Abstract
The Oxa1 translocase of the mitochondrial inner membrane facilitates the insertion of both mitochondrially and nuclear-encoded proteins from the matrix into the inner membrane. Most mitochondrially encoded proteins are hydrophobic membrane proteins which are integrated into the lipid bilayer during their synthesis on mitochondrial ribosomes. The molecular mechanism of this co-translational insertion process is unknown. Here we show that the matrix-exposed C-terminus of Oxa1 forms an α-helical domain that has the ability to bind to mitochondrial ribosomes. Deletion of this Oxa1 domain strongly diminished the efficiency of membrane insertion of subunit 2 of cytochrome oxidase, a mitochondrially encoded substrate of the Oxa1 translocase. This suggests that co-translational membrane insertion of mitochondrial translation products is facilitated by a physical interaction of translation complexes with the membrane-bound translocase.
Keywords: co-translational insertion/mitochondria/protein export/Oxa1/ribosomes
Introduction
The proteome of mitochondria represents a mosaic of dual genetic origin. The vast majority of mitochondrial proteins are encoded in the nucleus and imported into the organelle following synthesis in the cytosol. In addition, a small number of gene products are encoded by the mitochondrial genome (eight in the yeast Saccharomyces cerevisiae, 13 in humans). These mitochondrially encoded proteins are almost exclusively highly hydrophobic polypeptides which form membrane-embedded reaction centers of the oxidative phosphorylation complexes. During evolution, most genetic information for mitochondrial proteins was transferred from the organelle to the nuclear genome. The hydrophobic nature of some gene products and, as a consequence, their tendency to form unproductive aggregates might have prevented the transfer of their respective genes to the nucleus. The requirement for some proteins to be inserted co-translationally into the inner membrane therefore might have forced eukaryotic cells to maintain a protein-synthesizing machinery in mitochondria (for discussion see Herrmann, 2003). A co-translational mode of insertion of mitochondrial translation products was suggested mainly by two lines of evidence. First, electron microscopic studies revealed a location of translation-active ribosomes preferentially in close proximity to the inner membrane of mitochondria (Watson, 1972). Secondly, biochemical fractionation experiments indicated that synthesis and membrane integration of mitochondrial gene products occur in a simultaneous and coordinated manner (Sevarino and Poyton, 1980; Liu and Spremulli, 2000).
The molecular basis of the coupling of protein synthesis and membrane integration in mitochondria is not well understood. No homologs of signal recognition particles were found in mitochondria. Studies in the yeast S.cerevisiae revealed that membrane-associated protein translational activators physically and functionally interact with 5′-untranslated leaders of mitochondrial mRNAs (for a review see Fox, 1996). These translational activators thereby restrict the initiation of protein synthesis to a zone close to the inner membrane.
A central component in the insertion process of mitochondrially encoded proteins is the inner membrane protein Oxa1. Oxa1 belongs to the recently identified Oxa1/YidC/Alb3 family whose members mediate protein integration into membranes of mitochondria, bacteria and chloroplasts (Luirink et al., 2001). The Oxa1 protein of S.cerevisiae is the founding member of this family. It was identified originally as a component required for the biogenesis of respiratory chain complexes in mitochondria (Bauer et al., 1994; Bonnefoy et al., 1994). Oxa1 forms a homooligomeric protein complex that facilitates integration of several mitochondrially encoded proteins into the inner membrane (He and Fox, 1997; Hell et al., 2001; Nargang et al., 2002). In addition, Oxa1 mediates membrane insertion of some nuclear-encoded proteins which reach the inner membrane in an export-like step from the matrix (Hell et al., 1998). Oxa1 spans the inner membrane five times, exposing a long C-terminal domain into the matrix (Herrmann et al., 1997). In the current study, we present evidence that this Oxa1 C-terminal domain has the ability to bind mitochondrial ribosomes. Deletion of this domain abrogates the interaction of ribosomes with the Oxa1 translocase and leads to the accumulation of mitochondrial translation products in the matrix. In contrast, we observed only a minor effect on the Oxa1-dependent membrane integration of a nuclear-encoded precursor protein. This suggests that the C-terminal domain of Oxa1 serves as a ribosome-binding site that allows coupling of synthesis and membrane integration of mitochondrial translation products.
Results
The C-terminus of Oxa1 is required for biogenesis of the respiratory chain
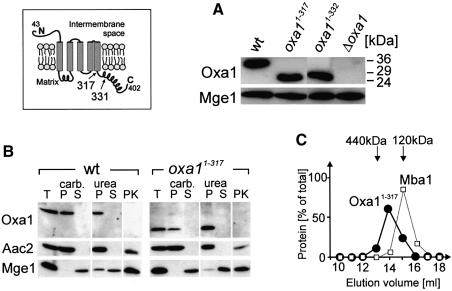
Oxa1 homologs comprise a well-conserved core domain of five transmembrane stretches which most probably performs the translocation function of these proteins (Luirink et al., 2001). This domain is flanked by less conserved regions. In contrast to their bacterial homologs, mitochondrial Oxa1 proteins contain a C-terminal domain of ∼100 amino acid residues that protrudes into the mitochondrial matrix (Herrmann et al., 1997). To assess the functional relevance of the Oxa1 C-terminal domain, yeast mutants were constructed in which the C-terminal matrix domain was completely (Oxa11–317) or partially truncated (Oxa11–331) (cf. Figure 1, inset). These mutations did not significantly change the steady-state levels of the Oxa1 protein (Figure 1A). Truncation of the C-terminus also did not affect the topology of the protein (Figure 1B) nor its assembly into an oligomeric complex (Figure 1C).
Fig. 1. Deletion of the C-terminal matrix domain of Oxa1 leads to deficiencies in respiratory chain complexes. The inset shows a model of the Oxa1 topology in the inner membrane. Numbers refer to amino acid residues of the sequence of the Oxa1 precursor. (A) Oxa1 levels are not significantly altered in truncation mutants. Mitochondrial protein (100 µg) of wild-type (wt), Oxa11–317, Oxa11–331 and Δoxa1 cells was subjected to SDS–PAGE and immunoblotted using antibodies against the N-terminus of mature Oxa1 and Mge1. The matrix protein Mge1 was used as a loading control. The positions of molecular size standards are indicated. (B) Localization of Oxa11–317. Proteins from wild-type (wt) and Oxa11–317 mitochondria were either loaded directly (T, total) or following fractionation into a membrane (P) and a soluble (S) fraction by extraction with sodium carbonate (carb.) or urea. To assess the accessibility of the N-terminus of Oxa1 from the intermembrane space, mitochondria were swollen and incubated with 50 µg of proteinase K for 30 min on ice (PK). Samples were analyzed by immunoblotting with antibodies against the N-terminus of Oxa1, the inner membrane protein ATP/ADP carrier (Aac2) and the matrix protein Mge1. (C) Oxa11–317 mitochondria were lysed with dodecyl maltoside. Proteins in the resulting extract were separated by gel filtration on a Superose 6 column (Pharmacia). The distribution of Oxa11–317 and Mba1 was analyzed by immunoblotting, and the resulting signals were quantified.
Whereas only a slight growth defect was observed for the Oxa11–331 mutant, deletion of the entire C-terminus prevented growth on a non-fermentable carbon source (Figure 2A). The growth defect of this mutant was restored upon transformation of an Oxa1-expressing plasmid (not shown). The growth defects in the Oxa11–317 strain were apparently more severe than that reported in the accompanying paper (Jia et al., 2003), which might be due to different genetic backgrounds of the mutants. Oxa1 is essential for the biogenesis of the cytochrome oxidase complex (Bauer et al., 1994; Bonnefoy et al., 1994). To assess the significance of the Oxa1 C-terminal domain in this process, the activities of respiratory chain complexes were measured in the oxa1 mutants (Figure 2B–D). No detectable cytochrome oxidase activity was found with mitochondria of an oxa1 deletion mutant. Truncation of the entire C-terminal matrix domain of Oxa1 likewise almost completely abolished the activity of this respiratory chain complex. Reduced functionality of the cytochrome oxidase was also observed in the Oxa11–331 mutant in which ∼35% cytochrome oxidase activity was retained. In addition, the activity levels of the cytochrome bc1 complex and ATP synthase were severely reduced in the Oxa1 truncation mutants. Thus, Oxa1 requires its C-terminal matrix domain to perform its role in the biogenesis of the oxidative phosphorylation complexes. The levels of malate dehydrogenase were assessed for control. This nuclear-encoded enzyme of the mitochondrial matrix showed comparable activities in the different mutant strains (Figure 2E).
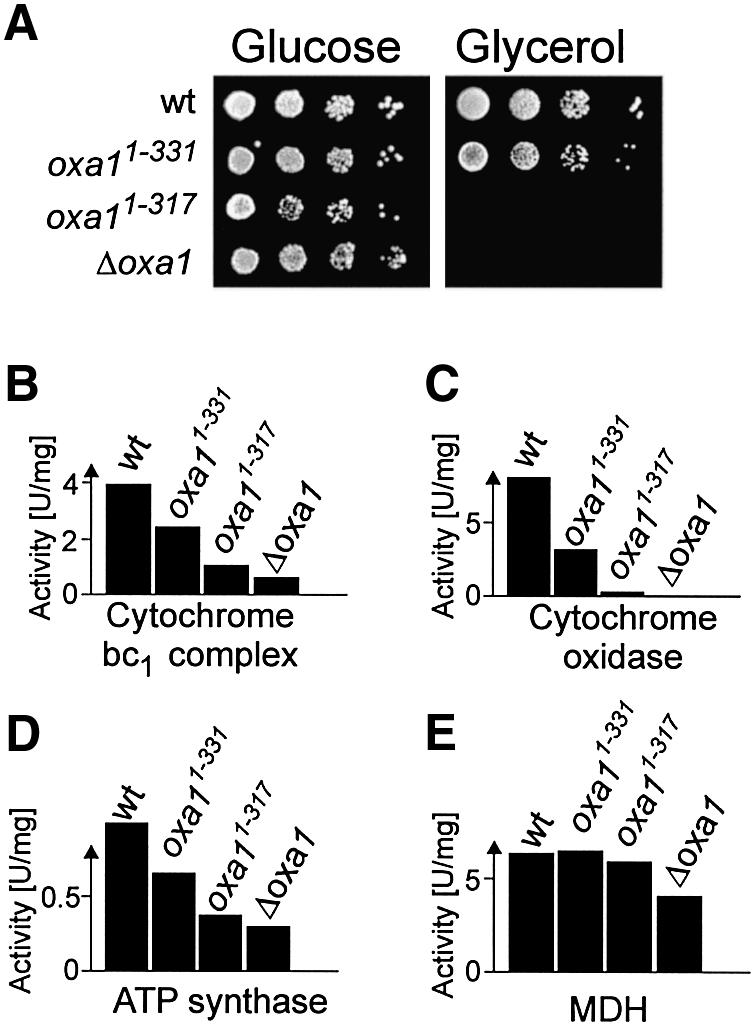
Fig. 2. Deletion of the entire Oxa1 C-terminal domain prevents growth on a non-fermentable carbon source. (A) Ten-fold serial dilutions of wild-type (wt), Oxa11–331, Oxa11–317 and Δoxa1 cells were spotted onto agar plates containing 2% peptone, 1% yeast extract containing 2% glucose or 2% glycerol as indicated. Plates were incubated for 2 days (glucose) or 3 days (glycerol) at 30°C. (B–E) The C-terminus of Oxa1 is required for the formation of respiratory chain complexes. Enzyme activities of cytochrome bc1 complex (B), cytochrome oxidase complex (C), ATP synthase (D) and malate dehydrogenase (E) were measured in wild-type and oxa1 mutant mitochondria.
The Oxa1 C-terminal domain is required for co-translational insertion of Cox2
The absence of cytochrome oxidase activity pointed to an insertion defect of cytochrome oxidase subunits in the Oxa11–317 mutant. Membrane integration of the mitochondrially encoded subunit 2 of cytochrome oxidase, Cox2, has been shown to occur in an Oxa1-dependent reaction (He and Fox, 1997; Hell et al., 1997). Cox2 is synthesized in the matrix with an N-terminal pre-sequence which is proteolytically removed following translocation into the intermembrane space. In the absence of Oxa1, the precursor form of Cox2 accumulates in the matrix and is degraded. As a consequence, no steady-state levels of Cox2 are found in mitochondria of oxa1 deletion strains (He and Fox, 1997; Hell et al., 1997). Immunoblotting of mitochondrial extracts of the Oxa11–317 mutant likewise revealed strongly reduced steady-state levels of Cox2 in mitochondria (Figure 3A). Besides very low amounts of mature Cox2, some Cox2 precursor was observed in the truncation mutant, suggesting that the defect in Cox2 biogenesis is caused by a block in membrane translocation of its N-terminus.
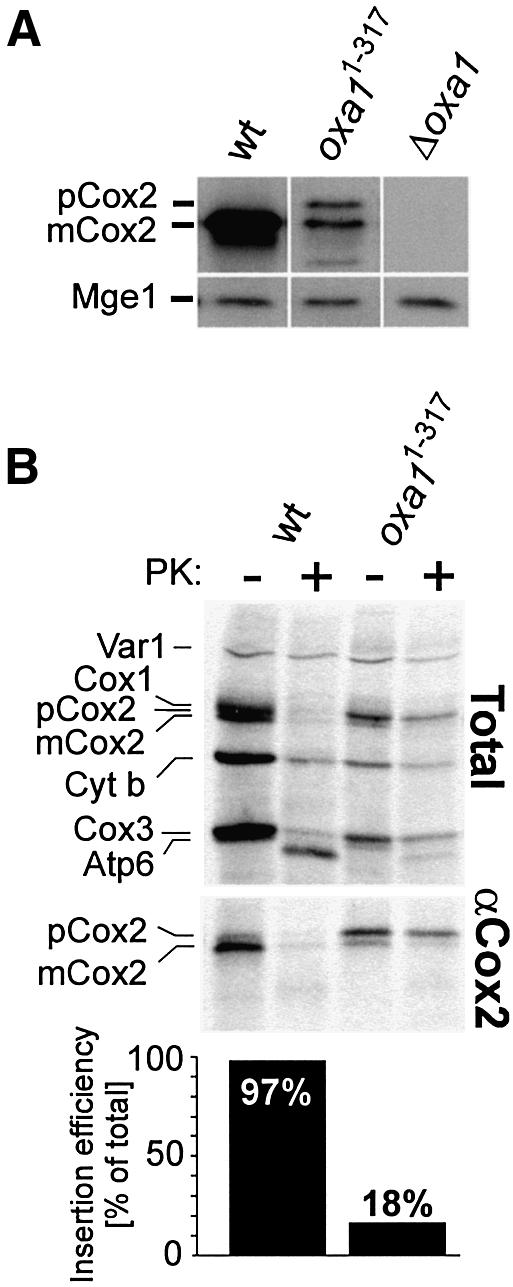
Fig. 3. Deletion of the C-terminus of Oxa1 blocks insertion of the mitochondrially encoded protein Cox2. (A) Deletion of the C-terminal domain of Oxa1 leads to strongly reduced steady-state levels of Cox2. A 100 µg aliquot of wild-type (wt), Oxa11–317 and Δoxa1 mitochondria was analyzed by immunoblotting using Cox2- and Mge1-specific antibodies. (B) Mitochondrial translation products were radiolabeled in wild-type or Oxa11–317 mitoplasts of the indicated strains for 30 min at 30°C. The samples were divided. One half was mock treated, and the other half was incubated with 50 µg/ml proteinase K (PK). The mitoplasts were washed and lysed in 1% SDS. The lysates were subjected either directly to SDS–PAGE (upper panel) or to immunoprecipitation using a C-terminal Cox2 antiserum (Herrmann et al., 1995) (middle panel). Efficient swelling was controlled by immunoblotting (not shown). The ratio of matured to total Cox2 was determined by densitometry as a measure of the insertion efficiency. The numbers are depicted in the lower panel following correction for the respective methionine content. pCox2, precursor form of Cox2; mCox2, mature form of Cox2; Cyt b, cytochrome b.
To analyze the significance of the Oxa1 C-terminal domain for membrane integration of Cox2 more directly, we radiolabeled mitochondrial translation products in wild-type and Oxa11–317 mitochondria and isolated the newly synthesized Cox2 protein by immunoprecipitation (Figure 3B). In the wild-type, Cox2 was almost entirely present in its mature form and correctly inserted. In contrast, in the Oxa11–317 mutant, Cox2 accumulated mostly in its precursor form. This precursor form of Cox2 remained inaccessible to protease after opening of the outer membrane, indicating its accumulation in the matrix. From this, we conclude that the Oxa1 C-terminal domain plays an important role in the integration of newly synthesized Cox2 protein into the inner membrane. In addition, in the Oxa11–317, mutant a significant fraction of the newly synthesized proteins Cox3 and cytochrome b remained protease resistant (Figure 3B), indicating that the Oxa1 C-terminal domain plays a general role in membrane integration of mitochondrially encoded Oxa1 substrates.
The C-terminus of Oxa1 is dispensable for membrane integration of Oxa1 and Cox18
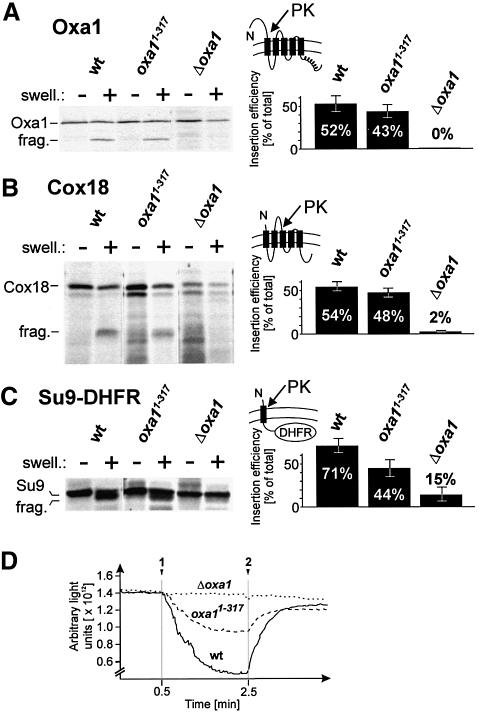
We asked whether the C-terminus of Oxa1 is likewise required for post-translational protein insertion of nuclear-encoded Oxa1 substrates. Previously characterized nuclear-encoded Oxa1 substrates are Oxa1 itself and Su9(1–112)-DHFR (Hell et al., 1998). In addition, we used the inner membrane protein Cox18 which also requires Oxa1 for membrane integration. The radiolabeled precursor proteins were imported into isolated mitochondria. Membrane insertion of the matured proteins from the matrix can be determined by protease treatment of mitoplasts which generates specific fragments from the membrane-embedded proteins (Rojo et al., 1995; Herrmann et al., 1997). In wild-type and Oxa11–317 mitochondria, Oxa1 and Cox18 were inserted efficiently (Figure 4A and B), whereas in the absence of Oxa1 insertion was completely blocked. Membrane insertion of Su9(1–112)-DHFR was reduced in the truncation mutant but was still significantly more efficient than in the absence of Oxa1 (Figure 4C). Since the export reaction of these proteins strongly depends on the membrane potential (Rojo et al., 1995; Herrmann et al., 1997), it is conceivable that the reduced insertion rates in Oxa11–317 mitochondria are caused by a lower membrane potential in this mutant due to its diminished activities of respiratory chain enzymes (Figure 4D). Alternatively, the C-terminus of Oxa1 might be more directly involved in the insertion of certain nuclear-encoded proteins. The ability of the truncated Oxa1 version to facilitate the insertion of some newly imported proteins indicates that deletion of the Oxa1 C-terminal domain does not cause a general insertion defect of Oxa1 substrates. It rather compromises the insertion of a subset of Oxa1 substrates including mitochondrial translation products.
Fig. 4. Deletion of the C-terminus of Oxa1 does not block membrane insertion of Oxa1 and Cox18. (A) Radiolabeled Oxa1 precursor was imported into 100 µg of wild-type (wt), Oxa11–317 or Δoxa1 mitochondria for 30 min at 30°C. The samples were split, and the mitochondria either were mock treated or the outer membrane was ruptured by hypotonic swelling (swell.). Then 50 µg/ml proteinase K was added to the samples. After 30 min on ice, the mitochondria were reisolated and proteins were resolved by SDS–PAGE and detected by autoradiography. From three independent experiments, the ratio of protease-accessible, i.e. inserted, to total imported Oxa1 was quantified by densitometry. The numbers were corrected for the methionine content of the proteins. Efficient swelling of the mitochondria was controlled by immunoblotting. frag., protease fragment of the inserted Oxa1 protein. The topology of the correctly inserted Oxa1 protein is depicted. (B and C) The insertion of Cox18 and Su9(1–112)-DHFR was analyzed as described in (A). (D) Deletion of the C-terminal domain of Oxa1 leads to a reduced membrane potential of mitochondria. The level of the membrane potential was determined as described in Hell et al. (1997). Additions of mitochondria (arrow 1) and of valinomycin and KCN (arrow 2) are indicated. Membrane potential-dependent uptake of the dye leads to a decrease in fluorescence. Upon dissipation of the membrane potential by addition of valinomycin, the dye is released and the fluorescent signal regained.
Oxa1 co-fractionates with mitochondrial ribosomes on sucrose gradients
The observed insertion defect of mitochondrial translation products in Oxa11–317 mitochondria could be explained by a compromised coordination of protein synthesis and protein translocation in this mutant. It was shown before that Oxa1 is in direct proximity to nascent polypeptide chains during their synthesis on mitochondrial ribosomes (Hell et al., 1998). This suggests that Oxa1 is close to translation-active ribosomes in mitochondria, which could be explained by the potential of the Oxa1 translocase to physically interact with ribosomes.
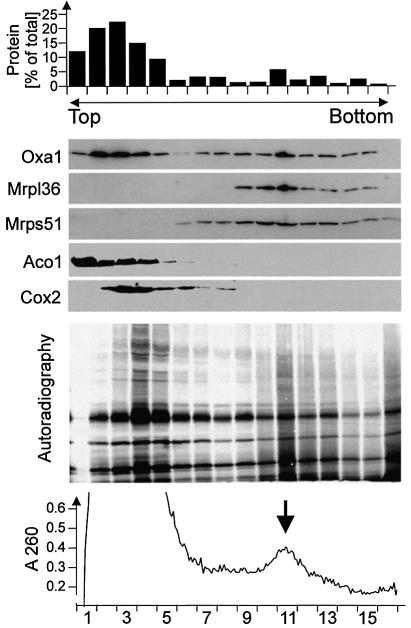
To assess an association of Oxa1 with the protein-synthesizing machinery in mitochondria, we tested whether Oxa1 co-purifies with mitochondrial ribosomes upon velocity centrifugation in sucrose gradients. In order to monitor the fractionation of mitochondrial translation products, isolated mitochondria were incubated in the presence of [35S]methionine under conditions stimulating protein synthesis in the matrix. Then the mitochondria were lysed in digitonin and subjected to sucrose gradient centrifugation. The majority of mitochondrial protein was recovered in the top fractions of the sucrose gradient (Figure 5, upper panel). The distribution of Oxa1, ribosomal proteins (Mrpl36 and Mrps51), Cox2 and the soluble matrix protein aconitase (Aco1) in the gradient was analyzed by immunoblotting (Figure 5, middle panel). The distribution profile of Oxa1 showed two different populations of the protein. A large proportion of Oxa1 migrated with the bulk of mitochondrial protein in the upper fractions of the gradient. In the same fractions, aconitase and Cox2 were found. About 15–20% of Oxa1 fractionated at higher sucrose densities. This Oxa1 population co-migrated with the ribosomal proteins Mrpl36 and Mrps51, indicating an association of this Oxa1 pool with mitochondrial ribosomes. The completed translation products fractionated mainly close to the top of the gradient, similar to the endogenous Cox2 (Figure 5, autoradiograph). In contrast, radiolabeled nascent polypeptide chains with a broad size distribution partially co-fractionated with mitochondrial ribosomes. We conclude that a fraction of the total Oxa1 protein is bound to ribosomes in mitochondria and, thus, that the machineries for synthesis and translocation of proteins in mitochondria are in physical contact.
Fig. 5. Oxa1 is associated with mitochondrial ribosomes. Mitochondrial translation products were radiolabeled in isolated wild-type mitochondria for 30 min. The mitochondria were washed and lysed in 2% digitonin. The extract was cleared by centrifugation and loaded onto a continuous 15–40% sucrose gradient. Following centrifugation for 16 h at 83 000 g, 16 fractions were collected. Total protein contents of the resulting fractions were determined (upper panel). Proteins of the fractions were precipitated by trichloroacetic acid and resolved by SDS–PAGE. The distribution of Oxa1, the ribosomal proteins Mrpl36 and Mrps51, and aconitase (Aco1) and Cox2 for control was analyzed by immunoblotting. The lower panel shows an autoradiograph of the same experiment to visualize the distribution of translation products. Note that completed translation products were predominantly present in the upper fractions of the gradient, whereas nascent polypeptides of less defined mass mainly migrated in fraction 11.
The Oxa1 C-terminal domain is required for binding to mitochondrial ribosomes
To assess whether deletion of the Oxa1 C-terminal domain domain influences the association of Oxa1 with mitochondrial ribosomes, we tested whether the C-terminally truncated mutant of Oxa1 still co-purifies with mitochondrial ribosomes. First translation products were radiolabeled in wild-type and Oxa11–331 mitochondria. The latter mutant was chosen due to its ability to grow on non-fermentable carbon sources, facilitating the isolation of decent amounts of mitochondria. After the labeling reaction, the mitochondria were lysed with digitonin and ribsomal pellet fractions were isolated by centrifugation through high-density sucrose cushions. The relative amounts of ribosomes (Mrpl36), Oxa1, ATP/ADP carrier (Aac2) and aconitase in the pellet fractions were assessed by immunoblotting (Figure 6A). Significant amounts of Oxa1 were found in the ribosomal fraction, confirming the physical association of Oxa1 with ribosomes (Figure 6A, lane 2). In mitochondrial extracts obtained from the Oxa11–331 mutant, no truncated Oxa1 protein was identified in the ribosomal fraction (Figure 6A, lane 6). From this, we conclude that the C-terminus of Oxa1 is required to allow binding of the Oxa1 translocase to ribosomes that is stable enough to survive the purification procedure. To test whether the presence of nascent chains on the ribosomes was essential for the interaction with Oxa1, mitochondria were treated with puromycin before lysis. This resulted in a complete release of the nascent chains from the ribosomes as no radioactivity was recovered in the ribosome-enriched fractions under these conditions (Figure 6A, lane 4, lower panel). Interestingly, a fraction of Oxa1 was still detectable in the ribosomal pellet, indicating that translation activity is not absolutely essential for the interaction of Oxa1 with ribosomes. This suggests that Oxa1 interacts with the ribosome not exclusively via contact with the nascent chains and points to a more direct role for the matrix-exposed C-terminus of Oxa1 in ribosome recruitment.
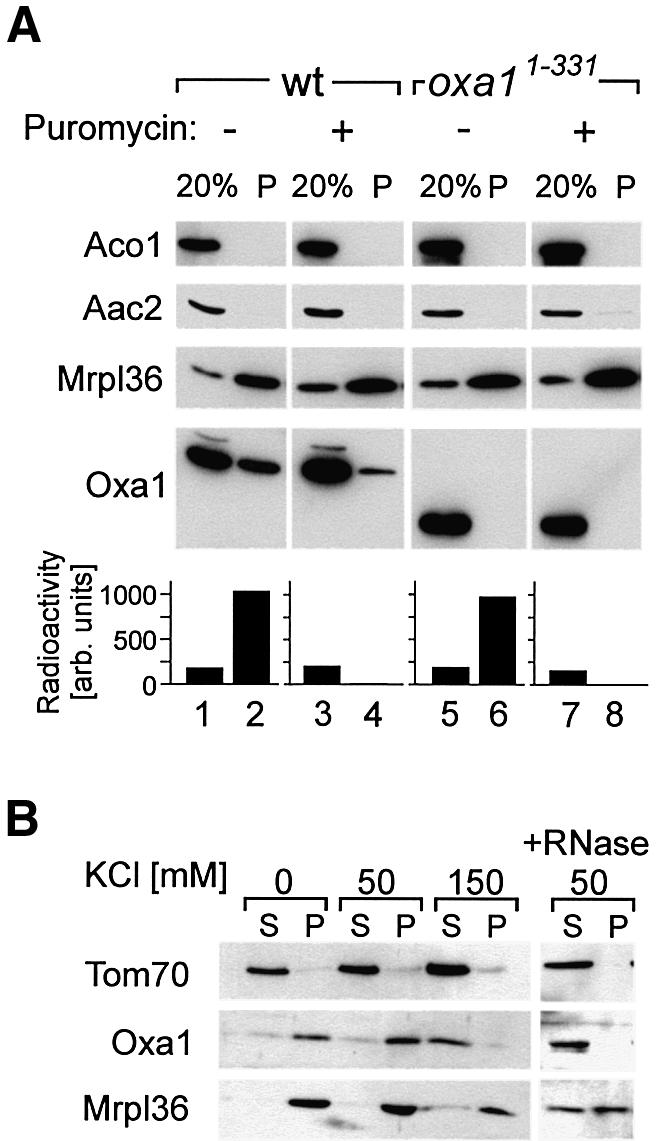
Fig. 6. The C-terminus of Oxa1 is required for stable binding of Oxa1 to mitochondrial ribosomes. (A) Translation products were radiolabeled in isolated wild-type or Oxa11–331 mitochondria prior to incubation with or without puromycin. Mitochondria were reisolated, washed and lysed. The extracts were cleared by centrifugation and ribosomes were sedimented by ultracentrifugation through a high-density sucrose cushion. The mitochondrial ribosomes and Oxa1 in the mitochondrial extract (20%) and in the ribosomal pellet (P) were analyzed by immunoblotting with antibodies specific for Mrpl36, Oxa1 and aconitase (Aco1). The lower panel shows the distribution of nascent chains in the gradients upon quantification of the autoradiographs by densitometry using a Pharmacia Image Scanner equipped with the Image Master 1D Elite software package. (B) The interaction of Oxa1 with the ribosomes is salt sensitive. Wild-type mitochondria (100 µg) were lysed in digitonin in the presence of the KCl concentrations indicated. A ribosomal fraction (P) was isolated as described in (A). Proteins in the supernatant (S) were precipitated. The distribution of Oxa1, Tom70 and Mrpl36 was analyzed by immunoblotting. For the rightmost lanes, the digitonin lysate was treated with 40 U/ml RNase for 60 min prior to the isolation of ribosomes.
The interaction of Oxa1 with mitochondrial ribosomes was salt sensitive as KCl concentrations of 150 mM caused the release of Oxa1 (Figure 6B). Similarly, when the integrity of the ribosomes was affected by the addition of RNase, Oxa1 did not sediment through the sucrose cushions (Figure 6B, right panel).
The C-terminal matrix domain of Oxa1 forms an α-helical structure
The C-terminal regions of mitochondrial Oxa1 homologs of plants, fungi and animals show no obvious conservation at the primary sequence level. However, coiled-coil structures are consistently predicted (Lupas, 1997) with high probability in this domain of Oxa1 homologs (see Figure 7A for yeast Oxa1) (Nargang et al., 2002). Coiled-coil-forming α-helices are common structural elements that mediate protein–protein interactions (Burkhard et al., 2001). For structural analysis, we expressed the C-terminal 89 residues of the yeast Oxa1 protein in Escherichia coli fused to GST (GST–Oxa1C) and purified it after cleavage of the fusion protein (Figure 7B). Circular dichroism spectroscopy of the C-terminal domain revealed a largely α-helical structure under the conditions tested, which is in agreement with the prediction of a coiled-coil interaction domain (Figure 7C). This led us to speculate that the Oxa1 C-terminal domain that protrudes into the matrix may play a direct role in the recruitment of the mitochondrial translation machinery.
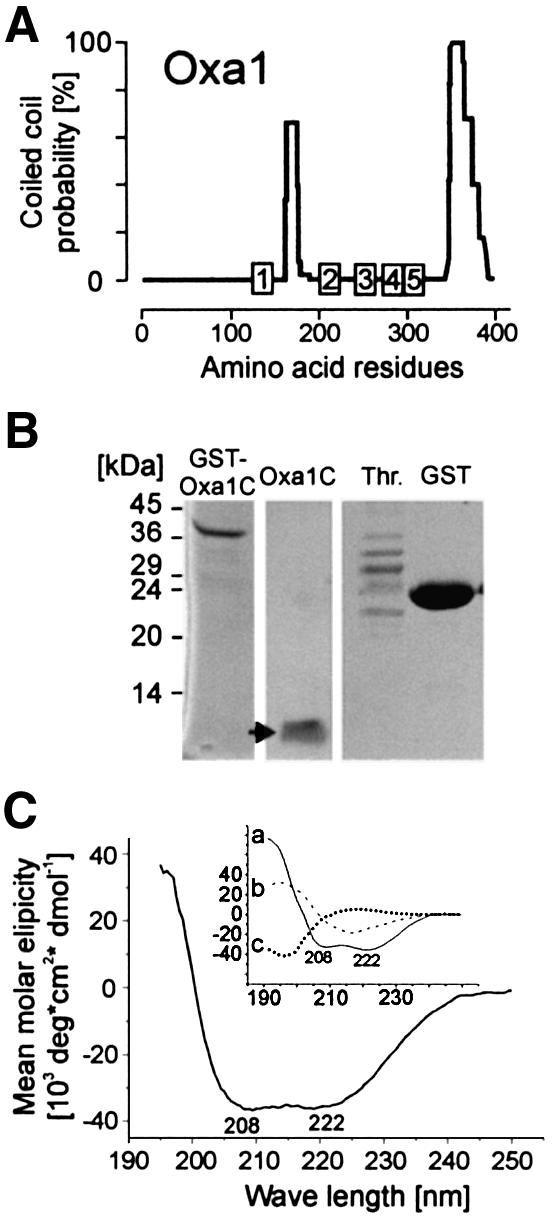
Fig. 7. The C-terminus of Oxa1 forms a coil structure. (A) Coiled-coil prediction for the S.cerevisiae Oxa1 sequence (Lupas, 1997). (B) The C-terminal 89 amino acid residues of the Oxa1 sequence (Oxa1C) were expressed in E.coli and purified without or with (arrow) proteolytic removal of the GST domain; thrombin (Th.) and GST are shown for control. (C) Circular dicroism spectrum of the purified Oxa1C domain (1 µM) in 50% trifluoroethanol, 20 mM potassium phosphate pH 7.2 (Kayed et al., 1999). The inset shows spectra of pure α-helical (a), β-sheet (b) or unstructured (c) sequences for comparison (Greenfield and Fasman, 1969).
The Oxa1 C-terminal domain binds to mitochondrial ribosomes in vitro
To test a putative ribosome binding ability of the Oxa1 C-terminus by affinity chromatography, GST–Oxa1C was immobilized on glutathione–Sepharose beads. For control, beads were used carrying either only the GST domain or GST fused to the C-terminal 110 residues of yeast Ssq1 (Lutz et al., 2001) (Figure 8A, upper panel). The latter domain shows a high probability score for coiled-coil formation similar to that of the C-terminus of Oxa1 (not shown). Translation products were radiolabeled in isolated mitochondria. The reaction was split. In one half of the reaction, nascent chains were stripped off from the ribosomes by incubation with puromycin following the labeling reaction. The mitochondria were reisolated, lysed with digitonin and the resulting extract applied on the immobilized fusion proteins. After extensive washing, the bound fractions were analyzed by immunoblotting with antibodies against ribosomal proteins (Mrpl36 and Mrps51) or aconitase (Aco1) for control (Figure 8A, middle panel). This revealed a specific interaction of GST–Oxa1C with mitochondrial ribosomes, regardless of the presence (lane 3) or absence (lane 7) of nascent chains on the ribosome. Autoradiography of the translation reaction (Figure 8A, lower panel, lanes 1 and 5) showed several distinct bands representing completed translation products and a background smear caused by incomplete nascent polypeptide chains of variable sizes. In the samples that were not pre-treated with puromycin, the nascent chains remained on the ribosomes and were specifically co-purifed with GST–Oxa1C, whereas completed translation products were not recovered with GST–Oxa1C (Figure 8A, lower panel, lane 3). Release of the nascent chains by puromycin abolished their interaction with GST–Oxa1C (lane 7). From this, we conclude that the C-terminal matrix domain of Oxa1 has the ability to bind to mitochondrial ribosomes. The presence of translation products on the ribosomes is not critical for this interaction.
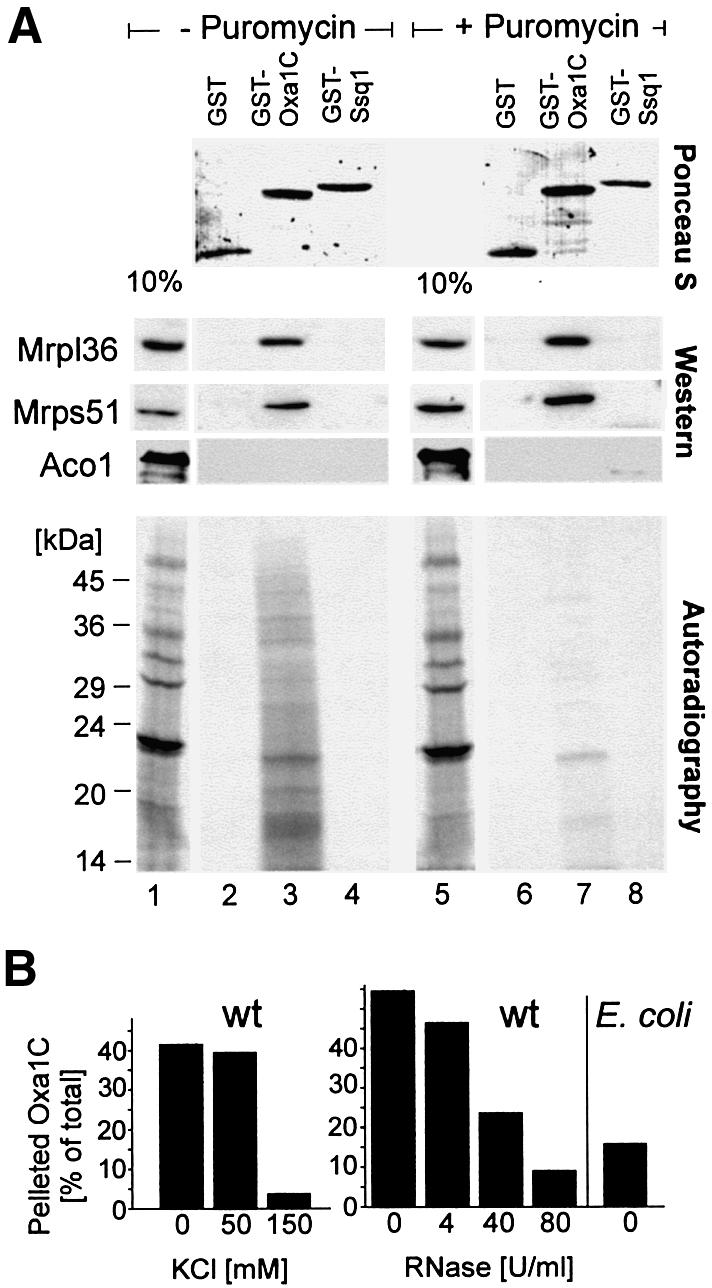
Fig. 8. The Oxa1 C-terminal domain specifically binds to ribosomes. (A) Mitochondrial translation products were radiolabeled in wild-type mitochondria prior to an incubation without (lanes 1–4) or with (lanes 5–8) puromycin. Mitochondria were lysed. The extracts were either applied directly to an SDS–polyacrylamide gel (lanes 1 and 5) or incubated with immobilized GST, GST–Oxa1C or GST–Ssq1. Following extensive washing, bound proteins were resolved by SDS–PAGE. Upper panel: bait proteins as detected by staining with Ponceau S. Middle panel: immunoblots using antisera against the ribosomal proteins Mrpl36 and Mrps51, or the matrix enzyme aconitase (Aco1) for control. Lower panel: autoradiography of the same membrane. The extracts applied in lanes 1 and 5 correspond to 10% of that used in the other lanes. The positions of molecular weight standards are indicated in the lower panel. (B) Wild-type mitochondria were lysed with digitonin as described for Figure 6B. The extracts were cleared by centrifugation and incubated with radiolabeled Oxa1314–402-DHFR in the absence or presence of KCl or RNase as indicated. Then, ribosomes were isolated by centrifugation and the amounts of Oxa1314–402-DHFR in the ribosomal fractions were quantified. For control, E.coli spheroplasts (500 µg) were lysed and incubated with Oxa1314–402-DHFR. Only low amounts of the fusion protein were recovered with the bacterial ribosomes.
To assess further the interaction of Oxa1 with ribosomes, a fusion protein containing the C-terminus of Oxa1 fused to DHFR was radiolabeled in vitro and incubated with mitochondrial or bacterial extracts. Following isolation of the ribosomes, the amounts of the fusion protein in the ribosomal fractions were assessed (Figure 8B). This again revealed the salt-sensitive interaction of Oxa1 with the ribosomes. The addition of increasing amounts of RNase led to a shift of ribosomal proteins to the supernatant fractions due to ribosomal disintegration (not shown). As a consequence, the radiolabeled protein was likewise shifted to the supernatant (Figure 8B, right). Only low amounts of the fusion protein sedimented with bacterial ribosomes, indicating a specific interaction of mitochondrial ribosomes with the C-terminus of Oxa1.
Discussion
The mitochondrial translation machinery of higher eukaryotes produces a small number of predominantly highly hydrophobic polypeptides. The Oxa1 translocase plays a pivotal role in the integration of these polypeptides into the lipid bilayer of the inner membrane. Here we show that the C-terminus of Oxa1 forms an α-helical domain that has the ability to bind to mitochondrial ribosomes. This interaction was observed both in vitro and in organello. The C-terminal domain of Oxa1 was both necessary and sufficient to bind mitochondrial ribosomes. Deletion of the entire C-terminus led to strong defects in the biogenesis of the respiratory chain and almost completely blocked insertion of mitochondrial translation products. The Oxa11–331 mutant, in which 15 amino acid residues of the C-terminal matrix domain were retained, was less severely affected, but also showed diminished activity levels of respiratory chain complexes. In extracts of this mutant, no Oxa1 co-purified with mitochondrial ribosomes, indicating that the truncation abolished or destabilized ribosome binding. The more moderate defects in this strain might be due to the differences in the generation of the mutants: whereas in the Oxa11–317 mutant the OXA1 reading frame was interrupted by insertion of a disruption cassette into the genome, the Oxa11–331 mutant was generated by introduction of a single stop codon into the OXA1 gene. The introduction of premature stop codons has been shown in other cases to lead to an incomplete termination (Fearon et al., 1994). This phenomenon, called termination read-through, might lead to low levels of wild-type Oxa1 protein in the Oxa11–331 mutant. This mutant still was valuable since its ability to grow on non-fermentable carbon sources allowed the purification of the rather large amounts of mitochondria required for the co-fractionation experiments.
The presence of nascent chains on the ribosomes was not a prerequisite for the association of ribosomes with Oxa1. Interestingly, it was reported before that the interaction of ribosomes with inverted inner membrane vesicles of mammalian mitochondria likewise was not abolished upon release of nascent chains by puromycin (Liu and Spremulli, 2000). In the same study, this interaction was shown to be highly sensitive to salt, and concentrations as low as 90 mM KCl released most of the ribosomes from the membrane. This is consistent with our observation of a rather salt-sensitive interaction of the ribosomes with Oxa1. However, the co-purification of Oxa1 with ribosomes on sucrose gradients demonstrated a highly persistent interaction under these conditions so that even after the 18 h of the experiment, a significant proportion the mitochondrial Oxa1 remained bound to ribosomes.
What is the function of a ribosome-binding site on Oxa1? Deletion of the entire C-terminal matrix domain of Oxa1 led to severe defects in the biogenesis of the respiratory chain and prevented growth on non-fermentable carbon sources. Insertion of Cox2 following its synthesis on mitochondrial ribosomes was almost completely blocked. In addition, the reduced levels of cytochrome bc1 and ATP synthase activity were consistent with a general defect in the biogenesis of mitochondrially encoded membrane proteins. In contrast, the nuclear-encoded proteins Oxa1 and Cox18 inserted into the inner membrane almost as efficiently as in wild-type. Thus, truncation of the C-terminal matrix domain of Oxa1 does not perturb the insertion activity of the Oxa1 translocase in general but causes a specific defect for a subset of Oxa1 substrates including proteins synthesized on mitochondrial ribosomes. This suggests that the C-terminus of Oxa1 plays a critical role in the co-translational insertion reaction of mitochondrially encoded proteins, most probably by a physical coupling of the protein synthesis and protein insertion machineries. The reduced insertion rates for Su9(1–112)-DHFR in a C-terminal Oxa1 deletion mutant might be due to the strong dependence of this substrate on the membrane potential (Rojo et al., 1995). Alternatively, the Oxa1 C-terminal domain may also play a role in the recruitment of some nuclear-encoded substrates.
How is a co-translational insertion mode achieved in mitochondria? Our current knowledge of this process leads us to propose the following model (Figure 9). Protein synthesis in mitochondria depends on mRNA-specific translational activators. Binding of these factors to the 5′-untranslated leader sequences of mitochondrial mRNAs is a prerequisite for initiation of the translation reaction (for a review see Fox, 1996). Several of these components are located at the matrix face of the inner membrane. We suggest that the close contact of ribosomes with the membrane then facilitates their binding to the Oxa1 complex via the C-terminal matrix domain of Oxa1. Alternatively, ribosomes might be tethered to Oxa1 even prior to the initiation process. The observation that more Oxa1 was found in association with ribosomes in the presence of nascent chains might be due to two reasons: (i) the interaction of Oxa1 with ribosomes might be dynamic and regulated by the activity of mitochondrial translation; and/or (ii) nascent polypeptides on the ribosomes might contribute to the association to Oxa1 thereby stabilizing the solubilized complex.
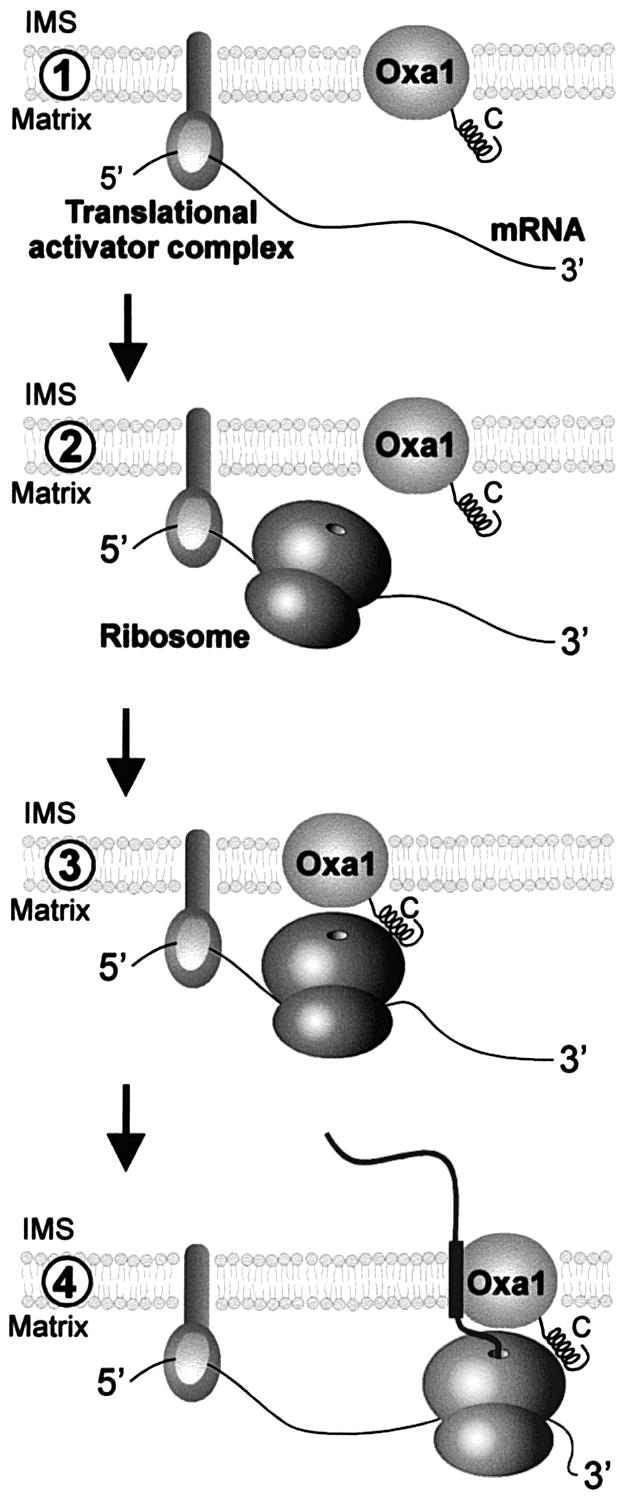
Fig. 9. Model of co-translational protein insertion into the inner membrane of mitochondria. (1) Membrane-bound translational activators bind to 5′-untranslated regions of mitochondrial mRNAs and recruit transcripts to the matrix face of the inner membrane. (2) Binding of translational activators is followed by initiation of translation. (3) The close proximity of the ribosomes to the membrane allows their binding to the Oxa1 translocase via the C-terminal matrix domain of Oxa1. Alternatively, ribosomes might already be associated with Oxa1 prior to translational initiation. (4) The association of the ribosome with the Oxa1 translocase facilitates co-translational insertion of nascent polypeptides. This physical coupling of translation and translocation reactions might support targeting of the translation products and/or prevent misfolding of the highly hydrophobic nascent polypeptides. IMS, intermembrane space.
In the last few years, several suppressor mutants were isolated which have the ability to grow on a non-fermentable carbon source in the absence of Oxa1 (Hamel et al., 1998; Saint-Georges et al., 2001). In these strains, membrane insertion of translation products must occur in an Oxa1-independent manner. Although these mutants are able to respire, the growth rates on glycerol medium are significantly lower than in the presence of Oxa1. In addition, a large fraction of newly synthesized Cox2 protein accumulated in its precursor form in the matrix following protein synthesis in mitochondria of these mutants (Saint-Georges et al., 2001; J.M.Herrmann, unpublished results). Thus, the loss of Oxa1 is only partially compensated by the suppressor mutations. Suppression of the defects of oxa1 mutants was dependent on the presence of the inner membrane protein Mba1 (Saint-Georges et al., 2001). Mba1 was proposed to function as a receptor component for mitochondrially encoded inner membrane proteins and it might cooperate in this respect with the C-terminal domain of Oxa1 (Preuss et al., 2001). Consistently, combination of mba1 deletion with the Oxa11–331 mutation led to a synergistic growth defect on non-fermentable carbon sources (not shown), supporting the idea of a partially overlapping function of Mba1 and the C-terminal matrix domain of Oxa1.
Co-translational association of cytosolic ribosomes with membranes of the endoplasmic reticulum has been known for a long time (Palade and Siekevitz, 1956). In recent years, biochemical experiments and microscopic examination revealed a direct binding of large ribosomal subunits to the Sec61 translocase (Prinz et al., 2000; Beckmann et al., 2001). The nascent polypeptides on the ribosomes most probably contribute to this interaction but ribosomes bind Sec61 even following removal of nascent chains by puromycin (Jungnickel and Rapoport, 1995; Prinz et al., 2000). As a consequence, large ribosomal subunits stay membrane associated following termination of protein synthesis (Seiser and Nicchitta, 2000). The ribosome and the Sec61 translocase thereby appear to form a continuous protein-conducting channel that might reduce the risk for nascent chains to misfold. The ability to bind ribosomes seems to be a conserved feature of Sec translocases. Escherichia coli ribosomes were shown to bind to the bacterial SecYEG complex in vitro (Prinz et al., 2000). In vivo, however, ribosomes were dissociated efficiently from bacterial membranes by puromycin treatment (Smith et al., 1978). This difference in the puromycin-insensitive binding of ribosomes to mitochondrial membranes (Liu and Spremulli, 2000) might be explained by the absence of the C-terminal ribosome-binding domain on the bacterial Oxa1 homolog YidC, which is in contact with the SecYEG complex during the process of protein insertion into the bacterial inner membrane (Scotti et al., 2000).
C-terminal matrix domains with high probabilities for coiled-coil formation are ubiquitously present in Oxa1 homologs of fungi, plants and animals. These domains typically contain patterns of positive charged residues, and an interaction of this helical domain with a protein or RNA component of the mitochondrial ribosomes is conceivable. These conserved structural characteristics of Oxa1 proteins suggest that the ability to bind to ribosomes might be a general property of this translocase in mitochondria. In a recent study, we reported that deletion of most of the Neurospora crassa Oxa1 C-terminal domain does not lead to significant growth defects of the fungus (Nargang et al., 2002). We assume that this is due to the presence of a second OXA1 gene in the Neurospora genome which we recently identified (our unpublished results).
A region similar in its amino acid sequence to the bacterial signal recognition particle subunit Ffh was found in the protein Mrpl36 which is part of the large ribosomal subunit of mitochondria (Bonnefoy et al., 2001). Interestingly, overexpression of Mrpl36 suppressed the membrane insertion defects of Cox2 variants containing mutations in their N-terminal leader peptides (Bonnefoy et al., 2001). This genetic interaction points to a role for this ribosomal protein in the process of Oxa1-dependent protein translocation. The generation of a ribosome-binding site on the Oxa1 translocase during evolution of mitochondria might have made a signal recognition particle unnecessary. Since mitochondrial translation products are almost entirely hydrophobic membrane proteins, a mechanism to differentiate between hydrophobic and soluble translation products might have become dispensable in mitochondria. In agreement with this hypothesis, it was reported recently that the only soluble protein synthesized in mitochondria, Var1, appears also to be produced in close proximity to the inner membrane (Fiori et al., 2003). Future studies will have to focus on the regulation of the interaction of Oxa1 with ribosomes and the identification of the Oxa1-binding site on the mitochondrial ribosome.
Materials and methods
Construction of strains and plasmids
For construction of GST–Oxa1, the sequence encoding the C-terminal 89 residues of Oxa1 was amplified from genomic S.cerevisiae DNA and cloned as described for GST–Ssq1 (Lutz et al., 2001). For generation of the Oxa11–331 strain, the codon 332 of the chromosomal OXA1 gene was changed to a stop codon by site-directed mutagenesis. Strain Oxa11–317 was constructed by insertion of a codon for lysine followed by a stop codon and a HIS3 cassette after codon 317 of the chromosomal OXA1 gene in strain YPH499 (Sikorski and Hieter, 1989). This mutant expressed a truncated version of Oxa1 consisting of residues 1–317 (arginine) followed by an additional lysine residue which was added to stabilize further the hydrophobic segment in the membrane. Generation of the oxa1 deletion mutant (Δoxa1) was described before (Hell et al., 1998). For in vitro translation of Oxa1C-DHFR, the sequence encoding the C-terminal 110 residues of Oxa1 was integrated into the BamHI site of pGEM3 vector (Promega) in front of the DNA encoding mouse dihydrofolate reductase (Herrmann et al., 1997). Due to reinitiation from the second methionine in the Oxa1 sequence, translation resulted in both Oxa1292–402-DHFR and Oxa1314–402-DHFR. The latter was used for the in vitro binding experiments.
Affinity chromatography with GST fusion proteins
GST fusion proteins were isolated from E.coli extracts as described (Lutz et al., 2001). Mitochondrial translation products were radiolabeled for 10 min at 30°C in 4 mg of wild-type mitochondria as described (Westermann et al., 2001). The sample was split and 40 µM puromycin was added to one half for 5 min. Mitochondria were reisolated and lysed in 1% digitonin, 30 mM MgSO4, 20% glycerol, 0.6 M sorbitol, 20 mM HEPES pH 7.4. The extract was cleared by centrifugation for 10 min at 18 000 g and divided into four aliquots. One aliquot was directly applied to an SDS–polyacrylamide gel. The other aliquots were incubated for 1 h at 4°C with 5 µg of purified GST, GST–Oxa1C or GST–Ssq1, respectively, that were immobilized on a glutathione–Sepharose resin. The beads were washed twice in lysis buffer and once with 20 mM Tris pH 7.4. Bound proteins were applied to an SDS–polyacrylamide gel and transferred to nitrocellulose. Following staining with Ponceau S, the nitrocellulose membrane was used for immunoblotting and autoradiography.
Isolation of a ribosomal protein fraction
For isolation of a ribosomal fraction, translation products were radiolabeled in wild-type or Oxa11–331 mitochondria (640 µg) for 5 min at 25°C. The reactions were split and further incubated in the absence or presence of puromycin for 25 min. The mitochondria were reisolated, washed in 0.6 M sorbitol, 80 mM KCl, 20 mM HEPES pH 7.5 and lysed in 2% digitonin, 20 mM Tris pH 7.4 for 20 min at 4°C in the presence of 10 mM MgSO4 or 5 mM EDTA. The extracts were cleared by centrifugation for 10 min at 18 000 g. To isolate a ribosomal pellet fraction, the sample was layered onto a high-density sucrose cushion consisting of 1.6 M sucrose, 20 mM Tris pH 7.4, and centrifuged for 1 h at 436 000 g.
To assess in vitro binding of the C-terminus of Oxa1 to ribosomes, mitochondrial extracts were incubated with urea-denatured Oxa1314–402-DHFR for 1 h at 4°C prior to the isolation of mitochondrial ribosomes.
For fractionation of ribosomes on sucrose gradients, translation products were radiolabeled for 30 min in 10 mg of freshly isolated mitochondria. The mitochondria were washed and lysed as above with the exception that 5 mM dithiothreitol (DTT) was added to the lysis buffer. The extract was layered onto a continuous 12 ml sucrose gradient [15–40% sucrose (w/v) in 0.1% digitonin, 5 mM DTT, 5 mM MgSO4, 20 mM Tris pH 7.4] and centrifuged at 83 000 g in a Beckman SW41 rotor for 16 h at 2°C. Then 16 fractions of 750 µl were collected and proteins were precipitated by addition of 12% trichloroacetic acid. The resulting pellets were dissolved in sample buffer and analyzed by immunoblotting and autoradiography.
Assessment of the mitochondrial inner membrane potential
The membrane potential of isolated mitochondria was measured in 0.6 M sorbitol containing 0.1% (w/v) bovine serum albumin (fatty acid free), 0.5 mM EDTA, 20 mM potassium phosphate pH 7.2 and 2 mM NADH at 25°C essentially as described in Hell et al. (1997). 3,3′-Dipropyl thiadicorbocyanine iodide, DiSC3(5), was added to a final concentration of 2 µM. After 0.5 min, mitochondria were added to a final mitochondrial protein concentration of 33 µg/ml. After 2 min of incubation, 3 µl of 1 M KCN and 3 µl of 10 mM valinomycin were added to dissipate the membrane potential. The increase of the fluorescence indicates dissipation of the membrane potential.
Miscellaneous
Import of in vitro synthesized proteins into isolated mitochondria was carried out essentially as described (Herrmann et al., 1997), with the exception that during the import reaction 2 mM ATP, 2 mM NADH, 1 mM creatine phosphate, 0.1 mg/ml creatine kinase, 2.5 mM malate and 2.5 mM succinate were added to warrant high membrane potential levels in the mitochondria. Subfractionation of mitochondria following import and in organello translation assays (Preuss et al., 2001), urea extraction (Borel and Simon, 1996), size exclusion chromatography of the Oxa1 complex (Preuss et al., 2001) and enzymatic measurement of the activities of the cytochrome bc1, cytochrome oxidase and ATP synthase complexes were performed as described before (Tzagoloff et al., 1975).
Acknowledgments
Acknowledgements
This work is dedicated to the memory of Gregor Szyrach who tragically died on June 17, 2003. We thank W.Voelter and A.Kapurniotu for assistance with the CD spectroscopy, K.Hell for the Cox18 expression plasmid, T.Fox for the Mrps51 antiserum, K.Nierhaus for antibodies against E.coli ribosomal proteins, T. Langer for the antiserum against the N-terminus of Oxa1, B.Westermann and F.Nargang for reading the manuscript, and the Deutsche Forschungsgemeinschaft (SFB 594) for financial support.
References
- Bauer M., Behrens,M., Esser,K., Michaelis,G. and Pratje,E. (1994) PET1402, a nuclear gene required for proteolytic processing of cytochrome oxidase subunit 2 in yeast. Mol. Gen. Genet., 245, 272–278. [DOI] [PubMed] [Google Scholar]
- Beckmann R., Spahn,C.M., Eswar,N., Helmers,J., Penczek,P.A., Sali,A., Frank,J. and Blobel,G. (2001) Architecture of the protein-conducting channel associated with the translating 80S ribosome. Cell, 107, 361–372. [DOI] [PubMed] [Google Scholar]
- Bonnefoy N., Chalvet,F., Hamel,P., Slominski,P.P. and Dujardin,G. (1994) OXA1, a Saccharomyces cerevisiae nuclear gene whose sequence is conserved from prokaryotes to eukaryotes controls cytochrome oxidase biogenesis. J. Mol. Biol., 239, 201–212. [DOI] [PubMed] [Google Scholar]
- Bonnefoy N., Bsat,N. and Fox,T.D. (2001) Mitochondrial translation of Saccharomyces cerevisiae COX2 mRNA is controlled by the nucleotide sequence specifying the pre-Cox2p leader peptide. Mol. Cell. Biol., 21, 2359–2372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borel A.C. and Simon,S.M. (1996) Biogenesis of polytopic membrane proteins: membrane segments assemble within translocation channels prior to membrane integration. Cell, 85, 379–389. [DOI] [PubMed] [Google Scholar]
- Burkhard P., Stetefeld,J. and Strelkov,S.V. (2001) Coiled coils: a highly versatile protein folding motif. Trends Cell Biol., 11, 82–88. [DOI] [PubMed] [Google Scholar]
- Fearon K., McClendon,V., Bonetti,B. and Bedwell,D.M. (1994) Premature translation termination mutations are efficiently suppressed in a highly conserved region of yeast Ste6p, a member of the ATP-binding cassette (ABC) transporter family. J. Biol. Chem., 269, 17802–17808. [PubMed] [Google Scholar]
- Fiori A., Mason,T.L. and Fox,T.D. (2003) Evidence that synthesis of the Saccharomyces cerevisiae mitochondrially encoded ribosomal protein Var1p may be membrane localized. Eukaryot. Cell, 2, 651–653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox T.D. (1996) Genetics of mitochondrial translation. In Hershey,J.W.B., Matthews,M.B. and Sonnenberg,N. (eds), Translational Control. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp. 733–758. [Google Scholar]
- Greenfield N. and Fasman,G.D. (1969) Computed circular dichroism spectra for the evaluation of protein conformation. Biochemistry, 8, 4108–4116. [DOI] [PubMed] [Google Scholar]
- Hamel P., Lemaire,C., Bonnefoy,N., Brivet-Chevillotte,P. and Dujardin,G. (1998) Mutations in the membrane anchor of yeast cytochrome c1 compensate for the absence of Oxa1p and generate cabonate-extractable forms of cytochrome c1. Genetics, 150, 601–611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He S. and Fox,T.D. (1997) Membrane translocation of mitochondrially coded Cox2p: distinct requirements for export of N and C termini and dependence on the conserved protein Oxa1p. Mol. Biol. Cell, 8, 1449–1460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hell K., Herrmann,J., Pratje,E., Neupert,W. and Stuart,R.A. (1997) Oxa1p mediates the export of the N- and C-termini of pCoxII from the mitochondrial matrix to the intermembrane space. FEBS Lett., 418, 367–370. [DOI] [PubMed] [Google Scholar]
- Hell K., Herrmann,J.M., Pratje,E., Neupert,W. and Stuart,R.A. (1998) Oxa1p, an essential component of the N-tail protein export machinery in mitochondria. Proc. Natl Acad. Sci. USA, 95, 2250–2255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hell K., Neupert,W. and Stuart,R.A. (2001) Oxa1p acts as a general membrane insertion machinery for proteins encoded by mitochondrial DNA. EMBO J., 20, 1281–1288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herrmann J.M. (2003) Converting bacteria to organelles: evolution of mitochondrial protein sorting. Trends Microbiol., 11, 74–79. [DOI] [PubMed] [Google Scholar]
- Herrmann J.M., Koll,H., Cook,R.A., Neupert,W. and Stuart,R.A. (1995) Topogenesis of cytochrome oxidase subunit II—mechanisms of protein export from the mitochondrial matrix. J. Biol. Chem., 270, 27079–27086. [DOI] [PubMed] [Google Scholar]
- Herrmann J.M., Neupert,W. and Stuart,R.A. (1997) Insertion into the mitochondrial inner membrane of a polytopic protein, the nuclear encoded Oxa1p. EMBO J., 16, 2217–2226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia L., Dienhart,M., Schramp,M., McCauley,M., Hill,K. and Stuart,R.A. (2003) Yeast Oxa1 interacts with mitochondrial ribosomes: the importance of the C-terminal region of Oxa1. EMBO J., 22, 6438–6447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jungnickel B. and Rapoport,T.A. (1995) A posttargeting signal sequence recognition event in the endoplasmic reticulum membrane. Cell, 82, 261–270. [DOI] [PubMed] [Google Scholar]
- Kayed R., Bernhagen,J., Greenfield,N., Sweimeh,K., Brunner,H., Voelter,W. and Kapurniotu,A. (1999) Conformational transitions of islet amyloid polypeptide (IAPP) in amyloid formation in vitro. J. Mol. Biol., 287, 781–796. [DOI] [PubMed] [Google Scholar]
- Liu M. and Spremulli,L. (2000) Interaction of mammalian mitochondrial ribosomes with the inner membrane. J. Biol. Chem., 275, 29400–29406. [DOI] [PubMed] [Google Scholar]
- Luirink J., Samuelsson,T. and de Gier,J.W. (2001) YidC/Oxa1p/Alb3: evolutionarily conserved mediators of membrane protein assembly. FEBS Lett., 501, 1–5. [DOI] [PubMed] [Google Scholar]
- Lupas A. (1997) Predicting coiled-coil regions in proteins. Curr. Opin. Struct. Biol., 7, 388–393. [DOI] [PubMed] [Google Scholar]
- Lutz T., Westermann,B., Neupert,W. and Herrmann,J.M. (2001) The mitochondrial proteins Ssq1 and Jac1 are required for the assembly of iron sulfur clusters in mitochondria. J. Mol. Biol., 307, 815–825. [DOI] [PubMed] [Google Scholar]
- Nargang F.E., Preuss,M., Neupert,W. and Herrmann,J.M. (2002) The Oxa1 protein forms a homooligomeric complex and is an essential part of the mitochondrial export translocase in Neurospora crassa. J. Biol. Chem., 277, 12846–12853. [DOI] [PubMed] [Google Scholar]
- Palade G. and Siekevitz,P. (1956) Liver microsomes: an integrated morphological and biochemical study. J. Biophys. Biochem. Cytol., 2, 171–201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Preuss M., Leonhard,K., Hell,K., Stuart,R.A., Neupert,W. and Herrmann,J.M. (2001) Mba1, a novel component of the mitochondrial protein export machinery of the yeast Saccharomyces cerevisiae. J. Cell Biol., 153, 1085–1096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prinz A., Behrens,C., Rapoport,T.A., Hartmann,E. and Kalies,K.U. (2000) Evolutionarily conserved binding of ribosomes to the translocation channel via the large ribosomal RNA. EMBO J., 19, 1900–1906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rojo E.E., Stuart,R.A. and Neupert,W. (1995) Conservative sorting of F0-ATPase subunit 9: export from matrix requires ΔpH across inner membrane and matrix ATP. EMBO J., 14, 3445–3451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saint-Georges Y., Hamel,P., Lemaire,C. and Dujardin,G. (2001) Role of positively charged transmembrane segments in the insertion and assembly of mitochondrial inner-membrane proteins. Proc. Natl Acad. Sci. USA, 98, 13814–13819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scotti P.A., Urbanus,M.L., Brunner,J., de Gier,J.L., von Heijne,G., van der Does,C., Driessen,A.J.M., Oudega,B. and Luirink,L. (2000) YidC, the Escherichia coli homologue of mitochondrial Oxa1p, is a component of the Sec translocase. EMBO J., 19, 542–549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seiser R.M. and Nicchitta,C.V. (2000) The fate of membrane-bound ribosomes following the termination of protein synthesis. J. Biol. Chem., 275, 33820–33827. [DOI] [PubMed] [Google Scholar]
- Sevarino K.A. and Poyton,R.O. (1980) Mitochondrial membrane biogenesis: identification of a precursor to yeast cytochrome c oxidase subunit II, an integral polypeptide. Proc. Natl Acad. Sci. USA, 77, 142–146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski R.S. and Hieter,P. (1989) A system of shuttle vectors and host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics, 122, 19–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith W.P., Tai,P.C. and Davis,B.D. (1978) Nascent peptide as sole attachment of polysomes to membranes in bacteria. Proc. Natl Acad. Sci. USA, 75, 814–817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tzagoloff A., Akai,A. and Needleman,R.B. (1975) Assembly of the mitochondrial membrane system. J. Biol. Chem., 250, 8228–8235. [PubMed] [Google Scholar]
- Watson K. (1972) The organization of ribosomal granules within mitochondrial structures of aerobic and anaerobic cells of Saccharomyces cerevisae. J. Cell Biol., 55, 721–726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westermann B., Herrmann,J.M. and Neupert,W. (2001) Analysis of mitochondrial translation products in vivo and in organello in yeast. Methods Cell Biol., 65, 429–438. [DOI] [PubMed] [Google Scholar]