Abstract
The production of blood cells depends on a rare hematopoietic stem-cell (HSC) population, but the molecular mechanisms underlying HSC biology remain incompletely understood. Here, we identify a subset of microRNAs (miRNAs) that is enriched in HSCs compared with other bone-marrow cells. An in vivo gain-of-function screen found that three of these miRNAs conferred a competitive advantage to engrafting hematopoietic cells, whereas other HSC miRNAs attenuated production of blood cells. Overexpression of the most advantageous miRNA, miR-125b, caused a dose-dependent myeloproliferative disorder that progressed to a lethal myeloid leukemia in mice and also enhanced hematopoietic engraftment in human immune system mice. Our study identifies an evolutionarily conserved subset of miRNAs that is expressed in HSCs and functions to modulate hematopoietic output.
Keywords: cancer, inflammation, myeloid, xenograft, noncoding RNA
Long-term production of blood cells depends on proper hematopoietic stem-cell (HSC) function (1). This involves a delicate balance between HSC self-renewal and differentiation into progenitor populations. The molecular networks that control these processes have begun to be elucidated in recent years, and several regulatory proteins have been shown to manage stem-cell physiology (2–6). Additionally, there is a substantial overlap between the proteins that regulate HSC biology and those that are dysregulated in cancers of immunological origin or other hematopoietic diseases, underlying the importance of these factors in maintaining homeostasis.
Mammalian noncoding microRNAs (miRNAs) have recently been identified as important repressors of gene expression through direct interactions with specific mRNA targets, and thus act to modulate gene function (7, 8). Several reports have found that miRNAs regulate the development of specific hematopoietic lineages (9). Conditional deletion of Dicer, which disrupts miRNA processing, has revealed critical roles for miRNAs during development of different hematopoietic lineages, including both B and T lymphocytes (10–12). Specific miRNAs have also been shown to impact development of distinct blood-cell lineages. Early B cell differentiation is modulated by miRNA (miR)-150 (13), miRNAs 17-5p-20a-106a direct human monocytes development (14), and miR-223 functions to restrict the granulocytic compartment in vivo (15).
Recent studies have shown a causative role for miRNAs in mediating malignant diseases in the hematopoietic system. For example, sustained expression of miR-155 or miR-29a in the mouse hematopoietic system causes a myeloproliferative disorder (16) or leukemia (17), respectively, whereas a B cell malignancy is triggered by constant expression of miR-155 in a B cell-restricted manner (18). However, tumor suppressor miRNAs like miR-15a/16-1 are deleted in a subset of lymphomas (19), which has been shown to cause chronic lymphocytic leukemia in mice (20).
Despite their being clearly linked to hematopoietic development and cancer, there is little known regarding the expression patterns and function of mammalian miRNAs in HSC populations. One study found that mice deficient in Ars2, which is required for proper miRNA processing, suffer from bone-marrow failure, suggesting a functional role for miRNAs in HSCs (21). Other groups have profiled miRNA expression in human CD34+ cells (22). However, many of the identified miRNAs are broadly expressed outside of HSCs, and their functional relevance in stem cells was not investigated in vivo. In the present study, we sought to both find specific miRNAs that are enriched in long-term HSCs and investigate their impact on long-term hematopoietic reconstitution.
Results
Identification of miRNAs Enriched in HSCs.
To identify miRNAs enriched in hematopoietic stem and progenitor cells (HSPCs), we isolated lineage-negative (lin−) cKit+Sca1+ (LKS) cells from adult C57BL6 mouse BM using FACS (Fig. 1A Upper). Total RNA was isolated from LKS cells and also from RBC-depleted total bone marrow (BM), and it was used to perform a microarray analysis to identify expression of mouse miRNAs. Some 137 miRNAs were expressed at detectable levels in the BM and/or LKS populations. Of these, 67 were expressed at higher levels in total BM vs. LKS cells, 59 were expressed at similar levels in the two cellular populations, and 11 were expressed at higher levels in the LKS compartment vs. total BM (Fig. 1A Lower and Table S1). Corroborating the microarray results, quantitative (q) PCR detected enrichment of these 11 miRNAs in LKS cells. They are miR125a-5p, miR-125b-5p, miR-155, miR-130a, miR196b, miR-99a, miR-126-3p, miR-181c, miR-193b, miR-542-5p, and let7e (Fig. 1B). A more extensive cellular fractionation of mouse BM further confirmed enrichment of most of these miRNAs in the LKS subset (Fig. 1C and Fig S1). Expression of miR-223, known to be enriched in mature myeloid cells, was measured as a control (Fig. 1C). The LKS population is comprised of both HSCs, capable of establishing long-term multilineage hematopoietic engraftment, and multipotent progenitors, with reduced capacity to reconstitute the hematopoietic system. LKS cells expressing the HSC marker endothelial protein C receptor (EPCR) (23) were first analyzed, and a higher relative expression of many of the miRNAs under study was found in EPCR+ vs. EPCR-LKS cells (Fig. 1C). Long-term HSCs have also been shown to be CD150+CD48-, whereas other LKS progenitor populations are negative for CD150 (24). Again, qPCR analysis found that most of the HSPC miRNAs were further enriched in CD150+CD48- long-term HSCs compared with CD150-LKS cells (Fig. 1D). These data indicate, using multiple criteria to define HSCs, that this subset of miRNAs is largely enriched in long-term HSCs.
Fig. 1.
Identification of miRNAs enriched in HSCs. (A) Microarray experiment comparing miRNA expression in the LKS compartment (FACS plot in Upper and subset boxed in red) with total bone marrow from C57BL6 mice. Number of miRNAs significantly enriched in total BM vs. the LKS compartment, unchanged between the two groups, or significantly enriched in LKS vs. total BM cells are shown (Lower). BM cells were pooled from 10 mice and sorted, and RNA was extracted for the microarray. (B) qPCR was used to determine the fold enrichment of the 11 miRNAs preferentially expressed in LKS cells vs. total BM. Data represent two independent experiments. (C) Expression levels of the 11 HSPC miRNAs were measured in hematopoietic cells from the different bone-marrow compartments shown in Fig. S1 and represent different stages of blood-cell development. (D) LKS-gated cells were further sorted by FACS into three groups according to their expression of the signaling lymphocytic activation molecule (SLAM) markers CD150 and CD48. Total RNA was collected from each group, and the relative expression levels of miRNAs from each LKS subpopulation were assayed by qPCR. Data represent two independent experiments, each starting with pooled BM from 20 mice. All data were normalized to sno202 and represent the mean + SEM.
Gain-of-Function Approach to Assess the Impact of HSC miRNAs on Long-Term Hematopoietic Engraftment.
The expression patterns of these particular miRNAs suggest that they have a specialized role in regulating hematopoiesis at the stem-cell level. To ascertain the functional impact of these HSC miRNAs on long-term hematopoietic reconstitution, we constructed a series of murine stem-cell virus (MSCV)-based miRNA-expression vectors that carry each of the 11 miRNAs under study. The vectors were formatted with mouse miR-155 arms and stem loop sequences, with each miRNA duplex encoded by the sense and antisense stem regions (Fig. 2A). This format was chosen because of its established ability to be processed in hematopoietic cells (25), including HSCs, where we have found mature miR-155 to be expressed (Fig. 1). Production of the different mature miRNAs using this format was confirmed in 293T cells by qPCR (Fig. S2).
Fig. 2.
Gain-of-function approach to assess the impact of HSC miRNAs on long-term hematopoietic reconstitution. (A) HSC miRNAs were cloned into an MSCV-based retroviral vector (MG) using an miR-155 arms and loop format. (B) Competitive BM reconstitutions were performed using C57BL6 mice to assess the engraftment potential of BM expressing each of the 11 HSC miRNAs (CD45.1 cells) compared with control BM (CD45.2 cells). GFP was used to identify cells containing a vector. A 4-mo time course showing the percentage of CD45.1+GFP+ cells in the peripheral blood of each group of mice is shown. The experiment was broken into two batches (Upper and Lower), each with its own negative control. (C) Lineage analysis of the CD45.1+GFP+ peripheral blood cells at the 4-mo time point was assessed by determining the percentages of B220+ (B cell), CD3+ (T cell), and CD11b+ (myeloid cell) in each group. Data represent the mean + SEM. Experiments had four mice per group.
Competitive repopulation experiments were performed by combining equal numbers of HSC-enriched CD45.1+ bone-marrow cells transduced with retroviruses encoding a specific HSC miRNA with HSC-enriched CD45.2+ bone marrow transduced with a control retrovirus. This cellular mixture was used to reconstitute lethally irradiated CD45.2+ C57BL6 recipients. Because the retrovectors coexpress GFP in addition to a miRNA, the ability of the CD45.1+ miRNA-expressing bone marrow to compete with CD45.2+ control vector-containing bone marrow during hematopoietic engraftment could be tracked by FACS analysis of peripheral blood cells. Mice were bled at 2 and 4 mo postreconstitution. When both vectors lacked a miRNA, there was similar representation of the two markers in the peripheral blood mononuclear cells (Fig. S3). In three cases, including miR-125b-5p, miR-126-3p, and miR-155, the miRNA conferred an evident and statistically significant (P < 0.05) competitive advantage to the engrafted bone marrow (Fig. 2B and Fig. S3). In contrast, overexpression of several of the miRNAs caused a significant (P < 0.05) disadvantage to the engrafted bone marrow, with miR-196b, miR-181c, let7e, and 542-5p having the largest impact (Fig. 2B and Fig. S3). FACS analysis of the peripheral blood was also used to determine the distribution of different hematopoietic lineages to assess whether engraftment was being skewed to a single lineage or was multilineage at 4 mo postreconstitution. The effects of the different miRNAs on engraftment were largely multilineage, consistent with these miRNAs regulating stem-cell homeostasis (Fig. 2C).
miR-125b Causes a Dose-Dependent Myeloproliferative Disorder That Progresses to a Myeloid Leukemia.
We next focused on further characterization of miR-125b, because it had the largest positive impact on competitive reconstitution among the miRNAs tested and is overexpressed in certain types of acute myeloid leukemia (AML), suggesting that it plays an active role in human malignancies (26, 27). miR-125b is expressed from two loci in the genome, and these sequences are referred to as miR-125b1 and miR-125b2. To assess the impact of expressing miR-125b with its native arms and loop sequences, we cloned segments of the miR-125b1 and miR-125b2 primary transcripts into our retroviral expression system (Fig. 3A). We found that these cassettes led to substantially increased expression levels of mature miR-125b in K562 cells compared with the miR-125b cassette formatted with miR-155 arms and loop sequences [used above and now referred to as 125b(f)], likely a result of more efficient processing (Fig. 3A). Furthermore, miR-125b1 expression was higher than miR-125b2, providing three vectors with conveniently graded expression of miR-125b.
Fig. 3.
miR-125b causes a dose-dependent MPD. (A) The three constructs used to enforce expression of miR-125b are shown. K562 cells were transduced with each retrovector, and miR-125b levels were assayed by qPCR. (B) Blood concentrations of the indicated cell types are shown for mice expressing MG, MG-125b1, MG-125b2, and in a separate experiment (separated by a dashed line), MG and MG-125b(f) 2-mo postreconstitution. (C) FACS plot showing a representative FACS plot of Gr1+CD11b+ cells in the peripheral blood of mice overexpressing miR-125b2, miR-125b1, or control vector 2 mo postreconstitution. Asterisk denotes a P value < 0.05.
Mice were subsequently reconstituted with bone marrow expressing miR-125b1, miR-125b2, or control vector, and a separate cohort of mice expressing miR-125b(f) or its vector control was also established. All mice from both groups were bled after 2 mo of reconstitution, and complete blood cell counts (CBCs) were recorded (Fig. 3B). Mice expressing miR-125b, regardless of the construct used, had subtly elevated total WBC counts compared with control mice. Reconstitution of mice with HSPCs expressing the lowest levels of miR-125b, miR-125b(f), resulted in a general increase in all WBC compartments analyzed, including myeloid, lymphoid, and platelets, but did not impact RBCs. However, mice with the more highly expressed endogenous miR-125b1 or b2 vectors exhibited a clear myeloproliferative disorder (MPD) that was characterized by increased absolute numbers of monocytes and neutrophils and decreased levels of lymphocytes, RBCs, and platelets. Peripheral blood from miR-125b-1– and miR-125b-2–expressing mice, compared with control mice, also had dramatic increases in the percentage of CD11b+ myeloid cells, many of which were also Gr1+ as assayed by FACS (Fig. 3C).
Analysis of Wright-stained bone-marrow smears from miR-125b(f) mice 2 mo postreconstitution revealed a mild expansion of myeloid cells in the bone marrow, whereas H&E-stained spleen sections were similar to those from control mice (Fig. S4). However, miR-125b1– and miR-125b2–expressing mice exhibited myeloid-dominated BM and spleens as well as myeloid-cell infiltration into the liver (Fig. S4). By 3.5 mo postreconstitution, miR-125b1 and miR-125b2 mice had substantially increased WBCs in their peripheral blood consisting mainly of immature granulocytic and monocytic cells (Fig. S4). Shortly after, the condition became lethal for most of the mice expressing the highest levels of miR-125b, with tumor infiltration evident in the spleen and liver (Fig. 4 A and B). The surviving miR-125b2 mice had progressively increasing WBC counts and higher levels of leukemic blasts in their peripheral blood from 3.5 to 4.5 mo postreconstitution and also eventually succumbed to leukemia (Fig. 4 A and C). Furthermore, transfer of malignant BM from moribund miR-125b1–expressing mice to immunodeficient Rag2−/−γc−/− secondary recipients resulted in death within weeks, whereas transfer of BM from control mice had no impact on survival (Fig. 4D). These findings show a dose-dependent sufficiency of miR-125b to drive pathological myeloid-cell expansion, a condition that progresses to an aggressive myeloid leukemia.
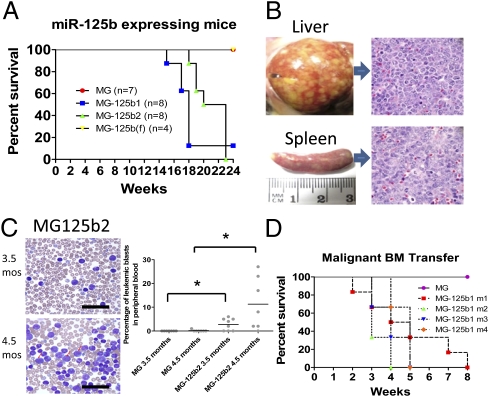
Fig. 4.
MPDs caused by miR-125b progress to myeloid leukemia in a dose-dependent manner. (A) Survival of mice expressing different levels of miR-125b. (B) Liver and spleen from miR-125b2–expressing mice with leukemia 4.5 mo after reconstitution. Representative H&E-stained tissue sections of each respective organ taken from miR-125b2–expressing mice are shown on Right. (C) Percentage of leukemic blasts in the peripheral blood of control or miR-125b2–expressing mice 3.5 and 4.5 mo postreconstitution, and representative Wright-stained blood smears from miR-125b2 mice at each time point. Examples of blasts are indicated by red arrows. (D) Survival of mice after i.v. transfer of malignant MG-125b1 (four different donors) or MG control bone marrow to Rag2−/−γc−/− recipients (n = 3–6 mice per group). Asterisk denotes a P value < 0.05.
Evolutionarily Conserved miRNA Expression and Function in Human CD34+ HSPCs.
We investigated whether the subset of miRNAs enriched in mouse HSCs was also enriched in human CD34+ HSPCs. RNA was prepared from CD34+ and CD34− umbilical cord blood (CB) peripheral blood mononuclear cell (PBMC)s and analyzed by qPCR. With the exception of miR-193b, all of the miRNAs showed significant enrichment in the CD34+ HSPC cellular subset compared with mature CD34− PBMCs (Fig. 5A). To determine whether miR-125b could also impact human hematopoietic engraftment, CD34+ CB cells were transduced with control or a miR-125b1–expressing lentiviral vector (Fig. 5B) and subsequently injected intrahepatically into newborn Rag2−/−γc−/− mice on a BALB/c genetic background generating the previously described human immune system (HIS) model (28). After 10 wk, peripheral blood was collected and analyzed for the presence of human CD45+ WBCs by FACS, and a dose-dependent enhancement in multilineage engraftment was observed in miR-125b1–expressing vs. control HIS mice (Fig. 5 C–E). At 12 wk postreconstitution, the high-dose (multiplicity of infection [MOI] 40) groups were harvested, and miR-125b–expressing HIS mice had significantly elevated levels of human (h) CD45+ cells and hCD34+ HSPCs in their bone marrow (Fig. 5 F and G). These data provide evidence that miR-125b also promotes hematopoietic engraftment of human HSCs.
Fig. 5.
Evolutionarily conserved miRNA expression and function in human CD34+ HSPCs. (A) Human CB was sorted into CD34+ (HSPC+) and CD34− (HSPC−) fractions. Expression of the miRNAs enriched in mouse HSCs was measured by qPCR. Data are normalized to RNU48 and presented as mean + SEM (n = 3 different donors). (B) Human CB CD34+ cells were transduced with control or miR-125b1–expressing lentiviral vectors at an MOI of ~4 or 40, and miR-125b levels were assayed by qPCR. (C) Representative FACS plots showing an increased ratio of human to mouse CD45+ WBCs in the peripheral blood of miR-125b–expressing HIS mice 10 wk after CD34+ cell injection. (D) A gray dot represents each mouse. (E) Human cells in the peripheral blood of high-dose (MOI 40) HIS mice were analyzed by FACS for the expression of different lineage markers including CD19 (B cells) and CD33 (myeloid). Data are represented as the mean ± SEM. (F) A representative FACS plot of hCD45+ and hCD34+ cells in the BM of miR-125b–expressing and control vector HIS mice 12 wk post-CD34+ injection. (G) A gray dot represents each mouse. Asterisk denotes a P value < 0.05.
Discussion
In this work, we have identified an evolutionarily conserved subset of miRNAs that is enriched in HSCs and have provided evidence that these miRNAs function to properly manage hematopoietic output. Because some miRNAs promoted whereas others diminished hematopoietic engraftment, it is possible that these HSC miRNAs work in a concerted manner to ensure proper output of blood cells. A similar paradigm has recently been shown for ES cells, where opposing miRNA families modulate ES cell self-renewal and differentiation (29). This suggests that regulatory networks involving multiple miRNAs participate in the control of both embryonic and adult stem cells.
Owing to their important developmental roles and ability to target known oncogenes and tumor suppressors (9), miRNAs are a key part of the intricate molecular networks that drive or suppress cancer development and progression. Certain hematopoietic cancers, such as chronic myeloid leukemia (CML), are sustained by rare cancer stem-cell populations that have significant mRNA expression overlap with normal HSCs, despite producing malignant progeny (30). Our newly identified set of HSC miRNAs might prove to be useful as biomarkers of such cancer stem-cell populations. In the case of miR-125b, it will also be of interest to assess whether it transforms HSCs or a committed myeloid progenitor and to characterize secondary mutations that may assist in this process.
Many of the miRNAs identified in our study, including miR-125b, miR-155, miR-126, miR-196b, and miR-99a, have been shown to be dysregulated in different subsets of AML (31). Because we have shown functional relevance for many of these HSC miRNAs in vivo, including the abilities of miR-155 (16) and now miR-125b to trigger aggressive MPDs and myeloid leukemia, respectively, therapeutic targeting of this subset may be effective in combating hematopoietic cancers.
Materials and Methods
All animal experiments were approved by the institution animal care and use committee (IACUC) at Caltech. HSPCs were isolated from C57BL6 bone marrow using MACS (Miltenyi) followed by FACS sorting (antibodies described in Table S2), and total RNA was collected from equal numbers of cells by miRNeasy (Qiagen) and was used to quantitate miRNA expression by microarray (Agilent) and qPCR (ABI Taqman). Cell culture, retroviral transduction, bone-marrow transplants, CBCs, FACS, and histopathology were performed as described previously (16). For competitive bone-marrow reconstitutions, C57BL6 mice expressing the congenic markers CD45.1 and CD45.2 were used as bone-marrow donors, and CD45.2 mice were used as recipients. Equal numbers of 5-Fluorouracil (5-FU)–treated bone-marrow cells from each group were infected with an miRNA-expressing (CD45.1 cells) or control vector (CD45.2 cells), and a 1:1 cell mixture was injected i.v. into lethally irradiated mice. miRNA-expression vectors formatted with miR-155 loop and arms were constructed as described previously (Table S3 shows oligonucleotide sequences) (25). Endogenous miR-125b sequences were obtained from System Biosciences and were PCR cloned into MG and MGP retrovectors or a third-generation replication-deficient lentiviral vector. Human cell work was approved by the Institute Biosafety Committee (IBC) at Caltech. HIS mice were generated by injecting 2 × 105 lentivirus transduced CD34+ CB cells i.h. into irradiated newborn Rag2−/−γc−/− mice on a BALB/c genetic background as described (28). Expanded methods are found in SI Materials and Methods.
Supplementary Material
Acknowledgments
We thank the Caltech FACS core facility for their assistance with cell sorting. R.M.O. was funded in part by the Irvington Institute Fellowship Program of the Cancer Research Institute and by Award K99HL102228 from the National Heart, Lung, and Blood Institute. A.A.C. was funded by the Graduate Research Fellowship Program of the National Science Foundation. D.S.R. was funded by Award 1K08CA133521 from the National Cancer Institute. A.B.B. is supported by American Foundation for AIDS Research (amfAR) fellowship #107756-47-RFVA. This work was also supported by National Institutes of Health Grant 1R01AI079243-01.
Footnotes
Conflict of interest statement: The authors declare that they have no competing financial interests except for D.B., who is a scientific advisor to Regulus Therapeutics, a company devoted to microRNA therapeutics.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1009798107/-/DCSupplemental.
References
- 1.Orkin SH, Zon LI. Hematopoiesis: An evolving paradigm for stem cell biology. Cell. 2008;132:631–644. doi: 10.1016/j.cell.2008.01.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Santaguida M, et al. JunB protects against myeloid malignancies by limiting hematopoietic stem cell proliferation and differentiation without affecting self-renewal. Cancer Cell. 2009;15:341–352. doi: 10.1016/j.ccr.2009.02.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zhang J, et al. PTEN maintains haematopoietic stem cells and acts in lineage choice and leukaemia prevention. Nature. 2006;441:518–522. doi: 10.1038/nature04747. [DOI] [PubMed] [Google Scholar]
- 4.Ramalho-Santos M, Yoon S, Matsuzaki Y, Mulligan RC, Melton DA. “Stemness:” Transcriptional profiling of embryonic and adult stem cells. Science. 2002;298:597–600. doi: 10.1126/science.1072530. [DOI] [PubMed] [Google Scholar]
- 5.Park IK, et al. Bmi-1 is required for maintenance of adult self-renewing haematopoietic stem cells. Nature. 2003;423:302–305. doi: 10.1038/nature01587. [DOI] [PubMed] [Google Scholar]
- 6.Deneault E, et al. A functional screen to identify novel effectors of hematopoietic stem cell activity. Cell. 2009;137:369–379. doi: 10.1016/j.cell.2009.03.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bartel DP, Chen CZ. Micromanagers of gene expression: The potentially widespread influence of metazoan microRNAs. Nat Rev Genet. 2004;5:396–400. doi: 10.1038/nrg1328. [DOI] [PubMed] [Google Scholar]
- 8.Winter J, Jung S, Keller S, Gregory RI, Diederichs S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol. 2009;11:228–234. doi: 10.1038/ncb0309-228. [DOI] [PubMed] [Google Scholar]
- 9.O'Connell RM, Rao DS, Chaudhuri AA, Baltimore D. Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol. 2010;10:111–122. doi: 10.1038/nri2708. [DOI] [PubMed] [Google Scholar]
- 10.Cobb BS, et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J Exp Med. 2005;201:1367–1373. doi: 10.1084/jem.20050572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Koralov SB, et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell. 2008;132:860–874. doi: 10.1016/j.cell.2008.02.020. [DOI] [PubMed] [Google Scholar]
- 12.Muljo SA, et al. Aberrant T cell differentiation in the absence of Dicer. J Exp Med. 2005;202:261–269. doi: 10.1084/jem.20050678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Xiao C, et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell. 2007;131:146–159. doi: 10.1016/j.cell.2007.07.021. [DOI] [PubMed] [Google Scholar]
- 14.Fontana L, et al. MicroRNAs 17-5p-20a-106a control monocytopoiesis through AML1 targeting and M-CSF receptor upregulation. Nat Cell Biol. 2007;9:775–787. doi: 10.1038/ncb1613. [DOI] [PubMed] [Google Scholar]
- 15.Johnnidis JB, et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature. 2008;451:1125–1129. doi: 10.1038/nature06607. [DOI] [PubMed] [Google Scholar]
- 16.O'Connell RM, et al. Sustained expression of microRNA-155 in hematopoietic stem cells causes a myeloproliferative disorder. J Exp Med. 2008;205:585–594. doi: 10.1084/jem.20072108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Han YC, et al. microRNA-29a induces aberrant self-renewal capacity in hematopoietic progenitors, biased myeloid development, and acute myeloid leukemia. J Exp Med. 2010;207:475–489. doi: 10.1084/jem.20090831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Costinean S, et al. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc Natl Acad Sci USA. 2006;103:7024–7029. doi: 10.1073/pnas.0602266103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Calin GA, et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2002;99:15524–15529. doi: 10.1073/pnas.242606799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Klein U, et al. The DLEU2/miR-15a/16-1 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell. 2010;17:28–40. doi: 10.1016/j.ccr.2009.11.019. [DOI] [PubMed] [Google Scholar]
- 21.Gruber JJ, et al. Ars2 links the nuclear cap-binding complex to RNA interference and cell proliferation. Cell. 2009;138:328–339. doi: 10.1016/j.cell.2009.04.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Georgantas RW, 3rd, et al. CD34+ hematopoietic stem-progenitor cell microRNA expression and function: A circuit diagram of differentiation control. Proc Natl Acad Sci USA. 2007;104:2750–2755. doi: 10.1073/pnas.0610983104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Balazs AB, Fabian AJ, Esmon CT, Mulligan RC. Endothelial protein C receptor (CD201) explicitly identifies hematopoietic stem cells in murine bone marrow. Blood. 2006;107:2317–2321. doi: 10.1182/blood-2005-06-2249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kiel MJ, et al. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell. 2005;121:1109–1121. doi: 10.1016/j.cell.2005.05.026. [DOI] [PubMed] [Google Scholar]
- 25.O'Connell RM, Chaudhuri AA, Rao DS, Baltimore D. Inositol phosphatase SHIP1 is a primary target of miR-155. Proc Natl Acad Sci USA. 2009;106:7113–7118. doi: 10.1073/pnas.0902636106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Klusmann JH, et al. miR-125b-2 is a potential oncomiR on human chromosome 21 in megakaryoblastic leukemia. Genes Dev. 2010;24:478–490. doi: 10.1101/gad.1856210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bousquet M, et al. Myeloid cell differentiation arrest by miR-125b-1 in myelodysplastic syndrome and acute myeloid leukemia with the t(2;11)(p21;q23) translocation. J Exp Med. 2008;205:2499–2506. doi: 10.1084/jem.20080285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Traggiai E, et al. Development of a human adaptive immune system in cord blood cell-transplanted mice. Science. 2004;304:104–107. doi: 10.1126/science.1093933. [DOI] [PubMed] [Google Scholar]
- 29.Melton C, Judson RL, Blelloch R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature. 2010;463:621–626. doi: 10.1038/nature08725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Savona M, Talpaz M. Getting to the stem of chronic myeloid leukaemia. Nat Rev Cancer. 2008;8:341–350. doi: 10.1038/nrc2368. [DOI] [PubMed] [Google Scholar]
- 31.Cammarata G, et al. Differential expression of specific microRNA and their targets in acute myeloid leukemia. Am J Hematol. 2010;85:331–339. doi: 10.1002/ajh.21667. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.