Abstract
ERAL1, a homologue of Era protein in Escherichia coli, is a member of conserved GTP-binding proteins with RNA-binding activity. Depletion of prokaryotic Era inhibits cell division without affecting chromosome segregation. Previously, we isolated ERAL1 protein as one of proteins which were associated with mitochondrial transcription factor A by using immunoprecipitation. In this study, we analysed the localization and function of ERAL1 in mammalian cells. ERAL1 was localized in mitochondrial matrix and associated with mitoribosomal proteins including the 12S rRNA. siRNA knockdown of ERAL1 decreased mitochondrial translation, caused redistribution of ribosomal small subunits and reduced 12S rRNA. The knockdown of ERAL1 in human HeLa cells elevated mitochondrial superoxide production and slightly decreased mitochondrial membrane potential. The knockdown inhibited the growth of HeLa cells with an accumulation of apoptotic cells. These results suggest that ERAL1 is localized in a small subunit of the mitochondrial ribosome, plays an important role in the small ribosomal constitution, and is also involved in cell viability.
INTRODUCTION
The circular 16.5 kb human mitochondrial DNA (mtDNA) molecule encodes 2 rRNAs, 22 tRNAs and 13 proteins of the mitochondrial oxidative phosphorylation system responsible for the production of cellular ATP. All the 13 proteins are considered to be essential subunits of a mitochondrial respiratory chain. The 2 rRNAs and 22 tRNAs, used for constructing mitochondrial translational machineries, are also essential for synthesis of the proteins encoded by mtDNA (1,2). Given that majority of ATP production depends on the respiratory chain, maintenance of the mitochondrial genome is critical for individuals to maintain normal health. The mitochondrial respiratory chain consists of five complexes, composed of more than 80 structural proteins. The 13 mtDNA-encoded polypeptides are integral parts of four mitochondrial respiratory chain complexes (I, III, IV and V). Complex II is an only respiratory chain complex that lacks mtDNA-encoded subunits (3).
Synthesis of these 13 proteins is carried out on a specialized translational apparatus located in the mitochondrial matrix. Components of the mitochondrial translational machinery (translational factors, elongation factor and mitoribosomes) are considered to generally resemble the bacterial translational machinery and are distinct from those in cytosol (4,5). Mitoribosome in mammalian is a 55S particle composed of a small 28S and a large 39S subunit and contains a much higher number of proteins and RNAs than those of the bacterial 70S ribosome (6). All of ∼80 mitochondrial ribosomal proteins are the products of nuclear genes and are imported into the mitochondrial matrix (7). To assemble the mitochondrial ribosome, the rRNA synthesis by the mitochondrial transcription machinery needs to coordinate with nuclear expression of the mitoribosomal proteins (6). Several mutations are reported in the nuclear-encoded translation machinery factors such as elongation factor EFG1 and mitochondrial small ribosomal protein (MRPS16 and MRPS22) of patients with autosomal recessive mitochondrial translation defects (8–11).
Mammalian cells contain up to thousands of copies of mtDNA. mtDNAs are organized in nucleoids (12,13). Mitochondrial transcription factor A (TFAM), which was cloned as a transcription factor for mtDNA (14), has known to be essential for the maintenance of mtDNA. mtDNA molecules are packaged into nucleoids. TFAM may be a main factor for the packaging (15). The nucleoids may dynamically change their structure and distribution within mitochondria undergoing fission and fusion and are involved in a variety of dynamic processes including mitochondrial replication, transcription and translation (16). Previously, we isolated ERAL1 as a protein associated with TFAM (17).
In Escherichia coli, Era (E. coli Ras-like protein), is an essential GTP-binding protein and exchanges guanine nucleotide rapidly within seconds (18,19). Era leads to accumulation of an unprocessed precursor of the 16S rRNA. It is shown by electron microscopy that Era locates to the cleft between the head and the platform of the 30S small ribosomal subunit. Furthermore, Era makes contacts with the several assembly elements of the 30S small ribosomal subunit. These observations suggest direct involvement of Era in the functional assembly of the 30S small ribosomal subunit of E. coli (20).
In eukaryote, depletion and mutation of ERA, a chicken homologue of Era, in chicken DT40 cells led to growth impairment, accompanied by an accumulation of apoptotic cells (21,22), suggesting that ERA also plays an essential role in eukaryotic cells. The chicken ERA protein regulates the G1 phase progression via an as yet unknown molecular mechanism, in which an RNA recognition domain of ERA protein was shown to be important. However, there are no investigations on its subcellular localization and function at mitochondria in DT40 cells.
We have previously found that ERAL1, a human orthologue of ERA, is associated with mitochondrial TFAM protein (17). ERAL1 has a putative N-terminal mitochondria targeting sequence of 59 amino acids. Here, we first show that ERAL1 is localized in the mitochondrial matrix. Then, we sought to figure out the roles of human ERAL1 in mitochondrial functions. We have examined what mitochondrial components are associated with ERAL1 protein. To know the physiological roles of ERAL1 in mitochondria, we have investigated the mitochondrial functions such as mitochondrial membrane potential and reactive oxygen species (ROSs) production after siRNA-mediated knockdown of ERAL1. Furthermore, we provide evidence that ERAL1 is required for proper assembly of mitochondrial ribosomal subunits and thus important for mitochondrial translation. Interestingly, the siRNA knockdown of ERAL1 led to cell growth retardation. These results suggest that mitochondrial ribosome-associated ERAL1 plays an important role in mitochondrial function as well as viability of human cells.
METHODS
Antibody
Anti-hemagglutinin (Anti-HA), TFAM, endonuclease G (EndoG), mitochondrial single-stranded binding protein (mtSSB), leucin rich protein 130 (LRP130), elongation factor-Tu (EF-Tu), ERAL1 were raised in our own laboratory. Anti-Calnexin, Histon H1, BAP37 were purchased from StressGen (AnnArber, MI, USA), Millipore (Millerica, MA, USA) and SantaCruze (SantaCruze, CA, USA), respectively. MitoTracker Red and H2DCFDA were purchased from Invitrogen (Carlsbad, CA, USA).
Cell culture
Human cervical cancer HeLa cells were cultured in Dulbecco Eagle’s minimal essential medium (Sigma, St Louis, MO, USA) with 10% heat-inactivated fetal bovine serum (FBS). Cell lines were maintained in a 5% CO2 atmosphere at 37°C. In some experiments, glucose-free Dulbecco's modified eagle medium was supplemented with 0.9 mg/ml galactose, 1 mM sodium pyruvate, 10% FBS and 2 mM l-glutamine.
Immunofluorescent imaging of HeLa cells
Human HeLa cells were incubated in the presence of 100 nM MitoTracker Red for 20 min. After washing with phosphate-buffered saline (PBS) three times, the cells were fixed with 3.7% formaldehyde for 30 min, then incubated with PBS containing 0.1% Triton X-100 for 30 min. After washing with PBS three times, the fixed cells were blocked with PBS containing 1% bovine serum albumin (BSA) for 30 min. Then the cells were incubated with 250-fold diluted anti-HA or -TFAM antibody in PBS/BSA for 1 h. After washing the cells with wash buffer (PBS containing
0.1% Tween-20) three times, the cells were incubated with 250-fold diluted Alexa Fluor 488 goat anti-rabbit IgG (Invitrogen) for 30 min. The glass slides were mounted using Slowfade mounting medium (Invitrogen). Fluorescence images were obtained under a confocal laser microscope (Nikon, Tokyo, Japan).
Immunoblot analysis
HeLa cells were lysed with Lysis buffer (50 mM Tris–HCl, pH 7.5, 1 mM EDTA, 150 mM NaCl and 0.5% NP-40) (TNE) and total lysate (20 µg) were subjected to immunoblot analysis as described previously (23) using polyclonal antibodies or monoclonal antibodies against COX II (Invitrogen), caspase 3, cleaved caspase 3 (Cell signaling, Danvers, MA, USA) and death associated protein 3 (DAP3) (SantaCruz).
Proteins (20 µg) were separated by SDS–polyacrylamide gel electrophoresis (SDS–PAGE) and immunoblotted with indicated specific antibodies. The signals were visualized with horseradish peroxidase-labelled anti-rabbit immunoglobulin G and an ECL reagent (GE healthcare). The chemiluminescence was recorded and quantified with a chilled charge-coupled device camera, LAS1000plus (Fuji Photo Film, Tokyo, Japan).
Knockdown analysis using siRNAs
The following double-stranded ERAL1 RNA 25 bp oligonucleotides were generated from Stealth Select RNAi (Invitrogen): 5′-UAACCAUAGCAACAGAGUGCGUUCC-3′ and 5′-GGAACGCACUCUGUUGCUAUGGUUA-3′. siRNA transfections were performed according to the manufacturer’s instructions (Invitrogen). Briefly, 2 µl Oligofectamine (Invitrogen) was diluted in 100 µl Opti-MEM I medium (Invitrogen) and incubated for 5 min at room temperature. Next, 20 pmol ERAL1 or control duplex Stealth RNA (Invitrogen) in 100 µl Opti-MEM I were added gently and incubated for 20 min at room temperature. Oligomer-Oligofectactamine complexes and aliquots of 2 × 105 HeLa cells in 2 ml culture medium were combined and incubated for 10 min at room temperature. The cells were seeded in six-well dishes with 2 ml culture medium and assayed at indicated time for western blotting and FACS analysis.
Cell proliferation assay
To determine cell proliferation, HeLa cells transfected with control- or ERAL1-siRNA were seeded in 24-well plates at a density of 2 × 104 cells per well. After 24 h, one set of cells was trypsinized, resuspended in PBS and counted by a cell counter (Beckman Coulter, Fullerton, CA, USA) (24). Cells were counted in a similar way every 24 h, up to 96 h.
Expression of ERAL1 in HeLa Tet-on cells
cDNA of ERAL1 containing the deduced first methionine site was amplified from a cDNA library of human HeLa cells by polymerase chain reaction (PCR) using a primer set: 5′-AT GGC TGC CCC CAG CTG GCG and 5′-TCA CTT GAG GAG CTT CAC AGA GAG. Then BamHI and SpeI sites were added to the 5′- and 3′-terminals, respectively, of the cDNA by a second PCR using primers, 5′-ATA TGG ATC CAT GGC TGC CCC CAG CTG GCG and 50-ATATACTAGT CTT GAG GAG CTT CAC AGA GAG. The PCR product was digested with BamHI and SpeI. The DNA fragment encoding ERAL1, a DNA fragment encoding an HA-tag, and a pcDNA5/FRT vector (Invitrogen) were digested with BamHI and NheI and then were ligated. The vector was named pERAL1-HA. We transfected HeLa/FRP/Tet-on cells with the pERAL1-HA vector and selected cells bearing the transgene in the presence of hygromycin B (200 µg/ml) as previously described (25).
Submitochondrial localization
HeLa cells (1 × 108 cells) cultured in 10-cm dishes were scraped off with a cell lifter, suspended in PBS, precipitated by centrifugation and washed with homogenizing buffer (10 mM HEPES-KOH, pH 7.4, 0.25 M sucrose and 1 mM EDTA). The cells were then suspended in four volumes of the same buffer, homogenized with a Potter-Elvehjem homogenizer, and centrifuged at 900g for 10 min. The supernatant was centrifuged at 10 000g for 6 min. The pellet was collected as a crude mitochondrial fraction. The crude mitochondria (20 µg) were suspended in hypotonic buffer (10 mM HEPES-KOH, pH 7.4 and 1 mM EDTA) at 4°C for disruption of the mitochondrial outer membranes. After the hypotonic treatment, the mitochondria were precipitated by centrifugation at 10 000g for 6 min. The mitochondria were resuspended in hypotonic buffer and then digested with proteinase K (200 µg/ml) with or without 1% Triton X-100 on ice for 20 min. The protein was precipitated by the addition of 15% trichloroacetic acid. The precipitates were solubilized in sample buffer (6% SDS, 150 mM Tris base, 10 mM EDTA and 25% glycerol), separated by SDS–PAGE, and analysed by immunoblotting.
Immunoprecipitation using anti-HA antibodies
Each step was done at 4°C or on ice. Doxycyclin-induced HeLa (ERAL1-HA) cells (1 × 108 cells) were homogenized with a Potter-Elvehjem homogenizer in 2 ml of homogenizing buffer (10 mM HEPES-KOH, pH 7.4, 0.25 M sucrose and 1 mM EDTA) and centrifuged at 900g for 10 min. The supernatant (∼2 ml) was diluted with 2 ml of 20% Percoll buffer [0.25 M sucrose, 20 mM Tris-HCl, pH 7.4, 1 mM EDTA and 20% Percoll (GE healthcare)], and the sample was overlaid on a discontinuous Percoll density gradient (4 ml of 40% and 4 ml of 20% Percoll buffer) in a 12-ml centrifugation tube. After centrifugation at 24 000 r.p.m. for 1 h using a SW41Ti rotor (Beckman Coulter), a mitochondrial band located in the middle of the tube was taken. Two to three milligrams of mitochondrial protein were solubilized in 1 ml of IP buffer (10 mM Tris–HCl, pH 7.4, 150 mM NaCl, 1 mM EDTA, 0.5% BSA and 0.5% NP-40) containing 40 µl of beads coated with anti-HA antibodies and control mouse IgG. After 12 h rotation, the beads were washed four times with wash buffer (IP buffer without BSA) and eluted with 1 M glycine (pH 2).
Immunoprecipitation after crosslinking
All procedures were done at 4°C or on ice. Two milligrams of mitochondrial protein were suspended in 1 ml of crosslinking buffer (0.25 M sucrose, 20 mM HEPES–KOH, pH 7.4, 2 mM EDTA and 25 mM NaCl). Then 27 µl of 37% formaldehyde or distilled water (as a control) were added and the mixture was kept on ice for 2 h. After incubation, 125 µl of 1M glycine was added to quench the crosslinking reaction, and the sample was centrifuged at 10 000g for 1 min to precipitate the mitochondria. The mitochondria were solubilized with 100 µl of IP buffer containing 1% SDS. After 10-fold dilution with 1 ml of IP buffer, the immunoprecipitation reaction was done as described above. The immunoprecipitates were heated at 95°C for 30 min to cleave the crosslinks and the proteins were separated by 12% SDS–PAGE. Bands were visualized by staining with Coomassie Brilliant Blue (CBB). The immunoprecipitates were separated by SDS–PAGE and immunoblotted with each antibody.
In-gel protein digestion and LC-MS/MS
A CBB-stained band was excised and the protein was digested with Lysyl Endopeptidase (WAKO, Japan) in digestion buffer (100 mM Tris–HCl, pH 9.0, 0.5% SDS and 1 mM EDTA) for 12 h. The supernatant containing the digested peptides was analysed by liquid chromatography and mass spectrometry (LC/MS/MS) (Agilent 1100 Series an LC/MS system). The peptide masses were assigned to the peptide mass databases using the MASCOT program (MATRIX SCIENCE). All proteins in the SWISS-PROT mammalian database were taken into account. The peptide mass error was limited to 1.0 Da and the MS/MS mass error was limited to 0.8 Da. Only one missed cleavage was accepted.
Analysis of ROS production
The intracellular H2O2 concentration was estimated by means of an oxidation-sensitive fluorescent probe dye, 2′7′-dichlorodihydrofluorescein diacetate (H2DCFDA) (Invitrogen). H2DCFDA was deacetylated intracellularly by non-specific esterase, which was further oxidized by cellular peroxides to the fluorescent compound, 2,7-dichlorofluorescein (DCF). In brief, cells were incubated with the ERAL1- or control siRNA for 72 h. The cells were then washed in PBS and incubated with 20 µM H2DCFDA at 37°C for 30 min according to the instructions of the manufacturer. DCF fluorescence was detected using a fluorescence activated cell sorting (FACS) caliber flow cytometry (Becton Dickinson, Flanklin Lakes, NJ, USA). For each sample, 10 000 events were collected. H2O2 production was expressed as mean fluorescence intensity, which was calculated by FlowJo software.
Pulse-labelling of mitochondrial translation products
In vitro pulse-labelling of mitochondrial translation with [35S]-(methionine and cysteine) (GE Healthcare) for 1 h was performed. In experiments where the label was chased, cells were incubated for 6 min in emetine (100 µg/ml) prior to labelling. The labelled cells were rinsed with isotonic buffer (25 mM Tris–HCl, pH 7.4, 137 mM NaCl, 10 mM KCl, 0.7 mM Na2HPO4). After centrifuge 115g for 5 min, the pellets were suspended in loading buffer containing 93 mM Tris–HCl, pH 6.7, 7.5% glyrerol, 1% SDS, 0.25 mg/ml of bromophenol blue and 3% mercaptoethanol. And the total lysate (50 µg) was loaded and run on 14% SDS–PAGE for 3 h at 180 V. The gels were fixed in a methanol–acetic acid solution, treated with Amplify and measured by BAS2500 (Fuji).
Sucrose gradient analysis of mitochondrial ribosomes
Cell lysates were prepared from HeLa cells treated with ERAL1 or non-targeted (Control) siRNA for 72 h. The HeLa cells were solubilized in a non-ionic detergent (1% lauryl maltoside). Total cell lysates (2 mg) were loaded on a 10–35% sucrose density gradient in 10 mM Tris–HCl, pH 7.2, 150 mM NaCl, 1 mM EDTA and centrifuged for 3 h at 1 00 000g at 4°C by using a swinging bucket SW 40.1 rotor (Beckman Coulter). After separation through 10–35% sucrose gradients, fractions were precipitated by 10% trichloroacetic acid and washed in acetone, and the entire fraction was resolved by SDS–PAGE. The small (Elongation Factor-Tu(EF-Tu) DAP3, MRPS22) and large (MRPL3) mitochondrial ribosomal subunits, ERAL1, cytochrome c oxidase (COX) II, TFAM, Lon and LRP130 were detected by immunoblotting. For real time PCR, RNA was extracted from 50 µl of fractions and was assayed by real time PCR.
To co-immunoprecipitate RNA with ERAL1-HA protein
We used a previously described procedure (24). Briefly, HeLa mitochondria lysates induced by doxycycline were first pre-cleared with 50 µg of rabbit IgG (Bio-Rad) by bath-sonicating for 15 min at 4°C, followed by binding to anti-HA agarose beads and control IgG for 12 h at 4°C. The complexes were pulled down and washed four times with 1 ml of immunoprecipitation buffer. Then, RNA was eluted from the beads with a QIAGEN RNAeasy kit (QIAGEN, Hilden, Germany). Two micrograms of the eluted mitochondrial RNA (mtRNA) were treated by DNase I (100 units) at 37°C for 30 min and then the mtRNA was cleaned with an RNAeasy kit. The RNA pellets were resuspended in water and stored at −70°C.
Quantification of mRNAs
Total RNA was extracted from HeLa cells or a fraction of sucrose density gradient with a QIAGEN RNAeasy kit (QIAGEN). Two micrograms of RNA were treated by DNase I (100 U) at 37°C for 30 min and then RNA was cleaned with an RNAeasy kit. The reverse transcription (RT) of 1 µg of the total RNA was performed with SuperScript II RT (Invitrogen) according to manufacturer’s instructions. Random 6-mer primers were used in the RT reaction. The expressions of mitochondrial genes were detected by quantitative PCR with a thermal cycler (StepOne plus, Invitogen). The PCR primers were listed in Supplementary Table S1.
RT-PCR
RT reactions were carried out using SuperScript II RT and random primers in a 20 : 1 volume following the manufacturer’s protocol. RT reaction mixtures were diluted 1 : 10 with water and the cDNAs were used for PCR. Primer pairs were shown in supplementary Data.
RESULTS
ERAL1 in mitochondria
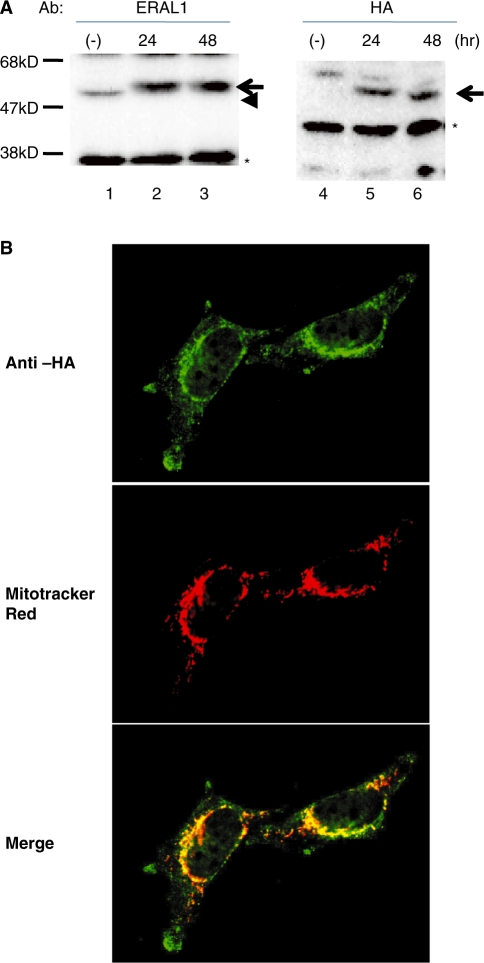
ERAL1 is a protein composed of 368 amino acids as deduced from its cDNA. The existence of a 59-amino acid N-terminal mitochondria targeting sequence was predicted by MitoProt II, a prediction program for mitochondrial localization. The probability of mitochondrial localization is 0.9998 for ERAL1. Then, we produced stable cell lines that express the recombinant full length ERAL1 with a C-terminal HA tag (ERAL1-HA) upon induction by doxycycline addition. The recombinant ERAL1-HA was seen only after the induction by doxycycline-addition at 24 and 48 h (Figure 1A). In immunocytochemistry, the ERAL1-HA was granularly stained with anti-HA antibodies after the induction (Figure 1B). The HA-staining is completely co-localized with the mitochondria visualized with a MitoTracker Red dye, suggesting that the expressed ERAL1 is exclusively localized at mitochondria.
Figure 1.
Inducible expression of recombinant ERAL1 with an HA tag. (A) ERAL1 with an HA tag was inducibly expressed in HeLa cells using a Tet-on system as described under ‘Methods’ section. ERAL1 was detected by immunoblotting with anti-ERAL1 (lanes 1–3) and anti-HA (lanes 4–6) antibodies. Lanes 1/4; no induction, lanes 2/5 and 3/6; 24 and 48 h after tetracycline treatment, respectively. Arrows and an arrowhead indicate recombinant and endogenous ERAL1, respectively. Asterisks show non-specific bands. (B) Immunocytochemistry of recombinant ERAL1. The ERAL1-HA-transfected cells were cultured in the presence of doxycyclin for 48 h. Mitochondria and the recombinant ERAL1 were visualized with a mitochondria-staining dye, MitoTracker Red, (middle panels) and anti-HA antibodies (upper panel), respectively. The lower panels are merged.
Subcellular localization of ERAL1
Then we examined the intracellular localization of ERAL1 by cellular fractionation. The HeLa cells were separated into Nuclear (Nu), Mitochondrial (Mt), microsomal (Micro) and cytosolic (Cyt) fractions. Calnexin, histoneH1, mtSSB and TFAM were detected by immunoblotting as markers for endoplasmic reticulum, nuclei, mitochondria and mitochondria, respectively. The mitochondrial fraction contained essentially no histoneH1, a nuclear protein, indicating minimal contamination of the mitochondrial fraction by nuclei (Supplementary Figure S1). ERAL1-HA was found mostly in the mitochondrial fraction as were TFAM and mtSSB (Supplementary Figure S1, third lane). TFAM was also very weakly detected in the nuclear fraction. Calnexin was detected in the microsomal fraction. These results suggest again that ERAL1 is localized at mitochondria.
Submitochondrial localization of ERAL1
Next we examined the submitochondrial localization of ERAL1-HA. The mitochondrial outer membranes were disrupted by hypotonic treatment (Figure 2, lanes 3 and 4) as verified by the observation that EndoG is localized in the intermembrane space (IMS), was completely digested by proteinase K (Figure2, lane 4). BAP37, which is an inner membrane (IM) protein and largely faces the IMS, was digested by proteinase K. On the other hand, TFAM, which localizes in the matrix, was resistant to the proteinase K digestion (Figure 2, lane 4 in lower panel), indicating that the IMs were intact. Under these hypotonic conditions, ERAL1 was not cleaved by proteinase K (Figure 2, lane 4 in upper panel). When the IMs were solubilized with a non-ionic detergent Triton X-100, both TFAM and ERAL1 were cleaved by proteinase K (Figure 2, lane 5). Taken together, these results suggest that ERAL1 is localized in the mitochondrial matrix side of IM.
Figure 2.
Submitochondrial localization of exogenous ERAL1-HA. Mitochondria were incubated in hypotonic buffer for disruption of the outer membranes without (lanes 3 and 4) or with a non-ionic detergent, Triton X-100 (lane 5). Then the mitochondria were digested with proteinase K (lanes 2, 4 and 5). The indicated proteins were detected by immunoblotting. EndoG, BAP37 and TFAM are markers for the mitochondrial IMS, IM and matrix, respectively.
RNA immunoprecipitation
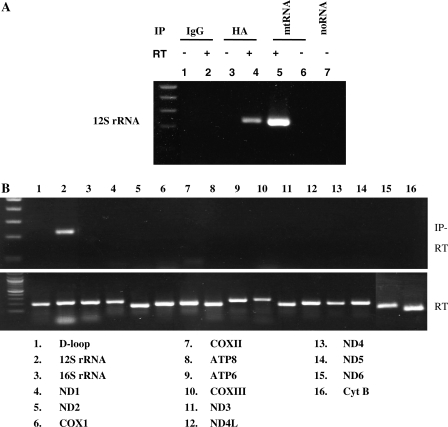
Escherichia coli Era protein has the KH domain which can bind to RNA. We performed RNA immunoprecipitation assay to test whether ERAL1 protein associates with mitochondrial ribosome including the mitochondrial ribosome RNA or mitochondrial transcripts in vivo. Mitochondrial lysates were prepared using immunoprecipitation buffer. After immunoprecipitation with anti-HA antibodies, RNA was extracted from the immunoprecipitates and the eluted mtRNA was used as a template in the RT reaction (Figure 3A). As a control, total mtRNA was used in the RT reaction (Figure 3A, lane 5). First, primer pairs covering 12S rRNA were used in the PCR. A PCR product with a predicted size was obtained for 12S mitochondrial rRNA from the mtRNA (Figure 3A, lane5) and the mtRNA obtained from the HA-immunoprecipitates (Figure 3A, lane 4). There were no PCR products irrespective of the RT reaction when the template was obtained by immunoprecipitation with control IgG (Figure 3A, lanes 1 and 2). There were also no PCR products without RT from the HA-immunoprecipitates (Figure 3A, lane 3), indicating no DNA contamination. Next we investigated whether ERAL1 protein was associated with the other mitochondrial transcripts. We examined 16 PCR primer sets for mitochondrial genes. Only 12S ribosomal RNA was PCR-amplified (Figure 3B). These results suggest that ERAL1 is associated with the mitochondrial ribosomal small subunit which contained 12S mitochondrial ribosomal RNA or directly associated with 12S rRNA. The latter is likely based on its homology to bacterial Era and chicken ERA.
Figure 3.
ERAL1 is associated with mitochondrial 12S rRNA in HeLa cells. (A) ERAL1-HA-transfected HeLa cells were treated by tetracycline for 48 h. The cells were lysed by TNE buffer and immunoprecipitated with control IgG (lanes 1 and 2) or anti-HA antibodies (lanes 3 and 4). RNAs were extracted using an RNAeasy kit from the immunoprecipitates (lanes 1–4) and whole cells (lanes 5 and 6) and then treated with DNaseI. The DNaseI-treated RNAs were subjected to RT (lanes 2, 4 and 5) or not subjected to RT (lanes 1, 3 and 6) and then amplified by PCR with a primer set for 12S rRNA. No RNA template in lane 7. (B) The RNA in lane 4 of (A) (IP-RT, upper panel) and that in lane 5 of (A) (RT, lower panel) were amplified similarly by PCR using indicated primer sets for 16 mitochondrial genes.
Proteins associated with ERAL1
To seek for proteins interacting with ERAL1 in mitochondria, we immunoprecipitated ERAL1-HA-associated proteins with anti-HA antibodies after cross-linking reactions (Figure 4, upper panel). Under these conditions, DAP3 (mitochondrial ribosome protein S29), LRP130 and EF-Tu (mitochondrial translation elongation factor) were detected in the immunoprecipitates (Figure 4), suggesting that ERAL1 is located close to mitoribosome or translation related machinery. We also detected TFAM in the immunoprecipitates.
Figure 4.
Mitochondrial proteins crosslinked to ERAL1-HA. ERAL1-HA-transfected HeLa cells were treated by tetracycline for 48 h. Mitochondrial fraction was purified by percoll density gradient and lysed by TNE buffer. Immunoprecipitates obtained with anti-HA and mouse-IgG were separated by SDS–PAGE and then immunoblotted with anti-EF-Tu, ERAL1, LRP130, TFAM, mtSSB and DAP3 antibodies. An asterisk indicates a heavy chain of IgG.
The other cross-linked proteins which were found by LC-MS/MS analysis are listed in Table 1. The LC-MS/MS analysis revealed three mitoribosomal proteins, presumably indicating the presence of ERAL1 in the small (28S) ribosomal subunits. The second group was proteins which were reported to be associated with human mitochondrial nucleoids. There were factors involved in intra-organellar translation, nucleic acid binding proteins and chaperones/proteases, all of which are expected to associate with ribosomal components or nascent peptides. Several polypeptides known to be involved in metabolism were also identified. In non-cross-linked experiments, we observed that ERAL1 was associated with two tRNA synthetase and Stress70 protein. In summary, many proteins related to chaperons and translation machinery were identified by LC-MS/MS.
Table 1.
List of proteins identified in complex with ERAL1
| Swiss-Prot accession number | Gene symbol | Name | No. of peptides matched | % Sequence coverage | MW | PI | Nucleoid type |
|---|---|---|---|---|---|---|---|
| P38646 | HSPA9 | Stress-70 protein | 8 | 17 | 73 920 | 5.87 | Class I |
| Q15031 | LARS2 | Leucyl-tRNA synthetase | 4 | 4 | 1 02 823 | 8.46 | |
| Q9BW92 | TARS2 | Threonyl-tRNA synthetase | 1 | 1 | 81 841 | 6.9 | |
| P31327 | CPS1 | Carbamoyl-phosphate synthase | 8 | 4 | 1 65 975 | 6.30 | Class I |
| P42704 | LRPPRC | Leucine-rich PPR motif-containing protein | 7 | 5 | 1 59 003 | 5.81 | Class I |
| Q00059 | TFAM | Transcription factor A | 1 | 4 | 29 306 | 9.74 | Class I |
| P34897 | SHMT2 | Serine hydroxymethyltransferase | 3 | 5 | 56 414 | 8.76 | Class I |
| P49411 | TUFM | Elongation factor Tu | 3 | 8 | 49 852 | 7.26 | Class I |
| P10809 | HSPD1 | 60 kDa heat shock protein | 24 | 37 | 61 187 | 5.70 | Class II |
| Q99623 | PHB2 | Prohibitin-2 | 7 | 24 | 33 276 | 9.83 | Class II |
| P40926 | MDH2 | Malate dehydrogenase | 2 | 7 | 35 965 | 8.92 | Class III |
| P54886 | ALDH18A1 | Delta-1-pyrroline-5-carboxylate synthetase | 3 | 3 | 87 989 | 6.66 | |
| Q12931 | TRAP1 | Heat shock protein 75 kDa | 3 | 4 | 80 345 | 8.30 | |
| P31040 | SDHA | Succinate dehydrogenase [ubiquinone] flavoprotein subunit | 1 | 2 | 73 672 | 7.06 | |
| Q5R1W8 | VIM | Vimentin | 6 | 17 | 53 677 | 5.03 | |
| P00367 | GLUD1 | Glutamate dehydrogenase 1 | 6 | 12 | 61 701 | 7.66 | |
| P55084 | HADHB | Trifunctional enzyme subunit γ | 1 | 2 | 51 547 | 9.45 | |
| O75616 | ERAL1 | GTP-binding protein era homologue | 20 | 32 | 48 833 | 9.05 | |
| P60709 | ACTB | Actin, cytoplasmic 1 | 5 | 15 | 42 052 | 5.29 | |
| P82650 | MRPS22 | 28S ribosomal protein S22 | 1 | 3 | 41 425 | 7.70 | |
| P04406 | GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | 3 | 12 | 36 201 | 8.57 | |
| P48047 | ATP5O | ATP synthase subunit O | 4 | 24 | 23 377 | 9.97 | |
| Q9UIK4 | DAPK2 | Death-associated protein kinase 2 | 2 | 4 | 43 042 | 6.45 | |
| Q92665 | MRPS31 | 28S ribosomal protein S31 | 2 | 6 | 45 405 | 9.32 | |
| Q46199 | MTIF2 | Translation initiation factor IF-2 | 3 | 2 | 81 837 | 6.71 | |
| P82909 | MRPS36 | 28S ribosomal protein S36 | 1 | 4 | 11 459 | 9.99 |
The columns correspond to their accession number, gene symbol, name, number of unique peptide matched, percent sequence coverage, estimated MW and PI identified by LC-MS/MS analysis. We observed top three proteins in non-crosslinkng condition. Putative nucleoid proteins as recently classified by Bogenhagen et al. (29). Class I: core nucleoid protein, Class II: proteins seen in native but not cross-linked nucleoids and Class III: proteins seen in cross-linked but not native nucleoids.
siRNA-mediated knockdown of ERAL1 induced mitochondrial dysfunction
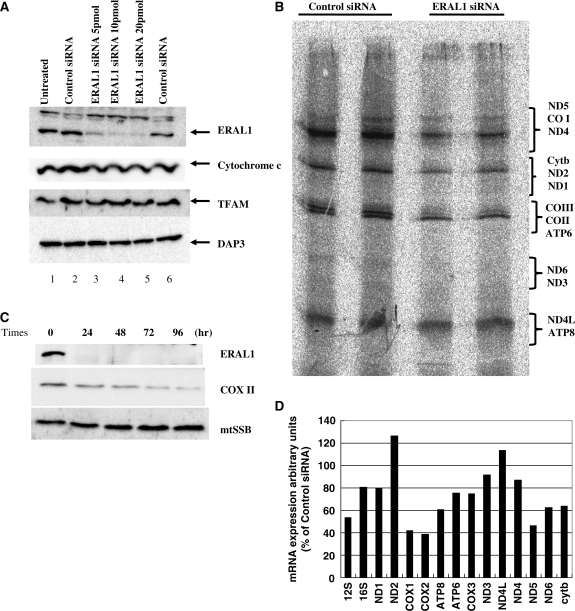
To identify the ERAL1 function in mitochondria matrix, we used an siRNA strategy to downregulate the ERAL1 expression. Because ERAL1 localized in mitochondrial matrix and associated with mitochondrial ribosomes, we hypothesized that suppression of the ERAL1 expression could have an impact on mitochondrial function, especially translation. The expression of ERAL1 was diminished by ERAL1-targeted siRNA treatment for 72 h. However, DAP3, cytochrome c and TFAM were not affected by the ERAL1 knockdown for 72 h (Figure 5A). A non-targeting siRNA (control siRNA) was used to confirm specificity.
Figure 5.
siRNA-mediated knockdown of ERAL1 and in vivo labelling. (A) HeLa cells were transfected with nothing (lane 1), control siRNA (lanes 2 and 6), and 5, 10, 20 pmol of ERAL1 siRNA (lanes 3–5) using oligofectamine. After 2 days of the transfection, the cells were lysed and followed by immunoblotting with anti-ERAL1, cytochrome c, TFAM and DAP3 antibodies. (B) Synthesis of mitochondrial polypeptides. HeLa cells were transfected with control and ERAL1 siRNAs for 72 h. These cells were pulse-labelled in the presence of emetine. The mitochondrial translation products were run on a 15–20% polyacrylamide gradient gel. The gel was processed for fluorography as detailed in the ‘Methods’ section. The seven complex I subunits (ND), one complex III subunit (cyt b), three complex IV subunits (COX) and two complex V subunits (ATP) are indicated on the right. (C) siRNA-mediated knockdown of ERAL1. HeLa cells were transfected with ERAL1 siRNA and harvested every 24 h. The cells were lysed and followed by immunoblotting with anti-ERAL1, COX II and mtSSB antibodies. (D) Quantification of mitochondria-encoded mRNA. HeLa cells were treated with control or ERAL1 siRNA for 72 h. The amounts of mRNAs were measured as described under ‘Methods’ section. The value in control siRNA-treated cells is 100% for each mRNA.
Reduced translation rate in vivo labelling
To investigate the defect in mitochondrial protein synthesis caused by the ERAL1 siRNA transfection, HeLa cells were pulse-labelled with a mixture of [35S] methionine and [35S] cystein in the presence of emetine, an inhibitor of cytoplasmic translation. The overall rate of protein synthesis in mitochondria of the ERAL1 siRNA transfectants was ∼50% of the control siRNA transfectants as shown in Figure 5B. Especially, the protein synthesis rate of COX I, II and III seemed more strongly decreased than the other proteins when ERAL1 was downregulated. These results suggest that ERAL1 protein is involved in mitochondrial translation.
Depletion of ERAL1 decreased COX II protein
To verify the selective effects on the protein synthesis by the ERAL1 knockdown, HeLa cells were transfected with the ERAL1 siRNA and the changes of several proteins were analysed by western blotting. ERAL1 protein was decreased by 24 h after the ERAL1 siRNA transfection. mtSSB was not altered up to 96 h (Figure 5C). Consistent with the stronger decrease of the COX II translation (Figure 5B), a COX II protein level was decreased at 48 h after the siRNA transfection (Figure 5C).
Next, to see if the overall decrease of the mitochondrial translation was affected by the amounts of mRNAs, we investigated effects of the ERAL1 depletion on expression of the mitochondrially encoded RNAs. The ERAL1-directed siRNA significantly lowered the mRNA levels of 12S rRNA, COX I, COX II and ND5 (Figure 5D), but at most moderately affected the 16S rRNA, ND1, ND2, ND3, ND4, cyt b and ND4L transcripts. This relatively stronger decrease of COX II mRNA might contribute in part to the apparent stronger decrease of the COX II translation. The decrease of 12S rRNA might be caused by disassembly of the mitoribosomes as shown later (Figure 6A). It is currently unknown how the other mRNAs was decreased by the ERAL1 depletion.
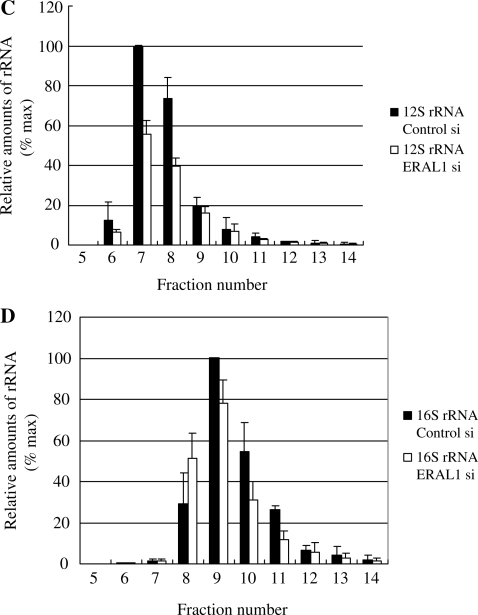
Figure 6.
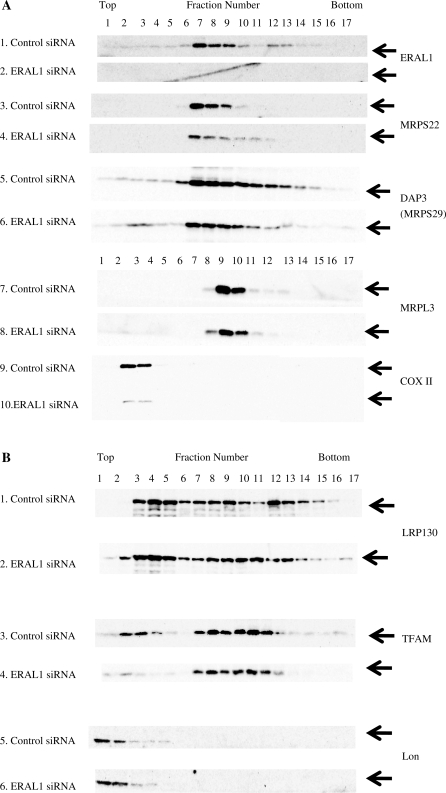
Reduction of ERAL1 results in redistribution of ribosomal protein and ribosomal RNA. (A and B) Cell lysates were prepared from HeLa cells treated with ERAL1 or non-targeted (control) siRNA for 72 h. After separation by centrifugation through 10–35% sucrose, the sample was divided into 17 fractions. Each fraction was analysed by western blotting using antibodies against the small (DAP3, MRPS22) and large (MRPL3) mitochondrial ribosomal subunits, ERAL1, COX II, TFAM, Lon and LRP130. (C and D) Ribosomal RNA levels of each fraction were analyzed by using real time PCR: 12S mitochondrial ribosomal RNA (C), 16S mitochondrial ribosomal RNA (D). The maximal value is 100% for each RNA.
Depletion of ERAL1 affects mitochondrial small subunit
The ERAL1 siRNA inhibited the mitochondrial protein synthesis. To further clarify a role of ERAL1 in the mitochondrial translation, we analysed sedimentation profiles in a sucrose density gradient (Figure 6A). The 3-days treatment with the ERAL1 siRNA significantly decreased ERAL1 in a HeLa cell (Figure 6A, panels 1 and 2) as well as COX II (Figure 6A, panels 9 and 10) as expected. Either the amounts or distribution profiles of the other proteins were not changed by the ERAL1 depletion (Figure 6B). Therefore, the same number of fraction may be comparable before and after the ERAL1 depletion.
ERAL1 largely co-migrated with the small ribosomal subunit proteins (MRPS22 and DAP3) (Figure 6A, panels 1, 3 and 5), suggesting that ERAL1 associates with the small subunit of mitochondrial ribosome (Figure 6A). The treatment caused a striking change in the distribution of MRPS22, proteins of the small ribosomal subunits. In spite of the decrease of overall amount, MRPS22 became evident also in fractions 11 and 12 (Figure 6A, compare panels 3 and 4). Thus, the ERAL1 siRNA treatment shifted the distribution of MRPS22 to heavier fractions. These results suggested that ERAL1 is required for proper assembly of mitochondrial ribosomal small subunits. Conversely, MRPL3 protein were distributed within fractions 9–11 (Figure 6A, panels 7 and 8) and the distribution of MRPL3 were barely changed by the ERAL1 depletion. Taken together, we consider that the ERAL1 associates with small ribosomal subunit and ERAL1 depletion compromises assembly of the small ribosomal subunit.
Next, we investigated the sedimentation profile of 12S rRNA (small subunit) and 16S rRNA (large subunit) in the sucrose density gradient (Figure 6C and D). Quantitative PCR analysis showed that the amount of 12S rRNA at fraction 7–8 was decreased after ERAL1 depletion. However, the abundance of 12S rRNA at fractions 9–10 was not changed even after the ERAL1 depletion (Figure 6C). The relative 12S rRNA distribution is consistent with the results of MRPS22 protein (Figure 6A and C). The distribution and the total amount of 16S rRNA were little changed even after the ERAL1 depletion, suggesting that the ERAL1 depletion did not affect the assembly of the large subunit itself (Figure 6D). Taken together, we consider that the ERAL1 depletion affected assembly of the small ribosomal subunit.
ERAL1 siRNA-mediated knockdown reduced MMP
The ERAL1 siRNA-mediated inhibition of the mitochondrial translation may influence mitochondrial functions. First, we measured mitochondrial membrane potential (MMP) (Δψm) using JC-1 that is a fluorescent dye sensitive to mitochondrial membrane potential. The Δψm value of ERAL1 siRNA transfectants was lower than those of normal as well as control siRNA transfected cells (Supplementary Figure S3). carbonyl cyanide m-chlorophenylhydrazone (CCCP)-treated cells served as a positive control for depolarization of the mitochondrial membrane.
The lower mitochondrial membrane potential in the ERAL1 siRNA-transfected cells was further confirmed by staining with MitoTracker Red, a fluorescent dye accumulating in mitochondria depending on the mitochondrial membrane potential. HeLa cells were transfected with control and ERAL1 siRNAs for 72 h. The cells pre-treated with MitoTracker Red for 20 min were fixed and then TFAM was stained with anti-TFAM antibodies (Supplementary Figure S4). The fluorescence intensity or pattern of TFAM staining was not changed (upper panel), however the ERAL1 siRNA-mediated knockdown cell were significantly weakly stained with MitoTracker Red (center and lower panels, right), suggesting that the ERAL1 depletion reduced the mitochondrial membrane potential.
Depletion of ERAL1 increases mitochondrial ROS
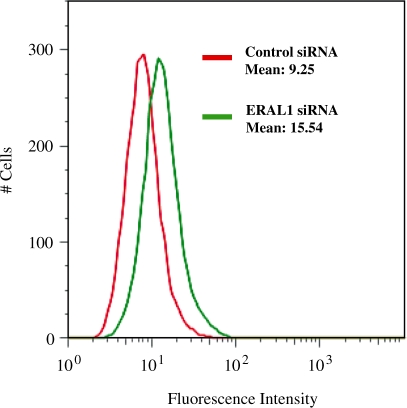
Stress conditions in mitochondria are frequently associated with elevated levels of ROS. A tight link exists between mitochondrial ROS generation and collapse of mitochondrial membrane potential. In subsequent experiments, therefore, the production of oxidants was tested by a redox-responsive fluorescent dye H2DCFDA to examine whether ERAL1 siRNA knockdown led to the generation of ROS. A statistically significant increase in the fluorescence over the control was seen at 72 h after the siRNA transfection (Figure 7).
Figure 7.
Increased ROS production by ERAL1 siRNA-mediated knockdown. HeLa cells were transfected with control (Green) and ERAL1 siRNAs (Red). After 72 h, the cells were treated 10 µM H2DCFDA for 30 min and were subjected to FACS analysis for quantitative estimation of ROS.
Cell growth retardation
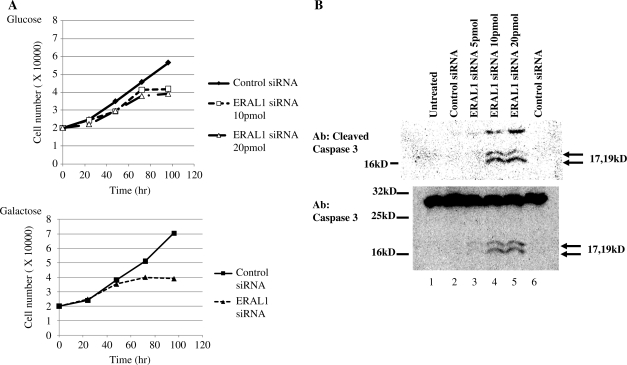
In order to investigate the effect of the elimination of ERAL1 on the cell growth, the number of living cells in each clone was monitored using a cell counter (Figure 8A, upper panel). Up to 2 days after siRNA transfection, there was no significant difference in the growth or morphology of the cells between ERAL1 siRNA and control siRNA transfectants. However, the increase in living cells was markedly suppressed at Day 3 in ERAL1 siRNA-treated cells and this was accompanied by an increment in the population of round and dead cells. These results indicate that ERAL1 is important for the normal cell proliferation and its loss affects mitochondrial functions. We cultured the cells in glucose-free media instead supplemented with galactose to force dependence on oxidative phosphorylation. Even in the galactose culture media, the inhibitory effect of ERAL1 depletion on the cell proliferation was little changed (Figure 8A, lower panel), raising the possibility that the inhibition of the cell proliferation is not much dependent on the mitochondrial dysfunction.
Figure 8.
Depletion of ERAL1 caused a growth defect and increased apoptosis. (A) Proliferation rate of siRNA-transfected HeLa cells. The cells were cultured with a normal glucose medium (upper) or a galactose medium (lower). HeLa cells were transfected with control and ERAL1 siRNAs at Day 0. At indicated time, the cells were harvested and counted by a cell counter. (B) HeLa cells were transfected with control and ERAL siRNAs for 72 h. Untransfected (lane1) and transfected HeLa cells were lysed and immunoblotted by anti-cleaved caspase 3 (upper panel) and anti-caspase 3 (lower panel) antibodies, respectively.
Induction of apoptosis in ERAL1 knockdown HeLa cells
The increase in the apparently apoptotic cells as described earlier prompted us to confirm apoptotic cell death by examining caspase 3 activation. After the treatment of ERAL1 siRNA for 72 h, the cells were lysed and subjected to western blotting. Cleaved caspase 3 or activation of caspase 3 was observed in the ERAL1 knockdown cells (Figure 8B).
DISCUSSION
Here we have shown that ERAL1 is a mitochondrial ribosome-associated factor and is important for mitochondrial function and cell viability based on the following evidence; (i) ERAL1 is a mitochondrial matrix protein, (ii) ERAL1 is associated with small mitoribosomes including the 12S rRNA, (iii) depletion of ERAL1 impairs the assembly of the small subunit of mitoribosome, inhibits the mitochondrial translation, decreases mitochondrial membrane potential and increases ROS generation and (iv) depletion of ERAL1 severely retards cell growth. One of primary roles of ERAL1 may be in the formation of functional ribosomal small subunits and might be involved in mitochondrial biogenesis and associated with the mitochondrial ribosome during the assembly, rRNA folding and RNA modification.
A direct link between mitochondrial transcription and translation has been demonstrated in Saccharomyces cerevisiae. An amino-terminal domain of mitochondrial RNA polymerase (Rpo41p) is a binding site for Nam1p that is proposed to deliver newly synthesized RNAs to the inner mitochondrial membrane and promote subsequent interactions between gene specific translational activators and ribosomes. Thus the processes of transcription and translation are physically and functionally coupled in S. cerevisiae (26,27).
Proteins co-immunoprecipitated with ERAL1 included putative nucleoid proteins in addition to mitochondrial ribosomal proteins and several translation-associated proteins in HeLa cells (Table 1). Nucleoids are dynamic structures containing mtDNA and proteins, which are involved in maintenance, replication and transcription of mtDNA (13). Coupling of transcription and translation in human mitochondria has been suggested (3). There has been reported interaction of mitochondrial RNA polymerase with a mitoribosomal protein MRPL12. This interaction was shown in vitro to stimulate transcription (28). In a recent report on proteins associated with mitochondrial nucleoids, 15 mitoribosomal proteins were identified along with assorted factors that also function in protein synthesis (29). These results raise the possibility that mitochondrial ribosomal proteins and associated factors are coupled with transcription via transcription-related factors such as TFAM also in higher eukaryotes.
Gohda et al. (21) reported that depletion of chicken ERA protein diminished the growth rate of the cell, accompanied by an accumulation of apoptosis cells. The analysis of cell cycle indicates that the elimination of chicken ERA caused arrest at G1 phase but not at M phase, which suggests a distinct role of chicken ERA in the cell cycle progression from that of bacterial Era protein (21,22). They showed that chicken ERA protein was localized in cytosol in DT40 cells and bound to RNA homopolymer (poly-U). Our experiment showed that human ERAL1 was localized at mitochondrial matrix and associated directly or indirectly with 12S mitochondrial rRNA. Depletion of human ERAL1 also diminished the growth rate of HeLa cell probably by apoptosis. In a eukaryotic cell, ERAL1 may be involved in cell viability and apoptosis at least in part through the mitochondrial function.
The human mitochondrial ribosome recycling factor is essential for cell viability (30). Rorbach et al. (30) cloned the putative mitochondrial recycling factor, mtRRF and showed that the protein could associate with mitoribosomes. Depletion of mtRRF was lethal, causing initially mitochondrial dysmorphism, aggregation of mitoribosome, elevated mitochondrial ROS and loss of OXPHOS complexes. The phenotypes caused by ERAL1 depletion resemble to those by mtRRF, suggesting that ERAL1 and mtRRF share the some functions or cooperate in mitochondrial ribosomes.
Mitochondrial nucleoids include not only factors involved in replication and transcription but also structural proteins required for mtDNA maintenance. Recently, Wang and Bogenhagen (31) reported that EF-Tu, LRP130, ATAD3 and DHX30 were TFAM- and mtSSB-associated proteins. EF-Tu is a translation elongation factor with significant homology to its prokaryote counterparts. Previously we have isolated several proteins such as ERAL1, EF-Tu, LRP130 and Hsp60 by co-immunoprecipitaion with TFAM (17). Thus, the nucleoides harbour the proteins related to translation in addition to transcription, replication and mtDNA maintenance.
In E. coli RbfA (ribosome-binding factor A) is essential for cell growth at low temperature (32,33). RbfA is a bacterial cold shock response protein, required for an efficient processing of the 5′ end of the 16S ribosomal RNA during assembly of the small (30S) ribosomal subunit. RbfA binds to the 30S subunit in a position overlapping the binding site of the A and P sites for tRNAs. RbfA has an important role in maturation of the 30S subunit and is involved in translational advantage under conditions of cold shock. It is demonstrated that Era and RbfA have an overlapping function that is essential for the ribosome biogenesis (32). Mammalian ERAL1 is also likely to bind 12S rRNA, is associated with the small subunit of mitochondrial ribosome, and is involved in the ribosomal assembly. Considering the importance of ERAL1 for cell viability, it might be also involved in stress response like RbfA.
Genetic investigation of patients with defective mitochondrial translation led to the discovery of novel mutations in EFG1 in one affected baby and in the mitochondrial EF-Tu in another one (34). Both patients were affected by severe lactic acidosis and rapidly progressive, fatal encephalopathy. The EFG1-mutant patient had early-onset Leigh syndrome, whereas the EF-Tu mutant patient had severe infantile macrocystic leukodystrophy with micropolygyria. ERAL1 is a mitoribosomal protein which is associated with a small subunit of ribosome. In our immunoprecipitation analysis, we showed the interaction of ERAL1 with EF-Tu. ERAL1 might be involved in the translation reaction itself in addition to the assembly of mitoribosomes.
DAP3, which exists in the mitochondrial matrix, is involved in apoptosis (35). DAP3 has been reported to be involved in both γ-interferon and tumour necrosis factor-α-induced apoptosis as well as staurosporine-induced mitochondrial fragmentation (36–38). hNOA1 (human nitric oxide-associated protein 1) interacts with complex I and DAP3 and regulates mitochondrial respiration and apoptosis (39). ERAL1 is also associated with DAP3 and depletion of ERAL1 causes apoptosis. Accordingly, ERAL1 could be involved in apoptosis through the DAP3 interaction.
So far, DAP3 has been the only GTP-binding protein found in the small subunit of the mammalian and yeast mitochondrial ribosomes (40). This protein accounts for the highly specific GTP-binding affinity of small subunits of mitochondrial ribosomes. An immuno-electron microscopic study showed that DAP3 is localized to the base of the lower lobe of the small subunit on the solvent side of the ribosome (41). ERAL1 is associated with the small ribosomal subunit and has a putative GTPase domain. There is the possibility that ERAL1 and DAP3 coordinately regulate the mitochondrial translation and apoptosis.
DAP3 is located at the lower bottom of the 28S subunit and it is not likely that it would interact with a protein that would bind to the platform of the small subunit. In our experiment by using the sucrose density gradient centrifugation, ERAL1 and DAP3 distributed broadly also in lighter fractions through fractions 1–6. However the distribution of other small subunit protein such as MRPS22 and large subunit protein MRPL3 were located through fractions 9–11. The RNA distribution analysis also showed that the ribosomes were located to fractions 7–9, raising the possibility that some DAP3 protein do not associate with the small ribosomal subunits and conducts another function for example as a death associated protein. We also observed that ERAL1 was associated with DAPK2 (death-associated protein kinase 2) (Table 1). We speculate that putative free DAP3 binds to free ERAL1 in mitochondria and the two proteins work cooperatively for apoptosis and/or other functions irrespective of the mitochondrial respiratory chain function. In this respect, it is noteworthy that the replacement of glucose by galactose in the culture medium did not augment the growth retardation induced by the ERAL1 siRNA (Figure 8A)
Several mitochondrial ribosome-related genes are known to be associated with mitochondrial disease and diseases characterized by reduced energy metabolism. For example, Weraarpachai et al. (42) have reported that TACO1 protein is a translational activator of COX I. Mutation of the TACO1 gene leads to cytochrome c oxidase deficiency and resultantly to late onset Leigh syndrome. The TACO1 protein is associated with EF-Ts and involved specifically in the COX I synthesis.
LRPPRC (LRP130), which is a yeast translational activator of Pet309 (43), was coimmunoprecipitated with ERAL1 (Table 1). Pet309 is a mitochondrial protein needed for the proper splicing and translational initiation of mtDNA-encoded COX mRNA (43). These proteins contain PPR motifs that consist of degenerate sequences of 35 amino acids and form antiparallel α helices. PPR motif proteins are thought to be involved in post-translational mRNA metabolism, especially in mitochondria and chloroplasts (44,45). Mutations in LRPPRC are found in French–Canadian individuals with Leigh syndrome harbouring an isolated COX deficiency (46). LRPPRC is reported to be involved in the stabilization of mRNAs for both COX I and COX III in mammals, without directly affecting their translation, suggesting that this protein is involved in processing of the primary RNA unit or in stabilizing mature transcripts with a very limited sequence selectivity (47). Mitochondrial ribosomal proteins and translation related proteins should be considered as important candidates for mitochondrial disease, especially when mtDNA mutations have been ruled out (47). Mutation of ERAL1 also could be a cause of some types of mitochondrial disease.
ERAL1 may be involved in diverse functions, such as mitoribosome assembly, mitochondrial translation, mitochondrial transcription, proliferation, apoptosis and so on, as a mitoribosome-associated protein and/or free protein.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
Ministry of Education, Science, Technology, Sports and Culture of Japan, Grants-in-Aid for Scientific Research (19209019 and 21590337). Funding for open access charge: Ministry of Education, Science, Technology, Sports and Culture of Japan.
Conflict of interest statement. None declared.
Supplementary Material
ACKNOWLEDGEMENTS
We wish to thank Ms Kawabe for helping the manuscript preparation. The authors would like to acknowledge the technical expertise of the Support Center for Education and Research, Kyushu University.
REFERENCES
- 1.Kang D, Hamasaki N. Mitochondrial transcription factor A in the maintenance of mitochondrial DNA: overview of its multiple roles. Ann. NY Acad. Sci. 2005;1042:101–108. doi: 10.1196/annals.1338.010. [DOI] [PubMed] [Google Scholar]
- 2.Asin-Cayuela J, Gustafsson CM. Mitochondrial transcription and its regulation in mammalian cells. Trends Biochem. Sci. 2007;32:111–117. doi: 10.1016/j.tibs.2007.01.003. [DOI] [PubMed] [Google Scholar]
- 3.Falkenberg M, Larsson NG, Gustafsson CM. DNA replication and transcription in mammalian mitochondria. Annu. Rev. Biochem. 2007;76:679–699. doi: 10.1146/annurev.biochem.76.060305.152028. [DOI] [PubMed] [Google Scholar]
- 4.Pel HJ, Grivell LA. Protein synthesis in mitochondria. Mol. Biol. Rep. 1994;19:183–194. doi: 10.1007/BF00986960. [DOI] [PubMed] [Google Scholar]
- 5.Rorbach J, Soleimanpour-Lichaei R, Lightowlers RN, Chrzanowska-Lightowlers ZM. How do mammalian mitochondria synthesize proteins? Biochem. Soc. Trans. 2007;35:1290–1291. doi: 10.1042/BST0351290. [DOI] [PubMed] [Google Scholar]
- 6.O'Brien TW. Evolution of a protein-rich mitochondrial ribosome: implications for human genetic disease. Gene. 2002;286:73–79. doi: 10.1016/s0378-1119(01)00808-3. [DOI] [PubMed] [Google Scholar]
- 7.Kenmochi N, Suzuki T, Uechi T, Magoori M, Kuniba M, Higa S, Watanabe K, Tanaka T. The human mitochondrial ribosomal protein genes: mapping of 54 genes to the chromosomes and implications for human disorders. Genomics. 2001;77:65–70. doi: 10.1006/geno.2001.6622. [DOI] [PubMed] [Google Scholar]
- 8.Coenen MJ, Antonicka H, Ugalde C, Sasarman F, Rossi R, Heister JG, Newbold RF, Trijbels FJ, van den Heuvel LP, Shoubridge EA, et al. Mutant mitochondrial elongation factor G1 and combined oxidative phosphorylation deficiency. N. Engl. J. Med. 2004;351:2080–2086. doi: 10.1056/NEJMoa041878. [DOI] [PubMed] [Google Scholar]
- 9.Antonicka H, Sasarman F, Kennaway NG, Shoubridge EA. The molecular basis for tissue specificity of the oxidative phosphorylation deficiencies in patients with mutations in the mitochondrial translation factor EFG1. Hum. Mol. Genet. 2006;15:1835–1846. doi: 10.1093/hmg/ddl106. [DOI] [PubMed] [Google Scholar]
- 10.Miller C, Saada A, Shaul N, Shabtai N, Ben-Shalom E, Shaag A, Hershkovitz E, Elpeleg O. Defective mitochondrial translation caused by a ribosomal protein (MRPS16) mutation. Ann. Neurol. 2004;56:734–738. doi: 10.1002/ana.20282. [DOI] [PubMed] [Google Scholar]
- 11.Emdadul Haque M, Grasso D, Miller C, Spremulli LL, Saada A. The effect of mutated mitochondrial ribosomal proteins S16 and S22 on the assembly of the small and large ribosomal subunits in human mitochondria. Mitochondrion. 2008;8:254–261. doi: 10.1016/j.mito.2008.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen XJ, Butow RA. The organization and inheritance of the mitochondrial genome. Nat. Rev. Genet. 2005;6:815–825. doi: 10.1038/nrg1708. [DOI] [PubMed] [Google Scholar]
- 13.Kucej M, Butow RA. Evolutionary tinkering with mitochondrial nucleoids. Trends Cell Biol. 2007;17:586–592. doi: 10.1016/j.tcb.2007.08.007. [DOI] [PubMed] [Google Scholar]
- 14.Parisi MA, Clayton DA. Similarity of human mitochondrial transcription factor 1 to high mobility group proteins. Science. 1991;252:965–969. doi: 10.1126/science.2035027. [DOI] [PubMed] [Google Scholar]
- 15.Ohno T, Umeda S, Hamasaki N, Kang D. Binding of human mitochondrial transcription factor A, an HMG box protein, to a four-way DNA junction. Biochem. Biophys. Res. Commun. 2000;271:492–498. doi: 10.1006/bbrc.2000.2656. [DOI] [PubMed] [Google Scholar]
- 16.Shadel GS. Expression and maintenance of mitochondrial DNA: new insights into human disease pathology. Am. J. Pathol. 2008;172:1445–1456. doi: 10.2353/ajpath.2008.071163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kanki T, Ohgaki K, Gaspari M, Gustafsson CM, Fukuoh A, Sasaki N, Hamasaki N, Kang D. Architectural role of mitochondrial transcription factor A in maintenance of human mitochondrial DNA. Mol. Cell. Biol. 2004;24:9823–9834. doi: 10.1128/MCB.24.22.9823-9834.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sullivan SM, Mishra R, Neubig RR, Maddock JR. Analysis of guanine nucleotide binding and exchange kinetics of the Escherichia coli GTPase Era. J. Bacteriol. 2000;182:3460–3466. doi: 10.1128/jb.182.12.3460-3466.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Meier TI, Peery RB, McAllister KA, Zhao G. Era GTPase of Escherichia coli: binding to 16S rRNA and modulation of GTPase activity by RNA and carbohydrates. Microbiology. 2000;146:1071–1083. doi: 10.1099/00221287-146-5-1071. [DOI] [PubMed] [Google Scholar]
- 20.Sharma MR, Barat C, Wilson DN, Booth TM, Kawazoe M, Hori-Takemoto C, Shirouzu M, Yokoyama S, Fucini P, Agrawal RK. Interaction of Era with the 30S ribosomal subunit implications for 30S subunit assembly. Mol Cell. 2005;18:319–329. doi: 10.1016/j.molcel.2005.03.028. [DOI] [PubMed] [Google Scholar]
- 21.Gohda J, Nomura Y, Suzuki H, Arai H, Akiyama T, Inoue J. Elimination of the vertebrate Escherichia coli Ras-like protein homologue leads to cell cycle arrest at G1 phase and apoptosis. Oncogene. 2003;22:1340–1348. doi: 10.1038/sj.onc.1206287. [DOI] [PubMed] [Google Scholar]
- 22.Akiyama T, Gohda J, Shibata S, Nomura Y, Azuma S, Ohmori Y, Sugano S, Arai H, Yamamoto T, Inoue J. Mammalian homologue of E. coli Ras-like GTPase (ERA) is a possible apoptosis regulator with RNA binding activity. Genes Cells. 2001;6:987–1001. doi: 10.1046/j.1365-2443.2001.00480.x. [DOI] [PubMed] [Google Scholar]
- 23.Shibahara K, Uchiumi T, Fukuda T, Kura S, Tominaga Y, Maehara Y, Kohno K, Nakabeppu Y, Tsuzuki T, Kuwano M. Targeted disruption of one allele of the Y-box binding protein-1 (YB-1) gene in mouse embryonic stem cells and increased sensitivity to cisplatin and mitomycin C. Cancer Sci. 2004;95:348–353. doi: 10.1111/j.1349-7006.2004.tb03214.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Uchiumi T, Fotovati A, Sasaguri T, Shibahara K, Shimada T, Fukuda T, Nakamura T, Izumi H, Tsuzuki T, Kuwano M, et al. YB-1 is important for an early stage embryonic development: neural tube formation and cell proliferation. J. Biol. Chem. 2006;281:40440–40449. doi: 10.1074/jbc.M605948200. [DOI] [PubMed] [Google Scholar]
- 25.Ohgaki K, Kanki T, Fukuoh A, Kurisaki H, Aoki Y, Ikeuchi M, Kim SH, Hamasaki N, Kang D. The C-terminal tail of mitochondrial transcription factor a markedly strengthens its general binding to DNA. J. Biochem. 2007;141:201–211. doi: 10.1093/jb/mvm020. [DOI] [PubMed] [Google Scholar]
- 26.Rodeheffer MS, Boone BE, Bryan AC, Shadel GS. Nam1p, a protein involved in RNA processing and translation, is coupled to transcription through an interaction with yeast mitochondrial RNA polymerase. J. Biol. Chem. 2001;276:8616–8622. doi: 10.1074/jbc.M009901200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wang Y, Shadel GS. Stability of the mitochondrial genome requires an amino-terminal domain of yeast mitochondrial RNA polymerase. Proc. Natl Acad. Sci. USA. 1999;96:8046–8051. doi: 10.1073/pnas.96.14.8046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang Z, Cotney J, Shadel GS. Human mitochondrial ribosomal protein MRPL12 interacts directly with mitochondrial RNA polymerase to modulate mitochondrial gene expression. J. Biol. Chem. 2007;282:12610–12618. doi: 10.1074/jbc.M700461200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bogenhagen DF, Rousseau D, Burke S. The layered structure of human mitochondrial DNA nucleoids. J. Biol. Chem. 2008;283:3665–3675. doi: 10.1074/jbc.M708444200. [DOI] [PubMed] [Google Scholar]
- 30.Rorbach J, Richter R, Wessels HJ, Wydro M, Pekalski M, Farhoud M, Kuhl I, Gaisne M, Bonnefoy N, Smeitink JA, et al. The human mitochondrial ribosome recycling factor is essential for cell viability. Nucleic Acids Res. 2008;36:5787–5799. doi: 10.1093/nar/gkn576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang Y, Bogenhagen DF. Human mitochondrial DNA nucleoids are linked to protein folding machinery and metabolic enzymes at the mitochondrial inner membrane. J. Biol. Chem. 2006;281:25791–25802. doi: 10.1074/jbc.M604501200. [DOI] [PubMed] [Google Scholar]
- 32.Inoue K, Chen J, Tan Q, Inouye M. Era and RbfA have overlapping function in ribosome biogenesis in Escherichia coli. J. Mol. Microbiol. Biotechnol. 2006;11:41–52. doi: 10.1159/000092818. [DOI] [PubMed] [Google Scholar]
- 33.Datta PP, Wilson DN, Kawazoe M, Swami NK, Kaminishi T, Sharma MR, Booth TM, Takemoto C, Fucini P, Yokoyama S, et al. Structural aspects of RbfA action during small ribosomal subunit assembly. Mol. Cell. 2007;28:434–445. doi: 10.1016/j.molcel.2007.08.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sasarman F, Antonicka H, Shoubridge EA. The A3243G tRNALeu(UUR) MELAS mutation causes amino acid misincorporation and a combined respiratory chain assembly defect partially suppressed by overexpression of EFTu and EFG2. Hum. Mol. Genet. 2008;17:3697–3707. doi: 10.1093/hmg/ddn265. [DOI] [PubMed] [Google Scholar]
- 35.Berger T, Brigl M, Herrmann JM, Vielhauer V, Luckow B, Schlondorff D, Kretzler M. The apoptosis mediator mDAP-3 is a novel member of a conserved family of mitochondrial proteins. J. Cell Sci. 2000;113:3603–3612. doi: 10.1242/jcs.113.20.3603. [DOI] [PubMed] [Google Scholar]
- 36.Kim HR, Chae HJ, Thomas M, Miyazaki T, Monosov A, Monosov E, Krajewska M, Krajewski S, Reed JC. Mammalian dap3 is an essential gene required for mitochondrial homeostasis in vivo and contributing to the extrinsic pathway for apoptosis. FASEB J. 2007;21:188–196. doi: 10.1096/fj.06-6283com. [DOI] [PubMed] [Google Scholar]
- 37.Kissil JL, Deiss LP, Bayewitch M, Raveh T, Khaspekov G, Kimchi A. Isolation of DAP3, a novel mediator of interferon-gamma-induced cell death. J. Biol. Chem. 1995;270:27932–27936. doi: 10.1074/jbc.270.46.27932. [DOI] [PubMed] [Google Scholar]
- 38.Mukamel Z, Kimchi A. Death-associated protein 3 localizes to the mitochondria and is involved in the process of mitochondrial fragmentation during cell death. J. Biol. Chem. 2004;279:36732–36738. doi: 10.1074/jbc.M400041200. [DOI] [PubMed] [Google Scholar]
- 39.Tang T, Zheng B, Chen SH, Murphy AN, Kudlicka K, Zhou H, Farquhar MG. hNOA1 interacts with complex I and DAP3 and regulates mitochondrial respiration and apoptosis. J. Biol. Chem. 2009;284:5414–5424. doi: 10.1074/jbc.M807797200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cavdar Koc E, Ranasinghe A, Burkhart W, Blackburn K, Koc H, Moseley A, Spremulli LL. A new face on apoptosis: death-associated protein 3 and PDCD9 are mitochondrial ribosomal proteins. FEBS Lett. 2001;492:166–170. doi: 10.1016/s0014-5793(01)02250-5. [DOI] [PubMed] [Google Scholar]
- 41.O'Brien TW, O'Brien BJ, Norman RA. Nuclear MRP genes and mitochondrial disease. Gene. 2005;354:147–151. doi: 10.1016/j.gene.2005.03.026. [DOI] [PubMed] [Google Scholar]
- 42.Weraarpachai W, Antonicka H, Sasarman F, Seeger J, Schrank B, Kolesar JE, Lochmuller H, Chevrette M, Kaufman BA, Horvath R, et al. Mutation in TACO1, encoding a translational activator of COX I, results in cytochrome c oxidase deficiency and late-onset Leigh syndrome. Nat. Genet. 2009;41:833–837. doi: 10.1038/ng.390. [DOI] [PubMed] [Google Scholar]
- 43.Manthey GM, Przybyla-Zawislak BD, McEwen JE. The Saccharomyces cerevisiae Pet309 protein is embedded in the mitochondrial inner membrane. Eur. J. Biochem. 1998;255:156–161. doi: 10.1046/j.1432-1327.1998.2550156.x. [DOI] [PubMed] [Google Scholar]
- 44.Lurin C, Andres C, Aubourg S, Bellaoui M, Bitton F, Bruyere C, Caboche M, Debast C, Gualberto J, Hoffmann B, et al. Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell. 2004;16:2089–2103. doi: 10.1105/tpc.104.022236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Delannoy E, Stanley WA, Bond CS, Small ID. Pentatricopeptide repeat (PPR) proteins as sequence-specificity factors in post-transcriptional processes in organelles. Biochem. Soc. Trans. 2007;35:1643–1647. doi: 10.1042/BST0351643. [DOI] [PubMed] [Google Scholar]
- 46.Mootha VK, Lepage P, Miller K, Bunkenborg J, Reich M, Hjerrild M, Delmonte T, Villeneuve A, Sladek R, Xu F, et al. Identification of a gene causing human cytochrome c oxidase deficiency by integrative genomics. Proc. Natl Acad. Sci. USA. 2003;100:605–610. doi: 10.1073/pnas.242716699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Xu F, Morin C, Mitchell G, Ackerley C, Robinson BH. The role of the LRPPRC (leucine-rich pentatricopeptide repeat cassette) gene in cytochrome oxidase assembly: mutation causes lowered levels of COX (cytochrome c oxidase) I and COX III mRNA. Biochem. J. 2004;382:331–336. doi: 10.1042/BJ20040469. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.