Abstract
With a realistic threat against biodiversity in rain forests and in the sea, a sustainable use of natural products is becoming more and more important. Basic research directed against different organisms in Nature could reveal unexpected insights into fundamental biological mechanisms but also new pharmaceutical or biotechnological possibilities of more immediate use. Many different strategies have been used prospecting the biodiversity of Earth in the search for novel structure–activity relationships, which has resulted in important discoveries in drug development. However, we believe that the development of multidisciplinary incentives will be necessary for a future successful exploration of Nature. With this aim, one way would be a modernization and renewal of a venerable proven interdisciplinary science, Pharmacognosy, which represents an integrated way of studying biological systems. This has been demonstrated based on an explanatory model where the different parts of the model are explained by our ongoing research. Anti-inflammatory natural products have been discovered based on ethnopharmacological observations, marine sponges in cold water have resulted in substances with ecological impact, combinatory strategy of ecology and chemistry has revealed new insights into the biodiversity of fungi, in depth studies of cyclic peptides (cyclotides) has created new possibilities for engineering of bioactive peptides, development of new strategies using phylogeny and chemography has resulted in new possibilities for navigating chemical and biological space, and using bioinformatic tools for understanding of lateral gene transfer could provide potential drug targets. A multidisciplinary subject like Pharmacognosy, one of several scientific disciplines bridging biology and chemistry with medicine, has a strategic position for studies of complex scientific questions based on observations in Nature. Furthermore, natural product research based on intriguing scientific questions in Nature can be of value to increase the attraction for young students in modern life science.
Keywords: Pharmacognosy, Geodia, COX-2, ChemGPS-NP, Chemical space, Phylogeny, Cyclotide, Viola, Truffle, Tuber, Lateral gene transfer, Trypanosoma
Modern life science
Many situations in the society of today have shown the need for science to achieve a sustainable future development. Among recent issues in this line we find the problems and implications related to global warming, such as the future demand of food and fodder for life stock. Other important problems are the threats against the biodiversity in the tropical rain forests. This is also the case for shallow water coral reefs and mangrove swamps, which are both threatened by higher temperature and a lowered pH due to an increase of carbon dioxide in the atmosphere. Another serious development is the increasing resistance of pathogenic bacteria against the available antibiotic used in treating a number of severe diseases.
In today’s society we see a renewed interest in green chemistry and a sustainable use of natural products. These tendencies are evident also in the search for new pharmaceutical and biotechnological possibilities, where the development of multidisciplinary incentives will probably be necessary. Such multidisciplinary research, for example directed on different organisms, could provide us with a more complete picture of the complex workings of the Earths biome, but also give unexpected insights into fundamental biological mechanisms of more immediate use.
Despite this, concern has been raised over the lowered interest in pursuing higher education in science among young people. In recent years the European commission has been investigating attitudes towards science and technology among European citizens. Their latest report “Young people and Science” (EC 2008) summarizes findings from a survey comprising interviews from close to 25,000 Europeans aged 15–25 from all 27 EU membership states, probing the views and interest of young citizens, regarding a future involvement in science and technology. In general the report presented a positive view concerning science and technology and an awareness of the fact that science is important for the development of the future society. However, in the responders own selection of study objects for the future, interest in natural science showed a surprisingly low preference in comparison to that of social sciences or economics.
This is daunting, as a future society will need politicians and citizens that have enough scientific knowledge to make critical decisions about important questions such as global warming, rain forest destruction, stem cell research, and nuclear power waste.
Furthermore, it seems that the number of students pursuing higher studies in basic science such as physics, chemistry, and biology are decreasing, while mirrored by an increased interest for applied aspects such as environmental chemistry, material physics, and biotechnology. The reason for a lowered interest in basic science has been discussed in recent editorials of the prestigious scientific journal Science (Cech and Kennedy 2005; Augustine 2008). Concern was raised about education methods and the failure to encourage the students for a scientific career but also about spreading knowledge about the importance of basic science in society.
We would argue that the multidisciplinary subject Pharmacognosy, as one of several scientific disciplines, has a strategic position in the studies of scientific questions based on observations in Nature. In addition to that, such a subject that itself is bridging biology and chemistry with medicine can be of value in the joint efforts to attract young students for studies in modern life science.
Natural products in relation to modern life science
For centuries the human race has constructed ingenious devices to solve problems and make products for our needs. Although this progress has resulted in the present day wealth and prosperity, there has also been another side to this, as discussed in the sections above. With the increasing understanding of why we find ourselves in this predicament, it is not surprising that along many paths of modern life science research we find attempts to understand and build on more natural or environmental friendly methods. However, the complexity of such development is shown by the old use of fossil fuels, which transcended in a first hype of technologically trivial but energetically unfavourable ethanol, and now exploring other paths more in line with a sustainable development and acceptable ethics. In the field of pharmaceutics, the synthetic chemistry of the twentieth century is gradually made more efficient and precise, but also gradually changing into bio-technological applications and a return to the infinitely more variable and complex chemistry of Nature.
Organisms in Nature produce secondary metabolites with the specific purpose to gain evolutionary advantages in the competition for example living space and in the search for nutrients. During evolution different enzymatic machineries have been developed, which has directed the biosynthesis of different molecular frameworks of secondary metabolites. With an estimated number of more than 300,000 species of plants and probably close to two million species of fungi, insects, and various marine organisms—not to mention bacteria—the biodiversity of Nature remains an unparalleled reservoir of biological and chemical diversity. However, most of the biodiversity is as yet unexplored.
Several different strategies, based on ethnopharmacological, ecological or toxicological observations together with random screening, and more recently phylogenetic frameworks, have been used in prospecting this biodiversity in the search for unique structure–activity relationships, and resulted in revolutionary discoveries in medicine. The ethnopharmacological approach has been the platform for research programs with the objective to study plants used in traditional medicine in different cultures. With the increased awareness of indigenous peoples rights to their traditional knowledge, established in several conventions, it has already become increasingly complicated to perform and finance such research. However, in the extension this will also develop more respectful insights into sharing of resources and traditional knowledge. Due to a very limited access to marine organisms for people in old times few reports exist where marine organisms have been used in traditional medicine. Bio-prospecting the marine environment is now rapidly expanding, in many cases driven by ecological observations, and the number of novel bioactive molecules isolated from marine organisms is increasing. Of importance for future natural products research is that in recent years microorganisms found in or associated with marine sponges have been shown to be the actual producers of isolated novel compounds. This can result in new methods for cultivation of these types of organisms as a sustainable resource of the bioactive compound.
Modern biotechnology has developed new possibilities and tools for elucidation of many biosynthetic pathways for natural products where enzymes involved have been isolated and characterized and in some cases even cloning of the coding genes. This development has created new strategies like combinatorial biosynthesis, which can lead to entirely new classes of polyketides synthesized by very large enzymes and non-ribosomal polypeptides. Another possibility to create new chemical structures, based on a natural scaffold, is the bioengineering of cyclic peptides such as cyclotides. A natural cyclotide consists of 28–37 conventional amino acids connected by amide bonds to a cyclic ring stabilized by three disulfide bridges which creates six loops of different size. It has been demonstrated that the amino acids in the different loops can be exchanged for new amino acids and in this way novel structure–activity relationships can be studied.
Instead of a specific observation in Nature, which defines the scientific question and strategy for a research project, extracts from randomly selected organisms can be screened for a specific activity. This approach has been successful in the discovery of novel antibiotics but less successful in other areas, even when technologically advanced methods such as combinatorial synthesis and industrial high throughput screening have been applied. One reason is that such endeavours soon become a numbers game, and hence methods to limit the number of compounds for testing and selecting the most promising candidates are imperative. One way to decrease the complexity and size of compound libraries necessary to transverse is to apply tools such as chemography to navigate chemical space (Oprea and Gottfries 2001). A recently described tool able to handle also the complex molecules of natural origin is the ChemGPS-NP global map of the biologically relevant volumes of multidimensional chemical property space. Combined with phylogenetic studies, which applies to the relationship of evolution of organisms, these methods can be used in silico to explore and attempt to understand how biosynthetic ability has developed among different organisms. Understanding such processes may lead to conceptually novel models for prediction and selection of organisms that produce compounds with specific biological activity but also to predict new targets, as discussed below.
A Linnaean view on science
There are many similarities with the present drive for more applied research and utilizable results, with the predominant views in the Linnaean eighteenth century Europe. In that époque the states and governments proved eager to explore Nature for its better use. The way this was pursued was through intense support of research funds to the universities, the erection of governmental offices and boards to survey and supervise mining, trade and agriculture—and as we can see on the numerous TV shows with research themes: a societal glorification of scientists. The major difference might be, that today the public appears less impressed and eager to participate. Perhaps the last centuries’ scientific and engineering advances have made us less prone to marvel at novel discoveries?
Linnaeus used a holistic approach in both his teaching and research. Together with his students he produced 186 theses during the years 1744–1776 in many different disciplines for example De methodo investigandi vires medicamentorum chemica (Regarding the chemical method to investigate the virtues of drugs) by Hiortzberg (1765). The multidisciplinary approach is obvious in his Materia Medica, first published in 1749, where he applies a surprisingly modern way of describing different aspects of medicinal plants and their preparation, use and effect.
Thunberg and Wahlenberg were the Linnaean successors at Uppsala University, followed in 1893 by Rosendahl who was first appointed to a chair in general and experimental Pharmcodynamics with Pharmacognosy. In the beginning of the twentieth century Rosendahl became the first Swedish professor in “Botany and Pharmacognosy” then affiliated to the Royal Institute of Pharmacy in Stockholm. During the first part of the twentieth century the Swedish education, training, and research in Pharmacognosy was botanically oriented with a focus on morphology and anatomy and the purpose of identification and quality control of crude drugs. During the past 60 years the subject of Pharmacognosy has developed towards a multidisciplinary scientific subject combining natural product chemistry with biology, and since 1973 forms a part of the revived Faculty of Pharmacy at Uppsala University.
Modern pharmacognosy—an integrated discipline
Most important in our research today is the development of strategies for selection, isolation and characterisation with the objective to discover unique bioactive chemical structures with drug potential, and to reveal unknown targets, by studying the evolutionary structure–activity optimization in Nature. In addition to the possibility to discover new drug candidates for drug development, bioactive natural projects have potential as pharmacological tools, intermediates, or templates for synthesis of drugs. The increased uses of herbal remedies, which contain complex mixtures of natural products, need intensified scientific studies to establish efficacy and safety of these types of products as well as clinical studies.
Thus, we would argue that our research represents a modernization and renewal of a venerable proven science, Pharmacognosy, without losing (or ignoring) the traditions in the field. With the increasing interest for environmental aspects, green chemistry, and a sustainable use of natural products, this renewal could have a strategic position in bridging chemistry and biology.
The interdisciplinary nature of the subject is explained in the model below (Fig. 1) where the starting point is an observation of an organism showing an interesting biological activity. The correlation between the chemical structures responsible for the shown bioactivity needs to be studied to understand the observation on a molecular level using both in vivo, in vitro and in silico methods (Larsson 2007).
Fig. 1.
Explanatory model the interdisciplinary nature of pharmacognosy interpreted in an explanatory model presented by Larsson and co-workers (2008). In this model a clearly defined role is presented for aspects of informatics, including bio- and chemoinformatics. Figure by Sonny Larsson, copyright Phytochemistry Letters, reproduced with permission
In the following sections, different parts of this model are explained in more detail by description of our research. Based on an observed ethnopharmacological use of plants against inflammation in depth studies of pure natural products against colon cancer are now in focus. Ecological relationships among organisms have resulted in studies of antifouling properties of molecules isolated from marine sponges and chemical profiling of fungi. Another project is focused on bioengineering of circular proteins, so called cyclotides, to create new structure–activity relationships. Novel strategies are developed for efficient prediction and selection of organisms and molecules and bioinformatic tools to predict novel targets based on lateral gene transfer.
Natural products with anti-inflammatory and anti-tumor activity
A scientific platform has been built in our long-term research on anti-inflammatory natural products as demonstrated in a number of publications and doctoral theses. Our earlier research started with ethnopharmacological observations and information of the use of medicinal plants against different forms of inflammation in different cultures. Many different chemical structures have been discovered, and chemically and pharmacologically characterised using bioassay-guided isolation procedures. In vivo methods such as rat paw and mouse ear oedema was used and later followed by in vitro enzyme and cell based methods. Two systems have been established to enable investigations of the effects of natural compounds on COX-2. The first method developed was an in vitro method suitable for measuring inhibition of COX-2 catalysed prostaglandin E2 biosynthesis, based on scintillation proximity assay technology (Huss et al. 2002). The second system comprises a cell model, suitable for studying the effects of compounds on COX-2 and inducible nitric oxide synthase (iNOS) at different cellular levels, including the effects on mRNA, protein, prostaglandin E2, and nitric oxide levels (Huss 2003).
In later years the project has developed towards enzyme inhibitors related to anti-tumor activity, especially in colon cancer. It has been shown that the process of inflammation and expression of cyclooxygenase-2 is important in colon carcinogenesis. Another important factor is diet. Many food phytochemicals have been shown to exert anti-inflammatory activity in vitro, and may act as cancer chemopreventive agents (Kim et al. 2003; Murakami and Ohigashi 2007). A vegetarian diet rich in phytochemicals may prevent colon carcinogenesis by affecting biochemical processes in the colonic mucosa. We have shown that intact faecal water (water phase) samples from human volunteers significantly decreased prostaglandin production and COX-2 expression in colonic cells. NMR spectroscopy and multivariate data analysis were later used for further analysis of the composition of the faecal waters and to trace the COX-2 inhibiting activity (Pettersson et al. 2008a, b). The wealth of different natural products with experimentally demonstrated COX inhibitory effects and an urge to understand and characterize their structural diversity was the starting-point for the application of chemography in our natural products research.
The identification from chemographic analyses of some specific groups of compounds, including a set of cardiac glycosides, as being of prime importance was further established in a screening of a large number of natural products for activity against colorectal cancer where several cardiac glycosides showed significant activity. This activity was further confirmed in primary cells from colon cancer patients. Cardiac glycosides have been reported to exhibit cytotoxic activity against several different cancer types, but studies against colorectal cancer are lacking. Drugs for clinical treatment of colon cancer are usually used in combination to overcome the problem with drug resistance and to increase the activity. Therefore, selected cardiac glycosides were tested in combination with four clinically relevant standard cytotoxic drugs (5-fluorouracil, oxaliplatin, cisplatin, irinotecan) to screen for synergistic effects. The combination of digitoxin and oxaliplatin exhibited synergism including the otherwise highly drug-resistant HT29 cell line (Felth et al. 2009). In depth studies are now in progress necessary to understand these effects on a molecular level.
Marine organisms—substances with ecological impact
The project is related to the sustainable use of natural products and development of “green chemistry”. The future society needs biodegradable natural products with specific actions and low residence times, e.g. for control of fouling organisms in the marine environment. Marine organisms have shown to contain a wealth of bioactive secondary metabolites with potential for new pharmaceutical or biotechnological applications. Marine sponges produce substances that have a key role in the defence against pathogens, parasites, predators and biofouling organisms.
In our earlier research we performed screening for biological activity, using both in vivo and in vitro methods, of marine organisms on the west coast of Sweden. This work resulted in several publications and two doctoral theses (Andersson 1987; Lidgren 1989). Extracts from the marine sponge Geodia barretti, distributed in the Norwegian fjords and the Swedish Kosterfjord, showed significant biological activity in several of the assays. These results combined with the observation that Geodia barretti possesses an almost completely fouling-free body surface prompted a study of possible chemical defense against and identification of pharmacologically active molecules.
In this project we have isolated, characterized and synthesized two cyclopeptides from the marine sponge Geodia barretti, with effect on cyprid larvae from Balanus improvisus that could explain why this sponge is free from ongrowth of other organisms. The objective of the studies was to further explore the chemical diversity in Geodia barretti. Furthermore, the aim was to understand the biological activity on different targets and to evaluate if the compounds produced by the sponge act in concert, either by synergistic or cooperative action, and to investigate a possible bacterial origin of the compounds.
For isolation of minor secondary metabolites state of the art methods for chemical analysis have been used, such as LC–MS, MS/MS and 2D-NMR. For establishing biological activity a barnacle settlement assay in vitro has been used to evaluate the effect of the isolated compounds on the behaviour on cyprid larvae. The brominated cyclopeptides have also been tested further for affinity to human serotonin receptors using an in vitro radioligand-binding assay based on displacement of radioligands from human 5HT-receptors expressed in HEK-293 cell membranes. A remotely operated vehicle has been used to evaluate the release of secondary metabolites in ambient water. Mass spectrometry has been used to understand the interaction of the different metabolites and dose–response calculations have been made to show the synergistic activity, and different standard protocols have been used to cultivate bacterial strains associated with Geodia barretti (Sjögren 2006; Fig. 2).
Fig. 2.
Geodia reef picture taken by remote operated vehicle at 120 m depth showing fouling-free marine sponges of the species Geodia barretti and G. macandrevia living in the Koster Fjord at the Swedish West-coast. Photo by Thomas Lundälv, Tjärnö Marine Biology Laboratory, Sweden, used with permission
A novel dibrominated cyclopeptide, bromobenzisoxalone barettin, was isolated from Geodia barretti using repeated extraction with acetonitrile and water, desalted with RP-SPE and then fractionated using RP-HPLC and detected with ESIMS. High resolution MS and 2D-NMR were used for elucidation of the chemical structure. The new compound displayed settlement inhibition of barnacle larvae with an EC50 value of 15 nM (Hedner et al. 2008).
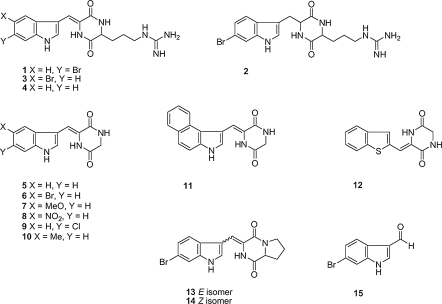
The brominated cyclodipeptides barettin and 8,9-dihydrobarettin have previously been shown to inhibit settlement of barnacle larvae in a dose-dependent manner in concentrations ranging from 0.5 to 25 μM. In order to further establish the molecular target and mode of action of these compounds, we investigated their affinity to human serotonin receptors. The two cyclopeptides selectively interacted with the serotonin receptors 5-HT2A, 5-HT2C and 5-HT4 at concentrations close to that of endogenous serotonin. Surprisingly, the novel barettin analogue did not show any effect on these receptors (Hedner et al. 2006; Hedner 2007). To gain further knowledge about the structure–activity basis for the shown antifouling activity and the effect on specific serotonin receptors, different analogs, based on the chemical core structures of combinations of barettin and dipodazin, were designed and synthesised (Sjögren et al. 2006; Fig. 3).
Fig. 3.
Structures compilation of structures of barettin (1), dipodazine (5) and 13 analogues synthesized and tested by Sjögren and co-workers (2006)
We have recently shown that the two congeneric defence cyclodipeptides, barettin and 8,9-dihydrobarettin, produced by the coldwater marine sponge Geodia barretti act in synergy to deter larvae of surface settlers. An in situ sampling using the remotely operated vehicle at a depth of 123 m revealed that the sponge continuously releases these two compounds to the ambient water. We suggest that the chemical defence in Geodia barretti involves synergistic action, with congeneric compounds produced by the same enzymatic pathway, where one of the targets is a 5-HT receptor and that the synergy of barettin and 8,9-dihydrobarettin have developed to reduce the cost for the sponge to uphold its chemical defence.
In conclusion, our results emphasize that also marine organisms in cold waters merits further attention. The performed studies of the cold water marine sponge Geodia barretti has not only given an increased understanding of antifouling substances and their interaction with cyprid larvae, but also further knowledge about cold water marine sponges and their possible role in the very old reef like fields of sponges, which have been pointed out as important hotspots of biodiversity in the North Atlantic (Klittgaard 1995).
Biodiversity of fungi—combinatory strategies of ecology and chemistry
Fungi are known for their peculiar chemistry, differing from that of both animals and higher plants. The secondary metabolites of fungi are produced to aid establishment in the various ecological niches of different fungal life forms. Several of these metabolites have played important roles as fungi-derived drugs revolutionizing the treatment of several serious conditions in man. Two such examples are penicillin and cyclosporin. In the antibiotic peptide penicillin, a β-lactam (4-membered cyclic amide) ring interferes with the synthesis of the bacterial cell wall. Fungi occasionally produce unusual amino acids and unlike animals and higher plants, fungi may not only produce and use the l- but the d-enantiomers in their biosynthesis of secondary metabolites resulting in anomalous compounds. One example of this is the immunosuppressant agent cyclosporin, which is essential during organ transplants to prevent rejection of the new organ by the immune system. Cyclosporin contains a number of unusual amino acids as well as both the l- and d-enantiomers of alanine (Traber et al. 1989). We still know relatively little about the general nature of fungal chemistry and in addition to this we only seem to know a fraction of the estimated number of existing fungal species on Earth. Today approximately 14,000 species of macrofungi are known to science, but estimations show that this may well be only a tenth of the actual number of all living macromycete species (Hawksworth 2001). For fungi in the widest sense, the total number of species on Earth has been estimated to 1.1 million (Mueller et al. 2007). Within the fungal kingdom lies a vast, and to a large extent, still unexplored potential for future pharmaceutical drugs.
In the section Fungi in Linnaeus’ Materia Medica (1749), the genus Lycoperdon is mentioned as an aphrodisiac and remedy against flatulence. Most probably this passage refers to what today is known as the deer truffle, Elaphomyces granulatus, earlier known under the name Boletus cervini. In more recent sources the deer truffle has been reputed for its abilities to induce oestrus manifestation in cows (Klintberg 1998), forming a somewhat unsuspected link to the Linnaean indication. The edible truffles of the genus Tuber have historically been used as an aphrodisiac by both men and women and in modern science small amounts of the steroid androstenol has been found in mature specimens of the black truffle Tuber melanosporum. Androstenol is a major component of the boar pheromone, excreted in the saliva during mating (Claus et al. 1981).
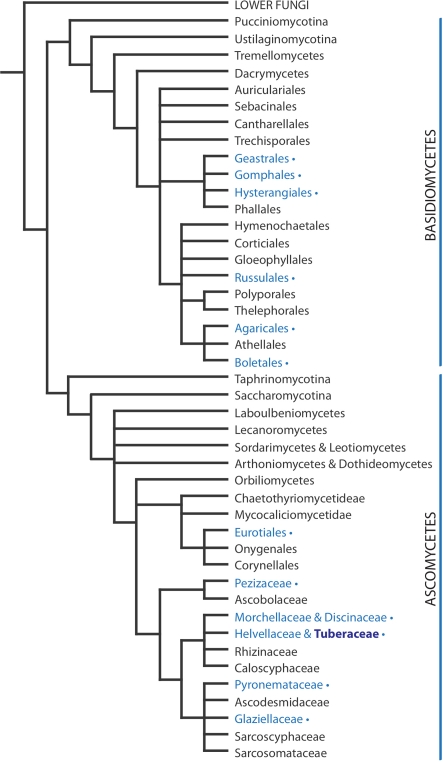
Truffles are an intricate fungal life form occurring in both the two major groups of macromycetes: asco- and basidiomycetes (see Fig. 4). Only within the ascomycetes, species that develop truffle fruit bodies are estimated to have evolved at least fifteen times in multiple families (Læssøe and Hansen 2007). Thus, from a modern phylogenetic perspective, truffles are regarded as a life form rather than a specific group of fungi. Despite this, all truffle species, regardless of phylogenetic placement, live in mycorrhizal symbiosis with vascular plants. Due to their hypogeous nature and subsequent loss of active spore dispersal, all truffle species are also dependant on an animal vector for spore dispersal. The evolution of a hypogeous fruit body might be an adaptation to dry climatic conditions, but the relationship between the hypogeous fungus, the tree and the animal vector seem to be imperative for the life and distribution of this life form suggesting a more sophisticated fungal niche against vertebrates. The dispersal vectors may be mice, squirrels, deer or wild boars (Trappe and Castellano 1991). Consequently the truffles, i.e. the hypogeous spore-producing fruiting bodies, have developed a diverse set of volatile substances to attract these vectors, and to make them unearth and eat the truffles, thereby spreading the spores. Depending on the animal vector’s lifestyle and ecology, these spores may be deposited in the shape of fecal pellets close to the roots of mycorrhizal trees (Trappe and Castellano 1991). Truffles also show high levels of endemism (Mueller et al. 2007), which may be explained by mammalian fecal pellets being their main mean of spore dispersal (Fig. 5).
Fig. 4.
Phylogeny of fungi phylogeny of asco- and basidiomycetes with focus on taxa with hypogeous fruiting bodies, truffles, based on a combination of several published studies (Læssøe and Hansen 2007; Hibbet et al. 2007; Celio et al. 2006). Taxa labeled in blue with ∙ indicate evolutionary groups that have developed truffles. The most well known truffle-producing family, Tuberaceae, comprising the edible white and Burgundy truffles is further emphasized in bold lettering. Figure by Christina Wedén and Anders Backlund
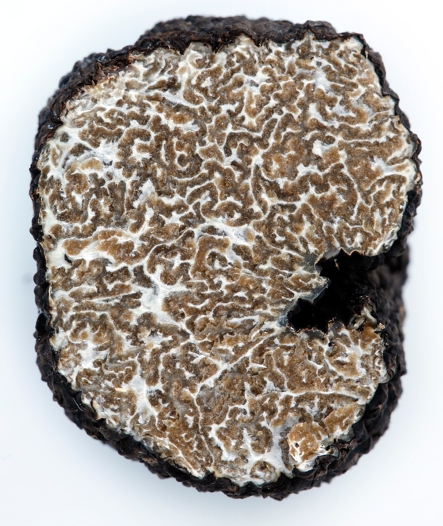
Fig. 5.
Section of truffle the characteristic scent of the Burgundy truffle does not develop until the entire spore mass has matured, indicating a sophisticated chemical control mechanism. In the picture above we see a fruit body of the truffle Tuber aestivum from the island of Gotland with completely matured spores, showing the interior with a characteristic marbled brown gleba. Photo by Erika Lidén ©2009, used with permission
The ascomycete truffle species Tuber aestivum provides a good model, with a narrow genetic variation (Wedén et al. 2005), enabling method development and an understanding of the type and range of intraspecific variation in secondary chemistry. On the Swedish island of Gotland, the population of T. aestivum poses several interesting challenges. After the last glaciation, the highest parts of the island of Gotland reappeared as an island above what was then a fresh water lake approximately 11,600 years ago. The European hazel, Corylus avellana was the first possible Tuber aestivum host tree to become establish on Gotland some 9,000 years ago (Påhlsson 1977). This detailed and well studied history of Gotland’s quartenary geology provides us with unique parameters for the calculation of the date for, and rate of, genetic divergence of Tuber aestivum, as well as possible island biogeographic hypotheses for the mode of introduction. Population dynamics and dispersal patterns on newly introduced land and over large bodies of water might also be tested.
By analysing secondary chemistry in combination with DNA sequence data we aim to find possible variation and congruencies within truffle populations. Tuber aestivum on the island of Gotland has been found to consist of a closely related population possibly indicating a single introduction or several introductions from a similar source (Wedén et al. 2004). Genotypes showed a normal distribution indicating sexual reproduction (Wedén 2004). Despite the genetic homogeneity, local truffle hunters have reported differences in the organoleptic properties of truffle fruit bodies between different sites. In an ongoing study of the Gotland T. aestivum population, we have aimed to investigate if chemical differences could be site specific or correlated to genetic traits. The molecular data set consists of sequences of the internal transcribed spacer region (ITS) from fresh fruit bodies of T. aestivum from 15 different localities on Gotland. Ethanol extracts of the same fruit bodies have then been analysed with LC–MS to create a data set of chemical profiles corresponding to that described by Ekenäs and co-authors (2009).
It is noteworthy that the volatile substances attracting the dispersal vectors are only produced when spore maturity in the truffle has reached 100% (Wedén 2004), thereby preventing premature dispersal. For some truffle species, this signalling mechanism acting as an olifactory triggering stimuli in vertebrates in nature, also appeals to the human pallet. A few truffle species are appreciated as delicacies, due to their appealing, intense and complex aroma. Volatile and non-volatile components of truffles are being studied to understand the nature of these interactions (Fig. 5).
Bioengineering—chemical structure and biological activity of cyclotides
Plant peptides and proteins may be considered an overlooked source for new chemical entities and novel bioactivites compared to low molecular natural products. The reason for this seems to be based on tradition and biased by the most commonly used techniques for natural products research. For example, most researchers in the field tend to avoid water as an extraction solvent, which results in a discrimination of peptides and proteins already at an initial stage. However, during the last decades, the number of reported plant peptides has grown substantially, and the field is about to mushroom. In our laboratory, plant peptides was a topic of research already in the 1950s through to the early 1980s, in the form of Professor Gunnar Samuelssons’ pioneering studies of mistletoe toxins (Samuelsson 1958). In the mid 1990s we made an attempt to assess plant peptides more broadly, with the design of an isolation protocol directly designed for their isolation (Claeson et al. 1998). One of the results of this effort was the discovery of a set of macrocyclic peptides in the plant family Violaceae (Göransson et al. 1999).
Strikingly, the peptides we characterised in Violaceae were found to be nearly identical with one of the most intriguing examples of pharmacognostic research in general, and plant peptides in particular, namely the peptide Kalata B1. The discovery of Kalata B1 was based on the ethnopharmacological use of the plant Oldenlandia affinis. During a Red Cross mission in Congo, the Norwegian physician Lorents Gran experienced a high frequency of complicated deliveries due to the use of this plant, which was locally known as “Kalata-Kalata” (Gran 1973a, b). Native women secretly used a decoction of this plant to facilitate childbirth, which they sipped as a tea but also applied directly at the birth canal (Gran et al. 2000). It induced extremely strong uterine contractions, which sometimes developed into cervical spasms necessitating acute caesarean section. On a second session in the same area some years later, the identity of the plant was revealed to Gran, who, triggered by his observations, brought the plant back to Norway for biological and chemical characterisation. He found that a peptide, Kalata B1, was the active component. The complete sequence and the cyclic structure was however not determined for more than 20 years later (Saether et al. 1995).
At the time of our report of the first cocktail of “kalata-like” peptides in Violaceae, four similar peptides had been reported in the literature as the serendipitous discoveries of three independent groups. When including those peptides in a sequence comparison, i.e. the anti-HIV circulins A and B, the neurotensin binding inhibitor cyclopsychotride A and the partially characterised violapeptide I, it was clear that they fell in two subgroups based on sequence similarity. This was confirmed by the landmark study by David Craik and coworkers that introduced the name cyclotides (as an acronym for cyclic peptides), and the two subfamilies were termed Möbius and bracelet cyclotides, based on the presence and absence, respectively, of a cis-proline bond.
Today, around 150 cyclotides have been reported from species of three plant families, Violaceae, Rubiaceae and Cucurbitaceae. The family Violaceae seems to be particularly rich in these proteins (Göransson et al. 2003, 2004b; Herrmann et al. 2008; Ireland et al. 2006; Simonsen et al. 2005), and a single Violaceae species may contain more than 60 different cyclotides. It has been suggested that there might be >9,000 cyclotides in the Violaceae alone (Simonsen et al. 2005).
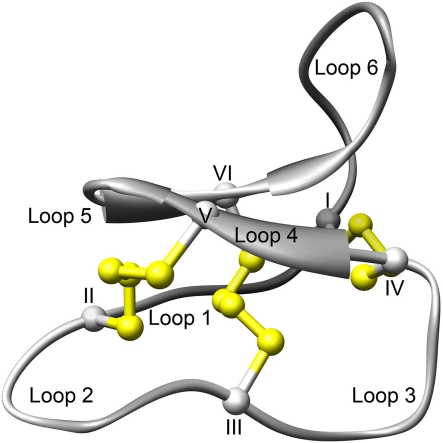
The structure of a representative cyclotide, varv F (Wang et al. 2009), is shown in Fig. 6. In addition to the amide bond that cyclise the backbone, cyclotides contain three stabilising disulfide bonds in a knotted arrangement, i.e. two disulfides form a ring together with their connecting protein backbone, which is threaded by the third disulfide (Göransson and Craik 2003; Rosengren et al. 2003; Saether et al. 1995). Together these features define the cyclic cystine knot (CCK) motif, and make cyclotides extraordinary stable protein structures (Colgrave and Craik 2004). In fact, they are impervious to enzymes, and, as indicated by the native’s use of decoction as an extraction method, cyclotides also withstand boiling water.
Fig. 6.
Cyclotide structure. The figure shows a ribbon representation of the crystal structure of varv peptide F (pdbcode 3E4H). Note the seamless peptide backbone and the three disulfides bonds (in yellow) that form the cystine knot: CysI–CysIV, II–V, III–VI. The six loops are marked with roman numerals. Figure by Ulf Göransson
The sequences between the cysteines, referred to as loops, are more or less variable. Notably, the loops confer additional characteristics that differentiate the typical Möbius and bracelet cyclotides; in particular the bracelet cyclotide contain a longer loop 5 that forms a short α-helix and a cluster of cationic residues in loop 3. As the cyclotide family grows however, more and more peptides are discovered that contain features from both subfamilies, so called hybrids. In addition, the peptides from the Cucurbitaceae plant family form a third subfamily that basically only has the CCK motif in common with Violaceae and Rubiaceae cyclotides. The meagre sequence similarity might indicate that these peptides evolved convergently to Möbius and bracelet cyclotides (instead they show homology to the linear trypsin inhibitory peptides from Cucurbitaceae).
As mentioned above, the early cyclotide discoveries were due to their biological effects. Although not all of the around 150 cyclotides that we know today have been tested in any assay, the number and types of biological effects are impressive. Besides being uterotonic, anti-HIV, haemolytic and neurotensin binding inhibitory, the list now includes antimicrobial (Tam et al. 1999), antifouling (Göransson et al. 2004a), antihelmintic (Colgrave et al. 2009), molluscicidal (Plan et al. 2008), cytotoxic (Lindholm et al. 2002; Svangård et al. 2004) and insecticidal (Jennings et al. 2001) activities. The latter effects are some of the most well studied and interesting effects: the cytotoxic effect together with the haemolytic effect have been the focus for detailed structure activity studies (Göransson et al. 2009; Herrmann et al. 2006), and the discovery of their insecticidal effect likely revealed cyclotides’ role in planta. Cyclotides’ mechanism of action is however yet unknown, but evidence is accumulating showing that membrane interactions followed by membrane pore or fissure formation are involved (Kamimori et al. 2005; Shenkarev et al. 2006; Svangård et al. 2007), which could provide an explanation to several of the reported effects.
Combined with the extraordinary CCK motif—with its conserved scaffold that can be dressed with variable loop sequences—the demonstrated biological activity of the cyclotides make them a first class target for protein engineering. To this end, inherent activites of native cyclotides can be reinforced or abolished, or new biologically active peptide epitopes can be grafted into the scaffold. For example, reinforcing the cytotoxic effect could potentially provide us with leads for anticancer drugs, or to completely remove that effect could provide us with an inert scaffold ideal for grafting. The first successful grafting of a biologically active epitope was reported just recently, showing proof of concept (Gunasekera et al. 2008).
The success of these strategies relies on the ability of efficient methods for production of cyclotides and cyclotide mutants. Being gene products, cell based production systems seem promising, but although cyclotide producing plant cell cultures have been established (Seydel and Dörnenburg 2006), solid phase peptide synthesis is still the method of choice (Gunasekera et al. 2006; Leta Aboye et al. 2008).
Our knowledge about their biosynthesis is yet scarce. We know the structural arrangement of the cyclotide precursor from cDNA, and that an asparaginyl endopeptidase has a likely role for cleaving at the N terminal side of the mature peptide and that protein disulfide isomerases seem to play a role for their successful folding. However, the order of the events to produce mature cyclotides is not yet known, i.e. if disulfide bonds are formed before or after excision and ligation, and nothing is known about how these processes are controlled. In the perspective of exploiting the cyclotide scaffold for engineering of bioactive peptides, the possibility of farming designed molecules in planta promises to be the optimal solution; the way there is still long though.
Strategies of selection—using phylogeny and chemography
More than two centuries ago in his Clavis Medicinæ Duplex Linnaeus wrote:
Plants which correspond to genus correspond also in their properties. Plants that stand close in their natural order also stand close to each other with regards to their virtues.
Linnaeus, 1766
[authors’ translation from latin]
Although void of knowledge regarding the concept of evolution, Linnaeus formulates this stipulate based on his general knowledge on plants and their application as drugs. This insight was revived and reformulated a century later by Helen Abbott, who concluded that:
The evolution of chemical constituents follows parallel lines with the evolutionary course of plant forms, the one being intimately connected with the other…
Helen Cecilia De Silver Abbot, 1887
Franklin Institute lecture:
The chemical basis of plant forms
Extrapolating from these clear-sighted suggestions one arrives at the thought that there ought to be a pattern of correlation between implications derived from exploration of chemical constituents, and those from evolutionary studies. Starting with the initial attempts to characterize chemical properties, such as in the thesis by Hiortzberg (1765)—one of the first in medicinal chemistry at Uppsala University—to present day exploration and charting of chemical space, and in analogy from the Linnaeus classifications of the eighteenth century to modern phylogenetic studies of evolutionary space.
The concept of chemical space is an attempt to transform physical–chemical data to the corresponding information. One way to pursue this task is explored in the area of chemography, where analogies are drawn to geography. As in geography the primary task in chemography is to produce a map to serve as a basis for exploration. Given that chemical space includes all known, and in principle also all possible compounds the sheer size gets woeful. It has been estimated that there are well above 1060 possible small carbon based compounds (Bohacek et al. 1996) and including also small peptides the number of compounds quickly rise to at least 10390 (Dobson 2004). But to further complicate the endeavour chemical space is, as Shoichet puts it: “…vast but most of it is biologically uninteresting; blank, lightless galaxies exist within it into which good ideas at their peril wander” (Shoichet 2004).
With this bleak perspective, it is comprehendible that a considerable effort has been invested in predicting and defining the sectors or multi-dimensional volumes having the best chance of containing the sought-for properties or molecules (Kirkpatrick and Ellis 2004). In pharmacognosy, as in many related fields of research such as chemical ecology, chemical biology and medicinal chemistry, the focus has been directed to sectors believed to contain molecules with biological activities. These are usually referred to as the “biologically relevant chemical space”, and the limits of this multidimensional subvolume are defined by properties and boundaries which allow for binding interactions between biological molecules, such as primary and secondary metabolites, polypeptides, enzymes, RNA, and DNA (Lipinski and Hopkins 2004). In populating the biologically relevant chemical space with molecules, evolutionary forces have led to the creation of an array of inspiring and unique structures—often combining properties that are difficult to access by synthetic chemistry (e.g. Hall et al. 2001; Kingston and Newman 2002; Booth and Zemmel 2004; Rawlins 2004; DiMasi et al. 2003; Pucheault 2008).
To facilitate this exploration, a global map of natural products chemical property has been developed following the principles for the Earth global positioning system (GPS). Starting with 1779 ‘satellite’ and ‘core’ compounds evaluated with 35 carefully selected chemical descriptors, and validated using more than 1.2 million compounds of natural and non-natural origin resulted in ChemGPS-NP. The development of this prediction model, as well as the public web-interface ChemGPS-NPweb available at http://chemgps.bmc.uu.se have been described in previous studies (Larsson et al. 2005, 2007; Rosén et al. 2009b). Starting with a chemical structure, related information can be retrieved in massive amounts by simple procedures, typically from a software package. Such computed information forms the raw material for a majority of the charting endeavours published (e.g. Kirkpatrick and Ellis 2004; Oprea 2002; Agrafiotis et al. 2002; Oprea and Gottfries 2001; Haggarty et al. 2004).
A central dogma of medicinal chemistry and chemical biology is that compounds with similar structures have similar activities. Although there are also numerous examples to the contrary (Kubinyi 1998; Martin et al. 2002) this appears in most cases to be true, and it is reassuring when different methods of exploration such as ethno-pharmacology, database exploitation, and molecular modelling converge on a common suggested lead compound (Bernard et al. 2001). To a large extent studies comparing similarity and estimating chemical diversity, have been pursued by the pharmaceutical industry. The initial driving forces were to be able to predict (Carhart et al. 1985) or expand (Willett et al. 1986) results from ongoing studies by comparing chemical similarity of compounds (Willett et al. 1998; Holliday et al. 2003). However, with the advent of large screening libraries and increasingly more complex targets, the need for stringent selection and experimental design to determine the diversity of a set of compounds became more important (Drie and Lajiness 1998).
In most studies on chemical diversity only the “chemical space of small molecules” or CSSM (Pollock et al. 2008) have been included, excluding more complex molecules such as polypeptides and enzymes which in our opinion ought to be included in the concept of chemical space (Walsh 2001). This poses important questions on data handling and descriptor selection, as there are inherent differences between these groups. While members of CSSM are biosynthesized via biosynthetic pathways, usually from a limited set of building blocks and in a chiral specific manner the polypeptides are in most, and enzymes in all, cases ribosomal products encoded by genes. This brings them closer to the evolutionary forces, as these processes act directly on the (genes coding for the) molecules, and their immediate expression (Kapralov and Filatov 2007; Andersson and Backlund 2008). On the other hand in the case of CSSM, the evolutionary forces act on the biosynthetic machinery, which may still after substantial modifications be able to perform the same synthesis resulting in a product indistinguishable from that of an unmodified enzyme. These differences have been explored by Larsson in a thesis on the toxic polypeptides of mistletoes (Larsson 2007).
What may then be the biological relevance of these chemical substances and their traits? In Nature virtually all (if not all) processes noticeable within and between organisms and their environment are fundamentally chemical reactions. These reactions are to an extensive degree mediated via elaborate enzymes, complex proteins and various high- and low molecular weight compounds, all of which are themselves synthesized ‘on purpose’. Even if we at present only understand a minute fraction of these interactions, we can be confident about the fact that they have been and continuously are evaluated and validated by evolutionary processes. They are all there for a reason—obvious or not.
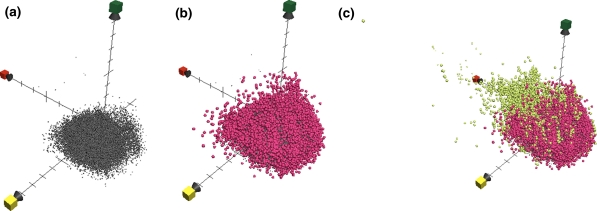
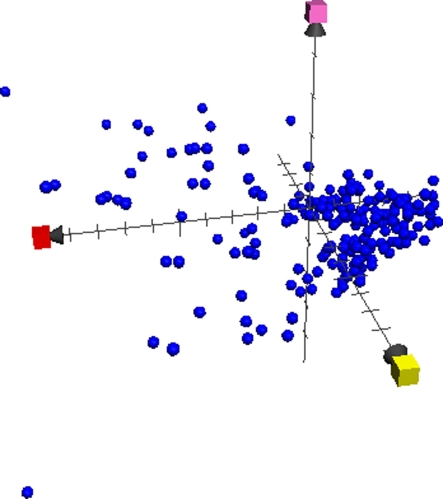
One application of ChemGPS-NP and its charting potential is demonstrated in the following example. A set of more than 49,554 molecules (Fig. 7a) has been screened for activity in a pyruvate kinase assay, with no more than 587 actives retrieved. Wondering to what extent the biologically relevant space has been sampled the physical–chemical properties of the tested molecules can easily become charted together with appropriate datasets for comparison. Introducing a sub-sample of 20,434 molecules from the ZINC database, labelled as ZINC-NP (Fig. 7b), as well as 178,210 compounds from the WOMBAT database (courtesy of Sunset Molecular) (Fig. 7c) we immediately make two conclusions. Firstly, that the tested sample only corresponds to a limited part of the experimentally determined and known biologically relevant chemical space. Secondly, that even the ZINC-NP subset of circa half the number of compounds, as compared to the test set, covers a significantly more diverse volume of chemical space. It is obvious, however, from Fig. 7c that the ZINC-NP set (pink) is comparably well represented among the low molecular compounds, while WOMBAT (lime) covers a much wider range among the larger molecules.
Fig. 7.
Comparison of chemical space plots (three subfigures) comparing chemical space coverage from three datasets, a 49,554 compounds screened in the pyruvate kinase assay [grey], b 20,434 compounds from the ZINC-NP subset [magenta], and c 178,210 compounds from WOMBAT 2007.1 corresponding to substances with demonstrated biological activity [lime]. Red axis = PC1, corresponding to size parameters, yellow axis = PC2, corresponding to aromaticity and conjugation related parameters, and green axis = PC3, corresponding mainly to lipophilicity. Figure by Anders Backlund, all predictions scores calculated using ChemGPS-NPweb, plotted using Apple MacOS X bundled software Grapher 2.1
While there is a general agreement on the concept of chemical space, opinions are much more diverse when it comes to biological space. Some authors see biological space as a subset of chemical space including the chemistry that is related to life, while others envisage a much broader view and even in some cases promote the opposite view—that chemical space is a subset of the biological space. Regardless of which, it is obvious that there is a tight link between parts of these two entities at least. As is the case also for the chemical space, exploration of the biological space can be done at different scale. Detailed knowledge on enzymes, structures, binding affinities and functions (Bologa et al. 2006; Blundell 2007), the patterns of change in proteins during evolution (Andersson and Backlund 2008; Taylor et al. 2001), or the over all patterns of change, speciation, and extinction that are the combined results of evolutionary forces (Angiosperm Phylogeny Group 1998, 2003), all form small contributions to an understanding. In this explorative process the concept of biological diversity has become central. In measuring and quantifying the elusive property “biological diversity”, the advances in phylogenetic reconstruction have become instrumental. With the initial premise set in the beginning of this section, that there are connections between evolution and chemistry, it can be inferred that a larger evolutionary or biological diversity should also indicate a potentially larger chemical diversity. This is one reason why the marine environment has attracted recent attention by natural product chemists in search for novel chemical entities (Hall et al. 2001; Lei and Zhou 2002; Newman and Hill 2006).
A phylogenetic hypothesis, in short a phylogeny, is an implicit hypothesis of evolutionary relationships. Such hypotheses can be erected based on intuition, as for example by Haeckel (1866), but are in a modern systematic or evolutionary biology context a result of careful analysis of scientific data. From a philosophical standpoint there is only one evolutionary history, and evolution hence ought to be represented as one single, bifurcating, tree diagram showing the evolution and succession of all species. The crux being to figure out which of the different possible trees that is ‘correct’, i.e. in the best way representing the result of the evolutionary processes. This is not a trivial problem, and to further complicate the situation, it is today widely accepted that only a bifurcating tree is not adequate for this purpose. This considering the known and well-studied processes of hybridization and lateral gene transfer—the latter explored in great detail for example through the research by Alsmark described in the following section dealing with target identification.
An immediate consequence of the relentless activities by evolutionary forces is that the biological space is at a constant state of change. This is also different to the chemical space where each substance as such (known or hypothesized) has its finite position, although different depending on the reference system applied. Evolution as such does not give room for nostalgia and retaining unnecessary and unimportant features. Everything in Nature is there and kept for a purpose—even if that purpose at present may be difficult for us to un-earthen. One such example can be seen in the enzyme ribulose-1,5-bisphoshate carboxylase/oxygenase, or rubisco, in which the history, development, and fine-tuning can be followed from possible Archaean enolizing enzymes via large subunit dimers in photosynthesizing purple and alpha bacteria to the ‘modern’ hexadecamer with both large and small subunits (Andersson and Backlund 2008). From this study, as well as previous work, features such as positive selection (Lodish et al. 1999), importance of the genetic code redundancy and rate changing separation of gene operons can be illuminated.
As pointed out above, correlation between the organisms and their chemistry is a result of their common history (Linnaeus 1766; Abbott 1887). Forming two sides of a coin, each will tell us something about the other. Presence of a unique compound would be an argument for a common ancestry or a result of sharing a common endogenous parasite or symbiont (el-Seedi et al. 2005). In the same way would close evolutionary kinship increase the chance of encountering similar chemistry (Backlund and Moritz 1998). The tools for biosynthesis are not only modular, flexible and adjustable, they are also at a normal state strongly regulated and perform their tasks following a complex array of control and feedback loops. Attempting to step out of our anthropocentric view this appears quite logical and consequently. Dobson (2004) conclude that one of the greatest challenges for the future of biosynthesis research is to understand how these systems could be influenced to perform in ways better suited for our needs for e.g. production of better drugs, food or building materials.
There is, as mentioned above, a fundamental difference between the chemical space and the evolutionary space. In the latter the single and same result is the expected outcome of different approaches to interpret evolution—as there is supposedly one single (albeit in some cases entangled and reticulate) evolutionary history for organisms on Earth. Chemical space, however, is characterized by an array of more or less well-suited molecular descriptors. When selecting a set of descriptors, this will influence the way in which the corresponding chemical space can be demonstrated. Hence, the process of selecting descriptors becomes central in chemical space exploration to a much greater and more direct extent that the selection of a particular method for phylogenetic analysis. The same comes true for selection of exemplar compounds or training set of compounds, as compared to which organisms are included in a phylogenetic study.
Even if many natural products display a wide range of biological activities, they are not honed by evolutionary forces with the purpose of becoming drugs for use in humans. What they may contribute with, however, is an amazing chemical diversity (Kingston and Newman 2002). This becomes very clear in cases such as the natural product cyclooxygenase (COX) inhibitors. The COX enzyme system of the inflammation cascade appears in at least two isoforms, COX 1 and COX 2, of which the latter is induced and involved in for example the complex of chronic inflammation. Compiling data on more than 200 published COX-1 and 2 inhibitors of natural origin, their mode of inhibition, and their organism of origin provide us with an intriguing pattern—both chemographic and phylogenetic—displayed in Fig. 8.
Fig. 8.
Chemical space plot of COX inhibitors chemical property space plot of 242 natural products with experimentally demonstrated cox-inhibitory effects. Red axis = PC1, corresponding to size parameters, yellow axis = PC2, corresponding to aromaticity and conjugation related parameters, and pink axis = PC8, corresponding primarily to the Lipinski alert index (LAI), positive values indicating Lipinski ‘rule of five’ violation. Note significant variation in size, and large proportion of compounds with positive LAI values. Picture by Anders Backlund, all predictions scores calculated using ChemGPS-NPweb, plotted using Apple MacOS X bundled software Grapher 2.1, data compiled by Maria Schröder-Vilar
The biologically relevant parts of chemical space could be defined as those in which we find natural products, as there is a general belief that evolutionary pressure would, with time, prohibit biosynthesis of compounds, which are not contributing to organism fitness. On the other hand, biologically relevant chemical space could be defined as those parts of chemical space in which substances with demonstrated biological activities are encountered. The latter definition would then include not only natural products but also semi-synthesized or completely synthetic drugs, hit substances from biological assays etc., and hence is different from the first. In an attempt to characterize the overlap or difference between these two different views of biologically relevant space, an experiment was designed by Rosén et al. (2009a). In this experiment more than 186,000 compounds from the database Wombat 2007.1 (‘World of Molecular Bio-Activity’), and 160,000 substances from ‘the Dictionary of Natural Products’ were extracted as SMILES. Both of these sets were subsequently predicted in the ChemGPS-NP chemical property space. The different distributions of the two datasets are very obvious, with features such as a large number of rigid compounds with a low proportion of aromaticity in the Wombat dataset, but not yet tested in biological assays. This fits well with what could be expected from the general section on differences between natural products and products from chemical synthesis, a cadre to which a comparably large portion of the compounds with experimentally investigated biological activity belong. In addition to these more general conclusions, the development and application of a novel similarity measure based on Euclidean distance in chemical property space is described. This approach is fundamentally different compared to structure based similarity measures such as the Tanimoto-index. Using the Euclidean distance measure, the closest neighbours to a set of well-known drugs can be retrieved and investigated from the available wealth of natural products.
Prediction of targets using bioinformatic tools—lateral gene transfer
Over the past few years it has become apparent that lateral gene transfer (LGT) has played an important role in the evolution of pathogenic prokaryotes. LGT is the transfer of genetic material between distinct evolutionary lineages. This contrasts with what is considered to be the ‘normal’ process of inheritance, id est the transmission of mutations and traits from parents to offspring, also known as vertical inheritance. While point mutations are quantitatively the major mode of evolution they generally exert their effects slowly. In contrast, LGT offers the opportunity of evolution by acquisition whereby radically new capabilities such as the ability to utilize new metabolites or the acquisition of drug resistance genes may be acquired quickly. While it is widely held that LGT has exerted a major quantitative and qualitative effect on prokaryotic genomes (Ochman et al. 2000), the role played by LGT—outside the major endosymbiosis events—in the evolution of eukaryotes, including pathogens and parasites of humans and their livestock, is still poorly understood (Doolittle et al. 2003). The main obstacles in analyses have been the reliable inference of LGT on a genomic scale as well as the functional assignment of genes in often very poorly studied organisms. Recent reports about failed treatment due to emerging resistant strains, highlights the urgent need for new drug targets.
In collaboration with TIGR (the Institute for Genomic Research) and the Sanger Institute we have made genome wide tree based screens for LGT in the genomes of Entamoeba histolytica (Loftus et al. 2005; Clark et al. 2007), the trypanosomatides (Berriman et al. 2005; El Sayed et al. 2005) and Trichomonas vaginalis (Carlton et al. 2007). These organisms are increasingly-difficult-to-treat parasites affecting millions of people and animals in the developed and developing world every year. The need for new drugs to treat infections by eukaryotic parasites is becoming increasingly evident as new reports of resistant strains continually emerge. At present metronidazole is the drug of choice used to treat Entamoeba histolytica, Giardia lamblia and Trichomonas vaginalis. However, it has numerous side effects such as nausea, vomiting, diarrhea, hypersensitivity and encephalopathy (Cudmore et al. 2004). Metronidazole has also been reported to be carcinogenic in rodents (Bendesky et al. 2002) and is known to pass the placental barrier in humans. In addition, strains resistant to metronidazole are being increasingly reported. The need for new drugs is therefore urgent.
Because these pathogens are also eukaryotes, they have only a few clear-cut metabolic and genetic differences to mammals. LGTs are attractive candidates for drug targets since their prokaryotic origin make them less likely to have a vulnerable homolog in mammalian genomes and hence the drugs may have less side effects on patients. Our previous analyses have shown that many of the metabolic differences between the parasites and their hosts are due to LGT into the parasite genomes. Many of the LGTs detected lack a homologue in mammalian genomes, for example 5-amino-6-(5-phosphoribosylamino) uracil reductase, is active in riboflavin metabolism in Trichomonas vaginalis, but not in humans. Other LGTs, inferred by phylogeny as bacterial like, are likely to be structurally different to the ancestral eukarytotic homologue, for example d-hydantoinase in Entamoeba histolytica. Better understanding of the metabolic impact of LGT in eukaryotes will guide us in the screen for potential drug targets.
A variety of methods, each with its strengths and weaknesses have been developed to infer the occurrence of LGT (Ragan 2001). Detailed phylogenetic analysis, using sophisticated models of sequence evolution, to detect topological incongruence with established relationships is considered to be the ‘gold standard’, but is too time consuming to apply directly to entire genomes. We have combined rapid screening methods for LGT followed by more detailed phylogenetic analysis of genes that pass the primary screen. The starting point is a Python programme for automated genome analysis called sPyPhy (Sicheritz-Pontén and Andersson 2001). The software sPyPhy automatically generates protein distance phylogenetic trees for all genes in a genome. All information generated, such as BLASTp hits, ClustalW alignments, database information and the bootstrap consensus trees are available for manual inspection through a user-friendly matrix of clickable gene entries. The matrix can be searched in a flexible way so that the best candidates for LGT can be selected. The second stage of the analysis for LGT involves piping alignments selected from the primary screen described above into the programme MrBayes (Huelsenbeck and Ronquist 2001). MrBayes carries out Bayesian phylogenetic analysis (Rannala and Yang 1996) of proteins using a range of sophisticated evolutionary models; the parameters of which can be estimated from the data during the analysis. Published simulations and empirical studies strongly suggest that these more sophisticated models produce more reliable trees, particularly for more divergent sequences like those found in parasites (Hirt et al. 1999). Each candidate LGT is analyzed by MrBayes using the WAG matrix (Whelan and Goldman 2001), a gamma correction for site rate variation and a proportion (pinvar) of invariable sites (Yang 1996). MrBayes uses the likelihood function and the Metropolis-coupled Markov Chain Monte Carlo method to sample model-parameter and tree space (including branch length), producing a set of trees and model parameter values from which a consensus tree is made. Because posterior probabilities, the support values used by Bayesian analysis to indicate confidence in groups, have been criticized (Cummings et al. 2003) we will also use bootstrapping to provide an additional indication of support for relationships. All cases where the tree topology shows one of our chosen parasites clustered with prokaryote sequences separated from any other eukaryote by at least one well supported (bootstrap 0.7, posterior probabilities 95) node was considered as a LGT in that species.
In Entamoeba histolytica a total of 5,740 trees were made and 548 of these were selected from the primary screen as representing potential LGT. The 548 candidate genes were then processed through the secondary screen to make better trees with a Bayesian approach. From the 548 trees LGT could be unambiguously inferred in 68 cases. The remaining 452 cases remained unresolved, but do not show vertical inheritance. Similar results were achieved from the trypanosomatides and Trichomonas vaginalis. These analyses are preliminary but a number of points can already be made. The first is that LGT into these protozoa genomes from bacteria does seem to have occurred, and potentially to a significant degree. The putative LGTs in Entamoeba histolytica are integrated into diverse metabolic pathways, including amino acid, nucleotide and amino acid metabolism (see Fig. 9).
Fig. 9.
Pathways predicted metabolic pathways of Entamoeba histolytica based on analysis of its genome showing inferred LGT (3). Glycolysis and fermentation are the major energy generation pathways. Bold gray arrows represent enzymes encoded by genes that are among the 68 candidates for HGT into the Entamoeba histolytica genome. Broken arrows indicate enzymes for which no gene could be identified from the genome data, although the activity is thought to be present. The framed arrow points to the target of Metronidazole, the major drug for treatment of amoebic liver abscess. Abbreviations: PEP phosphoenolpyruvate, GlcNAc N-acetylglucosamine, LCFA long chain fatty acid, VLCFA very long chain fatty acid, PRPP phosphoribosyl pyrophosphate, GPI glycosylphosphatidylinositol, PAPS phosphoadenosine phosphosulfate. [reproduced from previous publication, permission requested]
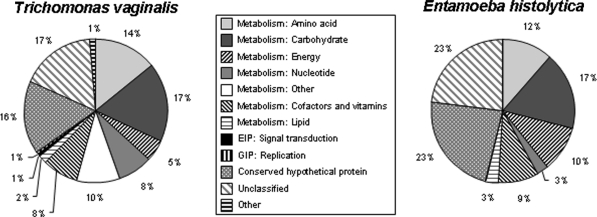
In Trichomonas, metabolic pathways affected include salvage pathways, amino acid metabolism, synthesis of lipophosphoglycan and many more (as indicated in Fig. 10).
Fig. 10.
Piecharts functional categories among Trichomonas vaginalis and Entamoeba histolytica candidate LGTs. Distribution of functional annotation from the KEGG database among candidate LGTs. [reproduced from previous publication, permission requested]
Thus, in the broadest sense LGT must be affecting the fitness of the recipient organism. For Entamoeba histolytica, 45% of the inferred transferred genes are hypothetical or unclassified proteins and in Trichomonas vaginalis the corresponding value is 33%. These values may simply reflect the observation that for Entamoeba histolytica and Trichomonas vaginalis around 30% of the proteins predicted from the genome sequence are also hypothetical or unclassified. The bacterial like-hemolysin acquired through LGT in Entamoeba may be directly involved in virulence; they are commonly transferred among bacterial pathogens (Alsmark et al. 2004).
These results indicate strongly that recent gene transfers are but the tip of a potentially very large iceberg of gene transfers which over time have fundamentally shaped the content of eukaryotic genomes. LGT has thus produced numerous pathways and genes that are present in individual parasitic protozoan species but that are not present on the mammalian genome, thus providing a large and variable pool of potential drug targets across protozoan diversity.
Future perspectives
With an estimated number of more than 300,000 species of plants and more than a million species of microorganisms, insects and marine organisms the biodiversity of Nature remains an unparalleled reservoir of biological and chemical diversity.
Biological diversity, including the variety of life in all its forms, will be of great importance in the future to secure the access of a range of different organisms for the survival of many people. The exploration of the genetic and biochemical diversity of natural resources can result in discovery of potential bioactive compounds and development and sustainable utilisation of these resources. Complex mixtures of natural products are marketed in the society as pharmaceutical products in the form of botanical drugs, nutraceuticals or even cosmeceuticals for enhancement of health. Scientific methods need to be developed for standardisation of bioactive components of this type of products together with clinical trials to prove the efficacy and safety.
An increased understanding that micro-organisms, found in or associated to different macro-organisms both in terrestrial and marine organisms, are the true producers of isolated novel compounds will result in development of new methodology for cultivation of such organisms and a sustainable resource for bioactive natural products.
The growing number of resistant pathogenic microorganisms due to an overuse of antibiotics and the problem to treat severe infectious diseases will accelerate the research to find novel effective molecules of natural origin.
In an earlier publication we have proposed the concept of molecular pharmacognosy, which describes techniques and methods to characterize the structure–activity reason for an observation in Nature (e.g. Bruhn and Bohlin 1997; Bohlin et al. 2007). The field of systems biology, which lies in line with this approach, is an integrated way of studying biological systems using several new technologies such as genomics with transcriptomics, proteomics and metabolomics. Applying these techniques will generate a vast amount of data, which requires informatics tools to be transformed into useful biological information. In future studies of complex mixtures of natural products these new techniques can reveal the influence of phytochemicals on the expression of genes that are involved in a specific pharmacological function using microarrays, evaluation of the enzymes involved in biosynthesis of bioactive constituents in different organisms, and correlation of the metabolic fingerprint of a crude extract to a specific biological activity.
Acknowledgments
The authors are grateful for economic support from the Swedish national research councils VR and FORMAS. All contributing graduate students, guest researchers, and other collaborators involved in the different projects are also greatly acknowledged.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
References
- Abbott HCdS (1887) The chemical basis of plant forms. Franklin Institute lecture
- Agrafiotis DK, Lobanov VS, Salemme FR. Combinatorial informatics in the post-genomics era. Nat Rev Drug Disc. 2002;1:337–346. doi: 10.1038/nrd791. [DOI] [PubMed] [Google Scholar]
- Alsmark CM, Frank AC, Karlberg EO, Legault BA, Ardell DH, Canback B, Eriksson AS, Näslund AK, Handley SA, Huvet M, La Scola B, Holmberg M, Andersson SG. The louse-borne human pathogen Bartonella quintana is a genomic derivative of the zoonotic agent Bartonella henselae. Proc Natl Acad Sci. 2004;101:9716–9721. doi: 10.1073/pnas.0305659101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersson L (1987) Biological screening of and chemical studies on some Swedish marine organisms. Acta Universitatis Upsaliensis. Comprehensives summaries of Uppsala dissertations from the Faculty of Pharmacy 29
- Andersson I, Backlund A. Structure and function of Rubisco. Plant Phys Biochem. 2008;46:275–291. doi: 10.1016/j.plaphy.2008.01.001. [DOI] [PubMed] [Google Scholar]
- Angiosperm Phylogeny Group I An ordinal classification of flowering plants. Ann Missouri Bot Gard. 1998;85:531–553. [Google Scholar]
- Angiosperm Phylogeny Group II An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG II. Bot J Linn Soc. 2003;141:399–436. [Google Scholar]
- Augustine NR. Silence—editorial. Science. 2008;321:1605. doi: 10.1126/science.1162924. [DOI] [PubMed] [Google Scholar]
- Backlund A, Moritz T. Phylogenetic implications of an expanded valepotriate distribution in the Valerianaceae. Biochem Syst Ecol. 1998;26:309–335. [Google Scholar]
- Bendesky A, Menendez D, Ostrosky-Wegman P. Is metronidazole carcinogenic? Mutat Res. 2002;2:133–144. doi: 10.1016/s1383-5742(02)00007-8. [DOI] [PubMed] [Google Scholar]
- Bernard P, Scior T, Didier B, Hibert M, Berthon J-Y. Ethnopharmacology and bioinformatic combinations for leads discovery: application to phospholipase A2 inhibitors. Phytochemistry. 2001;58:865–874. doi: 10.1016/s0031-9422(01)00312-0. [DOI] [PubMed] [Google Scholar]
- Berriman M, Ghedin E, Hertz-Fowler C, Blandin G, Renauld H, Bartholomeu DC, Lennard NJ, Caler E, Hamlin NE, Haas B, Böhme U, Hannick L, Aslett MA, Shallom J, Marcello L, Hou L, Wickstead B, Alsmark UC, Arrowsmith C, Atkin RJ, Barron AJ, Bringaud F, Brooks K, Carrington M, Cherevach I, Chillingworth TJ, Churcher C, Clark LN, Corton CH, Cronin A, Davies RM, Doggett J, Djikeng A, Feldblyum T, Field MC, Fraser A, Goodhead I, Hance Z, Harper D, Harris BR, Hauser H, Hostetler J, Ivens A, Jagels K, Johnson D, Johnson J, Jones K, Kerhornou AX, Koo H, Larke N, Landfear S, Larkin C, Leech V, Line A, Lord A, Macleod A, Mooney PJ, Moule S, Martin DM, Morgan GW, Mungall K, Norbertczak H, Ormond D, Pai G, Peacock CS, Peterson J, Quail MA, Rabbinowitsch E, Rajandream MA, Reitter C, Salzberg SL, Sanders M, Schobel S, Sharp S, Simmonds M, Simpson AJ, Tallon L, Turner CM, Tait A, Tivey AR, Van Aken S, Walker D, Wanless D, Wang S, White B, White O, Whitehead S, Woodward J, Wortman J, Adams MD, Embley TM, Gull K, Ullu E, Barry JD, Fairlamb AH, Opperdoes F, Barrell BG, Donelson JE, Hall N, Fraser CM, Melville SE, El-Sayed NM. The genome of the African trypanosome Trypanosoma brucei. Science. 2005;309:416–422. doi: 10.1126/science.1112642. [DOI] [PubMed] [Google Scholar]
- Blundell T. New dimensions of structural proteomics: exploring chemical and biological space. Structure. 2007;15:1342–1343. doi: 10.1016/j.str.2007.10.008. [DOI] [PubMed] [Google Scholar]
- Bohacek RS, McMartin C, Guida WC. The art and practice of structure-based drug design: a molecular modeling perspective. Med Res Rev. 1996;16:3–50. doi: 10.1002/(SICI)1098-1128(199601)16:1<3::AID-MED1>3.0.CO;2-6. [DOI] [PubMed] [Google Scholar]
- Bohlin L, Göransson U, Backlund A. Modern pharmacognosy: connecting biology and chemistry. Pure Appl Chem. 2007;79(4):763–774. [Google Scholar]
- Bologa CG, Revankar CM, Young SM, Edwards BS, Arterburn JB, Kiselyov AS, Parker MA, Tkachenko SE, Savchuck NP, Sklar LA, Oprea TI, Prossnitz ER. Virtual and biomolecular screening converge on a selective agonist for GPR30. Nat Chem Biol. 2006;2:207–212. doi: 10.1038/nchembio775. [DOI] [PubMed] [Google Scholar]
- Booth B, Zemmel R. Prospects for productivity. Nat Rev Drug Disc. 2004;3:451–456. doi: 10.1038/nrd1384. [DOI] [PubMed] [Google Scholar]
- Bruhn JG, Bohlin L. Molecular pharmacognasy: an explanatory model. Drug Disc Today. 1997;2:243–246. [Google Scholar]
- Carhart RE, Smith DH, Venkataraghavan R. Atom pairs as molecular features in structure–activity studies: definitions and applications. J Chem Inf Comput Sci. 1985;25:64–73. [Google Scholar]
- Carlton JM, Hirt RP, Silva JC, Delcher AL, Schatz M, Zhao Q, Wortman JR, Bidwell SL, Alsmark UC, Besteiro S, Sicheritz-Ponten T, Noel CJ, Dacks JB, Foster PG, Simillion C, Van de Peer Y, Miranda-Saavedra D, Barton GJ, Westrop GD, Müller S, Dessi D, Fiori PL, Ren Q, Paulsen I, Zhang H, Bastida-Corcuera FD, Simoes-Barbosa A, Brown MT, Hayes RD, Mukherjee M, Okumura CY, Schneider R, Smith AJ, Vanacova S, Villalvazo M, Haas BJ, Pertea M, Feldblyum TV, Utterback TR, Shu CL, Osoegawa K, de Jong PJ, Hrdy I, Horvathova L, Zubacova Z, Dolezal P, Malik SB, Logsdon JM, Jr, Henze K, Gupta A, Wang CC, Dunne RL, Upcroft JA, Upcroft P, White O, Salzberg SL, Tang P, Chiu CH, Lee YS, Embley TM, Coombs GH, Mottram JC, Tachezy J, Fraser-Liggett CM, Johnson PJ. Draft genome sequence of the sexually transmitted pathogen Trichomonas vaginalis. Science. 2007;315:207–212. doi: 10.1126/science.1132894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cech T, Kennedy D. Doing more for Kate—editorial. Science. 2005;310:1741. doi: 10.1126/science.1123580. [DOI] [PubMed] [Google Scholar]
- Celio GJ, Padamsee M, Dentinger BTM, Bauer R, McLaughlin DJ. Assembling the fungal tree of life: constructing the structural and biochemical database. Mycologia. 2006;98:850–859. doi: 10.3852/mycologia.98.6.850. [DOI] [PubMed] [Google Scholar]
- Claeson P, Göransson U, Johansson S, Luijendijk T, Bohlin L. Fractionation protocol for the isolation of polypeptides from plant biomass. J Nat Prod. 1998;61:77–81. doi: 10.1021/np970342r. [DOI] [PubMed] [Google Scholar]
- Clark CG, Alsmark UC, Tazreiter M, Saito-Nakano Y, Ali V, Marion S, Weber C, Mukherjee C, Bruchhaus I, Tannich E, Leippe M, Sicheritz-Ponten T, Foster PG, Samuelson J, Noël CJ, Hirt RP, Embley TM, Gilchrist CA, Mann BJ, Singh U, Ackers JP, Bhattacharya S, Bhattacharya A, Lohia A, Guillén N, Duchêne M, Nozaki T, Hall N. Structure and content of the Entamoeba histolytica genome. Adv Parasitol. 2007;65:51–69. doi: 10.1016/S0065-308X(07)65002-7. [DOI] [PubMed] [Google Scholar]
- Claus R, Hoppen HO, Karg H. The secret of truffles: a steroidal pheromone? Cell Mol Life Sci. 1981;37:1178–1179. [Google Scholar]
- Colgrave ML, Craik DJ. Thermal, chemical, and enzymatic stability of the cyclotide kalata B1: the importance of the cyclic cystine knot. Biochemistry. 2004;43:5965–5975. doi: 10.1021/bi049711q. [DOI] [PubMed] [Google Scholar]
- Colgrave ML, Kotze AC, Kopp S, McCarthy JS, Coleman GT, Craik DJ. Anthelmintic activity of cyclotides: in vitro studies with canine and human hookworms. Acta Trop. 2009;109:163–166. doi: 10.1016/j.actatropica.2008.11.003. [DOI] [PubMed] [Google Scholar]
- Cudmore SL, Delgaty KL, Hayward-McClelland SF, Petrin DP, Garber GE. Treatment of infections caused by metronidazole-resistant Trichomonas vaginalis. Clin Microbiol Rev. 2004;4:783–793. doi: 10.1128/CMR.17.4.783-793.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cummings MP, Handley SA, Myers DS, Reed DL, Rokas A, Winka K. Comparing bootstrap and posterior probability values in the four-taxon case. Syst Biol. 2003;52:477–487. doi: 10.1080/10635150390218213. [DOI] [PubMed] [Google Scholar]
- DiMasi JA, Hansen RW, Grabowski HG. The price of innovations: new estimates of drug development costs. J Health Econ. 2003;22:151–185. doi: 10.1016/S0167-6296(02)00126-1. [DOI] [PubMed] [Google Scholar]
- Dobson CM. Chemical space and biology. Nature. 2004;432:824–828. doi: 10.1038/nature03192. [DOI] [PubMed] [Google Scholar]
- Doolittle WF, Boucher Y, Nesbo CL, Douady CJ, Andersson JO, Roger AJ. How big is the iceberg of which organellar genes in nuclear genomes are but the tip? Philos Trans R Soc B-Biol Sci. 2003;358:39–58. doi: 10.1098/rstb.2002.1185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drie JHv, Lajiness MS. Approaches to virtual library design. Drug Disc Today. 1998;3:274–283. [Google Scholar]
- EC—European Commission (2008) Flash Eurobarometer 239—Young people and science, analytical report October. European Comission official documents
- Ekenäs C, Rosén J, Wagner S, Merfort I, Backlund A, Andreasen K. Secondary chemistry and ribosomal DNA data congruencies in Arnica (Asteraceae) Cladistics. 2009;25:78–92. doi: 10.1111/j.1096-0031.2008.00244.x. [DOI] [PubMed] [Google Scholar]
- El-Sayed NM, Myler PJ, Blandin G, Berriman M, Crabtree J, Aggarwal G, Caler E, Renauld H, Worthey EA, Hertz-Fowler C, Ghedin E, Peacock C, Bartholomeu DC, Haas BJ, Tran AN, Wortman JR, Alsmark UC, Angiuoli S, Anupama A, Badger J, Bringaud F, Cadag E, Carlton JM, Cerqueira GC, Creasy T, Delcher AL, Djikeng A, Embley TM, Hauser C, Ivens AC, Kummerfeld SK, Pereira-Leal JB, Nilsson D, Peterson J, Salzberg SL, Shallom J, Silva JC, Sundaram J, Westenberger S, White O, Melville SE, Donelson JE, Andersson B, Stuart KD, Hall N. Comparative genomics of trypanosomatid parasitic protozoa. Science. 2005;309:404–409. doi: 10.1126/science.1112181. [DOI] [PubMed] [Google Scholar]
- El-Seedi H, Larsson S, Backlund A. Chemosystematic value of cyclopeptide alkaloids from Heisteria nitida (Olacaceae) Biochem Syst Ecol. 2005;33:831–839. [Google Scholar]
- Felth J, Rickardson L, Rosén J, Wickström M, Fryknäs M, Lindskog M, Bohlin L, Gullbo J (2009) Cytotoxic effects of cardiac glycosides in colon cancer cells, alone and in combination with standard chemotherapeutic drugs. J Nat Prod 72:1969–1974 [DOI] [PubMed]
- Göransson U, Craik DJ. Disulfide mapping of the cyclotide kalata B1: chemical proof of the cyclic cystine knot motif. J Biol Chem. 2003;278:48188–48196. doi: 10.1074/jbc.M308771200. [DOI] [PubMed] [Google Scholar]
- Göransson U, Luijendijk T, Johansson S, Bohlin L, Claeson P. Seven novel macrocyclic polypeptides from Viola arvensis. J Nat Prod. 1999;62:283–286. doi: 10.1021/np9803878. [DOI] [PubMed] [Google Scholar]
- Göransson U, Broussalis AM, Claeson P. Expression of the Viola cyclotides by LC/MS and MS–MS sequencing of intercysteine loops after introduction of charges and cleavage sites by aminoethylation. Anal Biochem. 2003;318:107–117. doi: 10.1016/s0003-2697(03)00114-3. [DOI] [PubMed] [Google Scholar]
- Göransson U, Sjögren M, Svangård E, Claeson P, Bohlin L. Reversible antifouling effect of the cyclotide cycloviolacin O2 against barnacles. J Nat Prod. 2004;67:1287–1290. doi: 10.1021/np0499719. [DOI] [PubMed] [Google Scholar]
- Göransson U, Svangård E, Claeson P, Bohlin L. Novel strategies for isolation and characterization of cyclotides: the discovery of bioactive macrocyclic plant polypeptides in the Violaceae. Curr Protein Pept Sci. 2004;5:317–329. doi: 10.2174/1389203043379495. [DOI] [PubMed] [Google Scholar]
- Göransson U, Herrmann A, Burman R, Haugaard-Jönsson LM, Rosengren KJ. The conserved glu in the cyclotide cycloviolacin O2 has a key structural role. Chem Biochem. 2009;10:2354–2360. doi: 10.1002/cbic.200900342. [DOI] [PubMed] [Google Scholar]
- Gran L. On the effect of a polypeptide isolated from “Kalata–kalata” (Oldenlandia affinis DC.) on the oestrogen dominated uterus. Acta Pharmacol Toxicol. 1973;33:400–408. doi: 10.1111/j.1600-0773.1973.tb01541.x. [DOI] [PubMed] [Google Scholar]
- Gran L. Oxytocic principles of Oldenlandia affinis. Lloydia. 1973;36:174–178. [PubMed] [Google Scholar]
- Gran L, Sandberg F, Sletten K. Oldenlandia affinis (R&S) DC—a plant containing uteroactive peptides used in African traditional medicine. J Ethnopharmacol. 2000;70:197–203. doi: 10.1016/s0378-8741(99)00175-0. [DOI] [PubMed] [Google Scholar]
- Gunasekera S, Daly NL, Anderson MA, Craik DJ. Chemical synthesis and biosynthesis of the cyclotide family of circular proteins. IUBMB Life. 2006;58:515–524. doi: 10.1080/15216540600889532. [DOI] [PubMed] [Google Scholar]
- Gunasekera S, Foley FM, Clark RJ, Sando L, Fabri LJ, Craik DJ, Daly NL. Engineering stabilized vascular endothelial growth factor-A antagonists: synthesis, structural characterization, and bioactivity of grafted analogues of cyclotides. J Med Chem. 2008;51:7697–7704. doi: 10.1021/jm800704e. [DOI] [PubMed] [Google Scholar]
- Haeckel E. Generelle Morphologie der Organismen. Berlin: Georg Reiner; 1866. [Google Scholar]
- Haggarty SJ, Clemons PA, Wong JC, Schreiber SL. Mapping chemical space using molecular descriptors and chemical genetics: deacetylase inhibitors. Comb Chem HTS. 2004;7:669–676. doi: 10.2174/1386207043328319. [DOI] [PubMed] [Google Scholar]
- Hall DG, Manku S, Wang F. Solution- and solid-phase strategies for the design, synthesis, and screening of libraries based on natural product templates: a comprehensive survey. J Comb Chem. 2001;3:125–150. doi: 10.1021/cc0001001. [DOI] [PubMed] [Google Scholar]
- Hawksworth DL. Mushrooms: the extent of the unexplored potential. Int J Med Mushroom. 2001;3:333–337. [Google Scholar]
- Hedner E (2007) Bioactive compounds in the chemical defence on marine sponges—structure–activity relationships and pharmacological targets. Acta Universitatis Upsaliensis. Comprehensive summaries of Uppsala dissertations from the Faculty of Pharmacy 63
- Hedner E, Sjögren M, Frändberg P-A, Johansson T, Göransson U, Dahlström M, Jonsson P, Nyberg F, Bohlin L. Brominated cyclodipeptides from the marine sponge Geodia barretti as selective 5-HT ligands. J Nat Prod. 2006;69:1421–1424. doi: 10.1021/np0601760. [DOI] [PubMed] [Google Scholar]
- Hedner E, Sjögren M, Hodzic S, Andersson R, Göransson U, Jonsson PR, Bohlin L. Antifouling activity of a dibrominated cyclopeptide from the marine sponge Geodia barretti. J Nat Prod. 2008;71:330–333. doi: 10.1021/np0705209. [DOI] [PubMed] [Google Scholar]
- Herrmann A, Svangård E, Claeson P, Gullbo J, Bohlin L, Göransson U. Key role of glutamic acid for the cytotoxic activity of the cyclotide cycloviolacin O2. Cell Mol Life Sci. 2006;63:235–245. doi: 10.1007/s00018-005-5486-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herrmann A, Burman R, Mylne JS, Karlsson G, Gullbo J, Craik DJ, Clark RJ, Göransson U. The alpine violet, Viola biflora, is a rich source of cyclotides with potent cytotoxicity. Phytochemistry. 2008;69:939–952. doi: 10.1016/j.phytochem.2007.10.023. [DOI] [PubMed] [Google Scholar]
- Hibbett DS, Binder M, Bischoff JF, Blackwell M, Cannon PF, Eriksson OE, Huhndorf S, James T, Kirk PM, Lücking R, Thorsten Lumbsch H, Lutzoni F, Matheny PB, McLaughlin DJ, Powell MJ, Redhead S, Schoch CL, Spatafora JW, Stalpers JA, Vilgalys R, Aime MC, Aptroot A, Bauer R, Begerow D, Benny GL, Castlebury LA, Crous PW, Dai YC, Gams W, Geiser DM, Griffith GW, Gueidan C, Hawksworth DL, Hestmark G, Hosaka K, Humber RA, Hyde KD, Ironside JE, Kõljalg U, Kurtzman CP, Larsson KH, Lichtwardt R, Longcore J, Miadlikowska J, Miller A, Moncalvo JM, Mozley-Standridge S, Oberwinkler F, Parmasto E, Reeb V, Rogers JD, Roux C, Ryvarden L, Sampaio JP, Schüssler A, Sugiyama J, Thorn RG, Tibell L, Untereiner WA, Walker C, Wang Z, Weir A, Weiss M, White MM, Winka K, Yao YJ, Zhang N. A higher-level phylogenetic classification of the fungi. Mycol Res. 2007;111:509–547. doi: 10.1016/j.mycres.2007.03.004. [DOI] [PubMed] [Google Scholar]
- Hiortzberg L (1765) Methodo investigandi vires medicamentorum chemica. PhD dissertation in Medicinal Chemistry, Uppsala University
- Hirt RP, Logsdon JM, Healy B, Dorey MW, Doolittle WF, Embley TM. Microsporidia are related to fungi: evidence from the largest subunit of RNA polymerase II and other proteins. Proc Natl Acad Sci. 1999;96:580–585. doi: 10.1073/pnas.96.2.580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holliday JD, Salim N, Whittle M, Willett P. Analysis and display of the size dependence of chemical similarity coefficients. J Chem Inf Comput Sci. 2003;43:819–828. doi: 10.1021/ci034001x. [DOI] [PubMed] [Google Scholar]
- Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17:754–755. doi: 10.1093/bioinformatics/17.8.754. [DOI] [PubMed] [Google Scholar]
- Huss U (2003) Studies on the effects of plant and food constituents on cyclooxygenase-2. Aspects in inflammation and cancer. Acta Universitatis Upsaliensis. Comprehensive summaries of Uppsala dissertations from the faculty of Pharmacy 294:1–56
- Huss U, Ringbom T, Perera P, Bohlin L, Vasänge M. Screening of ubiquitous plant constituents for COX-2 inhibition with a scintillation proximity based assay. J Nat Prod. 2002;65:1571–1621. doi: 10.1021/np020023m. [DOI] [PubMed] [Google Scholar]
- Ireland DC, Colgrave ML, Craik DJ. A novel suite of cyclotides from Viola odorata: sequence variation and the implications for structure, function and stability. Biochem J. 2006;400:1–12. doi: 10.1042/BJ20060627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jennings C, West J, Waine C, Craik DJ, Anderson M. Biosynthesis and insecticidal properties of plant cyclotides: the cyclic knotted proteins from Oldenlandia affinis. Proc Natl Acad Sci. 2001;98:10614–10619. doi: 10.1073/pnas.191366898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamimori H, Hall K, Craik D, Aguilar M. Studies on the membrane interactions of the cyclotides kalata B1 and kalata B6 on model membrane systems by surface plasmon resonance. Anal Biochem. 2005;337:149–153. doi: 10.1016/j.ab.2004.10.028. [DOI] [PubMed] [Google Scholar]
- Kapralov MV, Filatov DA. Widespread positive selection in the photosynthetic Rubisco enzyme. BMC Evol Biol. 2007;7:73. doi: 10.1186/1471-2148-7-73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim DJ, Shin DH, Ahn B, Kang JS, Nam KT, Park CB, Kim CK, Hong JT, Kim TB. Chemoprevention of colon cancer by Korean food plant components. Mut Res. 2003;523(524):99–107. doi: 10.1016/s0027-5107(02)00325-1. [DOI] [PubMed] [Google Scholar]
- Kingston DG, Newman DJ. Mother Nature’s combinatorial libraries; their influence on the synthesis of drugs. Curr Opin Drug Disc Dev. 2002;5:304–316. [PubMed] [Google Scholar]
- Kirkpatrick P, Ellis C. Nature insight—chemical space. Nature. 2004;432:823. [Google Scholar]
- Klintberg B af (1998) Paddehattar: äldre folkliga uppfattningar om svamp i Sverige. Kuttrasju: folkloristiska och kulturhistoriska essäer. Stockholm
- Klitgaard AB. The fauna associated with outer shelf and upper slope sponges (Porifera, Demospongiae) at the Faroe Islands. Sarsia. 1995;80:1–22. [Google Scholar]
- Kubinyi H. Similarity and dissimilarity: A medicinal chemist’s view. Persp Drug Disc Design. 1998;9/10/11:225–252. [Google Scholar]
- Læssøe T, Hansen K. Truffle trouble: what happened to the Tuberales? Mycol Res. 2007;111:1075–1099. doi: 10.1016/j.mycres.2007.08.004. [DOI] [PubMed] [Google Scholar]
- Larsson S (2007) Mistletoes and thionins. As selection models in natural products drug discovery. Acta Universitatis Upsaliensis. Comprehensive summaries of Uppsala dissertations from the faculty of Pharmacy 49:1–65
- Larsson J, Gottfries J, Bohlin L, Backlund A. Expanding the ChemGPS chemical space with natural products. J Nat Prod. 2005;68:985–991. doi: 10.1021/np049655u. [DOI] [PubMed] [Google Scholar]
- Larsson J, Gottfries J, Muresan S, Backlund A. ChemGPS-NP: tuned for navigation in biologically relevant chemical space. J Nat Prod. 2007;70:789–794. doi: 10.1021/np070002y. [DOI] [PubMed] [Google Scholar]
- Lei J, Zhou J. A marine natural product database. J Chem Inf Comput Sci. 2002;42:742–748. doi: 10.1021/ci010111x. [DOI] [PubMed] [Google Scholar]
- Leta Aboye T, Clark RJ, Craik DJ, Göransson U. Ultra-stable peptide scaffolds for protein engineering-synthesis and folding of the circular cystine knotted cyclotide cycloviolacin O2. Chem Biochem. 2008;9:103–113. doi: 10.1002/cbic.200700357. [DOI] [PubMed] [Google Scholar]
- Lidgren G (1989) Pharmacognostic studies on some Swedish marine organisms with emphasis on the sponge Geodia barretti. Acta Universitatis Upsaliensis. Comprehensives summaries of Uppsala dissertations from the Faculty of Pharmacy
- Lindholm P, Göransson U, Johansson S, Claeson P, Gullbo J, Larsson R, Bohlin L, Backlund A. Cyclotides: a novel type of cytotoxic agents. Mol Cancer Ther. 2002;1:365–369. [PubMed] [Google Scholar]
- Linné C von (1749) Materia Medica. Laurentii Salvii, Holmiæ (Stockholm)
- Linné C von (1766) Clavis Medicinæ Duplex. Laurentii Salvii, Holmiæ (Stockholm)
- Lipinski CA, Hopkins AL. Navigating chemical space for biology and medicine. Nature. 2004;432:855–861. doi: 10.1038/nature03193. [DOI] [PubMed] [Google Scholar]
- Lodish H, Berk A, Zipursky L, Matsudaira P, Baltimore D, Darnell J. Molecular cell biology. 4. New York: W. H. Freeman; 1999. [Google Scholar]
- Loftus B, Anderson I, Davies R, Alsmark UC, Samuelson J, Amedeo P, Roncaglia P, Berriman M, Hirt RP, Mann BJ, Nozaki T, Suh B, Pop M, Duchene M, Ackers J, Tannich E, Leippe M, Hofer M, Bruchhaus I, Willhoeft U, Bhattacharya A, Chillingworth T, Churcher C, Hance Z, Harris B, Harris D, Jagels K, Moule S, Mungall K, Ormond D, Squares R, Whitehead S, Quail MA, Rabbinowitsch E, Norbertczak H, Price C, Wang Z, Guillén N, Gilchrist C, Stroup SE, Bhattacharya S, Lohia A, Foster PG, Sicheritz-Ponten T, Weber C, Singh U, Mukherjee C, El-Sayed NM, Petri WA, Jr, Clark CG, Embley TM, Barrell B, Fraser CM, Hall N. The genome of the protist parasite Entamoeba histolytica. Nature. 2005;433:865–868. doi: 10.1038/nature03291. [DOI] [PubMed] [Google Scholar]
- Martin YC, Kofron JL, Traphagen LM. Do structurally similar molecules have similar biological activity? J Med Chem. 2002;45:4350–4358. doi: 10.1021/jm020155c. [DOI] [PubMed] [Google Scholar]
- Mueller GM, Schmit JP, Leacock PR. Global diversity and distribution of macrofungi. Biodivers Conserv. 2007;16:37–48. [Google Scholar]
- Murakami A, Ohigashi H. Targeting NOX, INOS and COX-2 in inflammatory cells: chemoprevention using food phytochemicals. Int J Cancer. 2007;121:2357–2363. doi: 10.1002/ijc.23161. [DOI] [PubMed] [Google Scholar]
- Newman DJ, Hill RT. New drugs from marine microbes: the tide is turning. J Ind Microbiol Biotech. 2006;33:539–544. doi: 10.1007/s10295-006-0115-2. [DOI] [PubMed] [Google Scholar]
- Ochman H, Lawrence JG, Groisman EA. Lateral gene transfer and the nature of bacterial innovation. Nature. 2000;405:299–304. doi: 10.1038/35012500. [DOI] [PubMed] [Google Scholar]
- Oprea TI. Chemical space navigation in lead discovery. Curr Opin Chem Biol. 2002;6:384–389. doi: 10.1016/s1367-5931(02)00329-0. [DOI] [PubMed] [Google Scholar]
- Oprea TI, Gottfries J. Chemography: the art of navigating in chemical space. J Comb Chem. 2001;3:157–166. doi: 10.1021/cc0000388. [DOI] [PubMed] [Google Scholar]
- Påhlsson I. A standard pollen diagram from the Lojsta area of Central Gotland. Striae 3. Uppsala, Sweden: Societas Upsaliensis Pro Geologia Quarternaria; 1977. [Google Scholar]
- Pettersson J, Karlsson PC, Choi YH, Verpoorte R, Rafter JJ, Bohlin L. NMR metabolomic analysis of fecal water from subjects on a vegetarian diet. Biol Pharm Bull. 2008;31:1192–1198. doi: 10.1248/bpb.31.1192. [DOI] [PubMed] [Google Scholar]
- Pettersson J, Karlsson PC, Göransson U, Rafter JJ, Bohlin L. The flavouring phytochemical 2-pentanone reduces prostaglandin production and COX-2 expression in colon cancer cells. Biol Pharm Bull. 2008;31:534–537. doi: 10.1248/bpb.31.534. [DOI] [PubMed] [Google Scholar]
- Plan MR, Saska I, Cagauan AG, Craik DJ. Backbone cyclised peptides from plants show molluscicidal activity against the rice pest Pomacea canaliculata (golden apple snail) J Agric Food Chem. 2008;56:5237–5241. doi: 10.1021/jf800302f. [DOI] [PubMed] [Google Scholar]
- Pollock SN, Coutsias EA, Wester MJ, Oprea TI. Scaffold topologies I: exhaustive enumeration up to 8 rings. J Chem Inf Mod. 2008;48:1304–1310. doi: 10.1021/ci7003412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pucheault M. Natural products: chemical instruments to apprehend biological symphony. Org Biomol Chem. 2008;6:424–432. doi: 10.1039/b713022h. [DOI] [PubMed] [Google Scholar]
- Ragan MA. On surrogate methods for detecting lateral gene transfer. FEMS Microbiol Lett. 2001;201:187–191. doi: 10.1111/j.1574-6968.2001.tb10755.x. [DOI] [PubMed] [Google Scholar]
- Rannala B, Yang Z. Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol. 1996;43:304–311. doi: 10.1007/BF02338839. [DOI] [PubMed] [Google Scholar]
- Rawlins MD. Cutting the cost of drug development? Nat Rev Drug Disc. 2004;3:360–364. doi: 10.1038/nrd1347. [DOI] [PubMed] [Google Scholar]
- Rosén J, Gottfries J, Muresan S, Backlund A, Oprea TI. Novel chemical space exploration via natural products. J Med Chem. 2009;52:1953–1962. doi: 10.1021/jm801514w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosén J, Lövgren A, Kogej T, Muresan S, Gottfries J, Backlund A. ChemGPS-NPWeb-chemical space navigation online. J Comput Aid Mol Des. 2009;23:253–259. doi: 10.1007/s10822-008-9255-y. [DOI] [PubMed] [Google Scholar]
- Rosengren KJ, Daly NL, Plan MR, Waine C, Craik DJ. Twists, knots, and rings in proteins. Structural definition of the cyclotide framework. J Biol Chem. 2003;278:8606–8616. doi: 10.1074/jbc.M211147200. [DOI] [PubMed] [Google Scholar]
- Saether O, Craik DJ, Campbell ID, Sletten K, Juul J, Norman DG. Elucidation of the primary and three-dimensional structure of the uterotonic polypeptide kalata B1. Biochemistry. 1995;34:4147–4158. doi: 10.1021/bi00013a002. [DOI] [PubMed] [Google Scholar]
- Samuelsson G. Phytochemical and pharmacological studies on Viscum album L. Sve Farm Tidskr. 1958;62:1–21. [PubMed] [Google Scholar]
- Seydel P, Dörnenburg H. Establishment of in vitro plants, cell and tissue cultures from Oldenlandia affinis for the production of cyclic peptides. Plant Cell Tissue Org Cult. 2006;85:247–255. [Google Scholar]
- Shenkarev ZO, Nadezhdin KD, Sobol VA, Sobol AG, Skjeldal L, Arseniev AS. Conformation and mode of membrane interaction in cyclotides. Spatial structure of kalata B1 bound to a dodecylphosphocholine micelle. FEBS J. 2006;273:2658–2672. doi: 10.1111/j.1742-4658.2006.05282.x. [DOI] [PubMed] [Google Scholar]
- Shoichet BK. Virtual screening of chemical libraries. Nature. 2004;432:862–865. doi: 10.1038/nature03197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sicheritz-Pontén T, Andersson SG. A phylogenomic approach to microbial evolution. Nucleic Acids Res. 2001;29:545–552. doi: 10.1093/nar/29.2.545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simonsen SM, Sando L, Ireland DC, Colgrave ML, Bharathi R, Göransson U, Craik DJ. A continent of plant defense peptide diversity: cyclotides in Australian Hybanthus (Violaceae) Plant Cell. 2005;17:3176–3189. doi: 10.1105/tpc.105.034678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sjögren M (2006) Bioactive compounds from the marine sponge Geodia barretti. Acta Universitatis Upsaliensis. Comprehensive summaries of Uppsala dissertations from the Faculty of Pharmacy 32
- Sjögren M, Johnson A-L, Hedner E, Dahlström M, Göransson U, Shirani H, Bergman J, Jonsson PR, Bohlin L. Antifouling activity of synthesized peptide analogs of the sponge metabolite barettin. Peptides. 2006;27:2058–2064. doi: 10.1016/j.peptides.2006.03.027. [DOI] [PubMed] [Google Scholar]
- Svangård E, Göransson U, Hocaoglu Z, Gullbo J, Larsson R, Claeson P, Bohlin L. Cytotoxic cyclotides from Viola tricolor. J Nat Prod. 2004;67:144–147. doi: 10.1021/np030101l. [DOI] [PubMed] [Google Scholar]
- Svangård E, Burman R, Gunasekera S, Lövborg H, Gullbo J, Göransson U. Mechanism of action of cytotoxic cyclotides: cycloviolacin O2 disrupts lipid membranes. J Nat Prod. 2007;70:643–647. doi: 10.1021/np070007v. [DOI] [PubMed] [Google Scholar]
- Tam JP, Lu YA, Yang JL, Chiu KW. An unusual structural motif of antimicrobial peptides containing end-to-end macrocycle and cystine-knot disulfides. Proc Natl Acad Sci. 1999;96:8913–8918. doi: 10.1073/pnas.96.16.8913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor TC, Backlund A, Bjorhall K, Spreitzer RJ, Andersson I. First crystal structure of Rubisco from a green alga, Chlamydomonas reinhardtii. J Biol Chem. 2001;276:48159–48164. doi: 10.1074/jbc.M107765200. [DOI] [PubMed] [Google Scholar]
- Traber R, Hofmann H, Kobel H. Cyklosporins—new analogues by precursor directed biosynthesis. J Antibiot. 1989;42:591. doi: 10.7164/antibiotics.42.591. [DOI] [PubMed] [Google Scholar]
- Trappe JM, Castellano MA. Keys to the genera of truffles (Ascomycetes) McIlvanea. 1991;10:47–65. [Google Scholar]
- Walsh C. Enabling the chemistry of life. Nature. 2001;409:226–231. doi: 10.1038/35051697. [DOI] [PubMed] [Google Scholar]
- Wang CK, Hu SH, Martin JL, Sjögran T, Hajdu J, Bohlin L, Claeson P, Göransson U, Rosengren KJ, Tang J, Tan NH, Craik DJ. Combined X-ray and NMR analysis of the stability of the cyclotide cystine knot fold that underpins its insecticidal activity and potential use as a drug scaffold. J Biol Chem. 2009;284(16):10672–10683. doi: 10.1074/jbc.M900021200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wedén C (2004) Black truffles of Sweden. Systematics, Population Studies, Ecology and Cultivation of Tuber aestivum syn. T. uncinatum. Acta Universitatis Upsaliensis. Comprehensive Summaries of Uppsala Dissertations from the faculty of Science and Technology 1043
- Wedén C, Danell E, Camacho FJ, Backlund A. The population of the hypogeous fungus Tuber aestivum syn. T. uncinatum on the island of Gotland. Mycorrhiza. 2004;14:19–23. doi: 10.1007/s00572-003-0271-4. [DOI] [PubMed] [Google Scholar]
- Wedén C, Danell E, Tibell L. Species recognition in the truffle genus Tuber: The synonyms Tuber aestivum and Tuber uncinatum. Environ Microbiol. 2005;7:1535–1546. doi: 10.1111/j.1462-2920.2005.00837.x. [DOI] [PubMed] [Google Scholar]
- Whelan S, Goldman N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol. 2001;18:691–699. doi: 10.1093/oxfordjournals.molbev.a003851. [DOI] [PubMed] [Google Scholar]
- Willett P, Winterman V, Bawden D. Implementation of nearest-neighbor searching in an online chemical structure search system. J Chem Inf Comput Sci. 1986;26:36–41. [Google Scholar]
- Willett P, Barnard JM, Downs GM. Chemical similarity searching. J Chem Inf Comput Sci. 1998;38:983–996. [Google Scholar]
- Yang Z. Among-site rate variation and its impact on phylogenetic analyses. Trends Ecol Evol. 1996;11:367–372. doi: 10.1016/0169-5347(96)10041-0. [DOI] [PubMed] [Google Scholar]