Abstract
The maintenance of genome integrity requires a rapid and specific response to many types of DNA damage. The conserved and related PI3-like protein kinases, ataxia-telangiectasia mutated (ATM) and ATM-Rad3-related (ATR), orchestrate signal transduction pathways in response to genomic insults, such as DNA double-strand breaks (DSBs). It is unclear which proteins recognize DSBs and activate these pathways, but the Mre11/Rad50/NBS1 complex has been suggested to act as a damage sensor. Here we show that infection with an adenovirus lacking the E4 region also induces a cellular DNA damage response, with activation of ATM and ATR. Wild-type virus blocks this signaling through degradation of the Mre11 complex by the viral E1b55K/E4orf6 proteins. Using these viral proteins, we show that the Mre11 complex is required for both ATM activation and the ATM-dependent G2/M checkpoint in response to DSBs. These results demonstrate that the Mre11 complex can function as a damage sensor upstream of ATM/ATR signaling in mammalian cells.
Keywords: adenovirus/ATM/ATR/DNA double-strand breaks/Mre11 complex
Introduction
A central player in the cellular response to DSBs is the Mre11 complex, consisting of Mre11, Rad50 and NBS1 (Xrs2 in yeast) (Petrini, 1999; D’Amours and Jackson, 2002). Hypomorphic mutations in the genes for Mre11 and NBS1 result in the human genetic instability diseases ataxia-telangiectasia like disorder (A-TLD) and Nijmegen breakage syndrome (NBS), respectively (Carney et al., 1998; Stewart et al., 1999). Cell lines derived from these patients are sensitive to ionizing radiation (IR) and exhibit radioresistant DNA synthesis (RDS). Cytologic and genetic evidence suggests that the Mre11 complex acts as a DNA damage sensor (Mirzoeva and Petrini, 2001; Usui et al., 2001), but definitive evidence in mammalian cells has thus far remained elusive.
The related PI3-like kinases, ataxia-telangiectasia mutated (ATM) and ATM-Rad3-related (ATR), are signal transducers of the DNA damage response. Mutations in ATM lead to ataxia-telangiectasia (A-T), and cells from these patients are hypersensitive to DSBs and defective for induction of the G1/S, intra-S and G2/M cell-cycle checkpoints (Shiloh, 2003). Null mutations of ATR are lethal, but the protein has been studied using a conditional knockout allele or dominant-negative kinase dead mutants, which sensitize cells to all forms of DNA damage and affect the intra-S and G2/M checkpoint responses (Cliby et al., 1998; Nghiem et al., 2001; Brown and Baltimore, 2003). A multitude of DNA repair and checkpoint proteins have been identified as substrates for ATM and ATR kinase activity, including the checkpoint kinases Chk1 and Chk2, as well as 53BP1 (DiTullio et al., 2002; Fernandez-Capetillo et al., 2002), H2AX (Paull et al., 2000), BRCA1, p53 and RPA32 (reviewed in Kastan and Lim, 2000; Shiloh, 2003). Both Mre11 and NBS1 are also targets of ATM and possibly ATR (Gatei et al., 2000; Lim et al., 2000; Wu et al., 2000; Zhao et al., 2000; Costanzo et al., 2001; D’Amours and Jackson, 2001). Recent studies have begun to elucidate the cellular requirements for the activation of ATM and ATR upon DNA damage. It has been proposed that ATM is activated by intermolecular autophosphorylation on Ser1981 (Bakkenist and Kastan, 2003). In the case of ATR it has been suggested that RPA bound to single-stranded DNA is required for recruitment of ATR to sites of DNA damage (Zou and Elledge, 2003). The Mre11 complex has been proposed to facilitate ATM activation and functionally interact with ATR (Usui et al., 2001; Williams et al., 2002), but knowledge of molecular mechanisms to support these links is lacking in mammalian cells.
We have demonstrated that adenovirus (Ad) avoids detrimental genome concatemerization through degradation of the Mre11 complex by the viral E1b55K/E4orf6 proteins (Stracker et al., 2002). Here we show for the first time that infection with an adenovirus lacking the E4 region induces a cellular DNA damage response that requires ATM and ATR. We identified mutants of E1b55K that can selectively degrade different substrates. Using these E1b55K mutants we show that the Mre11 complex is required for the activation of ATM and ATR in response to Ad infection. In addition, we show that Mre11 degradation abrogates ATM autophosphorylation and signaling as well as the ATM-dependent G2/M checkpoint in response to DSBs. Together, these results support a role for the mammalian Mre11 complex in sensing DSBs and adenoviral infection.
Results
E4-deleted adenovirus elicits a cellular DNA damage response
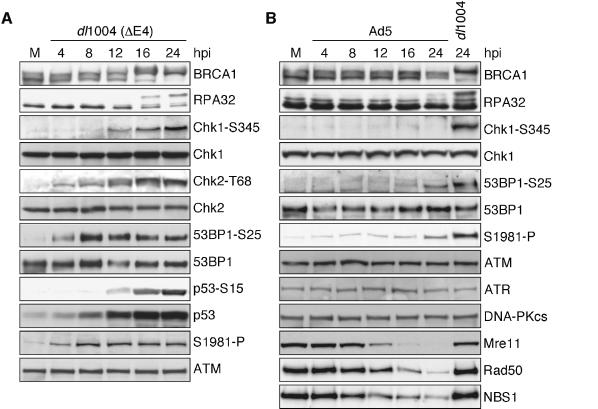
Infection with an E4-deleted Ad results in end-joining of viral genomes by cellular factors and this is accompanied by hyperphosphorylation of NBS1, which is indicative of a cellular DNA damage response (Stracker et al., 2002). To investigate this further, we examined whether other damage-response proteins become phosphorylated upon infection with the E4-deleted Ad dl1004 (Figure 1A). Immunoblotting of cell lysates over a time-course of infection revealed slower migrating forms of BRCA1 and RPA32. Antibodies that recognize specific phosphorylated residues demonstrated phosphorylation of Chk1, Chk2, 53BP1, p53 and ATM. The p53 protein also appeared to be stabilized, which may be due to E1a interference with proteasomes or the E6 protein expressed in the HeLa cells. These results indicate that infection with an E4-deleted virus leads to ATM activation and signaling to proteins involved in the cellular DNA damage response.
Fig. 1. Infection with the E4-deleted virus dl1004 activates signaling pathways involved in the cellular DNA damage response, but does not affect the steady-state levels of PI3-like kinases. (A) HeLa cells were mock infected (M) or infected with dl1004 for the indicated times and harvested for immunoblotting. (B) HeLa cells were mock infected (M), infected with dl1004 or Ad5 for the indicated times and harvested for immunoblotting.
These phosphorylation events were not detected to a significant level during infection with wild-type Ad (Figure 1B), suggesting that the wild-type virus can prevent activation of this cellular response. At late time points we observed slight activation of ATM and phosphorylation of 53BP1 and Chk2 (data not shown), but these signals were significantly lower than those observed during infection with the E4-deleted virus. Many of the phosphorylation events that we examined were on substrates of the PI3-like kinases, ATM, ATR and DNA-PKcs, and therefore we tested whether the virus indirectly affects the activities of these kinases by altering their steady-state levels (Figure 1B). We did not observe any alteration in the levels of ATM, ATR and DNA-PKcs over a 24 h time-course of infection with wild-type adenovirus type 5 (Ad5). In contrast, the Mre11 complex was degraded as the infection progressed.
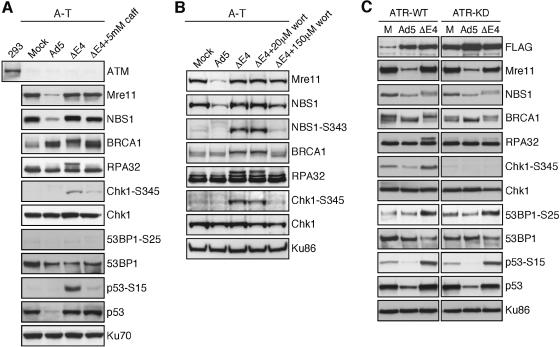
We next sought to determine the roles of ATM and ATR in the DNA damage response to E4-deleted adenovirus. Phosphorylation of substrates during infection with the E4-deleted virus dl1004 was investigated in cells deficient for ATM or ATR kinase activity. In ATM-deficient cells, phosphorylation of 53BP1 on Ser25 was abrogated, but all other phosphorylation events were detected (Figure 2A). These events were ablated by 5 mM caffeine, a dose that inhibits the kinase activity of both ATM and ATR in vitro (Sarkaria et al., 1999). These data suggest that while ATM signaling was not absolutely required, a caffeine-sensitive kinase was involved. We also performed the infections in A-T cells in the presence of 20 and 150 µM wortmannin (Figure 2B), which inhibits ATM and DNA-PKcs at the lower concentration and ATR at the higher concentration. We observed that treatment with 150 µM, but not 20 µM wortmannin prevented the phosphorylation events, suggesting that ATR may be involved in this signaling. The role of ATR was further examined using a cell line that expresses a doxycycline-inducible trans-dominant negative mutant (Nghiem et al., 2001). Infection with the E4-deleted virus led to phosphorylation of protein substrates in control cells (ATR-WT), but induction of the trans-dominant protein (ATR-KD) prevented phosphorylation of Chk1 on Ser345, and blocked hyperphosphorylation of RPA32 (Figure 2C). These phosphorylation events were not detected in either cell line during infection with wild-type Ad. These results suggest that ATM and ATR or another kinase sensitive to both caffeine and wortmannin may be redundant for the phosphorylation of damage response proteins during viral infection.
Fig. 2. ATM and ATR are activated by infection with E4-deleted adenovirus. Phosphorylation of cellular proteins was examined by immunoblotting of lysates from A-T cells (GM5849) (A and B) and cells expressing inducible ATR proteins (C), either wild-type (WT) or kinase-dead (KD) after doxycycline treatment. In each case cells were uninfected (M), infected with wild-type Ad5 or infected with the E4-deleted virus dl1004 (ΔE4). The infections in A-T cells were also performed in the presence of 5 mM caffeine (A) or 20 µM and 150 µM wortmannin (B) throughout infection. A-T cells were infected for 30 h and U2OS cells expressing ATR were infected for 24 h. Ku70 and Ku86 served as loading controls.
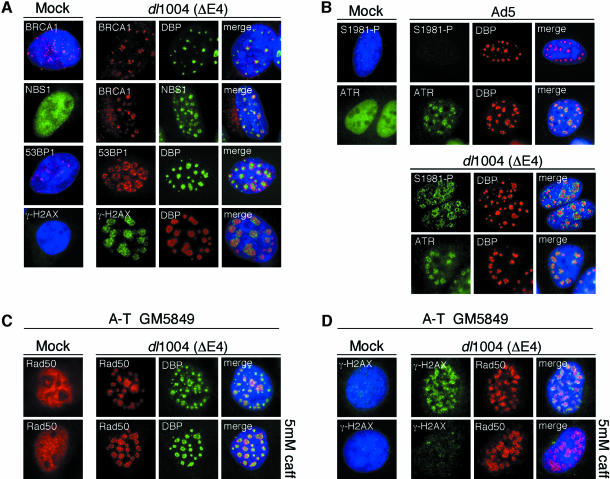
Many proteins involved in the DNA damage response accumulate in foci at sites of DSBs (Rouse and Jackson, 2002). In a similar fashion, the Mre11 complex accumulates at foci surrounding viral replication centers formed during infection with E4-deleted adenovirus (Stracker et al., 2002). Using immunofluorescence staining we examined whether other damage proteins also accumulated at viral centers during infection. Viral replication centers were identified with an antibody to the viral DNA binding protein (DBP), and we observed foci of BRCA1, 53BP1 and γ-H2AX (the phosphorylated form of H2AX) at these discrete sites (Figure 3A). We also examined localization of the ATM and ATR kinases (Figure 3B). Staining with a phosphospecific antibody to Ser1981 of ATM demonstrated accumulation of activated ATM at E4-deleted (dl1004) viral centers. This suggests that ATM is activated in the absence of E4 and accumulates at sites of virus replication. In contrast, ATM autophosphorylation was not observed in Ad5-infected cells (Figure 3B), although some staining was visible at late stages of infection (data not shown). Infection with both wild-type Ad5 and the E4-deleted virus resulted in accumulation of ATR, ATRIP and RPA at viral replication centers (Figure 3B and data not shown). These data suggest that ATR localization is independent of activation of the DNA damage response and the Mre11 complex (as the complex is degraded in wild-type Ad5 infection).
Fig. 3. DNA repair proteins accumulate at sites of viral replication during infection with a virus lacking E4. (A) U2OS cells were untreated (Mock) or infected with the E4-deleted virus dl1004 (m.o.i. = 25), and proteins were visualized at 23 h.p.i. by immunofluorescence. Viral replication centers were localized by staining for DBP. (B) Localization of kinases during virus infection. U20S cells were infected with wild-type Ad5 or dl1004, and stained with antibodies specific to the autophosphorylated site at S1981 of ATM or ATR. (C) Formation of Rad50 foci at viral replication centers is independent of signaling events and is caffeine resistant. The A-T cells were infected with the dl1004 virus in the presence and absence of 5 mM caffeine and harvested 30 h.p.i. (D) Foci of γ-H2AX at viral replication centers are sensitive to caffeine and are not required for Rad50 foci. The A-T cells were infected with the dl1004 virus as in (C).
We examined whether accumulation of the Mre11 complex at viral centers was dependent on the kinase activities of ATM/ATR. A-T cells were infected with the E4-deleted virus in the presence or absence of 5 mM caffeine. Recruitment of Rad50 to foci at viral replication centers occurred even after caffeine treatment of A-T cells (Figure 3C). We also examined the formation of γ-H2AX foci at viral centers (Figure 3D). Foci of γ-H2AX were observed at viral centers in A-T cells, but in contrast to Rad50 foci, the accumulation of γ-H2AX was abrogated in the presence of caffeine. This indicates that foci formation by the Mre11 complex is resistant to caffeine and is independent of γ-H2AX foci, consistent with previous reports showing that accumulation of the Mre11 complex at sites of damage is independent of ATM and γ-H2AX in response to DSBs (Mirzoeva and Petrini, 2001; Celeste et al., 2003). Together, these data suggest that the Mre11 complex can function upstream of, or coincident with, ATM and ATR activation.
It has previously been demonstrated that the ATM protein becomes associated with chromatin upon DNA damage (Andegeko et al., 2001). This was shown using a biochemical fractionation procedure in which a portion of the ATM pool was found to be resistant to detergent extraction after treating cells with agents that lead to DSBs (Andegeko et al., 2001). We used this assay to assess whether ATM retention also occurred during virus infection (supplementary figure 1, available at The EMBO Journal Online). Cells infected with either wild-type Ad5 or the E4-deleted virus dl1004 were harvested and fractionated by successive detergent extractions. Immunoblotting of cell-equivalent aliquots showed that the amount of loosely bound ATM removed by the first detergent extraction (FI) was similar between the samples. However, infection with the E4-deleted virus but not wild-type Ad5 led to a portion of the ATM protein being retained in fraction III (FIII). This assay supports the conclusion that infection with an E4-deleted adenovirus elicits a DNA damage response.
E1b55K/E4orf6 target the Mre11 complex for degradation
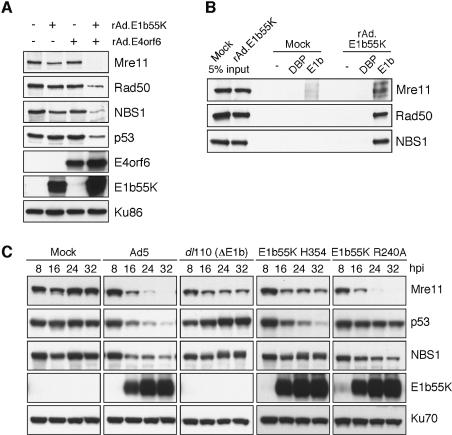
The data thus far demonstrated that removal of the Mre11 complex by wild-type adenovirus correlated with prevention of the DNA damage response to infection. We further characterized the mechanism and specificity of Mre11 complex degradation by the viral E1b55K/E4orf6 proteins. In order to study degradation induced by E1b55K and E4orf6 in multiple cell lines, we expressed them individually using replication-deficient, recombinant, E1-deleted adenovirus vectors (rAd.E1b55K and rAd.E4orf6). Degradation of Mre11, Rad50 and NBS1 proteins was only observed in cells infected with both vectors, whereas either vector alone had minimal effect (Figure 4A). Degradation of p53 was also observed, as previously reported (Cathomen and Weitzman, 2000; Querido et al., 2001a). These results show that the E1b55K/E4orf6 complex is sufficient for inducing degradation.
Fig. 4. The adenoviral E1b55K protein is important for substrate recognition and degradation of the Mre11 complex. (A) The E1b55K and E4orf6 proteins are required for degradation of the Mre11 complex. U2OS cells were untreated or infected with E1-deleted recombinant Ad vectors expressing E1b55K and E4orf6 alone or in combination (m.o.i. = 10 and 50 p.f.u., respectively). Cells were harvested at 30 h.p.i. for immunoblotting. Ku86 served as a loading control. (B) The E1b55K protein interacts with the Mre11 complex. U2OS cells were either untreated (Mock) or infected with rAd.E1b55K for 15 h. Cell lysates were immunoprecipitated with an antibody to E1b55K or a control antibody to adenovirus DBP. Immunoblotting is shown for the lysates (5% of input) or the precipitates. (C) Mutations in E1b55K separate degradation of p53 and the Mre11 complex. U2OS cells were infected with wild-type Ad5, a virus deleted of E1b55K (dl110) and two viruses expressing mutant E1b55K proteins. Cells were harvested at the indicated times for immunoblotting. Ku70 served as a loading control.
The E1b55K/E4orf6 complex has been shown to recruit cellular factors and function as a von Hippel-Lindau (VHL)-like ubiquitin ligase to mediate polyubiquitination of the p53 protein in vitro (Querido et al., 2001a; Harada et al., 2002). In the case of p53 it has been demonstrated that degradation requires E1b55K binding to the N-terminus (Cathomen and Weitzman, 2000; Querido et al., 2001b). Therefore we explored whether there is an interaction between E1b55K and the Mre11 complex. Immunoprecipitation of E1b55K from cells infected with rAd.E1b55K was followed by immunoblotting, and revealed pull-down of the Mre11, Rad50 and NBS1 proteins but not ATM (Figure 4B and data not shown). The reciprocal immunoprecipitations of NBS1 and Rad50 also pulled down E1b55K (supplementary figure 2). These data demonstrate that the E1b55K protein interacts with the Mre11 complex.
Degradation and substrate recognition was investigated further by screening a panel of viruses that express mutant forms of the E1b55K protein (R.A.S. and M.D.W., unpublished data). The E1b55K viruses that we tested harbor point mutations, linker insertions or large deletions, and have previously been well characterized for growth (Yew et al., 1990; Shen et al., 2001). We examined the effect of infection with these mutant viruses on the levels of p53 and Mre11 and compared them with infection with wild-type Ad5 and the E1b55K-deleted virus dl110 that does not degrade either protein. We identified two mutants that showed differential substrate specificity for Mre11 or p53 (Figure 4C). The E1b55K H354 mutant (Yew et al., 1990) degrades p53 but not the Mre11 complex, while the E1b55K R240A virus (Shen et al., 2001) degrades the Mre11 complex but not p53, demonstrating that E1b55K is involved in substrate recognition and that specificity for the Mre11 complex or p53 is separable.
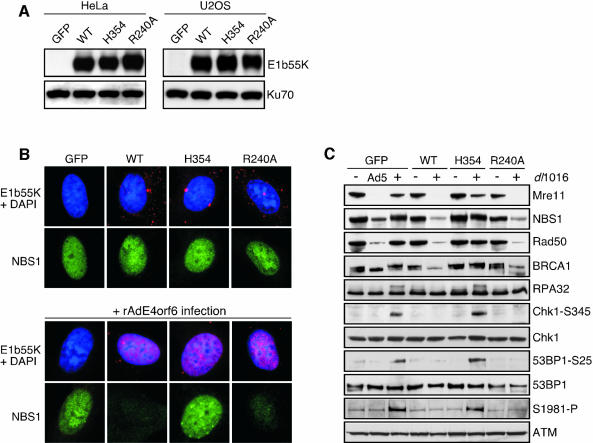
In order to study the effects of the mutant E1b55K proteins on signaling events in the absence of other viral proteins, we generated stable HeLa and U2OS cell lines expressing each mutant E1b55k protein from retrovirus vectors. The E1b55K proteins were stably expressed at similar levels in these cell lines (Figure 5A) and immunofluorescence revealed the wild-type and mutant E1b55K proteins at speckles in the cytoplasm (Figure 5B), consistent with previous reports of E1b55K localization (Zantema et al., 1985). Co-staining showed that the localization and levels of NBS1 were unaffected by E1b55K expression in these cells (Figure 5B). Co-expression of E4orf6 results in nuclear accumulation of E1b55K (Goodrum et al., 1996; Dobbelstein et al., 1997; Cathomen and Weitzman, 2000). The stable cell lines were infected with the rAd.E4orf6 virus, and in each case the E1b55K protein was found relocated to the nucleus in a predominantly diffuse pattern (Figure 5B). This indicates that the wild-type and mutant proteins all interact with E4orf6, consistent with published data from immunoprecipitation experiments (Rubenwolf et al., 1997; Shen et al., 2001). In the case of wild-type E1b55K and the R240A mutant, E4orf6 expression and nuclear accumulation were accompanied by degradation of Mre11, Rad50 and NBS1 as revealed by both immunofluorescence and immunoblotting (Figure 5B and C). The steady-state levels of members of the Mre11 complex were unaffected in cells expressing the H354 mutant or in a control cell line expressing green fluorescent protein (GFP). Similar results were observed for both the U2OS- and HeLa-derived cell lines. These cell lines thus provide a useful and controlled system to study cellular processes in the absence of the Mre11 complex.
Fig. 5. Degradation of the Mre11 complex prevents activation of the cellular DNA damage response during adenovirus infection. (A) Stable cell lines expressing E1b55K proteins. The E1b55K coding regions from wild-type and mutant viruses were cloned into retrovirus vectors and used to make stable cell lines. A GFP-expressing retrovirus was used as a control. Expression of E1b55K in HeLa and U2OS-derived cell lines was confirmed by immunoblotting. Ku70 served as a loading control. (B) E4orf6 recruits E1b55K into the nucleus and NBS1 is degraded. The U2OS cell lines described in (A) were uninfected (upper panels) or infected with rAd.E4orf6 (lower panels), and immunostained with antibodies to E1b55K and NBS1. Similar results were seen in the HeLa-derived cell lines. (C) Immunoblotting reveals that degradation of the Mre11 complex prevents cellular responses to viral infection. Cell lines derived from U2OS that express GFP or E1b55K proteins were infected with the virus dl1016 that is mutated for both E1b55K and E4orf3 genes. Infection of the GFP cell line with Ad5 served as a positive control. Cells were harvested at 30 h.p.i.
Mre11 degradation prevents ATM and ATR signaling in response to viral infection
We tested whether degradation of the Mre11 complex was essential in order to prevent the cellular damage response to virus infection. The U2OS cell lines expressing GFP or the E1b55K proteins were infected with an adenovirus mutant (dl1016) that was deleted for E1b55K but expressed functional E4orf6 (Figure 5C). This virus was used because it has an additional deletion in E4orf3, a gene that we previously showed to mislocalize and inactivate the Mre11 complex (Stracker et al., 2002). Infection of the GFP control cells and those expressing the H354 mutant did not result in degradation of Mre11, Rad50 and NBS1, and was accompanied by phosphorylation of ATM and ATM/ATR substrates, including NBS1, BRCA1, RPA32, Chk1 and 53BP1. Infection of cells expressing wild-type E1b55K (WT) and the R240A mutant resulted in degradation of the Mre11 complex, and there was no evidence of ATM activation or the phosphorylation of ATM/ATR substrates after dl1016 infection. This shows that targeted degradation of the Mre11 complex, and not p53, is important to prevent activation of ATM/ATR in response to viral infection. Similar results were observed for the HeLa-derived cell lines (data not shown).
Mre11 degradation prevents ATM activation and signaling in response to DSBs
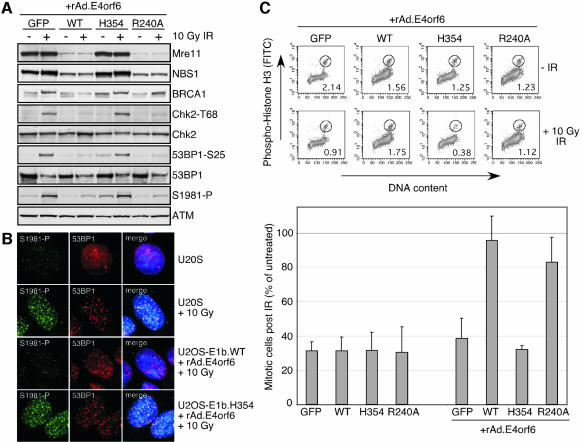
We hypothesized that by degrading the Mre11 complex, adenoviral E1b55K/E4orf6 proteins could prevent activation of signaling cascades induced by other forms of exogenous DNA damage. We therefore tested the effect of E1b55K/E4orf6 proteins on the cellular DNA damage response to IR. Cells expressing GFP or E1b55K proteins were infected with the recombinant vector rAd.E4orf6, prior to treatment with 10 Gy of IR. Expression of E4orf6 in cells containing wild-type E1b55K or the R240A mutant led to the degradation of the Mre11 complex, accompanied by greatly reduced ATM autophosphorylation and decreased phosphorylation of the ATM substrates BRCA1, Chk2 and 53BP1 (Figure 6A). In contrast, E4orf6 expression in cells containing the H354 mutant failed to degrade the Mre11 complex and had no effect on the damage response. Similar results were observed in the HeLa-derived cell lines (data not shown). In addition, foci of autophosphorylated ATM were detected after IR, but were absent with prior degradation of the Mre11 complex by E1b55K/E4orf6 (Figure 6B). These results suggest that the Mre11 complex is required for ATM activation and signaling after DSBs and that E1b55K/E4orf6 prevents the response by targeting the Mre11 complex for degradation.
Fig. 6. Degradation of the Mre11 complex by E1b55K/E4orf6 prevents the cell from activating a DNA damage response and the G2/M checkpoint after IR. (A) Degradation of the Mre11 complex prevents ATM activation and signaling upon IR. The stable U2OS cells lines were infected with rAd.E4orf6 for 24 h prior to 10 Gy IR, and were harvested for immunoblotting after a further 2 h. (B) Degradation of Mre11 prevents formation of autophosphorylated ATM foci in response to IR. Cells were either uninfected or infected with rAd.E4orf6, and then exposed to irradiation 1 h before immunofluorescence. (C) Degradation of the Mre11 complex abrogates the ATM-dependent early G2/M checkpoint in response to IR. Representative flow cytometry profiles of rAd.E4orf6-infected cells 1 h post-treatment, with or without 10 Gy IR. Cells were stained for DNA content (x-axis) and histone H3 phosphorylation (y-axis). The population of cells in mitosis is encircled and its percentage of the total cells is indicated. The change in the number of mitotic cells 1 h after IR is indicated in the graph below, in which the number of mitotic cells is presented as a percentage of the number detected in unirradiated samples of the same condition. The mean and standard deviation are shown for the results of three independent experiments for each condition.
Mre11 degradation abrogates the ATM-dependent early G2/M checkpoint
ATM signaling has been shown to be required for the early G2/M checkpoint in response to IR (Xu et al., 2002). This transient checkpoint is activated in cells that were in G2 when they incurred damage and prevents their progression into mitosis (M). Cells with functional ATM are prevented from entering M within the first 30–90 min after IR. However, ATM is not required for prolonged G2 accumulation after IR (Xu et al., 2002). There have been conflicting reports about the role of the Mre11 complex in the early G2/M checkpoint (Buscemi et al., 2001; Xu et al., 2001, 2002; Williams et al., 2002). We reasoned that if the Mre11 complex was required for ATM activation and signaling, cells expressing E1b55K/E4orf6 should exhibit a defect in the ATM-dependent G2 arrest after IR. In the absence of E4orf6, this checkpoint was intact in HeLa cells expressing either GFP or any of the E1b55K proteins (Figure 6C). However, in the wild-type E1b55K cell line infected with rAd.E4orf6, this checkpoint was abrogated, and cells continued to enter M phase after IR treatment. This was also seen in cells expressing the R240A mutant, but not in the GFP or H354 cell lines infected with rAd.E4orf6. These results show that the Mre11 complex is required for the early G2/M checkpoint, possibly via ATM activation. Mre11 degradation by E1b55K/E4orf6 did not abrogate ATM-independent long-term G2 accumulation after IR (data not shown).
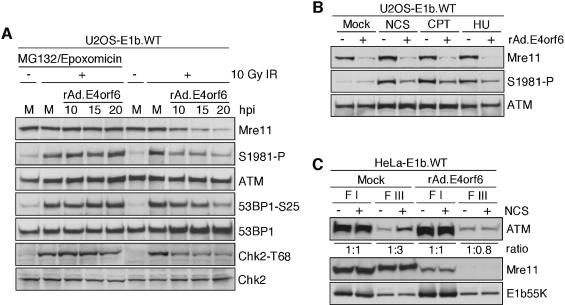
To demonstrate that the effect of E1b55K/E4orf6 on ATM activation and signaling is due to Mre11 complex degradation and not disruption of functional complexes or steric hindrance, we utilized proteasome inhibitors to block degradation (Figure 7A). Cells expressing wild-type E1b55K (U2OS-E1b.WT) were either untreated or infected with rAd.E4orf6 for various time periods prior to exposure with 10 Gy of IR. This experiment was performed in parallel in the presence and absence of the proteasome inhibitors MG132 and epoxomicin. The degree of ATM activation in response to IR diminished over time, with increased degradation of the Mre11 complex. The same was observed for signaling to downstream substrates, as measured by phosphorylation of 53BP1 and Chk2. In addition, ATM autophosphorylation and signaling were restored in the presence of proteasome inhibitors, demonstrating that E1b55K/E4orf6 did not directly block ATM kinase activity or the ability of Mre11 to lead to ATM activation.
Fig. 7. Degradation of the Mre11 complex prevents ATM activation and retention on chromatin in response to agents that cause DSBs. (A) Proteasome-mediated degradation is required for E1b55K/E4orf6 to prevent ATM activation. U2OS cells expressing wild-type E1b55K (U2OS-E1b.WT) were either mock infected (M) or infected with rAd.E4orf6 for the indicated times (h). Cells were irradiated with 10 Gy and harvested at 1 h post-treatment. This experiment was performed in parallel in the presence or absence of the proteasome inhibitors, MG132 (10 µM) and epoxomicin (1 µM). Inhibitors were added 2 h.p.i. and left on for the duration of the infection. (B) E1b55K/E4orf6 prevent ATM autophosphorylation in response to agents that lead to DSBs. U2OS cells expressing wild-type E1b55K (U2OS-E1b.WT) were either uninfected (–) or infected (+) with rAd.E4orf6. At 24 h.p.i. these cells were either untreated (Mock) or treated with 50 ng/ml neocarzinostatin (NCS) for 4 h, 1 µM camptothecin (CPT) for 2 h, or 2 mM hydroxyurea (HU) for 4 h. (C) Degradation of the Mre11 complex prevents ATM retention after treatment with NCS. HeLa cells expressing wild-type E1b55K (HeLa-E1b.WT) were either mock infected (Mock) or infected with rAd.E4orf6. At 24 h.p.i. the cells were treated with 200 ng/ml NCS for 30 min. The cells were harvested and biochemically fractionated as described in Materials and methods. The intensity of bands was quantitated and the ratios of ‘sample without NCS treatment’ to ‘sample with NCS’ are given below. Immunoblotting for Mre11 demonstrated degradation, and blotting for E1b55K served as an internal control for equal loading.
The effect of E1b55K/E4orf6 proteins on ATM activation was assessed following treatment with other DNA damaging agents that lead to DSBs (Figure 7B). Immunoblotting with the phosphospecific antibody to S1981 demonstrated that ATM was activated in U2OS cells expressing wild-type E1b55K after treatment with neocarzinostatin (NCS), camptothecin (CPT) and hydroxyurea (HU). Prior infection with rAd.E4orf6 led to Mre11 degradation and this was accompanied by abrogated ATM activation, demonstrating that the Mre11 complex is required for ATM activation in response to DSBs induced by a number of agents.
We also examined the effect of Mre11 degradation by E1b55K/E4orf6 on ATM chromatin retention in response to DSBs (Andegeko et al., 2001). We performed this assay using the HeLa-E1b.WT cell line in the absence or presence of E4orf6 (Figure 7C). We observed ATM retained in fraction III (FIII) of uninfected cells treated with 200 ng/ml NCS for 30 min, similar to published results in HeLa cells (Andegeko et al., 2001). This retention was abolished when the cells were infected with rAd.E4orf6 and the Mre11 complex was degraded prior to NCS treatment, suggesting that the Mre11 complex, and possibly ATM autophosphorylation, is required for the retention of ATM on chromatin after DSB induction.
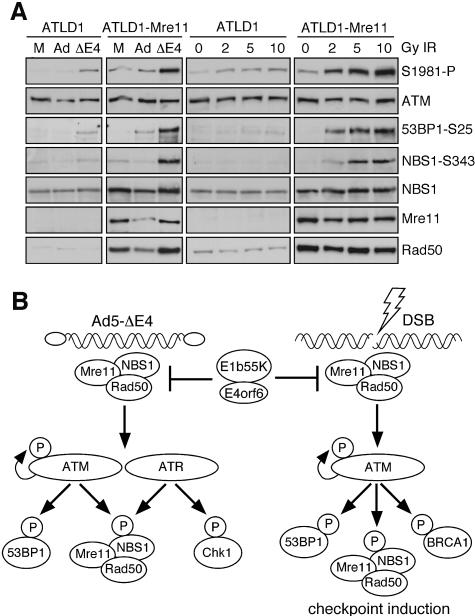
To confirm our conclusions from observations using E1b55K/E4orf6, we performed experiments in the A-TLD1 patient line that expresses a truncated, hypomorphic form of the Mre11 protein (Stewart et al., 1999). A-TLD1 cells were immortalized and then complemented with the wild-type Mre11 cDNA or transduced with an empty retrovirus vector. Expression of full-length Mre11 protein was confirmed by immunoblotting and resulted in stabilization of Rad50 and NBS1 proteins (Figure 8A), as well as relocalization of Rad50 into the nucleus (supplementary figure 3). Some autophosphorylation of ATM was observed in the A-TLD1 cells infected with the E4-deleted virus or exposed to IR, but it was highly abrogated. In contrast to the vector-transduced A-TLD1 cells, ATM autophosphorylation and signaling to downstream targets such as 53BP1 was robust in the Mre11-complemented cells. Together these data demonstrate that the Mre11 complex is required for full activation of the ATM response to virus infection and DSBs, and support the conclusions made from degradation of the Mre11 complex by the E1b55K/E4orf6 viral proteins.
Fig. 8. Complementation of A-TLD1 cells with Mre11 cDNA restores ATM activation and signaling in response to IR. (A) ATM activation and signaling is abrogated in A-TLD1 cells and can be restored by complementation with wild-type Mre11. Cells were either infected with virus (m.o.i. = 100) or exposed to IR. Cells were harvested 24 h.p.i. with wild-type Ad or the E4-deleted virus (ΔE4). Irradiated cells were harvested at 45 min after recovery from exposure to the indicated amount of IR. (B) Model for the role of the Mre11 complex in DNA damage response to virus infection and DSBs. Infection with an E4-deleted adenovirus leads to activation of ATM and ATR. These kinases are redundant for the phosphorylation of many substrates, including NBS1. However, these kinases also have specific targets, such as Ser25 on 53BP1 for ATM, and Ser345 on Chk1 for ATR. Degradation of the Mre11 complex by E1b55K/E4orf6 proteins prevents the activation of the ATM/ATR kinases. The Mre11 complex also plays a role as an upstream sensor in response to DSBs, and degradation of the Mre11 complex prevents ATM autophosphorylation and signaling, in addition to the ATM-dependent G2/M checkpoint.
Discussion
It has been shown that the Mre11 complex and ATM function in a common pathway, and that NBS1 in particular plays a role downstream as a substrate of ATM kinase activity (Gatei et al., 2000; Lim et al., 2000; Wu et al., 2000; Zhao et al., 2000). Our new data demonstrate that the Mre11 complex also functions upstream during ATM activation in mammalian cells. Recruitment of the Mre11 complex to viral centers (Figure 3C and 3D) and DSBs (Mirzoeva and Petrini, 2001; Celeste et al., 2003) was independent of ATM/ATR kinase activity or γ-H2AX foci, suggesting the Mre11 complex functions early in damage detection (as proposed in Nelms et al., 1998). In the case of both infection with the E4-deleted virus and induction of DSBs, degradation of the Mre11 complex prevented ATM autophosphorylation and chromatin retention after damage (supplementary figure 1; Figure 7C). This demonstrated that the Mre11 complex can function upstream of ATM activation as a damage sensor, in addition to acting as an effector of ATM signaling.
Cell-cycle checkpoints protect mammalian cells from the permanent genetic damage that can lead to transformation and oncogenesis. A requirement for the Mre11 complex in the S-phase checkpoint has been well established (D’Amours and Jackson, 2001; Usui et al., 2001; Falck et al., 2002), but a role in the G2/M checkpoint has been controversial. It has been shown that NBS cells have a defective G2/M transition immediately after a low dose of IR (Buscemi et al., 2001; Williams et al., 2002). However, other reports have suggested that the early G2/M checkpoint is independent of NBS1 (Xu et al., 2001, 2002). All these studies were performed using hypomorphic patient or mouse cell lines. The adenoviral system that we have employed provides an alternative means for studying these checkpoints in the absence of the Mre11 complex. Depletion of the cellular Mre11 complex by the E1b55K/E4orf6 proteins correlated with inhibition of the early G2/M checkpoint, consistent with the phenotype observed in cells lacking ATM and the idea that the Mre11 complex is required for ATM activation in response to DSBs.
Our data suggest a model in which the Mre11 complex plays a role as an upstream sensor of both Ad genomes and DSBs (Figure 8B). This conclusion is supported by a recent report showing a requirement for functional Mre11 in ATM activation to DSBs in hypomorphic A-TLD cell lines (Uziel et al., 2003). During adenovirus infection, degradation of the Mre11 complex prevents signaling through both ATM and ATR pathways. It will be interesting to determine whether there is also a requirement for the Mre11 complex in activation of ATR signaling in response to DNA damage other than viral infection. The mechanism by which the Mre11 complex leads to activation of signaling pathways in response to viral infection and DSBs remains to be elucidated. A conceivable model would be that the Mre11 complex is required to recognize adenovirus genomes and DSBs and then modify the DNA by resection to create a platform for ATM and other signaling factors (D’Amours and Jackson, 2002). This model is supported by data that demonstrate that the Mre11 complex can bind to DSBs in vitro (de Jager et al., 2001). In the case of Ad, this may include nucleolytic removal of the terminal protein to expose the dsDNA termini of the genome (Stracker et al., 2002). The interplay between adenovirus and host DNA repair pathways raises a cautionary note when interpreting studies using these viruses as vectors. Given the variety of replicative mechanisms employed by DNA viruses, we believe that there will be an emergence of other examples of viruses that must inactivate or take advantage of the DNA repair machinery to ensure a productive infection. By exploiting adenoviral proteins we have shown that the Mre11 complex can function upstream in ATM activation and signaling in response to DNA damage. This viral system provides a powerful tool to deplete the Mre11 complex in cells of any genetic background and will allow the assessment of functions of the complex that may not be perturbed in the hypomorphic NBS and A-TLD patient lines. Additionally, this system will allow the rapid analysis of the genetic interactions between the Mre11 complex and other repair factors, providing novel insights into the roles of cellular proteins in the DNA damage response.
Materials and methods
Cell lines
HeLa, U2OS and 293 cells were purchased from the American Tissue Culture Collection. Packaging cells for retroviruses were obtained from I.Verma (Salk Institute), W162 cells for growth of the E4-deleted virus were from G.Ketner (Johns Hopkins Bloomberg School of Health), and ATM cells (GM02052D and GM5849) were from Y.Shiloh (Tel Aviv University) or the Coriell Institute. The U2OS-derived cells that express inducible wild-type (GW33) and kinase-dead (GK41) ATR proteins were from S.Schreiber (Harvard University), and were induced with doxyclycine as described previously (Nghiem et al., 2001). A-TLD1 cells were from J.Petrini (Memorial Sloan-Kettering Cancer Center) and were immortalized by transduction with retroviruses expressing SV40 T-antigen and hTERT. These cells were then transduced with retrovirus expressing the wild-type Mre11 cDNA cloned into pLPC (Serrano et al., 1997) or the empty vector. Puromycin-resistant pools were selected and maintained at 1 µg/ml. All cells were maintained as monolayers in Dulbecco modified Eagle’s medium (DMEM) supplemented with 10 or 20% fetal bovine serum (FBS), at 37°C in a humidified atmosphere containing 5% CO2.
Viruses
Wild-type Ad5, E1b55K mutant viruses, the double mutant dl1016 and E1-deleted recombinant viruses were all propagated in 293 cells. The E4 mutant viruses dl1004 and dl1016 have been described previously (Bridge and Ketner, 1990), and were obtained from G.Ketner. The dl1004 virus was propagated on W162, a Vero-derived, E4-complementing cell line. The mutant E1b55K viruses, H354 and R240A, have been described previously (Yew et al., 1990; Shen et al., 2001) and were obtained from A.Berk (University of California-Los Angeles) and Y.Shen (Onyx Pharmaceuticals), respectively. The recombinant adenoviruses rAd.E4orf6 and rAd.E1b55K have been described previously (Querido et al., 2001a) and were obtained from P.Branton (McGill University). All viruses were purified by two sequential rounds of ultra-centrifugation in cesium chloride gradients and stored in 40% glycerol at –20°C.
Antibodies
Viral replication centers were visualized by staining for adenovirus DBP with either mouse monoclonal antibody (B6-8) or rabbit polyclonal antisera. Primary antibodies were purchased from Novus Biologicals Inc. (NBS1, NBS1-P-NB100-284, ATM), Genetex (Mre11-12D7, Rad50-13B3), Santa Cruz (Ku70, Ku86, DNA-PKcs, p53, ATR, Chk1), Upstate (Chk2), Cell Signaling (Chk1-S345, Chk2-T68, p53-S15), Oncogene (BRCA1) and Rockland (ATM S1981-P). Antisera to E4orf6 was from P.Branton, antibody to 53BP1 was from T.Halazonetiso (Wistar Institute) and the phosphospecific antibody was from P.Carpenter (University of Texas-Houston), antibody to RPA32 was from T.Melendy (University of Buffalo), antibody to phosphorylated H2AX was from W.Bonner (National Cancer Institute), antibodies to DBP and E1b55K were from A.Levine (Cancer Institute of New Jersey), and polyclonal rabbit antisera to DBP was from P.van der Vliet (University Medical Center of Utrecht). All secondary antibodies were supplied by Jackson Laboratories.
Generating E1b55K cell lines
To generate E1b55K cell lines, the gene was amplified by PCR using either wild-type Ad5 or the mutant viruses as template. The coding region was cloned under the control of the cytomegalovirus promoter in a modified version of the pCLNC retrovirus vector. Mutants were confirmed by DNA sequencing. Retroviruses were generated by transfection of packaging cell lines together with a plasmid expressing the VSV-G envelope protein. HeLa or U2OS cells were infected with retrovirus supernatants and selected in 900 µg/ml G418. Expression of E1b55K was assessed by immunoblotting and pools of resistant clones were used for experiments.
Infection, irradiation and drug treatments
Infections were performed on monolayers of cultured cells in DMEM supplemented with 2% FBS. After 2 h at 37°C, additional serum was added to a total of 10%. A Co60 irradiation chamber was used for irradiations. To block degradation, proteasome inhibitors MG132 (10 µM) and epoxomicin (1 µM) were added at 4 hours post-infection (h.p.i.) with rAd.E4orf6, and were left on for the duration of infection. Neocarzinostatin (NCS) was a gift from G.Wahl (Salk Institute). Caffeine, wortmannin, hydroxyurea (HU) and camptothecin (CPT) were obtained from Sigma.
ATM retention assay
The ATM retention assay was performed essentially as described previously (Andegeko et al., 2001). To assay the effect of virus infection on ATM retention, the HeLa cell line expressing GFP was infected with wild-type Ad5 and dl1004 for 18 h. Extraction with NP-40 extraction yielded three fractions (FI, FII and FIII). To test the effect of Mre11 degradation on ATM retention in response to DSBs, the HeLa cell line expressing wild-type E1b55K was infected with rAd.E4orf6 for 24 h and then treated with 200 ng/ml of neocarzinostatin for 30 min before harvesting.
Flow cytometry analysis
The G2/M checkpoint assay was performed essentially as described previously (Xu et al., 2002). Cell lines that express either GFP, or wild-type and mutant E1b55K proteins were either untreated or infected with rAd.E4orf6 (m.o.i. = 50) for 48 h prior to irradiation with 10 Gy, and then analyzed for mitosis and DNA content at 1 h post-IR. Phosphorylated histone H3 was detected with a phosphospecific polyclonal primary antibody (Upstate Biotechnology) and an FITC-conjugated anti-rabbit secondary antibody (Jackson Laboratories). To measure DNA content, cells were stained with propidium iodide (PI; Sigma). Fluorescence was measured by flow cytometry using a Becton-Dickinson FACScan. For each sample, 20 000 events were analyzed.
Immunoblotting and immunofluorescence
Western blotting was performed as described previously (Cathomen and Weitzman, 2000). For immunofluorescence, cells were grown on glass coverslips in 24-well dishes and infected with wild-type or mutant viruses at an m.o.i. of 25–100 p.f.u./cell. After 18–40 h the cells were washed with phosphate-buffered saline (PBS) and fixed at –20°C for 30 min with ice-cold methanol/acetone (1:1). For detection of ATR, cells were fixed in 3% paraformaldehyde for 20 min and extracted with 0.5% Triton X-100 in PBS for 10 min. In all cases, control staining experiments showed no cross-reaction between the fluorophores, and images obtained by staining with individual antibodies were the same as those shown for double-labeling. Nuclear DNA was stained with 4′,6-diamidino-2-phenylindol (DAPI) and coverslips were mounted using Fluoromount-G (Southern Biotechnology Associates). Immunoreactivity was visualized using a Nikon microscope in conjunction with a charge-coupled device camera (Cooke Sensicam). Images were obtained in double or triple excitation mode, and processed using SlideBook and Adobe Photoshop.
Acknowledgments
Acknowledgements
We are grateful to A.Berk, W.Bonner, P.Branton, P.Carpenter, T.Halazonetis, G.Ketner, A.Levine, T.Melendy, J.Petrini, S.Schreiber, Y.Shen, Y.Shiloh, P.van der Vliet and I.Verma for generous gifts of reagents. We thank J.Karlseder for help with immortalization of A-TLD1 cells. We thank A.Dillin, T.Hunter, J.Karlseder, J.Petrini, G.Wahl, J.Weitzman and members of the Weitzman lab for discussions and critical reading of the manuscript. We acknowledge the James B.Pendleton Charitable Trust for providing the Pendleton Microscopy Facility. This work was supported in part by National Institutes of Health (NIH) grants AI43341 and CA97093 (to M.D.W.), and by gifts from the Joe W. and Dorothy Dorsett Brown Foundation, and the Lebensfeld Foundation. C.T.C. was supported by the Timken-Sturgis Foundation and by an NIH Training Grant to the Salk Institute. R.S. was supported by The Hallock Foundation and by an NIH Graduate Training Grant to University of California, San Diego (UCSD). T.H.S. was supported by an NIH Graduate Training Grant to UCSD, and by fellowships from the H.A. and Mary K.Chapman Charitable Trust, the Legler Benbough Foundation and the Salk Institute Association. C.E.L. was supported by a Wellcome Trust International Research Fellowship (GR066559).
References
- Andegeko Y., Moyal,L., Mittelman,L., Tsarfaty,I., Shiloh,Y. and Rotman,G. (2001) Nuclear retention of ATM at sites of DNA double strand breaks. J. Biol. Chem., 276, 38224–38230. [DOI] [PubMed] [Google Scholar]
- Bakkenist C.J. and Kastan,M.B. (2003) DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature, 421, 499–506. [DOI] [PubMed] [Google Scholar]
- Bridge E. and Ketner,G. (1990) Interaction of adenoviral E4 and E1b products in late gene expression. Virology, 174, 345–353. [DOI] [PubMed] [Google Scholar]
- Brown E.J. and Baltimore,D. (2003) Essential and dispensable roles of ATR in cell cycle arrest and genome maintenance. Genes Dev., 17, 615–628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buscemi G. et al. (2001) Chk2 activation dependence on Nbs1 after DNA damage. Mol. Cell. Biol., 21, 5214–5222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carney J.P., Maser,R.S., Olivares,H., Davis,E.M., Le Beau,M., Yates,J.R.,III, Hays,L., Morgan,W.F. and Petrini,J.H. (1998) The hMre11/hRad50 protein complex and Nijmegen breakage syndrome: linkage of double-strand break repair to the cellular DNA damage response. Cell, 93, 477–486. [DOI] [PubMed] [Google Scholar]
- Cathomen T. and Weitzman,M.D. (2000) A functional complex of adenovirus proteins E1B-55kDa and E4orf6 is necessary to modulate the expression level of p53 but not its transcriptional activity. J. Virol., 74, 11407–11412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Celeste A., Fernandez-Capetillo,O., Kruhlak,M.J., Pilch,D.R., Staudt,D.W., Lee,A., Bonner,R.F., Bonner,W.M. and Nussenzweig,A. (2003) Histone H2AX phosphorylation is dispensable for the initial recognition of DNA breaks. Nature Cell Biol., 5, 675–679. [DOI] [PubMed] [Google Scholar]
- Cliby W.A., Roberts,C.J., Cimprich,K.A., Stringer,C.M., Lamb,J.R., Schreiber,S.L. and Friend,S.H. (1998) Overexpression of a kinase-inactive ATR protein causes sensitivity to DNA-damaging agents and defects in cell cycle checkpoints. EMBO J., 17, 159–169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Costanzo V., Robertson,K., Bibikova,M., Kim,E., Grieco,D., Gottesman,M., Carroll,D. and Gautier,J. (2001) Mre11 protein complex prevents double-strand break accumulation during chromosomal DNA replication. Mol. Cell, 8, 137–147. [DOI] [PubMed] [Google Scholar]
- D’Amours D. and Jackson,S.P. (2001) The yeast Xrs2 complex functions in S phase checkpoint regulation. Genes Dev., 15, 2238–2249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Amours D. and Jackson,S.P. (2002) The Mre11 complex: at the crossroads of DNA repair and checkpoint signalling. Nature Rev. Mol. Cell. Biol., 3, 317–327. [DOI] [PubMed] [Google Scholar]
- de Jager M., van Noort,J., van Gent,D.C., Dekker,C., Kanaar,R. and Wyman,C. (2001) Human Rad50/Mre11 is a flexible complex that can tether DNA ends. Mol. Cell, 8, 1129–1135. [DOI] [PubMed] [Google Scholar]
- DiTullio R.A. Jr, Mochan,T.A., Venere,M., Bartkova,J., Sehested,M., Bartek,J. and Halazonetis,T.D. (2002) 53BP1 functions in an ATM-dependent checkpoint pathway that is constitutively activated in human cancer. Nature Cell Biol., 4, 998–1002. [DOI] [PubMed] [Google Scholar]
- Dobbelstein M., Roth,J., Kimberly,W.T., Levine,A.J. and Shenk,T. (1997) Nuclear export of the E1B 55-kDa and E4 34-kDa adenoviral oncoproteins mediated by a rev-like signal sequence. EMBO J., 16, 4276–4284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falck J., Petrini,J.H., Williams,B.R., Lukas,J. and Bartek,J. (2002) The DNA damage-dependent intra-S phase checkpoint is regulated by parallel pathways. Nature Genet., 30, 290–294. [DOI] [PubMed] [Google Scholar]
- Fernandez-Capetillo O. et al. (2002) DNA damage-induced G2–M checkpoint activation by histone H2AX and 53BP1. Nat. Cell Biol., 4, 993–997. [DOI] [PubMed] [Google Scholar]
- Gatei M. et al. (2000) ATM-dependent phosphorylation of nibrin in response to radiation exposure. Nature Genet., 25, 115–119. [DOI] [PubMed] [Google Scholar]
- Goodrum F.D., Shenk,T. and Ornelles,D.A. (1996) Adenovirus early region 4 34-kilodalton protein directs the nuclear localization of the early region 1B 55-kilodalton protein in primate cells. J. Virol., 70, 6323–6335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harada J.N., Shevchenko,A., Pallas,D.C. and Berk,A.J. (2002) Analysis of the adenovirus E1B-55K-anchored proteome reveals its link to ubiquitination machinery. J. Virol., 76, 9194–9206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kastan M.B. and Lim,D.S. (2000) The many substrates and functions of ATM. Nat. Rev. Mol. Cell Biol., 1, 179–186. [DOI] [PubMed] [Google Scholar]
- Lim D.S., Kim,S.T., Xu,B., Maser,R.S., Lin,J., Petrini,J.H. and Kastan,M.B. (2000) ATM phosphorylates p95/nbs1 in an S-phase checkpoint pathway. Nature, 404, 613–617. [DOI] [PubMed] [Google Scholar]
- Mirzoeva O.K. and Petrini,J.H. (2001) DNA damage-dependent nuclear dynamics of the Mre11 complex. Mol. Cell. Biol., 21, 281–288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelms B.E., Maser,R.S., MacKay,J.F., Lagally,M.G. and Petrini,J.H. (1998) In situ visualization of DNA double-strand break repair in human fibroblasts. Science, 280, 590–592. [DOI] [PubMed] [Google Scholar]
- Nghiem P., Park,P.K., Kim,Y., Vaziri,C. and Schreiber,S.L. (2001) ATR inhibition selectively sensitizes G1 checkpoint-deficient cells to lethal premature chromatin condensation. Proc. Natl Acad. Sci. USA, 98, 9092–9097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paull T.T., Rogakou,E.P., Yamazaki,V., Kirchgessner,C.U., Gellert,M. and Bonner,W.M. (2000) A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr. Biol., 10, 886–895. [DOI] [PubMed] [Google Scholar]
- Petrini J.H. (1999) The mammalian Mre11-Rad50-nbs1 protein complex: integration of functions in the cellular DNA-damage response. Am. J. Hum. Genet., 64, 1264–1269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Querido E. et al. (2001a) Degradation of p53 by adenovirus E4orf6 and E1B55K proteins occurs via a novel mechanism involving a Cullin-containing complex. Genes Dev., 15, 3104–3117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Querido E., Morrison,M.R., Chu-Pham-Dang,H., Thirlwell,S.W., Boivin,D., Branton,P.E. and Morisson,M.R. (2001b) Identification of three functions of the adenovirus e4orf6 protein that mediate p53 degradation by the E4orf6-E1B55K complex. J. Virol., 75, 699–709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rouse J. and Jackson,S.P. (2002) Interfaces between the detection, signaling and repair of DNA damage. Science, 297, 547–551. [DOI] [PubMed] [Google Scholar]
- Rubenwolf S., Schutt,H., Nevels,M., Wolf,H. and Dobner,T. (1997) Structural analysis of the adenovirus type 5 E1B 55-kilodalton-E4orf6 protein complex. J. Virol., 71, 1115–1123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarkaria J.N., Busby,E.C., Tibbetts,R.S., Roos,P., Taya,Y., Karnitz,L.M. and Abraham,R.T. (1999) Inhibition of ATM and ATR kinase activities by the radiosensitizing agent, caffeine. Cancer Res., 59, 4375–4382. [PubMed] [Google Scholar]
- Serrano M., Lin,A.W., McCurrach,M.E., Beach,D. and Lowe,S.W. (1997) Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell, 88, 593–602. [DOI] [PubMed] [Google Scholar]
- Shen Y., Kitzes,G., Nye,J.A., Fattaey,A. and Hermiston,T. (2001) Analyses of single-amino-acid substitution mutants of adenovirus type 5 E1B-55K protein. J. Virol., 75, 4297–4307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiloh Y. (2003) ATM and related protein kinases: safeguarding genome integrity. Nat. Rev. Cancer, 3, 155–168. [DOI] [PubMed] [Google Scholar]
- Stewart G.S. et al. (1999) The DNA double-strand break repair gene hMRE11 is mutated in individuals with an ataxia-telangiectasia-like disorder. Cell, 99, 577–587. [DOI] [PubMed] [Google Scholar]
- Stracker T.H., Carson,C.T. and Weitzman,M.D. (2002) Adenovirus oncoproteins inactivate the Mre11-Rad50-NBS1 DNA repair complex. Nature, 418, 348–352. [DOI] [PubMed] [Google Scholar]
- Usui T., Ogawa,H. and Petrini,J.H. (2001) A DNA damage response pathway controlled by Tel1 and the Mre11 complex. Mol. Cell, 7, 1255–1266. [DOI] [PubMed] [Google Scholar]
- Uziel T., Lerenthal,Y., Moyal,L. andegeko,Y., Mittelman,L. and Shiloh,Y. (2003) Requirement of the MRN complex for ATM activation by DNA damage. EMBO J., 22, 5612–5621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams B.R., Miroeva,O.K., Morgan,W.F., Lin,J., Dunnick,W. and Petrini,J.H. (2002) A murine model of Nijmegen breakage syndrome. Curr. Biol., 12, 648–653. [DOI] [PubMed] [Google Scholar]
- Wu X. et al. (2000) ATM phosphorylation of Nijmegen breakage syndrome protein is required in a DNA damage response. Nature, 405, 477–482. [DOI] [PubMed] [Google Scholar]
- Xu B., Kim,S. and Kastan,M.B. (2001) Involvement of Brca1 in S-phase and G(2)-phase checkpoints after ionizing irradiation. Mol. Cell. Biol., 21, 3445–3450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu B., Kim,S.T., Lim,D.S. and Kastan,M.B. (2002) Two molecularly distinct G(2)/M checkpoints are induced by ionizing irradiation. Mol. Cell. Biol., 22, 1049–1059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yew P.R., Kao,C.C. and Berk,A.J. (1990) Dissection of functional domains in the adenovirus 2 early 1B 55K polypeptide by suppressor-linker insertional mutagenesis. Virology, 179, 795–805. [DOI] [PubMed] [Google Scholar]
- Zantema A., Fransen,J.A., Davis-Olivier,A., Ramaekers,F.C., Vooijs,G.P., DeLeys,B. and Van der Eb,A.J. (1985) Localization of the E1B proteins of adenovirus 5 in transformed cells, as revealed by interaction with monoclonal antibodies. Virology, 142, 44–58. [DOI] [PubMed] [Google Scholar]
- Zhao S. et al. (2000) Functional link between ataxia-telangiectasia and Nijmegen breakage syndrome gene products. Nature, 405, 473–477. [DOI] [PubMed] [Google Scholar]
- Zou L. and Elledge,S.J. (2003) Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science, 300, 1542–1548. [DOI] [PubMed] [Google Scholar]