Abstract
Cylindrospermopsin is a cytotoxin produced by Cylindrospermopsis raciborskii and other cyanobacteria that has been implicated in human intoxications. We report here the complete sequence of the gene cluster responsible for the biosynthesis of this toxin in Oscillatoria sp. strain PCC 6506. This cluster of genes was found to be homologous with that of C. raciborskii but with a different gene organization. Using an enzyme-linked immunosorbent assay and an optimized liquid chromatography analytical method coupled to tandem mass spectrometry, we detected 7-epicylindrospermopsin, cylindrospermopsin, and 7-deoxycylindrospermopsin in the culture medium of axenic Oscillatoria PCC 6506 at the following relative concentrations: 68.6%, 30.2%, and 1.2%, respectively. We measured the intracellular and extracellular concentrations, per mg of dried cells of Oscillatoria PCC 6506, of 7-epicylindrospermopsin (0.18 μg/mg and 0.29 μg/mg, respectively) and cylindrospermopsin (0.10 μg/mg and 0.11 μg/mg, respectively). We showed that these two toxins accumulated in the culture medium of Oscillatoria PCC 6506 but that the ratio (2.5 ± 0.3) was constant with 7-epicylindrospermopsin being the major metabolite. We also determined the concentrations of these toxins in culture media of other Oscillatoria strains, PCC 6407, PCC 6602, PCC 7926, and PCC 10702, and found that, except for PCC 6602, they all produced 7-epicylindrospermopsin and cylindrospermopsin, with the former being the major toxin, except for PCC 7926, which produced very little 7-epicylindrospermopsin. All the cylindrospermopsin producers studied gave a PCR product using specific primers for the amplification of the cyrJ gene from genomic DNA.
Cyanobacteria are known to produce a wide range of secondary metabolites (19), some of which are toxic to humans and animals, and cyanobacterial blooms are frequently associated with human- or animal-poisoning events. Several classes of cyanobacterial toxins have so far been identified, including hepatotoxins, neurotoxins, cytotoxins, and irritants (42). Cases of animal intoxication due to cyanobacterial toxin exposure are regularly reported in different places around the world, and it is now recognized that the release of cyanobacterial toxins in water reservoirs and water supplies has major implications for public health and for the environment (8). For instance, anatoxin-a and homoanatoxin-a, two cyanobacterial neurotoxins, provoke the rapid death of animals by acute asphyxia when ingested, because these alkaloids are potent agonists of the nicotinic acetylcholine receptor (6, 11, 43, 44).
In 1979, a significant human-poisoning incident was reported on Palm Island in Australia (14, 16). Afterwards, it was shown that the compound responsible for that poisoning was cylindrospermopsin (CYN) (Fig. 1) and that this cytotoxic molecule was produced by a cyanobacterium, Cylindrospermopsis raciborskii (28). Since then, it was shown that this molecule is also produced by cyanobacteria belonging to other genera such as Anabaena (38), Aphanizomenon (1, 29), Lyngbya (35), Raphidiopsis (20), and Umezakia (15).
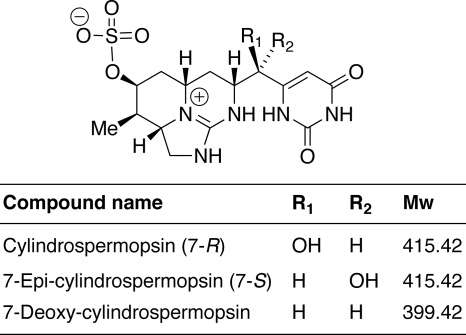
FIG. 1.
Structures of the cyanotoxin cylindrospermopsin (CYN) and of its natural analogs.
Two close analogs, 7-deoxycylindrospermopsin (7-deoxy-CYN) and 7-epicylindrospermopsin (7-epi-CYN) (Fig. 1), were later detected in cultures of CYN-producing cyanobacteria. 7-Deoxy-CYN was found in C. raciborskii (21, 27) as a minor metabolite (10% of the CYN concentration) and in Raphidiopsis curvata (20) and Lyngbya wollei (35) as the major metabolite. 7-Epi-CYN has so far been detected only in Aphanizomenon ovalisporum as a minor metabolite (2). These three closely related molecules were shown previously to be toxic using in vivo assays, and they were shown to inhibit protein synthesis with similar potencies (22, 32, 33).
Several analytical techniques have been developed to assay concentrations of CYN in cultures or in the environment: an enzyme-linked immunosorbent assay (ELISA) and assays based on liquid chromatography coupled either to a UV detector (40) or to a mass spectrometer (3, 12, 13).
The biosynthesis of CYN was studied by using feeding experiments (5), and a biosynthetic scheme was later proposed based on the bioinformatic analysis of the gene cluster responsible for the biosynthesis of CYN, recently identified in C. raciborskii (25). Interestingly, 7-deoxy-CYN was proposed to be a precursor of CYN, with the hydroxylation step likely being catalyzed by CyrI, a proline hydroxylase homolog. This pathway was thus compatible with the presence of CYN and 7-deoxy-CYN in CYN producers.
We have been interested for some time in characterizing cyanobacteria producing anatoxin-a and homoanatoxin-a (6, 7), and we have obtained a draft sequence of the genome of Oscillatoria sp. strain PCC 6506, a benthic cyanobacterium that produces these deadly neurotoxins. We have recently described the cluster of genes responsible for the biosynthesis of these two alkaloids (24) and proposed an original biosynthetic scheme that was substantiated by in vitro experiments (23). Surprisingly, in the genome of this neurotoxic strain, we found the genes responsible for the biosynthesis of CYN, which were described previously by Mihali et al. for C. raciborskii (25). We report here the complete characterization of this new cyr cluster. Furthermore, we show that Oscillatoria PCC 6506 and related Oscillatoria benthic strains, one of which was isolated in the Tarn River in France in 2006, produce CYN and 7-epi-CYN.
MATERIALS AND METHODS
Cyanobacterial strains and culture conditions.
Axenic Oscillatoria strains PCC 6407 (ATCC 27906), PCC 6506 (ATCC 29081 or UTEX 1547), PCC 6602 (ATCC 27935), and PCC 7926 were obtained from the Pasteur Culture Collection (PCC) of cyanobacteria (17). Oscillatoria strain PCC 10702 is not in the PCC catalog but is available upon request (6). All Oscillatoria strains were cultivated in BG11 medium under constant light illumination at 20°C as previously described (6). The purity of the cultures was routinely checked (30).
Genomic DNA preparation.
Genomic DNA was extracted from the cyanobacterial cells as previously described (26) by using a lysozyme-sodium dodecyl sulfate-proteinase K lysis protocol followed by phenol-chloroform extraction. The soluble DNA was then precipitated by ethanol, washed with 70% aqueous ethanol, and dissolved in 10 mM Tris-EDTA buffer (pH 8.0) (34). The purified DNA was stored at 4°C.
Sequencing and annotation of the genome of Oscillatoria PCC 6506.
High-quality genomic DNA from Oscillatoria PCC 6506 was prepared from fresh cells, grown as described above (500 ml), and harvested by centrifugation (4,000 × g for 20 min at 4°C), yielding 1.3 g wet cells. The cells were immediately treated for the isolation of genomic DNA as previously described (26). Sequencing of the genome is under way at the GATC Company (Konstanz, Germany) using 454 pyrosequencing technology, and a draft genome sequence has already been obtained. This genome has been automatically annotated by using the MaGe system (41; A. Méjean, O. Ploux, and C. Médigues, unpublished data). Expert annotation was performed by using Basic Local Alignment Search Tool (BLAST) software implemented with the National Center for Biotechnology Information (NCBI) Web server and by using the GenBank database. Specific sequence alignments were performed by using ClustalX software (39).
PCR amplifications and sequencing.
PCRs were performed with an Eppendorf Mastercycler (Hamburg, Germany). Oligonucleotides were purchased from Eurogentec (Angers, France). Taq polymerase, buffer, and deoxynucleoside triphosphates (dNTPs) were obtained from Promega (Paris, France), and LongAmp polymerase was obtained from New England Biolabs (Ozyme, France). Standard PCR conditions were used for all amplifications, as recommended by the manufacturer. Amplified DNA fragments were analyzed by 1% agarose gel electrophoresis and then purified (PCR Prep; Promega) and sequenced (GATC Company). The oligonucleotides used for the amplifications were as follows: cyrB1 (5′-TCAAAATGTGGTTGAATCGGCTCTGTGGGG) and cyrB2 (5′-TCGTCCGACCAGCACAAGGTGCCTGGCCCC) for cyrB amplification, cyrJF (5′-TTCTCTCCTTTCCCTATCTCTTTATC) and cyrJR (5′-TGCTACGGTGCTGTACCAAGGGGC) for cyrJ amplification, and 16S1 (5′-AGAGTTTGATCCTGGCTCAG) and 16S2 (5′-GGTCTCCCTAAAAGGAGGTG) for 16S rRNA gene amplification (9, 18). Amplifications using primers 16S1 and 16S2 were performed as positive controls for the amplification of genomic DNA.
CYN, 7-epi-CYN, and 7-deoxy-CYN standards.
CYN, 7-epi-CYN, and 7-deoxy-CYN standards were purchased from Novakits (Nantes, France). However, because the 7-epi-CYN and 7-deoxy-CYN standards were obtained in minute amounts, we calibrated their standard solutions with that of CYN by high-performance liquid chromatography (HPLC) coupled to a UV detector (dual wavelength; Varian, Les Ulis, France). The chromatographic conditions are described below. We assumed that these two analogs had the same molar extinction coefficients as CYN at 262 nm (2). This seems reasonable because the uracyl ring chromophore is unchanged in these compounds. Thus, all the concentrations reported here are concentrations relative to CYN rather than absolute concentrations.
Determination of CYN concentrations by ELISA.
The CYN concentration was determined by using the CYN ELISA kit (Novakits) as described by the manufacturer. The cyanobacteria were cultivated as described above for 30 to 80 days, and 1-ml aliquots were withdrawn for analysis at different time points. The samples were centrifuged (14,000 × g for 5 min), and the supernatant was then collected, filtered (0.2 μm, polyethersulfone; Nalgen), and diluted in water for analysis. Several dilutions were tested, and each dilution was assayed in triplicate. Standard curves were plotted, as described by the manufacturer, by using authentic CYN (Novakits). The relative responses of 7-epi-CYN and 7-deoxy-CYN over that of CYN, using this ELISA kit, were determined by using pure standards (Novakits).
Determination of concentrations of CYN and analogs by liquid chromatography coupled to tandem mass spectrometry (LC-MS2).
Chromatographic analyses were performed with an HPLC Varian system made up of a binary high-pressure pump for mobile-phase delivery (212 LC) and a Prostar 410 autosampler (Varian). The reversed-phase column was a C18 column (Atlantis, 3.5 μm, 2.1 by 150 mm; Waters, Saint-Quentin en Yvelines, France) for analyte separation. The mobile phase consisted of a gradient elution of solution B (methanol-water [90/10, vol/vol], both acidified with 5 mM ammonium acetate) in solution A (methanol-water [1/99, vol/vol]) (0% solution B for 0 to 3 min, linear gradient of 0 to 50% solution B during 3 to 13 min, and 50% solution B held for 2 min). The flow rate was set at 200 μl/min. A triple-quadrupole mass spectrometer (Varian) was coupled to the HPLC system with an electrospray ionization (ESI) source. Data were acquired in the positive-ion mode in the multiple-reaction monitoring mode (MRM). The compounds were detected following the transition 416.3→194.2 for CYN and 7-epi-CYN and 400.3→194.2 for 7-deoxy-CYN (10), with a capillary voltage of 80 V and with a collision energy of 25 eV. The quantification of compounds contained in the various samples was made by injecting nonextracted standard solutions to plot a calibration curve. Limits of quantification were estimated by measuring the signal-to-noise ratio with Varian Prostar software. The limit of quantification was less than 3.5 ng/ml with an injection volume of 10 μl.
A sample of an Oscillatoria PCC 6506 culture (7 weeks old; 2 ml) was centrifuged, and the supernatant was analyzed for CYN and 7-epi-CYN by LC-MS2 as described above. The corresponding cell pellet was then lyophilized and weighed to determine the metabolite concentration-over-biomass ratio. The intracellular concentrations of CYN and 7-epi-CYN were determined on a dry pellet of Oscillatoria PCC 6506 cells (10 mg suspended in 1 ml of water) after lysis of the cells by sonication (12 s at 400 W with a Misonix sonicator [3-mm probe] at room temperature) in water. After centrifugation (14,000 × g for 5 min), the supernatant was recovered, filtered (0.2 μm, polyethersulfone; Nalgen), and analyzed by LC-MS2 as described above.
Nucleotide sequence accession numbers.
The nucleotide sequences described in this work have been deposited in the GenBank database under the following accession numbers: FJ418586 for the Oscillatoria sp. PCC 6506 cylindrospermopsin biosynthetic gene cluster, complete sequence; HM008939 for the Oscillatoria sp. PCC 6506 adenylylsulfate kinase gene, cyrN, complete sequence; and HM008940 for the Oscillatoria sp. PCC 6506 abrB1 gene, complete sequence.
RESULTS AND DISCUSSION
Oscillatoria PCC 6506 cyr gene cluster.
We have obtained a draft sequence of the genome of Oscillatoria PCC 6506 (A. Méjean, S. Mann, R. Mazmouz, O. Ploux, and C. Médigue, unpublished data). This genome was automatically annotated by using the MaGe system (41), and in this draft sequence, we found two contigs that contained genes homologous to the cyr genes identified previously by Mihali et al. for C. raciborskii (25). The cyrB gene was in fact split into two parts in our draft sequence, because it contains a 150-bp-long fragment that is GC rich and contains repeated sequences (from position 7384 to position 7536 in the Oscillatoria PCC 6506 cyrB sequence). The repetition of the CCCCCGCT motif nine times resulted in a low accuracy of the sequencing using 454 technology. Attempts to amplify the region around this fragment using classical PCR conditions were unsuccessful, and we had to use specialized polymerase (LongAmp) to obtain a clean PCR product. The sequence of this product was then obtained by using optimized conditions for GC-rich sequences. The full cyr cluster sequence was then analyzed and annotated, and the results are summarized in Fig. 2 and Table 1. Figure 2B shows the Oscillatoria cyr gene cluster, which is flanked by a zinc finger SWIM domain protein gene and a putative rifampin ADP-ribosyltransferase gene at the 5′ end and 3′ end, respectively. These genes probably do not belong to the cyr gene cluster and are thus not represented here.
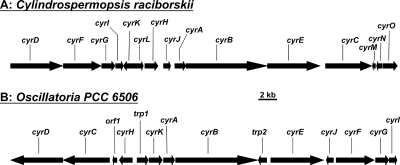
FIG. 2.
Comparison of the gene organizations of the cyr gene clusters from C. raciborskii (A) (25) and from Oscillatoria PCC 6506 (B) (this work). (Panel A was adapted from reference 25.)
TABLE 1.
Annotation of the genes of the cyr cluster from Oscillatoria PCC 6506
| Gene | Size (aa)b | BLAST similarity matcha | GenBank accession no. | Identity (%) |
|---|---|---|---|---|
| cyrD | 1,873 | Polyketide synthase (CyrD) (Cylindrospermopsis raciborskii AWT205) | ABX60152.1 | 83 |
| cyrC | 1,672 | Polyketide synthase (CyrC) (Cylindrospermopsis raciborskii AWT205) | ABX60163.1 | 86 |
| orf1 | 152 | RimK domain protein ATP-grasp (Herpetosiphon aurantiacus ATCC 23779) | ABX07651.1 | 41 |
| cyrH | 473 | Aminohydrolase (CyrH) (Cylindrospermopsis raciborskii AWT205) | ABX60158.1 | 91 |
| trp1 | 404 | Transposase, IS605 OrfB family (Cyanothece sp. strain PCC 7822) | EDX96268.1 | 53 |
| cyrK | 479 | Multidrug exporter MatE (CyrK) (Cylindrospermopsis raciborskii AWT205) | ABX60156.1 | 90 |
| cyrA | 398 | Amidinotransferase (CyrA) (Cylindrospermopsis raciborskii AWT205) | ABX60160.1 | 86 |
| cyrB | 2,957 | Mixed NRPSc/polyketide synthase (CyrB) (Cylindrospermopsis raciborskii AWT205) | ABX60161.1 | 79 |
| trp2 | 299 | Transposase, IS605 family (Synechococcus sp. strain PCC 7335) | EDX82555.1 | 56 |
| cyrE | 1,893 | Polyketide synthase (CyrE) (Cylindrospermopsis raciborskii AWT205) | ABX60162.1 | 84 |
| cyrJ | 261 | Putative sulfotransferase (CyrJ) (Cylindrospermopsis raciborskii AWT205) | ABX60159.1 | 85 |
| cyrF | 1,368 | Polyketide synthase (CyrF) (Cylindrospermopsis raciborskii AWT205) | ABX60153.1 | 84 |
| cyrG | 478 | Putative uracil ring formation (CyrG) (Cylindrospermopsis raciborskii AWT205) | ABX60154.1 | 88 |
| cyrI | 277 | Proline dioxygenase-like protein (CyrI) (Cylindrospermopsis raciborskii AWT205) | ABX60155.1 | 84 |
| cyrN | 175 | Adenylylsulfate kinase (Trichodesmium erythraeum IMS101) | ABG51178.1 | 74 |
The BLAST searches were performed by using the online tools available at the National Center for Biotechnology Information website and using the GenBank database (January 2010).
aa, amino acids.
NRPS, nonribosomal peptide synthase.
It can be seen in Table 1 that the annotation of the genes was unambiguous due to the high sequence identity (from 74% to 90%) with the corresponding C. raciborskii gene products. The Oscillatoria cyrA, cyrB, and cyrC gene products also showed high sequence identity with the A. ovalisporum aoaA, aoaB (partial), and aoaC gene products (36): 85%, 79%, and 85%, respectively. However, the comparison of the gene arrangements in the two cyr clusters so far identified, as shown in Fig. 2, indicated a substantial shuffling of the cyr genes in these two organisms. Interestingly, the three aoa genes that are likely cyr genes implicated in the biosynthesis of CYN in A. ovalisporum (36) show another arrangement with the following order: aoaB, aoaA, and aoaC (GenBank accession number AF395828.1). It is thus very likely that these cyr (aoa) clusters are evolutionarily related but that they substantially diverged from a common ancestor. The repeated sequence that we found in the Oscillatoria cyrB gene (see above) is translated in a rich proline protein sequence, with repeated PPLP motifs, that is not found in the homologous CyrB protein from C. raciborskii. This short sequence is in fact preceding the ketoreductase domain of the CyrB polyketide synthase, and it might serve as a linker between this domain and the preceding methyltransferase domain.
Besides the 11 cyr genes cyrA to cyrK, we found three other genes in the Oscillatoria cyr cluster that were not found in the C. raciborskii cyr cluster. The orf1 gene codes for a putative ATP-grasp protein for which no obvious function related to CYN biosynthesis was found. The two other genes, trp1 and trp2, code for transposases that are not related to the CyrL and CyrM (a partial N-terminal fragment of CyrL) transposases found in the cyr cluster from C. raciborskii. The sequence alignment of the two transposases from Oscillatoria PCC 6506 with CyrL from C. raciborskii did not reveal any significant identity. The cyrO gene that was annotated as a regulator by Mihali et al. (25) was found neither in our cyr cluster nor in the Oscillatoria PCC 6506 genome. Of course, this genome is not complete, and this gene might be present but unsequenced yet. The cyrN gene was not found in the Oscillatoria cyr cluster but was found on another contig of our draft sequence of the Oscillatoria PCC 6506 genome. This cyrN gene product belongs to the adenylylsulfate kinase family and presents 74% sequence identity with the enzyme from Trichodesmium erythraeum IMS101. It presented 52% sequence identity and 68% sequence similarity with the CyrN protein from C. raciborskii. However, because this gene was not found in the Oscillatoria cyr cluster and because the adenylylsulfate kinase is the only enzyme known to produce 3′-phosphoadenylylsulfate, the substrate of several sulfotransferases, we think that this gene should not be named as a cyr gene. In the C. raciborskii cyr cluster this gene is found at the 5′ end of the cluster (Fig. 2A) just preceding the cyrO gene, which was not found in our cluster, and thus, these two genes might not be part of the cyr cluster. Nevertheless, we used the nomenclature proposed previously by Mihali et al. (25) for this adenylylsulfate kinase gene to avoid confusion.
Shalev-Malul et al. (37) described in an elegant report that the expression of the aoaA and aoaC genes in A. ovalisporum was very likely regulated by a DNA binding protein of the AbrB family. Those authors identified this transcriptional regulator and showed that it binds to the DNA sequence found between the divergent aoaA and aoaC genes. In a BLAST search using this regulator sequence (GenBank accession number ABF29864.1) against the Oscillatoria PCC 6506 genome, we identified a homologous regulator, Oscillatoria PCC 6506 AbrB1 (79% sequence identity with the regulator from A. ovalisporum). A similar regulator was found in the genome of C. raciborskii (GenBank accession number EFA71458.1) (89% sequence identity with the regulator from A. ovalisporum and 77% sequence identity with the regulator from Oscillatoria PCC 6506). Interestingly, the alignment of the regions upstream of the ATG start codon of the aoaC (cyrC) genes from A. ovalisporum, C. raciborskii, and Oscillatoria PCC 6506 showed significant regions with high identities (Fig. 3), suggesting a common transcriptional regulation of the expression of these genes. The conserved regions might be the regulator-specific binding DNA sequences.
FIG. 3.
Alignment of the DNA sequence upstream of the aoaC and cyrC genes. The alignment was performed by using ClustalX software (39). The ATG start codon is underlined and in boldface type. A star indicates identity. The A. ovalisporum aoaC sequence was obtained from GenBank (accession number AF395828.1); the C. raciborskii cyrC sequence was obtained from GenBank (accession number EU140798.1); the Oscillatoria PCC 6506 cyrC sequence was from this work.
Toxin analysis by ELISA and LC-MS2.
Using a classical commercially available ELISA kit, we detected and determined the amount of CYN excreted by growing Oscillatoria PCC 6506 cells (Table 2). We obtained an average value of 1.12 ± 0.03 μg/ml of excreted CYN in the culture medium after 11 weeks of culture. In our hands, the ELISA determination was not very accurate (as shown in Table 2). This is probably due to matrix effects, as noted previously by others (3). Nevertheless, this extracellular CYN concentration is consistent with previously reported CYN concentrations determined by using other producing cyanobacteria or field samples (3, 4, 12, 31). Using pure 7-epi-CYN and 7-deoxy-CYN standards, we obtained the following relative responses using this ELISA kit: 1.0 for CYN, 0.5 for 7-epi-CYN, and 0.4 for 7-deoxy-CYN. Thus, this assay is not specific for CYN, and we decided to use an optimized LC-MS2 analytical method to determine the concentration of each of these compounds in our samples.
TABLE 2.
Concentrations of excreted CYN and 7-epi-CYN in different Oscillatoria strains as determined by ELISA and LC-MS2e
| Oscillatoria strain | Origin (yr of sampling) | Toxin concn (μg/ml) determined by:a |
Ratio of concn determined by ELISA/concn determined by LC-MS2 | cyrJ PCR amplificationb | Anatoxin-a producerc | ||
|---|---|---|---|---|---|---|---|
| ELISA for CYN (mean ± SD) | LC-MS2d |
||||||
| 7-epi-CYN | CYN | ||||||
| PCC 6407 | Freshwater, California (1964) | 0.64 ± 0.42 | 0.42 | 0.13 | 3.2 | + | + |
| PCC 6506 | Unknown | 1.12 ± 0.03 | 0.69 | 0.28 | 2.5 | + | + |
| PCC 6602 | Freshwater, California (1966) | ND | ND | ND | ND | − | − |
| PCC 7926f | Brackish water, Gulf of Finland, Helsinki, Finland (1978) | 0.39 ± 0.16 | 0.01 | 0.72 | 0.01 | + | − |
| PCC 10702 | Tarn river, Les Vignes, France (2006) | 0.83 ± 0.20 | 0.50 | 0.17 | 2.9 | + | + |
Measured in the culture medium after 11 weeks.
Presence (+) or absence (−) of a PCR product using specific primers to amplify an internal part of the cyrJ gene from genomic DNA of the strain.
Data previously reported (6).
The standard deviation was estimated to be ±10% for the LC-MS2 analyses.
ND, not detected.
For this strain we determined the metabolites using lyophilized cells rather than culture medium.
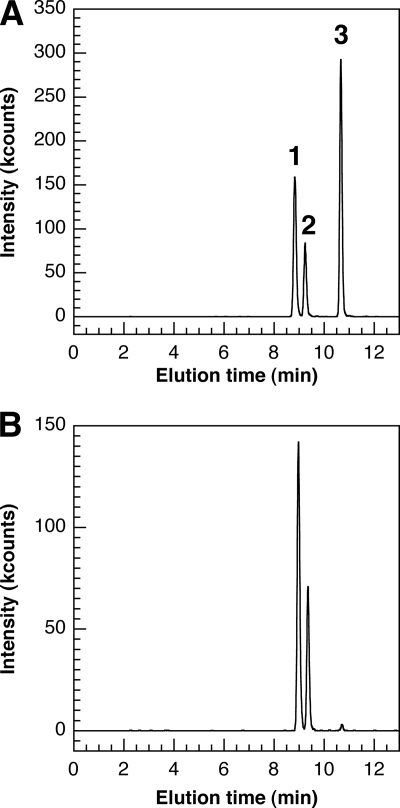
For the LC-MS2 method, we used a C18 reversed-phase HPLC column with an optimized gradient of methanol for separation, and we detected the three metabolites either by UV detection at 262 nm or by mass spectrometry using the MRM mode. As shown in Fig. 4 A, these conditions allowed the separation of the three natural metabolites 7-epi-CYN (eluting at 8.9 min), CYN (eluting at 9.3 min), and 7-deoxy-CYN (eluting at 10.6 min) and, thus, their relative quantifications. By injecting a sample of a culture medium of Oscillatoria PCC 6506 (Fig. 4B), we confirmed the presence of 7-epi-CYN, CYN, and 7-deoxy-CYN, with the following relative concentrations: 68.6% for 7-epi-CYN, 30.2% for CYN, and 1.2% for 7-deoxy-CYN.
FIG. 4.
LC-MS2 analysis of 7-epi-CYN, CYN, and 7-deoxy-CYN production by Oscillatoria PCC 6506. The analysis was performed by LC-MS2 in the MRM mode. (A) A mixture of pure standards (10 μl) was injected: 7-epi-CYN (8.9 min; 7.1 ng) (1), CYN (9.3 min; 3.3 ng) (2), and 7-deoxy-CYN (10.6 min; 11.0 ng) (3). (B) A sample (10 μl) of a culture medium (BG11 medium with 10 weeks of culture) of Oscillatoria PCC 6506 was injected. 7-Epi-CYN (8.9 min; 68.6%), CYN (9.3 min; 30.2%), and 7-deoxy-CYN (10.6 min; 1.2%) were detected.
Using pure standards, we estimated the following concentrations in the Oscillatoria PCC 6506 culture medium after 11 weeks of culture (Table 2): 0.69 μg/ml and 0.28 μg/ml for 7-epi-CYN and CYN, respectively. The sum of these values matches approximately that determined by the ELISA method, confirming that the antibodies used in the ELISA kit recognize both epimers with similar affinities. The extracellular and intracellular concentrations of 7-epi-CYN and CYN in Oscillatoria PCC 6506 were determined together with the mass of dried cells. We found 0.29 μg per mg of dried cells of excreted 7-epi-CYN and 0.11 μg per mg of dried cells of excreted CYN, thus giving a ratio of 2.6 in favor of 7-epi-CYN. The intracellular concentrations were 0.18 μg/mg of 7-epi-CYN and 0.10 μg/mg of CYN, thus giving a ratio of 1.8 in favor of intracellular 7-epi-CYN. The intracellular concentration of the metabolites and the ratio of the 7-epi-CYN concentration over that of CYN are thus slightly lower than the corresponding extracellular figures. A more quantitative study is needed before any conclusions are drawn.
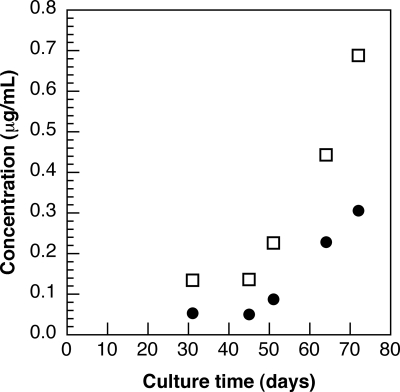
Using our LC-MS2 methodology, we have determined the concentrations of 7-epi-CYN and CYN in the culture medium of growing Oscillatoria PCC 6506 cells at different time points (Fig. 5). As shown in Fig. 5, both metabolites accumulated, but the ratio of the 7-epi-CYN concentration over that that of CYN was constant, with an average value of 2.5 ± 0.3. The concentration of the metabolites was proportional to the biomass, as determined by turbidimetry at 750 nm (data not shown). This finding well illustrates that the metabolites accumulate in the culture medium and that the extracellular fraction of these molecules is a far-better source than the intracellular fraction to obtain large quantities of these toxins as standards.
FIG. 5.
Accumulation of 7-epi-CYN and CYN in the culture medium of Oscillatoria PCC 6506. Oscillatoria PCC 6506 was cultivated in BG11 medium at 20°C under constant illumination, and aliquots of the culture medium were withdrawn at different time points. The concentrations of 7-epi-CYN (□) and CYN (•) excreted were then determined by LC-MS2 analysis. Each data point represents the average of data from two determinations. The experimental errors were less than 10%.
In A. ovalisporum, 7-epi-CYN was detected as a minor metabolite, and it was the first time that this metabolite was found as the major CYN analog produced by a cyanobacterium. In the biosynthetic scheme proposed previously by Mihali et al., CYN is formed by the hydroxylation of 7-deoxy-CYN catalyzed by CyrI, a protein of the α-ketoglutarate iron-dependent hydroxylase family. It is possible that this enzyme shows a relaxed stereospecificity and produces both stereoisomers at position 7. However, it is also possible that an unidentified protein produces the 7-epi-CYN from 7-deoxy-CYN. The difference in epimer concentrations would reflect the proportion of these two putative hydroxylases in a particular cyanobacterium. Further work is needed to answer that question more precisely.
Production of CYN and 7-epi-CYN by other Oscillatoria strains.
Intrigued by the fact that an Oscillatoria strain could produce CYN and 7-epi-CYN, we determined the concentrations of these two compounds in culture medium of four other Oscillatoria strains obtained from the PCC of cyanobacteria (17) by using ELISA and LC-MS2 methodologies. As shown in Table 2 we found three CYN and 7-epi-CYN Oscillatoria producers and one (PCC 6602) that did not produce these metabolites. As discussed above, the ELISA determinations gave an estimate of the total concentration, that is, 7-epi-CYN and CYN, while the LC-MS2 analysis gave a determination for each epimer. Interestingly, the ratios of the 7-epi-CYN concentration over that of CYN varied little for strains PCC 6407, PCC 6506, and PCC 10702, with ratios of 3.2, 2.5, and 2.9, respectively, but the ratio was very low for PCC 7926, with a ratio of 0.01. Thus, Oscillatoria PCC 7926, like A. ovalisporum, produces much more CYN than 7-epi-CYN, contrary to the other Oscillatoria strains tested here.
Using specific primers to amplify the internal DNA sequence of the cyrJ gene (25), we showed that the observation of a specific PCR product (563 bp) correlated with the CYN/7-epi-CYN production phenotype (Table 2). The sequence of these PCR products was determined and found to be identical (100% identity) to that of the corresponding cyrJ sequence of Oscillatoria PCC 6506 and similar (91% identity) to that of the corresponding cyrJ sequence of C. raciborskii. Thus, these strains have probably acquired the cyr cluster of genes by horizontal transfer in a recent time of the evolution.
In conclusion, we have identified the cyr gene cluster in Oscillatoria PCC 6506 and showed that these cyr genes are homologous to those of C. raciborskii but that the two clusters have probably diverged from a common ancestor. We have also shown that cyanobacteria of the genus Oscillatoria isolated from freshwater or brackish water are capable of producing CYN and 7-epi-CYN, the latter being the major product of several strains tested. This is the first time that Oscillatoria benthic cyanobacteria have been shown to produce CYN, and given the fact that these cyanobacteria are very commonly found in Europe, like in the Tarn River, France, we believe that CYN producers will be frequently detected in the future. It is thus likely that CYN is not a cyanobacterial toxin characteristic of hot-climate countries, as was first suggested, but a worldwide, widespread toxin. Using our sensitive LC-MS2 technology, we are currently testing other cyanobacteria and field samples to test these hypotheses.
Acknowledgments
We thank C. Médigue and her team (Génoscope, CNS, CEA, Evry, France) for help and advice concerning the annotation of the genome of Oscillatoria PCC 6506.
Footnotes
Published ahead of print on 4 June 2010.
REFERENCES
- 1.Banker, R., S. Carmeli, O. Hadas, B. Teltsch, R. Porat, and A. Sukenik. 1997. Identification of cylindrospermopsin in Aphanizomenon ovalisporum (Cyanophyceae) isolated from Lake Kinneret, Israel. J. Phycol. 33:613-616. [Google Scholar]
- 2.Banker, R., B. Teltsch, A. Sukenik, and S. Carmeli. 2000. 7-Epicylindrospermopsin, a toxic minor metabolite of the cyanobacterium Aphanizomenon ovalisporum from lake Kinneret, Israel. J. Nat. Prod. 63:387-389. [DOI] [PubMed] [Google Scholar]
- 3.Bláhová, L., M. Oravec, B. Marsálek, L. Sejnohová, Z. Simek, and L. Bláha. 2009. The first occurrence of the cyanobacterial alkaloid toxin cylindrospermopsin in the Czech Republic as determined by immunochemical and LC/MS methods. Toxicon 53:519-524. [DOI] [PubMed] [Google Scholar]
- 4.Brient, L., M. Lengronne, M. Bormans, and J. Fastner. 2009. First occurrence of cylindrospermopsin in freshwater in France. Environ. Toxicol. 24:415-420. [DOI] [PubMed] [Google Scholar]
- 5.Burgoyne, D. L., T. K. Hemscheidt, R. E. Moore, and M. T. Runnegar. 2000. Biosynthesis of cylindrospermopsin. J. Org. Chem. 65:152-156. [DOI] [PubMed] [Google Scholar]
- 6.Cadel-Six, S., C. Peyraud-Thomas, L. Brient, N. Tandeau de Marsac, R. Rippka, and A. Méjean. 2007. Different genotypes of anatoxin-producing cyanobacteria coexist in the Tarn River, France. Appl. Environ. Microbiol. 73:7605-7614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cadel-Six, S., I. Iteman, C. Peyraud-Thomas, S. Mann, O. Ploux, and A. Méjean. 2009. Identification of a polyketide synthase coding sequence specific for anatoxin-a-producing Oscillatoria cyanobacteria. Appl. Environ. Microbiol. 75:4909-4912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chorus, I., and J. Bartram (ed.). 1999. Toxic cyanobacteria in water: a guide to their public health consequences, monitoring and management. Spon Press, London, United Kingdom.
- 9.Comte, K., M. Sabacká, A. Carré-Mlouka, J. Elster, and J. Komárek. 2007. Relationships between the Arctic and the Antarctic cyanobacteria; three Phormidium-like strains evaluated by a polyphasic approach. FEMS Microbiol. Ecol. 59:366-376. [DOI] [PubMed] [Google Scholar]
- 10.Dörr, F. A., J. C. Tomaz, N. P. Lopes, and E. Pinto. 2008. Comparative analysis of the gas-phase reactions of cylindrospermopsin and the difference in the alkali metal cation mobility. Rapid Commun. Mass Spectrom. 22:2015-2020. [DOI] [PubMed] [Google Scholar]
- 11.Edwards, C., K. Beattie, C. Scrimgeour, and G. Codd. 1992. Identification of anatoxin-a in benthic cyanobacteria (blue-green algae) and in associated dog poisoning at Loch Insh, Scotland. Toxicon 30:1165-1175. [DOI] [PubMed] [Google Scholar]
- 12.Fastner, J., R. Heinze, A. R. Humpage, U. Mischke, G. K. Eaglesham, and I. Chorus. 2003. Cylindrospermopsin occurrence in two German lakes and preliminary assessment of toxicity and toxin production of Cylindrospermopsis raciborskii (Cyanobacteria) isolates. Toxicon 42:313-321. [DOI] [PubMed] [Google Scholar]
- 13.Gallo, P., S. Fabbrocino, M. G. Cerulo, P. Ferranti, M. Bruno, and L. Serpe. 2009. Determination of cylindrospermopsin in freshwaters and fish tissue by liquid chromatography coupled to electrospray ion trap mass spectrometry. Rapid Commun. Mass Spectrom. 23:3279-3284. [DOI] [PubMed] [Google Scholar]
- 14.Griffiths, D. J., and M. L. Saker. 2003. The Palm Island mystery disease 20 years on: a review of research on the cyanotoxin cylindrospermopsin. Environ. Toxicol. 18:78-93. [DOI] [PubMed] [Google Scholar]
- 15.Harada, K.-I., I. Ohtani, K. Iwamoto, M. Suzuki, M. F. Watanabe, M. Watanabe, and K. Terao. 1994. Isolation of cylindrospermopsin from a cyanobacterium Umezakia natans and its screening method. Toxicon 32:73-84. [DOI] [PubMed] [Google Scholar]
- 16.Hawkins, P. R., M. T. C. Runnegar, A. R. B. Jackson, and I. R. Falconer. 1985. Severe hepatotoxicity caused by the tropical cyanobacterium (blue-green alga) Cylindrospermopsis raciborskii (Woloszynska) Seenaya and Subba Raju isolated from a domestic water supply reservoir. J. Appl. Environ. Microbiol. 50:1292-1295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Herdman, M., I. Iteman, and R. Rippka. 2005. Catalogue of cyanobacterial strains, 2nd ed. Institut Pasteur, Paris, France.
- 18.Iteman, I., R. Rippka, N. Tandeau de Marsac, and M. Herdman. 2002. rDNA analyses of planktonic heterocystous cyanobacteria, including members of the genera Anabaenopsis and Cyanospira. Microbiology 148:481-496. [DOI] [PubMed] [Google Scholar]
- 19.Jones, A. C., L. Gu, C. M. Sorrels, D. H. Sherman, and W. H. Gerwick. 2009. New tricks from ancient algae: natural products biosynthesis in marine cyanobacteria. Curr. Opin. Chem. Biol. 13:216-223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li, R., W. W. Carmichael, S. Brittain, G. K. Eaglesham, G. R. Shaw, Y. Liu, and M. M. Watanabe. 2001. First report of the cyanotoxins cylindrospermopsin and deoxycylindrospermopsin from Raphidiopsis curvata (Cyanobacteria). J. Phycol. 37:1121-1126. [Google Scholar]
- 21.Li, R., W. W. Carmichael, S. Brittain, G. K. Eaglesham, G. R. Shaw, A. Mahakhant, N. Noparatnaraporn, W. Yongmanitchai, K. Kaya, and M. M. Watanabe. 2001. Isolation and identification of the cyanotoxin cylindrospermopsin and deoxy-cylindrospermopsin from a Thailand strain of Cylindrospermopsis raciborskii (Cyanobacteria). Toxicon 39:973-980. [DOI] [PubMed] [Google Scholar]
- 22.Looper, R. E., M. T. Runnegar, and R. M. Williams. 2005. Synthesis of the putative structure of 7-deoxycylindrospermopsin: C7 oxygenation is not required for the inhibition of protein synthesis. Angew. Chem. Int. Ed. Engl. 44:3879-3881. [DOI] [PubMed] [Google Scholar]
- 23.Méjean, A., S. Mann, G. Vassiliadis, B. Lombard, D. Loew, and O. Ploux. 2010. In vitro reconstitution of the first steps of anatoxin-a biosynthesis in Oscillatoria PCC 6506: from free L-proline to acyl carrier protein bound dehydroproline. Biochemistry 49:103-113. [DOI] [PubMed] [Google Scholar]
- 24.Méjean, A., S. Mann, T. Maldiney, G. Vassiliadis, O. Lequin, and O. Ploux. 2009. Evidence that biosynthesis of the neurotoxic alkaloids anatoxin-a and homoanatoxin-a in the cyanobacterium Oscillatoria PCC 6506 occurs on a modular polyketide synthase initiated by L-proline. J. Am. Chem. Soc. 131:7512-7513. [DOI] [PubMed] [Google Scholar]
- 25.Mihali, T. K., R. Kellmann, J. Muenchhoff, K. D. Barrow, and B. A. Neilan. 2008. Characterization of the gene cluster responsible for cylindrospermopsin biosynthesis. Appl. Environ. Microbiol. 74:716-722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mlouka, A., K. Comte, A. M. Castets, C. Bouchier, and N. Tandeau de Marsac. 2004. The gas vesicle gene cluster from Microcystis aeruginosa and DNA rearrangements that lead to loss of cell buoyancy. J. Bacteriol. 186:2355-2365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Norris, R. L., G. K. Eaglesham, G. Pierens, G. R. Shaw, M. J. Smith, R. K. Chiswell, A. A. Seawright, and M. R. Moore. 1999. Deoxycylindrospermopsin, an analog of cylindrospermopsin from Cylindrospermopsis raciborskii. Environ. Toxicol. 14:163-165. [DOI] [PubMed] [Google Scholar]
- 28.Ohtani, I., R. E. Moore, and M. T. C. Runnegar. 1992. Cylindrospermopsin, a potent hepatotoxin from the blue-green alga Cylindrospermopsis raciborskii. J. Am. Chem. Soc. 114:7942-7944. [Google Scholar]
- 29.Preussel, K., A. Stüken, C. Wiedner, I. Chorus, and J. Fastner. 2006. First report on cylindrospermopsin producing Aphanizomenon flos-aquae (Cyanobacteria) isolated from two German lakes. Toxicon 47:156-162. [DOI] [PubMed] [Google Scholar]
- 30.Rippka, R. 1988. Isolation and purification of cyanobacteria. Methods Enzymol. 167:3-27. [DOI] [PubMed] [Google Scholar]
- 31.Rücker, J., A. Stüken, B. Nixdorf, J. Fastner, I. Chorus, and C. Wiedner. 2007. Concentrations of particulate and dissolved cylindrospermopsin in 21 Aphanizomenon-dominated temperate lakes. Toxicon 50:800-809. [DOI] [PubMed] [Google Scholar]
- 32.Runnegar, M. T., C. Xie, B. B. Snider, G. A. Wallace, S. M. Weinreb, and J. Kuhlenkamp. 2002. In vitro hepatotoxicity of the cyanobacterial alkaloid cylindrospermopsin and related synthetic analogues. Toxicol. Sci. 67:81-87. [DOI] [PubMed] [Google Scholar]
- 33.Runnegar, M. T., S. M. Kong, Y. Z. Zhong, and S. C. Lu. 1995. Inhibition of reduced glutathione synthesis by cyanobacterial alkaloid cylindrospermopsin in cultured rat hepatocytes. Biochem. Pharmacol. 49:219-225. [DOI] [PubMed] [Google Scholar]
- 34.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- 35.Seifert, M., G. McGregor, G. Eaglesham, W. Wickramasinghe, and G. Shaw. 2007. First evidence for the production of cylindrospermopsin and deoxy-cylindrospermopsin by the freshwater benthic cyanobacterium, Lyngbya wollei (Farlow ex Gomont) Speziale and Dyck. Harmful Algae 6:73-80. [Google Scholar]
- 36.Shalev-Alon, G., A. Sukenik, O. Livnah, R. Schwarz, and A. Kaplan. 2002. A novel gene encoding amidinotransferase in the cylindrospermopsin producing cyanobacterium Aphanizomenon ovalisporum. FEMS Microbiol. Lett. 209:87-91. [DOI] [PubMed] [Google Scholar]
- 37.Shalev-Malul, G., J. Lieman-Hurwitz, Y. Viner-Mozzini, A. Sukenik, A. Gaathon, M. Lebendiker, and A. Kaplan. 2008. An AbrB-like protein might be involved in the regulation of cylindrospermopsin production by Aphanizomenon ovalisporum. Environ. Microbiol. 10:988-999. [DOI] [PubMed] [Google Scholar]
- 38.Spoof, L., K. A. Berg, J. Rapala, K. Lahti, L. Lepistö, J. S. Metcalf, G. A. Codd, and J. Meriluoto. 2006. First observation of cylindrospermopsin in Anabaena lapponica isolated from the boreal environment (Finland). Environ. Toxicol. 21:552-560. [DOI] [PubMed] [Google Scholar]
- 39.Thompson, J. D., T. J. Gibson, F. Plewniak, F. Jeanmougin, and D. G. Higgins. 1997. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25:4876-4882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Törökné, A., M. Asztalos, M. Bánkiné, H. Bickel, G. Borbély, S. Carmeli, G. A. Codd, J. Fastner, Q. Huang, A. Humpage, J. S. Metcalf, E. Rábai, A. Sukenik, G. Surányi, G. Vasas, and V. Weiszfeiler. 2004. Interlaboratory comparison trial on cylindrospermopsin measurement. Anal. Biochem. 332:280-284. [DOI] [PubMed] [Google Scholar]
- 41.Vallenet, D., L. Labarre, Z. Rouy, V. Barbe, S. Bocs, S. Cruveiller, A. Lajus, G. Pascal, C. Scarpelli, and C. Médigue. 2006. MaGe: a microbial genome annotation system supported by synteny results. Nucleic Acids Res. 34:53-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.van Apeldoorn, M. E., H. P. van Egmond, G. J. Speijers, and G. J. Bakker. 2007. Toxins of cyanobacteria. Mol. Nutr. Food Res. 51:7-60. [DOI] [PubMed] [Google Scholar]
- 43.Wonnacott, S., and T. Gallagher. 2006. The chemistry and pharmacology of anatoxin-a and related homotropanes with respect to nicotinic acetylcholine receptors. Mar. Drugs 4:228-254. [Google Scholar]
- 44.Wood, S. A., A. I. Selwood, A. Rueckert, P. T. Holland, J. R. Milne, K. F. Smith, B. Smits, L. F. Watts, and C. S. Cary. 2007. First report of homoanatoxin-a and associated dog neurotoxicosis in New Zealand. Toxicon 50:292-301. [DOI] [PubMed] [Google Scholar]