Addition of reversine to dividing cells ejects Mad1 and the RZZ complex from unattached kinetochores and prevents resolution of incorrect chromosome–microtubule attachments (see also related papers by Hewitt et al. and Maciejowski et al. in this issue).
Abstract
The catalytic activity of the MPS1 kinase is crucial for the spindle assembly checkpoint and for chromosome biorientation on the mitotic spindle. We report that the small molecule reversine is a potent mitotic inhibitor of MPS1. Reversine inhibits the spindle assembly checkpoint in a dose-dependent manner. Its addition to mitotic HeLa cells causes the ejection of Mad1 and the ROD–ZWILCH–ZW10 complex, both of which are important for the spindle checkpoint, from unattached kinetochores. By using reversine, we also demonstrate that MPS1 is required for the correction of improper chromosome–microtubule attachments. We provide evidence that MPS1 acts downstream from the AURORA B kinase, another crucial component of the error correction pathway. Our experiments describe a very useful tool to interfere with MPS1 activity in human cells. They also shed light on the relationship between the error correction pathway and the spindle checkpoint and suggest that these processes are coregulated and are likely to share at least a subset of their catalytic machinery.
Introduction
At each mitosis, cells face the tremendous challenge of separating the sister chromatids in two identical pools. This process, on which all cells rely to remain viable, is usually executed with great accuracy. Its perturbation results in aberrations in chromosome numbers (aneuploidies), which are a cause of disease and correlate with cellular transformation (Weaver and Cleveland, 2006).
Fidelity of cell division is the result of feedback controls. The first control mechanism halts the process of cell division and imposes a mitotic arrest when chromosome–microtubule attachment is perturbed in different ways (for review see Rieder and Palazzo, 1992). This ability of eukaryotic cells activates a checkpoint (for reviews see Hartwell and Weinert, 1989; McIntosh, 1991; Rieder and Palazzo, 1992), generally known as the spindle assembly checkpoint (for review see Musacchio and Salmon, 2007) and herewith often abbreviated as spindle checkpoint or simply checkpoint. The checkpoint cannot be satisfied under conditions that perturb chromosome–microtubule attachment, most typically the depolymerization of microtubules (lack of attachment). In humans, spindle checkpoint components include enzymes such as the BUB1, BUBR1, MPS1, and PRP4 kinases and protein–protein interaction devices such as BUB3, MAD1, MAD2, and the three-subunit ROD–ZWILCH–ZW10 (RZZ) complex (for review see Musacchio and Salmon, 2007).
During prometaphase, the checkpoint proteins are recruited to unattached kinetochores, which are large protein assemblies built on chromosomal loci known as centromeres (Cleveland et al., 2003). An ∼550-kD, 10-subunit assembly, the KMN network (from the initials of its subcomplexes the Knl1, Mis12, and Ndc80 complexes), provides the microtubule-binding core of the outer kinetochore (Cheeseman et al., 2006; for review see Santaguida and Musacchio, 2009). Kinetochore recruitment of the checkpoint proteins is an obligatory condition for sustained checkpoint signaling. Its impairment invariably leads to a failure in the checkpoint response (for examples and discussions see Meraldi et al., 2004).
Spindle checkpoint activity converges on the generation of an anaphase-promoting complex/cyclosome (APC/C) inhibitor known as the mitotic checkpoint complex (for review see Musacchio and Salmon, 2007). Mad2, BubR1, and Bub3 contribute in different ways to the formation of the mitotic checkpoint complex. Cdc20, the target of the checkpoint proteins in the mitotic checkpoint complex, is a positive regulator of the APC/C, an ubiquitin-ligase whose activity is required for progression into anaphase. By inhibiting Cdc20, the spindle checkpoint prevents APC/C activation toward crucial substrates for anaphase such as Cyclin B and Securin and, consequently, mitotic exit (for review see Musacchio and Salmon, 2007).
The second control mechanism, generally referred to as error correction, prevents the stabilization of kinetochore–microtubule attachments until they come under tension (Nicklas and Koch, 1969; Li and Nicklas, 1995). Improper kinetochore–microtubule attachments such as merotelic or syntelic attachments are probably distinguished from proper attachments (amphitelic attachment or biorientation) and corrected because they are not under full tension. The molecular basis of stabilization or destabilization of improper attachments is being actively investigated. The first protein to become clearly implicated in this process was the AURORA B kinase (for review see Ruchaud et al., 2007). AURORA B is a member of the AURORA family of S/T kinases, which also includes the ubiquitously expressed AURORA A, which is involved in spindle bipolarization, and AURORA C, whose role is poorly understood but likely limited to meiosis and early development (for review see Ruchaud et al., 2007). AURORA B is part of the chromosome passenger complex, whose subunits also include INCENP, SURVIVIN, and BOREALIN (for review see Ruchaud et al., 2007). Inactivation of Ipl1, the only AURORA kinase in Saccharomyces cerevisiae, leads to the stabilization of syntelic attachments, implicating Ipl1 in their correction (Tanaka et al., 2002). In vertebrates, inhibition of AURORA B by small molecules or RNAi leads to the accumulation of merotelic and syntelic attachments (Ditchfield et al., 2003; Hauf et al., 2003; Lampson et al., 2004; Cimini et al., 2006; Knowlton et al., 2006). The regulation of microtubule-destabilizing enzymes known as MCAK (mitotic centromere–associated kinase) and KIF2B by AURORA B may be important for correction (for reviews see Pinsky and Biggins, 2005; Vader and Lens, 2008; Kelly and Funabiki, 2009). Furthermore, AURORA B phosphorylates NDC80, a subunit of the KMN network, on at least six to eight sites near the microtubule-binding interface, causing a strong decrease of microtubule-binding affinity (Cheeseman et al., 2006; DeLuca et al., 2006; Ciferri et al., 2008; Guimaraes et al., 2008; Miller et al., 2008). Thus, stabilization of kinetochore–microtubule attachment might be concomitant with NDC80 dephosphorylation.
Besides being implicated in the spindle assembly checkpoint, BUB1, BUBR1, and MPS1 have also been shown to take part in biorientation and possibly in error correction (for review see Kang and Yu, 2009). The detailed mechanisms through which these proteins may contribute to these functions are being actively investigated. For instance, it was recently proposed that MPS1 acts upstream of AURORA B to control AURORA B function in biorientation (Jelluma et al., 2008b).
Reversine, a 2,6-disubsituted purine, has been originally identified for its ability to facilitate the dedifferentiation of C2C12 myoblasts into multipotent cells capable of redifferentiating into different cell types (Chen et al., 2004, 2007). Recently, this property of reversine was attributed to its ability to inhibit the AURORA B kinase (D’Alise et al., 2008; Amabile et al., 2009). This spurred our interest in testing the mitotic effects of reversine, and we set out to test whether reversine had additional mitotic targets besides AURORA B. In the course of this analysis, we realized that reversine is a very potent and relatively selective ATP-competitive inhibitor of human MPS1. The mitotic effects of reversine are consistent with the possibility that MPS1 is its principal target in mitosis. Our results demonstrate that MPS1 is indeed a checkpoint component required for the recruitment of other checkpoint proteins, including the subunits of the RZZ complex and MAD1–MAD2, to unattached kinetochores. We also show that MPS1 is implicated in biorientation and in error correction. Our results are consistent with a model in which MPS1 operates downstream from AURORA B and suggest that the error correction and the spindle checkpoint may respond to a single upstream sensor designed to detect lack of attachment and reduced or missing tension.
Results
Reversine is a potent MPS1 inhibitor
Reversine has been shown to target AURORA kinases in vitro and in living cells (D’Alise et al., 2008; Amabile et al., 2009). To assess the potency of reversine on AURORA kinases, we compared its effects with those of known AURORA inhibitors. Reversine inhibited AURORA B in vitro with an IC50 of 98.5 nM, ∼30-fold and twofold above the IC50 of hesperadin and ZM447439, respectively (Fig. 1 A; Fig. S1, A and B; and Table S1). In contrast, AURORA A was inhibited with an IC50 of 876 nM (Fig. S1, A and B; and Table S1).
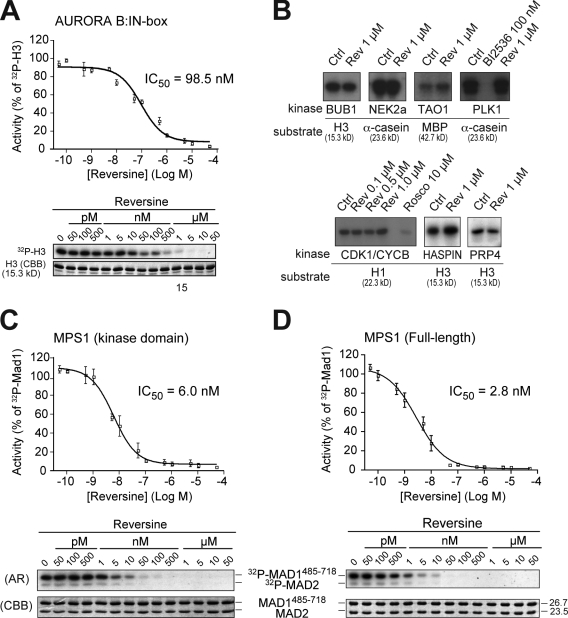
Figure 1.
Reversine inhibits MPS1 in vitro. (A) A kinase assay on the human AURORA B1–344–INCENP835–903 complex with the indicated concentrations of reversine. The substrate is histone H3. (B) The indicated recombinant, purified mitotic kinases were tested with the indicated substrates for their sensitivity to 1 µM reversine (Rev). None of the kinases were significantly inhibited. Specific inhibitors against PLK1 (BI2536) and CDK1 (roscovitine) were used as positive controls. (C and D) A reversine titration experiment on the kinase domain of MPS1 (C) or full-length MPS1 (D). The substrate is the MAD1–MAD2 complex (Sironi et al., 2001). Molecular mass is indicated in kilodaltons. (A, C, and D) Error bars are mean ± SEM. AR, autoradiography; CBB, Coomassie brilliant blue staining; Ctrl, control; CYCB, CYCLIN B.
To ascertain whether reversine is a selective AURORA B inhibitor, we set up an in vitro kinase assay with a battery of human mitotic kinases, including BUB1, CDK1–CYCLIN B, HASPIN, MPS1, NEK2A, PLK1, PRP4, and TAO1 (Fig. 1 B and Table S1). At 1 µM, reversine failed to alter the activity of all but one of these kinases. The MAPKs, which have also been implicated in mitotic control in vertebrates (e.g., Zhao and Chen, 2006), are not significantly inhibited at 1 µM reversine (D’Alise et al., 2008). The only kinase in our dataset to be effectively inhibited by reversine is MPS1, with an IC50 of 6 nM and 2.8 nM for its kinase domain and full-length versions, respectively (Fig. 1, C and D). The latter IC50 value indicates 35-fold selectivity over AURORA B in vitro (both kinases were tested at 50 µM ATP). As a comparison, we found that SP600125, which has been previously shown to inhibit MPS1 (Schmidt et al., 2005), has an IC50 for MPS1 of ∼2.5 µM (Fig. S1 C and Table S1). Surprisingly, we also found that this inhibitor has a significantly lower IC50 for AURORA B (Fig. S1 D and Table S1).
Mitotic phenotypes of reversine
Next, we tried to determine a working concentration of reversine that would inhibit MPS1 but not AURORA kinases. Inhibition of AURORA A or the Eg5 kinesin prevents spindle bipolarization, resulting in a monopolar spindle (Glover et al., 1995). Contrarily to the Eg5 inhibitor S-trityl-l-cysteine (STLC) and the pan-AURORA inhibitor VX680, used as positive controls, reversine did not inhibit spindle bipolarization at concentrations up to 10 µM (Fig. 2, A and B; and Fig. S2, A–D). Thus, AURORA A is unlikely to be a cellular target of reversine at concentrations up to 10 µM or above.
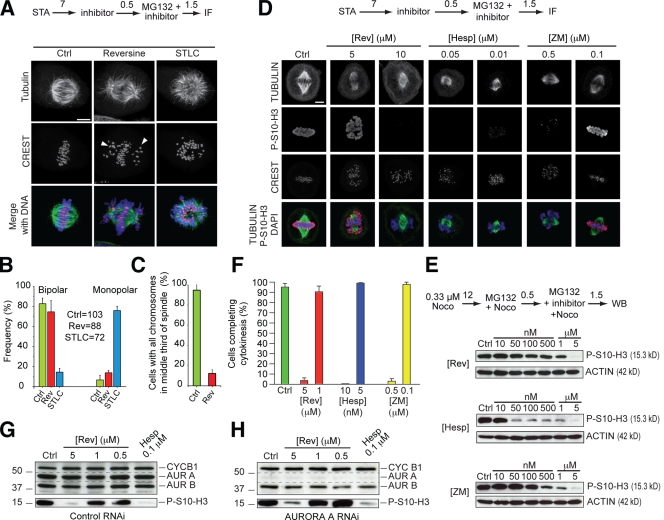
Figure 2.
Submicromolar reversine does not inhibit Aurora B in living cells. (A) Reversine does not prevent spindle bipolarization, but several chromosomes fail to congress (arrowheads). 7 h (numbers above arrows indicate time in hours) after a single thymidine arrest (STA), STLC (an Eg5 inhibitor causing spindle monopolarization) or reversine was added, and after an additional 0.5 h, MG132 was added to prevent mitotic exit. After 1.5 h, cells were then processed for immunofluorescence (IF). (B) Quantification of the experiment in A on the indicated number of cells. (C) Quantification of alignment defects in the same experiment. (D) After release from a single thymidine arrest, cells entered mitosis in the presence of reversine (Rev), hesperadin (Hesp), and ZM447439 (ZM) and were fixed and subjected to immunofluorescence to monitor the levels of P-S10-H3. (E) A comparison of the effects of the inhibitors tested in D on the levels of P-S10-H3 in total cell lysates of mitotic HeLa cells. (F) The effects of the same three inhibitors on cytokinesis were evaluated in a time-lapse experiment. (B, C, and F) Error bars are mean ± SEM. (G and H) AURORA A (AUR A) does not contribute to the generation of P-S10-H3. Under conditions of RNAi of AURORA A, the disappearance of P-S10-H3 in the presence of reversine (H) followed the same pattern as in the control experiments (G). A similar experiment testing the effects on P-S7–CENP-A is reported in Fig. S4 B. Dashed lines highlight the disappearance of AURORA A in H as the result of RNAi. Ctrl, control; CYC B1, CYCLIN B1; Noco, nocodazole; WB, Western blot. Bars, 5 µm.
Reversine did not inhibit kinetochore fiber formation, as assessed with a cold treatment microtubule depolymerization assay (Fig. S2 E). However, reversine had strong effects on chromosome congression. Many chromosomes failed to congress to the metaphase plate in the presence of reversine, a phenotype which was clearly visible already at 250 nM reversine (Fig. 2, A–C; and Fig. S2, A–D).
Based on previous analyses, the reversine phenotype is consistent with inhibition of MPS1 in mammalian cells (Jelluma et al., 2008b; Tighe et al., 2008). However, the phenotype is also reminiscent of phenotypes created by bona fide AURORA B inhibitors such as hesperadin and ZM447439 (Fig. S2 B; Ditchfield et al., 2003; Hauf et al., 2003). To assess the relative contribution of AURORA B or MPS1 inhibition to the chromosome congression problems described in the previous paragraph, we asked whether reversine affected other cellular functions known to implicate AURORA B activity. By immunofluorescence, the phosphorylation of Ser10 of H3 (P-S10-H3), a bona fide AURORA B substrate, was visible until concentrations of reversine >5 µM, whereas the same signal disappeared at significantly lower concentrations of hesperadin or ZM447439 (Fig. 2 D). Similarly, by Western blotting, reversine inhibited P-S10-H3 only at concentrations >2–5 µM, whereas ZM447439 affected significant inhibition of P-S10-H3 already at 500 nM (Fig. 2 E and see Fig. 4 D). With hesperadin, P-S10-H3 was strongly inhibited between 10 and 50 nM (Fig. 2 E).
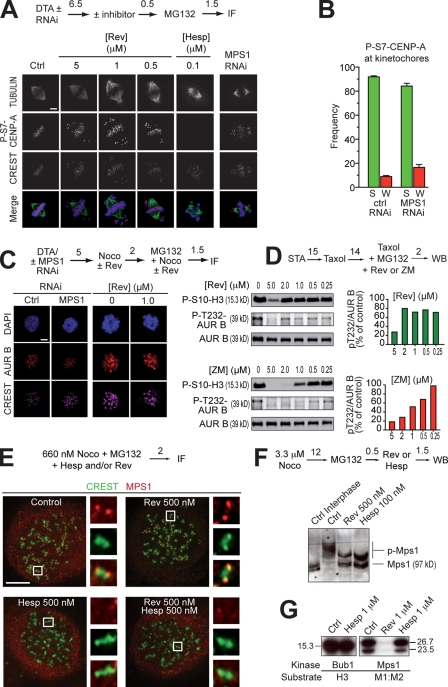
Figure 4.
MPS1 acts downstream from Aurora B. (A) P-S7–CENP-A in mitotic HeLa cells is unaltered even at 5 µM reversine (Rev). The antigen is present on centromeres/kinetochores of chromosomes near the poles, as well as of chromosomes at the equator. The antigen is invisible in the presence of 100 nM hesperadin (Hesp). No compensation from Aurora A was observed (Fig. S4 B). (B) A quantification of the results in A. “S” and “W” indicate strong and weak binding, respectively. These criteria are indicated in Fig. S4 A. Error bars are mean ± SEM. (C) Kinetochore localization of AURORA B (AUR B) in HeLa cells is unaffected after MPS1 RNAi or the addition of reversine. (D, top) Phosphorylation of the activation loop of AURORA B (P-T232) is not affected by reversine until above 2 µM. The pattern of loss of activation loop phosphorylation follows the pattern of loss of P-S10-H3 phosphorylation. (bottom) The same experiment with ZM447439 (ZM) as a positive control. (E) Kinetochore localization of MPS1 in 660 nM nocodazole (Noco) is enhanced by 0.5 µM reversine. If AURORA B is inhibited with 0.5 µM hesperadin, reversine-induced localization of MPS1 is abrogated. Images were taken on a Delta Vision microscope. The insets represent 10× zooms of the boxed areas interpolated using SoftWoRx. (F) Both MPS1 and AURORA B inhibitors reduce the phosphorylation of mitotic MPS1, as visualized through the PHOS tag method (Kinoshita et al., 2006). (A and C–F) Numbers above arrows indicate time in hours. (G) Hesperadin does not inhibit BUB1 or MPS1 in an in vitro kinase assay (see also Table S1). Molecular mass is indicated in kilodaltons. Ctrl, control; DTA, double thymidine arrest; IF, immunofluorescence; WB, Western blot. Bars, 5 µm.
We also tested the effects on cytokinesis, a stringent assay for AURORA B activity (Fig. 2 F). In the 5–10 nM range, hesperadin impaired cytokinesis in 100% of cells. Similar effects were observed in the 0.1–0.5 µM concentration range of ZM447439. However, cytokinesis appeared unaffected at 1 µM reversine and was only impaired at higher concentrations.
To test a possible compensatory role of AURORA A, which, as shown in Fig. S1 (A and B) and Table S1, is only modestly inhibited by reversine in vitro and does not appear to be inhibited in living cells by the criterion that spindles are bipolar, we lowered the levels of AURORA A by RNAi and tested the effects of reversine on P-S10-H3 (Fig. 2, G and H). This condition failed to exacerbate the effect of reversine on P-S10-H3, excluding the hypothesis that AURORA A compensates for AURORA B when reversine is present.
Collectively, these results justify the conclusion that inhibition of AURORA B is unlikely to be the cause of the effects of submicromolar concentrations of reversine in mitotic HeLa cells. Therefore, we decided to carry out additional characterization experiments on the effects of reversine at a reference working concentration of 0.5 µM or else at the concentrations indicated in each figure.
Reversine and RNAi of MPS1 produce similar phenotypes
To corroborate the idea that the observed effects of reversine can be ascribed to the inhibition of MPS1, we performed a systematic comparison of the effects from using 0.5 µM reversine or from ablating MPS1 by RNAi. Because the addition of 0.5 µM reversine or MPS1 depletion by RNAi overrides the spindle checkpoint response to 0.33 µM nocodazole (see Fig. 6), cells were kept in mitosis with 10 µM MG132. At least macroscopically, reversine and MPS1 RNAi caused identical alignment phenotypes (Fig. 3 A). No obvious additive effects on chromosome alignment from combining MPS1 RNAi with reversine were observed, suggesting that MPS1 is a target of submicromolar concentrations of reversine or, alternatively, that the target of reversine works in the same pathway as MPS1.
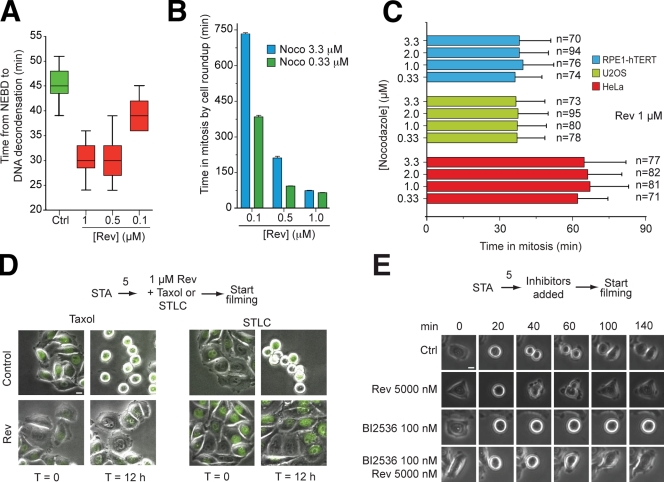
Figure 6.
Reversine is a spindle checkpoint inhibitor. (A) Reversine (Rev) causes normally cycling HeLa cells to exit mitosis prematurely, which is a consequence of spindle checkpoint inactivation. The plot is a quantification of a time-lapse video microscopy experiment. (B) As in A, the experiment quantifies the behavior of cells in time-lapse video microscopy experiments in which HeLa cells were treated with two concentrations of nocodazole (Noco). Additional values (including controls) are collected in Table S2. (C) The ability of reversine to drive HeLa cells out of mitosis extends to at least two additional cell types. (A–C) Error bars are mean ± SEM. (D) The ability of HeLa cells to arrest in mitosis in the presence of 500 nM Taxol or 10 µM STLC was tested in a time-lapse experiment in the presence of reversine. 12 h after the beginning of the video, control cells treated with Taxol or STLC were still in mitosis, whereas the presence of reversine caused a spindle assembly checkpoint override and mitotic exit. (E) Similar experiments were performed in the presence of the Polo kinase inhibitor BI2536, which causes a mitotic arrest (Lénárt et al., 2007). In this case, time 0 refers to the last video frame before mitotic rounding up. (D and E) Numbers above arrows indicate time in hours. Ctrl, control; NEBD, nuclear envelope breakdown. Bars, 10 µm.
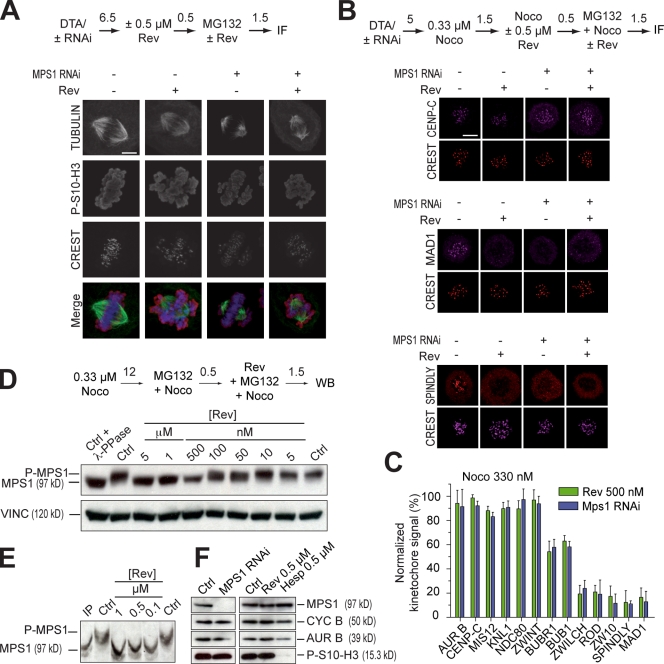
Figure 3.
Reversine inhibits MPS1 in living cells. (A) Chromosome alignment phenotypes of mitotic HeLa cells that were depleted of MPS1 by RNAi or treated with 0.5 µM reversine (Rev), or both. The levels of P-S10-H3 appeared unaltered in all three experiments. A representative RNAi-based depletion of MPS1 is shown in F. (B) Representative localization experiments on different kinetochore proteins including CREST, CENP-C, MAD1, and SPINDLY (Griffis et al., 2007; Chan et al., 2009). Results from the complete analysis are summarized in C and in Fig. S3. (A and B) Numbers above arrows indicate time in hours. (C) The RZZ subunits ROD, ZWILCH, and ZW10, as well as the RZZ-associated protein SPINDLY and MAD1 are all largely evicted from kinetochores when the spindle checkpoint is triggered with 330 nM nocodazole (Noco), with no significant difference between MPS1 RNAi or reversine treatment. The effects on localization are expressed as ratios of the fluorescence value of the indicated protein to the value of CREST (both background subtracted) normalized to the equivalent ratio in control cells. Error bars are mean ± SEM. (D) Dose-dependent inhibition of MPS1 phosphorylation in the presence of reversine. Vinculin (VINC) was used as a loading control. (E) Dose-dependent inhibition of MPS1. Samples were separated on an 8% gel with the PHOS tag method (Kinoshita et al., 2006). (F) Western blotting demonstrates that P-S10-H3 levels are untouched upon MPS1 RNAi or inhibition with reversine. The hesperadin control illustrates the effects from inhibiting AURORA B (AUR B). Ctrl, control; CYC B, CYCLIN B; DTA, double thymidine arrest; IF, immunofluorescence; WB, Western blot. Bars, 5 µm.
We extended the comparison to the localization of an array of a dozen centromere and kinetochore markers, including subunits of the inner and outer kinetochore, of the RZZ complex, and of the spindle checkpoint (Fig. 3, B and C; and Fig. S3). The experiments were performed in the presence of 0.33 µM nocodazole and MG132. In control cells, these conditions prevented satisfaction of the spindle checkpoint, and all checkpoint proteins were recruited to kinetochores (Fig. 3 B). Neither reversine nor RNAi treatment affected kinetochore recruitment of KMN network subunits, indicating that reversine does not grossly perturb the structure of the outer kinetochore (Fig. 3, B and C; Fig. S3; and not depicted).
We next tested the effects from adding reversine on MPS1 phosphorylation, which correlates with its mitotic activation (Kang et al., 2007; Mattison et al., 2007; Jelluma et al., 2008a; Xu et al., 2009). In agreement with the idea that MPS1 is a target of reversine, we observed a dose-dependent reversal of the electrophoretic mobility of MPS1, which reflects autophosphorylation (Fig. 3, D and E; Kang et al., 2007; Mattison et al., 2007; Jelluma et al., 2008a). At 0.5 µM reversine, a concentration that completely inhibits MPS1 autophosphorylation, no effects on P-S10-H3 were observed. Similarly, we did not observe effects on the level of P-S10-H3 upon RNAi-based depletion of MPS1 (Fig. 3 F).
MPS1 acts downstream of AURORA B
Our results so far suggest that reversine is an MPS1 inhibitor in vitro and in vivo. They also demonstrate that reversine does not cause a prominent reduction in the levels of P-S10-H3 in living cells at concentrations (e.g., 0.5 µM) that cause substantial problems in chromosome biorientation and on MPS1 autophosphorylation. Similarly, reversine does not significantly inhibit cytokinesis at 0.5 µM (Fig. 2 F). Overall, these results strongly suggest that MPS1 does not exercise a strong direct control over AURORA B activity. In agreement with this idea, the kinetochore levels of P–CENP-A were not influenced at concentrations of reversine up to 5 µM or above (Fig. 4 A and Fig. S4) and were also not inhibited upon MPS1 RNAi (Fig. 4, A and B). Incidentally, it is worth noting that these experiments were performed in nocodazole, i.e., in the presence of unattached kinetochores. The presence of an intense P–CENP-A signal in nocodazole and its disappearance in the presence of an AURORA B inhibitor such as hesperadin (Fig. S4) shows that, in agreement with a recent study (Liu et al., 2009), AURORA B is active on unattached kinetochores.
We also assessed whether reversine or MPS1 RNAi influenced the localization of AURORA B. In either case, we failed to observe defects in the localization of AURORA B (Fig. 4 C). Furthermore, the presence of reversine did not influence the state of activation of AURORA B, as monitored by activation loop autophosphorylation (P-T232), at least until concentrations at which reversine appeared to hit AURORA B directly (Fig. 4 D).
We monitored MPS1 localization in the presence of reversine and/or hesperadin. In unperturbed mitoses (not depicted) or in nocodazole (Fig. 4 E), we observed a significant cytosolic signal and relatively weak MPS1 kinetochore staining. However, strong kinetochore staining was observed when MPS1 activity was inhibited with 0.5 µM reversine. This result is inconsistent with a recent report that autophosphorylation of MPS1 is required for kinetochore localization (Xu et al., 2009). Inhibition of AURORA B with 0.5 µM hesperadin prevented kinetochore localization of MPS1 in nocodazole, as well as the kinetochore enrichment of MPS1 caused by reversine (Fig. 4 E). Similar results were obtained with 100 nM hesperadin at 3.3 µM nocodazole (Fig. S5). These results indicate that AURORA B may be required for kinetochore localization of MPS1.
Both reversine and hesperadin reduced the mitotic phosphorylation of MPS1 (Fig. 4 F). This was unlikely to be caused by a direct effect of hesperadin on MPS1 because we failed to observe significant MPS1 (and BUB1) inhibition at 1 µM hesperadin in vitro (Fig. 4 G). Collectively, the experiments in Fig. 4 support the idea that MPS1 acts downstream of AURORA B rather than upstream, as recently proposed (Jelluma et al., 2008b).
Role of MPS1 in error correction
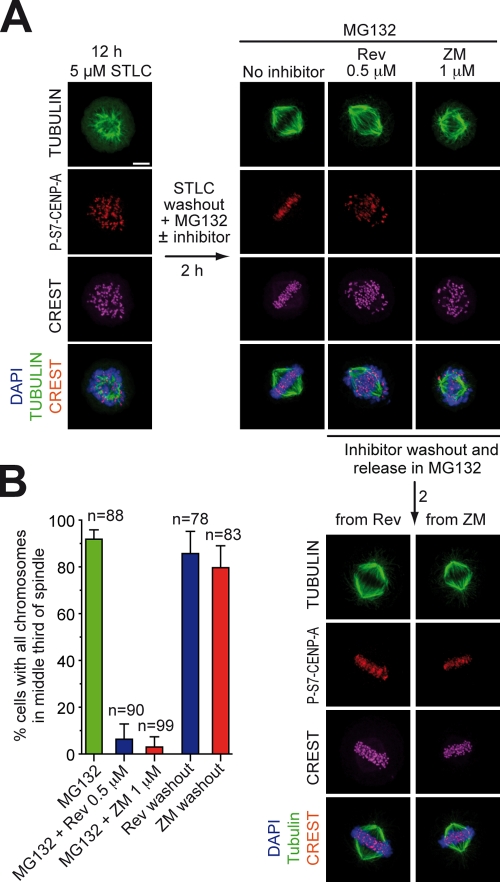
The work so far demonstrates that MPS1 is important for biorientation, which is in agreement with previous observations (Jones et al., 2005; Maure et al., 2007; Jelluma et al., 2008b; Tighe et al., 2008). We wished to exploit the availability of a small molecule inhibitor of MPS1 to test whether this kinase is implicated in error correction. For this, we applied an assay previously developed to test the implication of AURORA B in error correction (Lampson et al., 2004). HeLa cells were first treated with the Eg5 inhibitor STLC to induce a monopolar spindle and a large number of kinetochore–microtubule attachment errors (Fig. 5 A). Cells were then allowed to recover by washing out the Eg5 inhibitor in the presence of MG132. Control cells formed a bipolar spindle. If the recovery phase was performed in the presence of reversine to inhibit MPS1 or ZM447439 to inhibit AURORA B, bipolar spindles also formed, but several misaligned chromosomes were evident (as quantified in Fig. 5 B). Thus, both MPS1 and AURORA B activity are required to recover from the attachment errors induced by monopolarization. Of note, although the P–CENP-A signal disappeared in ZM447439, no inhibition of P–CENP-A was evident in the presence of reversine, indicating that the target of reversine in error correction is unlikely to be, or to act upstream of, AURORA B in this pathway. At 1 µM, ZM447439 did not inhibit MPS1 in vitro (Table S1). After washout of ZM447439 or reversine, normal metaphases with properly aligned chromosomes formed, indicating that the targets of these inhibitors are required for error correction. Overall, these results implicate MPS1, like AURORA B, in the correction of improper kinetochore–microtubule attachments.
Figure 5.
MPS1 is involved in error correction. (A) Cycling HeLa cells were treated with STLC for 12 h. Most cells arrest in mitosis with a monopolar spindle. After STLC washout in MG132, control cells bipolarize and form a normal metaphase. If STLC washout is performed in the presence of reversine (Rev) or ZM447439 (ZM), the spindle bipolarizes normally, but a large fraction of improper attachments are visible. P-S7–CENP-A, a bona fide AURORA B substrate, appears unaltered in reversine-treated cells but disappears in ZM447439. After removal of the inhibitors, a metaphase plate forms. P-S7–CENP-A reappears after washout of ZM447439. In vitro, 2 µM ZM447439 does not inhibit MPS1 (Table S1 and not depicted). (B) Quantification of results with number of cells monitored in the experiment. Error bars are mean ± SEM. Bar, 5 µm.
Effects of reversine on cell cycle progression
As expected for an MPS1 inhibitor, reversine caused HeLa cells to exit mitosis prematurely during an unperturbed mitosis (Fig. 6 A), as demonstrated previously for the ablation of additional checkpoint components such as MAD2 and BUBR1 (e.g., Meraldi et al., 2004). This was confirmed in experiments in which cells were treated with concentrations of nocodazole (0.33 µM or 3.3 µM) that cause partial or complete microtubule depolymerization, respectively (Fig. 7 A; Brito et al., 2008). The addition of reversine caused a dose-dependent reduction in the timing of mitotic arrest, and the override was complete at 1.0 µM reversine at either concentration of nocodazole (Table S2). At lower concentrations of reversine, the effects on the duration of the checkpoint were more explicit at 0.33 µM nocodazole. Similar trends were observed with AURORA kinase inhibitors (Table S2). Checkpoint overriding by reversine was not limited to HeLa cells, as it was also observed with comparable potency in U2OS and retinal pigment epithelial cells (Fig. 6 C). Reversine also caused an override of the checkpoint in the presence of Taxol, STLC, or the Plk1 inhibitor BI2536 (Fig. 6, D and E).
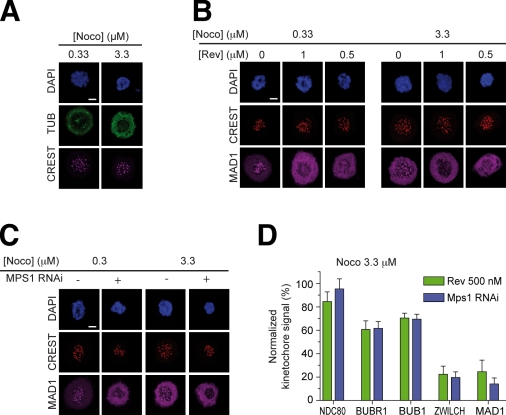
Figure 7.
MPS1 is required for kinetochore recruitment of the RZZ and MAD1 even in high nocodazole. (A) Immunofluorescence analysis of the distribution of TUBULIN (TUB) in the presence of 0.33 and 3.3 µM nocodazole (Noco). Residual foci of polymerized TUBULIN are visible in 0.33 µM nocodazole but not in 3.3 µM nocodazole. HeLa cells were incubated in the presence of nocodazole for 15 min before fixation for immunofluorescence. (B) 5 h after release from a double thymidine arrest, reversine (Rev; at the indicated concentrations) and nocodazole (0.33 or 3.3 µM) were added. MAD1 failed to localize to kinetochores at either nocodazole concentration. (C) The same experimental scheme as in B was used under conditions of RNAi-based depletion of MPS1. (D) The histogram summarizes results on localization experiments equivalent to those in B and C on MAD1 and the additional indicated kinetochore proteins. Localization data were quantified as in Fig. 3 C. Error bars are mean ± SEM. Bars, 5 µm.
MPS1 is required for localization of checkpoint proteins when microtubules are completely depolymerized
Kinetochore-bound microtubules contribute to removing the checkpoint proteins from kinetochores (for review see Musacchio and Salmon, 2007). A consequence of the artificial stabilization of kinetochore–microtubule attachment when the error correction pathway is inhibited is that the levels of checkpoint proteins at kinetochores are strongly reduced (Yang et al., 2009). To demonstrate beyond reasonable doubt that the inhibition of MPS1 causes a genuine checkpoint override rather than a mere satisfaction of the spindle checkpoint in the absence of error correction, as has been previously proposed for AURORA B inhibitors (Yang et al., 2009), we monitored the recruitment of the checkpoint proteins, an established hallmark of checkpoint activity, to kinetochores at 3.3 µM nocodazole (Fig. 7 A). Even at 3.3 µM nocodazole, both the RZZ and MAD1 were unable to localize to kinetochores (Fig. 7, B–D). Thus, the disappearance of checkpoint proteins from kinetochores when MPS1 is inhibited is not caused by satisfaction of the spindle checkpoint by residual kinetochore–microtubules in the absence of an error correction mechanism. Rather, this behavior reflects a genuine requirement of MPS1 in kinetochore recruitment of a subset of checkpoint components.
Reversine does not inhibit MEK1, nonmuscle Myosin II (NMMII), or phosphatidyl-inositol 3-kinase (PI3K) in mitosis
After their initial characterization of reversine in the dedifferentiation of lineage-committed mouse-derived C2C12 myoblasts (Chen et al., 2004), Chen et al. (2007) identified NMMII, MEK1, and PI3K as putative targets of reversine. Although our characterization of reversine strongly supports inhibition of MPS1 as the main mechanism of reversine action in mitosis, we wished to test the possibility that NMMII, MEK1, or PI3K are targets of reversine during mitosis.
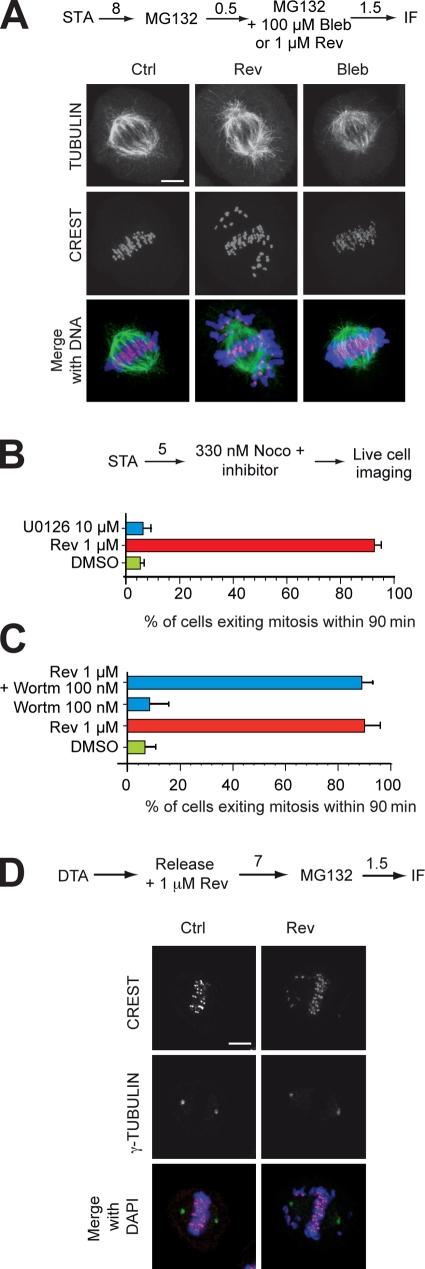
The effects of blebbistatin, an NMMII inhibitor (Straight et al., 2003), were compared with the effects of reversine. At 100 µM, blebbistatin did not cause any evident effects on chromosome alignment, suggesting that NMMII, the target of this inhibitor, does not contribute to chromosome alignment (Fig. 8 A). Blebbistatin did not significantly affect the ability of mitotic HeLa cells to maintain a nocodazole-mediated arrest (unpublished data). Because reversine does not have obvious effects on cytokinesis until concentrations of 2–5 µM, at which concentrations we show that it inhibits Aurora B (Fig. 2 F), we surmise that the mitotic phenotypes caused by submicromolar reversine are unlikely to be the result of the inhibition of NMMII and that if NMMII inhibition occurs, it does so at concentrations of reversine >2–5 µM. To assess whether NMMII is a target of reversine at high concentration in mitotic cells, it will be necessary to sort out the relative effects of reversine on Aurora B and NMMII, as both of these proteins work in cytokinesis.
Figure 8.
Reversine does not inhibit NMMII, MEK1, PI3K, or centrosome duplication. (A) At 100 µM, blebbistatin (Bleb) does not cause evident perturbations of chromosome alignment, contrarily to 1 µM reversine (Rev). (B) At 10 µM, the MEK1 inhibitor U0126 does not affect the duration of the spindle checkpoint. (C) At 100 nM, the PI3K inhibitor wortmannin (Wortm) does not affect the duration of the spindle checkpoint. (B and C) Error bars are mean ± SEM. (D) Cells arrested with a double thymidine arrest (DTA) procedure were released in the cell cycle, and the number of centrosomes was measured in the subsequent mitosis. In all cases, two centrosomes were measured, indicating that centrosome duplication takes place normally in the presence of reversine. (A, B, and D) Numbers above arrows indicate time in hours. Ctrl, control; IF, immunofluorescence; STA, single thymidine arrest. Bars, 5 µm.
We also compared the effects from adding MEK1 or PI3K inhibitors to the ability of HeLa cells to maintain a nocodazole-mediated arrest. Neither the MEK inhibitor U0126 nor the PI3K inhibitor wortmannin affected the duration of the spindle checkpoint in the presence of spindle poisons (Fig. 8, B and C). Overall, these results indicate that NMMII, MEK1, and PI3K are not prominent mitotic targets of reversine or else that their inhibition by reversine does not cause a prominent mitotic phenotype. In agreement with a previous study (Stucke et al., 2004), we also failed to see an effect of reversine on centrosome duplication (Fig. 8 D).
Discussion
In this study, we have demonstrated a role for the small molecule reversine in the mitotic inhibition of MPS1. After the discovery of cincreasin as an MPS1 inhibitor in budding yeast (Dorer et al., 2005), reversine now provides a small molecule tool for interfering with the spindle checkpoint in human cells, flanking additional recently described MPS1 inhibitors (Kwiatkowski et al., 2010; see Hewitt et al. in this issue). We show that reversine inhibits AURORA B in mitosis but at concentrations that are incompatible with the observed adverse effects of submicromolar reversine on biorientation, error correction, and the spindle checkpoint. However, the reported accumulation of polyploid cells at micromolar concentrations of reversine (presumably caused by a failure in cytokinesis) is consistent with AURORA B inhibition (D’Alise et al., 2008). Our systematic comparison of the effects from using reversine at submicromolar concentrations with the effects from ablating MPS1 by RNAi implies that MPS1 is the main mitotic target of reversine. Inhibition of additional targets in other cell cycle phases and in postmitotic cells may be responsible for the dedifferentiation function of reversine (Chen et al., 2004).
Our analysis indicates that the catalytic activity of MPS1 is implicated both in error correction and in the spindle checkpoint. We hypothesize that the error correction and spindle checkpoint pathways intersect at MPS1 when its kinase activity becomes activated at kinetochores so that substrates in both pathways become concomitantly phosphorylated. Although we support this hypothesis, it is formally possible that MPS1 is selectively activated to phosphorylate targets relevant to error correction or to the spindle checkpoint under different conditions (e.g., lack of attachment or reduced tension in the presence of attachment). Future studies will be required to distinguish between these two models.
Among the mechanisms through which MPS1 may contribute to biorientation and error correction is the ability of MPS1 to regulate the motor activity of CENP-E, a plus end–directed motor that crucially contributes to chromosome congression (Espeut et al., 2008). Furthermore, the ablation of kinetochore recruitment of the RZZ complex in the absence of MPS1 activity likely prevents kinetochore recruitment of Dynein, which also contributes to kinetochore–microtubule attachment (for review see Musacchio and Salmon, 2007). In yeast, Mps1 regulation of biorientation may proceed through phosphorylation of the subunits of the Dam1 and Ndc80 complexes (Shimogawa et al., 2006; Maure et al., 2007; Kemmler et al., 2009). However, MPS1 may control the spindle checkpoint by contributing, among additional functions, to kinetochore recruitment of the RZZ complex and MAD1.
It is important to characterize the hierarchical relationships at the apex of the sensory apparatus that distinguished correct from incorrect attachments and that ignites the error correction and checkpoint responses. Two recent studies demonstrated that intrakinetochore stretch upon microtubule binding, as opposed to interkinetochore stretch, correlates with the status of the checkpoint response (Maresca and Salmon, 2009; Uchida et al., 2009). Upon microtubule binding, the distance between specifically positioned fluorescence markers within the kinetochore, projected onto the interkinetochore axis, increases up to 35–40 nm (Maresca and Salmon, 2009; Uchida et al., 2009; Wan et al., 2009). These changes may reflect a distortion in the structure of kinetochores caused by the application of a physical force (tension) upon microtubule binding. Alternatively, they may reflect a conformational change in the kinetochore triggered by microtubule binding. The first hypothesis is reinforced by the observation that microtubule binding is by itself insufficient to cause full intrakinetochore stretching and that dynamic microtubules are required for full stretching (Maresca and Salmon, 2009, 2010).
The AURORA B kinase has emerged as a key regulator of the error correction pathway. It has been proposed that AURORA B may monitor variations in the distance from its substrates as microtubules attach to kinetochores (Tanaka et al., 2002; Andrews et al., 2004). Strong experimental evidence in favor of this idea is emerging (Vader et al., 2007; Liu et al., 2009, 2010; Welburn et al., 2010). Tension exerted by bound microtubules may contribute to the gradual displacement of substrates from AURORA B, resulting in turn in substrate dephosphorylation. We have recently proposed a speculative model picturing INCENP as a “dog leash” whose limited extension limits the ability of AURORA B to reach its substrates within the kinetochore (for review see Santaguida and Musacchio, 2009). Previous experiments with a deletion mutant of INCENP are indeed consistent with this idea (Vader et al., 2007).
We provide evidence that AURORA B acts upstream of MPS1 and that the perturbation of MPS1 activity does not grossly alter the phosphorylation of AURORA B substrates or the localization of AURORA B. Similar results are reported in an accompanying paper describing the effects from targeting an analogue-sensitized allele of MPS1 (see Maciejowski et al. in this issue). Similarly, no effects on the levels of AURORA B substrates are observed with an additional MPS1 inhibitor, AZ3146 (Hewitt et al., 2010). If inhibition of MPS1 does not result in overt changes in AURORA B activity, we show instead that inhibition of AURORA B causes a mislocalization of MPS1 and a reduction of its phosphorylation, suggesting that AURORA B acts upstream of MPS1. This possibility is also consistent with the pattern of recruitment of spindle checkpoint proteins in different systems (Vigneron et al., 2004; Famulski and Chan, 2007; Emanuele et al., 2008). Because MPS1 turns over rapidly at kinetochores (Howell et al., 2004), its activation at kinetochores, which probably involves dimerization and autophosphorylation (Kang et al., 2007; Mattison et al., 2007; Jelluma et al., 2008a), may precede its release in the cytosol in an active form.
Overall, these results may appear inconsistent with the recent proposal that MPS1 controls AURORA B through phosphorylation of BOREALIN, a subunit of the chromosome passenger complex (Jelluma et al., 2008b; Kwiatkowski et al., 2010; Sliedrecht et al., 2010). Because a phospho-mimicking mutant of BOREALIN simulating MPS1 phosphorylation rescues the effects on biorientation from loosing MPS1 (Jelluma et al., 2008b), MPS1 and BOREALIN may participate in an AURORA B–independent pathway implicated in biorientation. More studies will be required to assess this idea.
If MPS1, which is implicated in error correction and in the checkpoint, acts downstream from AURORA B and is activated by it, then AURORA B is also expected to control both error correction and the spindle checkpoint. Although the involvement of AURORA B in error correction is widely accepted, its participation in the spindle checkpoint is more controversial. In at least two model systems, Schizosaccharomyces pombe and Xenopus laevis, Aurora B is required for the checkpoint response to unattached kinetochores (Kallio et al., 2002; Petersen and Hagan, 2003; Vanoosthuyse and Hardwick, 2009). Direct involvement of AURORA B in checkpoint signaling has also been observed upon expression of an INCENP mutant deleted of the coiled-coil domain of INCENP (Vader et al., 2007). This mutant does not affect the ability of AURORA B to phosphorylate some of its centromeric substrates, suggesting that it is impairing a specific function of the chromosome passenger complex in spindle checkpoint control (Vader et al., 2007).
In many additional settings, including experiments with yeast temperature-sensitive mutants (Biggins and Murray, 2001) or small molecule inhibitors (Ditchfield et al., 2003; Hauf et al., 2003), the inhibition of AURORA B has been shown to reduce the strength of the checkpoint arrest to unattached kinetochores but not to lead to complete override. It is possible that these effects result from residual AURORA B activity as a consequence of incomplete depletion or inactivation. Small residual AURORA B activity may be sufficient to maintain the arrest under the strong checkpoint-activating conditions created by spindle-depolymerizing agents. However, the requirements on MPS1 may be more stringent, explaining why it is relatively easier to observe a checkpoint override when targeting MPS1.
A confusing aspect of the relationship between error correction and the spindle checkpoint is that the inhibition of error correction can influence the pattern of kinetochore localization of the spindle checkpoint proteins and therefore the strength of the checkpoint response at suboptimal concentrations of spindle-depolymerizing drugs such as nocodazole (Brito et al., 2008; Yang et al., 2009). Evidence of this can be extrapolated from Fig. 6 B: the same concentration of reversine (i.e., the same expected degree of target kinase inhibition) has significantly different effects on the duration of mitotic arrest at low or high nocodazole doses. Thus, residual microtubules may contribute to checkpoint satisfaction if kinetochores cannot let go of them because error correction is impaired (Yang et al., 2009). A pathway that removes the checkpoint proteins from microtubule-bound kinetochores (for review see Musacchio and Salmon, 2007) is likely responsible for this phenomenon. Future studies will have to refer to the rigorous test proposed by Yang et al. (2009) for evaluating the participation of MPS1, AURORA B, and other proteins in the checkpoint response. The test consists in evaluating the effects from ablating a putative checkpoint component when spindle-depolymerizing drugs are present at concentrations (3.2 µM nocodazole or more in HeLa cells) that remove any residual tubulin polymer. By applying this test to AURORA B, Yang et al. (2009) demonstrated that at 100 nM hesperadin, the presence or absence of residual microtubules results in dramatic differences in the localization of the checkpoint protein MAD2 to kinetochores. At high nocodazole concentrations (3.2 µM), MAD2 is retained on kinetochores despite the presence of hesperadin. Conversely, at low nocodazole concentrations and at the same concentration of hesperadin, MAD2 is absent from kinetochores (Yang et al., 2009).
This result predicts that previous studies implicating AURORA B in MAD2 recruitment might have been at least in part biased by the relatively low nocodazole concentrations used (e.g., Ditchfield et al., 2003; Hauf et al., 2003). However, we find that at higher hesperadin concentrations (0.5–1.0 µM), MAD1 (which is required for MAD2 recruitment) and the RZZ complex are lost from kinetochores even at high concentrations of nocodazole (unpublished data). Thus, AURORA B may be ultimately required for the recruitment of these checkpoint proteins, but higher levels of inhibition may be required for its involvement to become explicit. We show that at least in vitro, these higher concentrations of hesperadin do not inhibit BUB1 and MPS1, but it remains formally possible that hesperadin inhibits additional kinases in the MAD1 and RZZ recruitment pathway. We conclude that a formal assessment of the role of AURORA B in the checkpoint response will require more penetrant and selective inhibition of AURORA B.
Materials and methods
Cell culture and synchronization
HeLa cells and U2OS cells were grown in DME (EuroClone) supplemented with 10% fetal bovine serum (HyClone) and 2 mM l-glutamine. Human telomerase reverse transcriptase–retinal pigment epithelial cells were grown in minimal essential medium: Ham’s F12K medium 1:1 supplemented with 10% fetal bovine serum, 15 mM Hepes, and 0.5 mM Na pyruvate. 0.33 and 3.3 µM nocodazole, 0.5 µM Taxol, 5 µM STLC, and 2 mM thymidine were obtained from Sigma-Aldrich. MG132 (EMD) was used at 10 µM.
RNAi
siRNA duplexes used to repress AURORA A, AURORA B, BUB1, BUBR1, and MPS1 (Ditchfield et al., 2003) had the following sequences: AURORA A, 5′-AAGCACAAAAGCUUGUCUCCA-3′; AURORA B, 5′-AACGCGGCACUUCACAAUUGA-3′; BUB1, 5′-AAAUACCACAAUGACCCAAGA-3′; BUBR1, 5′-AACGGGCAUUUGAAUAUGAAA-3′; and MPS1, 5′-GACAGAUGAUUCAGUUGUA-3′. siRNA duplexes were purchased from Thermo Fisher Scientific and transfected using Lipofectamine 2000 reagent (Invitrogen) according to the manufacturer’s instructions.
Immunofluorescence microscopy and antibodies for immunofluorescence
In all cases except Fig. 4 E, immunofluorescence microscopy was performed on cells fixed using 4% PFA in PBS, permeabilized using 0.1% Triton X-100 in PBS, and then treated with 4% BSA in PBS as blocking agent and incubated with the proper antibodies diluted in 4% BSA in PBS. For MPS1 staining, cells grown on coverslips were washed in PBS, fixed in 1% formaldehyde for 5 min, quenched in glycine, pH 8.5, and then permeabilized with PBS plus 0.1% Triton X-100 (PBST) before incubation with primary and secondary antibodies (Taylor et al., 2001; Tighe et al., 2008; Trazzi et al., 2009).
The following antibodies were used for immunofluorescence: anticentromeric antibody (working dilution 1:50; Antibodies Inc.), mouse anti-HEC1 (human NDC80; working dilution 1:1,000; clone 9G3.23; GeneTex, Inc.), mouse anti–α-TUBULIN (working dilution 1:2,000; clone B512; Sigma-Aldrich), rabbit anti-SPINDLY (working dilution 1:250; Bethyl Laboratories, Inc.); rabbit anti–AURORA B (working dilution 1:1,000; Abcam), rabbit anti–PS10-H3 (working dilution 1:500; Abcam), and rabbit anti-P-S7–CENP-A Ser7 (working dilution 1:300; Abcam). Antibodies against BUB1, BUBR1, CENP-C, MAD1, MPS1, ZW10, and ZWILCH have been described previously (Taylor et al., 2001; De Antoni et al., 2005; Tighe et al., 2008; Trazzi et al., 2009; Civril et al., 2010). Antibody against ROD was a gift from T.J. Yen (Fox Chase Cancer Center, Philadelphia, PA). Antibodies against MIS12 and KNL1 were a gift from T. Kiyomitsu and M. Yanagida (Kyoto University, Sakyo-ku, Kyoto, Japan). Cy3- and Cy5-labeled and Alexa Fluor 488–labeled secondary antibodies for immunofluorescence were purchased from Jackson ImmunoResearch Laboratories, Inc. and Invitrogen, respectively. DNA was stained with DAPI. The coverslips were mounted using Mowiol mounting media.
Cells were imaged using a confocal microscope (TCS SP2; Leica) equipped with a 63× NA 1.4 objective lens using the LCS 3D software (Leica). Images were imported in Photoshop CS3 (Adobe Systems, Inc.), and levels were adjusted. Pixel intensity quantification has been performed using SoftWoRx (Applied Precision).
For Fig. 4 E, immunofluorescence images were acquired at room temperature on a restoration microscope (Delta Vision RT; Applied Precision) using a 100× NA 1.40 Plan-Apochromat objective and the Sedat Quad filter set (Chroma Technology Corp.). The images were collected using a charge-coupled device camera (CoolSNAP HQ; Photometrics) with a z-optical spacing of 0.2 mm. Raw images were then deconvolved using the SoftWoRx software, and maximum intensity projections of these deconvolved images are shown.
Antibodies for immunoblotting
The following antibodies were used for immunoblotting: mouse anti–AURORA A (working dilution 1:1,000; Sigma-Aldrich), rabbit anti–AURORA B (working dilution 1:1,000; Abcam), rabbit anti-BUB1 (working dilution 1:2,000; Abcam), mouse anti-BUBR1 (working dilution 1:1,000; BD), mouse anti-MPS1 (working dilution 1:2,000; Millipore), and rabbit anti–P-S10-H3 (working dilution 1:1,000; Abcam).
Video microscopy
Live cell imaging was performed using an inverted microscope (IX70; Nikon) equipped with an incubation chamber (Solent Scientific) maintained at 37°C in an atmosphere of 5% CO2. Videos were acquired using a magnification objective of 20 controlled by ScanR software (Olympus).
In vitro kinase assays
Kinase assays were performed in 30 µl (DeMoe et al., 2009). Reaction mixes contained 50 µM ATP, 1 mM DTT, 1 mM Na3VO4, 5 µCi γ-[32P]ATP, 1 µg of the appropriate substrate, 1 µl DMSO or drugs dissolved in DMSO, and the indicated kinases (typically at a concentration of 5 nM). Reaction mixes were incubated for 1 h at 30°C, quenched with SDS loading buffer, and resolved on 14% SDS-PAGE. Incorporation of 32P was visualized by autoradiography. Densitometry analysis was performed using ImageJ software (National Institutes of Health). IC50 values were calculated from log-dose response curves using Prism 4 software (GraphPad Software, Inc.).
Purification of mitotic kinases
Human Aurora B1–344–INCENP835–903 was expressed in and purified from Escherichia coli as a fusion to GST. The protein was purified on reduced glutathione (GSH) Sepharose Fast Flow (GE Healthcare), and the GST tag was cleaved using PreScission protease (GE Healthcare). The cleaved product was further purified by size exclusion chromatography in 50 mM Tris-HCl, pH 7.6, 150 mM NaCl, and 1 mM DTT. Aurora A–TPX2 (Bayliss et al., 2003), a gift of E. Conti (Max Planck Institute of Biochemistry, Martinsried, Germany) was assayed like Aurora B. Human Nek2A (residues 1–291) was expressed in E. coli as a fusion to GST. The protein was purified on GSH Sepharose Fast Flow, and the GST tag was cleaved using PreScission protease. The cleaved product was further purified by size exclusion chromatography. NEK2A assays were performed in 50 mM Tris-HCl, pH 7.5, 10 mM MgCl2, and 10 mM MnCl2 with α-casein as a substrate. The Bub1–Bub3 complex was expressed in and purified from Sf9 insect cells infected with recombinant baculoviruses. The complex was isolated on Ni–nitrilotriacetic acid beads and further purified by size exclusion chromatography. Bub1–Bub3 kinase reaction buffer contained 50 mM Tris-HCl, pH 7.6, 150 mM NaCl, 10 mM MgCl2, and 1 mM EDTA, and histone H3 was used as substrate. Full-length Mps1 was purchased from Invitrogen and assayed in 50 mM Tris-HCl, pH 7.5, 10 mM MgCl2, 10 mM MnCl2, and Mad1–Mad2 complex as a substrate. Human NEK2A (residues 1–291) was expressed in E. coli as a fusion to GST. The protein was purified on reduced glutathione (GSH) Sepharose Fast Flow, and the GST tag was cleaved using PreScission protease. The cleaved product was further purified by size exclusion chromatography. NEK2A assays were performed in 50 mM Tris-HCl, pH 7.5, 10 mM MgCl2, and 10 mM MnCl2 with α-casein as a substrate. Human Plk1 (a gift of A. Tarricone, European Institute of Oncology, Milan, Italy) was tested in 50 mM Tris-HCl, pH 7.6, 150 mM NaCl, 10 mM MgCl2, and 1 mM EDTA with α-casein as a substrate. The cDNA encoding human TAO1 (residues 1–356) was a gift of D. Alessi (University of Dundee, Dundee, Scotland, UK). TAO1 was expressed as an NH2-terminal GST fusion in E. coli and isolated on GSH Sepharose Fast Flow. GST-tagged TAO1 immobilized on GSH Sepharose beads was directly used in kinase assay in 40 mM Hepes, pH 7.5, 10 mM MgCl2, 1 mM EDTA, and myelin basic protein as a substrate. PRP4 kinase was expressed as a fusion to a hexahistidine tag in Hi5 insect cells infected with recombinant baculoviruses. The complex was isolated on Ni–nitrilotriacetic acid beads, eluted using 200 mM imidazole, and further dialyzed against PBS. PRP4 kinase reaction buffer contained 50 mM Tris-HCl, pH 7.6, 150 mM NaCl, 10 mM MgCl2, and 1 mM EDTA, and histone H3 was used as substrate. The HASPIN kinase domain (residues 452–798) was expressed in and purified from E. coli as a fusion to GST. GST–Haspin452–798 was affinity purified on GSH beads (Villa et al., 2009). After removal of the tag, the supernatant was further purified on Resource Q and a Superdex 200 column (GE Healthcare). Reactions were performed in a solution containing 50 mM Tris, pH 7.6, 10 mM MgCl2, 150 mM NaCl, and 1 mM EDTA. CDK1–CYCLIN B was a gift of A. Tarricone. Kinase assays were performed in 40 mM Hepes, pH 8, 40 µM potassium glutamate, 8 mM MgCl2, 1 mM EGDA, and 0.5 mM EDTA.
Online supplemental material
Fig. S1 shows additional kinase assays. Fig. S2 shows the characterization of the alignment phenotypes of different inhibitors. Fig. S3 shows additional kinetochore localization experiments. Fig. S4 shows that the levels of P-S7–CENP-A are not affected by reversine. Fig. S5 shows that AURORA B inhibition prevents accumulation of kinetochore MPS1. Table S1 shows IC50 values (in nM) for the combination of different inhibitors and kinases. Table S2 shows the duration of mitosis in cells treated with spindle poisons and kinase inhibitors. Online supplemental material is available at http://www.jcb.org/cgi/content/full/jcb.201001036/DC1.
Acknowledgments
We thank the members of the Musacchio laboratory and R. Cortese for many helpful discussions, L. Massimiliano for help with insect cell expression, G. Ossolengo for help with polyclonal antibodies, E. Conti, A. Tarricone, S. Plyte, T. Kiyomitsu, and M. Yanagida for sharing reagents, S. Lens, G. Kops, and T. Tanaka for critical reading of the manuscript, and S. Lens and M. Vromans for help with Fig. 4 D.
Work in the Musacchio laboratory is funded by the Association for International Cancer Research, the Telethon Foundation, the FP7 European Research Council grant KINCON, the FP7 Integrated Project MitoSys, the Italian Association for Cancer Research, the Fondo di Investimento per la Ricerca di Base, the Cariplo Foundation, and the Human Frontier Science Program. S. Santaguida is a graduate student of the European School of Molecular Medicine and is supported by a fellowship from the Italian Foundation for Cancer Research. S.S. Taylor is a Cancer Research UK Senior Fellow.
Footnotes
Abbreviations used in this paper:
- APC/C
- anaphase-promoting complex/cyclosome
- NMMII
- nonmuscle Myosin II
- PI3K
- phosphatidyl-inositol 3-kinase
- RZZ
- ROD–ZWILCH–ZW10
- STLC
- S-trityl-l-cysteine
References
- Amabile G., D’Alise A.M., Iovino M., Jones P., Santaguida S., Musacchio A., Taylor S., Cortese R. 2009. The Aurora B kinase activity is required for the maintenance of the differentiated state of murine myoblasts. Cell Death Differ. 16:321–330 10.1038/cdd.2008.156 [DOI] [PubMed] [Google Scholar]
- Andrews P.D., Ovechkina Y., Morrice N., Wagenbach M., Duncan K., Wordeman L., Swedlow J.R. 2004. Aurora B regulates MCAK at the mitotic centromere. Dev. Cell. 6:253–268 10.1016/S1534-5807(04)00025-5 [DOI] [PubMed] [Google Scholar]
- Bayliss R., Sardon T., Vernos I., Conti E. 2003. Structural basis of Aurora-A activation by TPX2 at the mitotic spindle. Mol. Cell. 12:851–862 10.1016/S1097-2765(03)00392-7 [DOI] [PubMed] [Google Scholar]
- Biggins S., Murray A.W. 2001. The budding yeast protein kinase Ipl1/Aurora allows the absence of tension to activate the spindle checkpoint. Genes Dev. 15:3118–3129 10.1101/gad.934801 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brito D.A., Yang Z., Rieder C.L. 2008. Microtubules do not promote mitotic slippage when the spindle assembly checkpoint cannot be satisfied. J. Cell Biol. 182:623–629 10.1083/jcb.200805072 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan Y.W., Fava L.L., Uldschmid A., Schmitz M.H., Gerlich D.W., Nigg E.A., Santamaria A. 2009. Mitotic control of kinetochore-associated dynein and spindle orientation by human Spindly. J. Cell Biol. 185:859–874 10.1083/jcb.200812167 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheeseman I.M., Chappie J.S., Wilson-Kubalek E.M., Desai A. 2006. The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell. 127:983–997 10.1016/j.cell.2006.09.039 [DOI] [PubMed] [Google Scholar]
- Chen S., Zhang Q., Wu X., Schultz P.G., Ding S. 2004. Dedifferentiation of lineage-committed cells by a small molecule. J. Am. Chem. Soc. 126:410–411 10.1021/ja037390k [DOI] [PubMed] [Google Scholar]
- Chen S., Takanashi S., Zhang Q., Xiong W., Zhu S., Peters E.C., Ding S., Schultz P.G. 2007. Reversine increases the plasticity of lineage-committed mammalian cells. Proc. Natl. Acad. Sci. USA. 104:10482–10487 10.1073/pnas.0704360104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciferri C., Pasqualato S., Screpanti E., Varetti G., Santaguida S., Dos Reis G., Maiolica A., Polka J., De Luca J.G., De Wulf P., et al. 2008. Implications for kinetochore-microtubule attachment from the structure of an engineered Ndc80 complex. Cell. 133:427–439 10.1016/j.cell.2008.03.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cimini D., Wan X., Hirel C.B., Salmon E.D. 2006. Aurora kinase promotes turnover of kinetochore microtubules to reduce chromosome segregation errors. Curr. Biol. 16:1711–1718 10.1016/j.cub.2006.07.022 [DOI] [PubMed] [Google Scholar]
- Civril F., Wehenkel A., Giorgi F.M., Santaguida S., Di Fonzo A., Grigorean G., Ciccarelli F.D., Musacchio A. 2010. Structural analysis of the RZZ complex reveals common ancestry with multisubunit vesicle tethering machinery. Structure. 18:616–626 10.1016/j.str.2010.02.014 [DOI] [PubMed] [Google Scholar]
- Cleveland D.W., Mao Y., Sullivan K.F. 2003. Centromeres and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell. 112:407–421 10.1016/S0092-8674(03)00115-6 [DOI] [PubMed] [Google Scholar]
- D’Alise A.M., Amabile G., Iovino M., Di Giorgio F.P., Bartiromo M., Sessa F., Villa F., Musacchio A., Cortese R. 2008. Reversine, a novel Aurora kinases inhibitor, inhibits colony formation of human acute myeloid leukemia cells. Mol. Cancer Ther. 7:1140–1149 10.1158/1535-7163.MCT-07-2051 [DOI] [PubMed] [Google Scholar]
- De Antoni A., Pearson C.G., Cimini D., Canman J.C., Sala V., Nezi L., Mapelli M., Sironi L., Faretta M., Salmon E.D., Musacchio A. 2005. The Mad1/Mad2 complex as a template for Mad2 activation in the spindle assembly checkpoint. Curr. Biol. 15:214–225 10.1016/j.cub.2005.01.038 [DOI] [PubMed] [Google Scholar]
- DeLuca J.G., Gall W.E., Ciferri C., Cimini D., Musacchio A., Salmon E.D. 2006. Kinetochore microtubule dynamics and attachment stability are regulated by Hec1. Cell. 127:969–982 10.1016/j.cell.2006.09.047 [DOI] [PubMed] [Google Scholar]
- DeMoe J.H., Santaguida S., Daum J.R., Musacchio A., Gorbsky G.J. 2009. A high throughput, whole cell screen for small molecule inhibitors of the mitotic spindle checkpoint identifies OM137, a novel Aurora kinase inhibitor. Cancer Res. 69:1509–1516 10.1158/0008-5472.CAN-08-3133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditchfield C., Johnson V.L., Tighe A., Ellston R., Haworth C., Johnson T., Mortlock A., Keen N., Taylor S.S. 2003. Aurora B couples chromosome alignment with anaphase by targeting BubR1, Mad2, and Cenp-E to kinetochores. J. Cell Biol. 161:267–280 10.1083/jcb.200208091 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorer R.K., Zhong S., Tallarico J.A., Wong W.H., Mitchison T.J., Murray A.W. 2005. A small-molecule inhibitor of Mps1 blocks the spindle-checkpoint response to a lack of tension on mitotic chromosomes. Curr. Biol. 15:1070–1076 10.1016/j.cub.2005.05.020 [DOI] [PubMed] [Google Scholar]
- Emanuele M.J., Lan W., Jwa M., Miller S.A., Chan C.S., Stukenberg P.T. 2008. Aurora B kinase and protein phosphatase 1 have opposing roles in modulating kinetochore assembly. J. Cell Biol. 181:241–254 10.1083/jcb.200710019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Espeut J., Gaussen A., Bieling P., Morin V., Prieto S., Fesquet D., Surrey T., Abrieu A. 2008. Phosphorylation relieves autoinhibition of the kinetochore motor Cenp-E. Mol. Cell. 29:637–643 10.1016/j.molcel.2008.01.004 [DOI] [PubMed] [Google Scholar]
- Famulski J.K., Chan G.K. 2007. Aurora B kinase-dependent recruitment of hZW10 and hROD to tensionless kinetochores. Curr. Biol. 17:2143–2149 10.1016/j.cub.2007.11.037 [DOI] [PubMed] [Google Scholar]
- Glover D.M., Leibowitz M.H., McLean D.A., Parry H. 1995. Mutations in aurora prevent centrosome separation leading to the formation of monopolar spindles. Cell. 81:95–105 10.1016/0092-8674(95)90374-7 [DOI] [PubMed] [Google Scholar]
- Griffis E.R., Stuurman N., Vale R.D. 2007. Spindly, a novel protein essential for silencing the spindle assembly checkpoint, recruits dynein to the kinetochore. J. Cell Biol. 177:1005–1015 10.1083/jcb.200702062 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guimaraes G.J., Dong Y., McEwen B.F., Deluca J.G. 2008. Kinetochore-microtubule attachment relies on the disordered N-terminal tail domain of Hec1. Curr. Biol. 18:1778–1784 10.1016/j.cub.2008.08.012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartwell L.H., Weinert T.A. 1989. Checkpoints: controls that ensure the order of cell cycle events. Science. 246:629–634 10.1126/science.2683079 [DOI] [PubMed] [Google Scholar]
- Hauf S., Cole R.W., LaTerra S., Zimmer C., Schnapp G., Walter R., Heckel A., van Meel J., Rieder C.L., Peters J.M. 2003. The small molecule Hesperadin reveals a role for Aurora B in correcting kinetochore–microtubule attachment and in maintaining the spindle assembly checkpoint. J. Cell Biol. 161:281–294 10.1083/jcb.200208092 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hewitt L., Tighe A., Santaguida S., White A.M., Jones C.D., Musacchio A., Green S., Taylor S.S. 2010. Sustained Mps1 activity is required in mitosis to recruit O-Mad2 to the Mad1–C-Mad2 core complex. J. Cell Biol. 190:25–34 10.1083/jcb.201002133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howell B.J., Moree B., Farrar E.M., Stewart S., Fang G., Salmon E.D. 2004. Spindle checkpoint protein dynamics at kinetochores in living cells. Curr. Biol. 14:953–964 10.1016/j.cub.2004.05.053 [DOI] [PubMed] [Google Scholar]
- Jelluma N., Brenkman A.B., McLeod I., Yates J.R., III, Cleveland D.W., Medema R.H., Kops G.J. 2008a. Chromosomal instability by inefficient Mps1 auto-activation due to a weakened mitotic checkpoint and lagging chromosomes. PLoS One. 3:e2415 10.1371/journal.pone.0002415 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jelluma N., Brenkman A.B., van den Broek N.J., Cruijsen C.W., van Osch M.H., Lens S.M., Medema R.H., Kops G.J. 2008b. Mps1 phosphorylates Borealin to control Aurora B activity and chromosome alignment. Cell. 132:233–246 10.1016/j.cell.2007.11.046 [DOI] [PubMed] [Google Scholar]
- Jones M.H., Huneycutt B.J., Pearson C.G., Zhang C., Morgan G., Shokat K., Bloom K., Winey M. 2005. Chemical genetics reveals a role for Mps1 kinase in kinetochore attachment during mitosis. Curr. Biol. 15:160–165 10.1016/j.cub.2005.01.010 [DOI] [PubMed] [Google Scholar]
- Kallio M.J., McCleland M.L., Stukenberg P.T., Gorbsky G.J. 2002. Inhibition of aurora B kinase blocks chromosome segregation, overrides the spindle checkpoint, and perturbs microtubule dynamics in mitosis. Curr. Biol. 12:900–905 10.1016/S0960-9822(02)00887-4 [DOI] [PubMed] [Google Scholar]
- Kang J., Yu H. 2009. Kinase signaling in the spindle checkpoint. J. Biol. Chem. 284:15359–15363 10.1074/jbc.R900005200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang J., Chen Y., Zhao Y., Yu H. 2007. Autophosphorylation-dependent activation of human Mps1 is required for the spindle checkpoint. Proc. Natl. Acad. Sci. USA. 104:20232–20237 10.1073/pnas.0710519105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelly A.E., Funabiki H. 2009. Correcting aberrant kinetochore microtubule attachments: an Aurora B-centric view. Curr. Opin. Cell Biol. 21:51–58 10.1016/j.ceb.2009.01.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kemmler S., Stach M., Knapp M., Ortiz J., Pfannstiel J., Ruppert T., Lechner J. 2009. Mimicking Ndc80 phosphorylation triggers spindle assembly checkpoint signalling. EMBO J. 28:1099–1110 10.1038/emboj.2009.62 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kinoshita E., Kinoshita-Kikuta E., Takiyama K., Koike T. 2006. Phosphate-binding tag, a new tool to visualize phosphorylated proteins. Mol. Cell. Proteomics. 5:749–757 [DOI] [PubMed] [Google Scholar]
- Knowlton A.L., Lan W., Stukenberg P.T. 2006. Aurora B is enriched at merotelic attachment sites, where it regulates MCAK. Curr. Biol. 16:1705–1710 10.1016/j.cub.2006.07.057 [DOI] [PubMed] [Google Scholar]
- Kwiatkowski N., Jelluma N., Filippakopoulos P., Soundararajan M., Manak M.S., Kwon M., Choi H.G., Sim T., Deveraux Q.L., Rottmann S., et al. 2010. Small-molecule kinase inhibitors provide insight into Mps1 cell cycle function. Nat. Chem. Biol. 6:359–368 10.1038/nchembio.345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lampson M.A., Renduchitala K., Khodjakov A., Kapoor T.M. 2004. Correcting improper chromosome-spindle attachments during cell division. Nat. Cell Biol. 6:232–237 10.1038/ncb1102 [DOI] [PubMed] [Google Scholar]
- Lénárt P., Petronczki M., Steegmaier M., Di Fiore B., Lipp J.J., Hoffmann M., Rettig W.J., Kraut N., Peters J.M. 2007. The small-molecule inhibitor BI 2536 reveals novel insights into mitotic roles of polo-like kinase 1. Curr. Biol. 17:304–315 10.1016/j.cub.2006.12.046 [DOI] [PubMed] [Google Scholar]
- Li X., Nicklas R.B. 1995. Mitotic forces control a cell-cycle checkpoint. Nature. 373:630–632 10.1038/373630a0 [DOI] [PubMed] [Google Scholar]
- Liu D., Vader G., Vromans M.J., Lampson M.A., Lens S.M. 2009. Sensing chromosome bi-orientation by spatial separation of aurora B kinase from kinetochore substrates. Science. 323:1350–1353 10.1126/science.1167000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu D., Vleugel M., Backer C.B., Hori T., Fukagawa T., Cheeseman I.M., Lampson M.A. 2010. Regulated targeting of protein phosphatase 1 to the outer kinetochore by KNL1 opposes Aurora B kinase. J. Cell Biol. 188:809–820 10.1083/jcb.201001006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maciejowski J., George K.A., Terret M.-E., Zhang C., Shokat K.M., Jallepalli P.V. 2010. Mps1 directs the assembly of Cdc20 inhibitory complexes during interphase and mitosis to control M phase timing and spindle checkpoint signaling. J. Cell Biol. 190:89–100 10.1083/jcb.201001050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maresca T.J., Salmon E.D. 2009. Intrakinetochore stretch is associated with changes in kinetochore phosphorylation and spindle assembly checkpoint activity. J. Cell Biol. 184:373–381 10.1083/jcb.200808130 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maresca T.J., Salmon E.D. 2010. Welcome to a new kind of tension: translating kinetochore mechanics into a wait-anaphase signal. J. Cell Sci. 123:825–835 10.1242/jcs.064790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattison C.P., Old W.M., Steiner E., Huneycutt B.J., Resing K.A., Ahn N.G., Winey M. 2007. Mps1 activation loop autophosphorylation enhances kinase activity. J. Biol. Chem. 282:30553–30561 10.1074/jbc.M707063200 [DOI] [PubMed] [Google Scholar]
- Maure J.F., Kitamura E., Tanaka T.U. 2007. Mps1 kinase promotes sister-kinetochore bi-orientation by a tension-dependent mechanism. Curr. Biol. 17:2175–2182 10.1016/j.cub.2007.11.032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McIntosh J.R. 1991. Structural and mechanical control of mitotic progression. Cold Spring Harb. Symp. Quant. Biol. 56:613–619 [DOI] [PubMed] [Google Scholar]
- Meraldi P., Draviam V.M., Sorger P.K. 2004. Timing and checkpoints in the regulation of mitotic progression. Dev. Cell. 7:45–60 10.1016/j.devcel.2004.06.006 [DOI] [PubMed] [Google Scholar]
- Miller S.A., Johnson M.L., Stukenberg P.T. 2008. Kinetochore attachments require an interaction between unstructured tails on microtubules and Ndc80(Hec1). Curr. Biol. 18:1785–1791 10.1016/j.cub.2008.11.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Musacchio A., Salmon E.D. 2007. The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8:379–393 10.1038/nrm2163 [DOI] [PubMed] [Google Scholar]
- Nicklas R.B., Koch C.A. 1969. Chromosome micromanipulation. 3. Spindle fiber tension and the reorientation of mal-oriented chromosomes. J. Cell Biol. 43:40–50 10.1083/jcb.43.1.40 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petersen J., Hagan I.M. 2003. S. pombe aurora kinase/survivin is required for chromosome condensation and the spindle checkpoint attachment response. Curr. Biol. 13:590–597 10.1016/S0960-9822(03)00205-7 [DOI] [PubMed] [Google Scholar]
- Pinsky B.A., Biggins S. 2005. The spindle checkpoint: tension versus attachment. Trends Cell Biol. 15:486–493 10.1016/j.tcb.2005.07.005 [DOI] [PubMed] [Google Scholar]
- Rieder C.L., Palazzo R.E. 1992. Colcemid and the mitotic cycle. J. Cell Sci. 102:387–392 [DOI] [PubMed] [Google Scholar]
- Ruchaud S., Carmena M., Earnshaw W.C. 2007. Chromosomal passengers: conducting cell division. Nat. Rev. Mol. Cell Biol. 8:798–812 10.1038/nrm2257 [DOI] [PubMed] [Google Scholar]
- Santaguida S., Musacchio A. 2009. The life and miracles of kinetochores. EMBO J. 28:2511–2531 10.1038/emboj.2009.173 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt M., Budirahardja Y., Klompmaker R., Medema R.H. 2005. Ablation of the spindle assembly checkpoint by a compound targeting Mps1. EMBO Rep. 6:866–872 10.1038/sj.embor.7400483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimogawa M.M., Graczyk B., Gardner M.K., Francis S.E., White E.A., Ess M., Molk J.N., Ruse C., Niessen S., Yates J.R., III, et al. 2006. Mps1 phosphorylation of Dam1 couples kinetochores to microtubule plus ends at metaphase. Curr. Biol. 16:1489–1501 10.1016/j.cub.2006.06.063 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sironi L., Melixetian M., Faretta M., Prosperini E., Helin K., Musacchio A. 2001. Mad2 binding to Mad1 and Cdc20, rather than oligomerization, is required for the spindle checkpoint. EMBO J. 20:6371–6382 10.1093/emboj/20.22.6371 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sliedrecht T., Zhang C., Shokat K.M., Kops G.J. 2010. Chemical genetic inhibition of Mps1 in stable human cell lines reveals novel aspects of Mps1 function in mitosis. PLoS One. 5:e10251 10.1371/journal.pone.0010251 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Straight A.F., Cheung A., Limouze J., Chen I., Westwood N.J., Sellers J.R., Mitchison T.J. 2003. Dissecting temporal and spatial control of cytokinesis with a myosin II Inhibitor. Science. 299:1743–1747 10.1126/science.1081412 [DOI] [PubMed] [Google Scholar]
- Stucke V.M., Baumann C., Nigg E.A. 2004. Kinetochore localization and microtubule interaction of the human spindle checkpoint kinase Mps1. Chromosoma. 113:1–15 10.1007/s00412-004-0288-2 [DOI] [PubMed] [Google Scholar]
- Tanaka T.U., Rachidi N., Janke C., Pereira G., Gálová M., Schiebel E., Stark M.J., Nasmyth K. 2002. Evidence that the Ipl1-Sli15 (Aurora kinase-INCENP) complex promotes chromosome bi-orientation by altering kinetochore-spindle pole connections. Cell. 108:317–329 10.1016/S0092-8674(02)00633-5 [DOI] [PubMed] [Google Scholar]
- Taylor S.S., Hussein D., Wang Y., Elderkin S., Morrow C.J. 2001. Kinetochore localisation and phosphorylation of the mitotic checkpoint components Bub1 and BubR1 are differentially regulated by spindle events in human cells. J. Cell Sci. 114:4385–4395 [DOI] [PubMed] [Google Scholar]
- Tighe A., Staples O., Taylor S. 2008. Mps1 kinase activity restrains anaphase during an unperturbed mitosis and targets Mad2 to kinetochores. J. Cell Biol. 181:893–901 10.1083/jcb.200712028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trazzi S., Perini G., Bernardoni R., Zoli M., Reese J.C., Musacchio A., Della Valle G. 2009. The C-terminal domain of CENP-C displays multiple and critical functions for mammalian centromere formation. PLoS One. 4:e5832 10.1371/journal.pone.0005832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uchida K.S., Takagaki K., Kumada K., Hirayama Y., Noda T., Hirota T. 2009. Kinetochore stretching inactivates the spindle assembly checkpoint. J. Cell Biol. 184:383–390 10.1083/jcb.200811028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vader G., Lens S.M. 2008. The Aurora kinase family in cell division and cancer. Biochim. Biophys. Acta. 1786:60–72 [DOI] [PubMed] [Google Scholar]
- Vader G., Cruijsen C.W., van Harn T., Vromans M.J., Medema R.H., Lens S.M. 2007. The chromosomal passenger complex controls spindle checkpoint function independent from its role in correcting microtubule kinetochore interactions. Mol. Biol. Cell. 18:4553–4564 10.1091/mbc.E07-04-0328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanoosthuyse V., Hardwick K.G. 2009. A novel protein phosphatase 1-dependent spindle checkpoint silencing mechanism. Curr. Biol. 19:1176–1181 10.1016/j.cub.2009.05.060 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vigneron S., Prieto S., Bernis C., Labbé J.C., Castro A., Lorca T. 2004. Kinetochore localization of spindle checkpoint proteins: who controls whom? Mol. Biol. Cell. 15:4584–4596 10.1091/mbc.E04-01-0051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villa F., Capasso P., Tortorici M., Forneris F., de Marco A., Mattevi A., Musacchio A. 2009. Crystal structure of the catalytic domain of Haspin, an atypical kinase implicated in chromatin organization. Proc. Natl. Acad. Sci. USA. 106:20204–20209 10.1073/pnas.0908485106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wan X., O’Quinn R.P., Pierce H.L., Joglekar A.P., Gall W.E., DeLuca J.G., Carroll C.W., Liu S.T., Yen T.J., McEwen B.F., et al. 2009. Protein architecture of the human kinetochore microtubule attachment site. Cell. 137:672–684 10.1016/j.cell.2009.03.035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weaver B.A., Cleveland D.W. 2006. Does aneuploidy cause cancer? Curr. Opin. Cell Biol. 18:658–667 10.1016/j.ceb.2006.10.002 [DOI] [PubMed] [Google Scholar]
- Welburn J.P.I., Vleugel M., Liu D., Yates J.R., III, Lampson M.A., Fukagawa T., Cheeseman I.M. 2010. Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore-microtubule interface. Mol. Cell. 38:383–392 10.1016/j.molcel.2010.02.034 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu Q., Zhu S., Wang W., Zhang X., Old W., Ahn N., Liu X. 2009. Regulation of kinetochore recruitment of two essential mitotic spindle checkpoint proteins by Mps1 phosphorylation. Mol. Biol. Cell. 20:10–20 10.1091/mbc.E08-03-0324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Z., Kenny A.E., Brito D.A., Rieder C.L. 2009. Cells satisfy the mitotic checkpoint in Taxol, and do so faster in concentrations that stabilize syntelic attachments. J. Cell Biol. 186:675–684 10.1083/jcb.200906150 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Y., Chen R.H. 2006. Mps1 phosphorylation by MAP kinase is required for kinetochore localization of spindle-checkpoint proteins. Curr. Biol. 16:1764–1769 10.1016/j.cub.2006.07.058 [DOI] [PubMed] [Google Scholar]