Abstract
Germ-line stem cells (GSCs) are maintained by the somatic microenvironment, or GSC niche, which ensures that GSCs can both self-renew and produce functional gametes. However, it remains unclear how the proper niche size and location are regulated within the developing gonads. In the Drosophila testis, the hub cells that form the GSC niche are derived from a subset of somatic gonadal precursors (SGPs) in the anterior portion of the embryonic gonad. Here we show that Notch signaling induces hub differentiation. Notch is activated in almost all SGPs in the male embryonic gonad, but Epidermal growth factor receptor (Egfr) is activated in posterior SGPs to repress hub differentiation, thereby restricting the expansion of hub differentiation in the embryonic gonad. We further show that Egfr is activated in posterior SGPs by Spitz ligand secreted from primordial germ cells (PGCs), whereas the Notch ligand Serrate is expressed in SGPs. This suggests that varying the number of PGCs alters niche size. Indeed, a decrease in the number of PGCs causes ectopic hub differentiation, which consequently increases their opportunity to recruit PGCs as GSCs. When ectopic hub differentiation is repressed, the decreased number of PGCs fails to become GSCs. Thus, we propose that SGPs sense PGC number via signals from PGCs to SGPs that modulate niche size, and that this serves as a mechanism for securing GSCs.
Keywords: primordial germ cell, pole cell, somatic gonadal precursor, hub cell
Stem cells contribute to a steady source of new cells to maintain tissue homeostasis. Many stem cells reside in local microenvironments, or niches, that act as sources of local extrinsic signals to stem cells to maintain their self-renewing potential. Recently, niches have been identified in various tissues, such as the Drosophila intestinal epithelium (1–3), ovary (4), and testis (5, 6), as well as the mammalian skin epidermis (7), hematopoietic system (8–10), and neural tissues (11–13). To maintain proper tissue architecture, the number of stem cells and the proper placement of their niche in a tissue are crucial. For example, misregulation of niche formation results in tissue degeneration or tumor-like morphological abnormalities (8, 9, 14–18). Although the role of niches in the maintenance of stem cells and tissue homeostasis has been well-studied, relatively little is known about how niche size and location are precisely regulated during development.
One of the best-characterized stem cell niches is found in the Drosophila testis, in which easily identifiable germ-line stem cells (GSCs) and their niche cells maintain the continuous production of sperm. At the apical tip of the testis, GSCs lie in intimate contact with somatic niche cells, or hub cells, which send the self-renewal signal Unpaired (Upd), a ligand that activates Jak/STAT signaling pathway in the neighboring GSCs (5, 6, 19). When a GSC divides, the daughter remaining in contact with the hub cells maintains stem cell identity, while the daughter that is displaced from the hub receives a weaker signal and initiates spermatogenesis.
GSCs are derived from pole cells that form in the posterior pole of early embryos (20). They migrate into the interior of the embryo and associate with somatic gonadal precursors (SGPs) to form the embryonic gonad, then become primordial germ cells (PGCs). At this early stage, the embryonic gonad is already sexually dimorphic (21, 22). In male embryos, hub cells are specified from the anterior SGPs at the end of embryogenesis (23, 24). These hub cells send a short-range signal, Upd, to the neighboring PGCs to recruit them as GSCs, whereas PGCs that do not associate with hub cells undergo spermatogenesis (24). In contrast, in females, GSCs and their niche are formed later in third-instar larvae (25, 26).
We have previously reported that the receptor tyrosine kinase (RTK) Sev is required to ensure that the niche develops in the anterior region of the male embryonic gonads (17). Sev is expressed in the posterior SGPs and is activated by the Boss ligand emanating from PGCs to prevent ectopic niche differentiation in the posterior SGPs. However, Sev is not sufficient to repress hub differentiation in the anterior gonad (17), suggesting that another signaling pathway has a central role in restricting hub differentiation to the anterior SGPs. These findings also suggest that both posterior and anterior SGPs have the capacity to contribute to hub differentiation, but the mechanisms by which they do this remain elusive.
In this study, we demonstrate that Notch and Epidermal growth factor receptor (Egfr) signaling act antagonistically to regulate hub differentiation in the male embryonic gonad. We show that Notch is activated in almost all SGPs to induce hub differentiation. Notch is activated by Ser ligand emanating from SGPs. In contrast, activation of Egfr in posterior SGPs by Spitz ligand secreted from PGCs is necessary and sufficient to repress hub differentiation. Thus, anterior and posterior SGPs become competent to differentiate into hub cells through Notch-mediated interactions between SGPs, whereas PGCs signal to SGPs through Egfr and Sev to restrict hub differentiation to anterior SGPs. We further show that a decrease in the number of PGCs enhances ectopic hub formation, which in turn increases the recruitment of PGCs as GSCs. Thus, regulatory interactions between PGCs and SGPs ensure that the proper number of GSCs is generated within the male gonads.
Results and Discussion
Notch Signaling Pathway Induces Hub Differentiation in the Male Embryonic Gonad.
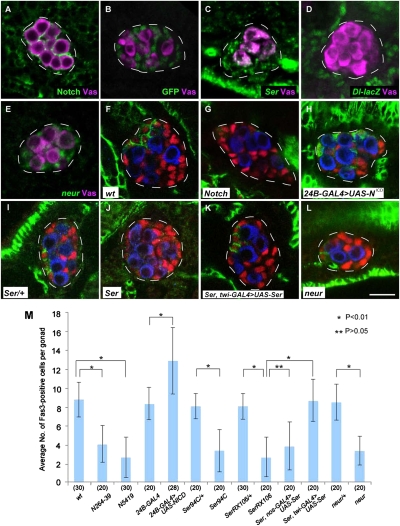
We tested the role of Notch signaling in hub differentiation within the male embryonic gonad, because Notch signaling pathway is required for stem cell maintenance and niche formation in various systems (3, 16, 26, 27). Notch expression was detectable in almost all SGPs from stage 12 until at least the end of embryogenesis in the male embryonic gonad (Fig. 1A). To determine whether Notch is required for hub differentiation, we examined the expression of Fasciclin 3 (Fas3) in Notch mutant embryos. Fas3 is expressed in hub cells from embryogenesis until adulthood (23, 28), and therefore is a good marker for hub cell identity (17). In wild-type embryos, Fas3-positive SGPs were observed only in the anterior region of the male embryonic gonad from stage 15 onward (Fig. 1F). Notch264-39 and Notch5419 null mutant gonads had a normal number of SGPs, and their overall morphology was indistinguishable from wild type during stage 13–16 (Fig. 1 G and M). However, they began to degenerate at stage 17. In Notch264-39 mutant embryonic gonads, hub differentiation was severely impaired, with the number of Fas3-positive cells reduced compared with wild type (Fig. 1 G and M). Conversely, when Notch activity was up-regulated in SGPs throughout the gonad by expression of the Notch intracellular domain (NotchICD), ectopic Fas3-positive cells were observed in the posterior of the gonad, and the number of Fas3-positive cells was significantly increased (Fig. 1 H and M). Based on these observations, we conclude that Notch is necessary and sufficient to induce hub differentiation throughout the male embryonic gonad.
Fig. 1.
Notch signaling is necessary and sufficient for inducing hub formation. (A) A male embryonic gonad (stage 15) stained for Notch (green) and the germ-line marker Vasa (Vas; magenta). Notch expression was observed in almost all SGPs in the male embryonic gonad. In all panels presented in this paper, anterior is to the left, and gonads are outlined by white lines. (B) A male embryonic hsp70-N-GV3/UAS-GFP gonad (stage 15) that was heat-treated as described in SI Materials and Methods and stained for GFP (green) and Vas (magenta). GFP expression was detected in almost all SGPs at stage 14–16. (C) A male embryonic gonad (stage 15) stained for Ser RNA (green) and Vas (magenta). Ser RNA was detected in almost all SGPs during stage 14–16. (D) A male embryonic gonad (stage 15) stained for β-galactosidase (green) and Vas (magenta). Expression of the Dl-lacZ enhancer trap (green) was undetectable within the male embryonic gonad. (E) A male embryonic gonad (stage 14) stained for neur RNA (green) and Vas (magenta). neur RNA was detected in almost all SGCs during stage 13–16. (F–L) Embryonic gonads (stage 16/17) stained for Fas3 (green), Tj (red; a marker for SGPs), and Vas (blue). (F) Wild-type embryo. (G) Notch264-39 mutant embryo. (H) Embryo expressing NotchICD in SGPs under the control of twist24B-GAL4 (24B-GAL4 > UAS-NotchICD). (I and J) SerRX106/+ (I) and SerRX106/SerRX106 (J) embryos. (K) SerRX106/SerRX106 embryo expressing Ser in SGPs (SerRX106/SerRX106, twist-GAL4 > UAS-Ser). (L) neur1/neur1 embryo. (Scale bar, 10 μm.) (M) The average number of Fas3-positive cells per gonad in wild-type (wt), N264-39 (N264-39), N5419 (N5419), twist24B-GAL4 (24B-GAL4), twist24B-GAL4 > UAS-NICD (24B-GAL4 > UAS-NCA), Ser94C/+ (Ser94C/+), Ser94C/Ser94C (Ser94C), SerRX106/+ (SerRX106/+), SerRX106/SerRX106 (SerRX106), neur1/+ (neur/+), and neur1/neur1 (neur) embryos, and in SerRX106/SerRX106 embryos expressing UAS-Ser in PGCs under the control of nanos-GAL4-VP16 (Ser, nos-GAL4 > UAS-Ser) and in SGCs under the control of twist-GAL4 (Ser, twi-GAL4 > UAS-Ser). Error bars represent SD. The number of gonads examined in each case is shown in parentheses. Significance was calculated using the Student's t test (*P < 0.01; **P > 0.05). The average number of SGPs (Tj-positive cells) ± SD per gonad at stage 16 was 34.9 ± 2.6 in wild-type embryos and 32.9 ± 2.6 in N264-39 embryos (20 gonads were examined in each case). These values were not significantly different (P > 0.05, Student's t test).
Because anterior SGPs contribute to hub differentiation during normal embryogenesis, we expected that Notch would be activated only in the anterior SGPs. To detect Notch activation, we expressed a NotchICD-GAL4 transgene under the control of a heat-shock promoter (hsp70-N-GV3) to activate a UAS-GFP reporter construct (29). Unexpectedly, GFP was detected in almost all SGPs in the embryonic gonad (Fig. 1B), indicating that Notch is active in both the posterior and anterior SGPs of the male embryonic gonad.
It has been reported that Notch is activated by binding to ligands encoded by Delta (Dl) or Serrate (Ser) (30). We detected Ser RNA in SGPs throughout the male embryonic gonad during stage 14–16 (Fig. 1C), whereas Dl-lacZ expression was undetectable (Fig. 1D). Consistent with these observations, the number of Fas3-positive cells was significantly decreased in Ser null (SerRX106 and Ser94C) mutant embryos, compared with control (Fig. 1 I, J, and M), whereas Dl mutations did not affect hub differentiation (Fig. S1 A and B). These results suggest that SGPs produce Ser ligand to activate Notch. This is further supported by several lines of evidence. First, hub differentiation occurs even in the absence of PGCs (17, 31). Second, forced expression of Ser in SGPs, but not in PGCs, is able to rescue hub differentiation defects in Ser mutants (Fig. 1 K and M). Third, neuralized (neur), which encodes an E3 ubiquitin ligase required for Ser activation in the signal-sending cells (30, 32), is expressed in almost all SGPs during stage 13–16, but not in PGCs (Fig. 1E). Finally, in the absence of neur activity, the number of Fas3-positive cells is significantly decreased, as observed in Ser and Notch mutant embryos (Fig. 1 L and M). Taken together, our observations strongly suggest that Ser/Notch signal transduction among SGPs induces hub differentiation in the male embryonic gonad.
Egfr Signaling from PGCs to SGPs Is Necessary and Sufficient to Repress Hub Differentiation.
The above observations suggest that SGPs are competent to become hub cells via the Ser/Notch signaling pathway throughout the male embryonic gonad. Because hub differentiation is restricted to the anterior SGPs during normal development, a repressive mechanism likely prevents posterior SGPs from becoming hub cells. We previously reported that Sev represses hub differentiation in posterior SGPs; in the absence of Sev activity, ectopic hub differentiation is observed. However, expression of a constitutively active form of Sev is unable to inhibit hub differentiation in the anterior SGPs (17), suggesting that Sev is not sufficient for repressing hub differentiation. Thus, we speculate that another RTK signaling pathway has a key role in restricting hub differentiation in anterior SGPs.
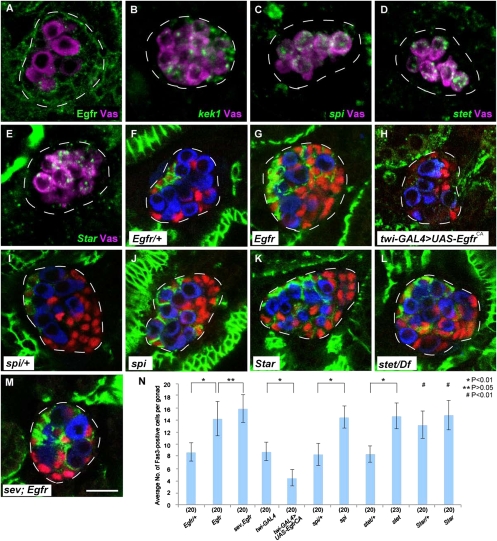
Here we focused on signaling via the RTK Egfr, because sev acts together with Egfr signaling to regulate eye development (33). We found that Egfr protein is expressed in almost all SGPs in the male embryonic gonad during stage 13–17 (Fig. 2A). To examine a potential role in hub differentiation, we used a temperature-sensitive allele of Egfr (Egfrts), because Egfr is known to be required for a variety of developmental processes, such as mesoderm development and muscle formation, during embryogenesis (34, 35). In Egfrts embryos cultured at a nonpermissive temperature (SI Materials and Methods), ectopic Fas3-positive cells were formed in the posterior region of the male embryonic gonad (Fig. 2 F, G, and N), whereas the morphology of the gonads and the total number of SGPs were unaffected (Fig. 2 F, G, and N). Moreover, hub differentiation, which is normally observed in anterior SGPs, was repressed by expressing a constitutively active form of Egfr (EgfrCA) throughout SGPs (Fig. 2 H and N). These observations show that Egfr is both necessary and sufficient for repressing hub differentiation.
Fig. 2.
Egfr is necessary and sufficient for repressing hub formation. (A–E) Male embryonic gonads (stage 15) stained for Vas (magenta) and Egfr (green) (A), kek1 RNA (green) (B), spi RNA (green) (C), stet RNA (green) (D), and Star RNA (green) (E). Egfr was expressed in SGPs during stage 13–17. kek1 RNA was expressed in posterior SGPs during stage 14–16. spi, stet, and Star RNA were expressed in PGCs during stage 13–16, 11–16, and 13–16, respectively. (F–M) Embryonic gonads (stage 16/17) stained for Fas3 (green), Tj (red), and Vas (blue). (F) Egfrtsla/+ embryo. (G) Egfrtsla/Egfrf24 (Egfrts) embryo. (H) Embryo expressing EgfrCA in SGPs (twist-GAL4 > UAS-EgfrCA). (I) spi1/+ embryo. (J) spi1/spi1 embryo. (K) StarIIN/StarIIN embryo. (L) stet871/Df(3L)PX62 embryo. (M) sevd2; Egfrts double-mutant embryo. (Scale bar, 10 μm.) (N) The average number of Fas3-positive cells per gonad in Egfrtsla/+ (Egfr/+), Egfrts (Egfr), sevd2; Egfrts (sev, Egfr), twist-GAL4 (twi-GAL4), twist-GAL4 > UAS-EgfrCA (twi-GAL4 > UAS-EgfrCA), spi1/+ (spi/+), spi1/spi1 (spi), stet871/+ (stet/+), stet871/Df(3L)PX62 (stet), StarIIN/+ (Star/+), and StarIIN/StarIIN (Star) embryos. Error bars represent SD. The number of gonads examined in each case is shown in parentheses. Significance was calculated using the Student's t test [*P < 0.01; **P > 0.05; #P < 0.01 (compared with wild type shown in Fig. 1M)]. The average number of SGPs ± SD per gonad at stage 16/17 was 33.9 ± 2.8 in Egfrtsla/+ embryos and 35.2 ± 3.0 in Egfrts embryos (20 gonads were examined in each case). These values were not significantly different (P > 0.05, Student's t test).
To determine whether Egfr signaling is activated in the posterior SGPs, we examined the expression of kek1, a known transcriptional target of Egfr signaling in various developmental contexts (36, 37). We found that kek1 RNA was expressed in the posterior but not in the anterior SGPs (Fig. 2B). This expression was abrogated by Egfrts mutation and, conversely, overexpression of EgfrCA caused ectopic kek1 expression in the anterior SGPs (Fig. S2 A and B), showing that kek1 expression is a readout of Egfr activation. From these observations, we propose that Egfr is activated only in the posterior SGPs in the male embryonic gonad.
We next asked which cell type (PGC or SGP) sends the signal that activates Egfr in SGPs. It has been reported that spitz (spi) encodes a ligand for Egfr, and its secretion requires stet and Star in the signal-sending cells (38–40). We found that spi expression was restricted to PGCs in the male embryonic gonad during stage 13–16 (Fig. 2C). Similarly, expression of stet and Star was detected in PGCs (Fig. 2 D and E). These results suggest that PGCs send a signal to activate Egfr in the SGPs.
To test whether reduction of spi, stet, and Star activity causes a similar phenotype as that observed in Egfr mutants, we used the loss-of-function mutations spi1, stet871, and StarIIN. In embryos homozygous for spi1 and StarIIN and transheterozygous for stet871 with a deficiency [Df(3L)PX62], Fas3-positive SGPs were observed ectopically in the posterior of the male embryonic gonad (Fig. 2 J–L). In spi and stet homozygous embryos, the number of Fas3-positive cells was significantly increased, compared with their heterozygous controls (Fig. 2 I, J, L, and N). These phenotypes were almost identical to that of Egfr mutant embryos (Fig. 2 G and N). In Star homozygous embryos, the number of Fas3-positive cells was increased to the level observed in Egfr, spi, and stet mutants (Fig. 2 K and N). We found that Star heterozygous gonads also showed a marked elevation in the number of Fas3-positive cells (Fig. 2N). This dosage sensitivity suggests that Star is a limiting factor for ligand production. Taken together, we conclude that the PGCs send a signal to activate Egfr signaling in the posterior SGPs, which in turn represses their differentiation into hub cells. Consequently, hub formation is restricted to the anterior portion of the male gonad.
Egfr Signaling Is Involved in the Mechanism Securing Germ-Line Stem Cells.
We have shown that Egfr is activated in posterior SGPs by Spi ligand from PGCs. Furthermore, we have reported that Boss/Sev signaling from PGCs to SGPs is also required to repress ectopic hub differentiation (17). It is interesting to note that both signaling pathways are activated by ligands emanating from PGCs. This implies that varying the number of PGCs alters the niche size. In the male gonad, only a subset of PGCs in the proximity of hub cells is recruited as GSCs at around hatching, whereas the others directly undergo spermatogenesis (24, 41). We speculate that a decrease in the number of PGCs should induce ectopic hub differentiation within the gonad, which consequently increases their opportunity to recruit PGCs as GSCs.
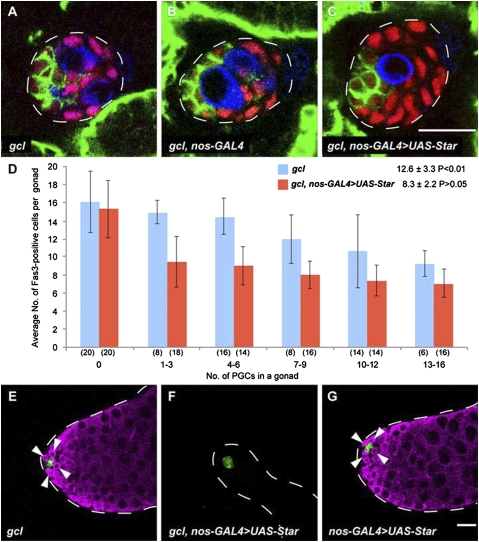
To address this, we decreased the number of PGCs by germ cell-less (gcl) mutation. In embryos with reduced maternal gcl activity (gcl embryos), a decreased number of pole cells are formed, but their subsequent development appears to be intact, except they have only a few PGCs in their gonads (42) (Fig. 3A). We found that reduction in the number of PGCs induced ectopic hub differentiation in the posterior SGPs within the embryonic gonad (Fig. 3 A and D). When these embryos were allowed to develop to adulthood, GSCs were normally observed in the testes (Fig. 3E and Table 1).
Fig. 3.
A decrease in the number of PGCs causes niche expansion. (A–C) Male embryonic gonads (stage 16/17) stained for Fas3 (green), Tj (red), and Vas (blue). (A) An embryo derived from a female homozygous for maternal gclrev390 mutation (gcl embryo). (B) gcl embryo carrying nos-GAL4-VP16. (C) gcl embryo carrying nos-GAL4-VP16 and UAS-Star (gcl, nos-GAL4 > UAS-Star embryo). UAS-Star was expressed in PGCs under the control of nos-GAL4-VP16. (Scale bar, 10 μm.) (D) The average number of Fas3-positive cells was plotted against the number of PGCs in a gonad of gcl (blue) and gcl, nos-GAL4 > UAS-Star embryos (red) at stage 16/17. The number of gonads examined in each case is shown in parentheses. Note that an increase in the number of Fas3-positive cells in gcl embryos was suppressed by UAS-Star expression in PGCs. When the gonad lacked pole cells (0), the values are not significantly different between gcl and gcl, nos-GAL4 > UAS-Star embryos. The average number of Fas3-positive cells per gonad carrying more than one pole cell ± SD is also shown. Significance was calculated using the Student's t test compared with wild type shown in Fig. 1M. (E–G) Testes from males (0–3 d after eclosion) derived from gcl (E), gcl, nos-GAL4 > UAS-Star (F), and nos-GAL4 > UAS-Star embryos (G). Testes were stained with anti-Fas3 (green) and anti-Vas (magenta). GSCs that were positive for Vas and associated with hub cells (green) were observed in the testes shown in E and G (arrowheads), but not in F. (Scale bar, 20 μm.)
Table 1.
Niche expansion is required to allocate GSCs
| Males | Number of testes examined | Number of testes lacking GSCs (%)* | Significance |
| gcl, nos-GAL4† | 200 | 0 (0) | |
| gcl, nos-GAL4 > UAS-Star‡ | 177 | 90 (50.8) | P < 0.001§ |
| nos-GAL4 > UAS-Star¶ | 200 | 0 (0) |
*For counting GSCs, adult testes were stained with anti-Vas, anti-Fas3, and 1B1 antibodies. The GSCs were identified as cells that were positive for Vas, contained a single spherical spectrosome marked by the 1B1 antibody, and were associated with Fas3-positive hub cells.
†The male flies were derived from gcl; nos-GAL4 homozygous mothers mated with wild-type males.
‡The male flies were derived from gcl; nos-GAL4 homozygous mothers mated with UAS-Star homozygous males.
§Significance was calculated versus gcl, nos-GAL4, and nos-GAL4 > UAS-Star by using the Fisher's exact probability test.
¶The male flies were derived from nos-GAL4 homozygous mothers mated with UAS-Star homozygous males.
These observations suggest that the ectopic hub cells increase the recruitment of PGCs as GSCs. If this is the case, repressing the niche expansion while reducing the number of PGCs produces adult males lacking GSCs. To repress niche expansion, we overexpressed Star in PGCs. We expected Star overexpression to cause up-regulation of Spi ligand production, which consequently activates Egfr in SGPs to prevent ectopic hub differentiation. Indeed, when Star was overexpressed in PGCs of gcl embryos, ectopic hub differentiation was repressed (Fig. 3 B and C), and the average number of Fas3-positive SGPs was statistically indistinguishable from that observed in wild-type embryos (Fig. 3D). In adult males derived from these embryos, approximately half of the testes lacked GSCs (Fig. 3F and Table 1). In contrast, Star overexpression alone did not affect GSC formation (Fig. 3G and Table 1). These observations show that niche expansion is required to recruit a decreased number of PGCs as GSCs in the developing male gonad.
Although a decrease in the number of PGCs induces ectopic hub differentiation in gcl embryos, the adults derived from these embryos have a normal number of hub cells and GSCs. We found that the average number of hub cells and GSCs per testis ± SD were 8.5 ± 0.9 and 7.5 ± 1.2 in gcl adults, and 8.9 ± 1.4 and 7.2 ± 1.0 in wild-type adults, respectively (20 testes were examined in each case). The values were not statistically different between gcl and wild type (P > 0.05, Student's t test). Thus, we speculate that there may be another mechanism regulating the proper number of hub cells and GSCs during postembryonic development. For example, it is possible that the hub cells which have successfully associated with PGCs (GSCs) may eliminate the extra hub cells.
Role of Notch, Egfr, and Sev Signaling Pathways in the Male Embryonic Gonad.
Our data show that Notch activation is necessary and sufficient to induce hub differentiation. Notch is activated in SGPs throughout the male embryonic gonad, and its activation requires Ser expression in the SGPs. Thus, Notch/Ser signaling among the SGPs induces hub differentiation. On the other hand, hub differentiation is normally repressed in the posterior SGPs both by Egfr and Sev. Reduction of Egfr or Sev activity results in ectopic hub differentiation in the posterior SGPs. Furthermore, Egfr alone, but not Sev, is sufficient to repress hub differentiation in the anterior SGPs. Thus, we propose that Egfr, rather than Sev, has a key role in repressing hub differentiation in SGPs.
Mutants that lack both Egfr and sev do not cause a significant increase in the number of Fas3-positive cells within the embryonic gonad, compared with Egfr single mutants (Fig. 2 M and N). This suggests that Egfr and Sev act synergistically, but not additively, to repress ectopic hub differentiation. In the developing eye, Sev sensitizes Egfr function by inactivating a repressor for a downstream gene of Egfr signaling (43). We speculate that Sev may potentiate Egfr signaling to repress hub differentiation in posterior SGPs. We further show that Egfr is necessary and sufficient for kek1 expression in the posterior SGPs, whereas kek1 expression is unaffected by sev mutation (Fig. S2 A–C). This suggests that Egfr and Sev differentially regulate transcription of target genes by using a different set of Ras/MAPK components. Further work will need not only to test this hypothesis but also to identify the target genes responsible for hub repression.
The question of how Egfr is activated in posterior SGPs still remains unanswered. Egfr is expressed in the anterior as well as the posterior SGPs, and spi, stet, and Star expression are all observed in PGCs. We found that kek1 expression requires the function of Abd-B (Fig. S2D), which specifies posterior cellular identities within the embryonic gonads (44, 45), suggesting that Abd-B is required to activate Egfr signaling pathway in the posterior SGPs. Because a constitutively active form of Egfr is able to induce kek1 expression in anterior SGPs lacking Abd-B function (Fig. S2B), it is likely that Abd-B is involved in Egfr activation itself, although we cannot rule out the possibility that Spi ligand production is enhanced in the posterior PGCs by Abd-B function. Abd-B may enhance a tight or stable physical association between PGCs and posterior SGPs which allows signal transfer between these cell types.
It is worthwhile to note that the number of PGCs incorporated into a pair of embryonic gonads varies from one embryo to another, and ranges from 10 to 40 (46–48). The total number of pole cells initially formed in blastodermal embryos is between 30 and 50. Of these, only a subset enters into the embryonic gonad, and the rest of the pole cells are eliminated by nonapoptotic cell death (49, 50). Furthermore, pole cells are competent to undergo apoptosis, and a reduction of maternal Nanos activity, which is essential for the establishment of germ-line fate, effectively triggers programmed cell death (51, 52). Thus, nonapoptotic and apoptotic death mechanisms eliminate aberrant and ectopic pole cells to maintain germ-line integrity. Here, we show that a decrease in the number of PGCs induces ectopic niche formation within the male embryonic gonad. The niche expansion could enhance their opportunity to recruit the decreased number of PGCs as GSCs. Indeed, we found that niche expansion is required to prevent loss of GSCs. We propose that signaling from PGCs to SGPs via Egfr acts as a key component of the mechanism securing GSCs.
It has been reported that interactions between the germ line and soma are essential for maintaining their proper ratio within the larval and adult gonad (20). In the larval ovary, signaling from PGCs to the somatic cells via Egfr positively regulates the number of somatic cells (53). In turn, the somatic cells negatively regulate PGC number by an unidentified signal transduction mechanism. Thus, positive and negative regulatory loops ensure that a sufficient number of PGCs occupy the niche to secure GSCs within the larval ovary. Furthermore, Egfr-mediated signaling from the germ line also facilitates their encapsulation by somatic support cells, which is essential for the germ-line cells to differentiate properly into gametes both in ovaries and testes (39, 54–56). In addition to Egfr pathway, Notch-mediated signal transduction from GSCs to the corresponding niche cells is essential for maintaining the number and function of the niche cells in ovaries (16). We now show that Notch and Egfr signaling act antagonistically to regulate proper formation of the GSC niche in male gonads earlier during embryogenesis. This study provides a striking example of how somatic gonad cells communicate with neighboring cells, including PGCs, to dynamically modulate niche formation during gonad development.
Materials and Methods
For fly stocks and in situ hybridization, see SI Materials and Methods.
Immunostaining.
Immunostaining of adult testes, embryos, and larvae was performed as previously described (17). Primary antibodies used were rabbit anti-Vas (a gift from P. Lasko, McGill University, Montreal, Canada, and A. Nakamura, RIKEN, Kobe, Japan) at 1:300, guinea pig anti-Tj (a gift from D. Godt, University of Toronto, Toronto, Canada) at 1:2,000, mouse anti-GFP (Sigma) at 1:300, rabbit anti-GFP (Molecular Probes) at 1:500, mouse anti-Egfr (E2906; Sigma) at 1:200, alkaline phosphatase-conjugated sheep anti-DIG (Roche) at 1:3,000, mouse anti-Sxl [M18; Developmental Studies Hybridoma Bank (DSHB), University of Iowa, Iowa City, IA] at 1:25, mouse anti-Fas3 (DSHB) at 1:50, mouse anti-Notch intracellular domain (C17.9C6; DSHB) at 1:200, mouse anti-β-galactosidase (40-1a; DSHB) at 1:50, and mouse anti-ADD87 antibodies (1B1; DSHB) at 1:50. Secondary antibodies used at 1:500 were goat anti-mouse, goat anti-rabbit, and goat anti-guinea pig conjugated to Alexa488, Alexa568, and Alexa647 (Molecular Probes). Sexual identity of embryos and larvae was determined by immunostaining for either Sxl expression, the presence of hub markers (Fas3), or by examining inheritance of an X-linked Kr-GFP transgene using an anti-GFP antibody (17).
Quantification of Fas3-Positive Cells and GSCs.
To count the number of Fas3-positive cells, embryos and adult testes were double-stained with anti-Fas3 antibody and anti-Tj, and were examined by confocal microscopy. The number of Fas3- and Tj-double-positive cells was then counted. The number of GSCs in adult testes was counted as described previously (17).
Supplementary Material
Acknowledgments
We thank the researchers who provided us with flies and antibodies. We also thank the Drosophila Genetic Resource Center, Kyoto, the Bloomington Stock Center, and the Developmental Studies Hybridoma Bank for fly stocks and antibodies. This work was partly supported by a Grant-in-Aid for Scientific Research on Innovative Areas, “Regulatory Mechanism of Gamete Stem Cells” (to S.K.), and a Grant-in-Aid for Research Activity Start-up (to Y.K.).
Footnotes
The authors declare no conflict of interest.
*This Direct Submission article had a prearranged editor.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1003462107/-/DCSupplemental.
References
- 1.Takashima S, Mkrtchyan M, Younossi-Hartenstein A, Merriam JR, Hartenstein V. The behaviour of Drosophila adult hindgut stem cells is controlled by Wnt and Hh signalling. Nature. 2008;454:651–655. doi: 10.1038/nature07156. [DOI] [PubMed] [Google Scholar]
- 2.Fox DT, Spradling AC. The Drosophila hindgut lacks constitutively active adult stem cells but proliferates in response to tissue damage. Cell Stem Cell. 2009;5:290–297. doi: 10.1016/j.stem.2009.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Mathur D, Bost A, Driver I, Ohlstein B. A transient niche regulates the specification of Drosophila intestinal stem cells. Science. 2010;327:210–213. doi: 10.1126/science.1181958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Xie T, Spradling AC. A niche maintaining germ line stem cells in the Drosophila ovary. Science. 2000;290:328–330. doi: 10.1126/science.290.5490.328. [DOI] [PubMed] [Google Scholar]
- 5.Kiger AA, Jones DL, Schulz C, Rogers MB, Fuller MT. Stem cell self-renewal specified by JAK-STAT activation in response to a support cell cue. Science. 2001;294:2542–2545. doi: 10.1126/science.1066707. [DOI] [PubMed] [Google Scholar]
- 6.Tulina N, Matunis E. Control of stem cell self-renewal in Drosophila spermatogenesis by JAK-STAT signaling. Science. 2001;294:2546–2549. doi: 10.1126/science.1066700. [DOI] [PubMed] [Google Scholar]
- 7.Tumbar T, et al. Defining the epithelial stem cell niche in skin. Science. 2004;303:359–363. doi: 10.1126/science.1092436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Calvi LM, et al. Osteoblastic cells regulate the haematopoietic stem cell niche. Nature. 2003;425:841–846. doi: 10.1038/nature02040. [DOI] [PubMed] [Google Scholar]
- 9.Zhang J, et al. Identification of the haematopoietic stem cell niche and control of the niche size. Nature. 2003;425:836–841. doi: 10.1038/nature02041. [DOI] [PubMed] [Google Scholar]
- 10.Li P, Zon LI. Resolving the controversy about N-cadherin and hematopoietic stem cells. Cell Stem Cell. 2010;6:199–202. doi: 10.1016/j.stem.2010.02.007. [DOI] [PubMed] [Google Scholar]
- 11.Mirzadeh Z, Merkle FT, Soriano-Navarro M, Garcia-Verdugo JM, Alvarez-Buylla A. Neural stem cells confer unique pinwheel architecture to the ventricular surface in neurogenic regions of the adult brain. Cell Stem Cell. 2008;3:265–278. doi: 10.1016/j.stem.2008.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Shen Q, et al. Adult SVZ stem cells lie in a vascular niche: A quantitative analysis of niche cell-cell interactions. Cell Stem Cell. 2008;3:289–300. doi: 10.1016/j.stem.2008.07.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tavazoie M, et al. A specialized vascular niche for adult neural stem cells. Cell Stem Cell. 2008;3:279–288. doi: 10.1016/j.stem.2008.07.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Brand U, Fletcher JC, Hobe M, Meyerowitz EM, Simon R. Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity. Science. 2000;289:617–619. doi: 10.1126/science.289.5479.617. [DOI] [PubMed] [Google Scholar]
- 15.Schoof H, et al. The stem cell population of Arabidopsis shoot meristems is maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell. 2000;100:635–644. doi: 10.1016/s0092-8674(00)80700-x. [DOI] [PubMed] [Google Scholar]
- 16.Ward EJ, et al. Stem cells signal to the niche through the Notch pathway in the Drosophila ovary. Curr Biol. 2006;16:2352–2358. doi: 10.1016/j.cub.2006.10.022. [DOI] [PubMed] [Google Scholar]
- 17.Kitadate Y, Shigenobu S, Arita K, Kobayashi S. Boss/Sev signaling from germline to soma restricts germline-stem-cell-niche formation in the anterior region of Drosophila male gonads. Dev Cell. 2007;13:151–159. doi: 10.1016/j.devcel.2007.05.001. [DOI] [PubMed] [Google Scholar]
- 18.Tanentzapf G, Devenport D, Godt D, Brown NH. Integrin-dependent anchoring of a stem-cell niche. Nat Cell Biol. 2007;9:1413–1418. doi: 10.1038/ncb1660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hardy RW, Tokuyasu KT, Lindsley DL, Garavito M. The germinal proliferation center in the testis of Drosophila melanogaster. J Ultrastruct Res. 1979;69:180–190. doi: 10.1016/s0022-5320(79)90108-4. [DOI] [PubMed] [Google Scholar]
- 20.Dansereau DA, Lasko P. The development of germline stem cells in Drosophila. Methods Mol Biol. 2008;450:3–26. doi: 10.1007/978-1-60327-214-8_1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wawersik M, et al. Somatic control of germline sexual development is mediated by the JAK/STAT pathway. Nature. 2005;436:563–567. doi: 10.1038/nature03849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Casper A, Van Doren M. The control of sexual identity in the Drosophila germline. Development. 2006;133:2783–2791. doi: 10.1242/dev.02415. [DOI] [PubMed] [Google Scholar]
- 23.Le Bras S, Van Doren M. Development of the male germline stem cell niche in Drosophila. Dev Biol. 2006;294:92–103. doi: 10.1016/j.ydbio.2006.02.030. [DOI] [PubMed] [Google Scholar]
- 24.Sheng XR, et al. Jak-STAT regulation of male germline stem cell establishment during Drosophila embryogenesis. Dev Biol. 2009;334:335–344. doi: 10.1016/j.ydbio.2009.07.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhu CH, Xie T. Clonal expansion of ovarian germline stem cells during niche formation in Drosophila. Development. 2003;130:2579–2588. doi: 10.1242/dev.00499. [DOI] [PubMed] [Google Scholar]
- 26.Song X, Call GB, Kirilly D, Xie T. Notch signaling controls germline stem cell niche formation in the Drosophila ovary. Development. 2007;134:1071–1080. doi: 10.1242/dev.003392. [DOI] [PubMed] [Google Scholar]
- 27.Chiba S. Notch signaling in stem cell systems. Stem Cells. 2006;24:2437–2447. doi: 10.1634/stemcells.2005-0661. [DOI] [PubMed] [Google Scholar]
- 28.Brower DL, Smith RJ, Wilcox M. Differentiation within the gonads of Drosophila revealed by immunofluorescence. J Embryol Exp Morphol. 1981;63:233–242. [PubMed] [Google Scholar]
- 29.Struhl G, Adachi A. Nuclear access and action of Notch in vivo. Cell. 1998;93:649–660. doi: 10.1016/s0092-8674(00)81193-9. [DOI] [PubMed] [Google Scholar]
- 30.Bray SJ. Notch signalling: A simple pathway becomes complex. Nat Rev Mol Cell Biol. 2006;7:678–689. doi: 10.1038/nrm2009. [DOI] [PubMed] [Google Scholar]
- 31.Gönczy P, DiNardo S. The germ line regulates somatic cyst cell proliferation and fate during Drosophila spermatogenesis. Development. 1996;122:2437–2447. doi: 10.1242/dev.122.8.2437. [DOI] [PubMed] [Google Scholar]
- 32.Glittenberg M, Pitsouli C, Garvey C, Delidakis C, Bray S. Role of conserved intracellular motifs in Serrate signalling, cis-inhibition and endocytosis. EMBO J. 2006;25:4697–4706. doi: 10.1038/sj.emboj.7601337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Freeman M. Reiterative use of the EGF receptor triggers differentiation of all cell types in the Drosophila eye. Cell. 1996;87:651–660. doi: 10.1016/s0092-8674(00)81385-9. [DOI] [PubMed] [Google Scholar]
- 34.Lüer K, Urban J, Klämbt C, Technau GM. Induction of identified mesodermal cells by CNS midline progenitors in Drosophila. Development. 1997;124:2681–2690. doi: 10.1242/dev.124.14.2681. [DOI] [PubMed] [Google Scholar]
- 35.Buff E, Carmena A, Gisselbrecht S, Jiménez F, Michelson AM. Signalling by the Drosophila epidermal growth factor receptor is required for the specification and diversification of embryonic muscle progenitors. Development. 1998;125:2075–2086. doi: 10.1242/dev.125.11.2075. [DOI] [PubMed] [Google Scholar]
- 36.Ghiglione C, et al. The transmembrane molecule kekkon 1 acts in a feedback loop to negatively regulate the activity of the Drosophila EGF receptor during oogenesis. Cell. 1999;96:847–856. doi: 10.1016/s0092-8674(00)80594-2. [DOI] [PubMed] [Google Scholar]
- 37.Ghiglione C, et al. Mechanism of inhibition of the Drosophila and mammalian EGF receptors by the transmembrane protein Kekkon 1. Development. 2003;130:4483–4493. doi: 10.1242/dev.00617. [DOI] [PubMed] [Google Scholar]
- 38.Urban S, Lee JR, Freeman M. A family of Rhomboid intramembrane proteases activates all Drosophila membrane-tethered EGF ligands. EMBO J. 2002;21:4277–4286. doi: 10.1093/emboj/cdf434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Schulz C, Wood CG, Jones DL, Tazuke SI, Fuller MT. Signaling from germ cells mediated by the rhomboid homolog stet organizes encapsulation by somatic support cells. Development. 2002;129:4523–4534. doi: 10.1242/dev.129.19.4523. [DOI] [PubMed] [Google Scholar]
- 40.Shilo BZ. Signaling by the Drosophila epidermal growth factor receptor pathway during development. Exp Cell Res. 2003;284:140–149. doi: 10.1016/s0014-4827(02)00094-0. [DOI] [PubMed] [Google Scholar]
- 41.Asaoka M, Lin H. Germline stem cells in the Drosophila ovary descend from pole cells in the anterior region of the embryonic gonad. Development. 2004;131:5079–5089. doi: 10.1242/dev.01391. [DOI] [PubMed] [Google Scholar]
- 42.Robertson SE, Dockendorff TC, Leatherman JL, Faulkner DL, Jongens TA. germ cell-less is required only during the establishment of the germ cell lineage of Drosophila and has activities which are dependent and independent of its localization to the nuclear envelope. Dev Biol. 1999;215:288–297. doi: 10.1006/dbio.1999.9453. [DOI] [PubMed] [Google Scholar]
- 43.Xu C, Kauffmann RC, Zhang J, Kladny S, Carthew RW. Overlapping activators and repressors delimit transcriptional response to receptor tyrosine kinase signals in the Drosophila eye. Cell. 2000;103:87–97. doi: 10.1016/s0092-8674(00)00107-0. [DOI] [PubMed] [Google Scholar]
- 44.Delorenzi M, Bienz M. Expression of Abdominal-B homeoproteins in Drosophila embryos. Development. 1990;108:323–329. doi: 10.1242/dev.108.2.323. [DOI] [PubMed] [Google Scholar]
- 45.DeFalco T, Le Bras S, Van Doren M. Abdominal-B is essential for proper sexually dimorphic development of the Drosophila gonad. Mech Dev. 2004;121:1323–1333. doi: 10.1016/j.mod.2004.07.001. [DOI] [PubMed] [Google Scholar]
- 46.Sonnenblick BP. In: Biology of Drosophila. Demerec M, editor. New York: Wiley; 1950. pp. 62–167. [Google Scholar]
- 47.Hay B, Ackerman L, Barbel S, Jan LY, Jan YN. Identification of a component of Drosophila polar granules. Development. 1988;103:625–640. doi: 10.1242/dev.103.4.625. [DOI] [PubMed] [Google Scholar]
- 48.Campos-Ortega JA, Hartenstein V. The Embryonic Development of Drosophila melanogaster. 2nd Ed. Heidelberg: Springer; 1997. [Google Scholar]
- 49.Sano H, Renault AD, Lehmann R. Control of lateral migration and germ cell elimination by the Drosophila melanogaster lipid phosphate phosphatases Wunen and Wunen 2. J Cell Biol. 2005;171:675–683. doi: 10.1083/jcb.200506038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Yamada Y, Davis KD, Coffman CR. Programmed cell death of primordial germ cells in Drosophila is regulated by p53 and the Outsiders monocarboxylate transporter. Development. 2008;135:207–216. doi: 10.1242/dev.010389. [DOI] [PubMed] [Google Scholar]
- 51.Sato K, et al. Maternal Nanos represses hid/skl-dependent apoptosis to maintain the germ line in Drosophila embryos. Proc Natl Acad Sci USA. 2007;104:7455–7460. doi: 10.1073/pnas.0610052104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Maezawa T, Arita K, Shigenobu S, Kobayashi S. Expression of the apoptosis inducer gene head involution defective in primordial germ cells of the Drosophila embryo requires eiger, p53, and loki function. Dev Growth Differ. 2009;51:453–461. doi: 10.1111/j.1440-169X.2009.01108.x. [DOI] [PubMed] [Google Scholar]
- 53.Gilboa L, Lehmann R. Soma-germline interactions coordinate homeostasis and growth in the Drosophila gonad. Nature. 2006;443:97–100. doi: 10.1038/nature05068. [DOI] [PubMed] [Google Scholar]
- 54.Kiger AA, White-Cooper H, Fuller MT. Somatic support cells restrict germline stem cell self-renewal and promote differentiation. Nature. 2000;407:750–754. doi: 10.1038/35037606. [DOI] [PubMed] [Google Scholar]
- 55.Tran J, Brenner TJ, DiNardo S. Somatic control over the germline stem cell lineage during Drosophila spermatogenesis. Nature. 2000;407:754–757. doi: 10.1038/35037613. [DOI] [PubMed] [Google Scholar]
- 56.Sarkar A, et al. Antagonistic roles of Rac and Rho in organizing the germ cell microenvironment. Curr Biol. 2007;17:1253–1258. doi: 10.1016/j.cub.2007.06.048. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.