Abstract
Intrapatient variability of the attachment (G) protein gene of respiratory syncytial virus (RSV) was examined using both population and single-genome sequencing. Samples from three patients infected with a group B virus variant which has a 60-nucleotide duplication in the G protein gene were examined. These samples were chosen because occasional mixed sequence bases were observed. In a minority of RSV genomes from these patients considerable variability was found, including point mutations, insertions, and deletions. Of particular note, the deletion of the exact portion of the gene which had been duplicated in some isolates was observed in viral RNAs from two patients.
Human respiratory syncytial virus (RSV) is a major cause of lower respiratory tract infection in infants and vulnerable adults (3, 9) and is unusual in that it can repeatedly reinfect individuals (5, 6). RSV isolates are classified into two groups, A and B, and the attachment (G) protein, a target for neutralizing antibodies, is the most variable of the viral proteins, showing considerable genetic and antigenic variability both within and between the groups (7, 8). The G protein is able to accommodate drastic changes, which have been observed both in culture during the selection of monoclonal antibody escape mutants (4, 12, 13) and in vivo with the emergence of new variants, including a group B strain with a duplication of 60 nucleotides (17). This strain with a 60-nucleotide duplication was first reported from Buenos Aires in 1999 (17) and then was subsequently detected in samples from 1998 in Madrid (16). The strain then became the dominant group B strain worldwide, indicating a selective advantage for this variant (16, 18). Thus, major genetic changes can be introduced into the G gene sequence while the virus replicates in its natural host, which can then be selected under favorable epidemiological conditions.
Previous investigations of the genetic diversity of RSV exploited direct sequencing of PCR-amplified products (2), which represent the population average of the in vivo variants. Such sequences are derived from multiple copies of cDNA and represent the dominant sequence, and they thus do not allow detection of minority populations below about 20% prevalence (11). Information on intrapatient viral diversity during infections may therefore be missed, knowledge of which could be important in the overall understanding of the genetic diversity of this virus. We report here the analysis of individual RSV RNA molecules derived by single-genome amplification (SGA) and sequencing from clinical samples using a methodology developed for the analysis of HIV genomes (11, 14).
RSV-positive samples were collected from infants admitted to Kilifi District Hospital, Kenya (10). Viral RNA extraction and cDNA synthesis were carried out as previously described (15). For population sequencing, a nested PCR was carried out on the cDNA using primers that amplified the ectodomain-coding part of the G protein gene, with the PCR product being directly sequenced. In the 2007-2008 RSV epidemic in Kilifi, group B viruses were predominant. By population sequencing of ∼100 group B samples, all were found to have the 60-nucleotide duplication observed in the Buenos Aires variant (data not shown). However, in some specimens there were some mixed bases at some positions, so the variability at the level of the single cDNA molecule was further investigated.
Three samples that gave occasional mixed signals in the sequence chromatograms were further analyzed by SGA and sequencing. For SGA the cDNAs were serially diluted 3-fold up to 1:6,361. Ten nested PCRs were carried out on each dilution using Platinum high-fidelity PCR Supermix (Invitrogen) (containing Taq polymerase together with the proofreading enzyme Pyrococcus species GB-D polymerase). Based on the Poisson distribution, it has been shown that for a sample dilution yielding approximately 30% positive PCRs there is an 80% likelihood that each PCR is derived from a single cDNA molecule (11). For each of the identified endpoint dilutions, the cDNA was amplified in 80 separate nested PCRs using the high-fidelity enzyme and the positive reaction products sequenced. The nomenclature for the sequences reported in this paper is place of isolation (Kenya [Ken])/year of isolation/strain number. For SGA sequences an additional Roman number is given.
The predicted length derived by population sequencing of the G proteins of the three samples examined by SGA was 310 amino acids, showing a 6-nucleotide deletion and a changed stop codon relative to the Buenos Aires strain (Fig. 1). The dominant sequences represented 60 to 88% of the sequences derived by SGA. The differences were due to point mutations, duplications, and deletions, as summarized in Table 1; the consequences of these changes for the predicted length of the G protein are shown in Table 2.
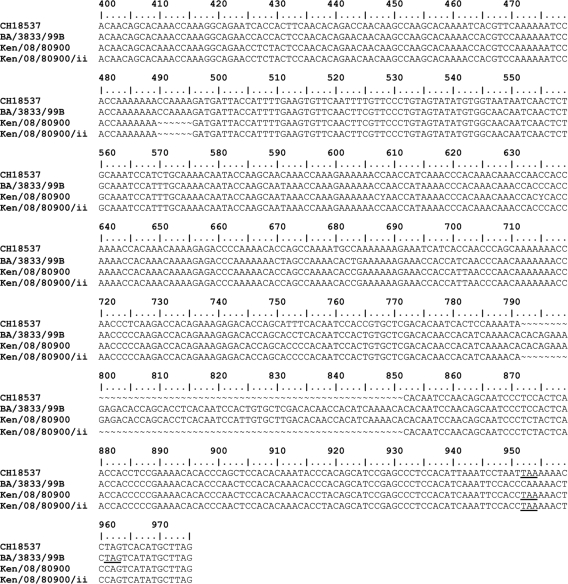
FIG. 1.
Nucleotide sequence alignment of part of the G protein gene (from nucleotide 400) of the sample 2 population sequence (Ken/08/80900) and a minority sequence (Ken/08/80900/ii), with the sequences of prototype group B strain CH18537 (accession number M17213) and Buenos Aires strain BA/3833/99B (accession number AY333362). This shows the duplication of 60 nucleotides in the Kenyan and Buenos Aires viruses relative to CH18537 and the loss of the same 60 nucleotides in the Kenyan minority sequence. Termination codons are underlined.
TABLE 1.
Diversity in the SGA-derived sequences
| Strain | Total no. of SGA-derived sequences | No. of: |
|||
|---|---|---|---|---|---|
| Variants | Substitutions | Insertions | Deletions | ||
| Ken/07/80395 | 17 | 3 | 1 | 1a | 0 |
| Ken/08/80900 | 12 | 3 | 1 | 0 | 1b |
| Ken/08/80767 | 15 | 6 | 3 | 0 | 4c |
The insertion involved an 8-nucleotide duplication.
The deletion was of 60 nucleotides.
The deletions ranged from 1 to 104 nucleotides, including one of 60 nucleotides.
TABLE 2.
Summary of nucleotide and predicted amino acid differences observed in single-genome amplification sequences
| Strain | Sequence no. | Mutation type (nucleotide position) | Effect on predicted polypeptide |
|---|---|---|---|
| Ken/07/80395 | Ken/07/80395/ii | Substitution (264) | None |
| Ken/07/80395/iii | Duplication (599-606) | Frameshift with a subsequent premature stop codon | |
| Ken/08/80900 | Ken/08/80900/ii | Deletion (792-851) | Reduction of length by 20 amino acids |
| Ken/08/80900/iii | Substitution (539) | Isoleucine-to-threonine change | |
| Ken/08/80767 | Ken/08/80767/ii | Substitution (592) | Immediate premature stop codon |
| Ken/08/80767/iii | Substitution (233) | Threonine-to-serine change | |
| Ken/08/80767/vi | Substitution (664) | Immediate premature stop codon | |
| Ken/08/80767/v | Deletion (599) | Frameshift with a subsequent premature stop codon | |
| Ken/08/80767/vi | Deletion (599) | Frameshift with a subsequent premature stop codon | |
| Deletion (792-851) | No effect due to earlier frameshift; otherwise this would cause a 20-amino-acid deletion | ||
| Ken/08/80767/vii | Deletion (836-939) | Frameshift, with no subsequent stop codon within the region sequenced |
For sample 1 (Ken/07/80395), 17 separate PCR products from SGA were sequenced. Sequence Ken/07/80395/i was the dominant sequence, occurring in 15/17 (88%) of the products. A synonymous mutation (T-C at 264) was seen in Ken/07/80395/ii, while Ken/07/80395/iii had an 8-nucleotide duplication starting at position 599, which resulted in a frameshift with a subsequent premature stop codon.
For sample 2 (Ken/08/80900), 12 separate SGA PCR products were sequenced, with the dominant sequence (Ken/08/80900/i) occurring in 9/12 (75%) of the products. One minority sequence (Ken/08/80900/ii) had a 60-nucleotide deletion starting at position 792. This deletion was identical to the duplication that characterizes this variant (Fig. 1) (16). It resulted in a reduction in the length of the predicted polypeptide by the encoded 20 amino acids. Finally, a nonsynonymous mutation was seen at position 539 (T-C), causing amino acid change I175T.
For sample 3 (Ken/08/80767), 15 SGA PCR products were sequenced, and the dominant sequence (Ken/08/80767/i) was seen in 9/15 (60%). The minority sequence Ken/08/80767/vi had a single adenosine deletion at nucleotide 599 together with the 60-nucleotide deletion identical to that observed for sample 2. In addition, sequence Ken/08/80767/vii had a 104-base deletion starting at position 836, which resulted in a frameshift with no subsequent stop codon in the region sequenced. Other changes for this sample are shown in Table 2.
The RSV G gene has previously been shown to be accumulating amino acid changes and associated antigenic changes in its variable regions in isolates collected over time (2, 20, 21). Sequence changes have also been observed after monoclonal antibody selection (4, 12, 13, 19), and these included point mutations and frameshifts due to insertions or deletions. In addition, analysis of molecular clones derived from cultured RSV showed that replication of the G gene, including in vitro, is prone to errors, again small insertions or deletions (1). This study reports changes similar to those described above detected directly within patients but also much more drastic changes such as the large deletions.
The samples examined by SGA in this study were originally selected because of mixed bases observed by population sequencing, so the results reported here may not be applicable to all RSV infections, and further studies will be required to ascertain how far these observations are generally applicable. The possibility of artifacts should be considered. However, a high-fidelity enzyme mix was used in these experiments, and the error rate in SGA analysis of HIV-1 has been found to be low (0.011%) and without major deletions and insertions(11). The possibility of PCR contamination or mixed infections contributing to the detection of the variant genomes with the exact 60-nucleotide deletion in the same position as the original duplication can be excluded because first, all the samples tested during the period in question had the duplication, and second, the rest of the sequences of the molecules matched most closely the “parent” population sequence (Fig. 1).
The results reported here confirm and extend the observations that the RSV G gene is highly prone to errors during replication, including within the patient, and so provides a pool of variants that can be subject to selection during transmission. The detection of minority genomes that showed a 60-nucleotide deletion at the exact position as the original duplication observed in the Buenos Aires strains was a surprising finding. The mechanisms for such a deletion may lie in the stem-loop structure of the viral RNA sequence that is duplicated, as described by Trento et al. (17). Such deletions provide the potential for emergence of viruses with the “normal” protein length (i.e., without the 20-amino-acid insertion), which, as such strains have been largely replaced in recent years, may be able to overcome the community immunity.
Nucleotide sequence accession numbers.
The GenBank accession numbers for the sequences determined in this study are GU811691 to GU811706.
Acknowledgments
We thank the Kilifi RSV project staff for collecting the samples analyzed here, and we also thank the study participants.
This study is published with permission of the Director of KEMRI.
This work was supported by a Wellcome Trust Masters Training Fellowship to C.N.A. (grant no. 085882) and program grant no. 084663.
Footnotes
Published ahead of print on 21 July 2010.
REFERENCES
- 1.Cane, P. A., D. A. Matthews, and C. R. Pringle. 1993. Frequent polymerase errors observed in a restricted area of clones derived from the attachment (G) protein gene of respiratory syncytial virus. J. Virol. 67:1090-1093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cane, P. A., and C. R. Pringle. 1995. Evolution of subgroup A respiratory syncytial virus: evidence for progressive accumulation of amino acid changes in the attachment protein. J. Virol. 69:2918-2925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Falsey, A. R., P. A. Hennessey, M. A. Formica, C. Cox, and E. E. Walsh. 2005. Respiratory syncytial virus infection in elderly and high-risk adults. N. Engl. J. Med. 352:1749-1759. [DOI] [PubMed] [Google Scholar]
- 4.Garcia-Barreno, B., A. Portela, T. Delgado, J. A. Lopez, and J. A. Melero. 1990. Frame shift mutations as a novel mechanism for the generation of neutralization resistant mutants of human respiratory syncytial virus. EMBO J. 9:4181-4187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Glezen, W. P., L. H. Taber, A. L. Frank, and J. A. Kasel. 1986. Risk of primary infection and reinfection with respiratory syncytial virus. Am. J. Dis. Child. 140:543-546. [DOI] [PubMed] [Google Scholar]
- 6.Henderson, F. W., A. M. Collier, W. A. Clyde, Jr., and F. W. Denny. 1979. Respiratory-syncytial-virus infections, reinfections and immunity. A prospective, longitudinal study in young children. N. Engl. J. Med. 300:530-534. [DOI] [PubMed] [Google Scholar]
- 7.Johnson, P. R., M. K. Spriggs, R. A. Olmsted, and P. L. Collins. 1987. The G glycoprotein of human respiratory syncytial viruses of subgroups A and B: extensive sequence divergence between antigenically related proteins. Proc. Natl. Acad. Sci. U. S. A. 84:5625-5629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mufson, M. A., C. Orvell, B. Rafnar, and E. Norrby. 1985. Two distinct subtypes of human respiratory syncytial virus. J. Gen. Virol. 66:2111-2124. [DOI] [PubMed] [Google Scholar]
- 9.Nair, H., D. J. Nokes, B. D. Gessner, M. Dherani, S. A. Madhi, R. J. Singleton, K. L. O'Brien, A. Roca, P. F. Wright, N. Bruce, A. Chandran, E. Theodoratou, A. Sutanto, E. R. Sedyaningsih, M. Ngama, P. K. Munywoki, C. Kartasasmita, E. A. Simoes, I. Rudan, M. W. Weber, and H. Campbell. 2010. Global burden of acute lower respiratory infections due to respiratory syncytial virus in young children: a systematic review and meta-analysis. Lancet 375:1545-1555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Nokes, D. J., M. Ngama, A. Bett, J. Abwao, P. Munywoki, M. English, J. A. Scott, P. A. Cane, and G. F. Medley. 2009. Incidence and severity of respiratory syncytial virus pneumonia in rural Kenyan children identified through hospital surveillance. Clin. Infect. Dis. 49:1341-1349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Palmer, S., M. Kearney, F. Maldarelli, E. K. Halvas, C. J. Bixby, H. Bazmi, D. Rock, J. Falloon, R. T. Davey, Jr., R. L. Dewar, J. A. Metcalf, S. Hammer, J. W. Mellors, and J. M. Coffin. 2005. Multiple, linked human immunodeficiency virus type 1 drug resistance mutations in treatment-experienced patients are missed by standard genotype analysis. J. Clin. Microbiol. 43:406-413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Rueda, P., T. Delgado, A. Portela, J. A. Melero, and B. Garcia-Barreno. 1991. Premature stop codons in the G glycoprotein of human respiratory syncytial viruses resistant to neutralization by monoclonal antibodies. J. Virol. 65:3374-3378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rueda, P., B. Garcia-Barreno, and J. A. Melero. 1994. Loss of conserved cysteine residues in the attachment (G) glycoprotein of two human respiratory syncytial virus escape mutants that contain multiple A-G substitutions (hypermutations). Virology 198:653-662. [DOI] [PubMed] [Google Scholar]
- 14.Salazar-Gonzalez, J. F., E. Bailes, K. T. Pham, M. G. Salazar, M. B. Guffey, B. F. Keele, C. A. Derdeyn, P. Farmer, E. Hunter, S. Allen, O. Manigart, J. Mulenga, J. A. Anderson, R. Swanstrom, B. F. Haynes, G. S. Athreya, B. T. Korber, P. M. Sharp, G. M. Shaw, and B. H. Hahn. 2008. Deciphering human immunodeficiency virus type 1 transmission and early envelope diversification by single-genome amplification and sequencing. J. Virol. 82:3952-3970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Scott, P. D., R. Ochola, M. Ngama, E. A. Okiro, D. J. Nokes, G. F. Medley, and P. A. Cane. 2004. Molecular epidemiology of respiratory syncytial virus in Kilifi district, Kenya. J. Med. Virol. 74:344-354. [DOI] [PubMed] [Google Scholar]
- 16.Trento, A., I. Casas, A. Calderon, M. L. Garcia-Garcia, C. Calvo, P. Perez-Brena, and J. A. Melero. 2010. Ten years of global evolution of the human respiratory virus BA genotype with a 60-nucleotide duplication in the G protein gene. J. Virol. 84:7500-7512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Trento, A., M. Galiano, C. Videla, G. Carballal, B. Garcia-Barreno, J. A. Melero, and C. Palomo. 2003. Major changes in the G protein of human respiratory syncytial virus isolates introduced by a duplication of 60 nucleotides. J. Gen. Virol. 84:3115-3120. [DOI] [PubMed] [Google Scholar]
- 18.Trento, A., M. Viegas, M. Galiano, C. Videla, G. Carballal, A. S. Mistchenko, and J. A. Melero. 2006. Natural history of human respiratory syncytial virus inferred from phylogenetic analysis of the attachment (G) glycoprotein with a 60-nucleotide duplication. J. Virol. 80:975-984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Walsh, E. E., A. R. Falsey, and W. M. Sullender. 1998. Monoclonal antibody neutralization escape mutants of respiratory syncytial virus with unique alterations in the attachment (G) protein. J. Gen. Virol. 79:479-787. [DOI] [PubMed] [Google Scholar]
- 20.Woelk, C. H., and E. C. Holmes. 2001. Variable immune-driven natural selection in the attachment (G) glycoprotein of respiratory syncytial virus (RSV). J. Mol. Evol. 52:182-192. [DOI] [PubMed] [Google Scholar]
- 21.Zlateva, K. T., P. Lemey, E. Moes, A. M. Vandamme, and M. Van Ranst. 2005. Genetic variability and molecular evolution of the human respiratory syncytial virus subgroup B attachment G protein. J. Virol. 79:9157-9167. [DOI] [PMC free article] [PubMed] [Google Scholar]