Abstract
We have previously isolated GRIM-19, a novel growth suppressor, using a genetic method. GRIM-19 ablates cell growth by inhibiting the transcription factor signal transducer and activator of transcription 3 (STAT3). Up-regulation of STAT3 and growth promotion were observed in a number of human tumors. Although the tumor-suppressive actions of GRIM-19 are known, the structural elements required for its antitumor actions are not understood. Mutational and protein sequence analyses identified a motif in the N terminus of GRIM-19 that exhibited similarity to certain RNA viral proteins. We show that disruption of specific amino acids within this motif cripples the antitumor actions of GRIM-19. These mutants fail to interact with STAT3 efficiently and consequently do not inhibit growth-promoting gene expression. More importantly, we show that a clinically observed mutation in the N terminus of GRIM-19 also weakened its interaction with STAT3 and antitumor action. Together, these studies identify a major role for the N terminus of GRIM-19 in mediating its tumor-suppressive actions.
Interferons (IFNs) regulate antiviral, antitumor, and immune responses in vertebrates by stimulating gene expression via Janus tyrosine kinase (JAK)-signal transducer and activator of transcription (STAT) pathways.1 The antitumor actions of IFNs involve induction of growth inhibitors and/or activators of apoptosis.2,3 Given the complexities of IFN actions and their pleiotropic effects on various cell types, a much more dynamic protein network that changes to the need, by using distinct gene products, can be envisaged. Despite the widespread use of IFNs for clinical applications, mechanisms of IFN-induced growth control are not fully clear. An understanding of these pathways might allow the development of pharmacological agents that exert synergistic activity. Although IFNs strongly inhibit cell growth, many tumors are insensitive to IFN-induced growth inhibition. However, combining IFNs with all-trans retinoic acid (RA) enhances IFN-induced growth inhibition.2
Our laboratory used a genetic method and identified novel mediators used by IFN/RA action. One such novel gene product, GRIM-19, caused apoptosis when overexpressed and promoted growth when down-regulated.4 GRIM-19 was later shown to be a subunit of mitochondrial complex I in bovine heart tissue by a biochemical approach,5 it was required for the assembly of complex I,6 and deletion of Grim19 caused embryonic lethality in mice.7 Generation of high levels of reactive oxygen species during IFN/RA treatment seems to be a mechanism for inducing cell death in some cells8 for which the C terminus of GRIM-19 is required.4 We have shown that GRIM-19 binds to the mitochondrial serine protease HtrA2 and augments its proteolytic activity on XIAP. Degradation of XIAP leading to activation of caspase-9 in an IFN/RA-dependent manner is another mechanism of apoptosis induction by GRIM-19.9 In addition to the apoptotic function, GRIM-19 seems to participate in an innate immune response,10 cell motility,11 and calcium homeostasis during frog heart development.12 We and others have shown a loss of GRIM-19 expression in renal cell carcinoma13 and in colorectal carcinoma,14 indicating a potential tumor suppressor-like function for GRIM-19. GRIM-19 targets the oncogenic transcription factor STAT3 for inhibition during growth suppression.15,16 On a similar line, a secreted glycoprotein, GW112, of unknown function antagonizes the role of GRIM-19 in gastrointestinal cancers.17 Viral gene products block GRIM-19 from triggering the apoptotic cascade.18,19 Although GRIM-19 possesses potent antitumor properties, the structural elements required for its activity are unknown. In this report, we have identified a short motif in the N terminus of GRIM-19 required for its antitumor activity. More importantly, a clinically identified tumor-derived mutation in this region disrupted its biological activity and promoted growth. Together with our earlier observations, GRIM-19 seems to be a new tumor suppressor.
Materials and Methods
Cell Lines and Antibodies
3Y1 (nononcogenic rodent fibroblasts) and HeLa cells were grown in Dulbecco’s modified Eagle’s medium with 10% fetal bovine serum. HSC2, HSC3, and HSC4 cell lines established from lymph node metastases originated from oral squamous cell carcinomas of three different Japanese patients.20 Primary sites of the tumors for HSC2, HSC3, and HSC4 were floor of mouth, tongue, and tongue, respectively. HSC cell lines were grown in RPMI 1640 with 10% fetal bovine serum. The Ca9-2 cell line was established from gingival squamous cell carcinoma.21 All of these cell lines have been shown to contain mutant p53 alleles.22 Polyclonal antibodies against STAT3 and β-actin and monoclonal antibodies against Myc-tag (Cell Signaling Technology, Danvers, MA), FLAG-tag (Sigma-Aldrich, St. Louis, MO), and GRIM-1923 were used in these studies.
Construction of GRIM-19 Expression Plasmids
Wild-type and mutants of GRIM-19 were generated using PCR and subsequently cloned into pIRES-Puro2 vector and expressed as C-terminal Myc-tagged proteins. Point mutation K5N, originally reported in a thyroid tumor,24 was reconstructed using PCR-directed mutagenesis. The primers used for generation of point mutants and deletions are presented in Table 1. Sequence and expression-verified clones were used for all experiments. An expression vector coding for a constitutively dimerized Stat3 (S3C) was described previously.25 Wild-type and mutant GRIM-19 constructs were electroporated into cells using Amaxa Nucleofector technology. Selection with puromycin was started 24 hours after transfection and continued for 1 week. The surviving colonies were pooled to avoid a clonal bias in further experiments.
Table 1.
Primers Used for Generating Mutations and Deletions in Human GRIM19 Open Reading Frame
| Primer ID | Sequence |
|---|---|
| WT | 5′-GCGGAATTCGCCACCATGGCGGCGTCAAAGGTG-3′ |
| K5N | 5′-GCGGAATTCGCCACCATGGCGGCGTCAAACGTGAAG-3′ |
| Q8A | 5′-GCGGAATTCGCCACCATGGCGGCGTCAAAGGTGAAGGCGGACATG-3′ |
| D9A | 5′-GCGGAATTCGCCACCATGGCGGCGTCAAAGGTGAAGCAGGCGATGCC-3′ |
| M10A | 5′-GCGGAATTCGCCACCATGGCGGCGTCAAAGGTGAAGCAGGACGCGCCTCC-3′ |
| P11A | 5′-GCGGAATTCGCCACCATGGCGGCGTCAAAGGTGAAGCAGGACATGGCGCCGCC-3′ |
| NΔ17 | 5′-GCGGAATTCGCCACCATGCCCATCGACTACAAGCGGAAC-3′ |
| NΔ34 | 5′-GCGGAATTCGCCACCATGCTGGCCATAGGGATTGGAACC-3′ |
| Rev* | 5′-CAGGGATCCCGTGTACCACATGAAGCCG-3′ |
Restriction enzyme sites are underlined in the primer sequence. WT, wild type.
, common reverse primer for cloning the GRIM19 open reading frame.
For some experiments, cells were infected with replication-defective lentiviral particles carrying GRIM-19 or its mutant open reading frame. GRIM-19 inserts were cloned into the XhoI and XbaI sites of pLVX-Puro (Clontech, Mountain View, CA). To produce lentiviral particles coding for GRIM-19 proteins variants, HEK-293T cells (3 × 106) were transfected with 5 μg of transfer vector (eg, pLVX-Puro-GRIM19-Myc), 3.75 μg of packaging plasmid (pCMV-dR8.2 dvpr), and 1.5 μg of envelope plasmid (pCMV-VSV-G) (Addgene Inc., Cambridge, MA) using Lipofectamine-Plus reagent (Invitrogen, Carlsbad, CA) for 6 hours in serum-free medium. Cells were then fed with 10 ml of regular growth medium (Dulbecco’s modified Eagle’s medium with 10% fetal bovine serum). Viral particles from the growth medium were collected between 48 and 96 hours after transfection. The medium was centrifuged to remove debris and filtered through a 0.45-μm filter. The viral titers were determined in each case by infecting HeLa cells with different dilutions of the virus preparation and then selecting the cells with puromycin. After 2 weeks, the number of infectious virus particles was estimated based on the numbers of puromycin-resistant colonies formed. Equal numbers of particles expressing various GRIM-19 mutants were used for infecting the cells.
Gene Expression Analysis
Transient transfection, luciferase, and β-galactosidase reporter assays were performed as in our earlier studies.16 Lysates were prepared 48 hours after transfection and used for further analyses. Western blot and immunoprecipitation analyses were conducted essentially as described in our earlier reports.26
Real-Time PCR Analysis
First-strand cDNAs were generated from total RNA using oligo(dT) primer and SuperScript III reverse transcriptase (Invitrogen). RT-PCR reactions were set up using an SYBR Green JumpStart Kit (Sigma-Aldrich), and reactions were monitored using the Mx3005P real-time PCR machine (Stratagene, La Jolla, CA). The comparative Ct (threshold cycle) method described in the manufacturer’s protocol was used for quantifying transcript levels. Transcripts coding for ribosomal proteins L32 and S24 were used as internal controls in these assays. Each experiment was repeated at least three times. Multiple samples (n = 6) from different experiments were used for measuring transcript levels, and the statistical significance of differences between various groups was assessed using Student’s t-test.
Immunofluorescence
Cells were infected with lentiviral particles coding for various GRIM-19 open reading frames. Thirty-six hours later, they were stained with MitoTracker (Invitrogen), fixed, and permeabilized. They were stained with an Myc-tag-specific antibody, fluorescein isothiocyanate-conjugated secondary antibody, and 4,6-diamidino-2-phenylindole. Images were captured using a fluorescence microscope (BX-FLA, Olympus, Osaka, Japan) fitted with a digital camera (QICAM) and processed by QCapture Pro 5.1 (QImaging, Surrey, BC, Canada).
Cell Growth and Apoptosis Assays
These assays were conducted as described earlier.27 Cells (1000 per well) were plated in 96-well plates. Each group had eight replicates. Cells were fixed with trichloroacetic acid (10% final) at 4°C for 1 hour at the end of the experiment and stained with 0.4% sulforhodamine B (Sigma-Aldrich). The bound dye was eluted with 100 μl of Tris-HCl (pH 10.5), and absorbance was measured at 570 nm. Eight hours after the cells were plated, one plate was fixed with trichloroacetic acid for determining the input. Absorbance obtained with this plate provided the starting (0 hours) cell numbers. Fas ligation with Jo-1 antibody (Clontech), annexin V staining assays were performed as described in our earlier studies.26
In Vitro Wound-Healing Assay
Cells lines expressing various genes were grown in a six-well plate. Confluent monolayers were scratched with a 200-μl pipette tip to make a wound. The monolayer was washed twice with PBS to remove the detached cells, supplemented with full medium, and incubated for 4 hours at 37°C. We first marked the injured front and measured space between the injured edges. Images were captured using a Nikon Eclipse TS-100 microscope fitted with a digital camera to monitor the cell movement into the wounded area. All micrographs were taken at the same time and at the same magnification. For quantification of wound closure, the width of each wound was measured after 4 hours and is expressed as a percentage of the original wound size as described elsewhere.28 Mean values from six different plates per cell line were used for calculating these data.
Subcellular Fractionation
Extracts from cells, expressing various mutants of GRIM-19, were fractionated into nuclear, cytoplasmic, and mitochondrial components using differential centrifugation as described elsewhere16,29,30 with certain modifications. All steps were performed at 4°C. Cells (5 × 107) were suspended in a fivefold hypotonic buffer (10 mmol/L Tris-HCl, pH 7.9; 10 mmol/L KCl, 0.5mmol/L dithiothreitol, 0.5 mmol/L EDTA, and 0.5 mmol/L phenylmethylsulfonyl fluoride) for 10 minutes followed by centrifugation and suspension in homogenization buffer (10 mmol/L Tris-HCl, pH 7.9, 0.5 mmol/L EDTA, and 0.5 mmol/L dithiothreitol) containing 8.5% sucrose. The cell suspension was homogenized using a glass pestle and centrifuged at 1000 × g for 5 minutes to remove debris. The supernatant was then layered over 50% sucrose prepared in the homogenization buffer and spun at 15,000 × g for 20 minutes. Nuclei form a pellet, whereas the other fractions stay in the top layer at this stage. Purity of the nuclear fraction was assessed by microscopy. After separation of nuclei, the top layer was collected and centrifuged at 39,000 × g for 45 minutes to pellet mitochondria and plasma membrane. The supernatant after this centrifugation was regarded as the cytosolic fraction. To separate the plasma membranes from the mitochondria, the pellet was homogenized in buffer containing 50% sucrose. Sucrose (8.5%) was layered on top of this suspension, and the resulting discontinuous gradient was centrifuged at 57,000 × g for 90 minutes. At this stage, the plasma membranes appear as a band at the sucrose interface, whereas the mitochondria form a pellet. The mitochondria were washed three times with homogenization buffer and then lysed in 1× SDS-polyacrylamide gel electrophoresis loading buffer. The nuclei were washed three times with homogenization buffer, suspended the same buffer containing 1 mmol/L MgCl2 and 10% glycerol; and incubated on ice for 10 minutes. Nuclear proteins were isolated by slow addition of the extraction buffer (10 mmol/L Tris-HCl, pH 7.9; 800 mmol/L KCl, 1 mmol/L MgCl2, 0.25 mmol/L EDTA, 0.5 mmol/L dithiothreitol, and 10% glycerol) and incubated for 30 minutes with gentle vortexing. After this, the mixture was centrifuged at 25,000 × g for 30 minutes, and the supernatant was collected. This was considered to be the nuclear extract. The protein content of each fraction was measured by Bradford reagent (Sigma-Aldrich). Organelle-specific markers used were c-Jun, cytochrome c, and β-tubulin, representing nuclear, mitochondrial, and cytoplasmic fractions, respectively.
Colony-Forming Ability in Soft Agar
These assays were described in our earlier reports.26 A bottom agar layer containing 0.7% w/v agarose with growth medium containing 10% fetal bovine serum was poured into six-well plates. Cells (104) were mixed with growth medium containing 0.35% w/v agarose and seeded on the solidified bottom agar layer. Plates were incubated for ∼3 weeks in a humidified incubator at 37°C. Colonies were counted in five random fields from each of the triplicate samples.
Tumor Growth Assays
Three- to four-week-old athymic nude (nu/nu) NCr mice (Taconic Farms, Germantown, NY) were used in the study. Procedures involving animals and their care were conducted in conformity with the institutional guidelines that are in compliance with national and international laws and policies (EEC Council Directive 86/609, OJL 358, 1 December 1987, and the National Institutes of Health Guide for the Care and Use of Laboratory Animals, NIH Publication No. 85-23, 1985). Each experimental group contained 10 mice, each bearing one tumor. Cells (106) prepared in 0.2 ml of PBS were inoculated into flanks in the mid-axillary line as described in our earlier studies.26,31 Cell viability was >98% at the time of inoculation. Tumor volume was calculated using caliper measurements and the formula V = (4/3)πa2b, where a and b are the minor and major axes of the prolate spheroid, respectively.
Statistical Analyses
Student’s t-test was used to assess the statistical significance of difference between pairs. A maximum P = 0.05 was considered significant.
Results
Identification of Human Cell Lines with Low GRIM-19
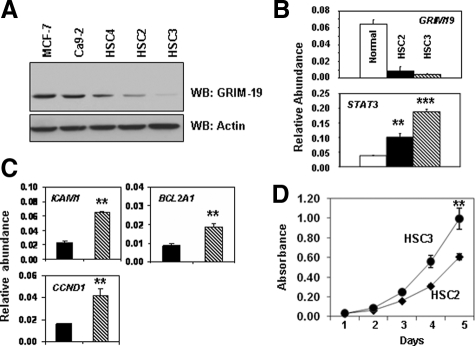
To analyze the structural elements of GRIM-19, we needed a cell line that almost lacked endogenous GRIM-19. Because many human cell lines express GRIM-19 to a variable extent, we screened for cell lines that express low levels of GRIM-19. Among the many cell lines screened (data not shown), two oral squamous cell carcinoma cell lines, HSC2 and HSC3, isolated from clinical tumors had the lowest level of GRIM-19, as determined by Western blot analyses with GRIM-19-specific monoclonal antibody (Figure 1A). Another oral squamous cancer cell line, Ca9-2 had the highest GRIM-19 level. MCF-7, a human breast carcinoma cell line, was used as a reference control. We also compared the levels of GRIM19 mRNA in normal human oral tissue (obtained from surgical waste) with those in HSC3 and HSC2 using real-time PCR analysis. The HSC2 and HSC3 cell lines expressed 6- and 12-fold lower levels of GRIM19, respectively, than normal oral tissue (Figure 1B, top).26 These cell lines had significantly (P < 0.005) higher STAT3 levels compared with normal oral tissue (Figure 1B, bottom). We observed a significant difference in STAT3 levels of HSC2 and HSC3. However, the GRIM-19 levels did not correspond to STAT3 levels in a 1:1 manner in these cells. These differences could be due to the fact that they are derived from different patients and originated from distinct regions of oral cavity (see Materials and Methods). It is also likely that other unknown genetic/epigenetic differences could contribute to different STAT3 levels in these cells. Most importantly, the lower level of GRIM-19 in HSC3 cells was associated with a higher steady-state level of STAT3-responsive genes (BCL2A1, CCND1, and ICAM1) than that in HSC2 cells (P < 0.005) (Figure 1C). These results are consistent with our earlier data that GRIM-19 is an inhibitor of STAT3. We next tested whether HSC3 and HSC2 cell lines grew differently from each other, because of the differences in GRIM-19 and STAT3 content (Figure 1D). Indeed, the HSC3 cell line grew significantly faster than HSC2 (P < 0.005). Because the HSC3 cell line had an extremely low level of GRIM-19, it was used for further experiments.
Figure 1.
Identification of human squamous cell carcinoma cell lines with low GRIM-19 expression. A: The indicated cell lysates (∼40 μg) was used for Western blot analyses with the indicated antibodies. The HSC3 cell line has the lowest GRIM-19 level. WB, Western blot. B: Levels of endogenous GRIM19 and STAT3 transcripts were quantified using real-time PCR analysis with gene-specific primers.26 RNA isolated from two pooled samples of normal buccal tissue from surgical waste was used as a control. Each bar represents the mean ± SE (n = 6). C: Real-time PCR analysis of steady-state STAT3-responsive gene levels in HSC2 and HSC3 cells. Each bar represents the mean ± SE (n = 4). Bar markings are same as in B. D: Cell growth pattern of HSC2 and HSC3 cells (n = 6). Statistical significance was obtained by Student’s t-test: **P < 0.005; ***P < 0.001.
Deletion of N Terminus Results in Loss of Antitumor Action of GRIM-19
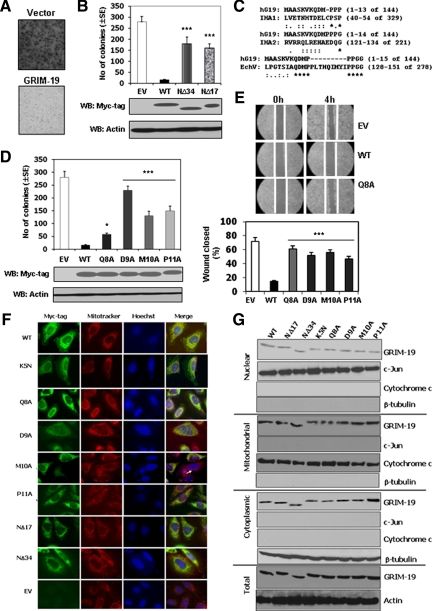
Although GRIM-19 is a potent growth suppressor; the critical domains required for this function have not been clearly defined. We first generated two N-terminal deletion mutants, NΔ34 and NΔ17, which lacked the N-terminal 34 and 17 amino acids of GRIM-19, respectively. These mutants were tested for their ability to suppress anchorage-independent growth of HSC3 cells. A typical pattern of soft agar colonies formed in the presence and absence of GRIM-19 is illustrated in Figure 2A. Whereas wild-type GRIM-19 robustly suppressed the formation of soft agar colonies, compared with empty expression vector (Figure 2B, top panel), the deletion mutants, NΔ17 and NΔ34, significantly lost such ability (P < 0.001). These differences are not due to differential expression of the mutants, as shown by a Western blot analysis of the cell extracts (Figure 2B, bottom panel). Based on the data obtained with the NΔ17 mutant, we searched the Protein Data Bank (http://www.rcsb.org/pdb/home/home.do, last accessed, December 12, 2005) for proteins with sequence matches using the first 17 amino acids of GRIM-19 as a query. These searches identified two viral proteins with good similarity (Figure 2C): i) the hemagglutinin of human influenza virus and ii) the capsid protein V of echovirus. In particular, the latter protein has a motif formed by the QDMP quartet. Therefore, we introduced point mutations in the QDMP motif to determine its role in the antitumor activity of GRIM-19.
Figure 2.
Identification of a growth-regulatory domain in GRIM-19 protein. A: Effect of wild-type GRIM-19 on soft-agar colony formation by HSC3 cells. Photomicrographs of representative fields are shown. Original magnification, ×20. B and D: Quantification of colony formation. Bars indicate the mean ± SE of triplicates in each experiment (n = 3). The Western blots (WB) below these graphs show the expression levels of GRIM-19 mutants and actin in these cells. A portion of the cells used for colony formation assays (in A) were plated in parallel, without soft agar, and selected with G418 for three weeks. An equal quantity of protein (50 μg) from each transfectant was used for Western blot analysis. C: Identification of proteins with GRIM-19-like domains. A homology map is shown: asterisk (∗), colon (:), and period (.) indicate identity, high similarity, and low similarity, respectively. E: Effect of QDMP mutations on cell motility. Assays were performed as described in Materials and Methods. Right panel shows a typical pattern of motility observed. White lines in these photomicrographs show the original boundaries of the scratched area. The graph below the pictures shows quantification of distances moved at the end of four hours after the initial introduction of a scratch into the monolayers. Each bar represents the mean ± SE obtained with quadruplicates in each experiment (n = 4). F: Cellular localization of GRIM-19 mutants in HCS3 cells. Immunofluorescent images of HSC3 cells infected with the indicated GRIM-19 construct (green channel) and mitochondria visualized using MitoTracker (red channel) are shown. The majority of the protein is found in the mitochondrion, but some nuclear fluorescence is also seen. Arrowhead indicates noninfected cell(s). G: Cellular subfractionation of HSC3 cells expressing the indicated GRIM-19 construct. Distribution of GRIM-19 in the cell is not exclusively mitochondrial as evidenced by its presence in nuclear and cytoplasmic fractions. c-Jun, cytochrome c, and β-tubulin were used as markers for nuclear, mitochondrial, and cytoplasmic fractions, respectively. A Myc-tag-specific antibody was used to detect GRIM-19. Statistical significance was obtained by Student’s t-test: *P < 0.01; **P < 0.005; ***P < 0.001. EV, empty vector control; WT, wild-type.
Mutations in QDMP Motif Cause a Loss of GRIM-19 Activity
The mutant constructs were individually expressed in HSC3 cells and compared with wild-type GRIM-19 for growth ability on soft agar. As anticipated, wild-type GRIM-19 suppressed colony formation strongly compared with the empty vector (Figure 2D, top panel). Under these conditions, mutations in the QDMP motif, in particular, the DMP residues, caused a significant increase in colony formation (P < 0.001). The Q8A mutant marginally, but significantly (P < 0.01), lost the ability to suppress anchorage-independent growth, and D9A mutant lost most of this ability. Thus, the QDMP motif seems to be important for inhibiting anchorage-independent growth of HSC3 cells. All mutant proteins expressed equivalently (Figure 2D, bottom panel), indicating that the differences are due to a functional difference in them.
One characteristic of cancer cells is their mobility, which is required for metastasis. We reported earlier that GRIM-19 inhibits cell motility.11 Therefore, we tested whether mutations in the QDMP motif affected the ability of GRIM-19 to suppress cell motility. HSC3 cells expressing either wild-type or mutant GRIM-19 were used for cell motility assays as described in Materials and Methods. A scratch was introduced into a confluent monolayer of cells, and mobilization of cells into the “denuded” area was monitored. As exemplified in Figure 2E (top panel), wild-type GRIM-19 suppressed the motility of cells, whereas the mutant Q8A failed to do so. A similar experiment was repeated with DMP mutants, and their motility was quantified (Figure 2E, bottom panel). All mutants lost the ability (P < 0.001) to suppress cell motility.
One possible cause for the loss of activity of the mutants is their differential localization in cells. To determine this aspect, we generated lentiviral particles, coding for the wild-type and mutant GRIM-19 proteins, and used them for infecting HSC3 cells. Our previous studies showed that GRIM-19 is present in both cytoplasmic and nuclear compartments. In particular, it was a part of the mitochondrion in the cytoplasm. In HSC3 cells, a major portion of GRIM-19 was found in the mitochondrion (Figure 2F), as revealed by colocalization with MitoTracker, a mitochondrion-specific dye. However, some minor portion was also present in the nucleus. The mutants were similar to the wild-type protein in terms of their cellular localization. The P11A mutant, in addition, exhibited a higher nonmitochondrial localization in many fields (Figure 2F). To further ascertain the in situ observations, we fractionated HSC3 cell extracts, expressing various mutants into nuclear, mitochondrial, and cytoplasmic components. Each of these fractions was subjected to Western blot analyses with antibodies that can detect GRIM-19 (ie, Myc-tag), cytochrome c (mitochondrial marker), c-Jun (nuclear marker), and β-tubulin (cytosolic marker). We have also used an additional cytoplasmic marker, JAK1 (a cytoplasmic tyrosine kinase). It was found in the cytosolic but not in the nuclear and mitochondrial fractions (data not shown). As shown in Figure 2G, all GRIM-19 mutants were present in the nuclear, mitochondrial, and cytoplasmic fractions, like the wild-type protein. However, a higher amount of P11A was present in the cytoplasmic fraction. All mutants expressed equivalently as revealed by total GRIM-19 protein.
A Tumor-Derived Mutation in the N Terminus of GRIM-19 Inactivates Its Antitumor Function
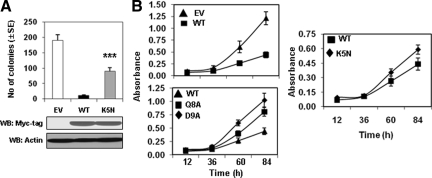
One recent study reported mutations in the GRIM19 gene in primary human Hürthle cell carcinomas.24 The biological relevance of this mutation was unclear. Because the N terminus was important for the antitumor effect of GRIM-19, we next tested the impact of a clinically observed mutation for which the fifth amino acid, lysine, was converted to an asparagine (K5N). This mutant was compared with wild-type GRIM-19 for its ability to suppress soft agar colony formation by HSC3 cells (Figure 3A, top panel). As expected, wild-type GRIM-19 protein robustly suppressed soft agar colony formation. However, the K5N mutant significantly lost the ability to inhibit soft agar colonies (P < 0.001). Thus, the N terminus of GRIM-19 harbors critical residues required for its antitumor action(s). There was no discernible difference with respect to the expression of wild-type and K5N mutant proteins (Figure 3A, bottom panel).
Figure 3.
Effect of N-terminal mutations on GRIM-19 activity. A: Quantitative representation of soft agar colony formation by HSC3 cells in the presence of K5N, a tumor-derived mutation. A Myc-tag-specific antibody was used in Western blots (WB) to detect GRIM-19. B: Effect of GRIM-19 mutants on cell growth. Equal numbers of HSC3 cells expressing the indicated mutants were used in a growth assay. Significant growth differences between wild-type GRIM-19 (WT) and empty-vector control (EV) (P < 0.001), D9A (P < 0.005), Q8A (P < 0.005), and K5N (P < 0.01) are seen. Each data point represents the mean ± SE of eight replicates in each case (n = 4). Statistical significance was obtained by Student’s t-test: ***P < 0.001. EV, empty vector control; WT, wild-type.
Impact of N-Terminal Mutations on Cell Growth
We next tested the effect of N-terminal mutants on growth of HSC3 cells compared with that of cells expressing wild-type GRIM-19. As shown in Figure 3B, wild-type GRIM-19 robustly suppressed cell growth compared with the empty vector (P < 0.001). The clinical mutant K5N (P < 0.01) and experimental mutants Q8Aand D9A (P < 0.005) significantly lost such ability compared with wild-type GRIM-19. These data support the importance of the N terminus of GRIM-19 for suppressing tumor cell growth.
Mutant GRIM-19 Proteins Fail to Inhibit STAT3-Induced Transcription
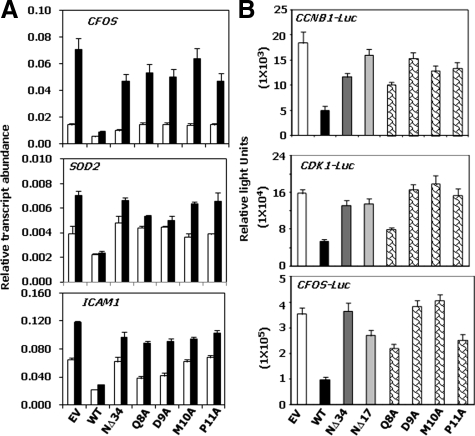
We have shown that GRIM-19 targets the transcription factor STAT3 for ablating the growth-promoting effects,16 and a low level of GRIM-19 in HSC3 cells correlated with a high STAT3 activity (Figure 1). To determine whether the mutants have lost the ability to suppress STAT3-responsive gene expression, we took the following approaches: i) quantified the expression of STAT3-responsive genes using real-time PCR and ii) measured the expression of luciferase reporter genes driven by STAT3-responsive gene promoters in HeLa cells. As shown in Figure 4A, wild-type GRIM-19 protein suppressed the expression of STAT3-responsive genes, CFOS, ICAM1, and SOD2, compared with empty vector-transfected cells (P < 0.001). In contrast, the deletion of the N-terminal region or mutations in the QDMP residues resulted in a significant (P < 0.001) loss of anti-STAT3 activity. A similar picture was obtained with the luciferase reporters, for which N-terminal mutants were found to be significantly defective (P < 0.005) at repressing gene expression (Figure 4B).16,26 All three reporters, cyclin B1-Luc, CDC2-Luc, and c-fos-Luc yielded a very similar pattern. Thus, mutations in the N terminus of GRIM-19 greatly diminished anti-STAT3 activity.
Figure 4.
Effect of GRIM-19 mutants on STAT3-responsive gene expression. A: Real-time PCR analysis with gene-specific primers was used for quantifying gene expression in HSC3 cells stably expressing the indicated mutants. White and black bars represent steady state and interleukin-6-induced mRNA levels. B: The indicated luciferase reporters driven by the promoters of various STAT3-inducible genes16,26 were used for studying the effect of GRIM-19 mutants on gene expression. Transfection efficiency was normalized to the expression of the internal control (β-galactosidase). Each data point represents the mean ± SE of triplicates in each experiment (n = 3). Statistical significance was obtained by Student’s t-test: **P < 0.005; ***P < 0.001. EV, empty vector control; WT, wild-type.
Effect of N-Terminal and Tumor-Derived Mutations of GRIM-19 on Anti-STAT3 Activity
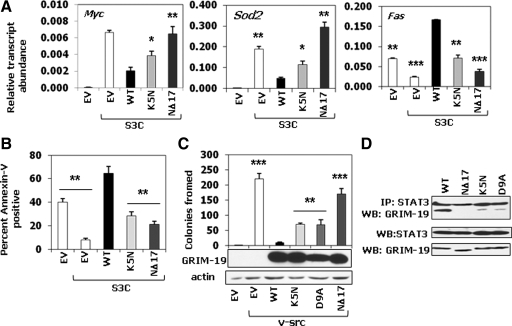
Although the above studies show the effect of GRIM-19 on STAT3-responsive genes, they do not prove whether these effects are directed against STAT3. To examine this aspect, we used a rodent fibroblast cell line, 3Y1, in which expression of a constitutively-dimerizing variant of Stat3, S3C, is sufficient to cause oncogenic transformation.25 Use of this cell line allowed us to clearly distinguish the anti-STAT3 effects of GRIM-19 mutants. For these studies, we used the S3C-expressing 3Y1 cell line26 and infection with lentiviral particles coding for wild-type, NΔ17, and K5N proteins. The empty expression vector was used as a control for all comparisons. In these experiments, we measured the expression of three STAT3-responsive growth-associated genes. Two such growth-promoting genes, Myc and Sod2, were up-regulated after expression of S3C (Figure 5A).26 In the presence of wild-type GRIM-19, S3C-induced expression of these transcripts was significantly inhibited. In sharp contrast to these, NΔ17 and K5N mutants lost the ability to suppress S3C-driven transcription. A converse effect was observed on Fas death receptor, another STAT3-regulated gene.32 Expression of Fas was inhibited in the presence of S3C, compared with empty vector-transfected cells. Fas expression was de-repressed in presence of wild-type GRIM-19 to a level higher than that of the empty vector-expressing cells. In contrast, both K5N (P < 0.01) and NΔ17 significantly lost their ability to de-repress Fas expression (P < 0.005). Thus, a clinically observed mutation in the N terminus of GRIM-19 also disrupted anti-STAT3 activity. To explore the significance of Fas up-regulation, we ligated these cells with an agonistic antibody that induces apoptosis. Cell death was measured as percentage of annexin V-positive cells using fluorescence-activated cell sorting. As shown in Figure 5B, the empty vector-expressing cells underwent apoptosis, whereas S3C-expressing cells were fourfold less sensitive to apoptosis. Expression of GRIM-19 restored apoptosis above the levels observed with the vector itself. The mutants, in contrast, failed to induce apoptosis like wild-type GRIM-19. Thus, Fas induction in the presence of GRIM-19 enhanced cellular sensitivity to apoptosis.
Figure 5.
Effect of GRIM-19 mutants on S3C-responsive gene expression, apoptosis, and transformation by v-Src. A: Stat3-responsive gene expression was measured using real-time PCR after infection of 3Y1 cells expressing S3C26 with lentiviral particles coding for the indicated GRIM-19 mutants. Each bar represents the mean ± SE of six replicates in each experiment (n = 3). B: Effect of Fas ligation on cell death in the presence of S3C and GRIM-19 mutants. The Jo-1 monoclonal antibody (200 ng/ml), which activates the Fas death receptor, was incubated for 16 hours before cells were stained with tetramethylrhodamine B isothiocyanate-labeled annexin V. The percentage of cells stained positive (apoptotic) was quantified after florescence-activated cell sorting and plotted. Each data point represents the mean ± SE of triplicates in each experiment (n = 3). C: Effect of GRIM-19 mutants on v-Src-induced cellular transformation. 3Y1 cells expressing v-Src were infected with lentiviral vectors coding for the indicated mutants. Soft agar colony formation was monitored. Mean ± SE were plotted in each case (n = 5). Myc-tag-specific antibody was used in Western blots to detect GRIM-19. D: Interaction of STAT3 with GRIM-19 mutants in HSC3 cells. Coimmunoprecipitation (IP) and Western blot (WB) analysis were performed after infection with lentiviral particles coding for GRIM-19. Statistical significance was obtained by Student’s t-test: *P < 0.01; **P < 0.005; ***P < 0.001. EV, empty vector control; WT, wild-type.
Last, we assessed the effects of select mutants (K5N, D9A, and NΔ17), as representatives of the N-terminal region, for their effects on cellular transformation induced by v-Src, a known activator of STAT3.33,34 We have recently shown that GRIM-19 suppressed v-Src-induced transformation.11 3Y1 cells have lower GRIM-19 levels than many human tumor cell lines (data not shown). Therefore, we infected v-Src-expressing 3Y1 cells with lentiviral particles coding for wild-type and mutant versions of GRIM-19. Soft agar colony formation was estimated in each case (Figure 5C). As expected, the NΔ17 mutant completely lost the antitransforming activity, although these cells expressed GRIM-19 at comparable levels. To understand the biochemical differences between wild-type and mutant GRIM-19 proteins we tested their ability to interact with endogenous STAT3 in HSC3 cells. Cell extracts were prepared, and total protein (200 μg) from each sample was used for immunoprecipitation analysis with a STAT3-specific antibody. The immunoprecipitation products were subjected to Western blot analysis with an Myc-tag-specific antibody. Although wild-type GRIM-19 was pulled down with the STAT3-specific antibody, mutants D9A and K5N poorly coimmunoprecipitated (12 to 15% of the wild-type level) with STAT3 (Figure 5D). The NΔ17 mutant did not have detectable interaction with STAT3. As expected, no GRIM-19 signals were seen in empty vector-transfected cells (data not shown). These cells expressed comparable levels of GRIM-19 mutants and endogenous STAT3. Thus, the weaker interaction of mutant GRIM-19 with STAT3 seems to be responsible for their ineffectiveness as growth suppressors.
Experimental and Clinical Mutations in N Terminus of GRIM-19 Cripple Its Antitumor Action
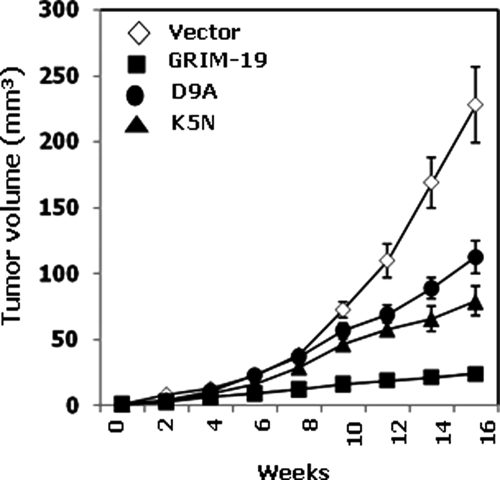
To test the biological relevance of the mutations, we transplanted the HSC3 cell line expressing K5N and D9A subcutaneously into athymic nude mice. In parallel, empty vector and wild-type GRIM-19-expressing cells were used as controls. As shown in Figure 6,31 the empty vector-transfected cells formed the largest tumors (P < 0.001) by the end of 16 weeks. These tumors grew differently from the GRIM-19-expressing tumors by the 8th week. Wild-type GRIM-19-expressing tumors grew weakly under these conditions. The K5N and D9A tumors grew differently from the empty vector- and wild-type GRIM-19-expressing tumors (P < 0.005). Unlike, wild-type GRIM-19-expressing tumors, the mutant-expressing tumors grew faster. However, they did not grow as aggressively as the empty vector-expressing tumors. Thus, experimental and clinical mutations in human GRIM-19 cripple the antitumor actions.
Figure 6.
Effect of GRIM-19 mutants on tumor growth in vivo. HSC3 cell lines expressing the indicated genes were transplanted into athymic nude mice (n = 10) subcutaneously, and tumor growth was monitored as in our earlier studies.31 Significant growth differences between wild-type GRIM-19 and empty vector control (P < 0.001), D9A (P < 0.005), and K5N (P < 0.005) are seen. Statistical significance was obtained by Student’s t-test.
Discussion
Activating mutations in oncogenes in conjunction with the inactivating mutations in tumor suppressor genes promote unregulated cell growth. Using a genetic method, we isolated a novel growth-suppressive gene GRIM19.4 One of the major cellular targets of GRIM-19 is transcription factor STAT3. STAT3 is transiently activated via phosphorylation at Y705 and S727 residues, which are required for dimerization and transcriptional activation, respectively, by a number of cytokines (using JAKs) and growth factors (using receptor tyrosine kinases) in normal cells.35 These signals are rapidly terminated by feedback inhibitory mechanisms.36,37 In contrast to this, STAT3 is constitutively activated by a number of viral and cellular oncogenes; and chronic STAT3 activation has been documented in a number of human cancers.35,38 STAT3 induces expression of a number of genes involved in tumor growth, eg, BclII, Mcl1, Bcl-XL (anti-apoptotic members of the BclII family); cyclins B1 and D1, cdc2 (cell division-associated proteins); c-fos; c-myc, M-Ras, and c-Met (oncoproteins)25,39,40; hypoxia-inducible factor and vascular endothelial growth factor (angiogenic factors)40,41; and metastasis-associated matrix metalloproteases.42,43,44,45,46 Indeed, an experimentally engineered constitutively-dimerizing Stat3 (S3C) oncogenically transforms cells by inhibiting apoptosis.25,47 Interference with STAT3 activity through the use of dominant-negative mutants, DNA oligonucleotides, and short hairpin RNAs causes growth suppression via induction of apoptosis.43,48,49,50 A compelling role for STAT3 in NPM-ALK oncogene-induced tumor formation has been shown.51 A recent study with skin-specific transgenic S3C mice showed that it promotes carcinogen-induced tumor formation.52 The requirement of carcinogenic treatment in these mice suggests that inactivation of STAT3 inhibitors may be necessary for promoting S3C-dependent tumor formation.
It is not clear what mechanisms deregulate STAT3 activity in tumors, apart from its phosphorylation by oncogenic tyrosine kinases. The cytokine-induced STAT3 activity is tightly regulated by feedback inhibitors.37 It is likely that inactivation of these inhibitors may be a key step in growth promotion. There are three known inhibitors of STAT3, including GRIM-19. One of them, suppressor of cytokine signaling 3 (SOCS3), inhibits the activation of JAKs to prevent STAT3 phosphorylation in cytokine-induced signal transduction pathways by acting as a feedback inhibitor.53,54 In some lung cancers, SOCS3 is hypermethylated,55 indicating a role for STAT inhibitors in growth regulation. However, STAT3 activation by endogenous activated oncogenes and autocrine growth factors seems to be insensitive to SOCS-mediated inhibition.56 In fact, STAT3 is constitutively active in some solid tumors despite the presence of normal SOCS3.40 This apparent insensitivity of STAT3 to SOCS3 could also be due to the fact that it can induce growth-promoting gene expression, independently of tyrosyl phosphorylation,40 by heterodimerizing with the RelA (p65) form of nuclear factor-κB.57 Furthermore, SOCS proteins target only JAKs, not receptor tyrosine kinases.37 Thus, unlike the transiently-regulated STAT3 dimer (formed in response to cytokine treatment), the chronic responses (oncogenic) are dependent on unique STAT3 complexes with other proteins. The physiological and antitumor role of another inhibitor, PIAS3, a protein inhibitor of the DNA-binding property of STAT3, is unknown.58 Its inhibitory effects on STAT3 have been primarily shown using simple reporter gene assays and by overexpression in cells. This protein seems to target steroid receptors more than STAT3 itself.59,60,61
Based on the data presented in this article, we believe that GRIM-19 is a novel tumor suppressor and its dysfunction could potentially deregulate STAT3 activity and tumor growth. We have shown that GRIM-19 suppresses oncogenic S3C- and v-Src-induced cellular transformation.11,26 Last, direct administration of a plasmid expressing GRIM-19 into tumors efficiently suppressed tumor growth in vivo.62 Consistent with these results, we reported a loss of GRIM19 expression in primary human renal cell carcinomas13 and prostate carcinomas.62 More importantly, GRIM-19 associates with STAT3, independently of its Y705 residue.16 As mentioned earlier, STAT3 can promote growth-associated gene expression without requiring this residue.40 Many STAT3-regulated genes studied in this report are from nuclear genomes and are regulated independently of Y705 phosphorylation.
Using growth assays, gene expression measurements, transformation, and soft agar growth assays, we present a prominent role for the N terminus of GRIM-19 in regulating antitumor activity. Experiments reported here were verified in three independent cell lines, HSC3 (squamous cell carcinoma), HeLa (cervical carcinoma), and 3Y1 (a nononcogenic murine fibroblast cell line), indicating the validity of observations in several cell types. The use of a GRIM-19-defective cell line, HSC3, for measuring the impact of GRIM19 mutants unequivocally defines their effects. We present three important pieces of evidence that qualify GRIM-19 to be a new tumor suppressor. i) We have identified multiple tumor-derived cell lines with extremely low levels of GRIM-19. These cell lines also had high STAT3 levels and/or activity. More importantly, restoration of GRIM-19 in such cell lines was accompanied by restrained STAT3-responsive gene expression and growth suppression. ii) We have defined a critical motif in the N terminus required for mediating anti-STAT3 activities. iii) Very importantly, we have defined a tumor-derived mutation that lost the antitumor activity. Together, these data establish the tumor suppressor-like credentials for GRIM-19. There is a significant difference in GRIM-19 levels between HSC2 and HSC3 cells (Figure 1A). This difference is consistent with higher STAT3 activity in these cells. However, the differences in GRIM-19 levels do not show a 1:1 correspondence to STAT3 levels in these cell lines for several reasons: i) these two cell lines are derived from different patients; ii) they originated from two different tissues in the mouth (HSC2 from the floor of the mouth and HSC3 from the tongue); and iii) it is likely that there may be other uncharacterized differences between these cell lines. Thus, the differences in STAT3 levels of HSC2 and HSC3 may be due to a combination of GRIM-19 levels and other unknown genetic/epigenetic changes. Remarkably, GRIM-19 seems to behave like the well known tumor suppressor p53.63 Like p53, GRIM-19 is also i) inhibited by viral gene products,18,19,64 ii) mutated/lost in human tumors,24 iii) a modulator of apoptosis by altering gene expression16,26, and iv) a modulator of mitochondria-dependent death pathways, ie, both use HtrA2, a serine protease for suppressing the IAP proteins.9,65 Like GRIM-19,5,7 p53 is located in the mitochondrion and regulates respiration.66,67,68 However, unlike p53, GRIM-19 is not a transcription factor. Whereas p53 induces proapoptotic proteins,69,70,71 GRIM-19 inhibits the expression of antiapoptotic proteins.16,26 The HSC3 cell line used in these studies carries mutant p53 alleles.22 Although Ca9-2 cells also bore mutant p53 alleles,22 it still expressed high amounts of GRIM-19. MCF-7 cells have a wild-type p53 and express higher levels of GRIM-19. Thus, there seems to be no direct correlation between p53 status and GRIM-19 expression. Strikingly, despite p53 mutations in HSC3 cells, restoration of GRIM-19 alone is sufficient for restraining growth. Thus, p53-independent tumor-suppressive pathways can control growth and GRIM-19 seems to promote one such critical pathway. Mechanistically, we have previously shown that S3C-expressing cells, compared with the control vector-transfected cells, did not undergo apoptosis on Fas ligation.26 However, S3C/GRIM-19-expressing cells regained their sensitivity to Fas-dependent apoptosis. Consistent with these observations, cells expressing GRIM-19 mutants had a significantly lower sensitivity to Fas-induced apoptosis, compared to wild-type GRIM-19 (Figure 5B).
GRIM-19 is also present in the mitochondrial compartment.5,7 A recent study showed that STAT3 is also present in the mitochondrion of pro-B cells.72 A mitochondrial targeting sequence in GRIM-19 was mapped to amino acid residues 20 to 60.73 However, the confocal images presented in that report still showed a mitochondrial presence of the mutants. Furthermore, the domain investigated in our study is outside such a motif. It should be noted that GRIM-19 is not exclusively mitochondrial (Figure 2G). In addition, our chromatin immunoprecipitation analyses have shown promoter-bound GRIM-19 in a complex with STAT3 in the context of many nuclear genes.26 Therefore, we believe that GRIM-19 targets nuclear STAT3 to repress gene expression. Moreover, GRIM-19 interacts with NOD2, a cytoplasmic protein, involved in antibacterial responses10 and GW112, a secreted glycoprotein up-regulated during the S phase of the cell cycle.17 Thus, all STAT3/GRIM-19 interactions and/or all GRIM-19 in a cell are not entirely mitochondrial.
The critical growth-regulatory motif mapped in this report is homologous to a motif found in some viral proteins, specifically influenza viral hemagglutinin and echovirus protein V. It is likely that these viral motifs may be necessary for an interaction with cellular proteins. Although the exact mechanisms and targets of this motif in hemagglutinin protein is unclear, the echovirus motif seems to interact with a cellular protein,74 the decay accelerating factor. This protein seems to promote complement-mediated lysis of cells.75 However, the biological effects of these interactions are far from clear. Because the GRIM-19 amino acid sequence shows no conserved motifs/signatures, we modeled its three-dimensional structure using RAPTOR software (Bioinformatics Solutions Inc., Waterloo, ON, Canada) that uses a protein-threading algorithm. When there is marginal sequence homology, the software scans the protein sequence against a database of known structures. By using a scoring function with compatibility analysis between three-dimensional structures and linear protein sequences, the best structural template is used to build the sequence’s structure. The model suggested that the N terminus of GRIM-19 acts a free structure away from the compacted central helical region (data not shown). Based on this, we suggest that the central α-helices of GRIM-19 form a dimerization domain, whereas the N termini permit its interaction with other proteins. The validity of this model needs to be further verified by structural studies. These are being pursued currently. In the meantime, the biochemical and biological data presented in this study represent compelling evidence for a critical function of the N terminus of GRIM-19.
Footnotes
Address reprint requests to Dhananjaya V. Kalvakolanu, Ph.D., 660 W. Redwood St., Howard Hall 350, University of Maryland School of Medicine, Baltimore, MD 21201. E-mail: dkalvako@umaryland.edu.
Supported by the National Institutes of Health (grant R01 CA105005 to D.V.K.).
References
- Schindler C, Levy DE, Decker T. JAK-STAT signaling: from interferons to cytokines. J Biol Chem. 2007;282:20059–20063. doi: 10.1074/jbc.R700016200. [DOI] [PubMed] [Google Scholar]
- Kalvakolanu DV. The GRIMs: a new interface between cell death regulation and interferon/retinoid induced growth suppression. Cytokine Growth Factor Rev. 2004;15:169–194. doi: 10.1016/j.cytogfr.2004.01.002. [DOI] [PubMed] [Google Scholar]
- Kimchi A. DAP genes: novel apoptotic genes isolated by a functional approach to gene cloning. Biochim Biophys Acta. 1998;1377:F13–F33. doi: 10.1016/s0304-419x(98)00002-x. [DOI] [PubMed] [Google Scholar]
- Angell JE, Lindner DJ, Shapiro PS, Hofmann ER, Kalvakolanu DV. Identification of GRIM-19, a novel cell death-regulatory gene induced by the interferon-β and retinoic acid combination, using a genetic approach. J Biol Chem. 2000;275:33416–33426. doi: 10.1074/jbc.M003929200. [DOI] [PubMed] [Google Scholar]
- Fearnley IM, Carroll J, Shannon RJ, Runswick MJ, Walker JE, Hirst J. GRIM-19, a cell death regulatory gene product, is a subunit of bovine mitochondrial NADH: ubiquinone oxidoreductase (complex I). J Biol Chem. 2001;276:38345–38348. doi: 10.1074/jbc.C100444200. [DOI] [PubMed] [Google Scholar]
- Vogel RO, Dieteren CE, van den Heuvel LP, Willems PH, Smeitink JA, Koopman WJ, Nijtmans LG. Identification of mitochondrial complex I assembly intermediates by tracing tagged NDUFS3 demonstrates the entry point of mitochondrial subunits. J Biol Chem. 2007;282:7582–7590. doi: 10.1074/jbc.M609410200. [DOI] [PubMed] [Google Scholar]
- Huang G, Lu H, Hao A, Ng DC, Ponniah S, Guo K, Lufei C, Zeng Q, Cao X. GRIM-19, a cell death regulatory protein, is essential for assembly and function of mitochondrial complex I. Mol Cell Biol. 2004;24:8447–8456. doi: 10.1128/MCB.24.19.8447-8456.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang G, Chen Y, Lu H, Cao X. Coupling mitochondrial respiratory chain to cell death: an essential role of mitochondrial complex I in the interferon-β and retinoic acid-induced cancer cell death. Cell Death Differ. 2007;14:327–337. doi: 10.1038/sj.cdd.4402004. [DOI] [PubMed] [Google Scholar]
- Ma X, Kalakonda S, Srinivasula SM, Reddy SP, Platanias LC, Kalvakolanu DV. GRIM-19 associates with the serine protease HtrA2 for promoting cell death. Oncogene. 2007;26:4842–4849. doi: 10.1038/sj.onc.1210287. [DOI] [PubMed] [Google Scholar]
- Barnich N, Hisamatsu T, Aguirre JE, Xavier R, Reinecker HC, Podolsky DK. GRIM-19 interacts with NOD2 and serves as down-stream effector of anti-bacterial function in intestinal epithelial cells. J Biol Chem. 2005;280:19021–19026. doi: 10.1074/jbc.M413776200. [DOI] [PubMed] [Google Scholar]
- Kalakonda S, Nallar SC, Gong P, Lindner DJ, Goldblum SE, Reddy SP, Kalvakolanu DV. Tumor suppressive protein gene associated with retinoid-interferon-induced mortality (GRIM)-19 inhibits src-induced oncogenic transformation at multiple levels. Am J Pathol. 2007;171:1352–1368. doi: 10.2353/ajpath.2007.070241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y, Yuen WH, Fu J, Huang G, Melendez AJ, Ibrahim FB, Lu H, Cao X. The mitochondrial respiratory chain controls intracellular calcium signaling and NFAT activity essential for heart formation in Xenopus laevis. Mol Cell Biol. 2007;27:6420–6432. doi: 10.1128/MCB.01946-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alchanati I, Nallar SC, Sun P, Gao L, Hu J, Stein A, Yakirevich E, Konforty D, Alroy I, Zhao X, Reddy SP, Resnick MB, Kalvakolanu DV. A proteomic analysis reveals the loss of expression of the cell death regulatory gene GRIM-19 in human renal cell carcinomas. Oncogene. 2006;25:7138–7147. doi: 10.1038/sj.onc.1209708. [DOI] [PubMed] [Google Scholar]
- Gong LB, Luo XL, Liu SY, Tao DD, Gong JP, Hu JB. Correlations of GRIM-19 and its target gene product STAT3 to malignancy of human colorectal carcinoma. Ai Zheng. 2007;26:683–687. [PubMed] [Google Scholar]
- Lufei C, Ma J, Huang G, Zhang T, Novotny-Diermayr V, Ong CT, Cao X. GRIM-19, a death-regulatory gene product, suppresses Stat3 activity via functional interaction. EMBO J. 2003;22:1325–1335. doi: 10.1093/emboj/cdg135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Yang J, Roy SK, Tininini S, Hu J, Bromberg JF, Poli V, Stark GR, Kalvakolanu DV. The cell death regulator GRIM-19 is an inhibitor of signal transducer and activator of transcription 3. Proc Natl Acad Sci USA. 2003;100:9342–9347. doi: 10.1073/pnas.1633516100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Huang Q, Yang Z, Li Y, Li CY. GW112, a novel antiapoptotic protein that promotes tumor growth. Cancer Res. 2004;64:2474–2481. doi: 10.1158/0008-5472.can-03-3443. [DOI] [PubMed] [Google Scholar]
- Seo T, Lee D, Shim YS, Angell JE, Chidambaram NV, Kalvakolanu DV, Choe J. Viral interferon regulatory factor 1 of Kaposi’s sarcoma-associated herpesvirus interacts with a cell death regulator, GRIM19, and inhibits interferon/retinoic acid-induced cell death. J Virol. 2002;76:8797–8807. doi: 10.1128/JVI.76.17.8797-8807.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reeves MB, Davies AA, McSharry BP, Wilkinson GW, Sinclair JH. Complex I binding by a virally encoded RNA regulates mitochondria-induced cell death. Science. 2007;316:1345–1348. doi: 10.1126/science.1142984. [DOI] [PubMed] [Google Scholar]
- Momose F, Araida T, Negishi A, Ichijo H, Shioda S, Sasaki S. Variant sublines with different metastatic potentials selected in nude mice from human oral squamous cell carcinomas. J Oral Pathol Med. 1989;18:391–395. doi: 10.1111/j.1600-0714.1989.tb01570.x. [DOI] [PubMed] [Google Scholar]
- Kamata N, Chida K, Rikimaru K, Horikoshi M, Enomoto S, Kuroki T. Growth-inhibitory effects of epidermal growth factor and overexpression of its receptors on human squamous cell carcinomas in culture. Cancer Res. 1986;46:1648–1653. [PubMed] [Google Scholar]
- Sakai E, Tsuchida N. Most human squamous cell carcinomas in the oral cavity contain mutated p53 tumor-suppressor genes. Oncogene. 1992;7:927–933. [PubMed] [Google Scholar]
- Hu J, Angell JE, Zhang J, Ma X, Seo T, Raha A, Hayashi J, Choe J, Kalvakolanu DV. Characterization of monoclonal antibodies against GRIM-19, a novel IFN-β and retinoic acid-activated regulator of cell death. J Interferon Cytokine Res. 2002;22:1017–1026. doi: 10.1089/107999002760624242. [DOI] [PubMed] [Google Scholar]
- Máximo V, Botelho T, Capela J, Soares P, Lima J, Taveira A, Amaro T, Barbosa AP, Preto A, Harach HR, Williams D, Sobrinho-Simoes M. Somatic and germline mutation in GRIM-19, a dual function gene involved in mitochondrial metabolism and cell death, is linked to mitochondrion-rich (Hürthle cell) tumours of the thyroid. Br J Cancer. 2005;92:1892–1898. doi: 10.1038/sj.bjc.6602547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bromberg JF, Wrzeszczynska MH, Devgan G, Zhao Y, Pestell RG, Albanese C, Darnell JE., Jr Stat3 as an oncogene. Cell. 1999;98:295–303. doi: 10.1016/s0092-8674(00)81959-5. [DOI] [PubMed] [Google Scholar]
- Kalakonda S, Nallar SC, Lindner DJ, Hu J, Reddy SP, Kalvakolanu DV. Tumor-suppressive activity of the cell death activator GRIM-19 on a constitutively active signal transducer and activator of transcription 3. Cancer Res. 2007;67:6212–6220. doi: 10.1158/0008-5472.CAN-07-0031. [DOI] [PubMed] [Google Scholar]
- Skehan P, Storeng R, Scudiero D, Monks A, McMahon J, Vistica D, Warren JT, Bokesch H, Kenney S, Boyd MR. New colorimetric cytotoxicity assay for anticancer-drug screening. J Natl Cancer Inst. 1990;82:1107–1112. doi: 10.1093/jnci/82.13.1107. [DOI] [PubMed] [Google Scholar]
- Scott LA, Vass JK, Parkinson EK, Gillespie DA, Winnie JN, Ozanne BW. Invasion of normal human fibroblasts induced by v-Fos is independent of proliferation, immortalization, and the tumor suppressors p16INK4a and p53. Mol Cell Biol. 2004;24:1540–1559. doi: 10.1128/MCB.24.4.1540-1559.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma X, Karra S, Guo W, Lindner DJ, Hu J, Angell JE, Hofmann ER, Reddy SP, Kalvakolanu DV. Regulation of interferon and retinoic acid-induced cell death activation through thioredoxin reductase. J Biol Chem. 2001;276:24843–24854. doi: 10.1074/jbc.M100380200. [DOI] [PubMed] [Google Scholar]
- Hoffmann K, Blaudszun J, Brunken C, Hopker WW, Tauber R, Steinhart H. New application of a subcellular fractionation method to kidney and testis for the determination of conjugated linoleic acid in selected cell organelles of healthy and cancerous human tissues. Anal Bioanal Chem. 2005;381:1138–1144. doi: 10.1007/s00216-004-3009-z. [DOI] [PubMed] [Google Scholar]
- Lindner DJ, Borden EC, Kalvakolanu DV. Synergistic antitumor effects of a combination of interferons and retinoic acid on human tumor cells in vitro and in vivo. Clin Cancer Res. 1997;3:931–937. [PubMed] [Google Scholar]
- Ivanov VN, Bhoumik A, Krasilnikov M, Raz R, Owen-Schaub LB, Levy D, Horvath CM, Ronai Z. Cooperation between STAT3 and c-jun suppresses Fas transcription. Mol Cell. 2001;7:517–528. doi: 10.1016/s1097-2765(01)00199-x. [DOI] [PubMed] [Google Scholar]
- Yu CL, Meyer DJ, Campbell GS, Larner AC, Carter-Su C, Schwartz J, Jove R. Enhanced DNA-binding activity of a Stat3-related protein in cells transformed by the Src oncoprotein. Science. 1995;269:81–83. doi: 10.1126/science.7541555. [DOI] [PubMed] [Google Scholar]
- Bromberg JF, Horvath CM, Besser D, Lathem WW, Darnell JE., Jr Stat3 activation is required for cellular transformation by v-src. Mol Cell Biol. 1998;18:2553–2558. doi: 10.1128/mcb.18.5.2553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bromberg J. Stat proteins and oncogenesis. J Clin Invest. 2002;109:1139–1142. doi: 10.1172/JCI15617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levy DE, Darnell JE. STATs: transcriptional control and biological impact. Nat Rev Mol Cell Biol. 2002;3:651–662. doi: 10.1038/nrm909. [DOI] [PubMed] [Google Scholar]
- Alexander WS, Hilton DJ. The role of suppressors of cytokine signaling (SOCS) proteins in regulation of the immune response. Annu Rev Immunol. 2004;22:503–529. doi: 10.1146/annurev.immunol.22.091003.090312. [DOI] [PubMed] [Google Scholar]
- Buettner R, Mora LB, Jove R. Activated STAT signaling in human tumors provides novel molecular targets for therapeutic intervention. Clin Cancer Res. 2002;8:945–954. [PubMed] [Google Scholar]
- Dauer DJ, Ferraro B, Song L, Yu B, Mora L, Buettner R, Enkemann S, Jove R, Haura EB. Stat3 regulates genes common to both wound healing and cancer. Oncogene. 2005;24:3397–3408. doi: 10.1038/sj.onc.1208469. [DOI] [PubMed] [Google Scholar]
- Yang J, Chatterjee-Kishore M, Staugaitis SM, Nguyen H, Schlessinger K, Levy DE, Stark GR. Novel roles of unphosphorylated STAT3 in oncogenesis and transcriptional regulation. Cancer Res. 2005;65:939–947. [PubMed] [Google Scholar]
- Gray MJ, Zhang J, Ellis LM, Semenza GL, Evans DB, Watowich SS, Gallick GE. HIF-1α, STAT3, CBP/p300 and Ref-1/APE are components of a transcriptional complex that regulates Src-dependent hypoxia-induced expression of VEGF in pancreatic and prostate carcinomas. Oncogene. 2005;24:3110–3120. doi: 10.1038/sj.onc.1208513. [DOI] [PubMed] [Google Scholar]
- Dechow TN, Pedranzini L, Leitch A, Leslie K, Gerald WL, Linkov I, Bromberg JF. Requirement of matrix metalloproteinase-9 for the transformation of human mammary epithelial cells by Stat3-C. Proc Natl Acad Sci USA. 2004;101:10602–10607. doi: 10.1073/pnas.0404100101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao L, Zhang L, Hu J, Li F, Shao Y, Zhao D, Kalvakolanu DV, Kopecko DJ, Zhao X, Xu DQ. Down-regulation of signal transducer and activator of transcription 3 expression using vector-based small interfering RNAs suppresses growth of human prostate tumor in vivo. Clin Cancer Res. 2005;11:6333–6341. doi: 10.1158/1078-0432.CCR-05-0148. [DOI] [PubMed] [Google Scholar]
- Song Y, Qian L, Song S, Chen L, Zhang Y, Yuan G, Zhang H, Xia Q, Hu M, Yu M, Shi M, Jiang Z, Guo N. Fra-1 and Stat3 synergistically regulate activation of human MMP-9 gene. Mol Immunol. 2008;45:137–143. doi: 10.1016/j.molimm.2007.04.031. [DOI] [PubMed] [Google Scholar]
- Wang L, Luo J, He S. Induction of MMP-9 release from human dermal fibroblasts by thrombin: involvement of JAK/STAT3 signaling pathway in MMP-9 release. BMC Cell Biol. 2007;8:14. doi: 10.1186/1471-2121-8-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Itoh M, Murata T, Suzuki T, Shindoh M, Nakajima K, Imai K, Yoshida K. Requirement of STAT3 activation for maximal collagenase-1 (MMP-1) induction by epidermal growth factor and malignant characteristics in T24 bladder cancer cells. Oncogene. 2006;25:1195–1204. doi: 10.1038/sj.onc.1209149. [DOI] [PubMed] [Google Scholar]
- Shen Y, Devgan G, Darnell JE, Jr, Bromberg JF. Constitutively activated Stat3 protects fibroblasts from serum withdrawal and UV-induced apoptosis and antagonizes the proapoptotic effects of activated Stat1. Proc Natl Acad Sci USA. 2001;98:1543–1548. doi: 10.1073/pnas.041588198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niu G, Heller R, Catlett-Falcone R, Coppola D, Jaroszeski M, Dalton W, Jove R, Yu H. Gene therapy with dominant-negative Stat3 suppresses growth of the murine melanoma B16 tumor in vivo. Cancer Res. 1999;59:5059–5063. [PubMed] [Google Scholar]
- DeArmond D, Brattain MG, Jessup JM, Kreisberg J, Malik S, Zhao S, Freeman JW. Autocrine-mediated ErbB-2 kinase activation of STAT3 is required for growth factor independence of pancreatic cancer cell lines. Oncogene. 2003;22:7781–7795. doi: 10.1038/sj.onc.1206966. [DOI] [PubMed] [Google Scholar]
- Leong PL, Andrews GA, Johnson DE, Dyer KF, Xi S, Mai JC, Robbins PD, Gadiparthi S, Burke NA, Watkins SF, Grandis JR. Targeted inhibition of Stat3 with a decoy oligonucleotide abrogates head and neck cancer cell growth. Proc Natl Acad Sci USA. 2003;100:4138–4143. doi: 10.1073/pnas.0534764100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiarle R, Simmons WJ, Cai H, Dhall G, Zamo A, Raz R, Karras JG, Levy DE, Inghirami G. Stat3 is required for ALK-mediated lymphomagenesis and provides a possible therapeutic target. Nat Med. 2005;11:623–629. doi: 10.1038/nm1249. [DOI] [PubMed] [Google Scholar]
- Chan KS, Sano S, Kataoka K, Abel E, Carbajal S, Beltran L, Clifford J, Peavey M, Shen J, Digiovanni J. Forced expression of a constitutively active form of Stat3 in mouse epidermis enhances malignant progression of skin tumors induced by two-stage carcinogenesis. Oncogene. 2008;27:1087–1094. doi: 10.1038/sj.onc.1210726. [DOI] [PubMed] [Google Scholar]
- Croker BA, Krebs DL, Zhang JG, Wormald S, Willson TA, Stanley EG, Robb L, Greenhalgh CJ, Forster I, Clausen BE, Nicola NA, Metcalf D, Hilton DJ, Roberts AW, Alexander WS. SOCS3 negatively regulates IL-6 signaling in vivo. Nat Immunol. 2003;4:540–545. doi: 10.1038/ni931. [DOI] [PubMed] [Google Scholar]
- Lang R, Pauleau AL, Parganas E, Takahashi Y, Mages J, Ihle JN, Rutschman R, Murray PJ. SOCS3 regulates the plasticity of gp130 signaling. Nat Immunol. 2003;4:546–550. doi: 10.1038/ni932. [DOI] [PubMed] [Google Scholar]
- He B, You L, Uematsu K, Zang K, Xu Z, Lee AY, Costello JF, McCormick F, Jablons DM. SOCS-3 is frequently silenced by hypermethylation and suppresses cell growth in human lung cancer. Proc Natl Acad Sci USA. 2003;100:14133–14138. doi: 10.1073/pnas.2232790100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwamoto T, Senga T, Naito Y, Matsuda S, Miyake Y, Yoshimura A, Hamaguchi M. The JAK-inhibitor. JAB/SOCS-1 selectively inhibits cytokine-induced, but not v-Src induced JAK-STAT activation. Oncogene. 2000;19:4795–4801. doi: 10.1038/sj.onc.1203829. [DOI] [PubMed] [Google Scholar]
- Yang J, Liao X, Agarwal MK, Barnes L, Auron PE, Stark GR. Unphosphorylated STAT3 accumulates in response to IL-6 and activates transcription by binding to NFκB. Genes Dev. 2007;21:1396–1408. doi: 10.1101/gad.1553707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung CD, Liao J, Liu B, Rao X, Jay P, Berta P, Shuai K. Specific inhibition of Stat3 signal transduction by PIAS3. Science. 1997;278:1803–1805. doi: 10.1126/science.278.5344.1803. [DOI] [PubMed] [Google Scholar]
- Gross M, Liu B, Tan J, French FS, Carey M, Shuai K. Distinct effects of PIAS proteins on androgen-mediated gene activation in prostate cancer cells. Oncogene. 2001;20:3880–3887. doi: 10.1038/sj.onc.1204489. [DOI] [PubMed] [Google Scholar]
- Junicho A, Matsuda T, Yamamoto T, Kishi H, Korkmaz K, Saatcioglu F, Fuse H, Muraguchi A. Protein inhibitor of activated STAT3 regulates androgen receptor signaling in prostate carcinoma cells. Biochem Biophys Res Commun. 2000;278:9–13. doi: 10.1006/bbrc.2000.3753. [DOI] [PubMed] [Google Scholar]
- Sentis S, Le Romancer M, Bianchin C, Rostan MC, Corbo L. Sumoylation of the estrogen receptor α hinge region regulates its transcriptional activity. Mol Endocrinol. 2005;19:2671–2684. doi: 10.1210/me.2005-0042. [DOI] [PubMed] [Google Scholar]
- Zhang L, Gao L, Li Y, Lin G, Shao Y, Ji K, Yu H, Hu J, Kalvakolanu DV, Kopecko DJ, Zhao X, Xu DQ. Effects of plasmid-based Stat3-specific short hairpin RNA and GRIM-19 on PC-3M tumor cell growth. Clin Cancer Res. 2008;14:559–568. doi: 10.1158/1078-0432.CCR-07-1176. [DOI] [PubMed] [Google Scholar]
- Riley T, Sontag E, Chen P, Levine A. Transcriptional control of human p53-regulated genes. Nat Rev Mol Cell Biol. 2008;9:402–412. doi: 10.1038/nrm2395. [DOI] [PubMed] [Google Scholar]
- Guerra S, Lopez-Fernandez LA, Pascual-Montano A, Munoz M, Harshman K, Esteban M. Cellular gene expression survey of vaccinia virus infection of human HeLa cells. J Virol. 2003;77:6493–6506. doi: 10.1128/JVI.77.11.6493-6506.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin S, Kalkum M, Overholtzer M, Stoffel A, Chait BT, Levine AJ. CIAP1 and the serine protease HTRA2 are involved in a novel p53-dependent apoptosis pathway in mammals. Genes Dev. 2003;17:359–367. doi: 10.1101/gad.1047003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matoba S, Kang JG, Patino WD, Wragg A, Boehm M, Gavrilova O, Hurley PJ, Bunz F, Hwang PM. p53 regulates mitochondrial respiration. Science. 2006;312:1650–1653. doi: 10.1126/science.1126863. [DOI] [PubMed] [Google Scholar]
- Mihara M, Erster S, Zaika A, Petrenko O, Chittenden T, Pancoska P, Moll UM. p53 has a direct apoptogenic role at the mitochondria. Mol Cell. 2003;11:577–590. doi: 10.1016/s1097-2765(03)00050-9. [DOI] [PubMed] [Google Scholar]
- Erster S, Mihara M, Kim RH, Petrenko O, Moll UM. In vivo mitochondrial p53 translocation triggers a rapid first wave of cell death in response to DNA damage that can precede p53 target gene activation. Mol Cell Biol. 2004;24:6728–6741. doi: 10.1128/MCB.24.15.6728-6741.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyashita T, Reed JC. Tumor suppressor p53 is a direct transcriptional activator of the human bax gene. Cell. 1995;80:293–299. doi: 10.1016/0092-8674(95)90412-3. [DOI] [PubMed] [Google Scholar]
- Nakano K, Vousden KH. PUMA, a novel proapoptotic gene is induced by p53. Mol Cell. 2001;7:683–694. doi: 10.1016/s1097-2765(01)00214-3. [DOI] [PubMed] [Google Scholar]
- Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T, Tokino T, Taniguchi T, Tanaka N. Noxa, a BH3-only member of the Bcl-2 family and candidate mediator of p53-induced apoptosis. Science. 2000;288:1053–1058. doi: 10.1126/science.288.5468.1053. [DOI] [PubMed] [Google Scholar]
- Wegrzyn J, Potla R, Chwae YJ, Sepuri NB, Zhang Q, Koeck T, Derecka M, Szczepanek K, Szelag M, Gornicka A, Moh A, Moghaddas S, Chen Q, Bobbili S, Cichy J, Dulak J, Baker DP, Wolfman A, Stuehr D, Hassan MO, Fu XY, Avadhani N, Drake JI, Fawcett P, Lesnefsky EJ, Larner AC. Function of mitochondrial Stat3 in cellular respiration. Science. 2009;323:793–797. doi: 10.1126/science.1164551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu H, Cao X. GRIM-19 is essential for maintenance of mitochondrial membrane potential. Mol Biol Cell. 2008;19:1893–1902. doi: 10.1091/mbc.E07-07-0683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuart AD, McKee TA, Williams PA, Harley C, Shen S, Stuart DI, Brown TD, Lea SM. Determination of the structure of a decay accelerating factor-binding clinical isolate of echovirus 11 allows mapping of mutants with altered receptor requirements for infection. J Virol. 2002;76:7694–7704. doi: 10.1128/JVI.76.15.7694-7704.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shafren DR, Williams DT, Barry RD. A decay-accelerating factor-binding strain of coxsackievirus B3 requires the coxsackievirus-adenovirus receptor protein to mediate lytic infection of rhabdomyosarcoma cells. J Virol. 1997;71:9844–9848. doi: 10.1128/jvi.71.12.9844-9848.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]