the reason for this variability. However, regardless of the extent of activation, in every experiment there was a synergy observed when both Elk-1 and p35-C/EBPβ are transfected in the presence of Ras. There is only a slight increase in CAT activity in the absence of Ras. Therefore, Elk-1 and p35-C/EBPβ are working synergistically to transactivate the reporter construct in the presence of SRF. This synergism is only observed in response to activation of mitogenic signaling pathways by Ras.
Elk-1 and p35-C/EBPβ synergize in transactivation of the SRE
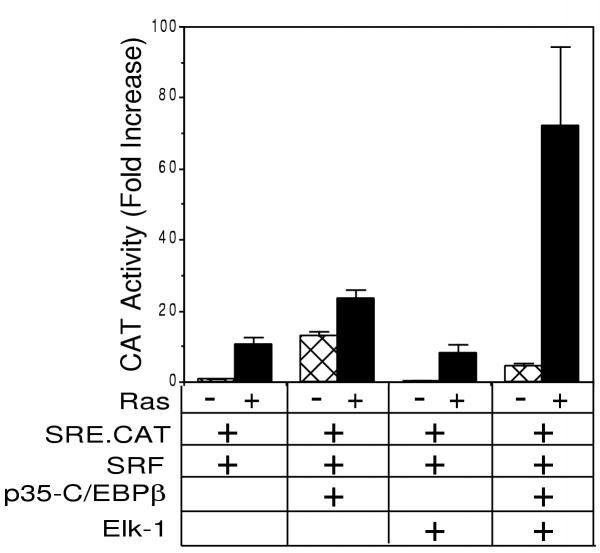
Since Elk-1 and p35-C/EBPβ synergize in transactivation of SRF-dependent transcription using a GAL4-dependent promoter, we next tested if Elk-1 and p35-C/EBPβ could also synergize in transactivation of a native SRF binding site, namely the c-fos SRE. To test this possibility, NIH 3T3 cells were transiently transfected with a CAT reporter gene driven by one copy of the wild type SRE upstream of the Rous sarcoma virus long terminal repeat minimal promoter. As shown in Fig. 2, when p35-C/EBPβ is co-transfected with the reporter construct, there is a 13-fold increase in CAT activity in the absence of activated Ras that is increased to 24-fold when CMV-Ras.V12 is co-transfected.
Figure 2.
Elk-1 and p35-C/EBPβ synergize in transactivation of the SRE. NIH 3T3 cells were transfected with 0.75 μg of SRE.CAT and 0.75 μg pCMV-SRF alone or with 0.75 μg pCMV-LAP or 0.75 μg pCMV-Elk1 in the absence or presence of 0.75 μg pCMV-Ras.V12. Cells were serum deprived for 40 h prior to harvesting. CAT activity was measured as previously described [33]. Data are the average of 4 determinations, and error bars represent standard error.
When CMV-Elk-1 is co-transfected with the SRE reporter construct, there is no additional stimulation in transactivation in either the absence or presence of Ras compared to the reporter alone. However, when both Elk-1 and p35-C/EBPβ are transfected with the SRE reporter construct, there is a synergistic effect in transactivation of the SRE, with a 72-fold increase in CAT activity over reporter construct alone. As was seen with the Gal4 reporter, this synergism is only observed in the presence of activated Ras. These data suggest that both Elk-1 and p35-C/EBPβ are necessary for maximal Ras-stimulated transactivation of the SRE.
The TCF family member Elk-1 and C/EBPβ interact in vitro
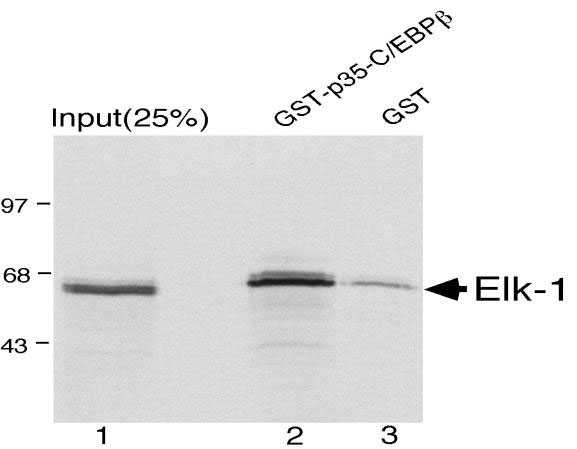
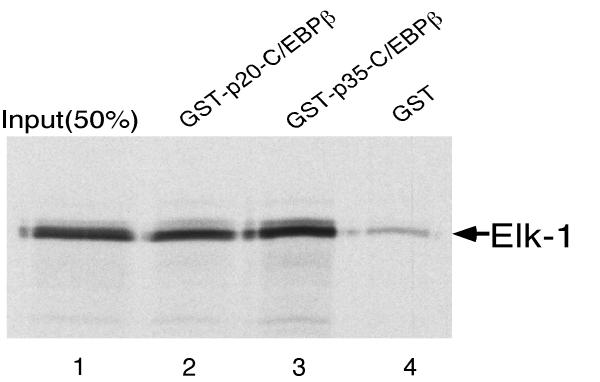
We next tested the possibility that there could be a direct protein-protein interaction between Elk-1 and p35-C/EBPβ based on the fact that they synergize in transactivation of the SRE. Therefore, we used a GST-pulldown assay to determine if the proteins could interact in vitro. p35-C/EBPβ was expressed as a chimeric GST protein and immobilized on glutathione-agarose beads. Beads containing GST-p35-C/EBPβ or GST alone were incubated with in vitro-translated Elk-1 labeled with [35S] methionine. As shown in Fig. 3, lane 2, approximately 35-45% of the input Elk-1 was retained on the GST-p35-C/EBPβ beads. A small amount of Elk-1 bound to the beads containing GST alone (Fig. 3, lane 3) which we have been unable to eliminate even after blocking with unprogrammed translation lysate. However, it is clear that the binding of Elk-1 is substantially increased when the GST-p35-C/EBPβ fusion protein is present on the beads. These data indicate that Elk-1 and p35-C/EBPβ are capable of interacting in vitro.
Figure 3.
Elk-1 binds to GST-p35-C/EBPβ in vitro. Radiolabeled Elk-1 was prepared by in vitro transcription and translation of CMV-Elk-1 expression construct and then incubated with equivalent amounts of GST or GST-p35-C/EBPβ protein immobilized on glutathione-Sepharose. Proteins eluted from the GST-p35-C/EBPβ beads (lanes 2) or GST beads (lanes 3) by boiling in Laemmli sample buffer were resolved on an SDS-10% polyacrylamide gel and detected by autoradiography. Lane 1 contains one-fourth of the amount of the radiolabeled proteins initially mixed with the beads.
Elk-1 and p35-C/EBPβ interact in vivo, but only in the presence of activated Ras
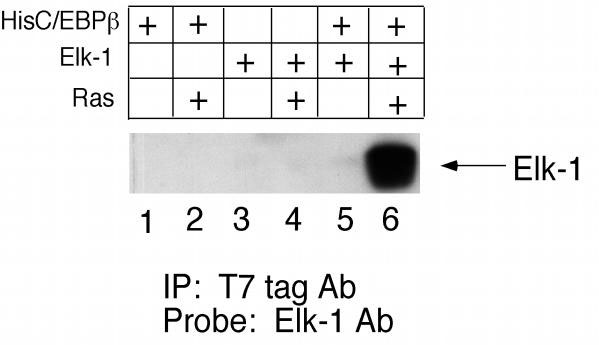
Since we observed an in vitro interaction between the Elk-1 and C/EBPβ proteins, we next tested whether the proteins could interact in vivo as well. To do this, we used a co-immunoprecipitation approach. COS-7 cells were transfected with an expression vector for a 6X histidine tagged construct of p35-C/EBPβ carrying the φ 10 (T7 tag) epitope sequence either in the presence or absence of Elk-1. Cell lysates were incubated with T7 tag Ab agarose beads followed by immunoblotting of the precipitated proteins with Elk-1 Ab. As shown in Fig. 4, there is no Elk-1 protein precipitated with the tagged C/EBPβ protein when both are transfected (lane 5). However, since the synergism of Elk-1 and p35-C/EBPβ is observed when activated Ras is present, we thought it likely that the interaction between the two proteins could be Ras-dependent. Indeed, when activated Ras is co-transfected along with Elk-1 and histidine-tagged p35-C/EBPβ, the Elk-1 protein is precipitated with the p35-C/EBPβ (lane 6). One possible explanation for this result is that Ras increases the amount of p35-C/EBPβ or Elk-1 protein in the COS-7 cells, but Western blot analysis showed that both p35-C/EBPβ and Elk-1 protein levels are the same in the absence and presence of Ras (data not shown). Therefore, p35-C/EBPβ and Elk-1 interact in vivo, but only in response to activation of Ras-dependent signaling pathways.
Figure 4.
Elk-1 coimmunoprecipitates with p35-C/EBPβ, but only in the presence of activated Ras. COS-7 cells were transfected with 10 μg of pCDNA3.1/His-p35-C/EBPβ (lanes 1, 2, 5, 6) and 10 μg of pCMV-Elk-1 (lanes 3, 4, 5, 6) in the presence and absence of 2 μg of pCMV-Ras.V12 as indicated. Cells were harvested 40 h post-transfection, and whole cell lysates were incubated with T7tag Ab-agarose beads, followed by immunoblotting of precipitated proteins with Elk-1 Ab.
The C-terminal domain of C/EBPβ is necessary to interact with Elk-1 in vitro
Since C/EBPβ and Elk-1 interact, we next wanted to narrow down the domains of the proteins that are required for their interaction. To determine the domain of C/EBPβ that is necessary to interact with Elk-1, a p20-C/EBPβ-GST fusion protein was constructed. p20-C/EBPβ encodes the 20 kDa form of C/EBPβ, which lacks the N-terminal transactivation domain of the longer p35-C/EBPβ isoform. This isoform, however, shares the C-terminal DNA binding and dimerization domain with p35-C/EBPβ. As shown in Fig. 5, approximately the same amount of [35S]-labeled Elk-1 is retained on both the GST-p20-C/EBPβ and GST-p35-C/EBPβ beads (compare lanes 2 and 3). Therefore, deletion of the N-terminus of C/EBPβ has no effect on its ability to interact with Elk-1. This data demonstrates that the C-terminal region of C/EBPβ is sufficient to mediate interaction with Elk-1 in vitro.
Figure 5.
The C-terminal domain of C/EBPβ interacts with Elk-1. Radiolabeled Elk-1 was prepared by in vitro transcription and translation of CMV-Elk-1 expression construct and then incubated with equivalent amounts of GST, GST-p35-C/EBPβ, or GST-p20-C/EBPβ protein immobilized on glutathione-Sepharose. Proteins eluted from the GST-p20-C/EBPβ beads (lane 2), the GST-p35-C/EBPβ beads (lane 3) or GST beads (lanes 4) were resolved on an SDS-10% polyacrylamide gel and detected by autoradiography. Lane 1 contains one-half of the amount of the radiolabeled proteins initially mixed with the beads.
The N-terminal A-box of Elk-1 is sufficient to interact with C/EBPβ in vitro
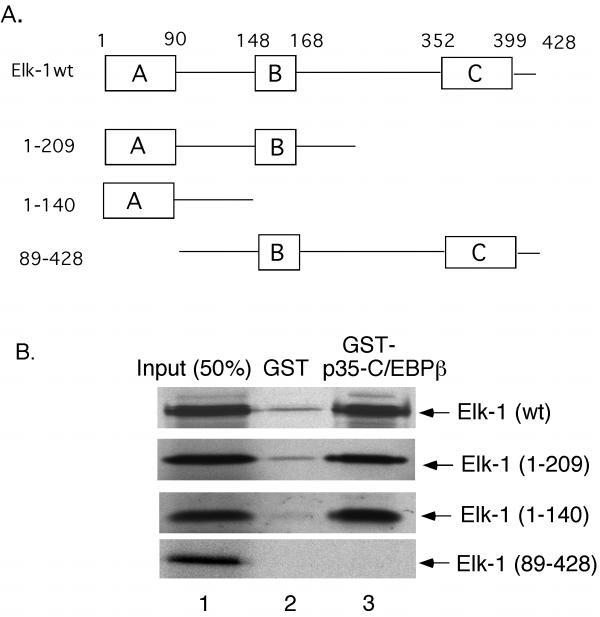
In order to narrow down the domain of Elk-1 that is necessary to interact with C/EBPβ in vitro, we made several Elk-1 deletion mutants (Fig. 6A) and tested these constructs in a pulldown assay with GST-p35-C/EBPβ. Elk-1(1-209) is a C-terminal deletion mutant that lacks the C-box. Elk-1(1-140) is also a C-terminal deletion, but it lacks both the B- and C-boxes. Therefore, this mutant contains neither a transactivation domain nor an SRF binding domain. Finally, Elk-1(89-428) is a deletion of the N-terminal A-box. Glutathione agarose beads containing GST-p35-C/EBPβ or GST alone were incubated with in vitro-translated Elk-1 mutants labeled with [35S] methionine. As shown in Fig. 6B, the C-terminal Elk-1 mutants, Elk-1(1-209) and Elk-1(1-140), bound to the GST-p35-C/EBPβ beads to the same extent as wild-type Elk-1 (compare lane 3 of top 3 panels). However, the A-box deletion mutant, Elk-1(89-428), no longer binds to the GST-p35-C/EBPβ (lane 3, bottom panel). Therefore, this data demonstrates that the A-box of Elk-1 is necessary to interact with C/EBPβ in vitro.
Figure 6.
The A-box of Elk-1 is necessary for interaction with p35-C/EBPβ in vitro. A) Schematic representation of Elk-1 wild-type and deletion mutants. B) Radiolabeled Elk-1(wt), Elk-1(1-209), Elk-1(1-140), and Elk-1(89-428) were prepared by in vitro transcription and translation of their respective expression constructs (see Mat. and Meth.). The proteins were incubated with equivalent amounts of GST or GST-p35-C/EBPβ protein immobilized on glutathione-Sepharose. Proteins eluted from the GST beads (lanes 2) or GST-p35-C/EBPβ beads (lanes 3) were resolved on an SDS-10% polyacrylamide gel and detected by autoradiography. Lane 1 contains one-half of the amount of the radiolabeled proteins initially mixed with the beads.
Discussion
We have shown that the transcription factors Elk-1 and p35-C/EBPβ synergize in transactivation of SRF-dependent transcription using both GAL4 and SRE-driven reporter constructs. Interestingly, this synergy is only observed in response to mitogenic stimulation by Ras. We have also demonstrated that the Elk-1 and p35-C/EBPβ proteins interact using both in vitro GST-pulldown and in vivo coimmunoprecipitation assays. The in vivo interaction, however, is dependent on the presence of activated Ras. Finally, we demonstrate that the in vitro interaction domains of the two proteins are the C-terminal domain of C/EBPβ and the N-terminal domain of Elk-1.
These results suggest a new mechanism for transactivation of the SRE, where a competent transcriptional complex consisting of at least SRF, TCF, and p35-C/EBPβ participate in transactivation in response to Ras. To this point, the TCF family members have been thought to be the main targets of Ras signaling to the SRE, and activation of TCF by Ras has been shown to result in transactivation of the SRE [16]. However, we show that Ras activation of TCF alone does not result in maximal SRE transactivation, but instead, is greatly enhanced in the presence of p35-C/EBPβ.
SRF and TCF have been shown to form a ternary complex at the SRE in vitro. We as yet have not been able to observe a ternary complex between SRF and p35-C/EBPβ at the SRE in vitro. The mechanisms of SRE transactivation described above may help to explain the in vitro results. The C/EBPβ recognition site in the SRE is a weak binding site [25]. Upon activation of Ras-dependent signaling pathways, the presence of p35-C/EBPβ at the SRE would be stabilized due to its strong protein-protein interactions not only with SRF, as we have shown previously [23], but also with Elk-1. It is possible that p35-C/EBPβ and SRF are not sufficient to form a ternary complex with the SRE in vitro, but Elk-1 may also be necessary. In addition, it is likely that p35-C/EBPβ, Elk-1, or both, may need to be activated by one or more MAPK family members for a multiprotein complex to form with SRF at the SRE. Further studies utilizing the information from the in vivo studies performed here may permit us to optimize the in vitro conditions necessary to observe such a multiprotein complex.
We observe a direct interaction between C/EBPβ and Elk-1 in a GST pulldown assay. Another ets family member, Ets-1, has previously been shown to interact with C/EBPβ in vitro [26]. We have not directly tested the ability of the other TCF famlily members, SAP1 and SAP2/ERP/NET, to interact with C/EBPβ. We find that the domain of Elk-1 that interacts with C/EBPβ is the ets domain, which is conserved among the other TCF family members [16]. We would therefore also expect SAP1 and SAP2/ERP/NET to interact with C/EBPβ. Interestingly, the in vivo interaction of Elk-1 and C/EBPβ is dependent on activation of Ras-dependent signaling pathways. The target(s) of the Ras signaling pathway is as yet unknown. It has been shown that transfection of activated Ras results in phosphorylation of Thr235 (numbering for human protein) in the C-terminal domain of C/EBPβ, and that Thr235 could be phosphorylated by a partially purified MAPK preparation in vitro [19]. We have also shown that this same residue is necessary for Ras stimultion of the interaction between C/EBPβ and SRF. Therefore, it will be interesting to determine if this site is also critical for stimulation of the interaction between C/EBPβ and Elk-1. Elk-1 also has several MAPK sites in its C-terminal domain, and therefore this region could also be a target for Ras.
We are currently investigating the mechanism of the observed synergy between Elk-1 and p35-C/EBPβ. We have shown that C/EBPβ interacts with the A-box of Elk-1, while SRF has previously been shown to interact with the B-box [16]. Therefore, C/EBPβ and SRF interact with two distinct domains of Elk-1, as well as with each other, which could allow formation of a complex of the three transcription factors. This would create an active transcriptional complex at the SRE and result in enhanced transactivation. It is possible that the p35-C/EBPβ-SRF-Elk-1 complex could interact more strongly with coactivators and/or the basal transcriptional machinery than each individual transcription factor. It has been shown that SRF, TCF, and C/EBPβ can all interact with components of transcriptional machinery such as p300/CBP, SRC-1, and TFII-I [27,28,29,30]. There is most likely a large protein complex assembled at the SRE composed of regulated transcription factors, coactivators, and the basal transcriptional machinery, resulting in rapid transcription of the c-fos gene in response to mitogenic stimulation.
Conclusions
This report demonstrates a new model for c-fos SRE activation in response to Ras-dependent signaling pathways. We show that SRF, Elk-1, and p35-C/EBPβ are all necessary for maximal Ras-stimulated transactivation of the SRE.
Materials and Methods
Cell Culture and Transfections
NIH 3T3 fibroblasts (from the American Type Culture Collection) and COS-7 cells (kindly provided by Dr. S. Hann, Vanderbilt University) were grown in Dulbecco's Modified Eagle Medium (DMEM) with 10% calf serum (Colorado Serum Company), 0.22% sodium bicarbonate, 4 mM L-glutamine, 25 U of penicillin G sodium per mL, and 25 mg of streptomycin per mL.
COS-7 cell transfections were performed by the calcium phosphate (CaPO4) technique [31]. Cells at 60-70% confluence were exposed to the CaPO4-DNA precipitate for 8 h. The medium was removed and replaced with complete medium for 36 h before harvesting. NIH 3T3 transfections were performed using NovaFector (Venn Nova) or Trans-IT LT1 (PanVera) as described by the manufacturers.
For NovaFector transfections, a 6:1 ratio of NovaFector:DNA was used for each transfection. Cells at 60-70% confluence were exposed to the NovaFector:DNA complex for 6-7 h in serum and antibiotic free medium. After the incubation, an equal volume of DMEM containing 20% calf serum was added for 24 h, bringing the final concentration of calf serum to 10%. Cells were then serum deprived for 36-40 h in DMEM supplemented as above except containing 0.5% calf serum before harvesting. For TransIT-LT1 transfections, a 6:1 ratio of TransIT-LT1:DNA was used in each transfection. Cells at 60-70% confluence were exposed to the TransIT-LT1:DNA complex for 8 h in complete medium plus freshly added 30 mg/mL polymyxin B antibiotic [32]. The medium was then removed and replaced with complete medium for 24 h. Cells were then serum deprived in DMEM containing 0.5% calf serum for 36-40 h before harvesting. Cell extracts were prepared and chloramphenicol acetyl transferase (CAT) assays were performed on extracts containing equivalent cell protein as previously described [33]. An internal control plasmid to measure transfection efficiency could not be used because C/EBPβ regulates transcrption from the control plasmid, and thus makes the internal control invalid. Therefore, the transfections were repeated multiple times to control for variability in transfection efficiency.
Plasmids
The pGAL4 and pG5CAT plasmids were obtained from the Mammalian MATCHMAKER Two-Hybrid Assay Kit (CLONTECH). pGAL4-SRF was constructed as previously described [23]. The SRE-CAT reporter gene was constructed as previously described [22]. pGST-p35-C/EBPβ was constructed by inserting the 1,739 bp EcoRI fragment from pRSETB-EFII [34] into EcoRI cut pGEX-4T-1 (Amersham-Pharmacia). pGST-p20-C/EBPβ was constructed by inserting the 581 bp BamHI/EcoRI fragment from pRsetA-LIP [34] into similarly digested pGEX4T-1. pcDNA3/Elk-1(1-209) and pcDNA3/Elk-1(1-140) were constructed by using the Erase-A-Base system (Promega) with pcDNA3/Elk-1 (gift of J. Schwartz, Univ. of Michigan) as described by the manufacturer. pcDNA3.1/His-p35-C/EBPβ was constructed by inserting an 1,739 bp EcoRI fragment from pRSETB-EFII into an EcoRI cut pcDNA3.1/HisC vector (Invitrogen). CMV-LAP was a gift of U. Schibler (Univ. of Geneva, Geneva, Switzerland) and CMV-Ras.V12 was a gift of E. Ruley (Vanderbilt Univ.).
Analysis of ELK-1 and C/EBPβ interaction in vitro
GST-p35-C/EBPβ, GST-p20-C/EBPβ, or GST alone was produced in BL21 E. Coli by 80 mM isopropyl-β-D-thiogalactopyranoside (IPTG) induction overnight at 37°C. Cells were harvested by centrifugation and washed once with phosphate-buffered saline (PBS) containing the following protease inhibitors: 2 mg/ml aprotinin, 2 mg/ml leupeptin, 2 mM phenylmethylsulfonyl fluoride (PMSF), and 0.2 mM pepstatin. The bacteria were lysed by sonication at 4oC in PBS containing the protease inhibitors described above. Triton X-100 was added to a final concentration of 0.1%, followed by gentle mixing for 30 min at 4°C. The lysate was clarified at 12,000 × g for 10 min at 4°C. The supernatant was gently mixed with glutathione-sepharose beads (Amersham-Pharmacia) at 4°C for 30 min. Beads containing GST proteins were collected by low speed centrifugation, followed by three successive washes with PBS containing 0.1% Triton X-100 and the protease inhibitors described above.
The ELK-1, ELK-1(1-209), ELK-1(1-140), and ELK-1(89-428) proteins were transcribed and translated in vitro using the TNT T7 coupled reticulocyte lysate system (Promega) according to the manufacture's instructions. Translated proteins were radiolabeled with the EXPRE35S35S protein labeling mix (Dupont NEN). The GST-pulldown assay was done as previously described [22].
Co-Immunoprecipitation
COS-7 cells were harvested in PBS containing 0.1 mM sodium vanadate and collected by low speed centrifugation. Cells were resuspended in lysis buffer (10 mM Tris (pH 7.5), 1 mM EDTA, 50 mM NaCl, 0.25% Nonidet P-40, 1 mM PMSF, 1 mg of aprotinin/mL, 0.1 mM sodium vanadate, 10 mM sodium molybdate, and 10 mM β-glycerol phosphate) and lysed by sonication with a microtip on setting 2 and 20% duty cycle for 10 s. After clarification by centrifugation at 12,000 × g for 10 min, the cell extract was incubated with T7 tag antibody (Ab)-agarose beads (Novagen) for 2 h. The beads were collected by low speed centrifugation and washed 3 times with lysis buffer. All steps were performed at 4°C. After washing, the beads were boiled for 5 min in Laemmli sample buffer.
Immunoblots
The samples from the coimmunoprecipitations described above were analyzed by electrophoresis on an SDS-12% polyacrylamide gel for 3 h at 160 V. The gel was equilibrated in transfer buffer (33 mM Tris base, 192 mM glycine, 20% methanol) for 15 min before transfer of proteins to Immobilon-P membrane (Millipore Corp.). After transfer, an immunoblot was performed as described previously [22]. A 1:2000 dilution of anti-Elk-1 Ab (Santa Cruz Biotechnology) and a 1:5000 dilution of goat anti-rabbit secondary Ab (Roche Molecular Biochemicals) were used. The secondary Ab was detected using SuperSignal Chemiluminescent Substrate (Pierce).
Abbreviations
aa, amino acid; Ab, antibody; bp, base pair; CaPO4, calcium phosphate; C/EBPβ, CCAAT-enhancer binding protein-beta; CAT, chloramphenicol acetyl transferase; DMEM, Dulbecco's Modified Eagle Medium; GST, glutathione-S-transferase; MAPK, mitogen activated protein kinase; PBS, phosphate buffered saline; PMSF, phenylmethylsulfonyl fluoride; SRE, serum response element; SRF, serum response factor; TCF, ternary complex factor.
Acknowledgments
Acknowledgments
We thank John van Doorninick and Michael Hann for expert technical assistance, and Robert Tilghman for critically reviewing the manuscript.
M. Hanlon was supported as a predoctoral trainee on the Cellular, Biochemical, and Molecular Sciences Training Grant 5T32GM08554. L. Bundy was supported by a postdoctoral fellowship from the Molecular Endocrinology Training Grant 5T32DK07563.
Contributor Information
Mary Hanlon, Email: marry.hanlon@mcmail.vanderbilt.edu.
Linda M Bundy, Email: linda.bundy@mcmail.vanderbilt.edu.
Linda Sealy, Email: linda.sealy@mcmail.vanderbilt.edu.
References
- Ransone LJ, Verma IM. Nuclear proto-oncogenes fos and jun. Annu Rev Cell Biol. 1990;6:539–557. doi: 10.1146/annurev.cellbio.6.1.539. [DOI] [PubMed] [Google Scholar]
- Treisman R. The SRE: a growth factor responsive transcriptional regulator. Semin Cancer Biol. 1990;1:47–58. [PubMed] [Google Scholar]
- Treisman R. Identification and purification of a polypeptide that binds to the c- fos serum response element. Embo J. 1987;6:2711–2717. doi: 10.1002/j.1460-2075.1987.tb02564.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norman C, Runswick M, Pollock R, Treisman R. Isolation and properties of cDNA clones encoding SRF, a transcription factor that binds to the c-fos serum response element. Cell. 1988;55:989–1003. doi: 10.1016/0092-8674(88)90244-9. [DOI] [PubMed] [Google Scholar]
- Johansen FE, Prywes R. Two pathways for serum regulation of the c-fos serum response element require specific sequence elements and a minimal domain of serum response factor. Mol Cell Biol. 1994;14:5920–5928. doi: 10.1128/mcb.14.9.5920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herrera RE, Shaw PE, Nordheim A. Occupation of the c-fos serum response element in vivo by a multi- protein complex is unaltered by growth factor induction. Nature. 1989;340:68–70. doi: 10.1038/340068a0. [DOI] [PubMed] [Google Scholar]
- Hipskind RA, Rao VN, Mueller CG, Reddy ES, Nordheim A. Ets-related protein Elk-1 is homologous to the c-fos regulatory factor p62TCF. Nature. 1991;354:531–534. doi: 10.1038/354531a0. [DOI] [PubMed] [Google Scholar]
- Dalton S, Treisman R. Characterization of SAP-1, a protein recruited by serum response factor to the c-fos serum response element. Cell. 1992;68:597–612. doi: 10.1016/0092-8674(92)90194-H. [DOI] [PubMed] [Google Scholar]
- Giovane A, Pintzas A, Maira SM, Sobieszczuk P, Wasylyk B. Net, a new ets transcription factor that is activated by Ras. Genes Dev. 1994;8:1502–1513. doi: 10.1101/gad.8.13.1502. [DOI] [PubMed] [Google Scholar]
- Lopez M, Oettgen P, Akbarali Y, Dendorfer U, Libermann TA. ERP, a new member of the ets transcription factor/oncoprotein family: cloning, characterization, and differential expression during B- lymphocyte development. Mol Cell Biol. 1994;14:3292–3309. doi: 10.1128/mcb.14.5.3292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janknecht R, Nordheim A. Elk-1 protein domains required for direct and SRF-assisted DNA-binding. Nucleic Acids Res. 1992;20:3317–3324. doi: 10.1093/nar/20.13.3317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw PE, Frasch S, Nordheim A. Repression of c-fos transcription is mediated through p67SRF bound to the SRE. Embo J. 1989;8:2567–2574. doi: 10.1002/j.1460-2075.1989.tb08395.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw PE. Ternary complex formation over the c-fos serum response element: p62TCF exhibits dual component specificity with contacts to DNA and an extended structure in the DNA-binding domain of p67SRF. Embo J. 1992;11:3011–3019. doi: 10.1002/j.1460-2075.1992.tb05371.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rao VN, Reddy ES. elk-1 domains responsible for autonomous DNA binding, SRE:SRF interaction and negative regulation of DNA binding. Oncogene. 1992;7:2335–2340. [PubMed] [Google Scholar]
- Treisman R, Marais R, Wynne J. Spatial flexibility in ternary complexes between SRF and its accessory proteins. Embo J. 1992;11:4631–4640. doi: 10.1002/j.1460-2075.1992.tb05565.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treisman R. Ternary complex factors: growth factor regulated transcriptional activators. Curr Opin Genet Dev. 1994;4:96–101. doi: 10.1016/0959-437X(94)90097-3. [DOI] [PubMed] [Google Scholar]
- Treisman R. Regulation of transcription by MAP kinase cascades. Curr Opin Cell Biol. 1996;8:205–215. doi: 10.1016/S0955-0674(96)80067-6. [DOI] [PubMed] [Google Scholar]
- Whitmarsh AJ, Davis RJ. Transcription factor AP-1 regulation by mitogen-activated protein kinase signal transduction pathways. J Mol Med. 1996;74:589–607. doi: 10.1007/s001090050063. [DOI] [PubMed] [Google Scholar]
- Akira S, Isshiki H, Sugita T, Tanabe O, Kinoshita S, Nishio Y, Nakajima T, Hirano T, Kishimoto T. A nuclear factor for IL-6 expression (NF-IL6) is a member of a C/EBP family. Embo J. 1990;9:1897–1906. doi: 10.1002/j.1460-2075.1990.tb08316.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Descombes P, Chojkier M, Lichtsteiner S, Falvey E, Schibler U. LAP, a novel member of the C/EBP gene family, encodes a liver-enriched transcriptional activator protein. Genes Dev. 1990;4:1541–1551. doi: 10.1101/gad.4.9.1541. [DOI] [PubMed] [Google Scholar]
- Descombes P, Schibler U. A liver-enriched transcriptional activator protein, LAP, and a transcriptional inhibitory protein, LIP, are translated from the same mRNA. Cell. 1991;67:569–579. doi: 10.1016/0092-8674(91)90531-3. [DOI] [PubMed] [Google Scholar]
- Sealy L, Malone D, Pawlak M. Regulation of the cfos serum response element by C/EBPbeta. Mol Cell Biol. 1997;17:1744–1755. doi: 10.1128/mcb.17.3.1744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanlon M, Sealy L. Ras regulates the association of serum response factor and CCAAT/enhancer-binding protein beta. J Biol Chem. 1999;274:14224–14228. doi: 10.1074/jbc.274.20.14224. [DOI] [PubMed] [Google Scholar]
- Johansen FE, Prywes R. Identification of transcriptional activation and inhibitory domains in serum response factor (SRF) by using GAL4-SRF constructs. Mol Cell Biol. 1993;13:4640–4647. doi: 10.1128/mcb.13.8.4640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boulden AM, Sealy LJ. Maximal serum stimulation of the c-fos serum response element requires both the serum response factor and a novel binding factor, SRE-binding protein. Mol Cell Biol. 1992;12:4769–4783. doi: 10.1128/mcb.12.10.4769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNagny KM, Sieweke MH, Doderlein G, Graf T, Nerlov C. Regulation of eosinophil-specific gene expression by a C/EBP-Ets complex and GATA-1. Embo J. 1998;17:3669–3680. doi: 10.1093/emboj/17.13.3669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramirez S, Ait-Si-Ali S, Robin P, Trouche D, Harel-Bellan A, Ait Si Ali S. The CREB-binding protein (CBP) cooperates with the serum response factor for transactivation of the c-fos serum response element [published erratum appears in J Biol Chem 1999 Jun 18;274(25):18140]. J Biol Chem. 1997;272:31016–31021. doi: 10.1074/jbc.272.49.31016. [DOI] [PubMed] [Google Scholar]
- Kim HJ, Kim JH, Lee JW. Steroid receptor coactivator-1 interacts with serum response factor and coactivates serum response element-mediated transactivations. J Biol Chem. 1998;273:28564–28567. doi: 10.1074/jbc.273.44.28564. [DOI] [PubMed] [Google Scholar]
- Janknecht R, Nordheim A. MAP kinase-dependent transcriptional coactivation by Elk-1 and its cofactor CBP. Biochem Biophys Res Commun. 1996;228:831–837. doi: 10.1006/bbrc.1996.1740. [DOI] [PubMed] [Google Scholar]
- Mink S, Haenig B, Klempnauer KH. Interaction and functional collaboration of p300 and C/EBPbeta. Mol Cell Biol. 1997;17:6609–6617. doi: 10.1128/mcb.17.11.6609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham FL, Eb AJvd. A new technique for the assay of infectivity of human adenovirus 5 DNA. Virology. 1973;52:456–467. doi: 10.1016/0042-6822(73)90341-3. [DOI] [PubMed] [Google Scholar]
- Cotten M, Saltik M. Intracellular delivery of lipopolysaccharide during DNA transfection activates a lipid A-dependent cell death response that can be prevented by polymyxin B. Hum Gene Ther. 1997;8:555–561. doi: 10.1089/hum.1997.8.5-555. [DOI] [PubMed] [Google Scholar]
- Mobley CM, Sealy L. Role of the transcription start site core region and transcription factor YY1 in Rous sarcoma virus long terminal repeat promoter activity. J Virol. 1998;72:6592–6601. doi: 10.1128/jvi.72.8.6592-6601.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sears RC, Sealy L. Multiple forms of C/EBP beta bind the EFII enhancer sequence in the Rous sarcoma virus long terminal repeat [published erratum appears in Mol Cell Biol 1994 Aug;14(8):5617]. Mol Cell Biol. 1994;14:4855–4871. doi: 10.1128/mcb.14.7.4855. [DOI] [PMC free article] [PubMed] [Google Scholar]