Abstract
Molecular interactions between the VAR2CSA protein, expressed on the surface of Plasmodium falciparum-infected erythrocytes, and placental chondroitin sulfate A (CSA) are primarily responsible for pregnancy-associated malaria (PAM). Interrupting these interactions may prevent or ameliorate the severity of PAM. Several of the Duffy binding-like (DBL) domains of VAR2CSA, including the DBL3x domain, have been shown to bind CSA in vitro, but a more detailed understanding of how DBL domains bind CSA is needed. In this study, we demonstrate that subdomain 3 (S3), one of the three subdomains of VAR2CSA DBL3x by itself, is the major contributor toward CSA binding. NMR spectroscopy and flow cytometry analyses show that S3 and the intact DBL3x domain bind CSA similarly. Mutations within the S3 portion of DBL3x markedly affect CSA binding. Both recombinant molecules, S3 and DBL3x, are recognized by antibodies in the plasma of previously pregnant women living in malaria-endemic regions of Mali, but much less so by plasma from men of the same regions. As the S3 sequence is highly conserved in all known VAR2CSA proteins expressed by different parasite isolates obtained from various malaria endemic areas of the world, the identification of S3 as an independent CSA-binding region provides a compelling molecular basis for designing interventions against PAM.
Keywords: Chondroitin Sulfate, Erythrocyte, Parasite, Protein Folding, Receptor Structure-Function, Malaria, Plasmodium Falciparum, Pregnancy-associated Malaria
Introduction
Each year, pregnancy-associated malaria (PAM)3 poses a risk to millions of women worldwide. Although women living in malaria-endemic areas eventually develop immunity to malaria after repeated exposure to Plasmodium falciparum, they become susceptible to severe disease during pregnancy. PAM is caused by the massive sequestration of P. falciparum-infected erythrocytes (IEs) that selectively bind to CSA in the placenta, leading to severe inflammation and life-threatening outcomes for both mother and child (1).
VAR2CSA, a member of the P. falciparum erythrocyte membrane protein 1 (PfEMP1) family, is primarily responsible for the binding of IEs to CSA (2). The var2csa gene is found in nearly all P. falciparum isolates (3) and is unusually conserved compared with other members of the var gene family that encode PfEMP1 molecules (4). VAR2CSA is specifically up-regulated in infected erythrocytes that are either isolated from the placenta (5) or selected in vitro for binding to CSA (2). On disruption of the var2csa gene, the ability of the infected erythrocytes to adhere to CSA is lost (6) or greatly reduced (7).
The risk of PAM is highest in the first pregnancy due to lack of VAR2CSA-specific antibodies, but with increasing parity, women acquire such antibodies that block the binding of IEs to CSA in the placenta. The naturally acquired antibodies from multiparous women collected from different malaria-endemic regions of the world recognize CSA-binding IEs in the placenta (8–12). The ability of such antibodies to block adhesion of IEs to CSA by recognizing conserved epitopes among different parasite clones, suggests that VAR2CSA-based vaccination against PAM is possible.
Expressing recombinant VAR2CSA proteins, which have molecular masses of ∼350 kDa and have six DBL domains with multiple disulfide bonds, is a major challenge. As reported earlier, four of the six DBL domains (DBL2x, DBL3x, DBL5, and DBL6) of VAR2CSA have been shown to bind to CSA (13, 14). Crystal structures have given insight into DBL3x (15, 16). The structure of DBL3x in complex with the CSA oligosaccharide enhanced our understanding of the molecular interactions between DBL3x and CSA, by showing electron density for CSA located close to the positively charged pocket between subdomain 2 (S2) and subdomain 3 (S3) of DBL3x (15).
Chondroitin sulfates are unbranched glycosaminoglycans of variable length, sulfate content, and sulfation pattern. They are composed of repeating disaccharides, which consist of glucuronic acid (GlcA) and N-acetylgalactosamine (GalNAc). Each GalNAc may be nonsulfated or sulfated at the O-4 or O-6 position. Human placenta contains very low sulfated chondroitin sulfate proteoglycan at high levels in the intervillous space, and only 10% of the disaccharide moieties of the chondroitin sulfate chains of chondroitin sulfate proteoglycan are sulfated specifically at O-4 (17). IEs bind mainly to O-4-sulfated chondroitin sulfate proteoglycan and thus sequester in the placenta, and this binding can be inhibited by soluble 4-O-sulfated chondroitin sulfate, but not by 6-O-sulfated chondroitin sulfate (18). It has been demonstrated earlier that a dodecasaccharide fragment of CSA comprises the minimal motif for optimal binding by the full-length VAR2CSA expressed on the surface of IEs (17–20). The DBL3x-CSA complex structure also shows that a hexasaccharide can be positioned at the electron density between S2 and S3 subdomains (15), indicating that DBL3x plays a major role in binding to CSA.
For designing molecular interventions against PAM, it is important to determine an independently folded and stable minimal CSA-binding region of DBL3x. In this study, we have used structure-based mutagenesis, NMR spectroscopy, and immunological studies to demonstrate that S3 is the minimal CSA-binding region, as suggested previously from the crystal structure of the DBL3x-CSA complex (15).
EXPERIMENTAL PROCEDURES
Protein Production
We cloned the S3 subdomain (residues 1446–1580, accession no. AAQ73926) from genomic DNA of the A4 strain (also called FCR3 or IT4) of P. falciparum in the pLM1 expression plasmid (21). The plasmid was transformed into Escherichia coli strain BL21 (DE3)-RIL (Stratagene, La Jolla, CA) cells that were then grown to an A600 of 0.6 at 37 °C and induced with 1 mm isopropyl-β-d-thiogalactopyranoside for 3 h to produce insoluble protein as inclusion bodies. The cells were harvested by centrifugation at 4000 × g and resuspended in 100 ml of lysis buffer (100 mm Tris-HCl, pH 8.0, 150 mm NaCl, 1% Triton X-100, and 1% (w/v) sodium deoxycholate). Cells were lysed by freezing and thawing in the presence of lysozyme (1 mg/ml).
S3 inclusion bodies were purified after multiple washes, centrifugation, and resuspension. The final pellet was dissolved in 6 m guanidine-HCl with 2 mm dithiothreitol. S3 was then refolded and purified as was wild-type DBL3x protein (15). Purified S3 was dialyzed against 10 mm Tris-HCl, pH 7.5, at 4 °C and concentrated in a centrifugal filter device (Centricon YM10, Millipore, Billerica, MA) to 10 mg/ml.
The His6 tag at the C terminus of S3 was added using standard techniques. The His-tagged protein was expressed and refolded under the same conditions as the wild-type protein. The S3 protein was dialyzed against 10 mm Tris-HCl, pH 7.5, at 4 °C and concentrated to 10 mg/ml.
The DBL3x domain was expressed and refolded as reported (15). The single mutants of DBL3x, R1467A, and R1503A and the double mutant K1324A,K1327A were produced using standard techniques and were confirmed by DNA sequencing. The DBL3x mutants were expressed, refolded, and purified similarly to the wild-type protein.
Preparation of Size-fractionated Oligosaccharides from Bovine Tracheal CSA
Bovine tracheal CSA, which consists of 53% 4-O-sulfated, 39% 6-O-sulfated, and 8% nonsulfated disaccharide moieties was partially depolymerized with bovine testicular hyaluronidase, and the oligosaccharides of varying sizes that formed were purified by size exclusion chromatography on Bio-Gel P-6 (18). The dodecasaccharide fragment was used for NMR studies.
Binding of S3 and the DBL3x Mutants to CSA on the Surface of CHO K1 Cells
The binding of S3 and DBL3x mutants to CSA expressed on the surface of Chinese hamster ovary (CHO) K1 cells (American Type Culture Collection) was carried out using flow cytometry as described previously (15, 22). About one million cells (100 μl) were incubated for 30 min with DBL3x (5 μm), mutant DBL3x (5 μm), or S3 (10 μm) either in the presence (50 μm) or absence of bovine tracheal CSA at room temperature. After incubation, cells were washed three times in FACS buffer and treated with anti-DBL3x monoclonal antibody or anti-His monoclonal antibody (MAB050, R&D Systems, Minneapolis, MN) for 30 min at 4 °C. The cells were washed twice in phosphate-buffered saline and incubated further with Alexa 488-labeled goat anti-mouse IgG (100 μl) (catalog no. A11029, Invitrogen) diluted 1:250 in FACS buffer. The cells were then incubated for 30 min, washed, resuspended in 200 μl FACS buffer and analyzed by FACSort (Becton Dickinson). All of the incubations and washing steps were performed at 4 °C. Events (75, 000) were acquired using CellQuest software (version 3.3; Becton Dickinson), and the data from the live cell gate were analyzed by FlowJo software (version 6.4.1; Tree Star, San Carlos, CA).
NMR Spectroscopy
Saturation transfer difference (STD) and two-dimensional transferred nuclear Overhauser effect (2D TrNOE) experiments were performed on a Bruker Avance DRX 500 MHz NMR spectrometer, equipped with a cryoprobe, at 20 °C (STD) and 30 °C (2D TrNOE). The NMR protein samples, containing 20 mm phosphate buffer, pH 7.0, and 50 mm NaCl were lyophilized three times and dissolved in 600 μl of D2O (deuterated water, 99.96%, Cambridge lsotope Laboratories, Inc., Andover, MA). Protein concentrations in the NMR samples were 6 and 60 μm. To assure minimal dilution effects, the lyophilized CSA dodecasaccharide was added to the D2O-exchanged concentrated protein stock solutions. High protein to CSA oligosaccharide molar ratios (1:100 and 1:20) were chosen to be optimal for STD and TrNOE effects, respectively.
The STD NMR experiments were performed as described by Mayer and Meyer (23). Selective protein saturation, off-resonance at 50.00 ppm, and on-resonance at 7.12 ppm, was achieved by a train of 40 Gaussian pulses of 50 ms each, with irradiation power set to ((γ/2π)B1 = 86 Hz), separated by a 1-ms delay for a total saturation time of 2.04 s. A T1ρ filter of a 30 ms spin-lock pulse with a strength of ((γ/2π)B1 = 4960 Hz) was applied to eliminate the background protein resonances. The WATERGATE 3-9-19 pulse sequence was used to suppress the residual water signal. The data were collected with 1024 scans, in an interleaved manner to compensate for the temperature and instrument instability effects. The free induction decays of the off- and on-resonance spectrum were stored and processed separately. Subtraction of the off- and on-resonance spectrum resulted in the STD NMR spectrum. The STD data were processed and analyzed using TopSpin software (Bruker Biospin Corp., Billerica, MA).
The 2D TrNOE experiments (24) of the CSA oligosaccharide only and the CSA oligosaccharide in the presence of either S3 or DBL3x protein (at 20:1 molar ratio) were carried out at 350- and 150-ms mixing times, respectively, with 32 scans and 384 complex data points in t1. The 2D TrNOE data were processed and analyzed using NMRPipe (25) and NMRView software (Bruce A. Johnson, One Moon Scientific, Inc.). Our naming of CSA hydrogens followed the report in which the original assignments were made (26).
ELISA with Plasma from a Malian Population
Human plasma samples were collected in 2008 from adults living in three malaria-endemic areas of Mali (54 males and 39 females ranging in age from 18 to 60 years). The study was approved by the ethical review committees of the Faculty of Medicine, Pharmacy, and Dentistry at the University of Bamako (Mali) and the NIAID, National Institutes of Health (Bethesda, MD). Individual written informed consent was obtained from all participants. The standardized methodology for performing the ELISA has been described previously (27). The minimal detection levels of this assay were 31 ELISA units for DBL3x and 3 ELISA units for S3. Data points below these limits of detection for each antigen were reassigned values of the respective limit of detection. The levels of antigen binding by antibodies in the male and female plasma were compared by an exact Wilcoxon-Mann-Whitney test. The Spearman rank correlation test was used to calculate the correlation between anti-S3 and anti-DBL3x titers. A p value of <0.05 was considered to be significant. ELISA data were analyzed using PRISM (GraphPad Software, Inc., CA).
RESULTS AND DISCUSSION
Residues on S3 Are Predicted to Be Involved in Binding CSA
Crystal structures show that DBL3x is made up of three subdomains (15, 16). Subdomain 1 (S1) (Fig. 1A, yellow) has little regular secondary structure and is held together by two disulfide bonds. S2 (Fig. 1A, blue) contains four helices, connected to each other by loops. S2 has one disulfide bond within the subdomain and a second disulfide bond that links S2 with S1, which holds the subdomains together (16). S3 (Fig. 1A, red) has two long antiparallel helices connected by a large loop. S3 has four disulfide bonds, but as none of these bonds link S3 with either S2 or S1, S3 is a highly cross-linked, independent subdomain.
FIGURE 1.
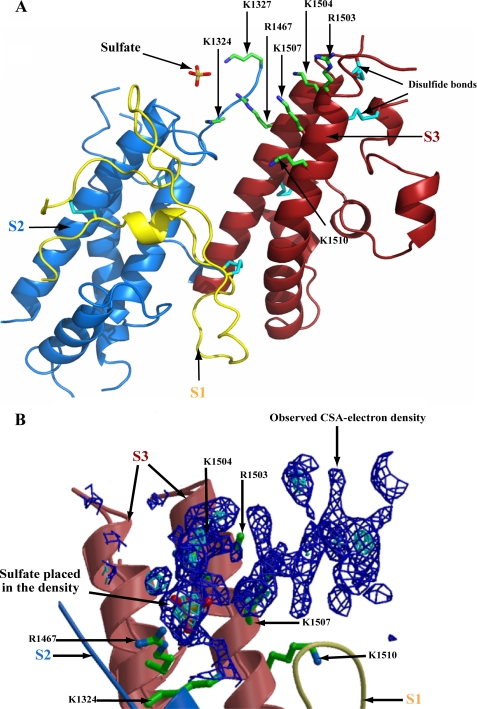
Residues comprising the CSA-binding pocket located on the subdomain structure of DBL3x. A, DBL3x (Protein Data Bank code 3CPZ) is composed of subdomain 1 (S1, yellow), subdomain 2 (S2, blue), and subdomain 3 (S3, red). Indicated are the sulfate group (yellow, sulfur; red, oxygen) that was fitted to CSA electron density and the residues (green) located within 5 Å of the CSA electron density (see B) on S2 (Lys1324, Lys1327) and S3 (Arg1467, Arg1503, Arg1504, Lys1507, and Lys1510). Disulfide bonds are shown in cyan, and three of the four S3 disulfides are visible in this view. B, electron density for CSA (blue) was located across the two long helices of S3. Residues within 5 Å of the CSA density are labeled. Compared with A, this panel is rotated by ∼90° around the vertical axis in the page. Lys1327 cannot be seen in this view.
Various complexes between DBL3x and CSA were obtained by cocrystallizing or soaking DBL3x crystals with different sizes of CSA oligosaccharides. After a detailed and careful analysis of many x-ray data sets, the structure of the DBL3x-CSA complex showed electron density (Fig. 1B) for a CSA oligosaccharide located at a positively charged pocket on the top surface of the molecule lying across the two long helices of S3 and at a loop of S2. The electron density for CSA was seen in 2Fobs − Fcalc maps contoured at 1.0σ and in Fobs − Fcalc maps contoured at 2.5σ above the mean density (Fig. 1B). The density was observed in the same area of DBL3x independent of the soaking or cocrystallization with various sizes or concentration of the oligosaccharides. The lack of a well defined and continuous electron density for various CSA oligosaccharides was likely due to multiple modes of binding, partial disordering, conformational fluctuations, or chemical heterogeneity. No electron density was observed without added oligosaccharide, which excluded the possibility that the density was due to buffer or water molecules. The electron density was of a size to fit four to six monosaccharides. A portion of this electron density (1.5σ in 2Fobs − Fcalc, 3.0σ in Fobs − Fcalc) was unambiguously assigned to a sulfate group of CSA as shown in Fig. 1B. The presence of this sulfate group was consistent with a model of a CSA hexasaccharide (Protein Data Bank code 1C4S) positioned at the electron density (15). Another x-ray structure of DBL3x (16), where crystals were soaked with ammonium sulfate or with the mixture of unsaturated CSA and CSC disaccharides obtained by bacterial chondroitinase ABC digestion of bovine trachea CSA, also showed a sulfate bound at the same location. The electron density (Fig. 1B) revealed the location where the CSA binds to DBL3x. The nature of the binding is such that it does not allow the placement of CSA in the electron density in a particular conformation. Such an interaction is not without precedent, it has been seen earlier in defining the binding of a heparin-like ligand to a thrombospondin domain. The density of the bound heparin-like molecule was also not well defined, but sulfates could be positioned in the largest electron density (28, 29). On examination of the residues within 5 Å of the CSA electron density (Fig. 1A) revealed that two S2 residues, Lys1324 and Lys1327, and five S3 residues, Arg1467, Arg1503, Lys1504, Lys1507, and Lys1510 probably have side chain interactions with CSA (15). Residues Lys1324, Lys1327, and Arg1467 appear to be coordinating the identified sulfate group.
Identification of Critical Amino Acid Residues in the CSA-binding Region by Structure-based Mutagenesis and CSA-binding Studies
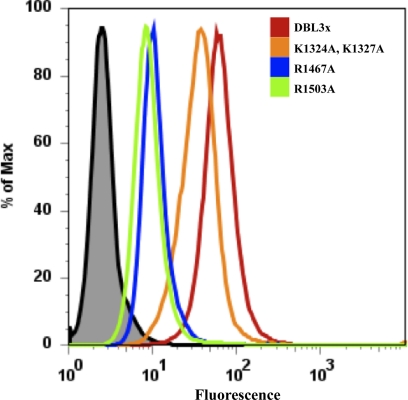
To identify residues that are crucial for CSA binding and that define the minimal CSA-binding region, we generated single and double mutations of the residues of the CSA-binding pocket. CSA binding by DBL3x and its mutants was assessed using flow cytometry. The DBL3x domain binds to CSA expressed on the surface of CHO K1 cells (Fig. 2, red peak). Mutant DBL3x proteins with mutations R1467A (Fig. 2, blue peak) or R1503A (Fig. 2, green peak), located within the S3 region, showed a considerable loss of binding to CSA on the surface of CHO K1 cells. This also highlights the importance of Arg1467, which coordinates the sulfate group in the binding pocket of the DBL3x-CSA complex structure. In contrast, the two mutations located in the loop of S2, K1324A and K1327A (Fig. 2, orange peak), were not critical for binding to CSA. In addition, as we have shown earlier (15), DBL3x bearing the S3 mutations K1507A and K1510A, exhibited a markedly decreased binding to CHO K1 cells. Neither DBL3x nor its mutants bind to CHO 745 cells, which do not express CSA on their surface (results not shown). Another study (30) also has confirmed by mutagenesis that the positively charged patch and sulfate-binding pocket contributes in the CSA binding site of DBL3x. Hence, these results demonstrate that the S3 residues Arg1467, Arg1503, Lys1507, and Lys1510 play a major role in binding CSA, but S2 residues Lys1324 and Lys1327 do not.
FIGURE 2.
The residues of DBL3x important for binding CSA. Wild-type DBL3x (red peak) binds to CSA-expressing CHO K1 cells in a flow cytometry assay. DBL3x proteins bearing the subdomain 3 mutations R1467A and R1503A (blue and green peaks) show significant reductions in binding to CSA on CHO K1 cells. In contrast, the Subdomain 2 double mutation K1324A,K1327A (orange peak) has little effect, binding CSA almost like wild-type DBL3x (red peak). Anti-DBL3x antibody alone (black peak) is shown as a negative control.
Characterization of Recombinant S3 Protein and Its Binding to CSA
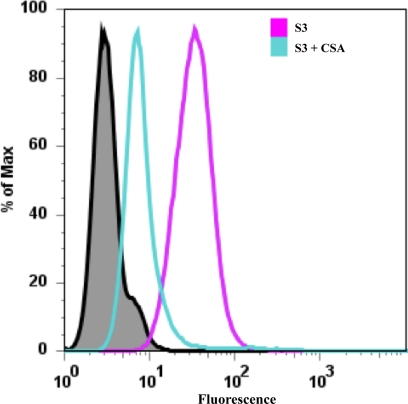
Recombinant S3 protein migrated as a single peak on size exclusion chromatography at a position consistent with its predicted molecular mass of 16,197 Da. SDS gel electrophoresis revealed a single band at the expected molecular mass, and mass spectroscopy confirmed its molecular mass (results not shown). S3 protein produced well shaped crystals upon crystallization but the crystals diffracted only to a resolution limit of 6–8 Å. CSA binding by S3 protein was assayed by flow cytometry with CHO K1 cells using the methods employed for the DBL3x domain. The refolded recombinant S3 protein bound to CSA (Fig. 3, magenta peak) on the surface of CHO K1 cells. This confirmed that recombinant S3 is correctly refolded and functional. Preincubating S3 with soluble bovine tracheal CSA (Fig. 3, cyan peak) inhibited the binding of S3 to the CHO K1 cells. CHO 745 cells did not support the binding of S3 (results not shown). These results demonstrate that S3, expressed as an independently refolding domain from E. coli, binds CSA. S3 binds CSA without participation of other DBL3x subdomains in a manner that is similar to DBL3x CSA binding. From these results, S3 must have a critical role in binding CSA, a conclusion that is consistent with the presence of conserved S3 sequences in all VAR2CSA DBL3x domains from different strains (31).
FIGURE 3.
Subdomain 3 of DBL3x alone can bind CSA. S3 binds to CSA expressed on the surface of CHO-K1 cells (magenta peak). Preincubating S3 with bovine tracheal CSA (S3 + CSA, cyan peak) considerably reduces S3 binding to the cells. Anti-DBL3x antibody alone (black peak) is shown as a negative control.
Binding of CSA Oligosaccharide to S3 and to DBL3x Is Almost Identical and Is Unaffected by NaCl Concentration
STD and 2D TrNOE NMR are ligand-based experiments that rely on magnetization transfer between the protein and the ligand under conditions of fast exchange. Both STD and 2D TrNOE NMR experiments were carried out to probe the binding of the CSA dodecasaccharide fragment, which is composed of six disaccharides ([-4)GlcA(β1–3)GalNAc(β1]6), to the S3 and DBL3x proteins.
An initial 1D 1H NMR spectrum of CSA dodecasaccharide (Fig. 4D) revealed chemical shifts consistent with previously reported values for chondroitin sulfates (26, 32) and indicated that the disaccharide units contain about ∼60% 4-O-sulfated N-acetyl-galactosamine (GalNAc-4S) and ∼40% 6-O-sulfated N-acetyl-galactosamine (GalNAc-6S) residues. The relative percentage of each GalNAc sulfation pattern was quantified by integration of the characteristic well resolved resonances of the H-4 proton of GalNAc-4S at 4.67 ppm (Fig. 4D, GalNAc H4) and the H-4 and H-6 protons of GalNAc-6S at 4.11 ppm and 4.14 ppm (Fig. 4D, each labeled with an x), in the 1D 1H NMR spectrum.
FIGURE 4.
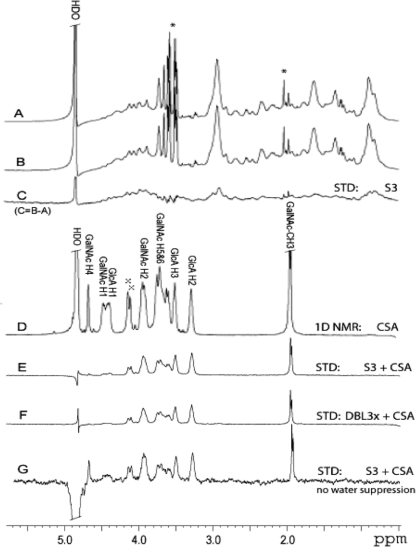
S3 and DBL3x binding to CSA oligosaccharide observed by STD NMR. A–C, STD spectrum (C) of 60 μm S3 protein without CSA oligosaccharide at 20 °C, 1024 scans, obtained by subtraction (C = B − A) of the on- (A) from the off- (B) saturation spectra. Present in solution are some low molecular weight contaminants (asterisk), but note that their signals are completely cancelled in the difference spectrum in C. D, 1D 1H NMR spectrum of CSA oligosaccharide (20 °C, 16 scans, no water suppression). Residual signals from chondroitin-6-O-sulfate are labeled with x. Due to overlap, only selected resonances are labeled. E, STD spectrum of 1:100 molar ratio of S3 and CSA oligosaccharide. F, STD spectrum of 1:100 molar ratio of DBL3x and CSA oligosaccharide at 20 °C using 1024 scans each. G, same as E but the STD data were collected without water suppression at 10 °C to observe the STD signals for the GalNAc-H4, GalNAc-H1, and GlcA-H1 resonances that were obscured by water suppression in E. HDO, residual partially deuterated water.
In an STD NMR experiment, the protein is selectively irradiated, and the magnetization is transferred via intramolecular spin diffusion, which leads to the saturation of the entire protein and of those parts of the bound ligand, which are in direct contact with the protein (23). Since bound ligands exchange with free ligands in solution, the magnetization transfers to free ligands and can be detected in a difference spectrum calculated by subtracting a spectrum with saturation from a reference spectrum. The reference spectrum is collected with off-resonance irradiation that is outside of the proton spectral window, which results in no protein saturation. As the saturation transfers most efficiently to ligand protons that are in direct contact with the protein, the difference spectrum displays signals that originate only from ligand protons that are near protein protons when the ligand was bound. The relative intensities of the signals reflect the relative distances of particular ligand protons from protein protons.
Fig. 4 shows the results from the STD NMR experiment obtained for the S3 protein in the absence of CSA oligosaccharide (Fig. 4, A–C) and STD spectra obtained for CSA oligosaccharide in the presence of either S3 protein (Fig. 4, E and G) or DBL3x protein (Fig. 4F). All STD spectra (Fig. 4, C and E–G) were obtained by subtraction of a corresponding spectrum with selective protein saturation at 7.12 ppm (on-resonance) from one without protein saturation at 50.0 ppm (off-resonance). The difference spectrum for the protein in the absence of CSA oligosaccharide shows complete cancellation of all sharp peaks, which are due to nonbinding, low molecular weight contaminants present in the protein solution (Fig. 4C).
A comparison of the 1D proton NMR spectrum of the CSA oligosaccharide reference in Fig. 4D with the STD spectra of the CSA in the presence of either S3 (Fig. 4E) or DBL3x (Fig. 4F) protein, shows STD signals that originate from CSA oligosaccharide protons at the ligand-protein binding interface. The signal intensities reflect the relative proximity of each proton to the protein surface. As evident by the equal STD effects in Fig. 4, E and F, CSA oligosaccharide binds both S3 and DBL3x in the same manner. The strong STD signal intensities (Fig. 4, E and F) for the CSA oligosaccharide resonance multiplets at 1.9 ppm (GalNAc-CH3), 3.28 ppm (GlcA-H2), 3.51 ppm (GlcA-H3), and 3.9 ppm (GalNAc-H2) indicate that these protons are in closest contact to the surface of both proteins. The rest of the CSA oligosaccharide protons show relatively smaller STD effects likely caused by their larger distance from protein protons. To observe STD enhancements for the GlcA-H1, GalNAc-H1, and GalNAc-H4 resonances that were obscured by the water suppression in Fig. 4, E and F, the STD data for the S3-CSA oligosaccharide complex also was collected without water suppression at 10 °C (Fig. 4G). The data in Fig. 4G show large STD intensities for the GalNAc-H4 protons and relatively smaller ones for the GalNAc-H1 and GlcA-H1 protons.
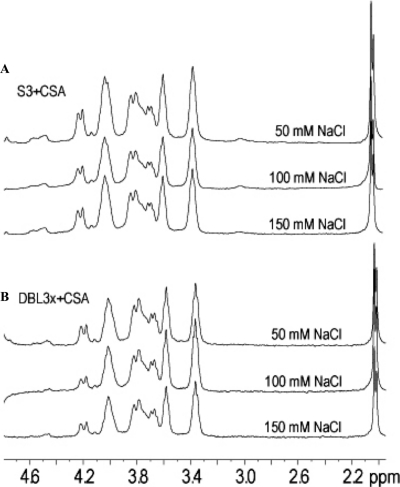
STD NMR data of CSA oligosaccharide in the presence of either S3 or DBL3x protein were collected at several NaCl concentrations (Fig. 5, A and B) to test the contributions of ionic interactions to ligand binding. The STD NMR signals acquired while CSA oligosaccharide is bound to the proteins are independent of the NaCl concentration up to the physiological concentration of 150 mm. These results imply that CSA binding is not to a fortuitous distribution of charged residues but is bound to a specific binding site, which is nearly identical on both proteins.
FIGURE 5.
Varying the NaCl concentration does not affect CSA binding. A and B, STD NMR spectra of S3+CSA oligosaccharide (A) and DBL3x+CSA oligosaccharide (B) at 1:100 molar excess of CSA oligosaccharide in D2O at 20 °C, 1024 scans each, collected in the presence of 50 mm NaCl (as in Fig. 4), 100 mm NaCl, and 150 mm NaCl.
Binding of S3 and DBL3x to CSA Oligosaccharide Observed in Solution by 2D TrNOE NMR
2D TrNOE experiments are routinely used to detect ligand binding to a target protein by examining the changes in the sign and/or intensity of the cross-peak in the ligand spectrum (33). Small ligands with a molecular mass of <1–2 kDa and a fast tumbling rate in solution exhibit positive NOE, which result in negative 2D TrNOE cross peaks, whereas large protein molecules (MW>10,000 Da) with a slower tumbling rate, exhibit negative NOEs, that yield positive 2D TrNOE cross-peaks. Upon binding, the ligand acquires the NOE characteristics of the large molecule and the sign of its 2D TrNOE peaks will change from negative (for the free ligand) through zero to positive. In cases where the free ligand exhibits positive cross-peaks, possibly due to its higher molecular weight or a restricted motion of individual protons, the ligand protons that are in direct contact with the protein are identified based only on the intensity increase of the corresponding positive TrNOE cross-peak. Thus, the ligand protons at the binding interface can be identified either by the change of both the sign and intensity or only intensity of the corresponding TrNOE cross-peaks.
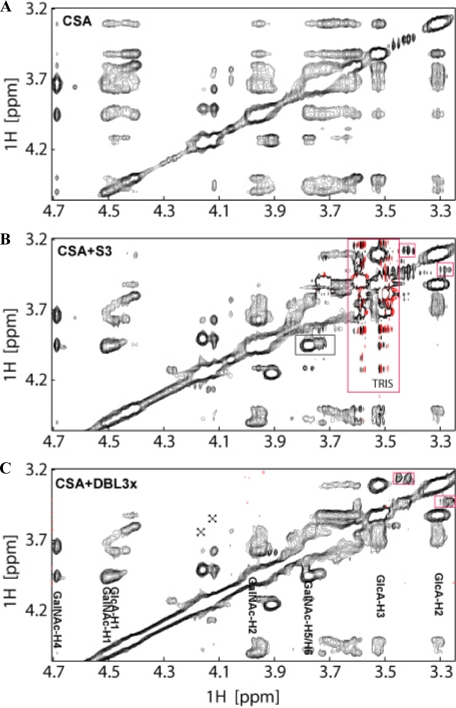
Fig. 6 shows an expanded region of the 2D TrNOE spectra of the free CSA oligosaccharide, (Fig. 6A) and of the CSA oligosaccharide in the presence of either S3 protein (Fig. 6B) or DBL3x protein (Fig. 6C). Consistent with the results from the STD data, large increases in the TrNOE cross-peak intensities, upon binding (Fig. 6, B and C), are observed for the GlcA-H2, GlcA-H3, and GalNAc-H2 protons, which confirm their direct contact with the protein. There are relatively smaller cross peak intensity changes for GalNAc-H5, GalNAc-H6, GlcA-H1, and GalNAc-H1. The non-detectable intensity change for the GalNAc-H4 proton resonances (4.7 ppm) can be attributed to partial saturation due to its proximity to the water resonance.
FIGURE 6.
CSA oligosaccharide binds in the same manner to S3 and to DBL3x as revealed by 2D TrNOE NMR spectra. A, a 2D TrNOE spectrum of free CSA oligosaccharide. B and C, CSA oligosaccharide in the presence of either S3 (B) or DBL3x (C) protein at a molar ratio of 20:1 in D2O at 30 °C. Note that the cross-peaks in B and in C are nearly identical, indicating that the conformations of CSA oligosaccharide that bind to S3 and to DBL3x are almost identical. Residual signals from chondroitin 6-O-sulfate are labeled with x. Cross-peaks due to low molecular weight contaminants are boxed in red. Due to overlap, only selected resonances are labeled.
The NOESY spectrum of the free ligand (Fig. 6A) was recorded with a standard NOESY pulse sequence. For the spectra of the ligand in the presence of protein (S3 or DBL3x) a modified NOESY pulse sequence, which utilized relaxation filter (T1ρ filter) (33–35) was applied to reduce the diagonal and cross-peak intensities of the protein (S3 or DBL3x) itself. Although the application of the filter results in partial decrease of the NOESY signals of the bound ligand (Fig. 6, B and C), the changes in the relative cross-peak intensities between bound and free ligand are readily observed.
Since the integrated volumes of the 2D TrNOE cross-peaks reflect the intra-ligand proton-proton distances, the almost identical relative intensity changes of TrNOE cross-peaks when CSA oligosaccharide is bound to either S3 or DBL3x suggest that changes in the conformation of CSA oligosaccharide upon binding to either protein are essentially the same (compare Fig. 6, A with B, also Fig. 6, A with C). Taken together, the NMR data indicate that CSA oligosaccharide binds specifically to S3 and to DBL3x in almost identical conformations on both proteins. This evidence supports the S3 subdomain as an independent CSA-binding region that binds CSA oligosaccharide almost exactly as does DBL3x.
Female Human Plasma from Malaria-endemic Areas of Mali Recognize DBL3x and S3 by ELISA
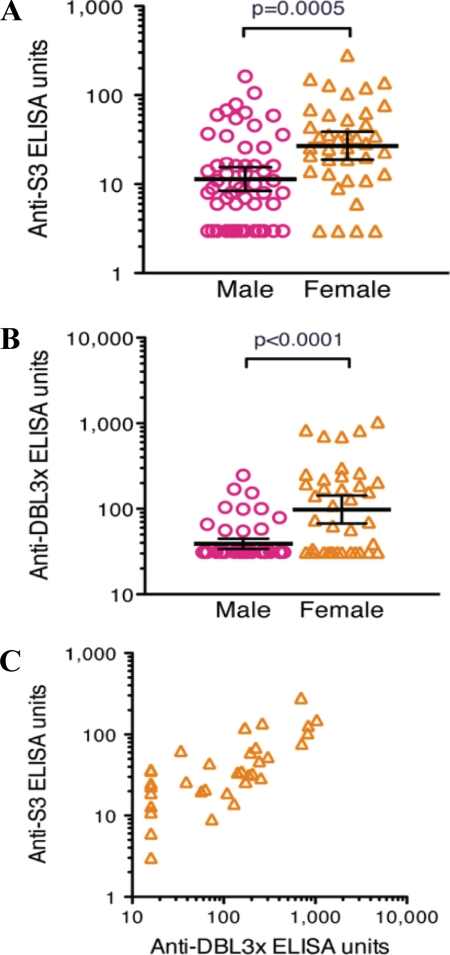
We have tested plasma samples from healthy adults living in malaria-endemic areas of Mali, West Africa, for their ability to recognize both S3 and DBL3x. When tested by ELISA, plasma from females who had prior pregnancy showed greater reactivity toward S3 (Fig. 7A) and DBL3x (Fig. 7B) than did male plasma. The reactivity of female plasma for both antigens is significantly higher compared with male plasma (p < 0.001 by exact Wilcoxon-Mann-Whitney tests). Furthermore, a significant positive correlation is seen between the S3 and DBL3x (Fig. 7C) responses of individual female plasma (Spearman rank test; r = 0.78 [95% confidence interval 0.60 to 0.90], p < 0.0001). These results show that plasma from females living in malaria-endemic areas recognize recombinant S3 and DBL3x, which supports that these recombinant refolded proteins display native antigenic conformations and are exposed on the surface of IEs which justifies their future study as potential vaccine candidates for PAM. Although in recent articles (36, 37), it has been reported that antibodies against full-length VAR2CSA inhibits binding of IE to chondroitin sulfate proteoglycans, there is evidence that antibodies raised against individual domains of VAR2CSA also show inhibition (38, 39). Thus, it is important to understand and study independent DBL domains and their minimal CSA-binding subdomain in detail from the therapeutic and vaccine point of view. Further structural and immunological studies are needed on individual domains as well as on full length VAR2CSA before we reach a conclusion about selecting the best vaccine target.
FIGURE 7.
Female human plasma from malaria-endemic areas of Mali recognizes S3 and DBL3x. Plasma collected from Malian women show significantly higher reactivity to S3 (A) and DBL3x (B) compared with plasma from Malian men (for both antigens, p < 0.001 by exact Wilcoxon-Mann-Whitney tests). C, positive correlation between the reactivity of Malian female plasma to S3 and DBL3x (r = 0.78, by Spearman rank test, p < 0.0001).
In conclusion, we have identified S3 as an independent stable minimal CSA-binding region of DBL3X that is highly conserved among different CSA-binding DBL domains. Based on previous knowledge of the crystal structure of DBL3x and the recent structure-based mutagenesis results, there is compelling evidence that the residues that contribute most substantially to CSA binding are in subdomain 3. S3 is held together by a high number of disulfide bonds, forming an independent minimal subunit of DBL3x. This observation suggests an important biological role for S3, as the disulfide bonds help to preserve the rigidity of the binding pocket. It has also been shown that S3 undergoes less diversifying selection as compared with other parts of DBL3x (31). Such conserved structures are attractive targets for vaccine development.
Acknowledgments
We thank Phuc Nguyen for preparing mutant plasmids, Michael Mohan for initial expression and refolding trials of the mutants, Satish Madala for help with flow cytometry, and Mark Garfield for mass spectrometry. We are grateful to Dr. Mahamadou Diakite and investigators in the Malaria Research and Training Center at the University of Bamako, Bamako, Mali for collection of adult plasma samples. We are also particularly grateful to these volunteers for donation of plasma.
This work was supported, in whole or in part, by Grant AI45086 from NIAID, National Institutes of Health and the Division of Intramural Research of the NIAID, National Institutes of Health.
- PAM
- pregnancy-associated malaria
- CSA
- chondroitin sulfate A
- DBL
- Duffy binding-like
- S1
- subdomain 1 of DBL3x
- S2
- subdomain 2 of DBL3x
- S3
- subdomain 3 of DBL3x
- PfEMP1
- Plasmodium falciparum erythrocyte membrane protein 1
- IE
- infected erythrocyte
- GlcA
- glucuronic acid
- GalNAc-4S
- 4-O-sulfated N-acetylgalactosamine
- FACS
- fluorescence-activated cell sorter
- GalNAc-6S
- 6-O-sulfated N-acetylgalactosamine
- CHO
- Chinese hamster ovary
- STD
- saturation transfer difference
- ppm
- parts per million
- 2D TrNOE
- two-dimensional transferred nuclear overhauser effect
- ELISA
- enzyme-linked immunosorbent assay
- NOESY
- nuclear overhauser effect spectroscopy.
REFERENCES
- 1.Brabin B. J., Romagosa C., Abdelgalil S., Menéndez C., Verhoeff F. H., McGready R., Fletcher K. A., Owens S., D'Alessandro U., Nosten F., Fischer P. R., Ordi J. (2004) Placenta 25, 359–378 [DOI] [PubMed] [Google Scholar]
- 2.Salanti A., Staalsoe T., Lavstsen T., Jensen A. T., Sowa M. P., Arnot D. E., Hviid L., Theander T. G. (2003) Mol. Microbiol. 49, 179–191 [DOI] [PubMed] [Google Scholar]
- 3.Trimnell A. R., Kraemer S. M., Mukherjee S., Phippard D. J., Janes J. H., Flamoe E., Su X. Z., Awadalla P., Smith J. D. (2006) Mol. Biochem. Parasitol. 148, 169–180 [DOI] [PubMed] [Google Scholar]
- 4.Rowe J. A., Kyes S. A., Rogerson S. J., Babiker H. A., Raza A. (2002) J. Infect. Dis. 185, 1207–1211 [DOI] [PubMed] [Google Scholar]
- 5.Fried M., Hixson K. K., Anderson L., Ogata Y., Mutabingwa T. K., Duffy P. E. (2007) Mol. Biochem. Parasitol. 155, 57–65 [DOI] [PubMed] [Google Scholar]
- 6.Viebig N. K., Gamain B., Scheidig C., Lépolard C., Przyborski J., Lanzer M., Gysin J., Scherf A. (2005) EMBO Rep. 6, 775–781 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Duffy M. F., Maier A. G., Byrne T. J., Marty A. J., Elliott S. R., O'Neill M. T., Payne P. D., Rogerson S. J., Cowman A. F., Crabb B. S., Brown G. V. (2006) Mol. Biochem. Parasitol. 148, 117–124 [DOI] [PubMed] [Google Scholar]
- 8.Fried M., Duffy P. E. (1998) J. Mol. Med. 76, 162–171 [DOI] [PubMed] [Google Scholar]
- 9.Maubert B., Fievet N., Tami G., Cot M., Boudin C., Deloron P. (1999) Infect. Immun. 67, 5367–5371 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Beeson J. G., Mann E. J., Byrne T. J., Caragounis A., Elliott S. R., Brown G. V., Rogerson S. J. (2006) J. Infect. Dis. 193, 721–730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ricke C. H., Staalsoe T., Koram K., Akanmori B. D., Riley E. M., Theander T. G., Hviid L. (2000) J. Immunol. 165, 3309–3316 [DOI] [PubMed] [Google Scholar]
- 12.Hviid L. (2010) Hum. Vaccin. 6, 84–89 [DOI] [PubMed] [Google Scholar]
- 13.Gamain B., Trimnell A. R., Scheidig C., Scherf A., Miller L. H., Smith J. D. (2005) J. Infect Dis. 191, 1010–1013 [DOI] [PubMed] [Google Scholar]
- 14.Avril M., Gamain B., Lépolard C., Viaud N., Scherf A., Gysin J. (2006) Microbes Infect. 8, 2863–2871 [DOI] [PubMed] [Google Scholar]
- 15.Singh K., Gittis A. G., Nguyen P., Gowda D. C., Miller L. H., Garboczi D. N. (2008) Nat. Struct. Mol. Biol. 15, 932–938 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Higgins M. K. (2008) J. Biol. Chem. 283, 21842–21846 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Achur R. N., Valiyaveettil M., Gowda D. C. (2003) J. Biol. Chem. 278, 11705–11713 [DOI] [PubMed] [Google Scholar]
- 18.Alkhalil A., Achur R. N., Valiyaveettil M., Ockenhouse C. F., Gowda D. C. (2000) J. Biol. Chem. 275, 40357–40364 [DOI] [PubMed] [Google Scholar]
- 19.Fried M., Lauder R. M., Duffy P. E. (2000) Exp. Parasitol. 95, 75–78 [DOI] [PubMed] [Google Scholar]
- 20.Chai W., Beeson J. G., Lawson A. M. (2002) J. Biol. Chem. 277, 22438–22446 [DOI] [PubMed] [Google Scholar]
- 21.Sodeoka M., Larson C. J., Chen L., LeClair K. P., Verdine G. L. (1993) Bioorg. Med. Chem. Lett. 3, 1089–1094 [Google Scholar]
- 22.Gamain B., Miller L. H., Baruch D. I. (2001) Proc. Natl. Acad. Sci. U.S.A. 98, 2664–2669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mayer M., Meyer B. (2001) J. Am. Chem. Soc. 123, 6108–6117 [DOI] [PubMed] [Google Scholar]
- 24.Clore G. M., Gronenborn A. M., Mitchinson C., Green N. M. (1982) Eur. J. Biochem. 128, 113–117 [PubMed] [Google Scholar]
- 25.Delaglio F., Grzesiek S., Vuister G. W., Zhu G., Pfeifer J., Bax A. (1995) J. Biomol. NMR 6, 277–293 [DOI] [PubMed] [Google Scholar]
- 26.Toida T., Toyoda H., Imanari T. (1993) Analytical Sciences 9, 53–58 [Google Scholar]
- 27.Miura K., Orcutt A. C., Muratova O. V., Miller L. H., Saul A., Long C. A. (2008) Vaccine 26, 193–200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tan K., Duquette M., Liu J. H., Zhang R., Joachimiak A., Wang J. H., Lawler J. (2006) Structure 14, 33–42 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Tan K., Duquette M., Liu J. H., Shanmugasundaram K., Joachimiak A., Gallagher J. T., Rigby A. C., Wang J. H., Lawler J. (2008) J. Biol. Chem. 283, 3932–3941 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Khunrae P., Philip J. M., Bull D. R., Higgins M. K. (2009) J. Mol. Biol. 393, 202–213 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dahlbäck M., Rask T. S., Andersen P. H., Nielsen M. A., Ndam N. T., Resende M., Turner L., Deloron P., Hviid L., Lund O., Pedersen A. G., Theander T. G., Salanti A. (2006) PLoS Pathog. 2, e124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Blanchard V., Chevalier F., Imberty A., Leeflang B. R., Basappa, Sugahara K., Kamerling J. P. (2007) Biochemistry 46, 1167–1175 [DOI] [PubMed] [Google Scholar]
- 33.Glaudemans C. P., Lerner L., Daves G. D., Jr., Kovác P., Venable R., Bax A. (1990) Biochemistry 29, 10906–10911 [DOI] [PubMed] [Google Scholar]
- 34.Scherf T., Anglister J. (1993) Biophys. J. 64, 754–761 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Cosman M., Krishnan V. V., Balhorn R. (2005) Methods Mol. Biol. 300, 141–163 [DOI] [PubMed] [Google Scholar]
- 36.Khunrae P., Dahlbäck M., Nielsen M. A., Andersen G., Ditlev S. B., Resende M., Pinto V. V., Theander T. G., Higgins M. K., Salanti A. (2010) J. Mol. Biol. 397, 826–834 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Srivastava A., Gangnard S., Round A., Dechavanne S., Juillerat A., Raynal B., Faure G., Baron B., Ramboarina S., Singh S. K., Belrhali H., England P., Lewit-Bentley A., Scherf A., Bentley G. A., Gamain B. (2010) Proc. Natl. Acad. Sci. U.S.A. 107, 4884–4889 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Salanti A., Resende M., Ditlev S. B., Pinto V. V., Dahlbäck M., Andersen G., Manczak T., Theander T. G., Nielsen M. A. (2010) Malar. J. 9, 11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Nielsen M. A., Pinto V. V., Resende M., Dahlbäck M., Ditlev S. B., Theander T. G., Salanti A. (2009) Infect. Immun. 77, 2482–2487 [DOI] [PMC free article] [PubMed] [Google Scholar]