Abstract
We recently reported identification of the bacterium Chlamydia pneumoniae in affected brain regions of patients with Alzheimer's disease (AD) (Balin et al., 1998). In this report, we extend those initial observations to demonstrate that, in addition to the frequently described, standard morphological forms of the organism, pleiomorphic forms are also present in the AD brain. All AD and control brain tissues examined were verified to be PCR-positive and negative, respectively, for the organism. DNA sequence determination of PCR products so derived from total DNA of infected AD brains, as well as from total DNA of cell lines infected with the organism following isolation from these same patient samples, confirmed the presence of organism in relevant samples. Various morphologic forms of C. pneumoniae were identified in PCR-positive tissues and these were characterized based on membrane structure, core density, size, and immunolabeling profiles. Structures identified include the typical pear-shaped elementary body, as well as larger, spherical and oblong reticulate bodies. Intact C. pneumoniae were found both intracellularly and extracellularly in the sampled autopsy brains. Intracellular organisms were located principally within microglia, astroglia, and presumptive pericytes. These results suggest that C. pneumoniae found in cells indigenous to the AD brain do not conform universally to the classical morphology observed in other infected cell types. This pleiomorphism may reflect an adaptive response and/or persistent state of infection for these organisms in Alzheimer's Disease.
Keywords: C. pneumoniae, Alzheimer's, Disease, Ultrastructure, Dementia
Introduction
A recent study from this group described an association between the obligate, intracellular bacterium Chlamydia pneumoniae and late-onset, sporadic Alzheimer's Disease (AD) (Balin et al., 1998). Although we have yet to establish a causal relationship between acute or chronic C. pneumoniae infection of the brain and development of late-onset AD, our prior immunohistochemistry results did demonstrate that microglia and astroglia in areas of typical AD neuropathology harbored this organism in a high proportion of patients. Ultrastructural analysis revealed that the organism exhibited morphological characteristics previously described for C. pneumoniae (Miyashita et al., 1993). In this regard, elementary bodies (EBs) and reticulate bodies (RBs) ranging in size from 0.2–1.2 μm in diameter were easily discernable. Moreover, we confirmed that the organisms identified in ultrastructural studies were indeed C. pneumoniae using a highly specific, monoclonal Ab (mAb) that targets an outer membrane protein (OMP) of the organism. A second mAb which specifically targets the lipopolysaccharide (LPS) of the genus Chlamydia strongly immunolabeled cells in AD brains as well. Double immunolabeling studies using glia-specific Ab, including those targeting glial fibrillary acidic protein (GFAP) and inducible nitric oxide synthase (iNOS), in conjunction with multiple Abs targeting C. pneumoniae, permitted identification of the specific infected cell-type found in areas demonstrating classical AD neuropathology (e.g., neuritic senile plaques)(Balin et al., 1998). This demonstration of infection by a respiratory pathogen in the AD brain poses a number of important questions regarding infection in this condition, not the least of which is the status of the organism in terms of its persistency in areas of AD neuropathology. One clue that is helpful in addressing the issue of persistency/chronicity of the infection is a morphological characterization of C. pneumoniae in this setting.
As with all members of the genus Chlamydia, C. pneumoniae exhibits two distinct morphologies during its developmental cycle: the EB and the RB (Miyashita et al., 1993). Appropriate host cells are infected by the small, extracellular, metabolically inactive form of the organism, the EB. C. pneumoniae EBs are approximately 0.2–1.2 μm in diameter, are often pear-shaped with a clear periplasmic space, and contain a condensed matrix of protein and nucleic acids (Miyashita et al., 1993). It is currently thought that EBs bind the prospective host cell via a glycosaminoglycan moiety in conjunction with an unidentified surface protein which is thought to be the product of the ompA gene (Matsumoto, 1988). Once bound, the bacterium undergoes endocytosis and is brought into a membrane-bound cytoplasmic vesicle, where it undergoes a developmental process resulting in the larger, vegetative growth form of the organism, the RB (Kuo et al., 1995). RBs undertake extremely active DNA, RNA, and protein synthesis (Ward, 1988), and each RB undergoes 7–8 mitotic divisions before final de-differentiation back to the EB form (Matsumoto, 1988). Newly-formed EBs exit the host cell following cell lysis, or via exocytosis (Matsumoto, 1988). Alternatively, the organisms may remain viable within host cells over extended periods of time, as described for C. trachomatis infected synovial tissues of patients with reactive arthritis and Reiter's syndrome (Nanagara et al., 1995). In such chronic infections, the chlamydial organisms show aberrant morphology while maintaining metabolic activity and viability (Gerard et al., 1998, Nanagara et al., 1995). In order to more fully define the morphological characteristics of C. pneumoniae during infection of the AD brain, we analyzed the organism in numerous AD affected and non-AD control cases. Utilizing both electron and immunoelectron microscopic techniques (EM, IEM respectively), we compared the morphology of the brain-derived organism with that of the typical laboratory strain of C. pneumoniae designated TW-183. The existence of several morphological chlamydial forms in the Alzheimer brain may be indicative of chronic infection, which may in turn contribute importantly to the pathogenesis of this neurodegenerative disease.
Materials and Methods
Tissue Samples
Postmortem tissue samples from various brain regions of patients with or without AD were obtained from the Harvard Brain Tissue Resource Center and from the MCP-Hahnemann University Hospital, Center City, Department of Pathology. All samples from patients indicated as having AD were confirmed histologically using standard accepted criteria for the disease (NINDS/CERAD)(Mirra et al., 1991). Samples from control (non-AD) patients were age matched to those of AD patients, and each set of such samples was examined histologically for senile plaques (SP), and neurofibrillary tangles (NFT) and confirmed as non-AD control brains.
We examined four AD brains, previously confirmed to be positive for C. pneumoniae by PCR and/or immunohistochemical analyses, and four non-AD brains previously confirmed to be negative for the organism by equivalent techniques (Balin et al., 1998). The mean age of death was 75 yr for AD patients and 74 yr for non-AD control patients. Mean post-mortem intervals were 15 hr and 7 hr for AD and control brains, respectively. Causes of death included cardiac arrest, pneumonia, lung carcinoma, and aspiration pneumonia for the AD patients, and respiratory failure, renal failure, aortic aneurysm, and adenocarcinoma for the non-AD, control individuals.
Electron and Immunoelectron Microscopy
Tissue sections from AD and control brains were cut from areas typically demonstrating neuropathology (e.g. hippocampus, temporal cortex) and from areas usually spared from AD pathology (e.g. cerebellum). These were prepared for EM and IEM analyses by fixation in 4% paraformaldehyde in 0.1M phosphate buffered saline (PBS) and osmication in 1% osmium tetroxide in PBS buffer prior to embedding in Epon-812 (EM Sciences, Fort Washington, PA). Tissue sections were cut on a Sorvall MT2B ultramicrotome with a Diatome diamond knife. These sections were etched for 60–75 sec with 7% hydrogen peroxide and immediately washed with filtered water. The sections were blocked with cold water fish gelatin (CWFG; 0.1% in PBS, pH 7.2) for 5 min prior to a 30 min incubation with the mAb specific for a C. pneumoniae outer membrane protein (DAKO RR402, Carpinteria, CA). Subsequently, the sections were blocked for 20 min prior to a 20 min incubation with an anti-mouse specific secondary Ab conjugated to 5 nm or 15 ran colloidal gold (Amersham, Arlington Heights, IL). Controls for immunolabeling used only secondary Ab or nonspecific primary Ab, such as anti-CD54 which recognizes intercellular cell adhesion molecule type I (ICAM-I; PharMingen, San Diego, CA).
Ultrastructural Analysis of C. pneumoniae from Infected THP-1 cells
The protocol for negative staining IEM consisted of adsorbing 5–10 μl of lysed THP-1 cells (ATCC) previously infected for 7–10 days with C. pneumoniae from AD homogenates onto carbon-coated copper EM grids (EM Sciences) followed by washing with PBS. The grids were blocked with 0.1% CWFG in PBS for 10 min prior to a 15–30 min incubation in the primary Ab, anti-OMP (DAKO) diluted as above. Grids were blocked again in 0.1% CWFG prior to a brief rinse in PBS, and subsequently incubated for 30 min in anti-mouse secondary Ab (1.0 mg/ml conjugated to 5 nm colloidal gold particles; Amersham). Controls were incubated solely in anti-mouse gold-conjugated secondary Ab. Prior to negative staining the grids with 2.0% aqueous uranyl acetate, grids were rinsed briefly in double-distilled H2O. Specimens were examined at 60–80 kV accelerating voltage on a Zeiss EM-10 electron microscope.
Preparation of Nucleic Acids
Nucleic acids were prepared from tissue samples as described (Branigan et al., 1996); 1 μg nucleic acid was used in the PCR assay. EBs of C. pneumoniae strain TW-183 were obtained from the ATCC, and DNA was prepared from them for control amplifications. PCR assays to screen for C. pneumoniae chromosomal DNA utilized a nested primer system targeting the major outer membrane protein gene (ompA) of C. pneumoniae (Melgosa Perez et al., 1991). Primers, synthesized by Genosys US, were designed using GeneRunner® software (Hastings Software, Hastings NY) and were derived from bases 26–43 (plus strand) and 567–548 (minus strand) external and bases 115–135 (plus strand) and 462–444 (minus strand) internal of the C. pneumoniae coding sequence (Melgosa-Perez et al., 1991). The primers were analyzed for sequence specificity via “Blast” comparison with all existing DNA sequences in the GenBank database. Cycling for PCR assays was done in a Perkin-Elmer 2400 thermocycler. External primer PCR conditions were 5 min at 94°C, and 35 cycles with 30 sec at 94°C, 30 sec at 52 °C, and 30 sec at 72 °C, and terminated with 5 min at 72 °C, Internal primer conditions were identical except that the annealing temperatures were ramped by 1°C for 30 sec each beginning at 59 °C to 50 °C. Products were analyzed on 1.8% agarose gels. This system produces an amplification product of 350 bp; these were ligated into an M13 vector and sequenced on an automated sequencer ABI Prizm model 377 V3.0.
Culture analyses for C. pneumoniae
Culture analyses for C. pneumoniae were done by infection of a human monocyte/macrophage cell line (THP-1; ATCC). Approximately 0.5 g of frozen brain tissue from the temporal cortex of two patients previously determined to be C. pneumoniae-positive by PCR and immunohistochemical analyses, as well as congruent tissue samples from two control brains, were subjected to three freeze-thaw cycles; tissues were homogenized and sonicated between cycles. From these samples, 100 μl of brain homogenate were mixed with 3 × 106 THP-1 cells in 2 ml of 10% fetal calf serum in RPMI-1640 (Sigma), 0.5% HEPES buffer (pH 7.0). The homogenate and cells were centrifuged at 500 × g for 30 min, diluted to 10 ml vol in fresh medium, then cultured for 72 hr. Following this initial culture incubation, 1 ml of culture supernatant was used for a second passage on 3 × 106 fresh THP-1 cells; remaining cells from the initial 72 hr incubation were separated from debris by centrifugation on Ficoll-Paque (Pharmacia-Biotech, Piscataway NJ) gradients. Cells from both first and second culture passages were subjected to immunocytochemical analyses to confirm infection or non-infection of the cultured THP-1 cells. Cells for immunocytochemistry were diluted to 2.5 × 105/ml in Hanks balanced salt solution, and 0.2 ml were cytospun onto ProbeOn+ slides (Fisher Biotech, Pittsburgh, PA) at 500 × g for 5 min, using a Shandon Cytocentrifuge III. Cells were fixed for at least 1 hr at room temperature with Streck Tissue Fixative (S.T.F.™ Streck Laboratories Inc., Omaha, NE), followed by a rinse with 1× Automation buffer (Biømeda, Foster City, CA). After endogenous peroxidase deactivation, immunocytochemistry was performed using the Biostain Super ABC Mouse/Rat kit (Biømeda) as described by the manufacturer. Anti-surface protein mAb specific for C. pneumoniae (anti-OMP, RR402; DAKO) and the genus-specific anti-LPS mAb (DAKO, IMAGEN-Chlamydia Kit) were each diluted 1: 10 in Ab Diluting buffer (Biømeda) and incubated for 30 min at 37°C, followed by nine rinses with Automation buffer (Biømeda). Similar incubations and washes were performed for the secondary anti-mouse biotinylated Ab and avidin-biotin conjugated (ABC) reagents. After a 1 min incubation with peroxidase enhancer, Ab binding was detected with the horseradish-peroxidase (HRP) conjugated chromagen Diaminobenzidine Cobalt (Biømeda) by 5–10 min incubation at room temperature. Slides were prepared with Crystal-Mount (Biømeda) for light microscopy.
Results
In an earlier communication (Balin et al., 1998), we reported immunohistochemical, PCR, and other analyses on tissues from the same brains examined in the present study. We used standard EM here to confirm and extend those initial studies by further and more detailed examination of samples from four AD and four control brains. As mentioned above, C. pneumoniae alternates between two distinct morphological forms in its life cycle (Miyashita et al., 1993): the EB and the RB. C. pneumoniae exhibited both forms of the organism in the AD brain (Figure 1). These forms were found in areas of AD brain showing typical AD neuropathology, i.e., temporal cortex and hippocampus. The cerebella of these AD brains were negative for the organism. EBs showed a dense central core and, at times, a clear periplasmic space. This form had a trilaminar membrane, ranged in size from 0.2–1.0 μm in diameter and occasionally was localized within an inclusional membrane. RBs were spherical and 0.5–1.2 μm in diameter. RBs alternated between forms exhibiting a dense, dark center and those exhibiting a mottled interior with a lighter, less electron-dense center (Figure 1A). RBs were also identified in various stages of binary fission and exhibited division planes that appeared to divide the organism into two halves, each containing a granular cytoplasm and a less dense center (Figure 1 B). Interestingly, some RBs also contained amorphous material of variable size within individual organisms; these structures may represent condensed genetic material consistent with plasmids, as has been noted for other chlamydial species (Storey et al., 1993). However, at this point, identification of plasmids in Chlamydia pneumoniae is inconclusive.
FIGURE 1.
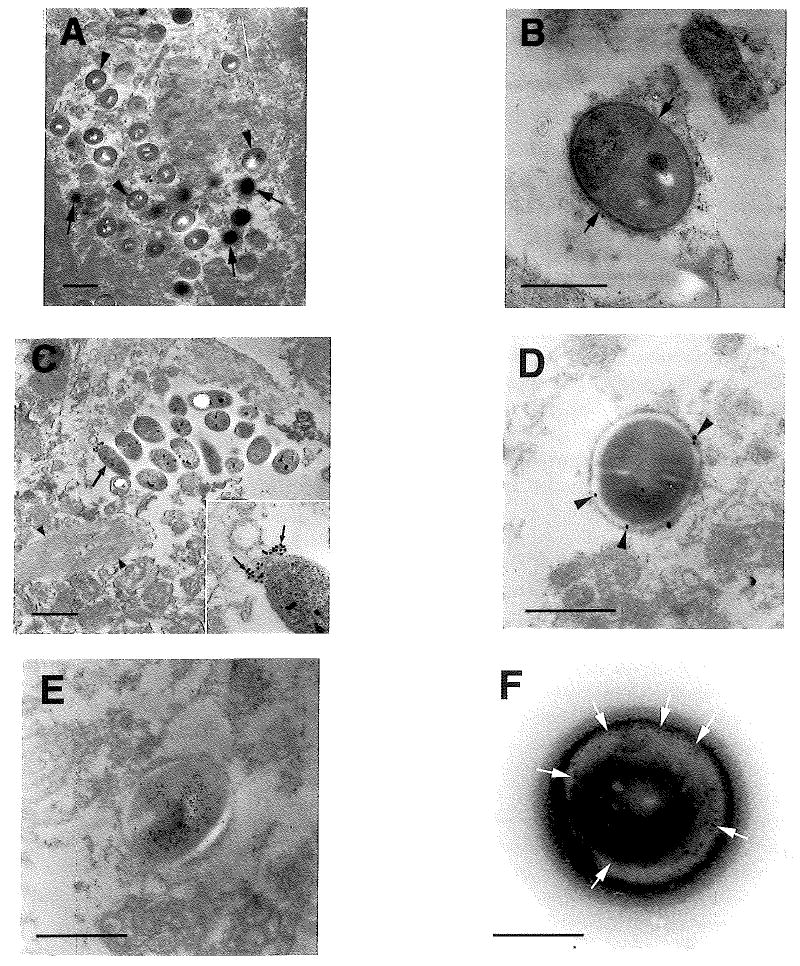
Ultrastructural analyses of brain tissues from AD patients to characterize the morphologies of C. pneumoniae. AD patient samples were prepared for electron microscopy and IEM as described in Methods; the anti-chlamydial Ab used was a mAb against a C. pneumoniae outer membrane protein (see text). Panel A: Multiple EBs (arrows) and RBs (arrowheads) forms of C. pneumoniae are apparent in the hippocampus of an AD brain. Panel B: Extracellular area of the same region in Panel A showing a dividing reticulate body undergoing binary fission and exhibiting a distinct division plane (arrows). Panel C: Within the temporal cortex of an AD brain, alternate phenotypes of Chlamydia pneumoniae (large arrow) are present; these forms were apparent in numerous AD brains. IEM analysis with the anti-OMP mAB shows polar labeling on one end only (inset, small arrows), but not all organisms are immunoreactive. A neurofibrillary tangle bundle can be seen proximal to the organisms (arrowheads). Panel D: IEM of C. pneumoniae labeled with anti-OMP mAb (arrowheads) within the hippocampus from an AD brain demonstrates circumferential immunolabeling. Panel E: Equivalent area to that shown in D exhibiting an organism with central immunolabeling by anti-OMP mAb. Panel F: IEM (arrows) using the anti-OMP mAb targeting C. pneumoniae isolated from THP-1 cells following infection with AD brain homogenate. Bars: A, C = 1 μm; B, D, E = 0.5 μm; F = 0.25 μm
In addition to the typical morphologies of chlamydial EBs and RBs, some AD samples contained organisms displaying atypical morphology (Figure 1C) These were either spherical or elongated and had a variable size of 0.2–1.2 μm in diameter, again fitting well within the size range typical of C. pneumoniae (Miyashita et al., 1993, Popov et al., 1991). These particular organisms, while clearly bacteria, exhibited a less distinct cell membrane when compared to the organisms previously described (see Figure 1A and B). They had a granular center which contained one or more possible nucleic acid dense bodies. The smaller, spherical organisms demonstrating this phenotype also appeared to be undergoing binary fission, yet did not exhibit definable division planes (data not shown).
Using a mAb against a Chlamydia pneumoniae OMP, we used IEM to confirm further the identity of the organism located in the AD brains. The atypical organisms immunolabeled in a polar manner, yet not every atypical organism was immunoreactive (Figure 1 C). Typical RBs exhibited two alternate immunolabeling profiles. Some were circumferentially labeled with the gold-conjugated secondary Ab bound to the cell surface (Figure 1 D), while others were immunolabeled over their central regions (Figure 1 E) probably as a result of the sectioning process and the specific expression of the OMP protein within and extending from the bacterium.
Using the same mAbs as above, we employed IEM to analyze C. pneumoniae pellets obtained from lysed THP-1 cells following 7–10 d infection with C. pneumoniae (Figure 1 F). In this context, negatively stained organisms displayed a spherical morphology which labeled in a manner analogous to that of bacteria observed in the AD brains. Central labeling occurred with both the anti-LPS and the anti-OMP mAbs. This further strengthens the identification of the brain-extracted organisms as Chlamydia pneumoniae.
C. pneumoniae were not only observed within glial cells and tissue parenchyma, but also were seen within monocyte/macrophage-like cells in small blood vessels in the brain and within presumptive pericytes and perivascular macrophages (Figure 2). This observation is consistent with our previously reported immunohistochemical findings (Balin et al., 1998) demonstrating immunolabeling for C. pneumoniae around small and mid-size blood vessels in the AD brain.
FIGURE 2.
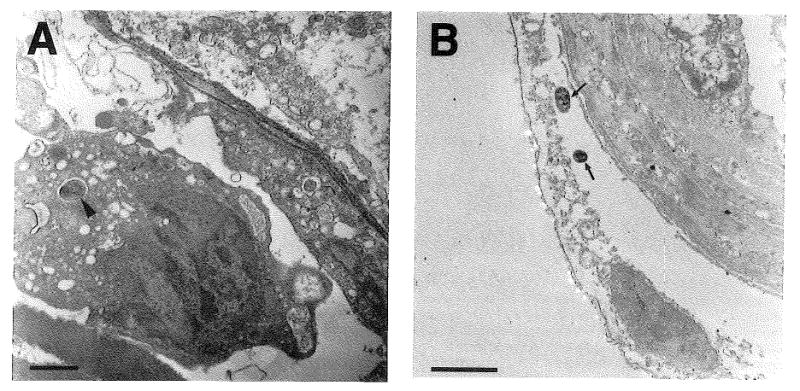
Analysis of C. pneumoniae-infected cells in AD blood vessels. Panel A: Blood vessel from the hippocampus from an AD brain showing an RB (arrowhead) within the cytoplasm of a presumptive monocyte/macrophage in the vessel lumen. Panel B: Blood vessel from the temporal cortex from an AD brain demonstrating two C. pneumoniae organisms (arrows) within the disrupted cytoplasm of a detached pericyte or perivascular macrophage on the abluminal surface of the vessel. Bars= 1 μm
Incubation of homogenized AD brain tissue with THP-1 cells revealed that C. pneumoniae was not only present within the samples, but also still viable. The THP-1 monocyte/macrophage cell line can act as a host cell for the organism isolated from the AD brain (Figure 3A). Immunocytochemical analyses using the mAb against an OMP of C. pneumoniae produced punctate labeling of the organism within the cytoplasm of infected THP-1 cells. Ultrastructural analysis revealed that the organisms had thickened cell surfaces, showed variable size, and were found within individual membranous structures within host cells (Figure 3B).
FIGURE 3.

Analysis of C. pneumoniae from AD brain homogenates. Panel A: THP-1 cells after infection with a homogenate from the temporal cortex from an AD brain 7 d post-incubation. Immunohistochemical analysis with an anti-OMP mAb shows organisms (arrows) within the cytoplasm of the cells. Bar=20 μm. Panel B: Ultrastructural analysis of infected THP-1 cells with C. pneumoniae from an AD brain homogenate 7 d post-incubation. Organisms demonstrating a more aberrant morphology can be seen in the cytoplasm of the host cell (arrow). The EBs and RBs show thickened cell walls and are variable in size. Since the bacteria are dependent on the host-cell's ATP, C. pneumoniae bodies are situated adjacent to mitochondria (arrowhead) which shows morphology indicative of mitochondria undergoing oxidative stress (Parker et al., 1990). Bar=1 μm. Panel C: Cloned 350 bp PCR product from the infected THP-1 cells shown in the previous two panels. (Left) Lane 1:100 bp standards. Lane 2: Uncut ligated M13 vector. Lane 3: Restriction digest of vector in lane 2; the lower band was sequenced and found to be Homologous to the C. pneumoniae omp A gene
To further confirm the identity of the organisms observed in the above studies, we extracted total DNA from the infected AD brain tissues, and from THP-1 cells infected with the organism isolated from AD brains, and we performed PCR analysis using previously published primers (Melgosa-Perez et al., 1991) for the C. pneumoniae ompA gene (Figure 3C). Following amplification, we cloned several of the PCR products from both the AD tissues and the THP-1 infected cells, determined the DNA sequence of those products, and compared those sequences against the non-redundant GenBank databases accession number m64064 (Melgosa-Perez et al., 1991). Our sequence showed 100% homology with bases 115–462 of the ompA gene which codes for a Chlamydia pneumoniae major outer membrane protein (Melgosa-Perez et al., 1991). Thus, the PCR product represented a significant portion of the ompA gene of Chlamydia pneumoniae. Both PCR products from the brain tissue and the infected THP-1 cells were identical to the Genbank sequences for the C. pneumoniae ompA gene.
Discussion
In studies reported here, we delineate the morphology of C. pneumoniae in situ during infection of the AD brain. We demonstrate the presence of typical and atypical morphologies of this organism in that context, in both EB and RB forms. The primary morphological variability was exhibited in overall shape and size of the organisms, consistent with an earlier EM study of the morphology of C. pneumoniae in another context (Carter et al., 1991); however, all morphologic forms were similar in immunoreactivity to C. pneumoniae laboratory strains as demonstrated by others (Christiansen et al., 1997). We identified the organism relatively rarely within identifiable membrane bound vacuoles, probably due to fixation of postmortem tissues. We also prepared total DNA from a cultured cell line infected with the organism isolated from the affected AD tissues; using that DNA and primers targeting the C. pneumoniae ompA gene, we amplified a product of appropriate size, which was cloned and sequenced. This PCR product was identifiable as a portion of the ompA gene, independently supporting the identity of the organism under study.
Localization of C. pneumoniae in the AD brain presents a number of possibilities regarding the organism's morphological form in that tissue. Since sporadic AD predominantly affects aged individuals, and because it is not possible to know precisely when any given individual was initially infected with the organism, either a chronic or an acute infection is possible in each patient. Previous studies have demonstrated that Chlamydia trachomatis bacteria involved in persistent infection are characterized by significantly altered morphologic and biochemical features (Beatty et al., 1995b). Medications (e.g. antibiotics and anti-inflammatory drugs) prescribed to older individuals may affect the persistent, and therefore the phenotypic, nature of the organism (Lorian, 1993, Matsumoto and Manire, 1970). Moreover, a host proinflammatory response following infection may result in pleiomorphism of C. pneumoniae, due to evoked bacteriostatic events such as free radical damage in the tissues and/or nutrient starvation of the organism (Igietseme, 1996).
The AD tissues examined included intracellular and extracellular organisms, with a number of bacteria identified in a state of binary fission. This observation suggests that C. pneumoniae engages in active infection of host cells in the brain, a situation that would result in the activation of astroglia/microglia; such activated cells have been described in the AD brain (Itagaki et al., 1989). Activation of neuroglia by C. pneumoniae in turn may suggest that the organism plays a role in AD pathogenesis. Multiple EBs and RBs were found not only within glial cells/neuronal tissue, but also in macrophages and presumptive pericytes. This disposition may indicate multiple modalities of infection. First, the bacterium may spread hematogenously either in the EB form or within monocytes/macrophages to invade endothelial and/or perivascular cells. Stasis due to vessel damage, ischemia, or presence of atherosclerotic plaques within the lumen of cerebral vessels may allow the organism to traverse the blood-brain barrier. Moreover, if active infection is the case in some anatomic context, whether chronic or acute, cells of the monocyte/macrophage lineage may spread scavenged organisms within the blood stream to susceptible areas, including the brain (Leinonen, 1993).
In our earlier study (Balin et al., 1998), we identified C pneumoniae in AD brain glial cells, and in two cultured cell lines within defined membranous structures; in some cases, we found organisms that appeared to be non-membrane bound within the host cell cytoplasm. Although the latter situation may result from compromised fixed tissue, both settings are consistent with previously published data (Miyashita et al., 1993, Ward, 1988). Since free EBs can infect cell lines when incubated in culture media without necessitating prior infection of carrier cells, glial cell infection within the brain parenchyma could be explained by the release of EBs from areas of active infection, such as atherosclerotic lesions, thereby allowing invasion of the surrounding brain tissues and cells. Although recent reports have begun to examine systemic dissemination of C. pneumoniae (Moazed et al., 1998), further investigation is required to define the extent to which other tissues in the Alzheimer's individual are infected with C. pneumoniae.
Previous studies have suggested a role for interferon-γ (IFN-γ) in the aberrant functioning of chlamydia (Ward, 1998). If grown in the presence of this compound, growth inhibition results, chlamydial differentiation is inhibited in vitro, and unusual intracellular morphologic forms develop. This growth inhibition results from induction of indoleamine-2,3-dioxygenase (IDO), which deprives the pathogen of tryptophan by degrading this amino acid. In vitro studies have shown that tryptophan deprivation leads to the formation of large, aberrantly-shaped chlamydial cells, representing an interrupted stage in the normal bacterial life cycle (Mehta et al., 1998). In addition, IFN-γ causes a selective down-regulation of bacterial LPS synthesis and of expression for several chlamydial genes, including ompA (Beatty et al., 1995b). OMP is a major structural component of all Chlamydiae, and both OMP and LPS are integral to the normal functioning of the organism (Miyashita et al., 1993). Attenuation of LPS synthesis, as well as decreased production of OMP may account for the modified chlamydial morphology observed in AD tissues, as well as for differential immunolabeling of organisms; i.e., bacteria expressing LPS or OMP will bind specific Ab, while organisms not expressing them will not display immunoreactivity. The immunolabeling pattern identified here is consistent with this explanation. Furthermore, the distribution of immunogold labeling of the organisms indicates both a typical expression pattern of OMP for some organisms as well as atypical dispositions on the more variant forms of the organism. Perhaps, long-term infection of the AD brain with C. pneumoniae, in which chronic cellular damage is apparent, reflects the morphologic variation observed for the infecting organism. Secretion of proinflammatory cytokines by infected cells may promote development of pleiomorphic forms via potential ionic fluxes, membrane lipid damage, and production of free oxygen radicals (Engelborghs and DeDeyn, 1997, Malinverni, 1996). Importantly, aberrant forms of C. trachomatis have been identified in vivo in synovial tissues of patients with reactive arthritis (Nanagara et al., 1995) and in various in vitro systems demonstrating chronic infection (Beatty et al., 1995a, Kohler et al., 1997). It will be intriguing to determine if C. pneumoniae infection in the AD brain results in attenuation of OMP and LPS synthesis in a manner similar to that seen in reactive arthritis with C. trachomatis.
In summary, we show that Chlamydia pneumoniae is present and active in the Alzheimer's brain, and in this context the organism often shows highly pleiomorphic morphology. The multiple forms of the bacteria and their localization within macrophages and glial cells may be indicative of chronic/persistent infection, as has been demonstrated in other chlamydia-associated diseases (Gerard et al., 1998, Malinverni et al., 1995, Nanagara et al., 1995). Our observations, in conjunction with C. pneumoniae's ability to infect cell lines in vitro following extraction from the AD brain, have important pathophysiologic implications for its role in AD. Ongoing studies in this group are addressing specific mechanisms of persistence that may facilitate the pleiomorphism of the organism and stimulate eukaryotic cellular changes consistent with AD neuropathology.
Acknowledgments
We thank the MCP-Hahnemann University School of Medicine, Department of Pathology and Laboratory Medicine for departmental funds and the Allegheny Health, Education, and Research Foundation, Neurosciences Research Institute for their support and funding for this project, and the Harvard Brain Tissue Resource Center (supported in part by PHS grant MH/NS31862). In addition, some of this study was supported by NIH grant AR-42541 (A.P.H.). Finally, we thank the families of all the Alzheimer's Disease individuals for their donations of tissues to the research efforts, without which, the work presented herein would not have been possible.
References
- Balin BJ, Gerard HC, Arking EJ, Appelt DM, Branigan PJ, Abrams JT, Whittum-Hudson JA, Hudson AP. Identification and Localization of Chlamydia pneumoniae in the Alzheimer's Brain. Med Microbiol Immunol. 1998;187:23–42. doi: 10.1007/s004300050071. [DOI] [PubMed] [Google Scholar]
- Beatty WL, Morrison RP, Byrne GI. Persistent Chlamydiae, from cell culture to a paradigm for chlamydial pathogenesis. Microbiol Reviews. 1995a;58:685–699. doi: 10.1128/mr.58.4.686-699.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beatty WL, Morrison RP, Byrne GI. Reactivation of Chlamydia trachomatis Infection in Cell Culture. Inject Immun. 1995b;63:199–205. doi: 10.1128/iai.63.1.199-205.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Branigan PJ, Gerard HC, Hudson AP, Schumacher HR. Comparison of synovial tissue and synovial fluid as a source of nucleic acids for detection of Chlamydia trachomatis by polymerase chain reaction. Arthritis Rheumat. 1996;39:1740–1746. doi: 10.1002/art.1780391018. [DOI] [PubMed] [Google Scholar]
- Carter MW, Al-Mahdawi SAH, Giles IG, Treharne JD, Ward ME, Clarke IN. Nucleotide sequence and taxonomic value of the major outer membrane protein gene of Chlamydia pneumoniae IOL-207. J Gen Microbiol. 1991;137:465–475. doi: 10.1099/00221287-137-3-465. [DOI] [PubMed] [Google Scholar]
- Christiansen G, Ostergaard L, Birkelund S. Molecular biology of the Chlamydia pneumoniae surface. Scand J Inject Dis Suppl. 1997;104:5–10. [PubMed] [Google Scholar]
- Engelborghs S, De Deyn Pp. The neurochemistry of Alzheimer's disease. Acta Neurologica Belgica. 1997;97:67–84. [PubMed] [Google Scholar]
- Gerard HC, Branigan PJ, Schumacher HR, Hudson AP. Synovial Chlamydia trachomatis in patients with reactive arthritis/Reiter's syndrome are viable but show aberrant gene expression. J Rheumatol. 1998;25:734–742. [PubMed] [Google Scholar]
- Igietseme JU. Molecular mechanism of T-cell control of Chlamydia in mice: role of nitic oxide in vivo. Immunol. 1996;88:1–5. [PMC free article] [PubMed] [Google Scholar]
- Itagaki S, McGeer PI, Akiyama H, Zhu S, Selkoe D. Relationship of microglia and astrocytes to amyloid deposits of Alzheimer disease. J Neuroimmunol. 1989;24:173–182. doi: 10.1016/0165-5728(89)90115-x. [DOI] [PubMed] [Google Scholar]
- Kohler L, Ott N, Netteinbreker E, Gerard HC, Drommer W, Hudson AP, Zeidler H. Ultrastructural analysis of the persistence of Chlamydia trachomatis serovar K in human monocytes. Microb Pathogen. 1997;22:133–142. doi: 10.1006/mpat.1996.0103. [DOI] [PubMed] [Google Scholar]
- Kuo CC, Jackson LA, Campbell LA, Grayston JT. Chlamydia pneumoniae (TWAR) Clin Microbiol Review. 1995;8:451–61. doi: 10.1128/cmr.8.4.451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leinonen M. Pathogenetic mechanisms and epidemiology of Chlamydia pneumoniae. Europ Heart J. 1993;14:57–61. [PubMed] [Google Scholar]
- Lorian V. Medical Relevance of Low Concentrations of Antibiotics. J Antimicrob Chemoth. 1993;31 D:137–148. doi: 10.1093/jac/31.suppl_d.137. [DOI] [PubMed] [Google Scholar]
- Malinverni R. The role of cytokines in chlamydial infection. Curr Opin Inject Dis. 1996;9:150–156. [Google Scholar]
- Malinverni R, Kuo CC, Campbell LA, Lee A, Grayston JT. Effects of two antibiotic regimens on course and persistence of experimental Chlamydia pneumoniae TWAR pneumonitis. Antimicrob Agents & Chemoth. 1995;39:45–9. doi: 10.1128/aac.39.1.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumoto A. Structural Characteristics of Chlamydial Bodies. In: Barron AL, editor. Microbiology of Chlamydia. CRC Press; 1988. pp. 21–46. [Google Scholar]
- Matsumoto A, Manire GP. Electron Microscopic Observations on the Effects of Penicillin on Chlamydia psittaci. J Bacteriol. 1970;101:278–285. doi: 10.1128/jb.101.1.278-285.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehta SJ, Miller RD, Ramirez JA, Summersgill JT. Inhibition of Chlamydia pneumoniae Replication in HEp-2 Cells by Interferon-Gamma: Role of Tryptophan Catabolism. J Infect Dis. 1998;177:1326–31. doi: 10.1086/515287. [DOI] [PubMed] [Google Scholar]
- Melgosa-Perez PM, Kuo CC, Campbell LA. Sequence analysis of the major outer membrane protein gene of Chlamydia pneumoniae. Infect Immun. 1991;59:2195–2199. doi: 10.1128/iai.59.6.2195-2199.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirra SS, Heyman A, McKeel D. Consortium to establish a registry for Alzheimer's disease (CERAD). Part II. Standardization of the neuropathologic assessment of Alzheimer's disease. Neurol. 1991;41:479–486. doi: 10.1212/wnl.41.4.479. [DOI] [PubMed] [Google Scholar]
- Miyashita N, Kanamoto Y, Matsumoto A. The morphology of Chlamydia pneumoniae. J Med Microbiol. 1993;38:418–25. doi: 10.1099/00222615-38-6-418. [DOI] [PubMed] [Google Scholar]
- Moazed TC, Kuo CC, Grayston JT, Campbell LA. Evidence of Systemic Dissemination of Chlamydia pneumoniae via Macrophages in the Mouse. J Infect Dis. 1998;177:1322–1325. doi: 10.1086/515280. [DOI] [PubMed] [Google Scholar]
- Nanagara R, Li F, Beutler AM, Hudson AP, Schumacher HR. Alteration of Chlamydia trachomatis biological behavior in synovial membranes: suppression of surface antigen production in reactive arthritis and Reiter's syndrome. Arthritis Rheumat. 1995;38:1410–1417. doi: 10.1002/art.1780381008. [DOI] [PubMed] [Google Scholar]
- Parker WD, Jr, Filley CM, Parks JK. Cytochrome Oxidase Deficiency in AD. Neurol. 1990;40:1302–1303. doi: 10.1212/wnl.40.8.1302. [DOI] [PubMed] [Google Scholar]
- Popov VL, Shatkin AA, Pankratova VN, Smirnova NS, von Bonsdorff CH, Ekman MR, Morttinen A, Saikku P. Ultrastructure of Chlamydia pneumoniae in cell culture. FEMS Microbiol Lett. 1991;68:129–34. doi: 10.1016/0378-1097(91)90115-q. [DOI] [PubMed] [Google Scholar]
- Storey C, Lusher M, Yates P, Richmond S. Evidence for Chlamydia pneumoniae of non-human origin. J Gen Microbiol. 1993;139:2621–6. doi: 10.1099/00221287-139-11-2621. [DOI] [PubMed] [Google Scholar]
- Ward ME. The Chlamydial Developmental Cycle. In: Barron AL, editor. Microbiology of Chlamydia. CRC Press; 1988. pp. 71–95. [Google Scholar]


